Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy)
Abstract
1. Introduction
2. Results
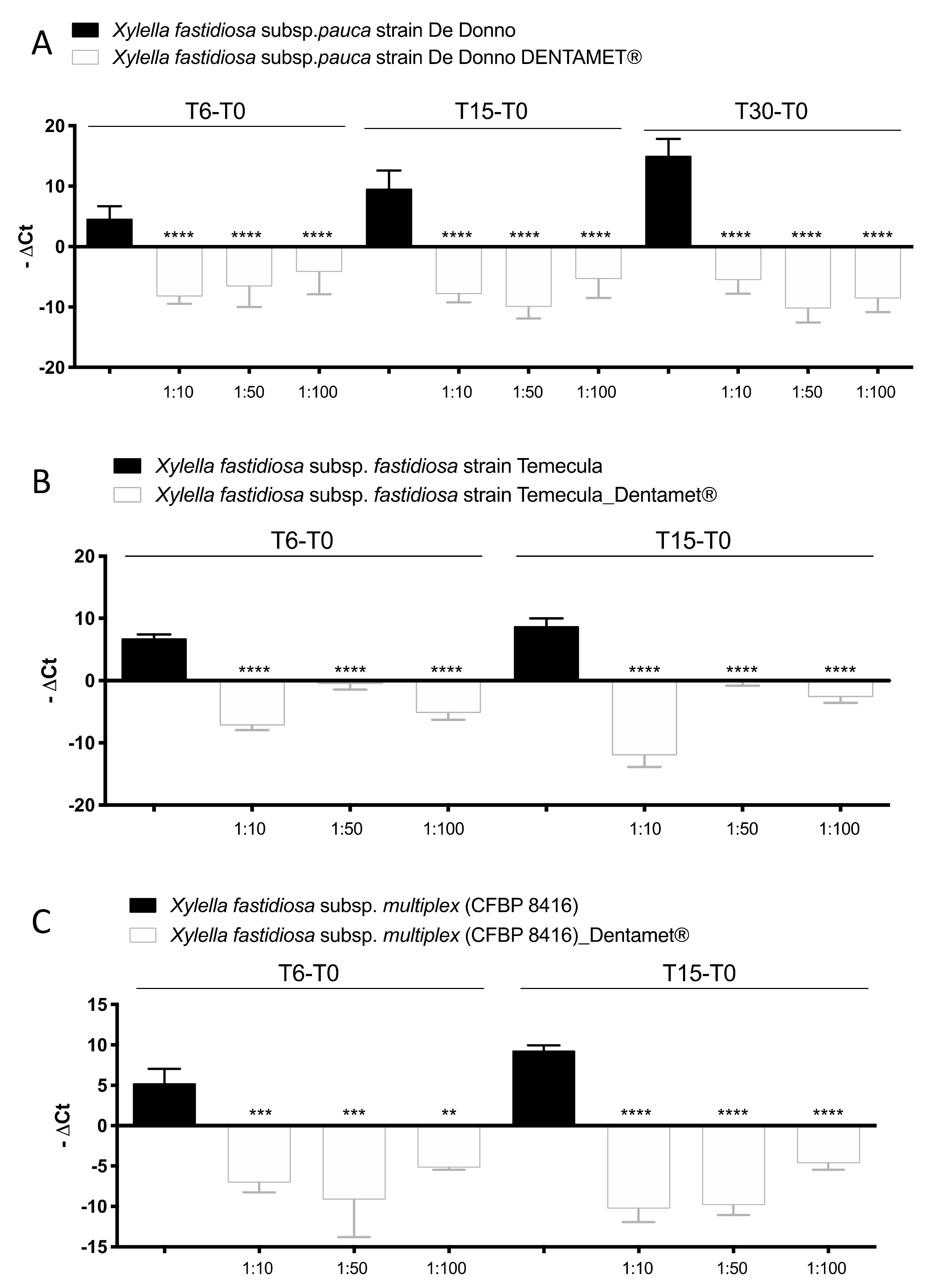
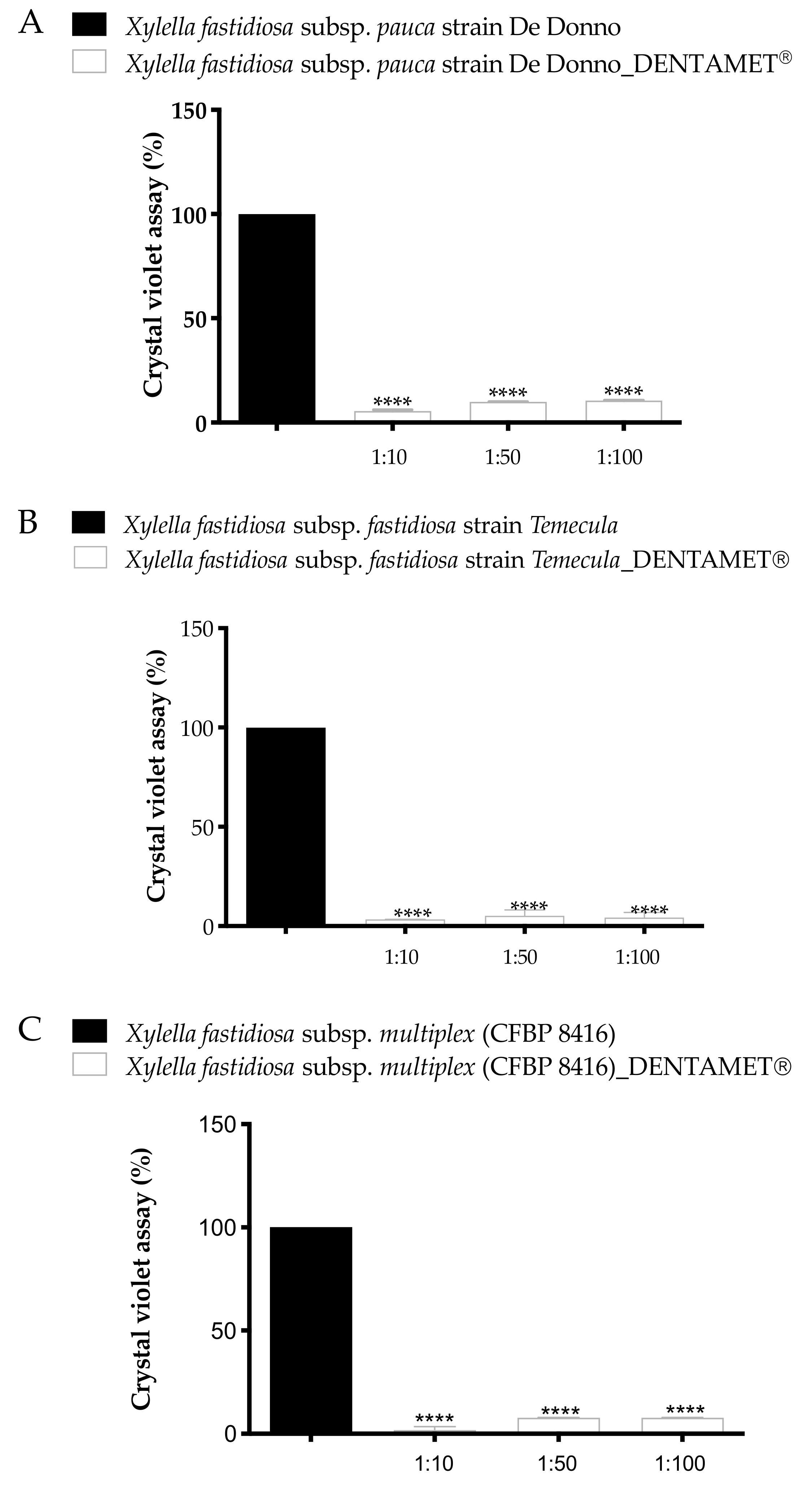
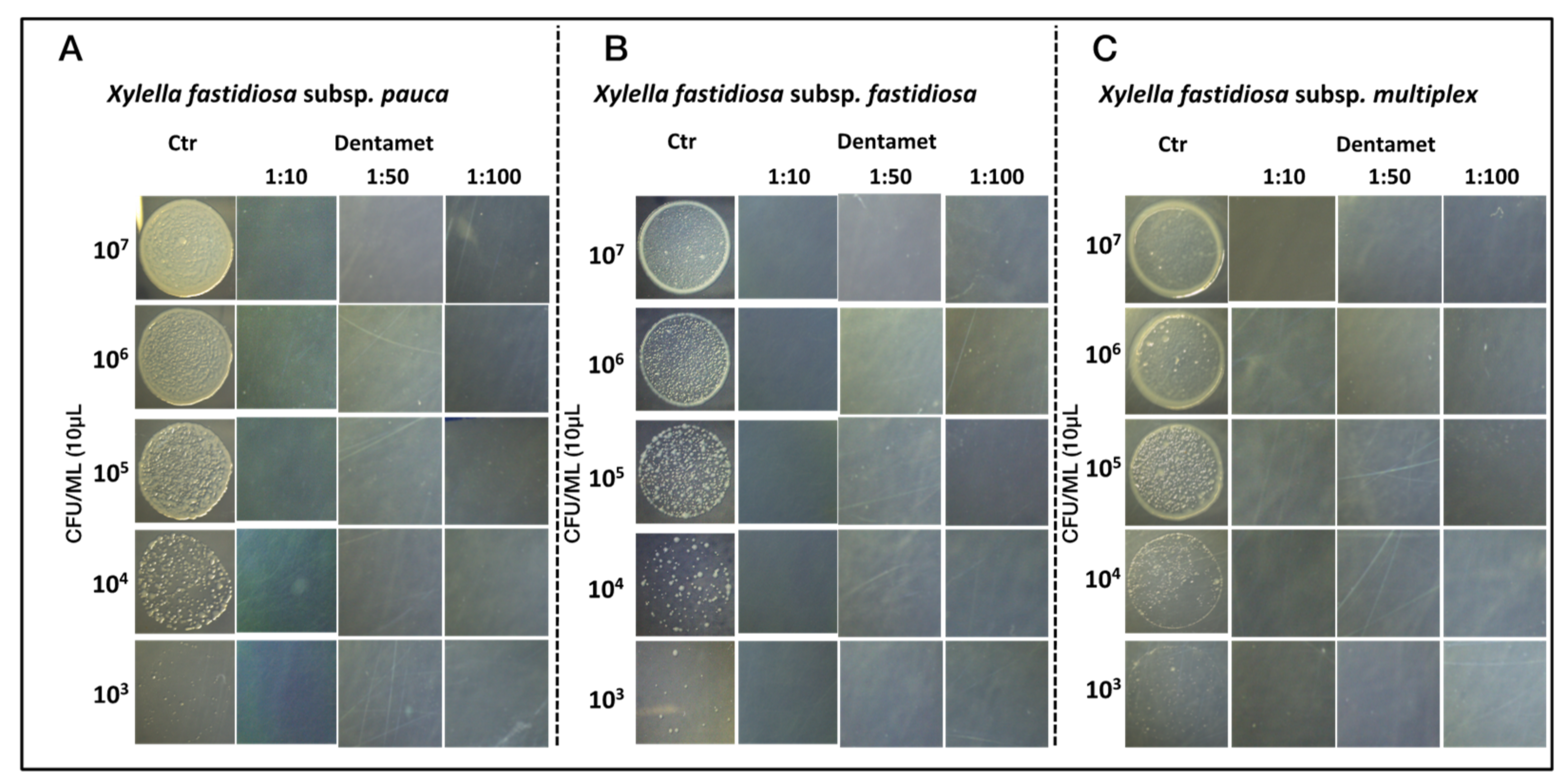
2.1. In Vitro Antibacterial Activity, Agar Dilution and Biofilm Inhibition Assays
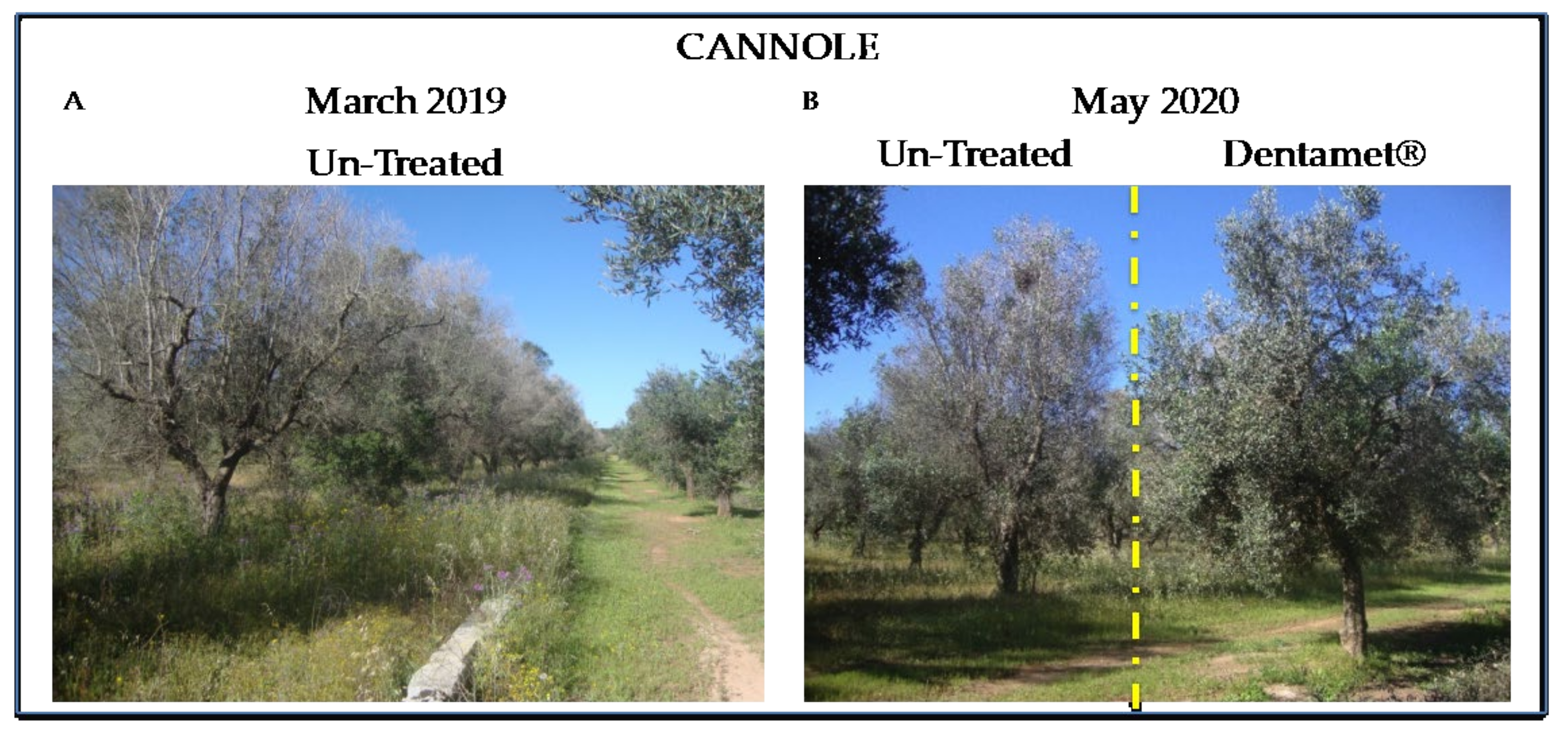
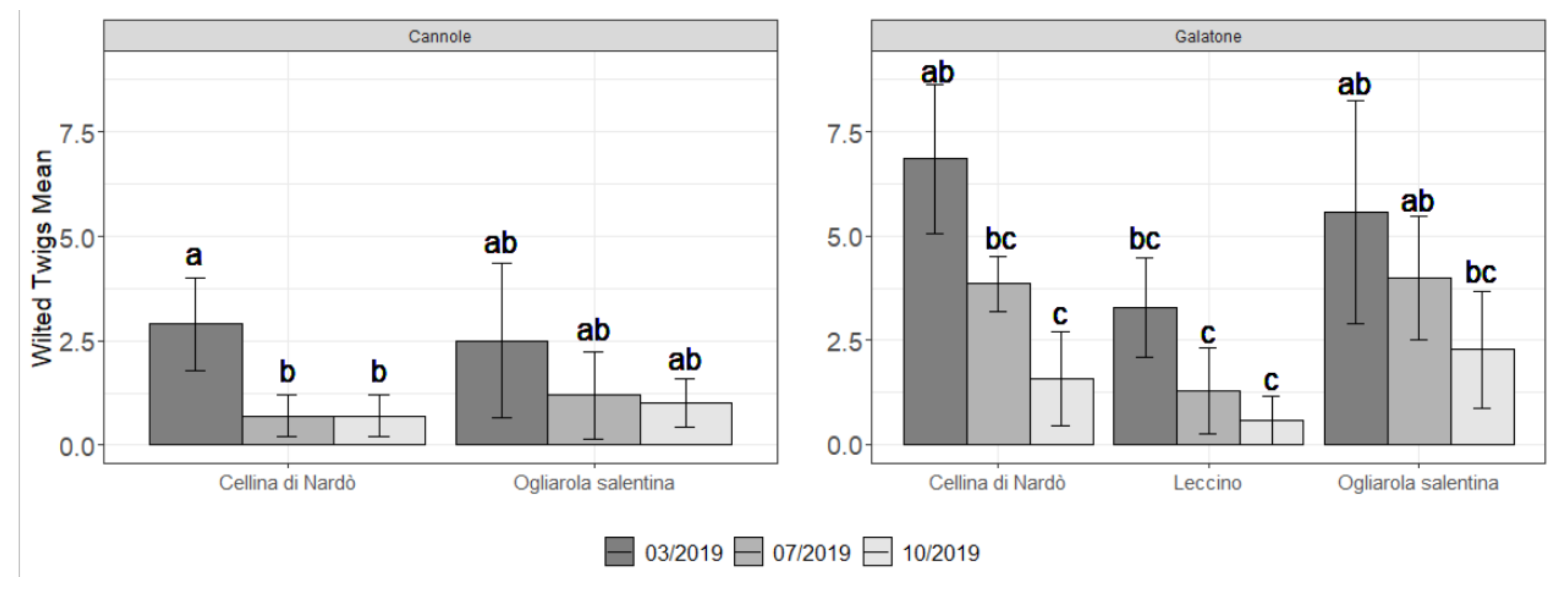
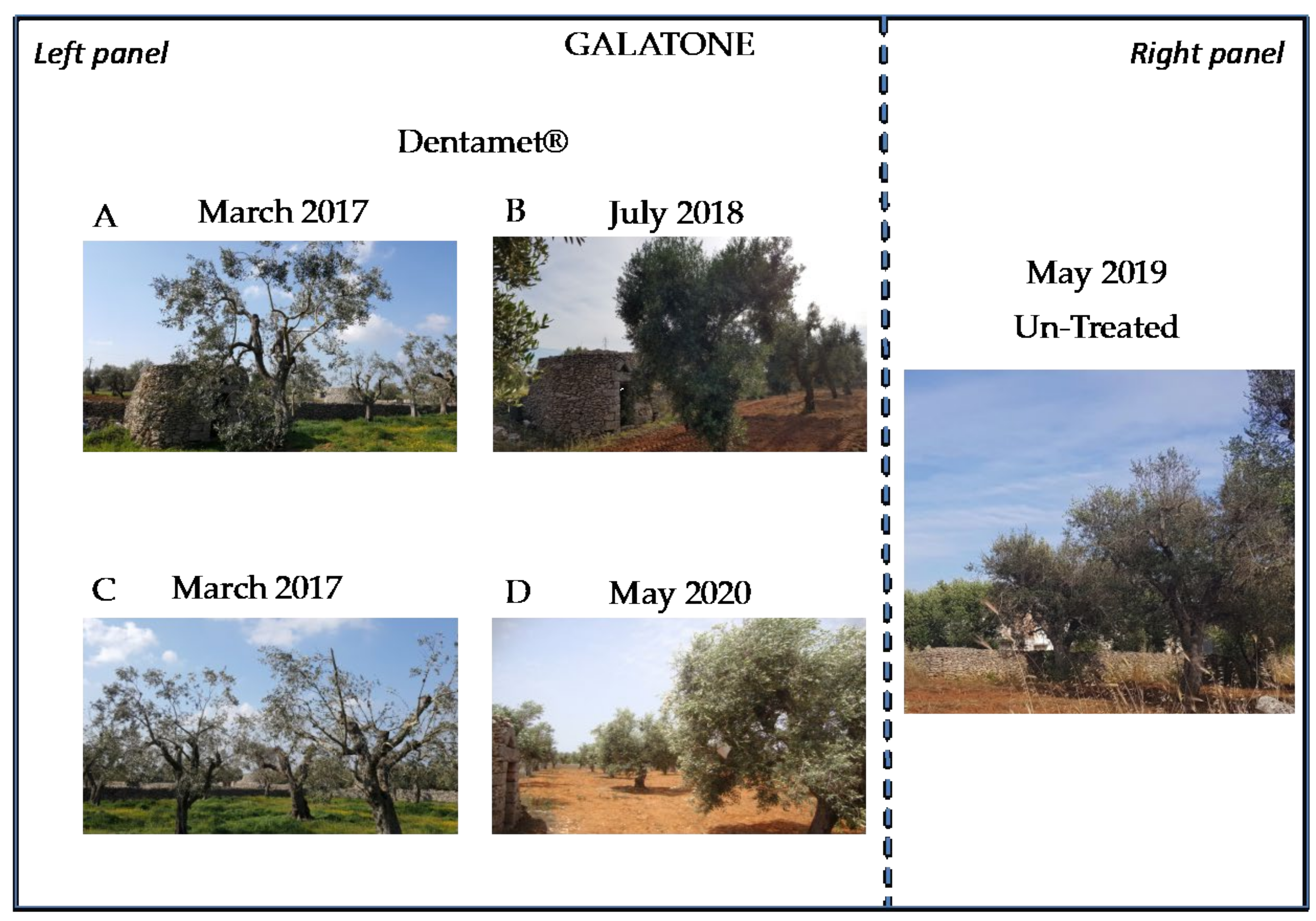
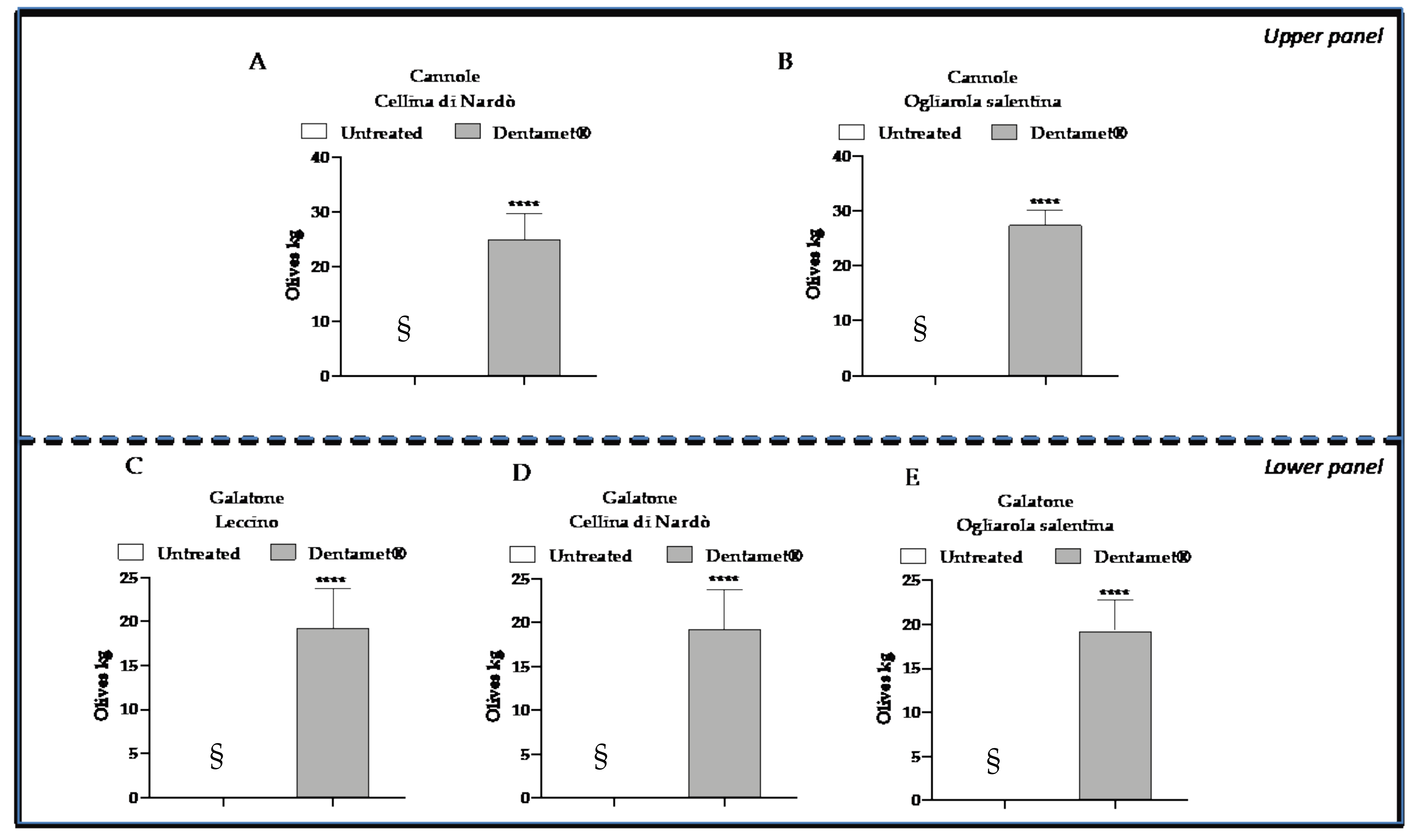
2.2. Field Symptoms Assessment and Tree Yield
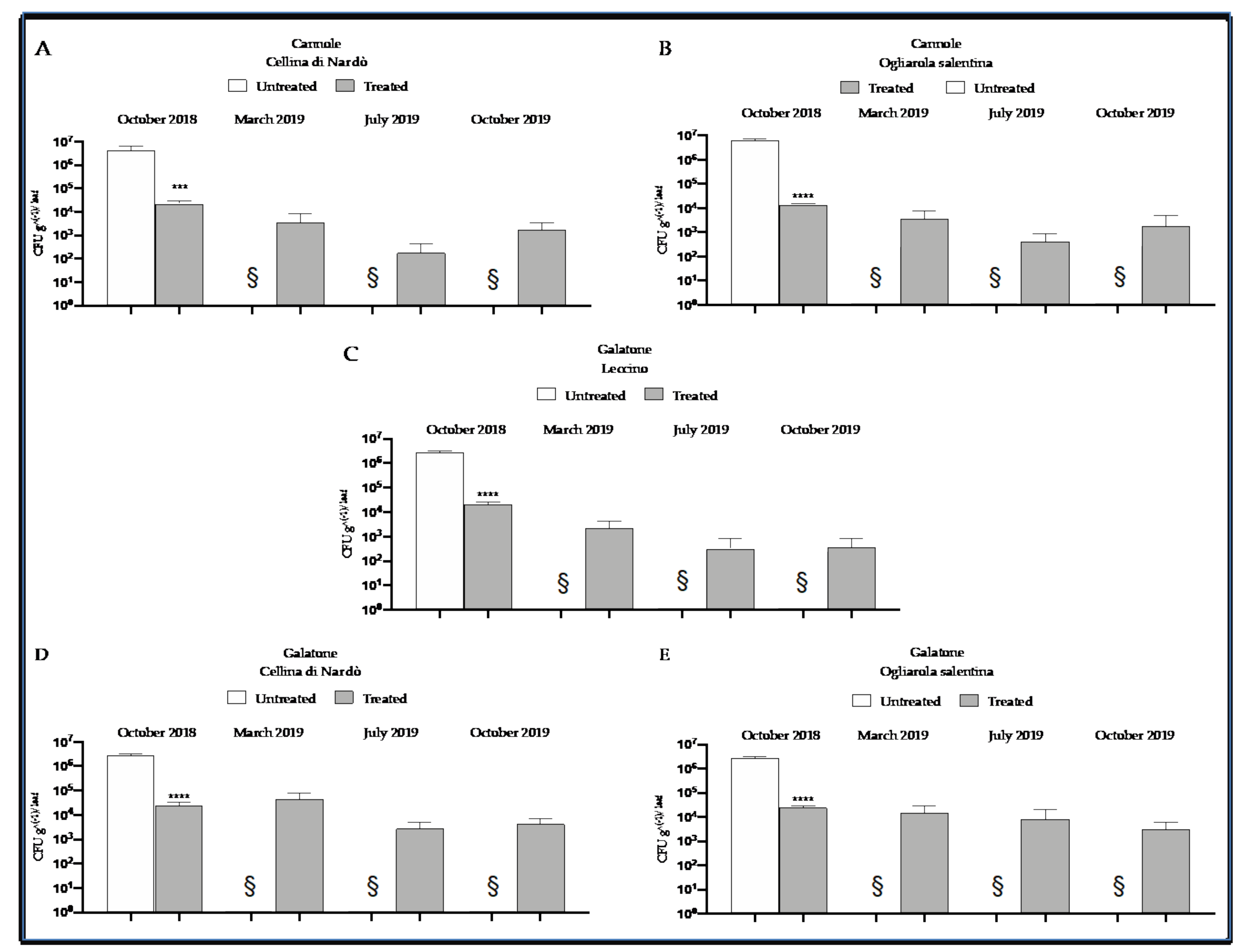
2.3. Quantitative Real-Time PCR Assessment of Xfp within Olive Leaves
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains Used in This Study
4.2. In Vitro Antibacterial Assays
4.3. Genomic DNA Extraction and Real-Time PCR
4.4. Agar Dilution and Biofilm Inhibition Assays
4.5. Statistical Analysis
4.6. Choice of the Olive Trees
4.7. Quantitative Real-Time Assessment of Xfp within Olive Leaves
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Saponari, M.; Boscia, D. Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (southern Italy). J. Plant. Pathol. 2013, 95, 668. [Google Scholar]
- Cariddi, C.; Saponari, M. Isolation of Xylella fastidiosa strain infecting olive and oleander in Apulia, Italy. Plant Pathol. J. 2014, 96, 425–429. [Google Scholar]
- Martelli, G.P. The current status of the quick decline syndrome of olive in southern Italy. Phytoparasitica 2015, 44, 1–10. [Google Scholar] [CrossRef]
- Scortichini, M.; Cesari, G. An Evaluation of Monitoring Surveys of the Quarantine Bacterium Xylella Fastidiosa Performed in Containment and Buffer Areas of Apulia, Southern Italy. Appl. Biosaf. 2019, 24, 96–99. [Google Scholar] [CrossRef]
- Scholten, R.; Martinez Sanchez, L. Monitoring the impact of Xylella on Apulia’s olive orchards using MODIS satellite data supported by weather data. 2019. 2nd European Conference on Xylella fastidiosa, Ajaccio 29–30 October. Available online: http://www.efsa.europa.eu/sites/default/files/event/191029-xylella/S6.P1_BECK.pdf (accessed on 29 October 2019).
- Saponari, M.; Loconsole, G.; Cornara, D.; Yokomi, R.K.; De Stradis, A.; Boscia, D.; Bosco, D.; Gp, G.P.M.; Krugner, R.; Porcelli, F. Infectivity and Transmission of Xylella fastidiosa by Philaenus spumarius (Hemiptera: Aphrophoridae) in Apulia, Italy. J. Econ. Èntomol. 2014, 107, 1316–1319. [Google Scholar] [CrossRef]
- Fierro, A.; Liccardo, A.; Porcelli, F. A lattice model to manage the vector and the infection of the Xylella fastidiosa on olive trees. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Strona, G.; Carstens, C.J.; Beck, P.S.A. Network analysis reveals why Xylella fastidiosa will persist in Europe. Sci. Rep. 2017, 7, 71. [Google Scholar] [CrossRef]
- Schneider, K.; Van der Werf, W. Impact of Xylella fastidiosa subspecies pauca in European olives. Proc. Natl. Acad. Sci. USA 2020, 117, 9250–9259. [Google Scholar] [CrossRef]
- Giampetruzzi, A.; Morelli, M.; Saponari, M.; Loconsole, G.; Chiumenti, M.; Boscia, D.; Savino, V.N.; Martelli, G.P.; Saldarelli, P. Transcriptome profiling of two olive cultivars in response to infection by the CoDiRO strain of Xylella fastidiosa subsp. pauca. BMC Genom. 2016, 17, 1–18. [Google Scholar] [CrossRef]
- Boscia, D.; Altamura, G. Resistenza a Xylella fastidiosa in diverse cultivar di olivo. L’Informatore Agrario 2017, 11, 59–63. [Google Scholar]
- Terra e vita. Available online: https://terraevita.edagricole.it/featured/xylella-mango-avocado/ (accessed on 29 October 2019).
- Catalano, L.; Al-Dobai, S. Guidelines for the Prevention, Eradication and Containment of Xylella Fastidiosa in Olive-Growing Areas; Food and Agriculture Organization of the United Nations: Cairo, Egypt, 2019; p. 55. [Google Scholar]
- Scortichini, M.; Chen, J. A zinc, copper and citric acid biocomplex shows promise for control of Xylella fastidiosa subsp. pauca in olive trees in Apulia region (southern Italy). Phytopathol. Mediterr. 2018, 57, 48–72. [Google Scholar]
- Del Coco, L.; De Pascali, S.A.; Fanizzi, F.P. NMR-Metabolomic Study on Monocultivar and Blend Salento EVOOs including Some from Secular Olive Trees. Food Nutr. Sci. 2014, 5, 89–95. [Google Scholar] [CrossRef]
- Negro, C.; Aprile, A.; Luvisi, A.; Nicolì, F.; Nutricati, E.; Vergine, M.; Miceli, A.; Blando, F.; Sabella, E.; De Bellis, L. Phenolic Profile and Antioxidant Activity of Italian Monovarietal Extra Virgin Olive Oils. Antioxidants 2019, 8, 161. [Google Scholar] [CrossRef] [PubMed]
- Colella, C.; Carradone, R. Problem Setting and Problem Solving in the Case of Olive Quick Decline Syndrome in Apulia, Italy: A Sociological Approach. Phytophatology 2019, 109, 187–199. [Google Scholar] [CrossRef] [PubMed]
- PM 7/24 (4) Xylella fastidiosa. Eppo Bull. 2019, 49, 175–227. [CrossRef]
- D’Attoma, G.; Morelli, M.; De La Fuente, L.; Cobine, P.A.; Saponari, M.; De Souza, A.A.; De Stradis, A.; Saldarelli, P. Phenotypic Characterization and Transformation Attempts Reveal Peculiar Traits of Xylella fastidiosa Subspecies pauca Strain De Donno. Microorganisms 2020, 8, 1832. [Google Scholar] [CrossRef]
- Marcelletti, S.; Scortichini, M. Genome-wide comparison and taxonomic relatedness of multiple Xylella fastidiosa strains reveal the occurrence of three subspecies and a new Xylella species. Arch. Microbiol. 2016, 198, 803–812. [Google Scholar] [CrossRef]
- Denancé, N.; Briand, M.; Gaborieau, R.; Gaillard, S.; Jacques, M.A. Identification of genetic relationships and subspecies signatures in Xylella fastidiosa. BMC Genom. 2019, 20, 1–21. [Google Scholar] [CrossRef]
- Gonella, E.; Orrù, B.; Alma, A. Egg masses treatment with micronutrient fertilizers has a suppressive effect on newly-emerged nymphs of the brown marmorated stink bug Halyomorpha halys. Èntomol. Gen. 2019, 39, 231–238. [Google Scholar] [CrossRef]
- Macomber, L.; Imlay, J.A. The Iron-Sulfur clusters of dehydratases are primary intracellular targets of copper toxicity. Proc. Natl. Acad. Sci. USA 2009, 106, 8344–8349. [Google Scholar] [CrossRef]
- Lemire, J.A.; Harrison, J.J.; Turner, R.J. Antimicrobial activity of metals: Mechanisms, molecular targets and applications. Nat. Rev. Genet. 2013, 11, 371–384. [Google Scholar] [CrossRef] [PubMed]
- Ishida, T. Antibacterial Mechanism of Bacteriolyses of Bacterial Cell Walls by Zinc(2+) Ion Induced Activations of PGN Autolysins, and DNA damages. J. Genes Proteins. 2017, 1, 1–7. [Google Scholar]
- Cobine, P.A.; Cruz, L.F.; Navarrete, F.; Duncan, D.; Tygart, M.; De La Fuente, L. Xylella fastidiosa Differentially Accumulates Mineral Elements in Biofilm and Planktonic Cells. PLoS ONE 2013, 8, e54936. [Google Scholar] [CrossRef] [PubMed]
- Navarrete, F.; De La Fuente, L. Response of Xylella fastidiosa to Zinc: Decreased Culturability, Increased Exopolysaccharide Production, and Formation of Resilient Biofilms under Flow Conditions. Appl. Env. Microbiol. 2014, 80, 1097–1107. [Google Scholar] [CrossRef]
- Navarrete, F.; De La Fuente, L. Zinc Detoxification Is Required for Full Virulence and Modification of the Host Leaf Ionome by Xylella fastidiosa. Mol. Plant Microbe Interact. 2015, 28, 497–507. [Google Scholar] [CrossRef]
- Phan, T.-N.; Buckner, T.; Sheng, J.; Baldeck, J.D.; Marquis, R.E. Physiologic actions of zinc related to inhibition of acid and alkali production by oral streptococci in suspensions and biofilms. Oral Microbiol. Immunol. 2004, 19, 31–38. [Google Scholar] [CrossRef]
- Baldassarre, F.; Tatulli, G.; Vergaro, V.; Mariano, S.; Scala, V.; Nobile, C.; Pucci, N.; Dini, L.; Loreti, S.; Ciccarella, G. Sonication-Assisted Production of Fosetyl-Al Nanocrystals: Investigation of Human Toxicity and In Vitro Antibacterial Efficacy against Xylella fastidiosa. Nanomaterials 2020, 10, 1174. [Google Scholar] [CrossRef]
- Ge, Q.; Cobine, P.A.; De La Fuente, L. Copper Supplementation in Watering Solution Reaches the Xylem But Does Not Protect Tobacco Plants Against Xylella fastidiosa Infection. Plant Dis. 2020, 104, 724–730. [Google Scholar] [CrossRef]
- Shuttleworth, K.L.; Unz, R.F. Influence of metals and metal speciation on the growth of filamentous bacteria. Water Res. 1991, 25, 1177–1186. [Google Scholar] [CrossRef]
- Cabrero, A.; Fernandez, S.; Mirada, F.; Garcia, J.; García, J. Effects of copper and zinc on the activated sludge bacteria growth kinetics. Water Res. 1998, 32, 1355–1362. [Google Scholar] [CrossRef]
- Utgikar, V.; Chaudhary, N.; Koeniger, A.; Tabak, H.H.; Haines, J.R.; Govind, R. Toxicity of metals and metal mixtures: Analysis of concentration and time dependence for zinc and copper. Water Res. 2004, 38, 3651–3658. [Google Scholar] [CrossRef] [PubMed]
- Şengör, S.S.; Gikas, P.; Moberly, J.G.; Peyton, B.M.; Ginn, T.R. Comparison of single and joint effects of Zn and Cu in continuous flow and batch reactors. J. Chem. Technol. Biotechnol. 2011, 87, 374–380. [Google Scholar] [CrossRef]
- Zhitnisky, D.; Rose, J. The higly synergistic, broad spectrum, antibacterial activity of organic acids and transition metals. Sci. Rep. 2017, 7, 44554. [Google Scholar] [CrossRef] [PubMed]
- Roper, C.; Castro, C. Xylella fastidiosa: Bacterial parasitism with hallmarks of commensalism. Curr. Opin. Plant Biol. 2019, 50, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Cardinale, M.; Luvisi, A.; Meyer, J.B.; Sabella, E.; De Bellis, L.; Cruz, A.C.; Ampatzidis, Y.; Cherubini, P. Specific Fluorescence in Situ Hybridization (FISH) Test to Highlight Colonization of Xylem Vessels by Xylella fastidiosa in Naturally Infected Olive Trees (Olea europaea L.). Front. Plant Sci. 2018, 9, 431. [Google Scholar] [CrossRef]
- Del Coco, L.; Migoni, D.; Girelli, C.R.; Angilè, F.; Scortichini, M.; Fanizzi, F.P. Soil and Leaf Ionome Heterogeneity in Xylella fastidiosa Subsp. Pauca-Infected, Non-Infected and Treated Olive Groves in Apulia, Italy. Plants 2020, 9, 760. [Google Scholar] [CrossRef]
- De La Fuente, L.; Parker, J.K. The Bacterial Pathogen Xylella fastidiosa Affects the Leaf Ionome of Plant Hosts during Infection. PLoS ONE 2013, 8, e62945. [Google Scholar] [CrossRef]
- D’Attoma, G.; Morelli, M.; Saldarelli, P.; Saponari, M.; Giampetruzzi, A.; Boscia, D.; Savino, V.N.; De La Fuente, L.; Cobine, P.A. Ionomic Differences between Susceptible and Resistant Olive Cultivars Infected by Xylella fastidiosa in the Outbreak Area of Salento, Italy. Pathogens 2019, 8, 272. [Google Scholar] [CrossRef]
- Cao, T.; Connell, J.H. Influence of inoculation date on the colonisation of Xylella fastidiosa and the persistence of almond leaf scorch disease among almond cultivars. Plant Dis. 2011, 95, 158–165. [Google Scholar] [CrossRef]
- Saponari, M.; Boscia, D.; Altamura, G.; Loconsole, G.; Zicca, S.; D’Attoma, G.; Morelli, M.; Palmisano, F.; Tavano, D.; Savino, V.N.; et al. Isolation and pathogenicity of Xylella fastidiosa associated to the olive quick decline syndrome in southern Italy. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]
- Luvisi, A.; Aprile, A. Xylella fastidiosa subsp. pauca (CoDiRO strain) infection in four olive (Olea europaea L.) cultivars: Profile of phenolic compounds in leaves and progression of leaf scorch symptoms. Phytopathol. Mediterr. 2017, 56, 259–273. [Google Scholar]
- Dongiovanni, C.; Cavalieri, V.; Bodino, N.; Tauro, D.; Di Carolo, M.; Fumarola, G.; Altamura, G.; Lasorella, C.; Bosco, D. Plant Selection and Population Trend of Spittlebug Immatures (Hemiptera: Aphrophoridae) in Olive Groves of the Apulia Region of Italy. J. Econ. Èntomol. 2018, 112, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Bodino, N.; Cavalieri, V.; Dongiovanni, C.; Plazio, E.; Saladini, M.A.; Volani, S.; Simonetto, A.; Fumarola, G.; Di Carolo, M.; Porcelli, F.; et al. Phenology, seasonal abundance and stage-structure of spittlebug (Hemiptera: Aphrophoridae) populations in olive groves in Italy. Sci. Rep. 2019, 9, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Weaver, C.R.; King, D.R. Meadow Spittlebug; Ohio Agriculture Experimental Station Research: Wooster, OH, USA, 1954. [Google Scholar]
- Cornara, D.; Saponari, M.; Zeilinger, A.R.; De Stradis, A.; Boscia, D.; Loconsole, G.; Bosco, D.; Martelli, G.P.; Almeida, R.P.P.; Porcelli, F. Spittlebugs as vectors of Xylella fastidiosa in olive orchards in Italy. J. Pest Sci. 2017, 90, 521–530. [Google Scholar] [CrossRef]
- Gucci, R.; Cantini, C. Pruning and Training Systems for Modern Olive Growing; CSIRO Publishing: Collingwood, Australia, 2000; p. 147. [Google Scholar]
- Berendson, R.L.; Pieterse, C.M.J. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef]
- Scortichini, M. The multi-millenial olive agroecosystem of Salento (Apulia, Italy) threatened by Xylella fastidiosa subsp. pauca: A working possibility of restoration. Sustainability 2020, 12, 6700. [Google Scholar] [CrossRef]
- Bleve, G.; Gallo, A.; Altomare, C.; Vurro, M.; Maiorano, G.; Cardinali, A.; D’Antuono, I.; Marchi, G.; Mita, G. In vitro activity of antimicrobial compounds against Xylella fastidiosa, the causal agent of the olive quick decline syndrome in Apulia (Italy). FEMS Microbiol. Lett. 2018, 365, 281. [Google Scholar] [CrossRef]
- Zaini, P.A.; De La Fuente, L. Grapevine Xylem Sap Enhances Biofilm Development by Xylella Fastidiosa. FEMS Microbiol. Lett. 2009, 295, 129–134. [Google Scholar] [CrossRef]
- Harper, S.J.; Ward, L.I.; Clover, G.R.G. Development of LAMP and Real-Time PCR Methods for the Rapid Detection of Xylella fastidiosa for Quarantine and Field Applications. Phytopathology 2010, 100, 1282–1288. [Google Scholar] [CrossRef]
- Modesti, V.; Pucci, N.; Lucchesi, S.; Campus, L.; Loreti, S. Experience of the Latium region (Central Italy) as a pest-free area for monitoring of Xylella fastidiosa: Distinctive features of molecular diagnostic methods. Eur. J. Plant Pathol. 2016, 148, 557–566. [Google Scholar] [CrossRef]


| CFU g^ (−1) 7 leaf | ||||||
|---|---|---|---|---|---|---|
| Cultivar | Month a | Plot | Mean | SD b | Median | IQR c |
| Ogliarola d | March | Galatone | 1.5 × 104 | 1.5 × 104 | 1.2 × 104 | 1.6 × 104 |
| Ogliarola | July | Galatone | 8.3 × 103 | 1.6 × 103 | 2.7 × 103 | 5.3 × 103 |
| Ogliarola | October | Galatone | 3.1 × 103 | 3.7 × 103 | 1.4 × 103 | 4.6 × 103 |
| Ogliarola | March | Cannole | 3.7 × 103 | 5.9 × 103 | 2.6 × 103 | 4.0 × 103 |
| Ogliarola | July | Cannole | 4.4 × 102 | 5.9 × 102 | 1.4 × 102 | 7.2 × 102 |
| Ogliarola | October | Cannole | 1.9 × 103 | 4.4 × 103 | 3.9 × 102 | 5.0 × 103 |
| Cellina e | March | Galatone | 4.5 × 104 | 4.4 × 104 | 2.6 × 104 | 4.0 × 104 |
| Cellina | July | Galatone | 4.4 ×102 | 5.9 × 102 | 1.4 × 102 | 7.2 × 102 |
| Cellina | October | Galatone | 4.4 × 103 | 3.3 × 103 | 3.9 × 103 | 5.0 × 103 |
| Cellina | March | Cannole | 3.6 × 103 | 6.2 × 103 | 1.9 × 100 | 4.9 × 103 |
| Cellina | July | Cannole | 1.2 × 102 | 3.5 × 102 | 9.2 × 100 | 1.2 × 102 |
| Cellina | October | Cannole | 1.7 × 103 | 2.6 × 103 | 7.5 × 102 | 1.8 × 102 |
| Leccino | March | Cannole | 2.2 × 103 | 3.1 × 103 | 7.5 × 102 | 3.7 × 103 |
| Leccino | July | Cannole | 1.8 × 102 | 3.5 × 102 | 9.2 × 100 | 1.3 × 102 |
| Leccino | October | Cannole | 3.5 × 102 | 7.9 × 102 | 0 | 1.8 × 102 |
| March f | 1.2 × 104 | 2.5 × 104 | 4.5 × 103 | 1.4 × 104 | ||
| July f | 2.1 × 103 | 7.4 × 103 | 1.2 × 102 | 1.1 × 102 | ||
| October f | 2.2 × 103 | 3.4 × 103 | 7.8 × 102 | 2.4 × 103 | ||
| Galatone g | 9.0 × 103 | 2.1 × 104 | 1.4 × 103 | 7.5 × 103 | ||
| Cannole g | 1.9 × 103 | 4.2 × 103 | 2.4 × 102 | 1.5 × 103 | ||
| Ogliarola in Galatone h | 8.7 × 103 | 1.4 × 104 | 4.0 × 103 | 1.0 × 104 | ||
| Cellina in Galatone h | 1.7 × 104 | 3.2 × 104 | 4.6 × 103 | 1.6 × 104 | ||
| Leccino in Galatone h | 9.1 × 102 | 2.1 × 103 | 4.4 × 100 | 5.9 × 102 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tatulli, G.; Modesti, V.; Pucci, N.; Scala, V.; L’Aurora, A.; Lucchesi, S.; Salustri, M.; Scortichini, M.; Loreti, S. Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy). Pathogens 2021, 10, 85. https://doi.org/10.3390/pathogens10010085
Tatulli G, Modesti V, Pucci N, Scala V, L’Aurora A, Lucchesi S, Salustri M, Scortichini M, Loreti S. Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy). Pathogens. 2021; 10(1):85. https://doi.org/10.3390/pathogens10010085
Chicago/Turabian StyleTatulli, Giuseppe, Vanessa Modesti, Nicoletta Pucci, Valeria Scala, Alessia L’Aurora, Simone Lucchesi, Manuel Salustri, Marco Scortichini, and Stefania Loreti. 2021. "Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy)" Pathogens 10, no. 1: 85. https://doi.org/10.3390/pathogens10010085
APA StyleTatulli, G., Modesti, V., Pucci, N., Scala, V., L’Aurora, A., Lucchesi, S., Salustri, M., Scortichini, M., & Loreti, S. (2021). Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy). Pathogens, 10(1), 85. https://doi.org/10.3390/pathogens10010085






