Early-Life Exposure to Environmental Contaminants Perturbs the Sperm Epigenome and Induces Negative Pregnancy Outcomes for Three Generations via the Paternal Lineage
Abstract
1. Introduction
2. Materials and Methods
2.1. POPs Mixture and Animals
2.2. F1, F2 and F3 Progeny Outcome and Male Fertility
2.3. Male Reproductive Assessment for F1, F2, F3
2.3.1. Sperm Analysis
2.3.2. Daily Testicular Spermatid Production
2.3.3. Histological Evaluation of Follicle Numbers
2.3.4. Testosterone Assay
2.3.5. Sperm DNA Methylation Assays
2.4. Statistical Analyses
3. Results
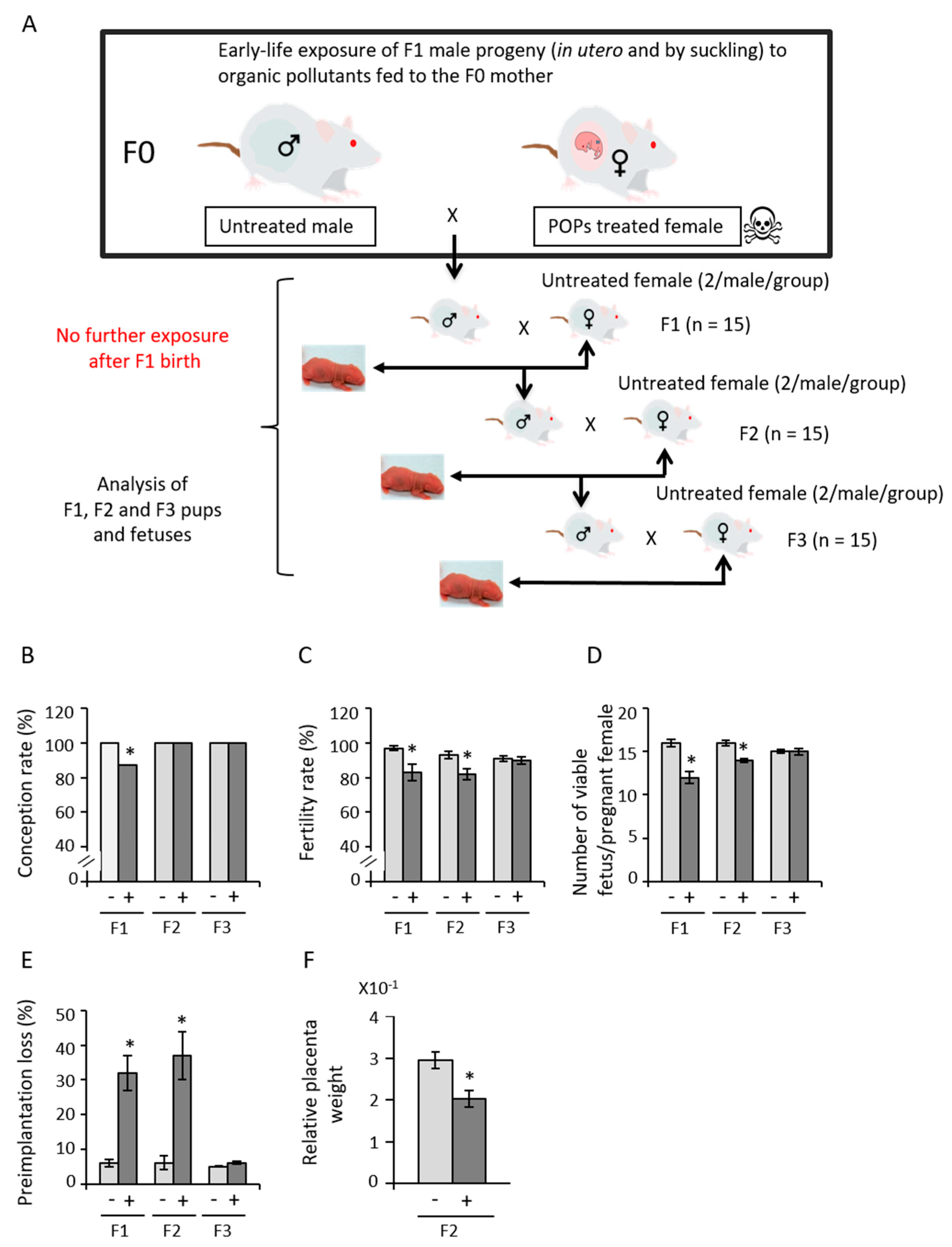
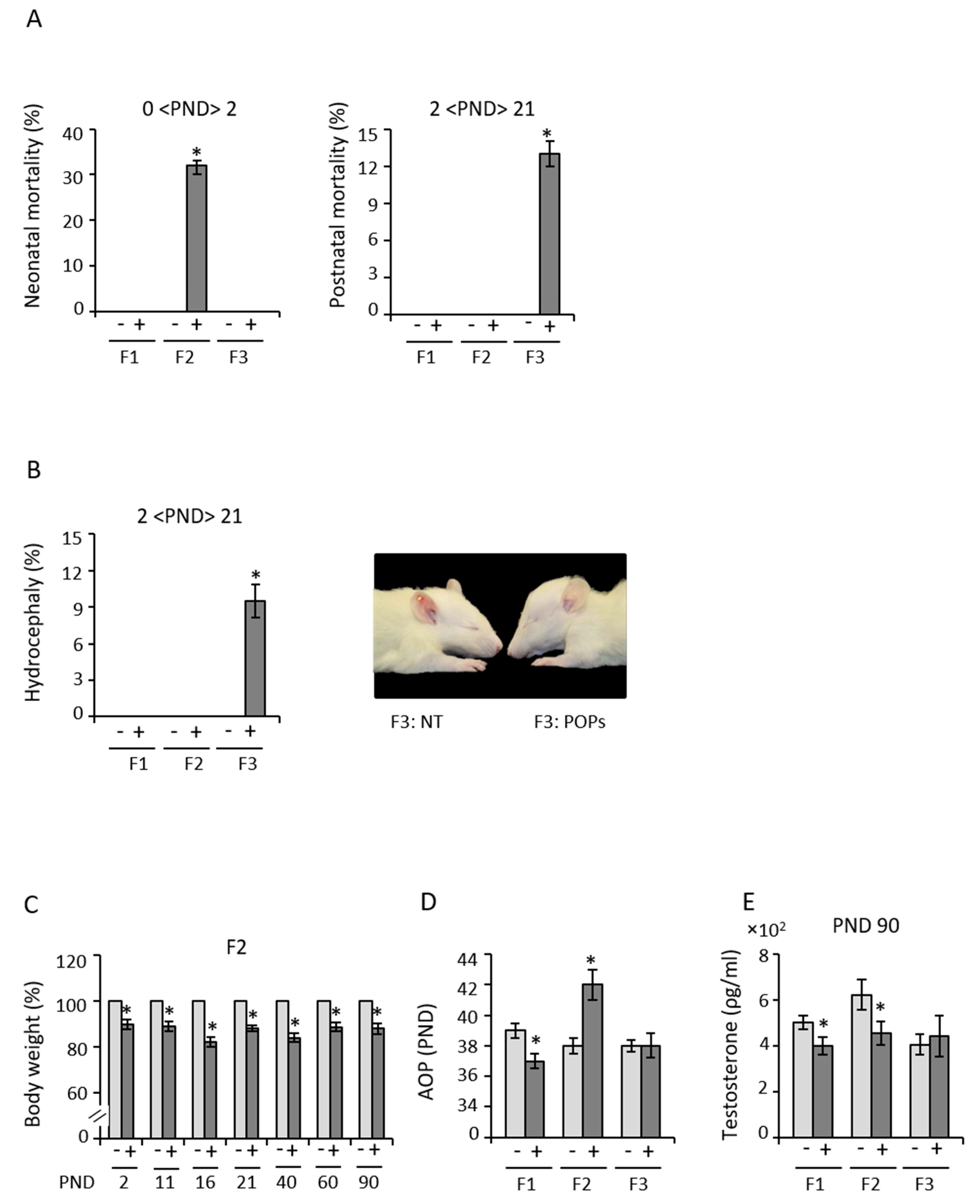
3.1. Early Exposure to POPs Produces Heritable Developmental Abnormalities
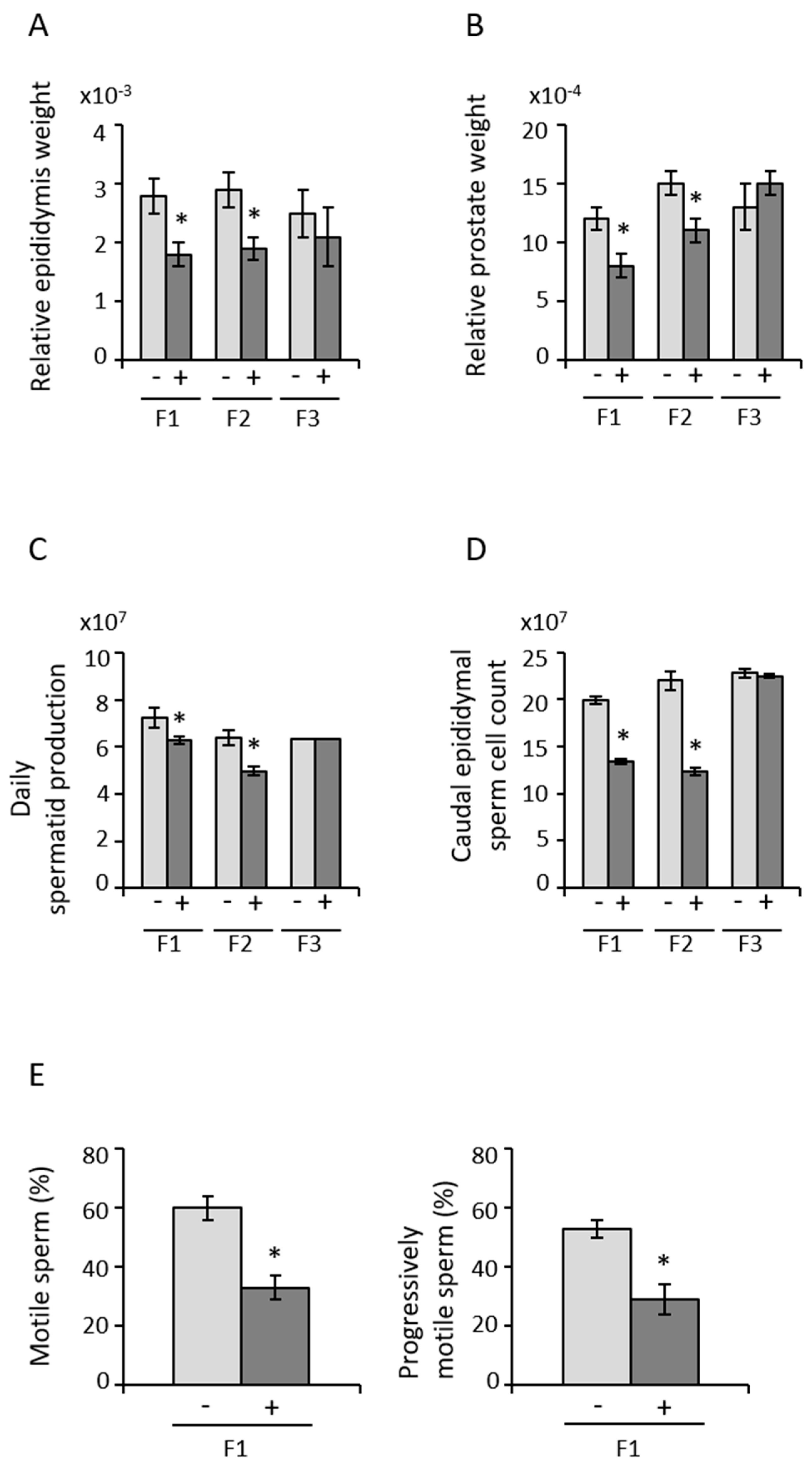
3.2. Paternal Transmission of POPs-Associated Anomalies in Reproductive Tract Development and Sperm Parameters
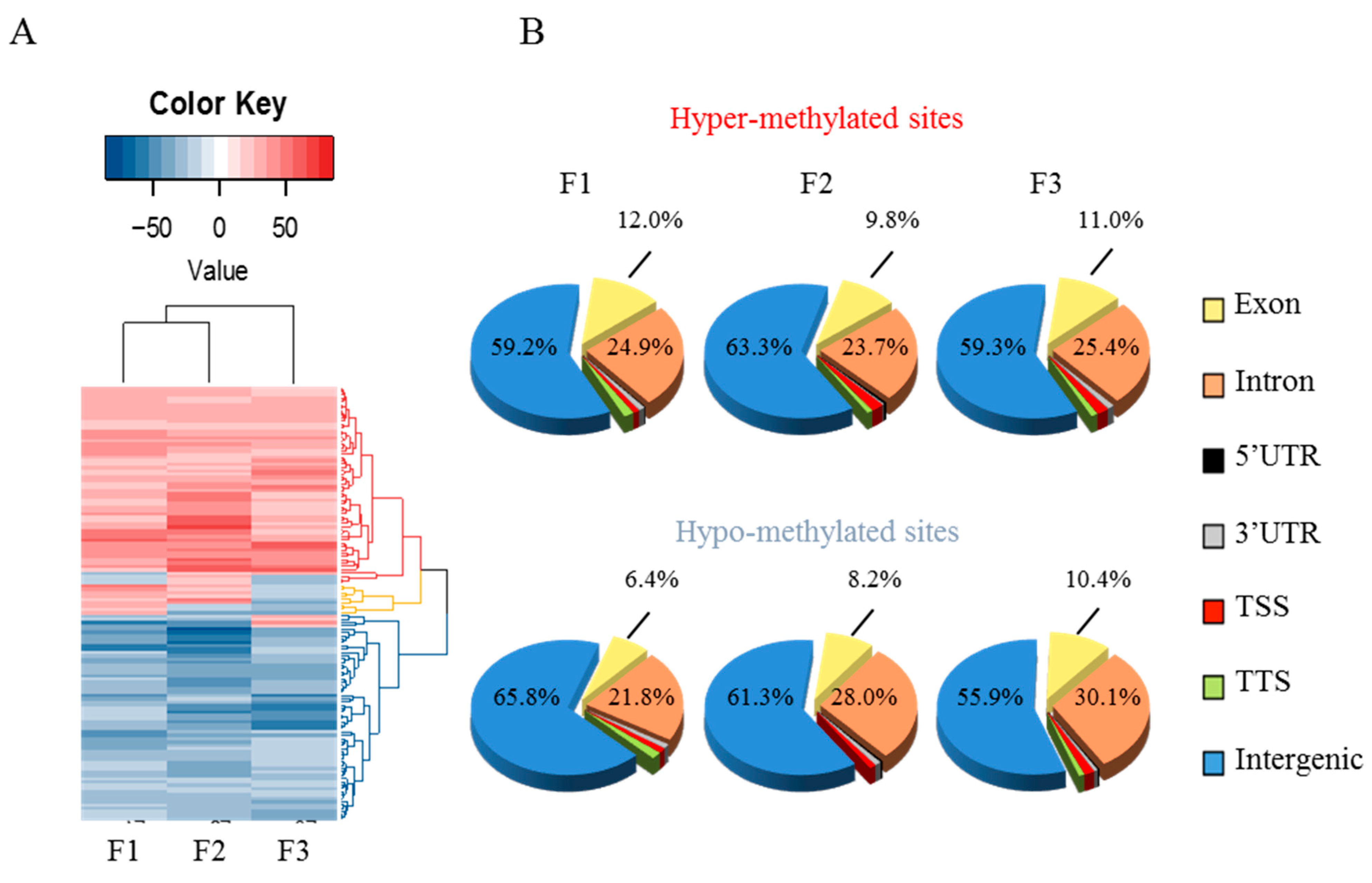
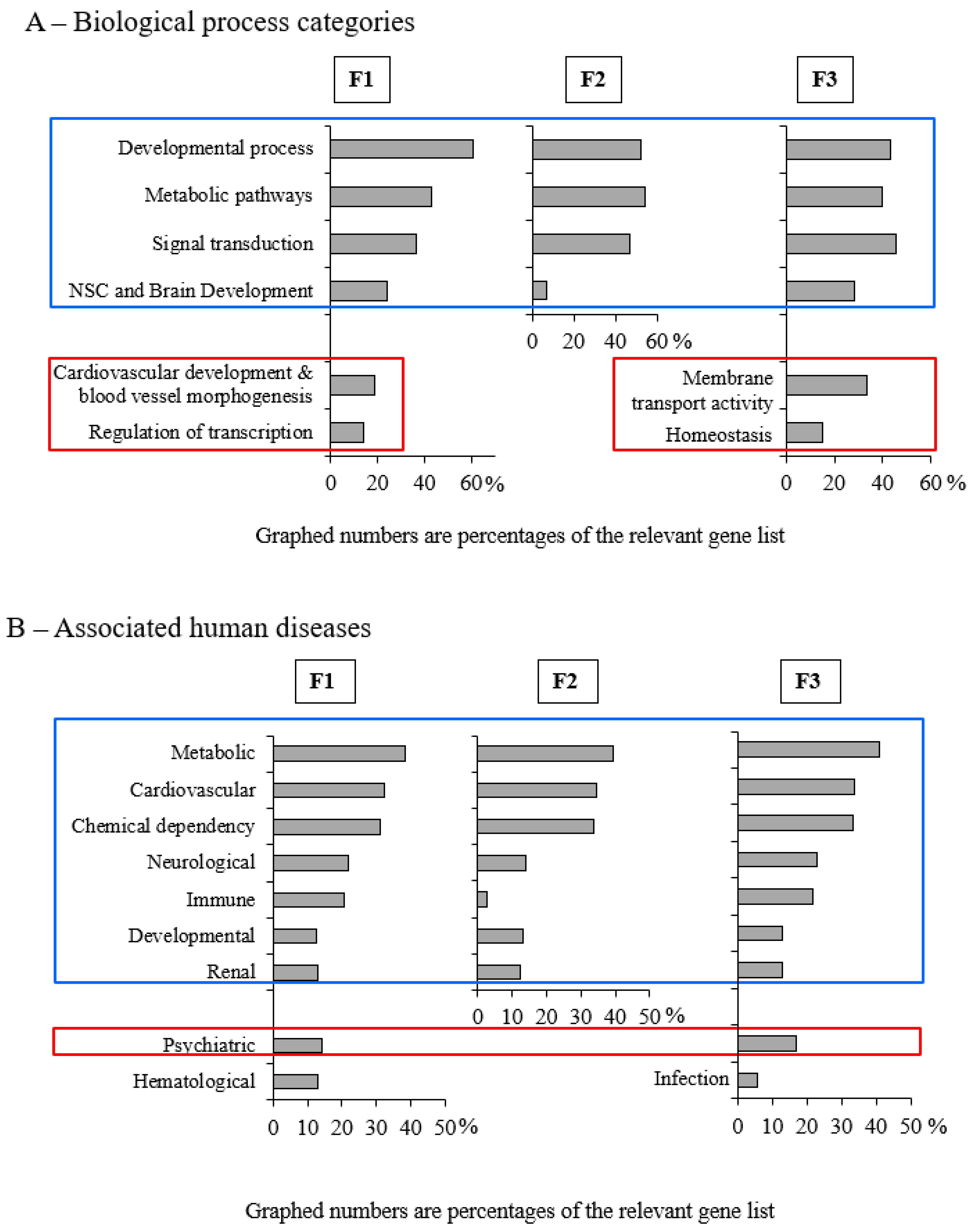
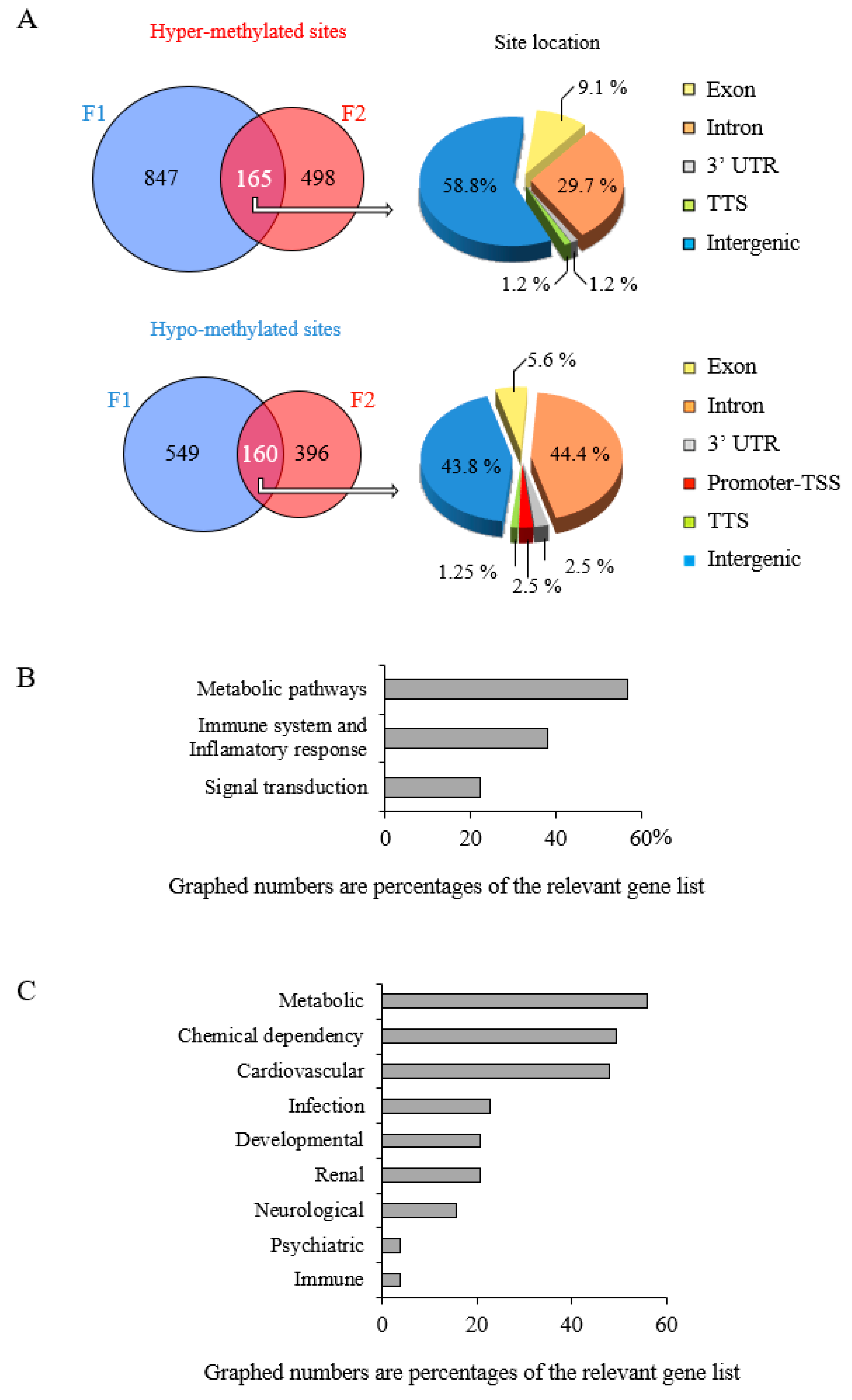
3.3. Early-Life Exposure to POPs Induced Modifications of the Sperm DNA Methylome Linked to Human Diseases
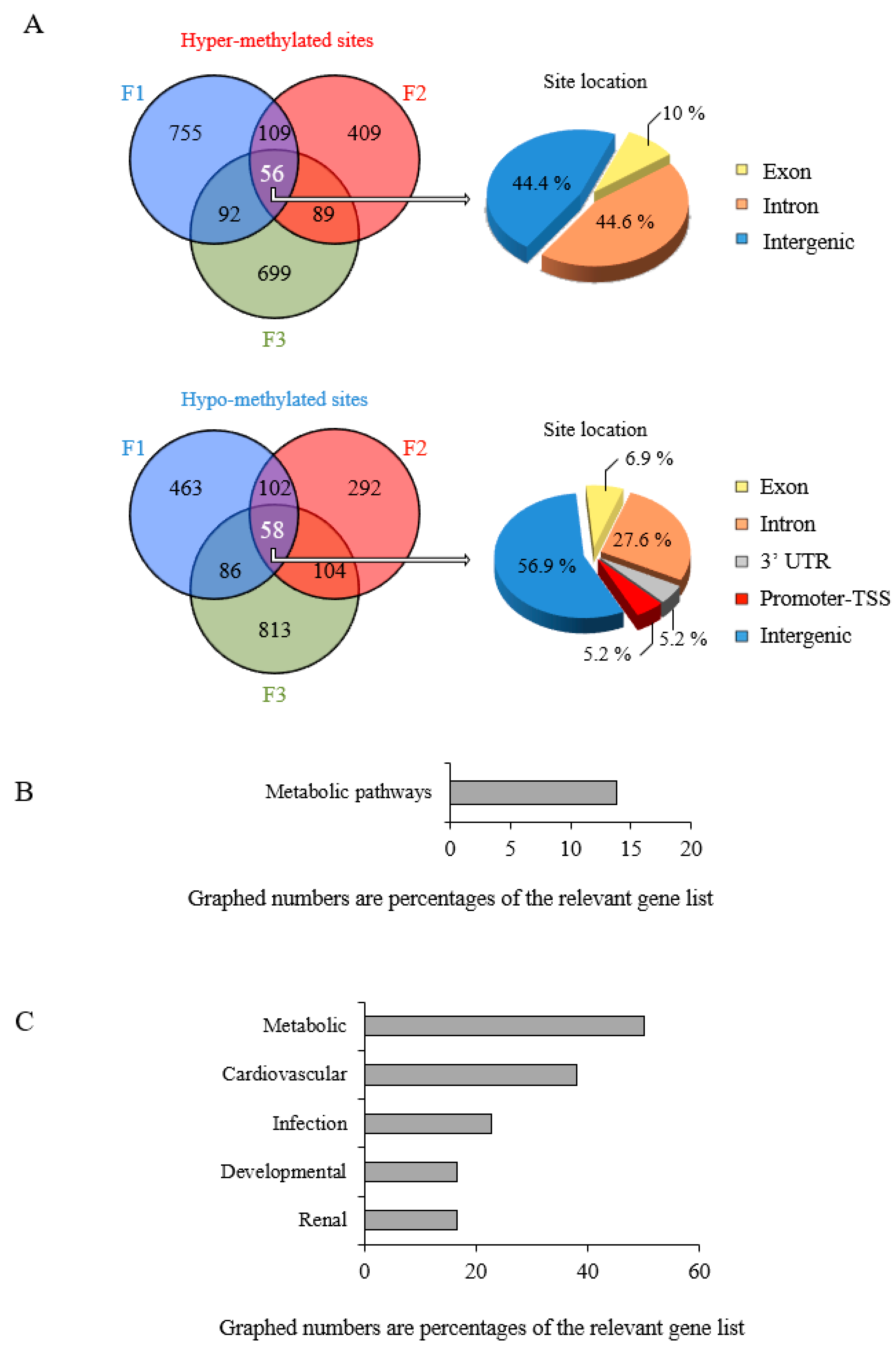
3.4. Epigenetic Marks Modified by POPs Are Heritable
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Androutsopaulos, V.P.; Hernandez, A.F.; Liesivuori, J.; Tsatsakis, A.M. A mechanistic overview of health associated effects of low levels of organochlorine and organophosphorous pesticides. Toxicology 2013, 307, 89–94. [Google Scholar] [CrossRef] [PubMed]
- Longnecker, M.P.; Rogan, W.J.; Lucier, G. The human health effects of DDT (dichlorodiphenyl-trichloroethane) and PCBs (polychlorinated biphenyls) and an overview of organochlorines in public health. Annu. Rev. Public Health 1997, 18, 211–244. [Google Scholar] [CrossRef] [PubMed]
- The World Health Organization. Persistent Organic Pollutants (POPs). Available online: http://chm.pops.int/TheConvention/ThePOPs/The12InitialPOPs/tabid/296/Default.aspx (accessed on 1 March 2021).
- Bonde, J.P.; Toft, G.; Rylander, L.; Rignell-Hydbom, A.; Giwercman, A.; Spano, M.; Manicardi, G.C.; Bizzaro, D.; Ludwicki, J.; Zvyezday, V.; et al. Fertility and markers of male reproductive function in Inuit and European populations spanning large contrasts in blood levels of persistent organochlorines. Environ. Health Perspect. 2008, 116, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Butler Walker, J.; Seddon, L.; McMullen, E.; Jan, H.; Tofflemire, K.; Corriveau, A.; Weber, J.; Mills, C.; Smith, S.; van Oostdam, J. Organochlorine levels in maternal and umbilical cord blood plasma in Arctic Canada. Sci. Total Environ. 2003, 302, 27–52. [Google Scholar] [CrossRef]
- Dewailly, E.; Ayotte, P.; Bruneau, S.; Laliberté, C.; Muir, D.C.; Norstrom, R.J. Inuit exposure to organochlorines through the aquatic food chain in arctic quebec. Environ. Health Perspect. 1993, 101, 618–620. [Google Scholar] [CrossRef]
- Brown, T.M.; Macdonald, R.W.; Muir, D.C.G.; Letcher, R.J. The distribution and trends of persistent organic pollutants and mercury in marine mammals from Canada’s Eastern Arctic. Sci. Total Environ. 2018, 618, 500–517. [Google Scholar] [CrossRef] [PubMed]
- Rigét, F.; Bignert, A.; Braune, B.; Dam, M.; Dietz, R.; Evans, M.; Green, N.; Gunnlaugsdóttir, H.; Hoydal, K.S.; Kucklick, J.; et al. Temporal trends of persistent organic pollutants in Arctic marine and freshwater biota. Sci. Total. Environ. 2019, 649, 99–110. [Google Scholar] [CrossRef]
- Donaldson, S.; Van Oostdam, J.; Tikhonov, C.; Feeley, M.; Armstrong, B.; Ayotte, P.; Boucher, O.; Bowers, W.; Chan, L.; Dallaire, F.; et al. Environmental contaminants and human health in the Canadian Arctic. Sci. Total. Environ. 2010, 408, 5165–5234. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, A.J.; Hetherington, R.A. Decade of research in Inuit children, youth, and maternal health in Canada: Areas of concentrations and scarcities. Int. J. Circumpolar Health 2012, 71, 18383. [Google Scholar] [CrossRef][Green Version]
- Auger, N.; Park, A.L.; Zoungrana, H.; Gros-Louis McHugh, N.; Luo, Z.C. Rates of stillbirth by gestational age and cause in Inuit and First Nations populations in Quebec. CMAJ. 2013, 185, E256–E262. [Google Scholar] [CrossRef]
- Dallaire, R.; Dewailly, E.; Ayotte, P.; Forget-Dubois, N.; Jacobson, S.W.; Jacobson, J.L.; Muckle, G. Exposure to organochlorines and mercury through fish and marine mammal consumption: Associations with growth and duration of gestation among Inuit newborns. Environ. Int. 2013, 54, 85–91. [Google Scholar] [CrossRef]
- Verner, M.A.; Plisquellec, P.; Muckle, G.; Ayotte, P.; Dewailly, E.; Jacobson, S.W.; Jacobson, J.L.; Charbonneau, M.; Haddad, S. Alteration of infant attention and activity by polychlorinated biphenyls: Unravelling critical windows of susceptibility using physiologically based pharmacokinetic modeling. Neurotoxicology 2010, 31, 424–431. [Google Scholar] [CrossRef] [PubMed]
- Dallaire, F.; Dewailly, E.; Vézina, C.; Muckle, G.; Weber, J.-P.; Bruneau, S.; Ayotte, P. Effect of prenatal exposure to polychlorinated biphenyls on incidence of acute respiratory infections in preschool Inuit children. Environ. Health Perspect. 2006, 114, 1301–1305. [Google Scholar] [CrossRef] [PubMed]
- Plusquellec, P.; Muckle, G.; Dewailly, E.; Ayotte, P.; Bégin, G.; Desrosiers, C.; Després, C.; Saint-Amour, D.; Poitras, K. The relation of environmental contaminants exposure to behavioral indicators in Inuit preschoolers in Arctic Quebec. Neurotoxicology 2010, 31, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Kaati, G.; Bygren, L.O.; Edvinsson, S. Cardiovascular and diabetes mortality determined by nutrition during parents’ and grandparents’ slow growth period. Eur. J. Hum. Genet. 2002, 10, 682–688. [Google Scholar] [CrossRef]
- Kaati, G.; Bygren, L.O.; Pembrey, M.; Sjostrom, M. Transgenerational response to nutrition, early life circumstances and longevity. Eur. J. Hum. Genet. 2007, 15, 784–790. [Google Scholar] [CrossRef]
- Pembrey, M.E.; Bygren, L.O.; Kaati, G.; Edvinsson, S.; Northstone, K.; Sjöström, M.; Golding, J.; ALSPAC Study Team. Sex-specific, male-line transgenerational responses in humans. Eur. J. Hum. Genet. 2006, 14, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.F.; Lin, R.C.Y.; Laybutt, R.; Barres, R.; Owens, J.A.; Morris, M.J. Chronic high-fat diet in fathers programs beta-cell dysfunction in female rat offspring. Nature 2010, 467, 963–966. [Google Scholar] [CrossRef]
- Watkins, A.J.; Sinclair, K.D. Paternal low protein diet affects adult offspring cardiovascular and metabolic function in mice. Am. J. Physiol. Heart Circ. Physiol. 2014, 306, H1444–H1452. [Google Scholar] [CrossRef]
- Anway, M.D.; Cupp, A.S.; Uzumcu, M.; Skinner, M.K. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 2005, 308, 1466–1469. [Google Scholar] [CrossRef]
- Guerrero-Bosagna, C.; Covert, T.R.; Haque, M.M.; Settles, M.; Nilsson, E.E.; Anway, M.D.; Skinner, M.K. Epigenetic transgenerational inheritance of vinclozolin induced mouse adult onset disease and associated sperm epigenome biomarkers. Reprod. Toxicol. 2012, 34, 694–707. [Google Scholar] [CrossRef] [PubMed]
- Lambrot, R.; Xu, C.; Saint-Phar, S.; Chountalos, G.; Cohen, T.; Paquet, M.; Suderman, M.; Hallett, M.; Kimmins, S. Low paternal dietary folate alters the mouse sperm epigenome and is associated with negative pregnancy outcomes. Nat. Commun. 2013, 4, 2889. [Google Scholar] [CrossRef] [PubMed]
- Henríquez-Hernández, L.A.; Luzardo, O.P.; Zumbado, M.; Serra-Majem, L.; Valeron, P.F.; Camacho, M.; Álavarez-Pérez, J.; Salas-Salvadó, J.; Boada, L.D. Determinants of increasing serum POPs in a population at high risk for cardiovascular disease. Results from the PREDIMED-CANARIAS study. Environ. Res. 2017, 156, 477–484. [Google Scholar] [CrossRef]
- La Merrill, M.A.; Lind, P.M.; Salihovic, S.; van Bavel, B.; Lind, L. The association between p,p′-DDE levels and left ventricular mass is mainly mediated by obesity. Environ. Res. 2018, 160, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Vafeiadi, M.; Georgiou, V.; Chalkiadaki, G.; Rantakokko, P.; Kiviranta, H.; Karachaliou, M.; Fthenou, E.; Venihaki, M.; Sarri, K.; Vassilaki, M.; et al. Association of Prenatal Exposure to Persistent Organic Pollutants with Obesity and Cardiometabolic Traits in Early Childhood: The Rhea Mother–Child Cohort (Crete, Greece). Environ. Heal. Perspect. 2015, 123, 1015–1021. [Google Scholar] [CrossRef]
- Raffetti, E.; Donato, F.; Spezianu, F.; Scarcella, C.; Gaia, A.; Magoni, M. Polychlorinated bisphenyls (PCBs) exposure and cardiovascular, endocrine and matabolic diseases: A population-based cohort study in a North Italian highly polluted area. Environ. Int. 2018, 120, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Salihovic, S.; Ganna, A.; Fall, T.; Broeckling, C.D.; Prenni, J.E.; van Bavel, B.; Lind, M.P.; Ingelsson, E.; Lind, L. The metabolic gingerprint of p.p′-DDE and HCB exposure in humans. Environ. Int. 2016, 88, 60–66. [Google Scholar] [CrossRef]
- Coole, J.B.; Burr, S.S.; Kay, A.M.; Singh, J.A.; Kondakala, S.; Yang, E.-J.; Kaplan, B.L.F.; Howell, G.E., III; Stewart, J.A., Jr. Persistent organic pollutants (POPs) increase rage signalling to promote downstream cardiovascular remodelling. Environ. Toxicol. 2019, 34, 1149–1159. [Google Scholar] [CrossRef] [PubMed]
- King, S.E.; McBirney, M.; Beck, D.; Sadler-Riggleman, I.; Nilsson, E.; Skinner, M.K. Sperm epimutation biomarkers of obesity and pathologies following DDT induced epigenetic transgenerational inheritance of disease. Environ. Epigenet. 2019, 5, dvz008. [Google Scholar] [CrossRef]
- Skinner, M.K.; Ben Maamar, M.; Sadler-Riggleman, I.; Beck, D.; Nilsson, E.; McBirney, M.; Klukovich, R.; Xie, Y.; Tang, C.; Yan, W. Alterations in sperm DNA methylation, non-coding RNA and histone retention associate with DDT-induced epigenetic transgenerational inheritance of disease. Epigenetics Chromatin 2018, 11, 1–24. [Google Scholar] [CrossRef]
- Anas, M.K.; Guillemette, C.; Ayotte, P.; Pereg, D.; Giguère, F.; Bailey, J.L. In utero and lactational exposure to an environmentally relevant organochlorine mixture disrupts reproductive development and function in male rats. Biol. Reprod. 2005, 73, 414–426. [Google Scholar] [CrossRef] [PubMed]
- Maurice, C.; Kaczmarczyk, M.; Côté, N.; Tremblay, Y.; Kimmins, S.; Bailey, J.L. Prenatal exposure to an environmentally relevant mixture of Canadian Arctic contaminants decrease male reproductive function in an aging rat model. J. Dev. Orig. Health Dis. 2018, 9, 511–518. [Google Scholar] [CrossRef]
- Ostby, J.S.; Gray, L.E., Jr. Transgenerational (in utero/lactational) exposure to investigate the effects of endocrine disrupting compounds (EDCS) in rats. Curr. Protoc. Toxicol. 2004, 19, 1–16. [Google Scholar] [CrossRef]
- Bianchi, P.G.; Manicardi, G.C.; Bizzaro, D.; Bianchi, U.; Sakkas, D. Effect of deoxyribonucleic acid protamination on fluorochrome staining and in situ nick-translation of murine and human mature spermatozoa. Biol. Reprod. 1993, 49, 1083–1088. [Google Scholar] [CrossRef]
- Maurice, C.; O’Brien, J.M.; Yauk, C.L.; Marchetti, F. Integration of sperm DNA damage assessment into OECD test guidelines for genotoxicity testing using the MutaMouse model. Toxicol. Appl. Pharmacol. 2018, 357, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Borgeest, C.; Symonds, D.; Mayer, L.P.; Hoyer, P.B.; Flaws, J.A. Methoxychlor may cause ovarian follicular atresia and proliferation of the ovarian epithelium in the mouse. Toxicol. Sci. 2002, 68, 473–478. [Google Scholar] [CrossRef]
- Smith, B.J.; Plowchalk, D.R.; Sipes, I.G.; Mattison, D.R. Comparison of random and serial sections in assessment of ovarian toxicity. Reprod. Toxicol. 1991, 5, 379–383. [Google Scholar] [CrossRef]
- Boyle, P.; Clement, K.; Gu, H.; Smith, Z.D.; Ziller, M.; Fostel, J.L.; Holmes, L.; Meldrim, J.; Kelley, F.; Gnirke, A.; et al. Gel-free multiplexed reduced representation bisulfite sequencing for large-scale DNA methylation profiling. Genome Biol. 2012, 13, R92. [Google Scholar] [CrossRef]
- Gu, H.; Smith, Z.D.; Boyle, P.; Gnirke, A.; Meissner, A. Preparation of reduced representation bisulfite sequencing libraries for genome-scale DNA methylation profiling. Nat. Protoc. 2011, 6, 468–481. [Google Scholar] [CrossRef]
- Krueger, F.; Andrews, S.R. Bismark: A flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 2011, 27, 1571–1572. [Google Scholar] [CrossRef]
- Akalin, A.; Kormaksson, M.; Li, S.; Garrett-Bakelman, F.E.; Figuerosa, M.E.; Melnick, A.; Mason, C.E. methylKit: A comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 2012, 13, R87. [Google Scholar] [CrossRef]
- Heinz, S.; Benner, C.; Spann, N.; Bertolino, E.; Lin, Y.C.; Laslo, P.; Cheng, J.X.; Murre, C.; Singh, H.; Glass, C.K. Simple Combinations of Lineage-Determining Transcription Factors Prime cis-Regulatory Elements Required for Macrophage and B Cell Identities. Mol. Cell 2010, 38, 576–589. [Google Scholar] [CrossRef] [PubMed]
- Durinck, S.; Spellman, P.T.; Birney, E.; Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 2009, 4, 1184–1191. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Mudunuri, U.; Che, A.; Yi, M.; Stephens, R.M. bioDBnet: The biological database network. Bioinformatics 2009, 25, 555–556. [Google Scholar] [CrossRef]
- Belleau, P.; Deschênes, A.; Scott-Boyer, M.P.; Lambrot, R.; Dalvai, M.; Kimmins, S.; Bailey, J.L.; Droit, A. Inferring and modeling inheritance of differentially methylated changes across multiple generations. Nucleic Acids Res. 2018, 46, e85. [Google Scholar] [CrossRef] [PubMed]
- Bourc’his, D.; Bestor, T.H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 2004, 431, 96–99. [Google Scholar] [CrossRef]
- Bourc’his, D.; Xu, G.L.; Lin, C.S.; Bollman, B.; Bestor, T.H. Dnmt3L and the establishment of maternal genomic imprints. Science 2001, 294, 2536–2539. [Google Scholar] [CrossRef]
- Itoh, H.; Iwasaki, M.; Kasuga, Y.; Yokoyama, S.; Onuma, H.; Nishimura, H.; Kusama, R.; Yoshida, T.; Yokoyama, K.; Tsugane, S. Association between serum organochlorines and global methylation level of leukocyte DNA among Japanese women: A cross-sectional study. Sci. Total. Environ. 2014, 490, 603–609. [Google Scholar] [CrossRef]
- Kim, K.-Y.; Kim, D.-S.; Lee, S.-K.; Lee, I.-K.; Kang, J.-H.; Chang, Y.-S.; Jacobs, D.R.; Steffes, M.; Lee, D.-H. Association of low-dose exposure to persistent organic pollutants with global DNA hypomethylation in healthy Koreans. Environ. Health Perspect. 2010, 118, 370–374. [Google Scholar] [CrossRef]
- Rusiecki, J.A.; Baccarelli, A.; Bollati, V.; Tarantini, L.; Moore, L.E.; Bonefeld-Jorgensen, E.C. Global DNA hypomethylation is associated with high serum-persistent organic pollutants in Greenlandic Inuit. Environ. Health Perspect. 2008, 116, 1547–1552. [Google Scholar] [CrossRef]
- Blattler, A.; Yao, L.; Witt, H.; Guo, Y.; Nicolet, C.M.; Berman, B.P.; Farnham, P.J. Global loss of DNA methylation uncovers intronic enhancers in genes showing expression changes. Genome Biol. 2014, 15, 469. [Google Scholar] [CrossRef]
- Cheung, H.H.; Davis, A.J.; Lee, T.-L.; Pang, A.L.; Nagrani, S.; Rennert, O.M.; Chan, W.-Y. Methylation of an intronic region regulates miR-199a in testicular tumor malignancy. Oncogene 2011, 30, 3404–3415. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Epigenetics in cancer. N. Engl. J. Med. 2008, 358, 1148–1159. [Google Scholar] [CrossRef]
- Ogawa, O.; Eccles, M.R.; Szeto, J.; McNoe, L.A.; Yun, K.; Maw, M.A.; Smith, P.J.; Reeve, A.E. Relaxation of insulin-like growth factor II gene imprinting implicated in Wilms’ tumour. Nature 1993, 362, 749–751. [Google Scholar] [CrossRef]
- Xue, Q.; Zhou, Y.F.; Zhu, S.N.; Bulun, S.E. Hypermethylation of the CpG island spanning from exon II to intron III is associated with steroidogenic factor 1 expression in stromal cells of endometriosis. Reprod. Sci. 2011, 18, 1080–1084. [Google Scholar] [CrossRef]
- Chen, Q.; Yan, M.; Cao, Z.; Li, X.; Zhang, Y.; Shi, J.; Feng, G.-H.; Peng, H.; Zhang, X.; Qian, J.; et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 2016, 351, 397–400. [Google Scholar] [CrossRef]
- Sharma, U.; Conine, C.C.; Shea, J.M.; Boskovic, A.; Derr, A.G.; Bing, X.Y.; Belleannee, C.; Kucukural, A.; Serra, R.W.; Sun, F.; et al. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 2016, 351, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Siklenka, K.; Erkek, S.; Godmann, M.; Lambrot, R.; McGraw, S.; LaFleur, C.; Cohen, T.; Xia, J.; Suderman, M.; Hallett, M.; et al. Disruption of histone methylation in developing sperm impairs offspring health transgenerationally. Science 2015, 350, aab2006. [Google Scholar] [CrossRef] [PubMed]
- Sifakis, S.; Androutsopoulos, V.P.; Tsatsakis, A.M.; Spandidos, D.A. Human exposure to endocrine disrupting chemicals: Effects on the male and female reproductive systems. Environ. Toxicol. Pharmacol. 2017, 51, 56–70. [Google Scholar] [CrossRef] [PubMed]
- Skakkebaek, N.E.; Jørgensen, N.; Main, K.M.; Meyts, E.R.-D.; Leffers, H.; Andersson, A.-M.; Juul, A.; Carlsen, E.; Mortensen, G.K.; Jensen, T.K.; et al. Is human fecundity declining? Int. J. Androl. 2006, 29, 2–11. [Google Scholar] [CrossRef]
- Toft, G.; Thulstrup, A.M.; Jönsson, B.A.; Pedersen, H.S.; Ludwicki, J.K.; Zvezday, V.; Bonde, J.P. Fetal loss and maternal serum levels of 2,2′,4,4′,5,5′-hexachlorbiphenyl (CB-153) and 1,1-dichloro-2,2-bis(p-chlorophenyl)ethylene (p,p′-DDE) exposure: A cohort study in Greenland and two European populations. Environ. Health 2010, 9, 22. [Google Scholar] [CrossRef] [PubMed]
- Baschat, A.A.; Hecher, K. Fetal growth restriction due to placental disease. Semin. Perinatol. 2004, 28, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Sandovici, I.; Hoelle, K.; Angiolini, E.; Constancia, M. Placental adaptations to the maternal-fetal environment: Implications for fetal growth and developmental programming. Reprod. Biomed. Online 2012, 25, 68–89. [Google Scholar] [CrossRef]
- Cox, B.; Kotlyar, M.; Evangelou, A.I.; Ignatchenko, V.; Ignatchenko, A.; Whiteley, K.; Jurisica, I.; Adamson, S.L.; Rossant, J.; Kislinger, T. Comparative systems biology of human and mouse as a tool to guide the modeling of human placental pathology. Mol. Syst. Biol. 2009, 5, 279. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, P.; Aitken, R.P.; Rhind, S.M.; Racey, P.A.; Wallace, J.M. Influence of placentally mediated fetal growth restriction on the onset of puberty in male and female lambs. Reproduction 2001, 122, 375–383. [Google Scholar] [CrossRef]
- Fisher, M.M.; Eugster, E.A. What is in our environment that effects puberty? Reprod. Toxicol. 2014, 44, 7–14. [Google Scholar] [CrossRef]
- Tinggaard, J.; Mieritz, M.G.; Sorensen, K.; Mouritsen, A.; Hagen, C.P.; Aksglaede, L.; Wohlfart-Veje, C.; Juul, A. The physiology and timing of male puberty. Curr. Opin. Endocrinol. Diabetes Obes. 2012, 19, 197–203. [Google Scholar] [CrossRef]
- Gilbert, K.M.; Whitlow, A.B.; Pumford, N.R. Environmental contaminant and disinfection by-product trichloroacetaldehyde stimulates T cells in vitro. Int. Immunopharmacol. 2004, 4, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Kovesi, T.A.; Cao, Z.; Osborne, G.; Egeland, G.M. Severe early lower respiratory tract infection is associated with subsequent respiratory morbidity in preschool Inuit children in Nunavut, Canada. J. Asthma 2011, 48, 241–247. [Google Scholar] [CrossRef]
- Carpenter, D.O. Polychlorinated biphenyls (PCBs): Routes of exposure and effects on human health. Rev. Environ. Health 2006, 21, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Dirinck, E.L.; Dirtu, A.C.; Govindan, M.; Covaci, A.; van Gaal, L.F.; Jorens, P.G. Exposure to persistent organic pollutants: Relationship with abnormal glucose metabolism and visceral adiposity. Diabetes Care 2014, 37, 1951–1958. [Google Scholar] [CrossRef]
- Rylander, L.; Rignell-Hydbom, A.; Hagmar, L. A cross-sectional study of the association between persistent organochlorine pollutants and diabetes. Environ. Health 2005, 4, 28. [Google Scholar] [CrossRef]
- Kuhnlein, H.V.; Receveur, O.; Soueida, R.; Egeland, G.M. Arctic indigenous peoples experience the nutrition transition with changing dietary patterns and obesity. J. Nutr. 2004, 134, 1447–1453. [Google Scholar] [CrossRef]
- Singh, K.; Chan, H.M. Persistent organic pollutants and diabetes among Inuit in the Canadian Arctic. Environ. Int. 2017, 101, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Okajima, D.; Kudo, G.; Yokota, H. Antidepressant-like behavior in brain-specific angiogenesis inhibitor 2-deficient mice. J. Physiol. Sci. 2011, 61, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Lischinsky, J.E.; Sokolowski, K.; Li, P.; Esumi, S.; Kamal, Y.; Goodrich, M.; Oboti, L.; Hammond, T.R.; Krishnamoorthy, M.; Feldman, D.; et al. Embryonic transcription factor expression in mice predicts medial amygdala neuronal identity and sex-specific responses to innate behavioral cues. eLife 2017, 6, 21012. [Google Scholar] [CrossRef]
- Asbreuk, C.H.; van Schaick, H.S.; Co, J.J.; Kromkamp, M.; Smidt, M.P.; Burbach, J.P.H. The homeobox genes Lhx7 and Gbx1 are expressed in the basal forebrain cholinergic system. Neuroscience 2002, 109, 287–298. [Google Scholar] [CrossRef]
- Ochalek, A.; Szczesna, K.; Petazzi, P.; Kobolak, J.; Dinnyes, A. Generation of cholinergic and dopaminergic interneurons from human pluripotent stem cells as a relevant tool for in vitro modeling of neurological disorders pathology and therapy. Stem Cells Int. 2016, 2016, 5838934. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Liu, L.; Chen, Y.; Li, H.; Yang, L.; Wang, Y.; Qian, Q. Synaptosome-related (SNARE) genes and their interactions contribute to the susceptibility and working memory of attention-deficit/hyperactivity disorder in males. Prog. Neuropsychopharmacol. Biol. Psychiatry 2015, 57, 132–139. [Google Scholar] [CrossRef]
- Fitzgerald, E.F.; Belanger, E.E.; Gomez, M.I.; Cayo, M.; McCaffrey, R.J.; Seegal, R.F.; Jansing, R.L.; Hwang, S.-A.; Hicks, H.E. Polychlorinated biphenyl exposure and neuropsychological status among older residents of upper Hudson River communities. Environ. Health Perspect. 2008, 116, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Roegge, C.S.; Schantz, S.L. Motor function following developmental exposure to PCBS and/or MEHG. Neurotoxicol. Teratol. 2006, 28, 260–277. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maurice, C.; Dalvai, M.; Lambrot, R.; Deschênes, A.; Scott-Boyer, M.-P.; McGraw, S.; Chan, D.; Côté, N.; Ziv-Gal, A.; Flaws, J.A.; et al. Early-Life Exposure to Environmental Contaminants Perturbs the Sperm Epigenome and Induces Negative Pregnancy Outcomes for Three Generations via the Paternal Lineage. Epigenomes 2021, 5, 10. https://doi.org/10.3390/epigenomes5020010
Maurice C, Dalvai M, Lambrot R, Deschênes A, Scott-Boyer M-P, McGraw S, Chan D, Côté N, Ziv-Gal A, Flaws JA, et al. Early-Life Exposure to Environmental Contaminants Perturbs the Sperm Epigenome and Induces Negative Pregnancy Outcomes for Three Generations via the Paternal Lineage. Epigenomes. 2021; 5(2):10. https://doi.org/10.3390/epigenomes5020010
Chicago/Turabian StyleMaurice, Clotilde, Mathieu Dalvai, Romain Lambrot, Astrid Deschênes, Marie-Pier Scott-Boyer, Serge McGraw, Donovan Chan, Nancy Côté, Ayelet Ziv-Gal, Jodi A. Flaws, and et al. 2021. "Early-Life Exposure to Environmental Contaminants Perturbs the Sperm Epigenome and Induces Negative Pregnancy Outcomes for Three Generations via the Paternal Lineage" Epigenomes 5, no. 2: 10. https://doi.org/10.3390/epigenomes5020010
APA StyleMaurice, C., Dalvai, M., Lambrot, R., Deschênes, A., Scott-Boyer, M.-P., McGraw, S., Chan, D., Côté, N., Ziv-Gal, A., Flaws, J. A., Droit, A., Trasler, J., Kimmins, S., & Bailey, J. L. (2021). Early-Life Exposure to Environmental Contaminants Perturbs the Sperm Epigenome and Induces Negative Pregnancy Outcomes for Three Generations via the Paternal Lineage. Epigenomes, 5(2), 10. https://doi.org/10.3390/epigenomes5020010






