On the Relationships between LncRNAs and Other Orchestrating Regulators: Role of the Circadian System
Abstract
1. Introduction
2. Orchestrating Regulators: Melatonin, Sirtuins and the Circadian Oscillator System
3. Cycling LncRNAs
4. Functional Diversity of Cycling LncRNAs and Their Regulators
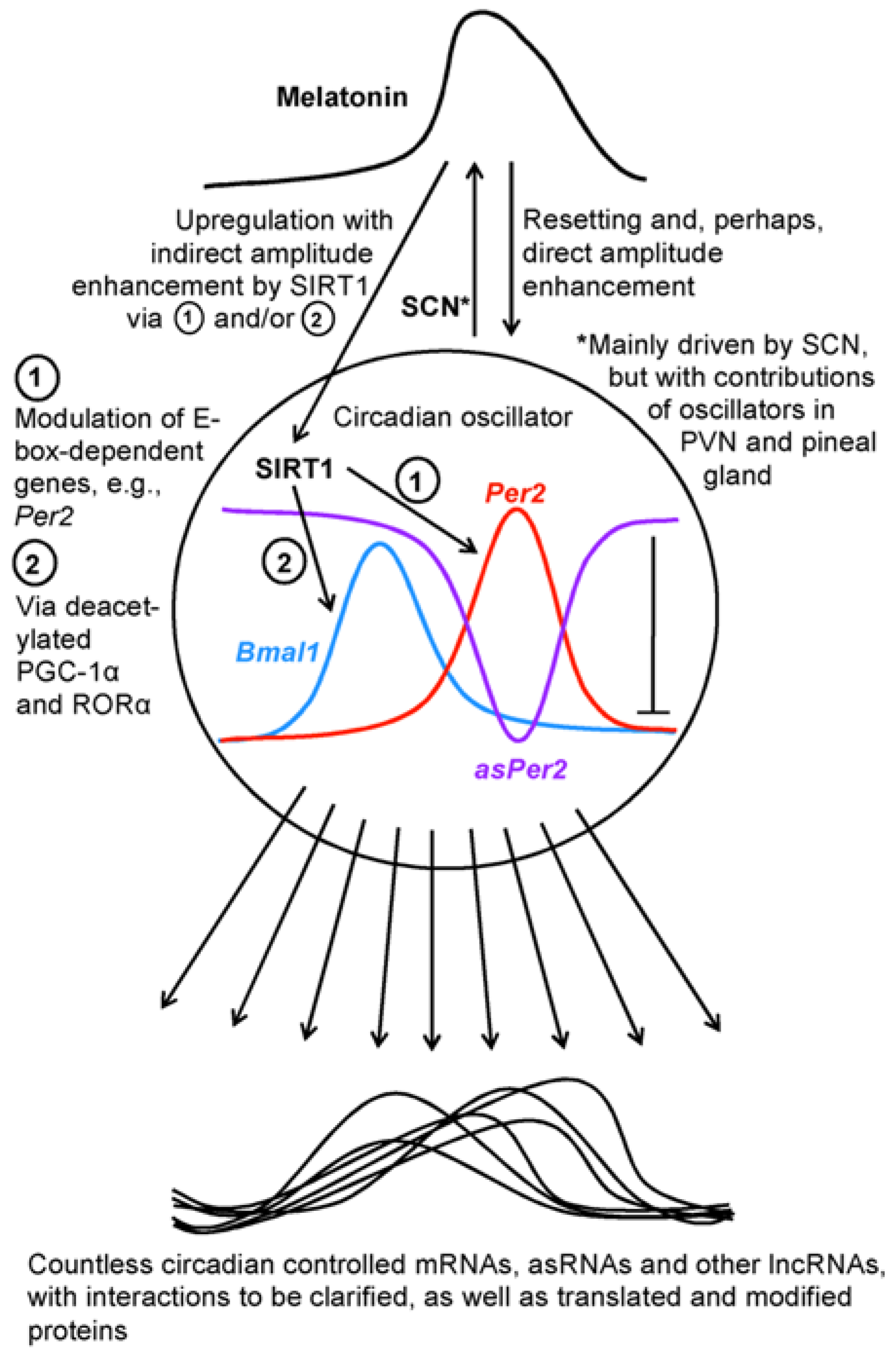
5. Sirtuin 1 as a Downstream Factor of Melatonin and a Circadian Regulator
6. Conclusions
Conflicts of Interest
References
- Noh, J.H.; Kim, K.M.; McClusky, W.G.; Abdelmohsen, K.; Gorospe, M. Cytoplasmic functions of long noncoding RNAs. Wiley Interdiscip. Rev. RNA 2018, 9, e1471. [Google Scholar] [CrossRef] [PubMed]
- Ard, R.; Allshire, R.C.; Marquardt, S. Emerging properties and functional consequences of noncoding transcription. Genetics 2017, 207, 357–367. [Google Scholar] [PubMed]
- Pefanis, E.; Wang, J.; Rothschild, G.; Lim, J.; Kazadi, D.; Sun, J.; Federation, A.; Chao, J.; Elliott, O.; Liu, Z.P.; et al. RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity. Cell 2015, 161, 774–789. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Dominguez, J.R.; Knoll, M.; Gromatzky, A.A.; Lodish, H.F. The super-enhancer-derived alncRNA-EC7/Bloodlinc potentiates red blood cell development in trans. Cell Rep. 2017, 19, 2503–2514. [Google Scholar] [CrossRef] [PubMed]
- Soibam, B. Super-lncRNAs: Identification of lncRNAs that target super-enhancers via RNA:DNA:DNA triplet formation. RNA 2017, 23, 1729–1742. [Google Scholar] [CrossRef] [PubMed]
- Rashid, F.; Shah, A.; Shan, G. Long non-coding RNAs in the cytoplasm. Genom. Proteom. Bioinform. 2016, 14, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Ahadi, A.; Brennan, S.; Kennedy, P.J.; Hutvagner, G.; Tran, N. Long non-coding RNAs harboring miRNA seed regions are enriched in prostate cancer exosomes. Sci. Rep. 2016, 6, 24922. [Google Scholar] [CrossRef] [PubMed]
- Salviano-Silva, A.; Lobo-Alves, S.C.; Almeida, R.C.; Malheiros, D.; Petzl-Erler, M.L. Besides pathology: Long non-coding RNA in cell and tissue homeostasis. Noncoding RNA 2018, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Pereira Fernandes, D.; Bitar, M.; Jacobs, F.M.J.; Barry, G. Long non-coding RNAs in neuronal aging. Noncoding RNA 2018, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R.; Madrid, J.A.; Tan, D.X.; Reiter, R.J. Melatonin, the circadian multioscillator system and health: The need for detailed analyses of peripheral melatonin signaling. J. Pineal Res. 2012, 52, 139–166. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Melatonin and the pathologies of weakened or dysregulated circadian oscillators. J. Pineal Res. 2017, 62, e12377. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R.; Cardinali, D.P.; Srinivasan, V.; Spence, D.W.; Brown, G.M.; Pandi-Perumal, S.R. Melatonin—A pleiotropic, orchestrating regulator molecule. Prog. Neurobiol. 2011, 93, 350–384. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.; Son, G.H.; Kim, K. Adrenal peripheral oscillator in generating the circadian glucocorticoid rhythm. Ann. N. Y. Acad. Sci. 2011, 1220, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Spencer, R.L.; Chun, L.E.; Hartsock, M.J.; Woodruff, E.R. Glucocorticoid hormones are both a major circadian signal and major stress signal: How this shared signal contributes to a dynamic relationship between the circadian and stress systems. Front. Neuroendocrinol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Moreira, A.C.; Antonini, S.R.R.; De Castro, M. Mechanisms in endocrinology: A sense of time of the glucocorticoid circadian clock: From the ontogeny to the diagnosis of Cushing’s syndrome. Eur. J. Endocrinol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Brain inflammaging: Roles of melatonin, circadian clocks and sirtuins. J. Clin. Cell. Immunol. 2018, 9, 543. [Google Scholar] [CrossRef]
- Sahar, S.; Sassone-Corsi, P. The epigenetic language of circadian clocks. Handb. Exp. Pharmacol. 2013, 217, 29–44. [Google Scholar]
- Chang, H.C.; Guarente, L. SIRT1 mediates central circadian control in the SCN by a mechanism that decays with aging. Cell 2013, 153, 1448–1460. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.; Becquet, D.; Franc, J.L.; François-Bellan, A.M. Circadian processes in the RNA life cycle. Wiley Interdiscip. Rev. RNA 2018, 9, e1467. [Google Scholar] [CrossRef] [PubMed]
- Honma, S. The mammalian circadian system: A hierarchical multi-oscillator structure for generating circadian rhythm. J. Physiol. Sci. 2018, 68, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Melatonin, noncoding RNAs, messenger RNA stability and epigenetics—Evidence, hints, gaps and perspectives. Int. J. Mol. Sci. 2014, 15, 18221–18252. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.M.; McIntyre, R.S.; Rosenblat, J.; Hardeland, R. Depressive disorders: Processes leading to neurodegeneration and potential novel treatments. Prog. Neuropsychopharmacol. Biol. Psychiatry 2018, 80, 189–204. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Circadian disruption, sleep loss, and low-grade inflammation. Res. Rev. Healthc. Open Access J. 2018, 1, 000109. [Google Scholar]
- Coon, S.L.; Munson, P.J.; Cherukuri, P.F.; Sugden, D.; Rath, M.F.; Møller, M.; Clokie, S.J.H.; Fu, C.; Olanich, M.E.; Rangel, Z.; et al. Circadian changes in long noncoding RNAs in the pineal gland. Proc. Natl. Acad. Sci. USA 2012, 109, 13319–13324. [Google Scholar] [CrossRef] [PubMed]
- Vollmers, C.; Schmitz, R.J.; Nathanson, J.; Yeo, G.; Ecker, J.R.; Panda, S. Circadian oscillations of protein-coding and regulatory RNAs in a highly dynamic mammalian liver epigenome. Cell Metab. 2012, 16, 833–845. [Google Scholar] [CrossRef] [PubMed]
- Mustafi, D.; Kevany, B.M.; Genoud, C.; Bai, X.; Palczewski, K. Photoreceptor phagocytosis is mediated by phosphoinositide signaling. FASEB J. 2013, 27, 4585–4595. [Google Scholar] [CrossRef] [PubMed]
- Powell, W.T.; Coulson, R.L.; Crary, F.K.; Wong, S.S.; Ach, R.A.; Tsang, P.; Yamada, N.A.; Yasui, D.H.; LaSalle, J.M. A Prader-Willis locus lncRNA cloud modulates diurnal genes and energy expenditure. Hum. Mol. Genet. 2013, 22, 4318–4328. [Google Scholar] [CrossRef] [PubMed]
- Fan, Z.; Zhao, M.; Joshi, P.D.; Li, P.; Zhang, Y.; Guo, W.; Xu, Y.; Wang, H.; Zhao, Z.; Yan, J. A class of circadian long non-coding RNAs mark enhancers modulating long-range circadian gene regulation. Nucleic Acids Res. 2017, 45, 5720–5738. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Du, G.; Song, X.; Li, L. Non-coding transcripts from enhancers: New insights into enhancer activity and gene expression regulation. Genom. Proteom. Bioinform. 2017, 15, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Battogtokh, D.; Kojima, S.; Tyson, J.J. Modeling the interactions of sense and antisense Period transcripts in the mammalian circadian clock network. PLoS Comput. Biol. 2018, 14, e1005957. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Liu, Z.; Song, R. Antisense lncRNA As-SLC7A11 suppresses epithelial ovarian cancer progression mainly by targeting SLC7A11. Pharmazie 2017, 72, 402–407. [Google Scholar] [PubMed]
- Zhao, Y.; Liu, Y.; Lin, L.; Huang, Q.; He, W.; Zhang, S.; Dong, S.; Wen, Z.; Rao, J.; Liao, W.; et al. The lncRNA MACC1-AS1 promotes gastric cancer cell metabolic plasticity via AMPK/Lin28 mediated mRNA stability of MACC1. Mol. Cancer 2018, 17, 69. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.Q.; Wang, Y.; Xiong, Y.; Chen, X.C.; Ma, M.L.; Cai, R.; Gao, Y.; Sun, Y.M.; Yang, G.S.; Pang, W.J. Sirt1 AS lncRNA interacts with its mRNA to inhibit muscle formation by attenuating function of miR-34a. Sci. Rep. 2016, 6, 21865. [Google Scholar] [CrossRef] [PubMed]
- Rapicavoli, N.A.; Qu, K.; Zhang, J.; Mikhail, M.; Laberge, R.M.; Chang, H.Y. A mammalian pseudogene lncRNA at the interface of inflammation and anti-inflammatory therapeutics. Elife 2013, 2, e00762. [Google Scholar] [CrossRef] [PubMed]
- Korkmaz, A.; Rosales-Corral, S.; Reiter, R.J. Gene regulation by melatonin linked to epigenetic phenomena. Gene 2012, 503, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Nopparat, C.; Sinjanakhom, P.; Govitrapong, P. Melatonin reverses H2O2-induced senescence in SH-SY5Y cells by enhancing autophagy via sirtuin 1 deacetylation of the RelA/p65 subunit of NF-κB. J. Pineal Res. 2017, 63, e12407. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Sun, J.; Li, T.; Zhang, Q.; Bu, S.; Wang, Q.; Lai, D. Melatonin ameliorates restraint stress-induced oxidative stress and apoptosis in testicular cells via NF-κB/iNOS and Nrf2/HO-1 signaling pathway. Sci. Rep. 2017, 7, 9599. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Jiang, Z.; Li, X.; Zhang, X. SIRT1 overexpression protects non-small cell lung cancer cells against osteopontin-induced epithelial-mesenchymal transition by suppressing NF-κB signaling. OncoTargets Ther. 2018, 11, 1157–1171. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Guang, H.; Zhang, H.; Chen, D.; Ding, L.; Fan, X.; Xue, F.; Gan, Z.; Wang, Y.; Mao, S.; et al. SIRT1 mediates apelin-13 in ameliorating chronic normobaric hypoxia-induced anxiety-like behavior by suppressing NF-κB pathway in mice hippocampus. Neuroscience 2018, 381, 22–34. [Google Scholar] [CrossRef] [PubMed]
- Su, S.C.; Reiter, R.J.; Hsiao, H.Y.; Chung, W.H.; Yang, S.F. Functional interaction between melatonin signaling and noncoding RNAs. Trends Endocrinol. Metab. 2018. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Wang, X.; Geng, P.; Tang, X.; Xiang, L.; Lu, X.; Li, J.; Ruan, Z.; Chen, J.; Xie, G.; et al. Melatonin regulates PARP1 to control the senescence-associated secretory phenotype (SASP) in human fetal lung fibroblast cells. J. Pineal Res. 2017, 63, e12405. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, X.; Bai, X.; Lin, Y.; Li, Z.; Fu, J.; Li, M.; Zhao, T.; Yang, H.; Xu, R.; et al. Melatonin prevents endothelial cell pyroptosis via regulation of long noncoding RNA MEG3/miR-223/NLRP3 axis. J. Pineal Res. 2018, 64, e12449. [Google Scholar] [CrossRef] [PubMed]
- Cai, B.; Ma, W.; Bi, C.; Yang, F.; Zhang, L.; Han, Z.; Huang, Q.; Ding, F.; Li, Y.; Yan, G.; et al. Long noncoding RNA H19 mediates melatonin inhibition of premature senescence of c-kit+ cardiac progenitor cells by promoting miR-675. J. Pineal Res. 2016, 61, 82–95. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.H.; Wu, C.H.; Yeh, C.T.; Su, S.C.; Hsia, S.M.; Liang, K.H.; Chen, C.C.; Hsueh, C.; Chen, C.Y. Melatonin suppresses hepatocellular carcinoma progression via lncRNA-CPS1-IT-mediated HIF-1α inactivation. Oncotarget 2017, 8, 82280–82293. [Google Scholar] [PubMed]
- Zhao, L.; An, R.; Yang, Y.; Yang, X.; Liu, H.; Yue, L.; Li, X.; Lin, Y.; Reiter, R.J.; Qu, Y. Melatonin alleviates brain injury in mice subjected to cecal ligation and puncture via attenuating inflammation, apoptosis, and oxidative stress: The role of SIRT1 signaling. J. Pineal Res. 2015, 59, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Jiang, S.; Dong, Y.; Fan, C.; Zhao, L.; Yang, X.; Li, J.; Di, S.; Yue, L.; Liang, G.; et al. Melatonin prevents cell death and mitochondrial dysfunction via a SIRT1-dependent mechanism during ischemic-stroke in mice. J. Pineal Res. 2015, 58, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Carloni, S.; Favrais, G.; Saliba, E.; Albertini, M.C.; Chalon, S.; Longini, M.; Gressens, P.; Buonocore, G.; Balduini, W. Melatonin modulates neonatal brain inflammation through ER stress, autophagy and miR-34a/SIRT1 pathway. J. Pineal Res. 2016, 61, 370–380. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Liu, H.; Yue, L.; Zhang, J.; Li, X.; Wang, B.; Lin, Y.; Qu, Y. Melatonin attenuates early brain injury via the melatonin receptor/Sirt1/NF-κB signaling pathway following subarachnoid hemorrhage in mice. Mol. Neurobiol. 2017, 54, 1612–1621. [Google Scholar] [CrossRef] [PubMed]
- Jenwitheesuk, A.; Boontem, P.; Wongchitrat, P.; Tocharus, J.; Mukda, S.; Govitrapong, P. Melatonin regulates the aging mouse hippocampal homeostasis via the sirtuin1-FOXO1 pathway. EXCLI J. 2017, 16, 340–353. [Google Scholar] [PubMed]
- Yang, W.; Kang, X.; Qin, N.; Li, F.; Jin, X.; Ma, Z.; Qian, Z.; Wu, S. Melatonin protects chondrocytes from impairment induced by glucocorticoids via NAD+-dependent SIRT1. Steroids 2017, 126, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Liu, Y.; Zhuang, H.; Yang, B.; Hei, K.; Xiao, M.; Hou, C.; Gao, H.; Zhang, X.; Jia, C.; et al. Quantitative proteomics reveals that long non-coding RNA MALAT1 interacts with DBC1 to regulate p53 acetylation. Nucleic Acids Res. 2017, 45, 9947–9959. [Google Scholar] [CrossRef] [PubMed]
- Chini, C.C.; Escande, C.; Nin, V.; Chini, E.N. DBC1 (Deleted in Breast Cancer 1) modulates the stability and function of the nuclear receptor Rev-erbα. Biochem. J. 2013, 451, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Giguère, S.S.; Guise, A.J.; Jean Beltran, P.M.; Joshi, P.M.; Greco, T.M.; Quach, O.L.; Kong, J.; Cristea, I.M. The proteomic profile of Deleted in Breast Cancer 1 (DBC1) interactions points to a multifaceted regulation of gene expression. Mol. Cell. Proteom. 2016, 15, 791–809. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Pang, W.J.; Wei, N.; Xiong, Y.; Wu, W.J.; Zhao, C.Z.; Shen, Q,W.; Yang, G.S. Identification, stability and expression of Sirt1 antisense long non-coding RNA. Gene 2014, 539, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Ming, G.F.; Wu, K.; Hu, K.; Chen, Y.; Xiao, J. NAMPT regulates senescence, proliferation, and migration of endothelial progenitor cells through the SIRT1 AS lncRNA/miR-22/SIRT1 pathway. Biochem. Biophys. Res. Commun. 2016, 478, 1382–1388. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.; Ni, Z.; He, J.; Jiang, S.; Li, X.; He, J.; Gong, W.; Zheng, L.; Chen, S.; Li, B.; et al. LncRNA HULC triggers autophagy via stabilizing Sirt1 and attenuates the chemosensitivity of HCC cells. Oncogene 2017, 36, 3528–3540. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.Q.; Gu, X.Y.; Hu, J.X.; Ping, Y.; Li, H.; Yan, J.Y.; Li, J.; Sun, R.; Yu, Z.J.; Zhang, Y. Hepatitis C virus core protein impairs metabolic disorder of liver cell via HOTAIR-Sirt1 signalling. Biosci. Rep. 2016, 36, e00336. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.; Li, Y.; Liu, J.; Ying, X.; Liu, Y.; Yan, J.; Chen, C.; Zhou, H.; Cao, L.; Ma, Y. LncRNA-mediated SIRT1/FoxO3a and SIRT1/p53 signaling pathways regulate type II alveolar epithelial cell senescence in patients with chronic obstructive pulmonary disease. Mol. Med. Rep. 2017, 15, 3129–3134. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wang, S.; Tang, C.; Chen, W. Upregulation of long non-coding RNA HIF 1α-anti-sense 1 induced by transforming growth factor-β-mediated targeting of sirtuin 1 promotes osteoblastic differentiation of human bone marrow stromal cells. Mol. Med. Rep. 2015, 12, 7233–7238. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, X.; Wu, X.; Dai, Y.; Chen, P.; Xie, L. MicroRNA-182 mediates Sirt1-induced diabetic corneal nerve regeneration. Diabetes 2016, 65, 2020–2031. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hardeland, R. On the Relationships between LncRNAs and Other Orchestrating Regulators: Role of the Circadian System. Epigenomes 2018, 2, 9. https://doi.org/10.3390/epigenomes2020009
Hardeland R. On the Relationships between LncRNAs and Other Orchestrating Regulators: Role of the Circadian System. Epigenomes. 2018; 2(2):9. https://doi.org/10.3390/epigenomes2020009
Chicago/Turabian StyleHardeland, Rüdiger. 2018. "On the Relationships between LncRNAs and Other Orchestrating Regulators: Role of the Circadian System" Epigenomes 2, no. 2: 9. https://doi.org/10.3390/epigenomes2020009
APA StyleHardeland, R. (2018). On the Relationships between LncRNAs and Other Orchestrating Regulators: Role of the Circadian System. Epigenomes, 2(2), 9. https://doi.org/10.3390/epigenomes2020009




