High Frequency of Cryptosporidium hominis Infecting Infants Points to A Potential Anthroponotic Transmission in Maputo, Mozambique
Abstract
1. Introduction
2. Results
2.1. Characteristics of the Participants
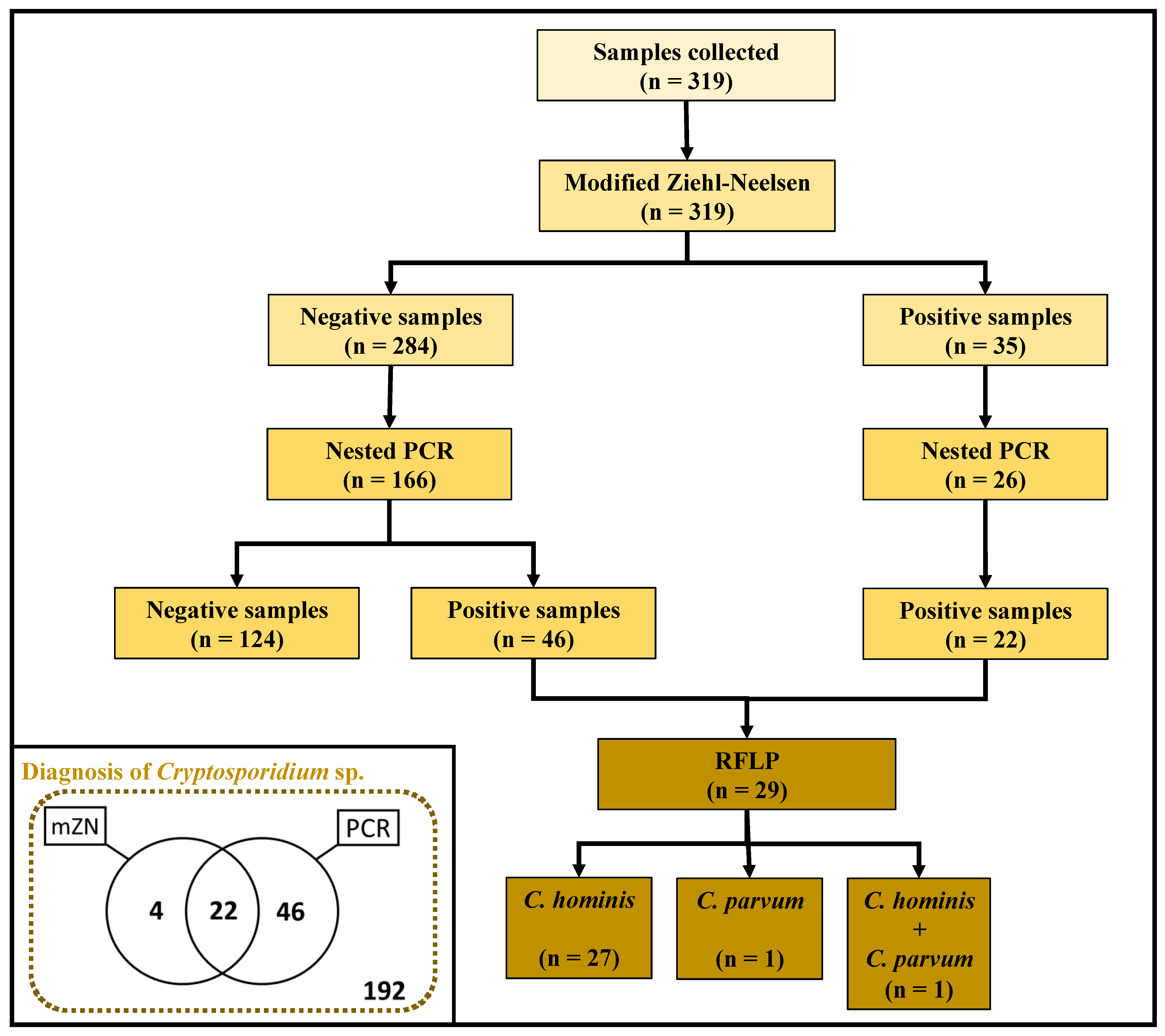
2.2. Frequency of Cryptosporidium spp. Infection
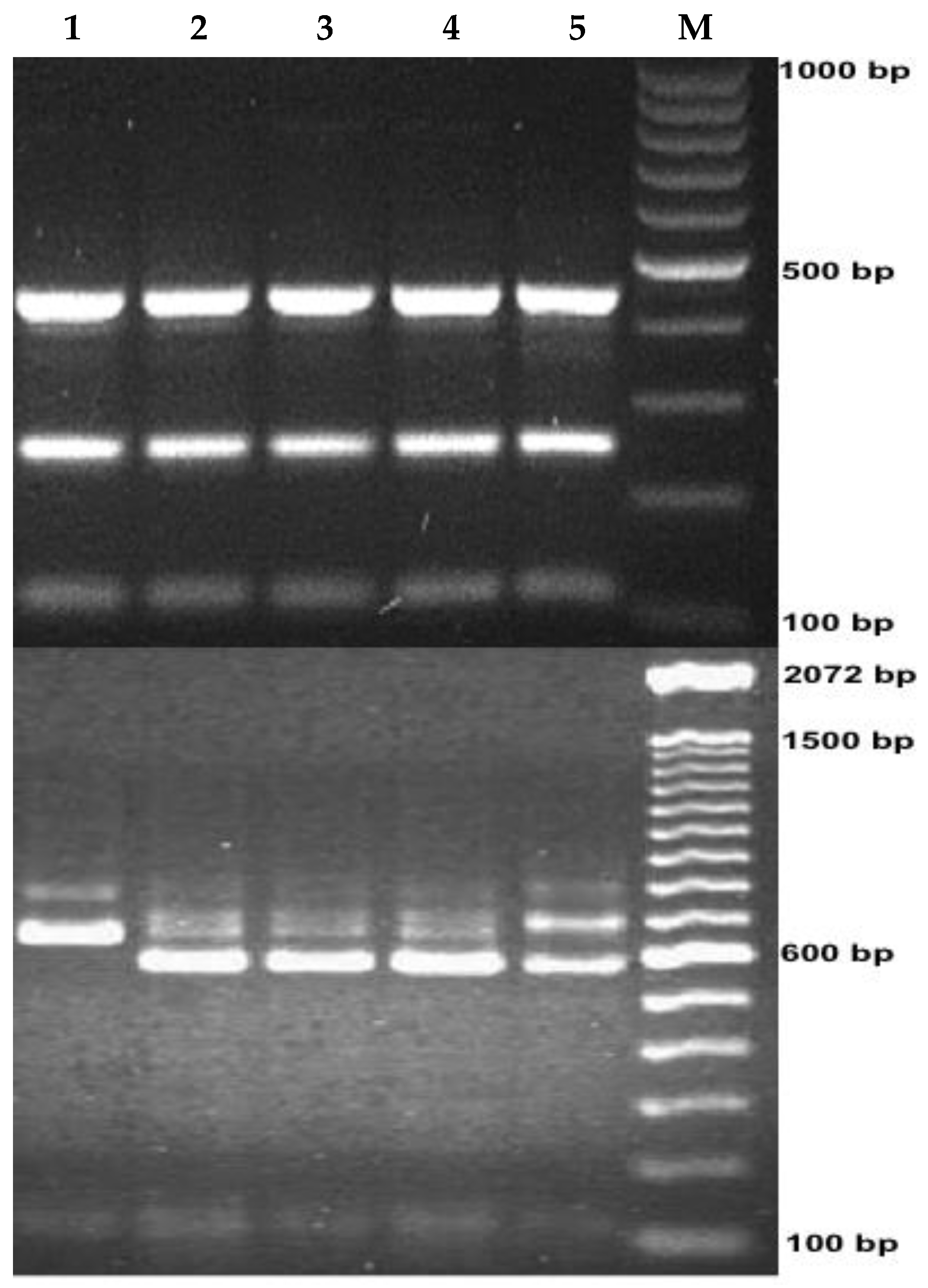
2.3. Molecular Characterization of Cryptosporidium Species
2.4. Risk Factors for Cryptosporidium spp. Infection
3. Discussion
4. Materials and Methods
4.1. Ethics Statements
4.2. Study Design, Site and Population
4.3. Sample Size Calculation
4.4. Data Collection
4.5. Sample Collection and Management
4.6. Laboratory Sample Processing
4.6.1. Direct Microscopy
4.6.2. DNA Extraction
4.6.3. Molecular Detection by Conventional Polymerase Chain Reaction (PCR)
4.6.4. Characterization of Cryptosporidium spp. Isolates by Nested PCR and Restriction Fragment Polymorphism (RFLP) Analysis
4.7. Data Management and Statistical Procedures
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kotloff, K.L.; Nataro, J.P.; Blackwelder, W.C.; Nasrin, D.; Farag, T.H.; Panchalingam, S.; Wu, Y.; Sow, S.O.; Sur, D.; Breiman, R.F.; et al. Burden and Aetiology of Diarrhoeal Disease in Infants and Young Children in Developing Countries (the Global Enteric Multicenter Study, GEMS): A Prospective, Case-Control Study. Lancet Lond. Engl. 2013, 382, 209–222. [Google Scholar] [CrossRef]
- GBD 2016 Causes of Death Collaborators. Global, Regional, and National Age-Sex Specific Mortality for 264 Causes of Death, 1980-2016: A Systematic Analysis for the Global Burden of Disease Study 2016. Lancet Lond. Engl. 2017, 390, 1151–1210. [Google Scholar] [CrossRef]
- Ministério da Saúde (MISAU); Instituto Nacional de Estatística (INE); ICF International (ICFI). Inquérito Demográfico e de Saúde (IDS); MISAU: Calverton, MD, USA, 2011.
- Rey, L. Bases Da Parasitologia Médica, 3rd ed.; Guanabara Koogan: Rio de Janeiro, Brazil, 2010. [Google Scholar]
- Chalmers, R.M.; Davies, A.P. Minireview: Clinical Cryptosporidiosis. Exp. Parasitol. 2010, 124, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Tzipori, S.; Ward, H. Cryptosporidiosis: Biology, Pathogenesis and Disease. Microbes Infect. 2002, 4, 1047–1058. [Google Scholar] [CrossRef]
- Widmer, G.; Carmena, D.; Kváč, M.; Chalmers, R.M.; Kissinger, J.C.; Xiao, L.; Sateriale, A.; Striepen, B.; Laurent, F.; Lacroix-Lamandé, S.; et al. Update on Cryptosporidium Spp.: Highlights from the Seventh International Giardia and Cryptosporidium Conference. Parasite Paris Fr. 2020, 27, 14. [Google Scholar] [CrossRef]
- Peletz, R.; Mahin, T.; Elliott, M.; Montgomery, M.; Clasen, T. Preventing Cryptosporidiosis: The Need for Safe Drinking Water. Bull. World Health Organ. 2013, 91, 238A. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Ryan, U.M.; Xiao, L. Genetic Diversity and Population Structure of Cryptosporidium. Trends Parasitol. 2018, 34, 997–1011. [Google Scholar] [CrossRef]
- Cunha, F.S.; Peralta, R.H.S.; Peralta, J.M. New Insights into the Detection and Molecular Characterization of Cryptosporidium with Emphasis in Brazilian Studies: A Review. Rev. Inst. Med. Trop. Sao Paulo 2019, 61, e28. [Google Scholar] [CrossRef]
- Xiao, L.; Morgan, U.M.; Limor, J.; Escalante, A.; Arrowood, M.; Shulaw, W.; Thompson, R.C.; Fayer, R.; Lal, A.A. Genetic Diversity within Cryptosporidium Parvum and Related Cryptosporidium Species. Appl. Environ. Microbiol. 1999, 65, 3386–3391. [Google Scholar] [CrossRef]
- Xiao, L.; Bern, C.; Limor, J.; Sulaiman, I.; Roberts, J.; Checkley, W.; Cabrera, L.; Gilman, R.H.; Lal, A.A. Identification of 5 Types of Cryptosporidium Parasites in Children in Lima, Peru. J. Infect. Dis. 2001, 183, 492–497. [Google Scholar] [CrossRef]
- Kotloff, K.L. The Burden and Etiology of Diarrheal Illness in Developing Countries. Pediatr. Clin. N. Am. 2017, 64, 799–814. [Google Scholar] [CrossRef] [PubMed]
- Sow, S.O.; Muhsen, K.; Nasrin, D.; Blackwelder, W.C.; Wu, Y.; Farag, T.H.; Panchalingam, S.; Sur, D.; Zaidi, A.K.M.; Faruque, A.S.G.; et al. The Burden of Cryptosporidium Diarrheal Disease among Children <24 Months of Age in Moderate/High Mortality Regions of Sub-Saharan Africa and South Asia, Utilizing Data from the Global Enteric Multicenter Study (GEMS). PLoS Negl. Trop. Dis. 2016, 10, e0004729. [Google Scholar] [CrossRef] [PubMed]
- Acácio, S.; Mandomando, I.; Nhampossa, T.; Quintó, L.; Vubil, D.; Sacoor, C.; Kotloff, K.; Farag, T.; Nasrin, D.; Macete, E.; et al. Risk Factors for Death among Children 0-59 Months of Age with Moderate-to-Severe Diarrhea in Manhiça District, Southern Mozambique. BMC Infect. Dis. 2019, 19, 322. [Google Scholar] [CrossRef]
- Nhampossa, T.; Mandomando, I.; Acacio, S.; Quintó, L.; Vubil, D.; Ruiz, J.; Nhalungo, D.; Sacoor, C.; Nhabanga, A.; Nhacolo, A.; et al. Diarrheal Disease in Rural Mozambique: Burden, Risk Factors and Etiology of Diarrheal Disease among Children Aged 0-59 Months Seeking Care at Health Facilities. PLoS ONE 2015, 10, e0119824. [Google Scholar] [CrossRef]
- Bauhofer, A.F.L.; Cossa-Moiane, I.; Marques, S.; Guimarães, E.L.; Munlela, B.; Anapakala, E.; Chilaúle, J.J.; Cassocera, M.; Langa, J.S.; Chissaque, A.; et al. Intestinal Protozoan Infections among Children 0-168 Months with Diarrhea in Mozambique: June 2014–January 2018. PLoS Negl. Trop. Dis. 2020, 14, e0008195. [Google Scholar] [CrossRef]
- Ferreira, F.S.; Pereira, F.d.L.M.; Martins, M.d.R.O. Intestinal Parasitic Infections in Children under Five in the Central Hospital of Nampula, Northern Mozambique. J. Infect. Dev. Ctries. 2020, 14, 532–539. [Google Scholar] [CrossRef] [PubMed]
- Cossa-Moiane, I.; Chilaúle, J.J.; Cossa Herminio, H.; Cassocera, M.; Guimarães, E.L.; de Deus, N. Parasitic Infections in Children Presenting with Acute Diarrhea in Mozambique: National Surveillance Data (2013–2015). Int. J. Infect. Dis. 2016, 45, 356. [Google Scholar] [CrossRef]
- Knee, J.; Sumner, T.; Adriano, Z.; Berendes, D.; de Bruijn, E.; Schmidt, W.-P.; Nalá, R.; Cumming, O.; Brown, J. Risk Factors for Childhood Enteric Infection in Urban Maputo, Mozambique: A Cross-Sectional Study. PLoS Negl. Trop. Dis. 2018, 12, e0006956. [Google Scholar] [CrossRef] [PubMed]
- Casmo, V.; Lebbad, M.; Maungate, S.; Lindh, J. Occurrence of Cryptosporidium Spp. and Cystoisospora Belli among Adult Patients with Diarrhoea in Maputo, Mozambique. Heliyon 2018, 4, e00769. [Google Scholar] [CrossRef]
- Muadica, A.S.; Balasegaram, S.; Beebeejaun, K.; Köster, P.C.; Bailo, B.; Hernández-de-Mingo, M.; Dashti, A.; Dacal, E.; Saugar, J.M.; Fuentes, I.; et al. Risk Associations for Intestinal Parasites in Symptomatic and Asymptomatic Schoolchildren in Central Mozambique. Clin. Microbiol. Infect. 2020. [Google Scholar] [CrossRef]
- Gatei, W.; Wamae, C.N.; Mbae, C.; Waruru, A.; Mulinge, E.; Waithera, T.; Gatika, S.M.; Kamwati, S.K.; Revathi, G.; Hart, C.A. Cryptosporidiosis: Prevalence, Genotype Analysis, and Symptoms Associated with Infections in Children in Kenya. Am. J. Trop. Med. Hyg. 2006, 75, 78–82. [Google Scholar] [CrossRef] [PubMed]
- Irisarri-Gutiérrez, M.J.; Mingo, M.H.; de Lucio, A.; Gil, H.; Morales, L.; Seguí, R.; Nacarapa, E.; Muñoz-Antolí, C.; Bornay-Llinares, F.J.; Esteban, J.G.; et al. Association between Enteric Protozoan Parasites and Gastrointestinal Illness among HIV- and Tuberculosis-Infected Individuals in the Chowke District, Southern Mozambique. Acta Trop. 2017, 170, 197–203. [Google Scholar] [CrossRef]
- Mbae, C.K.; Nokes, D.J.; Mulinge, E.; Nyambura, J.; Waruru, A.; Kariuki, S. Intestinal Parasitic Infections in Children Presenting with Diarrhoea in Outpatient and Inpatient Settings in an Informal Settlement of Nairobi, Kenya. BMC Infect. Dis. 2013, 13, 243. [Google Scholar] [CrossRef]
- Osman, M.; El Safadi, D.; Cian, A.; Benamrouz, S.; Nourrisson, C.; Poirier, P.; Pereira, B.; Razakandrainibe, R.; Pinon, A.; Lambert, C.; et al. Prevalence and Risk Factors for Intestinal Protozoan Infections with Cryptosporidium, Giardia, Blastocystis and Dientamoeba among Schoolchildren in Tripoli, Lebanon. PLoS Negl. Trop. Dis. 2016, 10, e0004496. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Breastfeeding Support: Close to Mothers. Available online: https.//www.afro.who.int/pt/news/breastfeeding-support-close-mothers (accessed on 26 November 2020).
- Tellevik, M.G.; Moyo, S.J.; Blomberg, B.; Hjøllo, T.; Maselle, S.Y.; Langeland, N.; Hanevik, K. Prevalence of Cryptosporidium Parvum/Hominis, Entamoeba Histolytica and Giardia Lamblia among Young Children with and without Diarrhea in Dar Es Salaam, Tanzania. PLoS Negl. Trop. Dis. 2015, 9, e0004125. [Google Scholar] [CrossRef] [PubMed]
- Checkley, W.; White, A.C.; Jaganath, D.; Arrowood, M.J.; Chalmers, R.M.; Chen, X.-M.; Fayer, R.; Griffiths, J.K.; Guerrant, R.L.; Hedstrom, L.; et al. A Review of the Global Burden, Novel Diagnostics, Therapeutics, and Vaccine Targets for Cryptosporidium. Lancet Infect. Dis. 2015, 15, 85–94. [Google Scholar] [CrossRef]
- Cacciò, S.M.; Pozio, E. Advances in the Epidemiology, Diagnosis and Treatment of Cryptosporidiosis. Expert Rev. Anti. Infect. Ther. 2006, 4, 429–443. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, A.M.; Fernandes, N.; Ferreira, F.S.; Gomes, J.; Centeno-Lima, S. Intestinal Parasites in Children Hospitalized at the Central Hospital in Maputo, Mozambique. J. Infect. Dev. Ctries. 2014, 8, 786–789. [Google Scholar] [CrossRef]
- Paulos, S.; Mateo, M.; de Lucio, A.; Hernández-de Mingo, M.; Bailo, B.; Saugar, J.M.; Cardona, G.A.; Fuentes, I.; Mateo, M.; Carmena, D. Evaluation of Five Commercial Methods for the Extraction and Purification of DNA from Human Faecal Samples for Downstream Molecular Detection of the Enteric Protozoan Parasites Cryptosporidium Spp., Giardia Duodenalis, and Entamoeba Spp. J. Microbiol. Methods 2016, 127, 68–73. [Google Scholar] [CrossRef]
- World Health Organizatio (WHO). Diarrhoeal Disease. Available online: http.//www.who.int/news-room/fact-sheets/detail/diarrhoeal-disease (accessed on 2 December 2020).
- OpenEpi-Toolkit Shell for Developing New Applications. Available online: https://www.openepi.com/SampleSize/SSPropor.htm (accessed on 10 May 2020).
- World Health Organizatio (WHO). Manual of BasicTecnhiques for Health Laboratory, 2nd ed.; World Health Organizatio: Geneva, Switzerland, 2003. [Google Scholar]
- Xiao, L.; Escalante, L.; Yang, C.; Sulaiman, I.; Escalante, A.A.; Montali, R.J.; Fayer, R.; Lal, A.A. Phylogenetic Analysis of Cryptosporidium Parasites Based on the Small-Subunit RRNA Gene Locus. Appl. Environ. Microbiol. 1999, 65, 1578–1583. [Google Scholar] [CrossRef]
| Characteristics | N = 319 | Frequency (%) |
|---|---|---|
| Provenience | ||
| HGM | 156 | 48.9 |
| HGJM | 163 | 51.1 |
| Sex | ||
| Female | 135 | 42.3 |
| Male | 184 | 57.7 |
| Age (in months), categorized | ||
| 0–6 | 63 | 19.7 |
| 7–12 | 130 | 40.8 |
| 13–18 | 81 | 25.4 |
| 19–24 | 30 | 9.4 |
| 25–60 | 15 | 4.7 |
| Animal contact | ||
| No | 195 | 61.1 |
| Yes | 124 | 38.9 |
| Caregiver literacy status | ||
| Illiterate | 133 | 41.7 |
| Literate | 185 | 58.0 |
| Unknown/missing | 1 | 0.3 |
| Child HIV status | ||
| Negative | 168 | 52.7 |
| Positive | 41 | 12.9 |
| Unknown/missing | 110 | 34.5 |
| Characteristics | N = 319 | n = 81 | % | Crude OR (95% CI) | Adjusted OR (95% CI) |
|---|---|---|---|---|---|
| Provenience | |||||
| HGM | 156 | 41 | 26.3 | NA | NA |
| HGJM | 163 | 40 | 24.5 | NA | NA |
| Sex | |||||
| Female | 135 | 31 | 23.0 | 1 | |
| Male | 184 | 50 | 27.2 | 1.252 (0.747–2.097) | |
| Age (in months), categorized | |||||
| 0–6 | 63 | 6 | 9.5 | 1 | 1 |
| 7–12 | 130 | 37 | 28.5 | 3.780 (1.501–9.517) ** | 3.604 (1.426–9.112) ** |
| 13–18 | 81 | 25 | 30.9 | 4.241 (1.617–11.124) ** | 4.170 (1.584–10.979) ** |
| 19–24 | 30 | 7 | 23.3 | 2.891 (0.877–9.533) | 2.503 (0.727–8.618) |
| 25–60 | 15 | 6 | 40.0 | 6.333 (1.671–23.999) ** | 5.861 (1.532–22.417) ** |
| Animal contact | |||||
| No | 195 | 53 | 27.2 | 1.280 (0.756–2.166) | |
| Yes | 124 | 28 | 22.6 | 1 | |
| Caregiver literacy status | |||||
| Illiterate | 133 | 42 | 31.6 | 1.785 (1.071–2.976) ** | 1.688 (1.001–2.845) * |
| Literate | 185 | 38 | 20.5 | 1 | 1 |
| Unknown/missing | 1 | ||||
| Child HIV status | |||||
| Negative | 168 | 40 | 23.8 | 1 | |
| Positive | 41 | 10 | 24.4 | 1.032 (0.466–2.289) | |
| Unknown/missing | 110 |
| Specie | Band Expected (in bp) | |
|---|---|---|
| SspI Digestion | AseI Digestion | |
| C. hominis | 11, 12,111, 254, 449 | 70, 102, 104, 561 |
| C. parvum | 11, 12, 108, 254, 449 | 102, 104, 628 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cossa-Moiane, I.; Cossa, H.; Bauhofer, A.F.L.; Chilaúle, J.; Guimarães, E.L.; Bero, D.M.; Cassocera, M.; Bambo, M.; Anapakala, E.; Chissaque, A.; et al. High Frequency of Cryptosporidium hominis Infecting Infants Points to A Potential Anthroponotic Transmission in Maputo, Mozambique. Pathogens 2021, 10, 293. https://doi.org/10.3390/pathogens10030293
Cossa-Moiane I, Cossa H, Bauhofer AFL, Chilaúle J, Guimarães EL, Bero DM, Cassocera M, Bambo M, Anapakala E, Chissaque A, et al. High Frequency of Cryptosporidium hominis Infecting Infants Points to A Potential Anthroponotic Transmission in Maputo, Mozambique. Pathogens. 2021; 10(3):293. https://doi.org/10.3390/pathogens10030293
Chicago/Turabian StyleCossa-Moiane, Idalécia, Hermínio Cossa, Adilson Fernando Loforte Bauhofer, Jorfélia Chilaúle, Esperança Lourenço Guimarães, Diocreciano Matias Bero, Marta Cassocera, Miguel Bambo, Elda Anapakala, Assucênio Chissaque, and et al. 2021. "High Frequency of Cryptosporidium hominis Infecting Infants Points to A Potential Anthroponotic Transmission in Maputo, Mozambique" Pathogens 10, no. 3: 293. https://doi.org/10.3390/pathogens10030293
APA StyleCossa-Moiane, I., Cossa, H., Bauhofer, A. F. L., Chilaúle, J., Guimarães, E. L., Bero, D. M., Cassocera, M., Bambo, M., Anapakala, E., Chissaque, A., Sambo, J., Langa, J. S., Manhique-Coutinho, L. V., Fantinatti, M., Lopes-Oliveira, L. A., Da-Cruz, A. M., & de Deus, N. (2021). High Frequency of Cryptosporidium hominis Infecting Infants Points to A Potential Anthroponotic Transmission in Maputo, Mozambique. Pathogens, 10(3), 293. https://doi.org/10.3390/pathogens10030293






