Survey of Ticks and Tick-Borne Rickettsial and Protozoan Pathogens in Eswatini
Abstract
:1. Introduction
2. Results
2.1. Tick Diversity
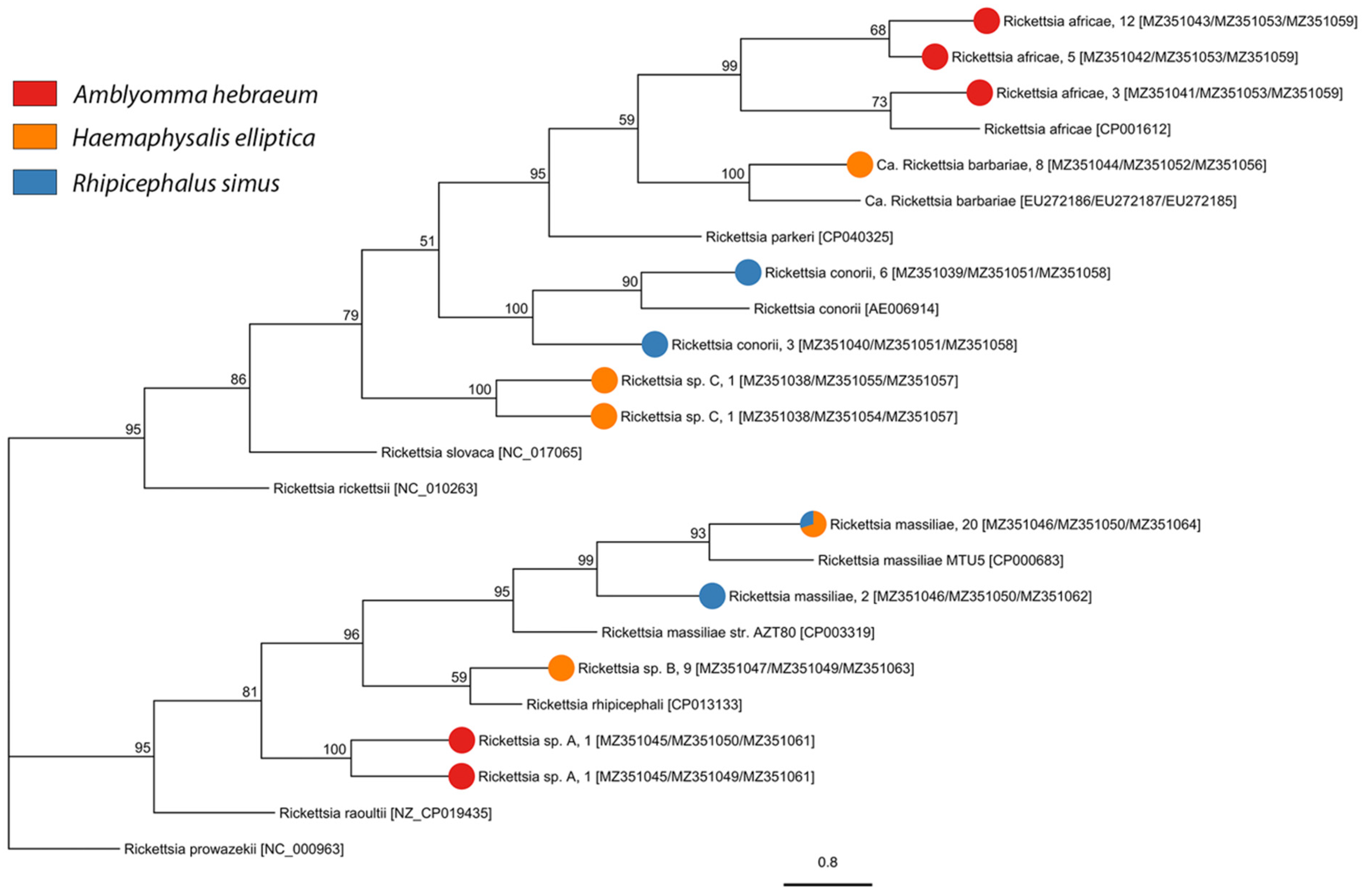
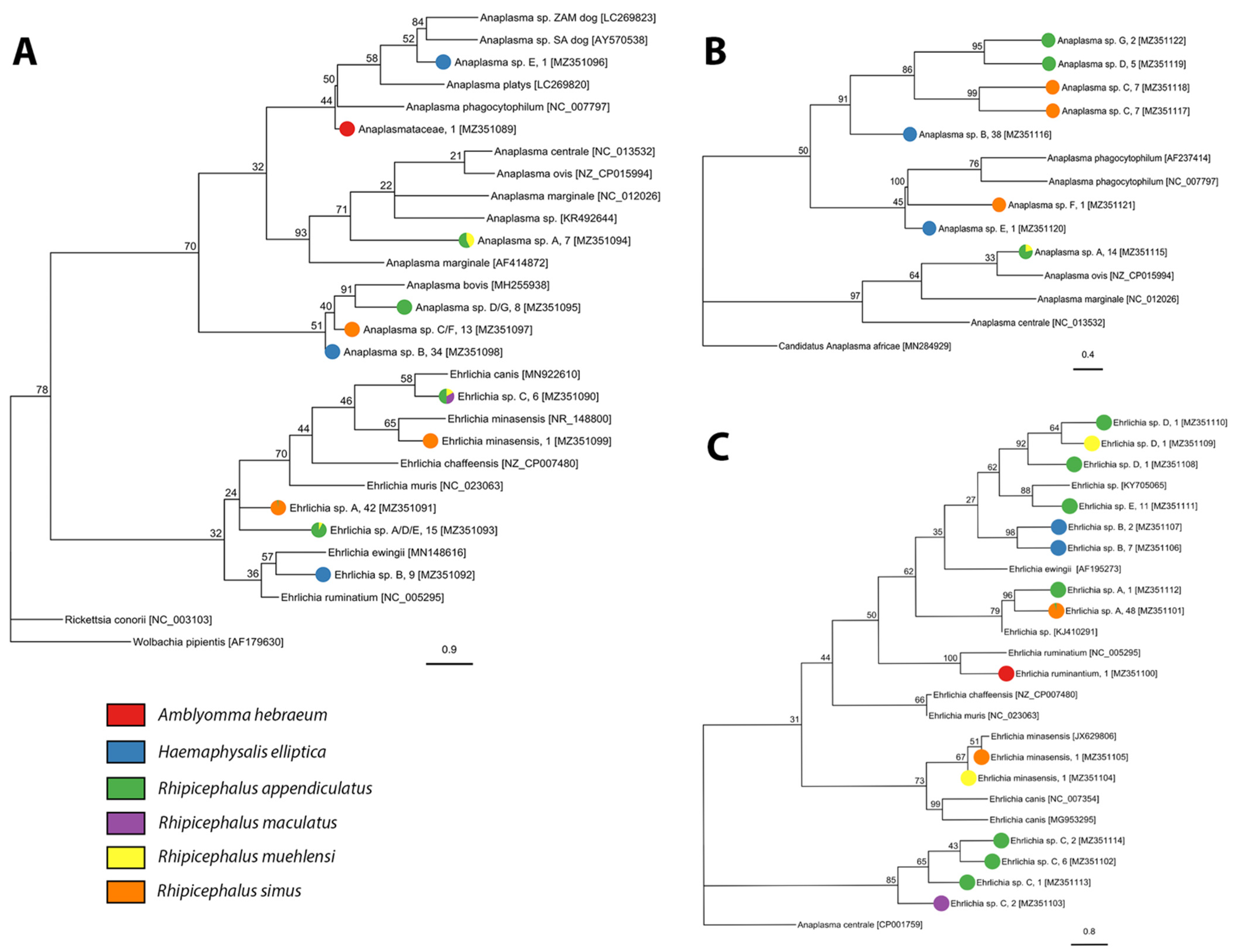
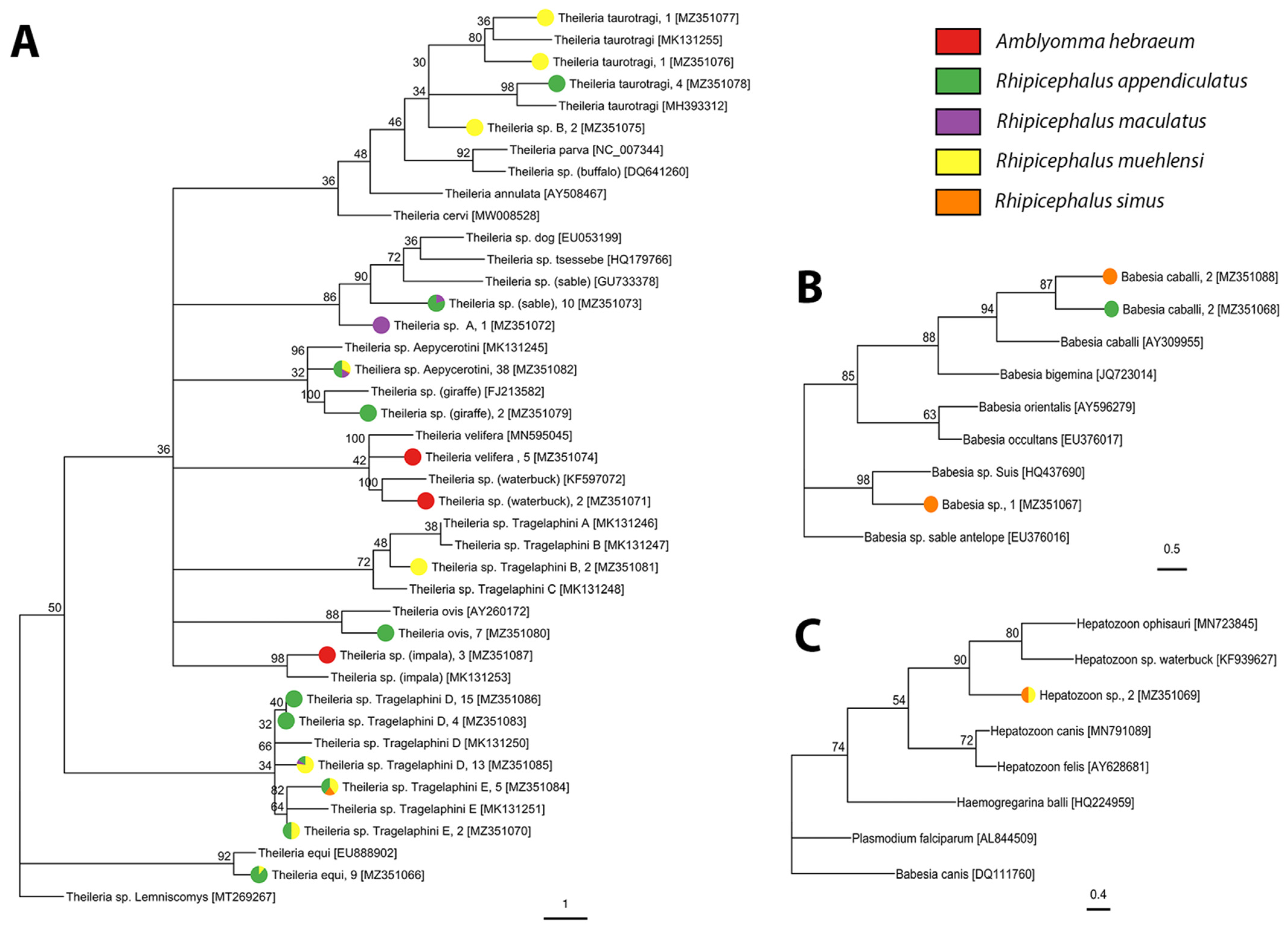
2.2. Tick-Borne Pathogen Diversity
3. Discussion
4. Materials and Methods
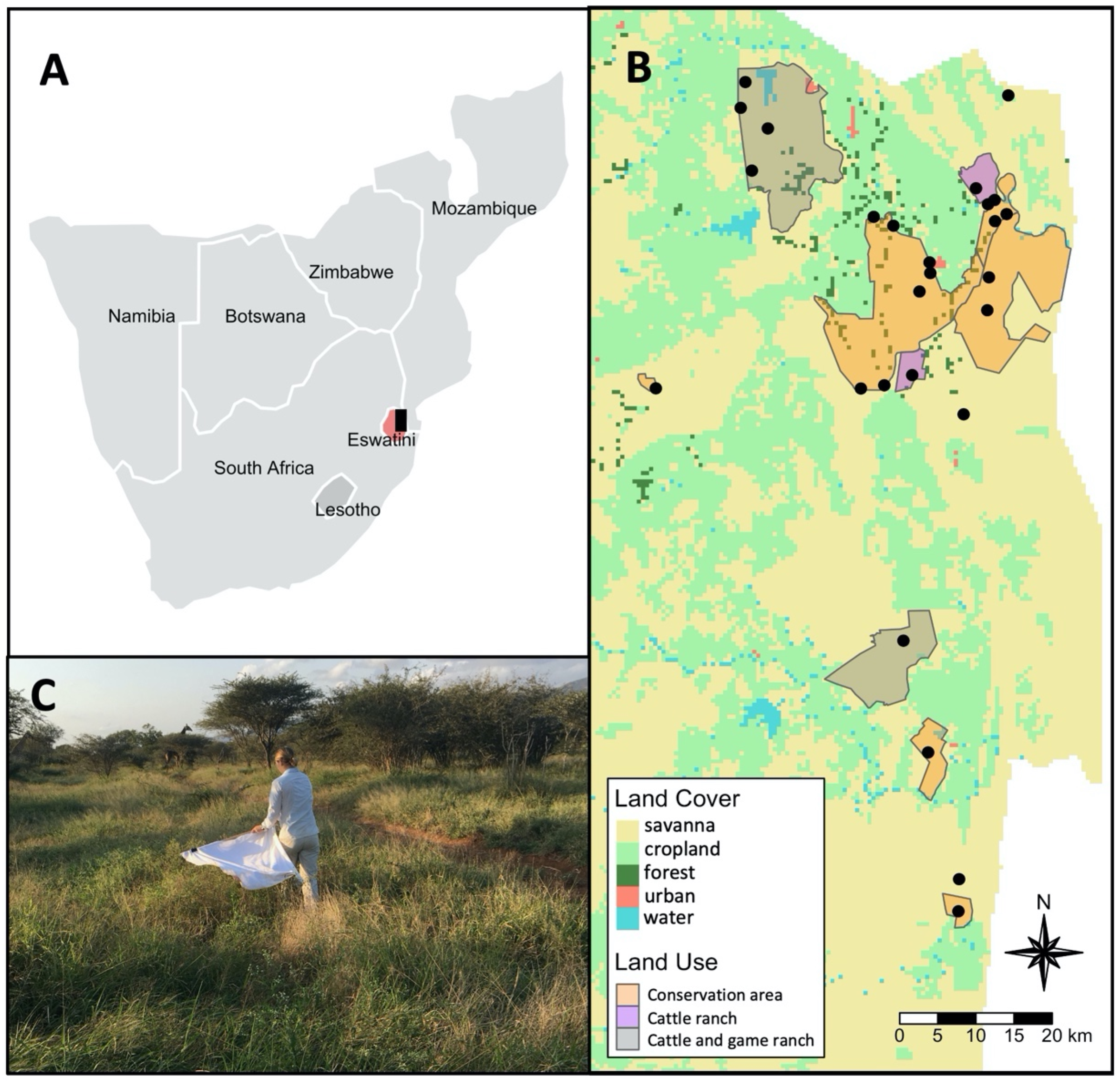
4.1. Study Area and Sampling Design
4.2. DNA Extraction
4.3. Molecular Tick Identification
4.4. Pathogen Detection
4.4.1. Spotted Fever Group Rickettsia
4.4.2. Anaplasmataceae
4.4.3. Apicomplexa
4.5. Phylogenetic Data Analysis
4.6. Statistical Analyses
4.7. Voucher Tick and Pathogen Sequences
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De la Fuente, J. Overview: Ticks as vectors of pathogens that cause disease in humans and animals. Front. Biosci. 2008, 13, 6938–6946. [Google Scholar] [CrossRef] [Green Version]
- Jongejan, F.; Uilenberg, G. The global importance of ticks. Parasitology 2004, 129, S3–S14. [Google Scholar] [CrossRef]
- Ryser-Degiorgis, M.-P. Wildlife health investigations: Needs, challenges and recommendations. BMC Vet. Res. 2013, 9, 223. [Google Scholar] [CrossRef] [Green Version]
- Allan, B.F.; Keesing, F.; Ostfeld, R.S. Effect of forest fragmentation on lyme disease risk. Conserv. Biol. 2003, 17, 267–272. [Google Scholar] [CrossRef] [Green Version]
- Perez, G.; Bastian, S.; Agoulon, A.; Bouju, A.; Durand, A.; Faille, F.; Lebert, I.; Rantier, Y.; Plantard, O.; Butet, A. Effect of landscape features on the relationship between Ixodes ricinus ticks and their small mammal hosts. Parasites Vectors 2016, 9, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, M.R.; Wilson, M.L. Microclimate-dependent survival of unfed adult Ixodes scapularis (acari: Ixodidae) in nature: Life cycle and study design implications. J. Med. Entomol. 1996, 33, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Randolph, S.E.; Storey, K. Impact of Microclimate on immature tick-rodent host interactions (acari: Ixodidae): Implications for parasite transmission. J. Med. Entomol. 1999, 36, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Burri, C.; Bastic, V.; Maeder, G.; Patalas, E.; Gern, L. Microclimate and the zoonotic cycle of tick-borne encephalitis virus in Switzerland. J. Med. Entomol. 2011, 48, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Bailey, K.M.; McCleery, R.; Binford, M.W.; Zweig, C. Land-cover change within and around protected areas in a biodiversity hotspot. J. Land Use Sci. 2015, 11, 154–176. [Google Scholar] [CrossRef]
- Lambin, E.F.; Geist, H.J.; Lepers, E. Dynamics ofland-use and land-coverchange intropicalregions. Annu. Rev. Environ. Resour. 2003, 28, 205–241. [Google Scholar] [CrossRef] [Green Version]
- Cumming, D.H.M.; Osofsky, S.A.; Atkinson, S.J.; Atkinson, M.W. Beyond fences: Wildlife, livestock and land use in Southern Africa. In One Health: The Theory and Practice of Integrated Health Approaches; CABI Publishing: Wallingford, UK, 2015; pp. 243–257. [Google Scholar]
- Ledger, K.J.; Keenan, R.M.; Sayler, K.A.; Wisely, S.M. Multi-scale patterns of tick occupancy and abundance across an agricultural landscape in southern Africa. PLoS ONE 2019, 14, e0222879. [Google Scholar] [CrossRef]
- Nijhof, A.M.; Penzhorn, B.; Lynen, G.; Mollel, J.O.; Morkel, P.; Bekker, C.P.J.; Jongejan, F. Babesia bicornis sp. nov. and Theileria bicornis sp. nov.: Tick-borne parasites associated with mortality in the black rhinoceros (Diceros bicornis). J. Clin. Microbiol. 2003, 41, 2249–2254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nijhof, A.M.; Pillay, V.; Steyl, J.; Prozesky, L.; Stoltsz, W.H.; Lawrence, J.A.; Penzhorn, B.; Jongejan, F. Molecular characterization of theileria species associated with mortality in four species of african antelopes. J. Clin. Microbiol. 2005, 43, 5907–5911. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matjila, P.; Leisewitz, A.; Oosthuizen, M.; Jongejan, F.; Penzhorn, B. Detection of a Theileria species in dogs in South Africa. Veter. Parasitol. 2008, 157, 34–40. [Google Scholar] [CrossRef]
- Oosthuizen, M.C.; Allsopp, B.A.; Troskie, M.; Collins, N.; Penzhorn, B. Identification of novel Babesia and Theileria species in South African giraffe (Giraffa camelopardalis, Linnaeus, 1758) and roan antelope (Hippotragus equinus, Desmarest 1804). Veter. Parasitol. 2009, 163, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Penzhorn, B.L.; Netherlands, E.C.; Cook, C.A.; Smit, N.; Vorster, I.; Harrison-White, R.F.; Oosthuizen, M.C. Occurrence of Hepatozoon canis (Adeleorina: Hepatozoidae) and Anaplasma spp. (Rickettsiales: Anaplasmataceae) in black-backed jackals (Canis mesomelas) in South Africa. Parasit. Vectors 2018, 11, 158. [Google Scholar] [CrossRef] [Green Version]
- Pfitzer, S.; Oosthuizen, M.; Bosman, A.-M.; Vorster, I.; Penzhorn, B. Tick-borne blood parasites in nyala (Tragelaphus angasii, Gray 1849) from KwaZulu-Natal, South Africa. Veter. Parasitol. 2010, 176, 126–131. [Google Scholar] [CrossRef] [Green Version]
- Berggoetz, M.; Schmid, M.; Ston, D.; Wyss, V.; Chevillon, C.; Pretorius, A.-M.; Gern, L. Tick-borne pathogens in the blood of wild and domestic ungulates in South Africa: Interplay of game and livestock. Ticks Tick Borne Dis. 2014, 5, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Adamu, M.; Troskie, M.; Oshadu, O.D.; Malatji, D.P.; Penzhorn, B.L.; Matjila, P.T. Occurrence of tick-transmitted pathogens in dogs in Jos, Plateau State, Nigeria. Parasites Vectors 2014, 7, 119. [Google Scholar] [CrossRef] [Green Version]
- Chaisi, M.; Sibeko-Matjila, K.; Collins, N.; Potgieter, F.T.; Oosthuizen, M. Identification of Theileria parva and Theileria sp. (buffalo) 18S rRNA gene sequence variants in the African Buffalo (Syncerus caffer) in southern Africa. Veter. Parasitol. 2011, 182, 150–162. [Google Scholar] [CrossRef] [Green Version]
- Brothers, P.; Peter, S.; Collins, N.; Oosthuizen, M.; Bhoora, R.; Troskie, M.; Penzhorn, B. Occurrence of blood-borne tick-transmitted parasites in common tsessebe (Damaliscus lunatus) antelope in Northern Cape Province, South Africa. Veter. Parasitol. 2011, 183, 160–165. [Google Scholar] [CrossRef] [Green Version]
- Berggoetz, M.; Schmid, M.; Ston, D.; Wyss, V.; Chevillon, C.; Pretorius, A.-M.; Gern, L. Protozoan and bacterial pathogens in tick salivary glands in wild and domestic animal environments in South Africa. Ticks Tick Borne Dis. 2014, 5, 176–185. [Google Scholar] [CrossRef]
- Maina, A.N.; Jiang, J.; Omulo, S.A.; Cutler, S.J.; Ade, F.; Ogola, E.; Feikin, D.R.; Njenga, M.K.; Cleaveland, S.; Mpoke, S.; et al. High Prevalence of Rickettsia africae Variants in Amblyomma variegatum Ticks from Domestic Mammals in Rural Western Kenya: Implications for Human Health. Vector Borne Zoonotic Dis. 2014, 14, 693–702. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aydin, M.F.; Aktas, M.; Dumanli, N.; Aydın, M.F. Molecular identification of Theileria and Babesia in ticks collected from sheep and goats in the Black Sea region of Turkey. Parasitol. Res. 2014, 114, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Halajian, A.; Palomar, A.M.; Portillo, A.; Heyne, H.; Luus-Powell, W.J.; Oteo, J.A. Investigation of Rickettsia, Coxiella burnetii and Bartonella in ticks from animals in South Africa. Ticks Tick Borne Dis. 2016, 7, 361–366. [Google Scholar] [CrossRef] [PubMed]
- Dahmani, M.; Davoust, B.; Rousseau, F.; Raoult, D.; Fenollar, F.; Mediannikov, O. Natural Anaplasmataceae infection in Rhipicephalus bursa ticks collected from sheep in the French Basque Country. Ticks Tick Borne Dis. 2017, 8, 18–24. [Google Scholar] [CrossRef]
- Mtshali, K.; Nakao, R.; Sugimoto, C.; Thekisoe, O. Occurrence of Coxiella burnetii, Ehrlichia canis, Rickettsia species and Anaplasma phagocytophilum-like bacterium in ticks collected from dogs and cats in South Africa. J. S. Afr. Veter. Assoc. 2017, 88, 1–6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iweriebor, B.C.; Mmbaga, E.J.; Adegborioye, A.; Igwaran, A.; Obi, L.C.; Okoh, A.I. Genetic profiling for Anaplasma and Ehrlichia species in ticks collected in the Eastern Cape Province of South Africa. BMC Microbiol. 2017, 17, 45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halajian, A.; Palomar, A.M.; Portillo, A.; Heyne, H.; Romero, L.; Oteo, J.A. Detection of zoonotic agents and a new Rickettsia strain in ticks from donkeys from South Africa: Implications for travel medicine. Travel Med. Infect. Dis. 2018, 26, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Beati, L.; Meskini, M.; Thiers, B.; Raoult, D. Rickettsia aeschlimannii sp. nov., a new spotted fever group rickettsia associated with hyalomma marginatum ticks. Int. J. Syst. Bacteriol. 1997, 47, 548–554. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pretorius, A.-M.; Birtles, R. Rickettsia aeschlimannii: A new pathogenic spotted fever group rickettsia, South Africa. Emerg. Infect. Dis. 2002, 8, 874. [Google Scholar] [CrossRef] [PubMed]
- Pretorius, A.-M.; Jensenius, M.; Birtles, R. Update on spotted fever group Rickettsiae in South Africa. Vector Borne Zoonotic Dis. 2004, 4, 249–260. [Google Scholar] [CrossRef]
- Pretorius, A.-M.; Birtles, R. Rickettsia mongolotimonaeInfection in South Africa. Emerg. Infect. Dis. 2004, 10, 126–129. [Google Scholar] [CrossRef]
- Parola, P.; Paddock, C.D.; Socolovschi, C.; Labruna, M.B.; Mediannikov, O.; Kernif, T.; Abdad, M.Y.; Stenos, J.; Bitam, I.; Fournier, P.-E.; et al. Update on tick-borne rickettsioses around the world: A geographic approach. Clin. Microbiol. Rev. 2013, 26, 657–702. [Google Scholar] [CrossRef] [Green Version]
- Woolf, D.; Jordaan, M.; Maartens, G. Acute disseminated encephalomyelitis due to Rickettsia conorii infection. S. Afr. Med. J. 2021, 111, 307–308. [Google Scholar] [CrossRef]
- Jensenius, M.; Fournier, P.-E.; Kelly, P.; Myrvang, B.; Raoult, D. African tick bite fever. Lancet Infect. Dis. 2003, 3, 557–564. [Google Scholar] [CrossRef]
- Lee, W.; Seong, H.; Kim, J.H.; Choi, H.; Kim, J.H.; Ahn, J.Y.; Jeong, S.J.; Ku, N.S.; Choi, J.Y.; Kim, C.-M.; et al. A Case of African tick-bite fever in a returning traveler from Southern Africa. Infect. Chemother. 2020, 52, e39. [Google Scholar] [CrossRef]
- Haemel, A.K.; Bearden, A.; Longley, B.J.; Crnich, C. Black spots in the returning traveler. Dermatol. Online J. 2013, 19, 20393. [Google Scholar] [CrossRef] [PubMed]
- Oostvogel, P.M.; Van Doornum, G.J.; Ferreira, R.; Vink, J.; Fenollar, F.; Raoult, D. African tickbite fever in travelers, Swaziland. Emerg. Infect. Dis. 2007, 13, 353–355. [Google Scholar] [CrossRef] [PubMed]
- Swantje Buechau, A.; Wurthner, J.U.; Reifenberger, J.; Ruzicka, T. Fever, episcleritis, epistaxis, and rash after safari holiday in Swaziland. Arch. Dermatol. 2006, 142, 1365–1366. [Google Scholar]
- Neal, S.; Cieslak, P.; Hedberg, K. African tick-bite fever among international travelers—Oregon, 1998. MMWR Morb. Mortal. Wkly. Rep. 1998, 47, 950–952. [Google Scholar]
- Simpson, G.; Quan, V.; Frean, J.; Knobel, D.L.; Rossouw, J.; Weyer, J.; Marcotty, T.; Godfroid, J.; Blumberg, L.H. Prevalence of selected zoonotic diseases and risk factors at a human-wildlife-livestock interface in Mpumalanga Province, South Africa. Vector Borne Zoonotic Dis. 2018, 18, 303–310. [Google Scholar] [CrossRef] [PubMed]
- Socolovschi, C.; Huynh, T.; Davoust, B.; Gómez, J.; Raoult, D.; Parola, P. Transovarial and trans-stadial transmission of Rickettsiae africae in Amblyomma variegatum ticks. Clin. Microbiol. Infect. 2009, 15, 317–318. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saraiva, D.G.; Bastos, F.A.N.; Horta, M.C.; Soares, H.S.; Nicola, P.; Pereira, L.C.M.; Labruna, M.B. Rickettsia amblyommii Infecting Amblyomma auricularium ticks in Pernambuco, Northeastern Brazil: Isolation, transovarial transmission, and transstadial perpetuation. Vector Borne Zoonotic Dis. 2013, 13, 615–618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsumoto, K.; Ogawa, M.; Brouqui, P.; Raoult, D.; Parola, P. Transmission of Rickettsia massiliae in the tick, Rhipicephalus turanicus. Med. Veter. Entomol. 2005, 19, 263–270. [Google Scholar] [CrossRef]
- Dumler, J.S.; Barbet, A.F.; Bekker, C.P.J.; Dasch, G.A.; Palmer, G.H.; Ray, S.; Rikihisa, Y.; Rurangirwa, F.R. Reorganization of genera in the families Rickettsiaceae and Anaplasmataceae in the order Rickettsiales: Unification of some species of Ehrlichia with Anaplasma, Cowdria with Ehrlichia and Ehrlichia with Neorickettsia, descriptions of six new species combinations and designation of Ehrlichia equi and ‘HGE agent’ as subjective synonyms of Ehrlichia phagocytophila. Int. J. Syst. Evol. Microbiol. 2001, 51, 2145–2165. [Google Scholar] [CrossRef] [Green Version]
- Rar, V.; Golovljova, I. Anaplasma, ehrlichia, and “candidatus neoehrlichia” bacteria: Pathogenicity, biodiversity, and molecular genetic characteristics, a review. Infect. Genet. Evol. 2011, 11, 1842–1861. [Google Scholar] [CrossRef]
- Machado, R.Z.; Teixeira, M.M.G.; Rodrigues, A.C.; André, M.R.; Gonçalves, L.R.; Da Silva, J.B.; Pereira, C.L. Molecular diagnosis and genetic diversity of tick-borne Anaplasmataceae agents infecting the African buffalo Syncerus caffer from Marromeu Reserve in Mozambique. Parasites Vectors 2016, 9, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Vlahakis, P.A.; Chitanga, S.; Simuunza, M.C.; Simulundu, E.; Qiu, Y.; Changula, K.; Chambaro, H.M.; Kajihara, M.; Nakao, R.; Takada, A.; et al. Molecular detection and characterization of zoonotic Anaplasma species in domestic dogs in Lusaka, Zambia. Ticks Tick Borne Dis. 2018, 9, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Kolo, A.O.; Sibeko-Matjila, K.; Maina, A.N.; Richards, A.L.; Knobel, D.; Matjila, P.; Paul, T. Molecular Detection of Zoonotic Rickettsiae and Anaplasma spp. in domestic dogs and their ectoparasites in Bushbuckridge, South Africa. Vector Borne Zoonotic Dis. 2016, 16, 245–252. [Google Scholar] [CrossRef] [Green Version]
- Slodki, J.; Jasik, K.P.; Kepa, M.; Idzik, D.; Wojtyczka, R.D. Tick-transmitted diseases caused by Apicomplexa. Acta Protozool. 2011, 50, 155–161. [Google Scholar] [CrossRef]
- Reichard, M.V.; Edwards, A.C.; Meinkoth, J.H.; Snider, T.A.; Meinkoth, K.R.; Heinz, R.E.; Little, S.E. Confirmation of Amblyomma americanum (Acari: Ixodidae) as a vector for Cytauxzoon felis (Piroplasmorida: Theileriidae) to domestic cats. J. Med. Entomol. 2010, 47, 890–896. [Google Scholar] [CrossRef] [PubMed]
- Mans, B.J.; Pienaar, R.; Latif, A. A review of Theileria diagnostics and epidemiology. Int. J. Parasitol. Parasites Wildl. 2015, 4, 104–118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, T.G. The Genus Hepatozoon (apicomplexa: Adeleina). J. Parasitol. 1996, 82, 565. [Google Scholar] [CrossRef] [PubMed]
- Despommier, D.; Ellis, B.R.; Wilcox, B.A. The role of ecotones in emerging infectious diseases. EcoHealth 2007, 3, 281–289. [Google Scholar] [CrossRef]
- Theiler, G. Zoological survey of the union of South Africa. Tick survey—Part, I. Distribution of Amblyomma hebraeum, the Heartwater Tick. Onderstepoort J. Vet. Res. Anim. Ind. 1948, 23, 217–231. [Google Scholar]
- Theiler, G. Zoological survey of the union of South Africa: Tick survey. Part II. Distribution of Boophilus (Palpoboophilus) decoloratus, the blue tick. Onderstepoort J. Vet. Res. Anim. Ind. 1949, 22, 255–268. [Google Scholar]
- Theiler, G. Zoological survey of the union of South Africa: Tick survey. Part III. Distribution of Rhipicephalus appendiculatus, the brown tick. Onderstepoort J. Vet. Res. Anim. Ind. 1949, 22, 269–284. [Google Scholar]
- Theiler, G. Zoological survey of the union of South Africa. Tick survey. Part, V. Distribution of Rhipicephalus evertsi, the red tick. Onderstepoort J. Vet. Res. Anim. Ind. 1950, 19, 33–36. [Google Scholar]
- Theiler, G. Zoological survey of the union of South Africa. Tick survey. Part IX. The distribution of the three South African Hyalommas or bontpoots. Onderstepoort J. Vet. Res. Anim. Ind. 1956, 27, 239–269. [Google Scholar]
- Walker, J.B.; Keirans, J.E.; Horak, I.G. The Genus Rhipicephalus (Acari, Ixodidae): A Guide to the Brown Ticks of the World; Cambridge University Press: Cambridge, UK, 2005. [Google Scholar]
- Mtambo, J.; Madder, M.; Van Bortel, W.; Berkvens, D.; Backeljau, T. Rhipicephalus appendiculatus and R. zambeziensis (Acari: Ixodidae) from Zambia: A molecular reassessment of their species status and identification. Exp. Appl. Acarol. 2007, 41, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Beati, L.; Keirans, J.E. Analysis of the systematic relationships among ticks of the Genera Rhipicephalus and Boophilus (Acari: Ixodidae) based on mitochondrial 12S ribosomal DNA gene sequences and morphological characters. J. Parasitol. 2001, 87, 32–48. [Google Scholar] [CrossRef]
- Bakkes, D.K.; Ropiquet, A.; Chitimia-Dobler, L.; Matloa, D.E.; Apanaskevich, D.A.; Horak, I.G.; Mans, B.J.; Matthee, C.A. Adaptive radiation and speciation in Rhipicephalus ticks: A medley of novel hosts, nested predator-prey food webs, off-host periods and dispersal along temperature variation gradients. Mol. Phylogenet. Evol. 2021, 162, 107178. [Google Scholar] [CrossRef]
- Dlamini, B.N.; Mdluli, S.; Mudyanavana, C.; Chikuni, N.E.; Masarirambi, M.T. Rabies in Eswatini: What are the Issues and Challenges? J. Adv. Microbiol. 2020, 20, 21–28. [Google Scholar] [CrossRef]
- Horak, I.G.; Heyne, H.; Williams, R.; Gallivan, G.J.; Spickett, A.M.; Bezuidenhout, J.D.; Estrada-Peña, A. The Ixodid Ticks (Acari: Ixodidae) of Southern Africa; Springer: Berlin/Heidelberg, Germany, 2018. [Google Scholar]
- Horak, I.G.; Emslie, F.R.; Spickett, A.M. Parasites of domestic and wild animals in South Africa. XL. Ticks on dogs belonging to people in rural communities and carnivore ticks on the vegetation. Onderstepoort J. Veter. Res. 2001, 68, 135–141. [Google Scholar]
- Horak, I.; Fourie, L.; Heyne, H.; Walker, J.B.; Needham, G. Ixodid Ticks Feeding on Humans in South Africa: With notes on preferred hosts, geographic distribution, seasonal occurrence and transmission of pathogens. Exp. Appl. Acarol. 2002, 27, 113–136. [Google Scholar] [CrossRef]
- Vitale, G.; Mansueto, S.; Rolain, J.-M.; Raoult, D. Rickettsia massiliae human isolation. Emerg. Infect. Dis. 2006, 12, 174–175. [Google Scholar] [CrossRef]
- Cascio, A.; Torina, A.; Valenzise, M.; Blanda, V.; Camarda, N.; Bombaci, S.; Iaria, C.; De Luca, F.; Wasniewska, M. Scalp eschar and neck lymphadenopathy caused by rickettsia massiliae. Emerg. Infect. Dis. 2013, 19, 836–837. [Google Scholar] [CrossRef] [Green Version]
- Harris, D.J.; Halajian, A.; Santos, J.L.; Swanepoel, L.H.; Taylor, P.J.; Xavier, R. Diversity of haemoprotozoan parasites infecting the wildlife of South Africa. Folia Parasitol. 2018, 65, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Beati, L.; Kelly, P.J.; Matthewman, L.A.; Mason, P.R.; Raoult, D. Prevalence of Rickettsia-like organisms and spotted fever group rickettsiae in ticks (acari: Ixodidae) from Zimbabwe. J. Med. Entomol. 1995, 32, 787–792. [Google Scholar] [CrossRef] [PubMed]
- Sarih, M.; Socolovschi, C.; Boudebouch, N.; Hassar, M.; Raoult, D.; Parola, P. Spotted fever group rickettsiae in ticks, Morocco. Emerg. Infect. Dis. 2008, 14, 1067–1073. [Google Scholar] [CrossRef] [PubMed]
- Berrelha, J.; Briolant, S.; Muller, F.; Rolain, J.-M.; Marie, J.-L.; Pagés, F.; Raoult, D.; Parola, P. Rickettsia felis and rickettsia massiliae in Ivory Coast, Africa. Clin. Microbiol. Infect. 2009, 15, 251–252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cabezas-Cruz, A.; Zweygarth, E.; Vancová, M.; Broniszewska, M.; Grubhoffer, L.; Passos, L.M.F.; Ribeiro, M.F.B.; Alberdi, P.; De La Fuente, J. Ehrlichia minasensis sp. nov., isolated from the tick Rhipicephalus microplus. Int. J. Syst. Evol. Microbiol. 2016, 66, 1426–1430. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cabezas-Cruz, A.; Zweygarth, E.; Aguiar, D.M. Ehrlichia minasensis, an old demon with a new name. Ticks Tick Borne Dis. 2019, 10, 828–829. [Google Scholar] [CrossRef]
- Lobanov, V.A.; Gajadhar, A.A.; Al-Adhami, B.; Schwantje, H.M. Molecular study of free-ranging mule deer and white-tailed deer from British Columbia, Canada, for evidence of anaplasma spp. and ehrlichia spp. Transbound. Emerg. Dis. 2011, 59, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Thomson, K.; Yaaran, T.; Belshaw, A.; Curson, L.; Tisi, L.; Maurice, S.; Kiddle, G. A new TaqMan method for the reliable diagnosis of ehrlichia spp. in canine whole blood. Parasites Vectors 2018, 11, 350. [Google Scholar] [CrossRef]
- Oura, C.A.L.; Tait, A.; Asiimwe, B.; Lubega, G.W.; Weir, W. Theileria parva genetic diversity and haemoparasite prevalence in cattle and wildlife in and around Lake Mburo National Park in Uganda. Parasitol. Res. 2010, 108, 1365–1374. [Google Scholar] [CrossRef] [PubMed]
- Mans, B.J.; Pienaar, R.; Latif, A.A.; Potgieter, F.T. Diversity in the 18S SSU rRNA V4 hyper-variable region of Theileria spp. in Cape buffalo (Syncerus caffer) and cattle from southern Africa. Parasitology 2011, 138, 766–779. [Google Scholar] [CrossRef] [Green Version]
- Njiiri, N.E.; Bronsvoort, M.; Collins, N.; Steyn, H.C.; Troskie, M.; Vorster, I.; Thumbi, S.; Sibeko-Matjila, K.; Jennings, A.; van Wyk, I.C.; et al. The epidemiology of tick-borne haemoparasites as determined by the reverse line blot hybridization assay in an intensively studied cohort of calves in western Kenya. Veter. Parasitol. 2015, 210, 69–76. [Google Scholar] [CrossRef] [Green Version]
- Steyl, J.C.; Prozesky, L.; Stoltsz, W.H.; Lawrence, J.A. Theileriosis (Cytauxzoonosis) in Roan antelope (Hippotragus equinus): Field exposure to infection and identification of potential vectors. Onderstepoort J. Veter. Res. 2012, 79, 8. [Google Scholar] [CrossRef]
- Altay, K.; Dumanli, N.; Aktas, M. Molecular identification, genetic diversity and distribution of Theileria and Babesia species infecting small ruminants. Veter. Parasitol. 2007, 147, 161–165. [Google Scholar] [CrossRef]
- Gholami, S.; Laktarashi, B.; Shiadeh, M.M.; Spotin, A. Genetic variability, phylogenetic evaluation and first global report of Theileria luwenshuni, T. buffeli, and T. ovis in sheepdogs in Iran. Parasitol. Res. 2016, 115, 2125–2130. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, J.; Byaruhanga, C.; Oosthuizen, M.; Mans, B. Theileriosis of sheep and goats. In Infectious Diseases of Livestock; Coetzer, J., Thomson, G., Maclachlan, J., Eds.; Anipedia: Pretoria, South Africa, 2017. [Google Scholar]
- Pienaar, R.; Josemans, A.; Latif, A.A.; Mans, B.J. The host-specificity of Theileria sp. (sable) and Theileria sp. (sable-like) in African Bovidae and detection of novel Theileria in antelope and giraffe. Parasitology 2019, 147, 213–224. [Google Scholar] [CrossRef] [Green Version]
- Scoles, G.A.; Ueti, M.W. Vector Ecology of Equine Piroplasmosis. Annu. Rev. Entomol. 2015, 60, 561–580. [Google Scholar] [CrossRef]
- De Sousa, K.C.M.; Fernandes, M.P.; Herrera, H.; Freschi, C.R.; Machado, R.Z.; André, M.R. Diversity of piroplasmids among wild and domestic mammals and ectoparasites in Pantanal wetland, Brazil. Ticks Tick Borne Dis. 2018, 9, 245–253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dahmana, H.; Granjon, L.; Diagne, C.; Davoust, B.; Fenollar, F.; Mediannikov, O. Rodents as hosts of pathogens and related zoonotic disease risk. Pathogens 2020, 9, 202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mangombi, J.B.; N’Dilimabaka, N.; Lekana-Douki, J.-B.; Banga, O.; Maghendji-Nzondo, S.; Bourgarel, M.; Leroy, E.; Fenollar, F.; Mediannikov, O. First investigation of pathogenic bacteria, protozoa and viruses in rodents and shrews in context of forest-savannah-urban areas interface in the city of Franceville (Gabon). PLoS ONE 2021, 16, e0248244. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.R. Ticks of Domestic Animals in Africa: A Guide to Identification of Species; Bioscience Reports: Edinburgh, UK, 2003; pp. 3–210. [Google Scholar]
- Smith, T.; Kilbourne, F. Investigations into the Nature Causation and Prevention of Texas or Southern Cattle Fever; US Department of Agriculture, Bureau of Animal Industry: Washington, DC, USA, 1893.
- Gray, J.S.; Estrada-Peña, A.; Zintl, A. Vectors of Babesiosis. Annu. Rev. Entomol. 2019, 64, 149–165. [Google Scholar] [CrossRef] [PubMed]
- Motloang, M.Y.; Thekisoe, O.M.; Alhassan, A.; Bakheit, M.; Motheo, M.P.; Masangane, F.E.; Thibedi, M.L.; Inoue, N.; Igarashi, I.; Sugimoto, C.; et al. Prevalence of theileria equi and babesia caballi infections in horses belonging to resource-poor farmers in the north-eastern free state province, South Africa. Onderstepoort J. Veter. Res. 2008, 75, 141–146. [Google Scholar] [CrossRef]
- Zobba, R.; Parpaglia, M.L.P.; Spezzigu, A.; Pittau, M.; Alberti, A. First Molecular identification and phylogeny of a babesia sp. from a symptomatic sow (sus scrofa linnaeus 1758). J. Clin. Microbiol. 2011, 49, 2321–2324. [Google Scholar] [CrossRef] [Green Version]
- Avenant, A.; Park, J.Y.; Vorster, I.; Mitchell, E.P.; Arenas-Gamboa, A.M. Porcine babesiosis caused by babesia sp. suis in a pot-bellied pig in South Africa. Front. Veter. Sci. 2021, 7, 1129. [Google Scholar] [CrossRef]
- De Waal, D.T.; Rebollar, L.M.L.; Potgieter, F.T. The transovarial transmission of Babesia trautmanni by Rhipicephalus simus to domestic pigs. Onderstepoort J. Veter. Res. 1992, 59, 219–221. [Google Scholar]
- Starkey, L.A.; Panciera, R.J.; Paras, K.; Allen, K.E.; Reiskind, M.; Reichard, M.V.; Johnson, E.M.; Little, S.E. Genetic diversity of hepatozoon spp. in coyotes from the south-central United States. J. Parasitol. 2013, 99, 375–378. [Google Scholar] [CrossRef]
- Maia, J.P.; Álvares, F.; Boratynski, Z.; Brito, J.; Leite, J.V.; Harris, D.J. Molecular assessment of hepatozoon (apicomplexa: Adeleorina) infections in wild canids and rodents from North Africa, with implications for transmission dynamics across taxonomic groups. J. Wildl. Dis. 2014, 50, 837–848. [Google Scholar] [CrossRef] [PubMed]
- Levi, M.M.; Nachum-Biala, Y.; King, R.; Baneth, G. A survey of babesia spp. and hepatozoon spp. in wild canids in Israel. Parasites Vectors 2018, 11, 150. [Google Scholar] [CrossRef] [PubMed]
- Clark, G.M. Hepatozoon griseisciuri n. sp.; a new species of hepatozoon from the grey squirrel (sciurus carolinensis gmelin, 1788), with studies on the life cycle. J. Parasitol. 1958, 44, 52. [Google Scholar] [CrossRef] [PubMed]
- Silaghi, C.; Woll, D.; Hamel, D.; Pfister, K.; Mahling, M.; Pfeffer, M. Babesia spp. and Anaplasma phagocytophilum in questing ticks, ticks parasitizing rodents and the parasitized rodents—Analyzing the host-pathogen-vector interface in a metropolitan area. Parasites Vectors 2012, 5, 191. [Google Scholar] [CrossRef] [Green Version]
- Kamani, J.; Harrus, S.; Nachum-Biala, Y.; Gutiérrez, R.; Mumcuoglu, K.Y.; Baneth, G. Prevalence of hepatozoon and sarcocystis spp. in rodents and their ectoparasites in Nigeria. Acta Trop. 2018, 187, 124–128. [Google Scholar] [CrossRef]
- Maia, J.P.M.C.; Harris, D.J.; Perera, A. Molecular survey of hepatozoon species in lizards from North Africa. J. Parasitol. 2011, 97, 513–517. [Google Scholar] [CrossRef]
- Vilcins, I.-M.E.; Ujvari, B.; Old, J.M.; Deane, E. Molecular and morphological description of a hepatozoon species in reptiles and their ticks in the Northern Territory, Australia. J. Parasitol. 2009, 95, 434–442. [Google Scholar] [CrossRef] [Green Version]
- Haklová, B.; Majlathova, V.; Majláth, I.; Harris, D.J.; Petrilla, V.; Litschka-Koen, T.; Oros, M.; Peťko, B. Phylogenetic relationship of Hepatozoon blood parasites found in snakes from Africa, America and Asia. Parasitology 2014, 141, 389–398. [Google Scholar] [CrossRef] [PubMed]
- Viljoen, S.; O’Riain, M.J.; Penzhorn, B.L.; Drouilly, M.; Serieys, L.E.K.; Cristescu, B.; Teichman, K.J.; Bishop, J.M. Molecular detection of tick-borne pathogens in caracals (Caracal caracal) living in human-modified landscapes of South Africa. Parasites Vectors 2020, 13, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Matjila, P.T.; Leisewitz, A.; Jongejan, F.; Bertschinger, H.J.; Penzhorn, B. (Banie) Molecular detection of Babesia rossi and Hepatozoon sp. in African wild dogs (Lycaon pictus) in South Africa. Veter. Parasitol. 2008, 157, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Harris, D.J.; Pereira, A.; Halajian, A.; Luus-Powell, W.J.; Kunutu, K.D. Screening for Hepatozoon parasites in gerbils and potential predators in South Africa. J. South. Afr. Veter. Assoc. 2017, 88, 1–4. [Google Scholar] [CrossRef]
- Netherlands, E.C.; Cook, C.A.; Kruger, D.J.; du Preez, L.H.; Smit, N.J. Biodiversity of frog haemoparasites from sub-tropical northern KwaZulu-Natal, South Africa. Int. J. Parasitol. Parasites Wildl. 2015, 4, 135–141. [Google Scholar] [CrossRef] [Green Version]
- Allen, K.E.; Johnson, E.M.; Little, S.E. Hepatozoon spp Infections in the United States. Veter. Clin. North. Am. Small Anim. Pr. 2011, 41, 1221–1238. [Google Scholar] [CrossRef]
- Demoner, L.D.C.; Rubini, A.S.; Paduan, K.D.S.; Metzger, B.; Antunes, J.M.A.D.P.; Martins, T.F.; Camargo-Mathias, B.; O’Dwyer, L.H. Investigation of tick vectors of hepatozoon canis in Brazil. Ticks Tick Borne Dis. 2013, 4, 542–546. [Google Scholar] [CrossRef]
- McCully, R.M.; Basson, A.P.; Bigalke, R.D.; De Vos, V.; Young, E. Observations on naturally acquired hepatozoonosis of wild carnivores and dogs in the Republic of South Africa. Onderstepoort J. Veter. Res. 1975, 42, 117–134. [Google Scholar]
- Stampa, S. Tick paralysis in the Karoo areas of South Africa. Onderstepoort J. Vet. Res. 1959, 28, 169–227. [Google Scholar]
- Zhang, H.-M.; Huang, B.; Lawrimore, J.; Menne, M.; Smith, T.M. NOAA Global Surface Temperature Dataset (NOAAGlobalTemp); Version 5; Physical Sciences Laboratory: New York, NY, USA, 2018. [Google Scholar] [CrossRef]
- Tsao, I.J.; Hamer, A.S.; Han, S.; Sidge, J.L.; Hickling, G.J. The Contribution of wildlife hosts to the rise of ticks and tick-borne diseases in North America. J. Med. Entomol. 2021, 58, 1565–1587. [Google Scholar] [CrossRef]
- Levin, M.L.; Killmaster, L.F.; Zemtsova, G.E. Domestic dogs (Canis familiaris) as reservoir hosts forrickettsia conorii. Vector Borne Zoonotic Dis. 2012, 12, 28–33. [Google Scholar] [CrossRef]
- Salje, J.; Weitzel, T.; Newton, P.N.; Varghese, G.M.; Day, N. Rickettsial infections: A blind spot in our view of neglected tropical diseases. PLoS Negl. Trop. Dis. 2021, 15, e0009353. [Google Scholar] [CrossRef]
- Goudie, A. The Atlas of Swaziland; Swaziland National Trust Commission: Lobamba, Eswatini, 1983. [Google Scholar]
- Monadjem, A. Full Terrestrial Vertebrate Survey of IYSIS Ranch; Queensland Government: Queensland, Australia, 2017.
- Soto-Shoender, J.R.; McCleery, R.A.; Monadjem, A.; Gwinn, D.C. The importance of grass cover for mammalian diversity and habitat associations in a bush encroached savanna. Biol. Conserv. 2018, 221, 127–136. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Penton, E.H.; Burns, J.M.; Janzen, D.H.; Hallwachs, W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci. USA 2004, 101, 14812–14817. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barrett, R.; Hebert, P. Identifying spiders through DNA barcodes. Can. J. Zool. 2005, 83, 481–491. [Google Scholar] [CrossRef]
- Loftis, A.D.; Abbassy, M.M.; Dasch, G.; Szumlas, D.E.; Helmy, I.M.; Moriarity, J.R.; Reeves, W.K. Surveillance of Egyptian fleas for agents of public health significance: Anaplasma, bartonella, coxiella, ehrlichia, rickettsia, and yersinia pestis. Am. J. Trop. Med. Hyg. 2006, 75, 41–48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Regnery, R.L.; Spruill, C.L.; Plikaytis, B.D. Genotypic identification of rickettsiae and estimation of intraspecies sequence divergence for portions of two rickettsial genes. J. Bacteriol. 1991, 173, 1576–1589. [Google Scholar] [CrossRef] [Green Version]
- Roux, V.; Fournier, E.P.; Raoult, D. Differentiation of spotted fever group rickettsiae by sequencing and analysis of restriction fragment length polymorphism of PCR-amplified DNA of the gene encoding the protein rOmpA. J. Clin. Microbiol. 1996, 34, 2058–2065. [Google Scholar] [CrossRef] [Green Version]
- Roux, V.; Raoult, D. Phylogenetic analysis of members of the genus Rickettsia using the gene encoding the outer-membrane protein rOmpB (ompB). Int. J. Syst. Evol. Microbiol. 2000, 50, 1449–1455. [Google Scholar] [CrossRef] [Green Version]
- Roux, V.; Rydkina, E.; Eremeeva, M.; Raoult, D. Citrate synthase gene comparison, a new tool for phylogenetic analysis, and its application for the rickettsiae. Int. J. Syst. Bacteriol. 1997, 47, 252–261. [Google Scholar] [CrossRef] [Green Version]
- Dahmani, M.; Davoust, B.; Benterki, M.S.; Fenollar, F.; Raoult, D.; Mediannikov, O. Development of a new PCR-based assay to detect anaplasmataceae and the first report of anaplasma phagocytophilum and anaplasma platys in cattle from Algeria. Comp. Immunol. Microbiol. Infect. Dis. 2015, 39, 39–45. [Google Scholar] [CrossRef]
- Parola, P.; Cornet, J.-P.; Sanogo, Y.O.; Miller, R.S.; Van Thien, H.; Gonzalez, J.-P.; Raoult, D.; Telford, S.R.; Wongsrichanalai, C. Detection of Ehrlichia spp., Anaplasma spp., Rickettsia spp., and Other Eubacteria in Ticks from the Thai-Myanmar Border and Vietnam. J. Clin. Microbiol. 2003, 41, 1600–1608. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Vienna, Austria, 2021. [Google Scholar]
- Wang, L.-G.; Lam, T.T.-Y.; Xu, S.; Dai, Z.; Zhou, L.; Feng, T.; Guo, P.; Dunn, C.W.; Jones, B.R.; Bradley, T.; et al. Treeio: An R package for phylogenetic tree input and output with richly annotated and associated data. Mol. Biol. Evol. 2020, 37, 599–603. [Google Scholar] [CrossRef]
- Yu, G.; Smith, D.K.; Zhu, H.; Guan, Y.; Lam, T.T.Y. Ggtree: An r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 2016, 8, 28–36. [Google Scholar] [CrossRef]
- Essbauer, S.; Hofmann, M.; Kleinemeier, C.; Wölfel, S.; Matthee, S. Rickettsia diversity in southern Africa: A small mammal perspective. Ticks Tick Borne Dis. 2018, 9, 288–301. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, E.; Kock, R.; McKeever, D.; Gakuya, F.; Musyoki, C.; Chege, S.M.; Mutinda, M.; Kariuki, E.; Davidson, Z.; Low, B.; et al. Prevalence of theileria equi and babesia caballi as well as the identification of associated ticks in sympatric grevy’s zebras (equus grevyi) and donkeys (equus africanus asinus) in Northern Kenya. J. Wildl. Dis. 2015, 51, 137–147. [Google Scholar] [CrossRef]
- Greay, T.L.; Zahedi, A.; Krige, A.-S.; Owens, J.M.; Rees, R.L.; Ryan, U.M.; Oskam, C.L.; Irwin, P.J. Endemic, exotic and novel apicomplexan parasites detected during a national study of ticks from companion animals in Australia. Parasites Vectors 2018, 11, 197. [Google Scholar] [CrossRef] [Green Version]
- Han, R.; Yang, J.-F.; Mukhtar, M.U.; Chen, Z.; Niu, Q.-L.; Lin, Y.-Q.; Liu, G.-Y.; Luo, J.-X.; Yin, H.; Liu, Z.-J. Molecular detection of Anaplasma infections in ixodid ticks from the Qinghai-Tibet Plateau. Infect. Dis. Poverty 2019, 8, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Stevenson, M.; Nunes, T.; Sanchez, J.; Thornton, R.; Reiczigel, J.; Robison-Cox, J.; Sebastiani, P.; Solymos, P.; Yoshida, K.; Jones, G.; et al. epiR: An R Package for the Analysis of Epidemiological Data; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
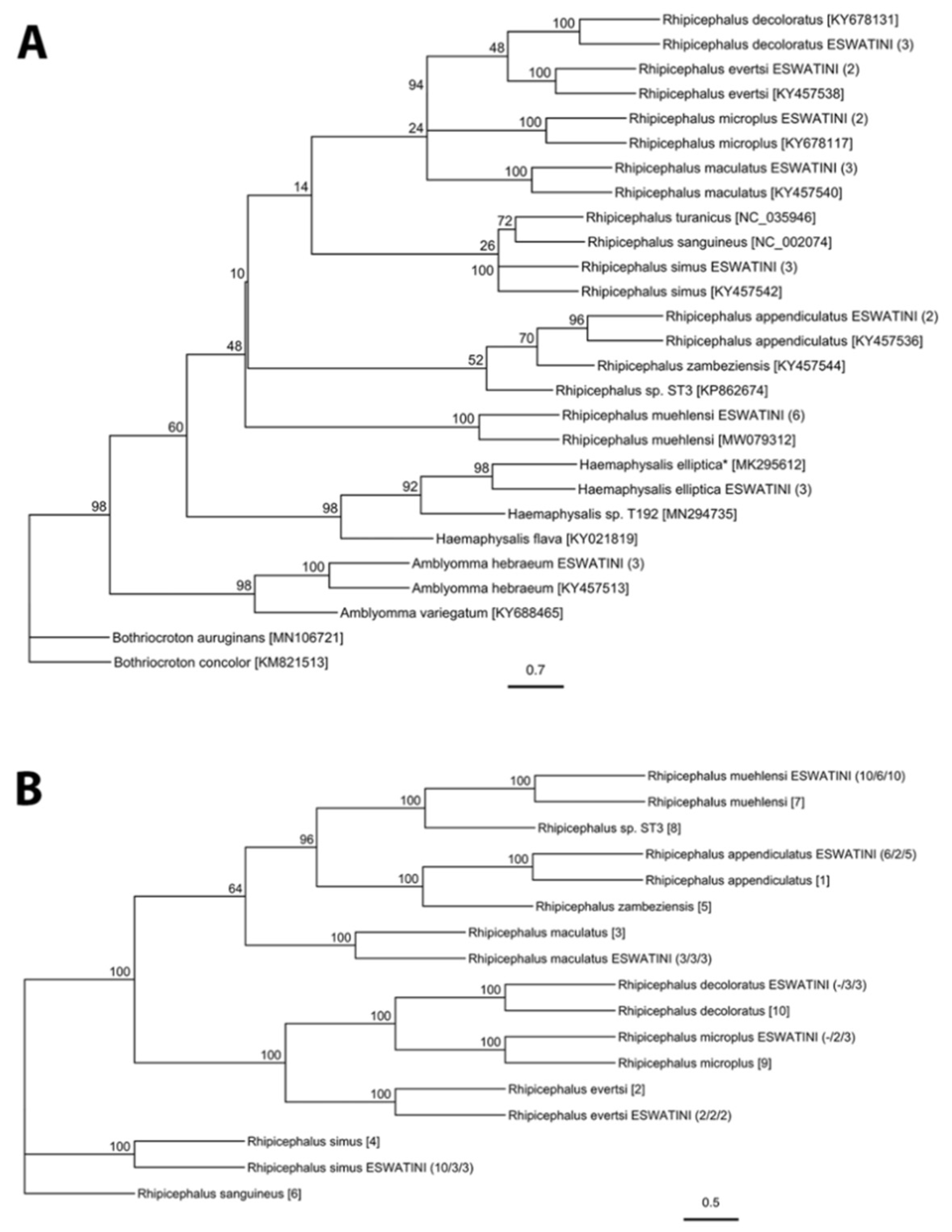

| Tick Species | ITS2 | CO1 | 12S |
|---|---|---|---|
| Amblyomma hebraeum | Amblyomma hebraeum: KY457490 (1059/1064 = 99.5%) | Amblyomma hebraeum: KY457513 (629/629 = 100%) | Amblyomma hebraeum: KY457513 (378/379 = 99.7%) |
| Haemaphysalis elliptica | Haemaphysalis elliptica: MK295615 (416/432 = 96.3%) Haemaphysalis sp. T192: MN266944 (1042/1049 = 99.3) | Haemaphysalis ellitptica *: MK295612 (630/660 = 95.5%) Haemaphysalis sp. T192: MN294735 (598/668 = 89.5%) | Haemaphysalis elliptica: HM068953 (373/378 = 98.7%) |
| Rhipicephalus appendiculatus | Rhipicephalus appendiculatus: KY457500 (1199/1200 = 99.9%) | Rhipicephalus appendiculatus: KY457536 (645/645 = 100%) | Rhipicephalus appendiculatus: KY457536 (374/276 = 99.5%) |
| Rhipicephalus evertsi | Rhipicephalus evertsi: KY457503 (1204/1212 = 99.3%) | Rhipicephalus evertsi: KY457538 (702/702 = 99.3%) | Rhipicephalus evertsi: KY457538 (384/388 = 99.0%) |
| Rhipicephalus maculatus | Rhipicephalus maculatus: KY457505 (1081/1081 = 100%) | Rhipicephalus maculatus: KY457540 (629/631 = 99.7%) | Rhipicephalus maculatus: KY457540 (381/385 = 99.0%) |
| Rhipicephalus muehlensi | Rhipicephalus sp. ST3: KP862668 (1109/1112 = 99.7%) Rhipicephalus appendiculatus: KY457500 (1123/1218 = 92.2%) | Rhipicephalus muehlensi: MW079312 (496/497 = 99.8%) Rhipicephalus sp. ST3: KP862674 (571/645 = 88.5%) Rhipicephalus appendiculatus: KY457536 (569/647 = 87.9%) | Rhipicephalus muehlensi: MW080169 (317/318 = 99.7%) Rhipicephalus appendiculatus: KY457536 (347/377 = 92.0%) |
| Rhipicephalus simus | Rhipicephalus simus: KY457508 (1228/1228 = 100%) | Rhipicephalus simus: KY457542 (631/634 = 99.5%) | Rhipicephalus simus: KY457542 (375/377 = 99.5%) |
| Amblyomma larvae | Amblyomma hebraeum: KY457490 (1079/1079 = 100%) | Amblyomma hebraeum: KY457513 (629/629 = 100%) | Amblyomma hebraeum: KY457513 (378/379 = 99.7%) |
| Rhipicephalus (Boophilus) larvae | Rhipicephalus decoloratus: MN266919 (1016/1054 = 96.4%) | Rhipicephalus decoloratus: KY678131 (634/636 = 99.7%) | NT |
| Rhipicephalus microplus: KY457506 (1246/1247= 99.9%) | Rhipicephalus microplus: KY678117 (635/635 = 100%) | NT | |
| Rhipicephalus larvae | Rhipicephalus appendiculatus: KY457500 (1159/1160 = 99.9%) | NT | NT |
| Rhipicephalus sp. ST3: KP862668 (1109/1112 = 99.7%) Rhipicephalus appendiculatus: KY457500 (1120/1215 = 92.2%) | NT | NT | |
| Rhipicephalus maculatus: KY457505 (1108/1108 = 100%) | NT | NT |
| Pathogen Species/Genotype | Amblyomma hebraeum | Haemaphysalis elliptica | Rhipicephlaus simus | |
|---|---|---|---|---|
| Adult (n = 12) | Nymph (n = 38) | Adult (n = 219) | Adult (n = 494) | |
| Rickettsia africae | 1; 8.33% (0.2–38.5%) | 19; 50% (33.4–66.6%) | 0 | 0 |
| Candidatus Rickettsia barbariae | 0 | 0 | 0 | 8; 1.6% (0.7–3.1%) |
| Rickettsia conorii | 0 | 0 | 9; 4.1% (1.9–7.7%) | 0 |
| Rickettsia massiliae | 0 | 0 | 8; 3.7% (1.6–7.1%) | 14; 2.8% (1.6–4.7%) |
| Rickettsia sp. A | 0 | 2; 5.26% (0.6–17.9%) | 0 | 0 |
| Rickettsia sp. B | 0 | 0 | 0 | 9; 1.8% (0.8–3.4%) |
| Rickettsia sp. C | 0 | 0 | 0 | 2; 0.4% (0.05–1.5%) |
| Richness | 2 | 2 | 4 | |
| A. hebraeum | H. elliptica | R. appendiculatus | R. maculatus | R. simus | R. muehlensi | ||
|---|---|---|---|---|---|---|---|
| Pathogen Species/Genotype | Nymph (n = 38) | Adult (n = 219) | Adult (n = 146) | Nymph (n = 554) | Adult (n = 93) | Adult (n = 494) | Adult (n = 39) |
| Anaplasmataceae | 1; 2.6% (0.1–13.8%) | 0 | 0 | 0 | 0 | 0 | 0 |
| Anaplasma sp. A | 0 | 0 | 4; 2.8% (0.8–6.9%) | 7; 1.3% (0.5–2.6%) | 0 | 0 | 3; 7.7% (1.6–20.1%) |
| Anaplasma sp. B | 0 | 38; 17.4% (12.6–23.0%) | 0 | 0 | 0 | 0 | 0 |
| Anaplasma sp. C | 0 | 0 | 0 | 0 | 0 | 13; 2.6% (1.4–4.5%) | 0 |
| Anaplasma sp. D | 0 | 0 | 3; 2.1% (0.4–5.9%) | 3; 0.5% (0.1–1.6%) | 0 | 0 | 0 |
| Anaplasma sp. E | 0 | 1; 0.5% (0.01–2.5%) | 0 | 0 | 0 | 0 | 0 |
| Anaplasma sp. F | 0 | 0 | 0 | 0 | 0 | 1; 0.2% (0.01–1.1%) | 0 |
| Anaplasma sp. G | 0 | 0 | 0 | 2; 0.4% (0.04–1.4%) | 0 | 0 | 0 |
| Ehrlichia minasensis | 0 | 0 | 0 | 0 | 0 | 1; 0.2% (0.01–1.12%) | 1; 2.6% (0.06–13.5%) |
| Ehrlichia ruminantium | 1; 2.6% (0.1–13.8%) | 0 | 0 | 0 | 0 | 0 | 0 |
| Ehrlichia sp. A | 0 | 0 | 2; 1.4% (0.2–4.9%) | 0 | 0 | 47; 9.5% (7.1–12.5%) | 0 |
| Ehrlichia sp. B | 0 | 9; 4.2% (1.9–7.6%) | 0 | 0 | 0 | 0 | 0 |
| Ehrlichia sp. C | 0 | 0 | 3; 2.1% (0.4–5.9%) | 6; 1.1%; (0.4–2.3%) | 2; 2.2% (0.3–7.6) | 0 | 0 |
| Ehrlichia sp. D | 0 | 0 | 1; 0.7% (0.2–3.8%) | 2; 0.4% (0.04–1.3%) | 0 | 0 | 1; 2.6% (0.06–13.5%) |
| Ehrlichia sp. E | 0 | 0 | 9; 6.2% (2.9–11.5%) | 2; 0.4% (0.04–1.3%) | 0 | 0 | 0 |
| Richness | 2 | 3 | 7 | 1 | 4 | 3 | |
| A. hebraeum | R. appendiculatus | R. maculatus | R. simus | R. muehlensi | ||||
|---|---|---|---|---|---|---|---|---|
| Pathogen Species/Genotype | Adult (n = 12) | Nymph (n = 38) | Adult (n = 146) | Nymph (n = 554) | Nymph (n = 41) | Adult (n = 494) | Adult (n = 39) | Nymph (n = 74) |
| Babesia caballi | 0 | 0 | 1; 0.7% (0.02–3.8%) | 1; 0.2% (0.005–1.0%) | 0 | 2; 0.4% (0.05–1.5%) | 0 | 0 |
| Babesia sp. | 0 | 0 | 0 | 0 | 0 | 1; 0.2% (0.01–1.1%) | 0 | 0 |
| Hepatozoon sp. | 0 | 0 | 0 | 0 | 0 | 1; 0.2% (0.01–1.1%) | 1; 2.6% (0.1–13.5%) | 0 |
| Theileria equi | 0 | 0 | 0 | 8; 1.4% (0.6–2.8%) | 0 | 0 | 1; 2.6% (0.1–13.5%) | 0 |
| Theileria ovis | 0 | 0 | 0 | 7; 1.2% (0.5–2.6%) | 0 | 0 | 0 | 0 |
| Theiliera sp. Aepycerotini | 0 | 0 | 0 | 19; 3.4% (2.1–5.3%) | 6; 14.6% (5.6–29.2%) | 1; 0.2% (0.01–1.1%) | 1; 2.6% (0.1–13.5%) | 11; 14.9% (7.7–25.0%) |
| Theileria sp. giraffe | 0 | 0 | 2; 1.4% (0.2–4.9%) | 0 | 0 | 0 | 0 | 0 |
| Theileria sp. impala | 0 | 3; 7.9% (1.7–21.4%) | 0 | 0 | 0 | 0 | 0 | 0 |
| Theileria sp. rodent | 0 | 0 | 0 | 0 | 0 | 1; 0.2% (0.01–1.1%) | 0 | 0 |
| Theileria sp. sable | 0 | 0 | 7; 4.8% (1.9–9.6%) | 1; 0.2% (0.005–1.0%) | 0 | 2; 0.4% (0.05–1.5%) | 0 | 0 |
| Theileria sp. Tragelaphini B | 0 | 0 | 0 | 0 | 0 | 0 | 2; 5.1% (0.6–17.3%) | 0 |
| Theileria sp. Tragelaphini D | 0 | 0 | 8; 5.5% (2.4–10.5%) | 14; 2.5% (1.4–4.2%) | 1; 2.4% (0.06–12.9%) | 0 | 1; 2.6% (0.1–13.5%) | 9; 12.1% (5.7–21.8%) |
| Theileria sp. Tragelaphini E | 0 | 0 | 1; 0.7% (0.02–3.8%) | 2; 0.4% (0.04–1.3%) | 0 | 1; 0.2% (0.01–1.1%) | 3; 7.7% (1.6–20.9%) | 0 |
| Theileria sp. waterbuck | 0 | 2; 5.3% (0.6–17.7%) | 0 | 0 | 0 | 0 | 0 | 0 |
| Theileria taurotragi | 0 | 0 | 4; 2.7% (0.8–6.9%) | 0 | 0 | 0 | 2; 5.1% (0.6–17.3%) | 0 |
| Theileria velifera | 1; 8.3% (0.2–38.5%) | 4; 10.5% (2.9–24.8%) | 0 | 0 | 0 | 0 | 0 | 0 |
| Theileria sp. A | 0 | 0 | 0 | 0 | 1; 2.4% (0.06–12.9%) | 0 | 0 | 0 |
| Theileria sp. B | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2; 2.7% (0.3–9.4%) |
| Richness | 3 | 9 | 3 | 7 | 8 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ledger, K.J.; Beati, L.; Wisely, S.M. Survey of Ticks and Tick-Borne Rickettsial and Protozoan Pathogens in Eswatini. Pathogens 2021, 10, 1043. https://doi.org/10.3390/pathogens10081043
Ledger KJ, Beati L, Wisely SM. Survey of Ticks and Tick-Borne Rickettsial and Protozoan Pathogens in Eswatini. Pathogens. 2021; 10(8):1043. https://doi.org/10.3390/pathogens10081043
Chicago/Turabian StyleLedger, Kimberly J., Lorenza Beati, and Samantha M. Wisely. 2021. "Survey of Ticks and Tick-Borne Rickettsial and Protozoan Pathogens in Eswatini" Pathogens 10, no. 8: 1043. https://doi.org/10.3390/pathogens10081043
APA StyleLedger, K. J., Beati, L., & Wisely, S. M. (2021). Survey of Ticks and Tick-Borne Rickettsial and Protozoan Pathogens in Eswatini. Pathogens, 10(8), 1043. https://doi.org/10.3390/pathogens10081043







