Cooccurrence of Five Pathogenic Legionella spp. and Two Free-Living Amoebae Species in a Complete Drinking Water System and Cooling Towers
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Site Location and Sampling
4.2. Chemical and Physical Analyses
4.3. Microbiological Analysis
4.4. Water Sample Processing and DNA Extraction
4.5. Molecular Analysis
4.6. Molecular Analysis of Acanthamoeba spp., N. fowleri, General Legionella spp., and Four Pathogenic Legionella Species
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Armand, B.; Motazedian, M.H.; Asgari, Q. Isolation and identification of pathogenic free-living amoeba from surface and tap water of Shiraz City using morphological and molecular methods. Parasitol. Res. 2016, 115, 63–68. [Google Scholar] [CrossRef]
- Lares-García, C.; Otero-Ruiz, A.; Gonzalez-Galaviz, J.R.; Ibarra-Gámez, J.C.; Lares-Villa, F. Potentially pathogenic genera of free-living amoebae coexisting in a thermal spring. Exp. Parasitol. 2018, 195, 54–58. [Google Scholar] [CrossRef]
- Baquero, R.A.; Reyes-Batlle, M.; Nicola, G.G.; Martín-Navarro, C.M.; López-Arencibia, A.; Guillermo Esteban, J.; Valladares, B.; Martínez-Carretero, E.; Piñero, J.E.; Lorenzo-Morales, J. Presence of potentially pathogenic free-living amoebae strains from well water samples in Guinea-Bissau. Pathog. Glob. Health 2014, 108, 206–211. [Google Scholar] [CrossRef]
- Delafont, V.; Brouke, A.; Bouchon, D.; Moulin, L.; Héchard, Y. Microbiome of free-living amoebae isolated from drinking water. Water Res. 2013, 47, 6958–6965. [Google Scholar] [CrossRef]
- Retana-Moreira, L.; Abrahams-Sandí, E.; Cabello-Vílchez, A.M.; Reyes-Batlle, M.; Valladares, B.; Martínez-Carretero, E.; Piñero, J.E.; Lorenzo-Morales, J. Isolation and molecular characterization of Acanthamoeba and Balamuthia mandrillaris from combination shower units in Costa Rica. Parasitol. Res. 2014, 113, 4117–4122. [Google Scholar] [CrossRef]
- Canals, O.; Serrano-Suárez, A.; Salvadó, H.; Méndez, J.; Cervero-Aragó, S.; Ruiz de Porras, V.; Dellundé, J.; Araujo, R. Effect of chlorine and temperature on free-living protozoa in operational man-made water systems (cooling towers and hot sanitary water systems) in Catalonia. Environ. Sci. Pollut. Res. Int. 2015, 22, 6610–6618. [Google Scholar] [CrossRef]
- Ren, K.; Xue, Y.; Rønn, R.; Liu, L.; Chen, H.; Rensing, C.; Yang, J. Dynamics and determinants of amoeba community, occurrence, and abundance in subtropical reservoirs and rivers. Water Res. 2018, 146, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Shaheen, M.; Scott, C.; Ashbolt, N.J. Long-term persistence of infectious Legionella with free-living amoebae in drinking water biofilms. Int. J. Hyg. Environ. Health 2019, 222, 678–686. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.S.; Vaccaro, L.; Magnet, A.; Izquierdo, F.; Ollero, D.; Martínez-Fernández, C.; Mayo, L.; Moran, M.; Pozuelo, M.J.; Fenoy, S.; et al. Presence and interaction of free-living amoebae and amoeba-resisting bacteria in water from drinking water treatment plants. Sci. Total. Environ. 2020, 719, 137080. [Google Scholar] [CrossRef] [PubMed]
- Michel, R.; Hauröder-Philippczyk, B.; Müller, K.-D.; Weishaar, I. Acanthamoeba from human nasal mucosa infected with an obligate intracellular parasite. Eur. J. Protistol. 1994, 30, 104–110. [Google Scholar] [CrossRef]
- Rowbotham, T.J. Preliminary report on the pathogenicity of Legionella pneumophila for freshwater and soil amoebae. J. Clin. Pathol. 1980, 33, 1179–1183. [Google Scholar] [CrossRef]
- Liu, Z.; Lin, Y.E.; Stout, J.E.; Hwang, C.C.; Vidic, R.D.; Yu, V.L. Effect of flow regimes on the presence of Legionella within the biofilm of a model plumbing system. J. Appl. Microbiol. 2006, 101, 437–442. [Google Scholar] [CrossRef] [PubMed]
- Thomas, V.; Herrera-Rimann, K.; Blanc, D.S.; Greub, G. Biodiversity of amoebae and amoeba-resisting bacteria in a hospital water network. Appl. Environ. Microbiol. 2006, 72, 2428–2438. [Google Scholar] [CrossRef] [PubMed]
- Thomas, V.; Loret, J.-F.; Jousset, M.; Greub, G. Biodiversity of amoebae and amoebae-resisting bacteria in a drinking water treatment plant. Environ. Microbiol. 2008, 10, 2728–2745. [Google Scholar] [CrossRef]
- Rohr, U.; Weber, S.; Michel, R.; Selenka, F.; Wilhelm, M. Comparison of Free-Living Amoebae in Hot Water Systems of Hospitals with Isolates from Moist Sanitary Areas by Identifying Genera and Determining Temperature Tolerance. Appl. Environ. Microbiol. 1998, 64, 1822–1824. [Google Scholar] [CrossRef] [PubMed]
- Buse, H.Y.; Lu, J.; Struewing, I.T.; Ashbolt, N.J. Eukaryotic diversity in premise drinking water using 18S rDNA sequencing: Implications for health risks. Environ. Sci. Pollut. Res. 2013, 20, 6351–6366. [Google Scholar] [CrossRef]
- Kilvington, S.; Price, J. Survival of Legionella pneumophila within cysts of Acanthamoeba polyphaga following chlorine exposure. J. Appl. Bacteriol. 1990, 68, 519–525. [Google Scholar] [CrossRef] [PubMed]
- De Giglio, O.; Napoli, C.; Apollonio, F.; Brigida, S.; Marzella, A.; Diella, G.; Calia, C.; Scrascia, M.; Pacifico, C.; Pazzani, C.; et al. Occurrence of Legionella in groundwater used for sprinkler irrigation in Southern Italy. Environ. Res. 2019, 170, 215–221. [Google Scholar] [CrossRef]
- Shen, S.-M.; Chou, M.-Y.; Hsu, B.-M.; Ji, W.-T.; Hsu, T.-K.; Tsai, H.-F.; Huang, Y.-L.; Chiu, Y.-C.; Kao, E.-S.; Kao, P.-M.; et al. Assessment of Legionella pneumophila in recreational spring water with quantitative PCR (Taqman) assay. Pathog. Glob. Health 2015, 109, 236–241. [Google Scholar] [CrossRef]
- Ishizaki, N.; Sogawa, K.; Inoue, H.; Agata, K.; Edagawa, A.; Miyamoto, H.; Fukuyama, M.; Furuhata, K. Legionella thermalis sp. Nov., isolated from hot spring water in Tokyo, Japan. Microbiol. Immunol. 2016, 60, 203–208. [Google Scholar] [CrossRef]
- Leoni, E.; Catalani, F.; Marini, S.; Dallolio, L. Legionellosis Associated with Recreational Waters: A Systematic Review of Cases and Outbreaks in Swimming Pools, Spa Pools, and Similar Environments. Int. J. Environ. Res. Public Health 2018, 15, 1612. [Google Scholar] [CrossRef] [PubMed]
- Dey, R.; Bodennec, J.; Mameri, M.O.; Pernin, P. Free-living freshwater amoebae differ in their susceptibility to the pathogenic bacterium Legionella pneumophila. FEMS Microbiol. Lett. 2009, 290, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Greub, G.; Raoult, D. Microorganisms Resistant to Free-Living Amoebae. Clin. Microbiol. Rev. 2004, 17, 413–433. [Google Scholar] [CrossRef]
- McDade, J.E.; Shepard, C.C.; Fraser, D.W.; Tsai, T.R.; Redus, M.A.; Dowdle, W.R. Legionnaires’ disease: Isolation of a bacterium and demonstration of its role in other respiratory disease. N. Engl. J. Med. 1977, 297, 1197–1203. [Google Scholar] [CrossRef]
- Kaufmann, A.F.; McDade, J.E.; Patton, C.M.; Bennett, J.V.; Skaliy, P.; Feeley, J.C.; Anderson, D.C.; Potter, M.E.; Newhouse, V.F.; Gregg, M.B.; et al. Pontiac fever: Isolation of the etiologic agent (Legionella pneumophilia) and demonstration of its mode of transmission. Am. J. Epidemiol. 1981, 114, 337–347. [Google Scholar] [CrossRef]
- Abu Kwaik, Y.; Gao, L.-Y.; Stone, B.J.; Venkataraman, C.; Harb, O.S. Invasion of Protozoa by Legionella pneumophila and Its Role in Bacterial Ecology and Pathogenesis. Appl. Environ. Microbiol. 1998, 64, 3127–3133. [Google Scholar] [CrossRef]
- Albert-Weissenberger, C.; Cazalet, C.; Buchrieser, C. Legionella pneumophila—A human pathogen that co-evolved with fresh water protozoa. Cell. Mol. Life Sci. 2006, 64, 432. [Google Scholar] [CrossRef] [PubMed]
- Cirillo, J.D.; Falkow, S.; Tompkins, L.S. Growth of Legionella pneumophila in Acanthamoeba castellanii enhances invasion. Infect. Immun. 1994, 62, 3254–3261. [Google Scholar] [CrossRef] [PubMed]
- Nora, T.; Lomma, M.; Gomez-Valero, L.; Buchrieser, C. Molecular mimicry: An important virulence strategy employed by Legionella pneumophila to subvert host functions. Future Microbiol. 2009, 4, 691–701. [Google Scholar] [CrossRef]
- Dupuy, M.; Mazoua, S.; Berne, F.; Bodet, C.; Garrec, N.; Herbelin, P.; Ménard-Szczebara, F.; Oberti, S.; Rodier, M.-H.; Soreau, S.; et al. Efficiency of water disinfectants against Legionella pneumophila and Acanthamoeba. Water Res. 2011, 45, 1087–1094. [Google Scholar] [CrossRef]
- Hwang, M.G.; Katayama, H.; Ohgaki, S. Effect of intracellular resuscitation of Legionella pneumophila in Acanthamoeba polyphage cells on the antimicrobial properties of silver and copper. Environ. Sci. Technol. 2006, 40, 7434–7439. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.M.N.; Ilef, D.; Jarraud, S.; Rouil, L.; Campese, C.; Che, D.; Haeghebaert, S.; Ganiayre, F.; Marcel, F.; Etienne, J.; et al. A community-wide outbreak of legionnaires disease linked to industrial cooling towers—How far can contaminated aerosols spread? J. Infect. Dis. 2006, 193, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Buse, H.Y.; Schoen, M.E.; Ashbolt, N.J. Legionellae in engineered systems and use of quantitative microbial risk assessment to predict exposure. Water Res. 2012, 46, 921–933. [Google Scholar] [CrossRef]
- Ferré, M.R.S.; Arias, C.; Oliva, J.M.; Pedrol, A.; García, M.; Pellicer, T.; Roura, P.; Domínguez, A. A community outbreak of Legionnaires’ disease associated with a cooling tower in Vic and Gurb, Catalonia (Spain) in 2005. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2009, 28, 153–159. [Google Scholar] [CrossRef]
- Türetgen, I.; Sungur, E.I.; Cotuk, A. Enumeration of Legionella pneumophila in cooling tower water systems. Environ. Monit. Assess. 2005, 100, 53. [Google Scholar] [CrossRef]
- Bollin, G.E.; Plouffe, J.F.; Para, M.F.; Hackman, B. Aerosols containing Legionella pneumophila generated by shower heads and hot-water faucets. Appl. Environ. Microbiol. 1985, 50, 1128–1131. [Google Scholar] [CrossRef]
- Borella, P.; Montagna, M.T.; Romano-Spica, V.; Stampi, S.; Stancanelli, G.; Triassi, M.; Neglia, R.; Marchesi, I.; Fantuzzi, G.; Tatò, D.; et al. Legionella Infection Risk from Domestic Hot Water. Emerg. Infect. Dis. 2004, 10, 457–464. [Google Scholar] [CrossRef]
- Fraser, D.W.; Tsai, T.R.; Orenstein, W.; Parkin, W.E.; Beecham, H.J.; Sharrar, R.G.; Harris, J.; Mallison, G.F.; Martin, S.M.; McDade, J.E.; et al. Legionnaires’ disease: Description of an epidemic of pneumonia. N. Engl. J. Med. 1977, 297, 1189–1197. [Google Scholar] [CrossRef]
- Visvesvara, G.S.; Moura, H.; Schuster, F.L. Pathogenic and opportunistic free-living amoebae: Acanthamoeba spp., Balamuthia mandrillaris, Naegleria fowleri, and Sappinia diploidea. FEMS Immunol. Med. Microbiol. 2007, 50, 1–26. [Google Scholar] [CrossRef]
- Yu, V.L.; Plouffe, J.F.; Pastoris, M.C.; Stout, J.E.; Schousboe, M.; Widmer, A.; Summersgill, J.; File, T.; Heath, C.M.; Paterson, D.L.; et al. Distribution of Legionella Species and Serogroups Isolated by Culture in Patients with Sporadic Community-Acquired Legionellosis: An International Collaborative Survey. J. Infect. Dis. 2002, 186, 127–128. [Google Scholar] [CrossRef] [PubMed]
- Visvesvara, G. Free-Living Amebae as Opportunistic Agents of Human Disease. J. Neuroparasitol. 2010, 1, 1689–1693. [Google Scholar] [CrossRef]
- Booton, G.C.; Visvesvara, G.S.; Byers, T.J.; Kelly, D.J.; Fuerst, P.A. Identification and Distribution of Acanthamoeba Species Genotypes Associated with Nonkeratitis Infections. J. Clin. Microbiol. 2005, 43, 1689–1693. [Google Scholar] [CrossRef] [PubMed]
- Valciņa, O.; Pūle, D.; Mališevs, A.; Trofimova, J.; Makarova, S.; Konvisers, G.; Bērziņš, A.; Krūmiņa, A. Cooccurrence of Free-Living Amoeba and Legionella in Drinking Water Supply Systems. Medicina 2019, 55, 492. [Google Scholar] [CrossRef]
- Muchesa, P.; Leifels, M.; Jurzik, L.; Barnard, T.G.; Bartie, C. Detection of amoeba-associated Legionella pneumophila in hospital water networks of Johannesburg. S. Afr. J. Infect. Dis. 2018, 33, 72–75. [Google Scholar] [CrossRef]
- Scheikl, U.; Sommer, R.; Kirschner, A.; Rameder, A.; Schrammel, B.; Zweimüller, I.; Wesner, W.; Hinker, M.; Walochnik, J. Free-living amoebae (FLA) co-occurring with Legionellae in industrial waters. Eur. J. Protistol. 2014, 50, 422–429. [Google Scholar] [CrossRef]
- Scheikl, U.; Tsao, H.-F.; Horn, M.; Indra, A.; Walochnik, J. Free-living amoebae and their associated bacteria in Austrian cooling towers: A 1-year routine screening. Parasitol. Res. 2016, 115, 3365–3374. [Google Scholar] [CrossRef]
- Edagawa, A.; Kimura, A.; Doi, H.; Tanaka, H.; Tomioka, K.; Sakabe, K.; Nakajima, C.; Suzuki, Y. Detection of culturable and nonculturable Legionella species from hot water systems of public buildings in Japan. J. Appl. Microbiol. 2008, 105, 2104–2114. [Google Scholar] [CrossRef]
- Barrette, I. Comparison of Legiolert and a Conventional Culture Method for Detection of Legionella pneumophila from Cooling Towers in Québec. J. AOAC Int. 2019, 102, 1235–1240. [Google Scholar] [CrossRef] [PubMed]
- Steinert, M.; Emödy, L.; Amann, R.; Hacker, J. Resuscitation of viable but nonculturable Legionella pneumophila Philadelphia JR32 by Acanthamoeba castellanii. Appl. Environ. Microbiol. 1997, 63, 2047–2053. [Google Scholar] [CrossRef]
- Whiley, H.; Taylor, M. Legionella detection by culture and qPCR: Comparing apples and oranges. Crit. Rev. Microbiol. 2016, 42, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Young, C.; Smith, D.; Wafer, T.; Crook, B. Rapid Testing and Interventions to Control Legionella Proliferation Following a Legionnaires’ Disease Outbreak Associated with Cooling Towers. Microorganisms 2021, 9, 615. [Google Scholar] [CrossRef]
- Logan-Jackson, A.R.; Flood, M.; Rose, J.B. Enumeration and characterization of five pathogenic Legionella species from large research and educational buildings. Environ. Sci. Water Res. Technol. 2020. [Google Scholar] [CrossRef]
- Falzone, L.; Gattuso, G.; Lombardo, C.; Lupo, G.; Grillo, C.M.; Spandidos, D.A.; Libra, M.; Salmeri, M. Droplet digital PCR for the detection and monitoring of Legionella pneumophila. Int. J. Mol. Med. 2020, 46, 1777–1782. [Google Scholar] [CrossRef]
- Schneiders, S.; Hechard, T.; Edgren, T.; Avican, K.; Fällman, M.; Fahlgren, A.; Wang, H. Spatiotemporal Variations in Growth Rate and Virulence Plasmid Copy Number during Yersinia Pseudotuberculosis Infection. Infect. Immun. 2021, 89, e00710-20. [Google Scholar] [CrossRef]
- Li, H.; Bai, R.; Zhao, Z.; Tao, L.; Ma, M.; Ji, Z.; Jian, M.; Ding, Z.; Dai, X.; Bao, F.; et al. Application of droplet digital PCR to detect the pathogens of infectious diseases. Biosci. Rep. 2018, 38, BSR20181170. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Benson, R.; Pelish, T.; Brown, E.; Winchell, J.M.; Fields, B. Dual detection of Legionella pneumophila and Legionella species by real-time PCR targeting the 23S-5S rRNA gene spacer region. Clin. Microbiol. Infect. 2010, 16, 255–261. [Google Scholar] [CrossRef]
- Abdul Majid, M.A.; Mahboob, T.; Mong, B.G.; Jaturas, N.; Richard, R.L.; Tian-Chve, T.; Phimphila, A.; Mahaphonh, P.; Aye, K.N.; Aung, W.L.; et al. Pathogenic waterborne free-living amoebae: An update from selected Southeast Asian countries. PLoS ONE 2017, 12, e0169448. [Google Scholar] [CrossRef]
- Gabriel, S.; Khan, N.A.; Siddiqui, R. Occurrence of free-living amoebae (Acanthamoeba, Balamuthia, Naegleria) in water samples in Peninsular Malaysia. J. Water Health 2019, 17, 160–171. [Google Scholar] [CrossRef]
- Ambrose, M.; Kralovic, S.M.; Roselle, G.A.; Kowalskyj, O.; Rizzo, V.J.; Wainwright, D.L.; Gamage, S.D. Implementation of Legionella Prevention Policy in Health Care Facilities: The United States Veterans Health Administration Experience. J. Public Health Manag. Pract. 2020, 26, E1. [Google Scholar] [CrossRef] [PubMed]
- Donohue, M.J.; King, D.; Pfaller, S.; Mistry, J.H. The sporadic nature of Legionella pneumophila, Legionella pneumophila Sg1 and Mycobacterium avium occurrence within residences and office buildings across 36 states in the United States. J. Appl. Microbiol. 2019, 126, 1568–1579. [Google Scholar] [CrossRef]
- Kozak, N.A.; Lucas, C.E.; Winchell, J.M. Identification of Legionella in the environment. Methods Mol. Biol. 2013, 954, 3–25. [Google Scholar] [CrossRef] [PubMed]
- Breiman, R.F. Impact of Technology on the Emergence of Infectious Diseases. Epidemiol. Rev. 1996, 18, 4–9. [Google Scholar] [CrossRef]
- Marciano-Cabral, F.; Jamerson, M.; Kaneshiro, E.S. Free-living amoebae, Legionella and Mycobacterium in tap water supplied by a municipal drinking water utility in the USA. J. Water Health 2010, 8, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Masters, S.; Hong, Y.; Stallings, J.; Falkinham, J.O.; Edwards, M.A.; Pruden, A. Effect of disinfectant, water age, and pipe material on occurrence and persistence of Legionella, mycobacteria, Pseudomonas aeruginosa, and two amoebas. Environ. Sci. Technol. 2012, 46, 11566–11574. [Google Scholar] [CrossRef] [PubMed]
- Rivière, D.; Szczebara, F.M.; Berjeaud, J.-M.; Frère, J.; Héchard, Y. Development of a real-time PCR assay for quantification of Acanthamoeba trophozoites and cysts. J. Microbiol. Methods 2006, 64, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Pélandakis, M.; Serre, S.; Pernin, P. Analysis of the 5.8S rRNA gene and the internal transcribed spacers in Naegleria spp. and in N. fowleri. J. Eukaryot. Microbiol. 2000, 47, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Grimm, D.; Ludwig, W.; Brandt, B.; Michel, R.; Schleifer, K.-H.; Hacker, J.; Steinert, M. Development of 18S rRNA-Targeted Oligonucleotide Probes for Specific Detection of Hartmannella and Naegleria in Legionella—Positive Environmental Samples. Syst. Appl. Microbiol. 2001, 24, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Nazarian, E.J.; Bopp, D.J.; Saylors, A.; Limberger, R.J.; Musser, K.A. Design and implementation of a protocol for the detection of Legionella in clinical and environmental samples. Diagn. Microbiol. Infect. Dis. 2008, 62, 125–132. [Google Scholar] [CrossRef]
- Cross, K.E.; Mercante, J.W.; Benitez, A.J.; Brown, E.W.; Diaz, M.H.; Winchell, J.M. Simultaneous detection of Legionella species and L. anisa, L. bozemanii, L. longbeachae and L. micdadei using conserved primers and multiple probes in a multiplex real-time PCR assay. Diagn. Microbiol. Infect. Dis. 2016, 85, 295–301. [Google Scholar] [CrossRef][Green Version]
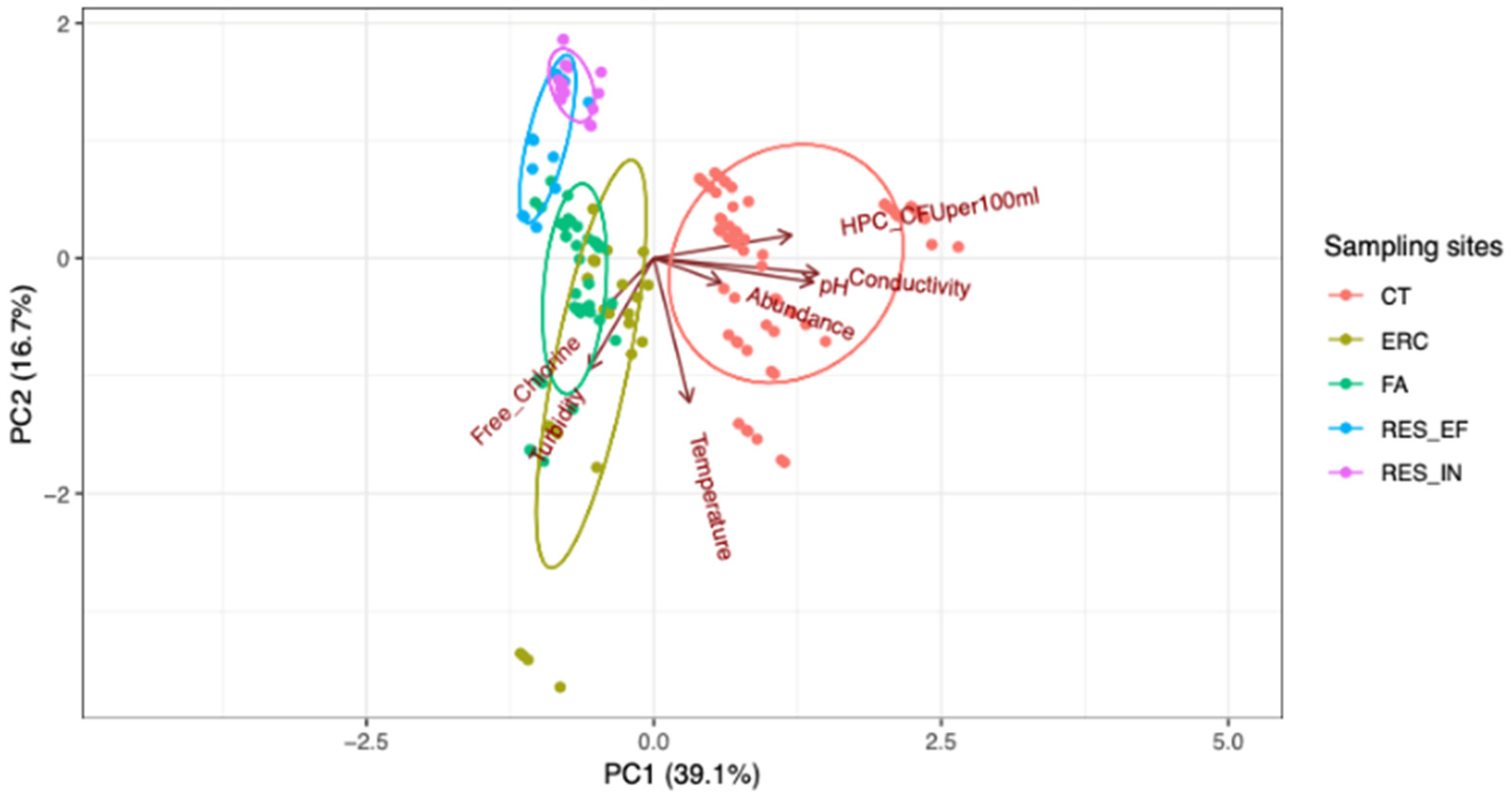
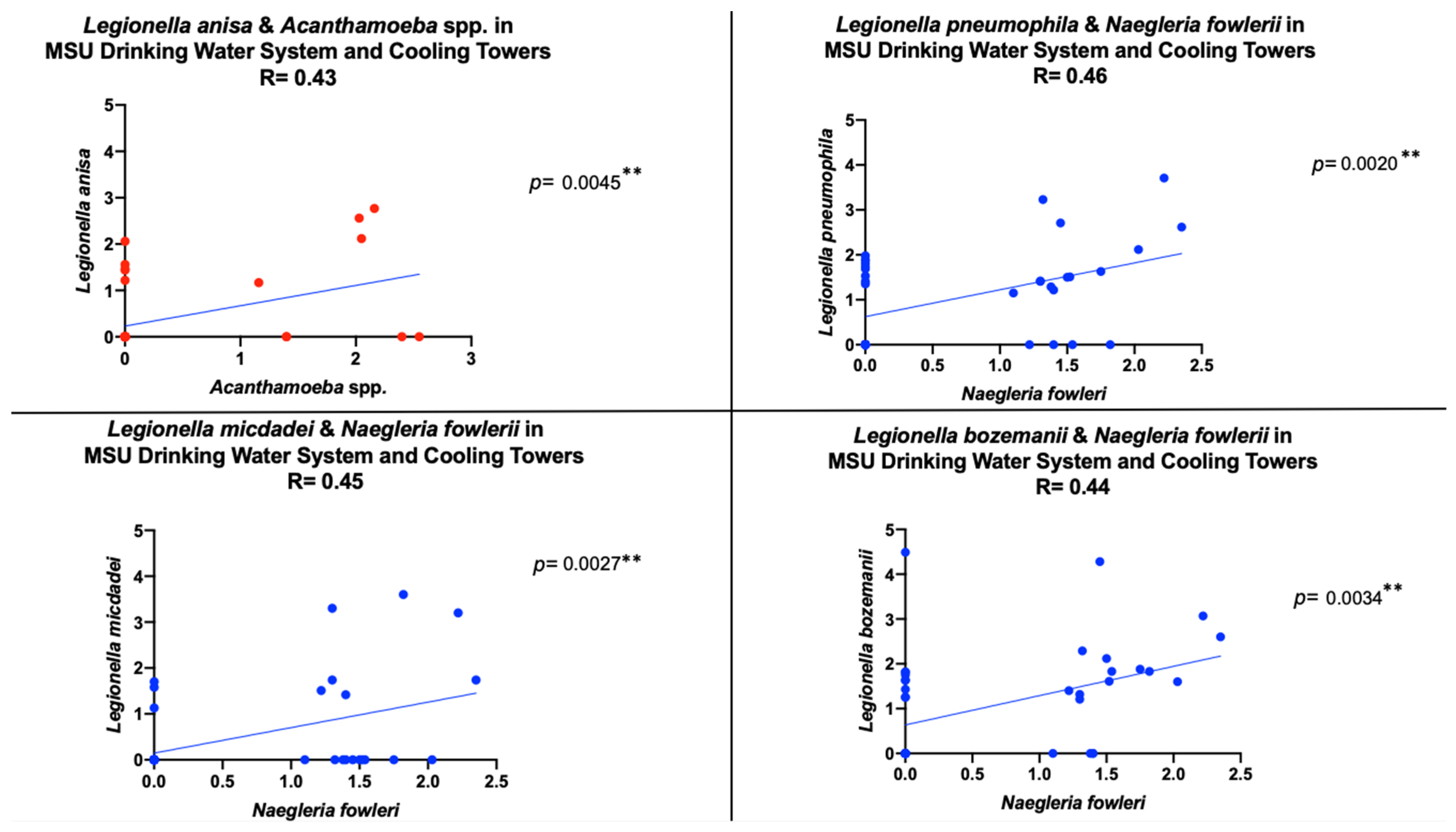
| Free-Living Amoebae | Site Location | ||||
|---|---|---|---|---|---|
| Res_In | Res_EF | Fa | ERC | CT | |
| Acanthamoeba spp. (%) | 0/6 (0%) | 0/6 (0%) | 3/15 (20%) | 0/9 (0%) | 5/6 (83%) |
| Acanthamoeba spp. Min, and Max Geomean (Log10 GC/100 mL) | ND | ND | 1.1, 1.4 | ND | 2.0, 2.5 |
| N.fowleri (%) | 5/6 (83%) | 1/6 (16%) | 0/15 (0%) | 6/9 (66%) | 5/6 (83%) |
| N.fowleri Min, and Max Geomean (Log10 GC/100 mL) | 1.3, 1.7 | 1.5 a | ND | 1.1, 1.8 | 1.3, 2.4 |
| Legionella Species | |||||
| Legionella spp. (23S rRNA) (%) | (6/6) (100%) | (6/6) (100%) | 15/15 (100%) | 9/9 (100%) | 6/6 (100%) |
| Legionella spp. (23S rRNA) Geomean (Log10 GC/100 mL) | 3.1 | 2.7 | 2.3 | 4.3 | 4.5 |
| L. pneumophila (%) | 5/6 (83%) | 5/6 (83%) | 6/15 (40%) | 3/9 (33%) | 5/6 (83%) |
| L. pneumophila Geomean (Log10 GC/100 mL) | 1.6 | 1.8 | 1.6 | 1.4 | 2.8 |
| L. micdadei (%) | 1/6 (17%) | 0/6 (0%) | 3/15 (20%) | 4/9 (44%) | 2/6 (33%) |
| L. micdadei Geomean (Log10 GC/100 mL) | 1.5 | ND | 1.5 | 1.6 | 2.4 |
| L. bozemanii (%) | 4/6 (67%) | (6/6) (100%) | 2/15 (13%) | 4/9 (44%) | 6/6 (100%) |
| L. bozemanii Geomean (Log10 GC/100 mL) | 1.5 | 1.7 | 1.4 | 1.5 | 2.9 |
| L. longbeachae (%) | 0/6 (0%) | 0/6 (0%) | 8/15 (53%) | 4/9 (44%) | 3/6 (50%) |
| L. longbeachae Geomean (Log10 GC/100 mL) | ND | ND | 1.4 | 1.4 | 1.5 |
| L. anisa (%) | 0/6 (0%) | 0/6 (0%) | 5/15 (33%) | 0/6 (0%) | 4/6 (67%) |
| L. anisa Geomean (Log10 GC/100 mL) | ND | ND | 1.4 | ND | 2.1 |
| Temperature (°C) | Total Chlorine (mg/L) | Free Chlorine (mg/L) | Turbidity NTU | pH | Conductivity (mS) | HPC (CFU/100 mL) | Water Age (h) |
|---|---|---|---|---|---|---|---|
| RES_IN (N = 6) | |||||||
| 12.1 | 0 | 0 | 4.1 | 7.2 | 851 | 3.52 × 101 | 4.5 |
| RES_EF (N = 6) | |||||||
| 11.9 | 0.64 | 0.33 | 3.85 | 7.2 | 855 | 2.10 × 100 | 3.4 |
| Building F Influent (N = 3) | |||||||
| 26.8 | 0.41 | 0.35 | 8.4 | 7.3 | 897 | 8.57 × 104 | 9.2 |
| Building Fa 1st Floor Cold; N = 3 (Hot Taps; N = 3) | |||||||
| 26.7 (28.6) | 0.16 (0.04) | 0.14 (0.02) | 3.06 (0.53) | 7.2 (7.1) | 867 (815) | 1.02 × 104 (7.3 × 103) | – |
| Building Fa 2nd Floor Cold; N = 3 (Hot Taps; N = 3) | |||||||
| 26.8 (28.8) | 0.05 (0.02) | 0.03 (0) | 3.37 (0.67) | 7.0 (6.9) | 856 (822) | 2.00 × 104 (3.15 × 103) | – |
| Building ERC Influent (N = 3) | |||||||
| 31.5 | 0.31 | 0.20 | 12.5 | 7.4 | 883 | 4.32 × 105 | 20.8 |
| Building ERC 1st Floor Cold (N = 3) (Hot Taps; N = 3) | |||||||
| 23.5 (24.5) | 0.09 (0.04) | 0.03 (0) | 5.97 (6.27) | 7.6 (7.5) | 866 (847) | 4.38 × 105 (6.80 × 105) | – |
| Cooling Towers (N = 6) | |||||||
| 25.3 | 0.49 | 0.08 | 1.94 | 8.2 | 2564 | 2.35 × 107 | 23.2 |
| ap Value = 0.0286 * | L. anisa Present | L. anisa Absent | Total |
|---|---|---|---|
| Acanthamoeba spp. Present | 4 (9.52%) | 4 (9.52%) | 8 |
| Acanthamoeba spp. Absent | 5 (11.90%) | 29 (69.05%) | 34 |
| Total | 9 (21.42%) | 33 (78.57%) | 42 |
| ap Value = 0.0293 * | L. micdadei Present | L. micdadei Absent | Total |
|---|---|---|---|
| N.fowleri Present | 7 (16.67%) | 10 (23.81%) | 17 |
| N.fowleri Absent | 3 (7.14%) | 22 (52.38%) | 25 |
| Total | 10 (23.8%) | 32 (76.19%) | 42 |
| ap Value = 0.0099 ** | L. bozemanii Present | L. bozemanii Absent | Total |
|---|---|---|---|
| N.fowleri Present | 13 (30.95%) | 4 (9.52%) | 17 |
| N.fowleri Absent | 9 (21.43%) | 16 (38.10%) | 25 |
| Total | 22 (52.38%) | 20 (47.61%) | 42 |
| ap Value = 0.0369 * | L. pneumophila Present | L. pneumophila Absent | Total |
|---|---|---|---|
| N.fowleri Present | 13 (30.95%) | 4 (9.52%) | 17 |
| N.fowleri Absent | 11 (26.19%) | 14 (33.33%) | 25 |
| Total | 24 (57.14%) | 18 (42.85%) | 42 |
| Target Species | Primer/Probe Name | Primer/Probe Sequence | Amplicon Length (bp) | Reference |
|---|---|---|---|---|
| Acanthamoeba spp. | 18S rRNAF 18S rRNAR 18S rRNAP | 5′-CGACCAGCGATTAGGAGACG-3′ 5′-CCGACGCCAAGGACGAC-3′ 5′-FAM-TGAATACAAAACACCACCATCGGCGC-BHQ1-3′ | 63 | [65] |
| N. fowleri | ITSF ITSR ITSP | 5′-GTGAAAACCTTTTTTCCATTTACA-3′ 5′-AAATAAAAGATTGACCATTTGAAA-3′ 5′-HEX-GTGGCCCACGACAGCTTT-BHQ1-3′ | 69 | [66,67] |
| Legionella species | 23SF 23SR 23SP | 5′-CCCATGAAGCCCGTTGAA-3′ 5′-ACAATCAGCCAATTAGTACGAG TTAGC-3′ 5′-HEX-TCCACACCTCGCCTATCAACGTCGTAGT-BHQ1-3′ | 92 | [68] |
| L. pneumophila | mipF mipR mipP | 5′-AAAGGCATGCAAGACGCTATG-3′ 5′-GAAACTTGTTAAGAACGTCTTTCATTTG-3′ 5′-FAM-TGGCGCTCAATTGGCTTTAACCGA-BHQ1-3′ | 78 | [68] |
| L. micdadei L. anisa L. bozemanii L. longbeachae | Pan-Legionella F Pan-Legionella R LmicdadeiP LanisaP Lbozemanii LlongbeachaeP | 5′-GTACTAATTGGCTGATTGTCTTG-3′ 5′-TTCACTTCTGAGTTCGAGATGG-3′ 5′-FAM-AGCTGATTGGTTAATAGCCCAATCGG-BHQ1-3′ 5′-HEX-CTCAACCTACGCAGAACTACTTGAGG-BHQ1-3′ 5′-FAM-TACGCCCATTCATCATGCAAACCAGnT-BHQ1-3′ 5′-HEX-CTGAGTATCATGCCAATAATGCGCGC-BHQ1-3′ | Not available | [69] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Logan-Jackson, A.; Rose, J.B. Cooccurrence of Five Pathogenic Legionella spp. and Two Free-Living Amoebae Species in a Complete Drinking Water System and Cooling Towers. Pathogens 2021, 10, 1407. https://doi.org/10.3390/pathogens10111407
Logan-Jackson A, Rose JB. Cooccurrence of Five Pathogenic Legionella spp. and Two Free-Living Amoebae Species in a Complete Drinking Water System and Cooling Towers. Pathogens. 2021; 10(11):1407. https://doi.org/10.3390/pathogens10111407
Chicago/Turabian StyleLogan-Jackson, Alshae, and Joan B. Rose. 2021. "Cooccurrence of Five Pathogenic Legionella spp. and Two Free-Living Amoebae Species in a Complete Drinking Water System and Cooling Towers" Pathogens 10, no. 11: 1407. https://doi.org/10.3390/pathogens10111407
APA StyleLogan-Jackson, A., & Rose, J. B. (2021). Cooccurrence of Five Pathogenic Legionella spp. and Two Free-Living Amoebae Species in a Complete Drinking Water System and Cooling Towers. Pathogens, 10(11), 1407. https://doi.org/10.3390/pathogens10111407






