DNA Methylation Suppression by Bhendi Yellow Vein Mosaic Virus
Abstract
:1. Introduction
2. Results
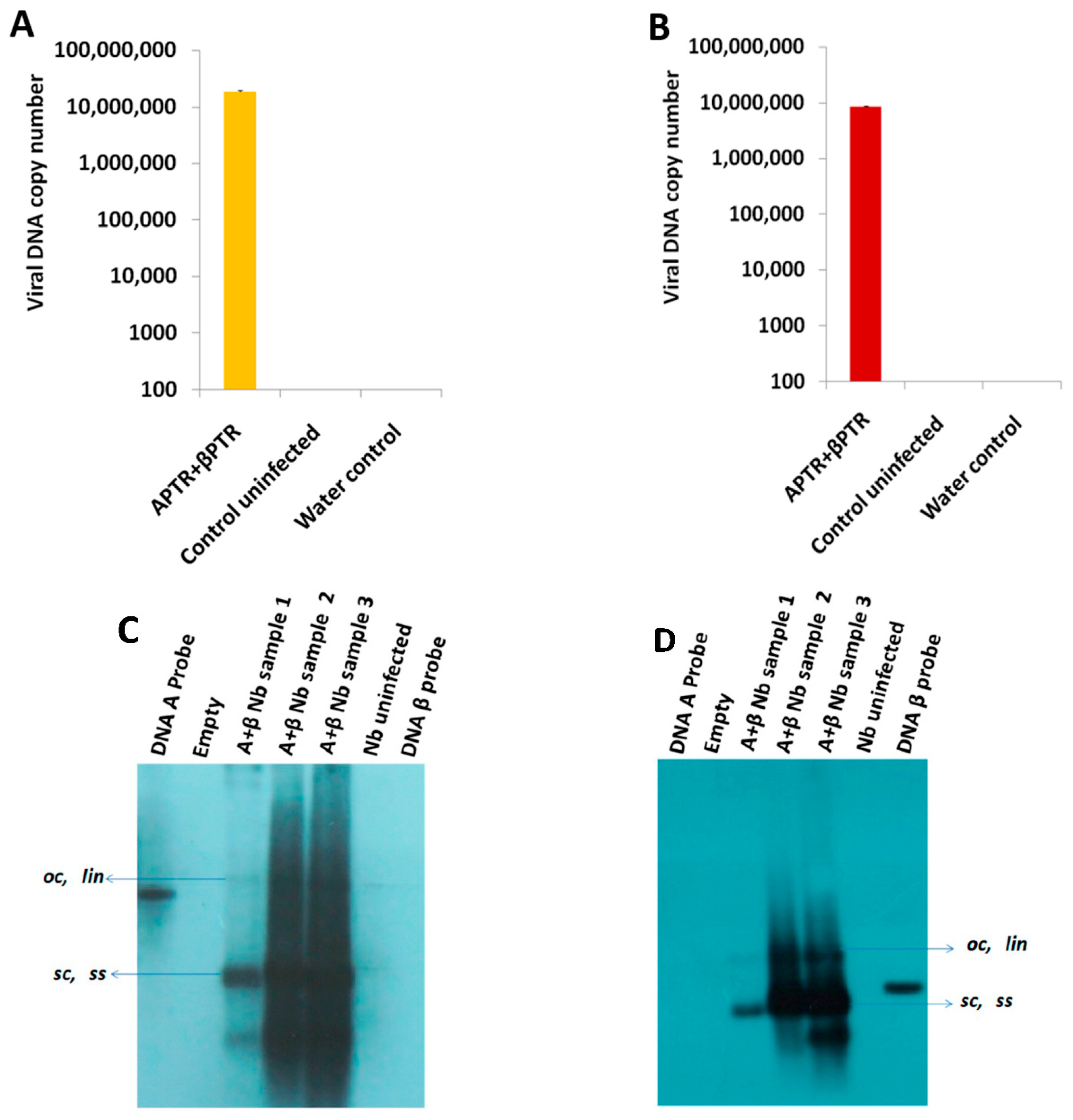
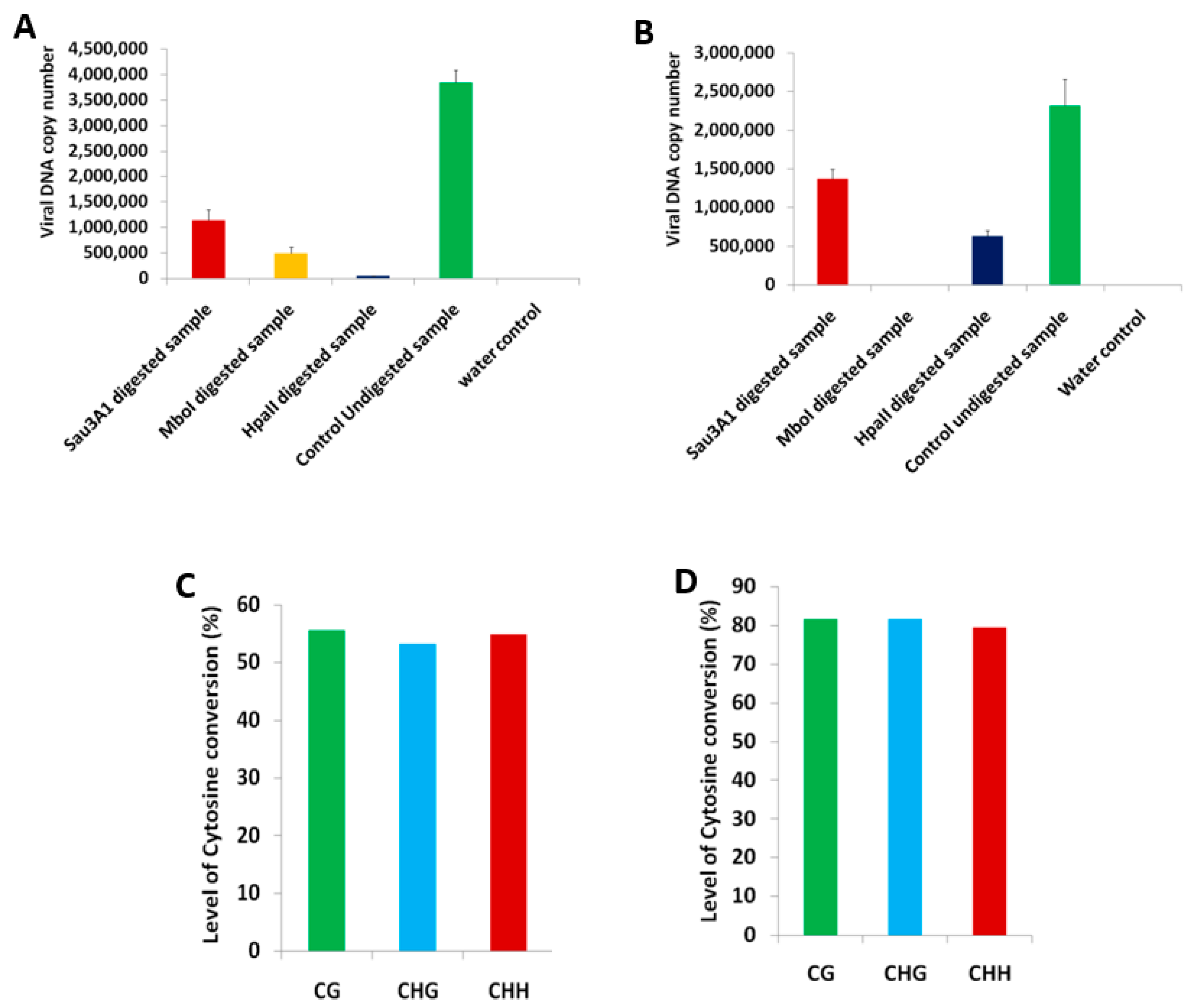
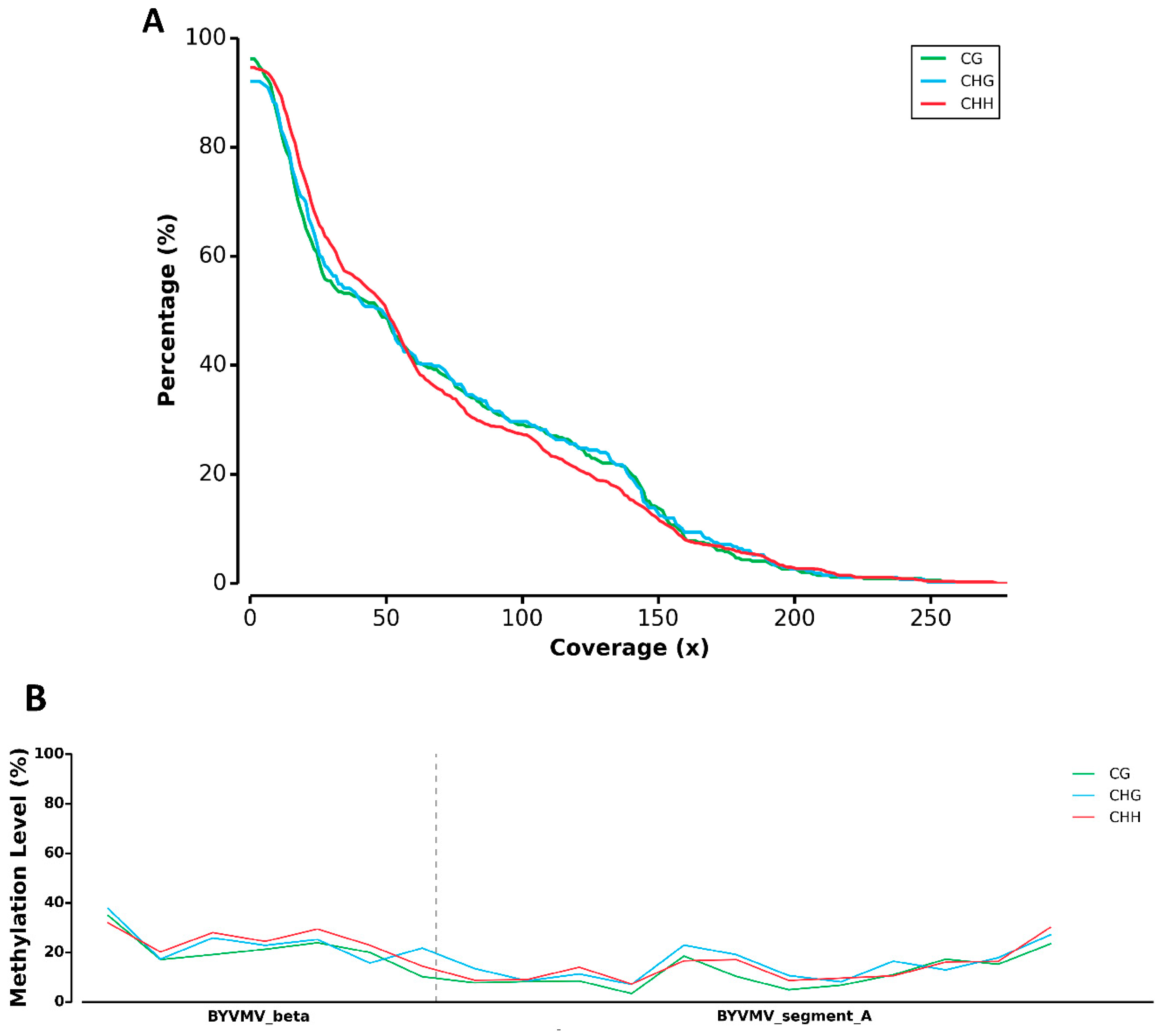
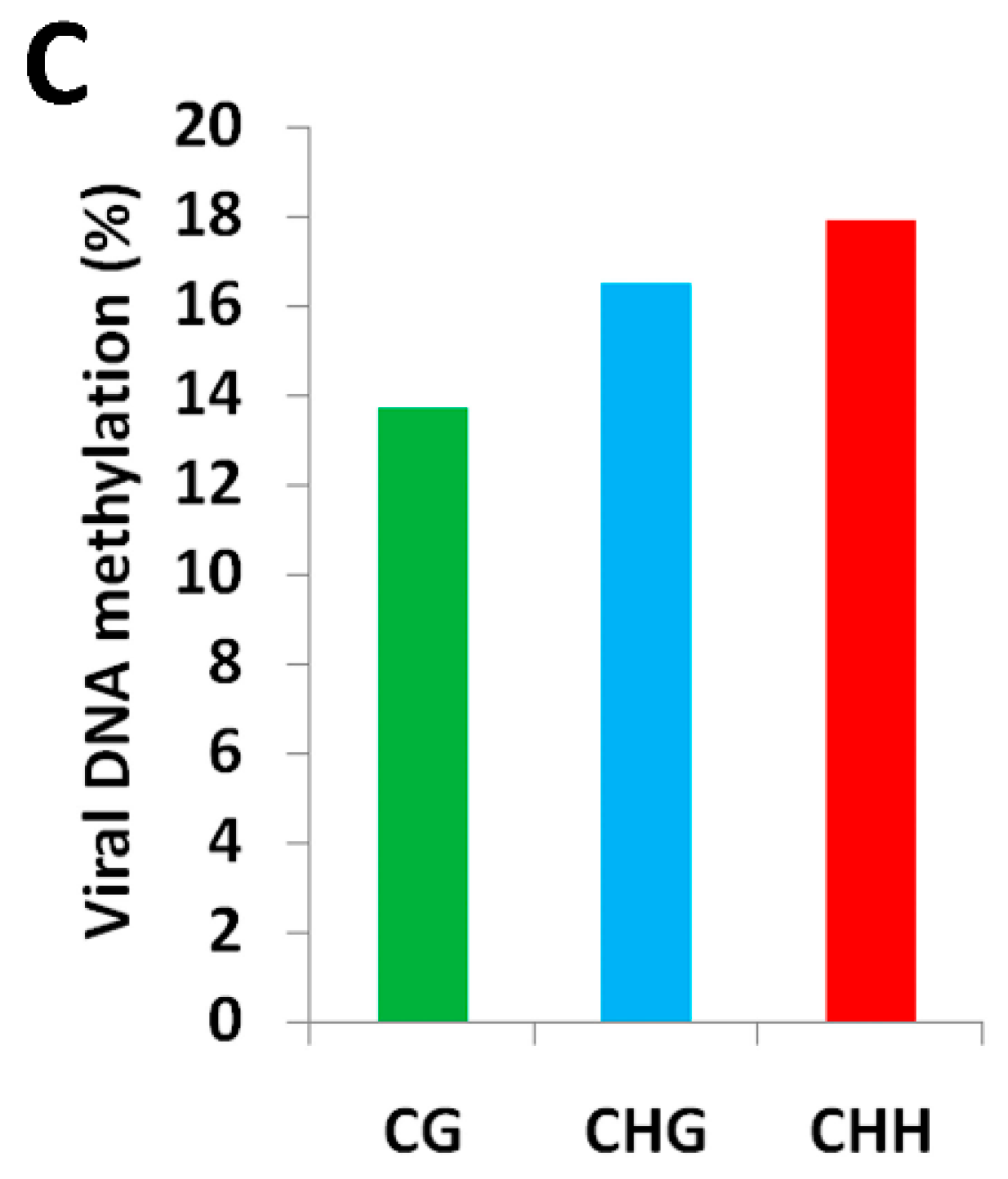
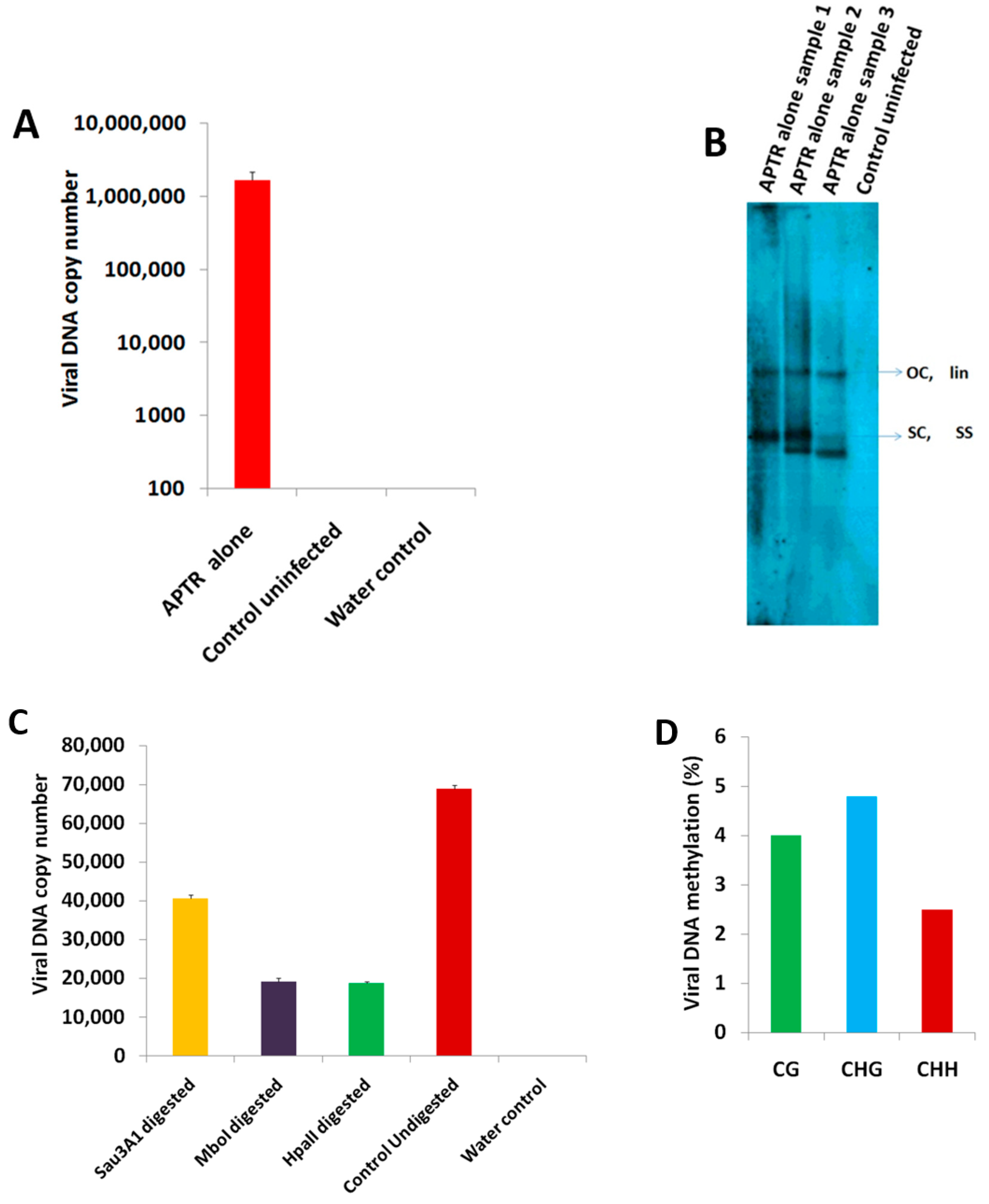
2.1. BYVMV Interferes with DNA Methylation
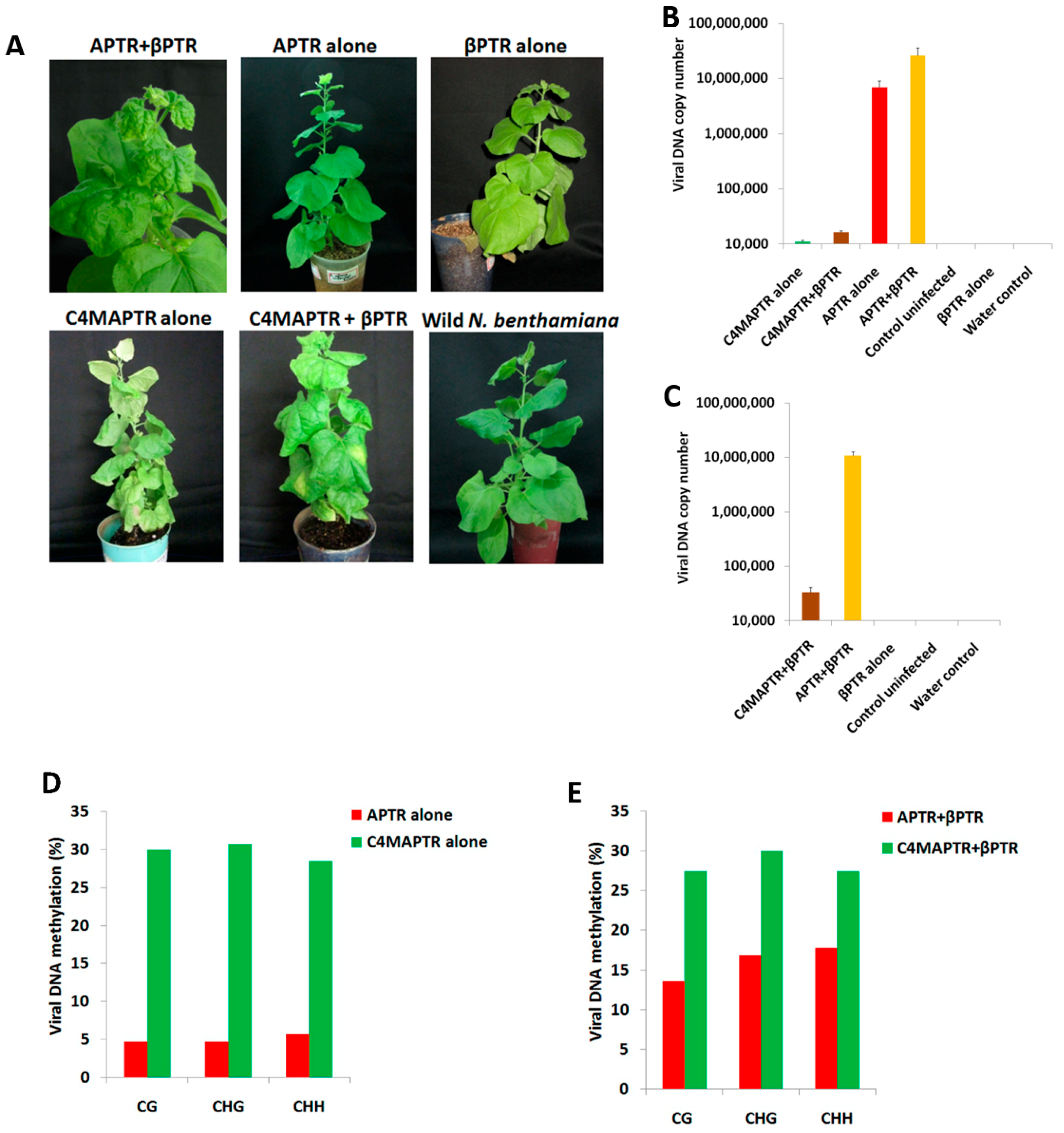
2.2. BYVMV Gene(s) Involved in Methylation Suppression
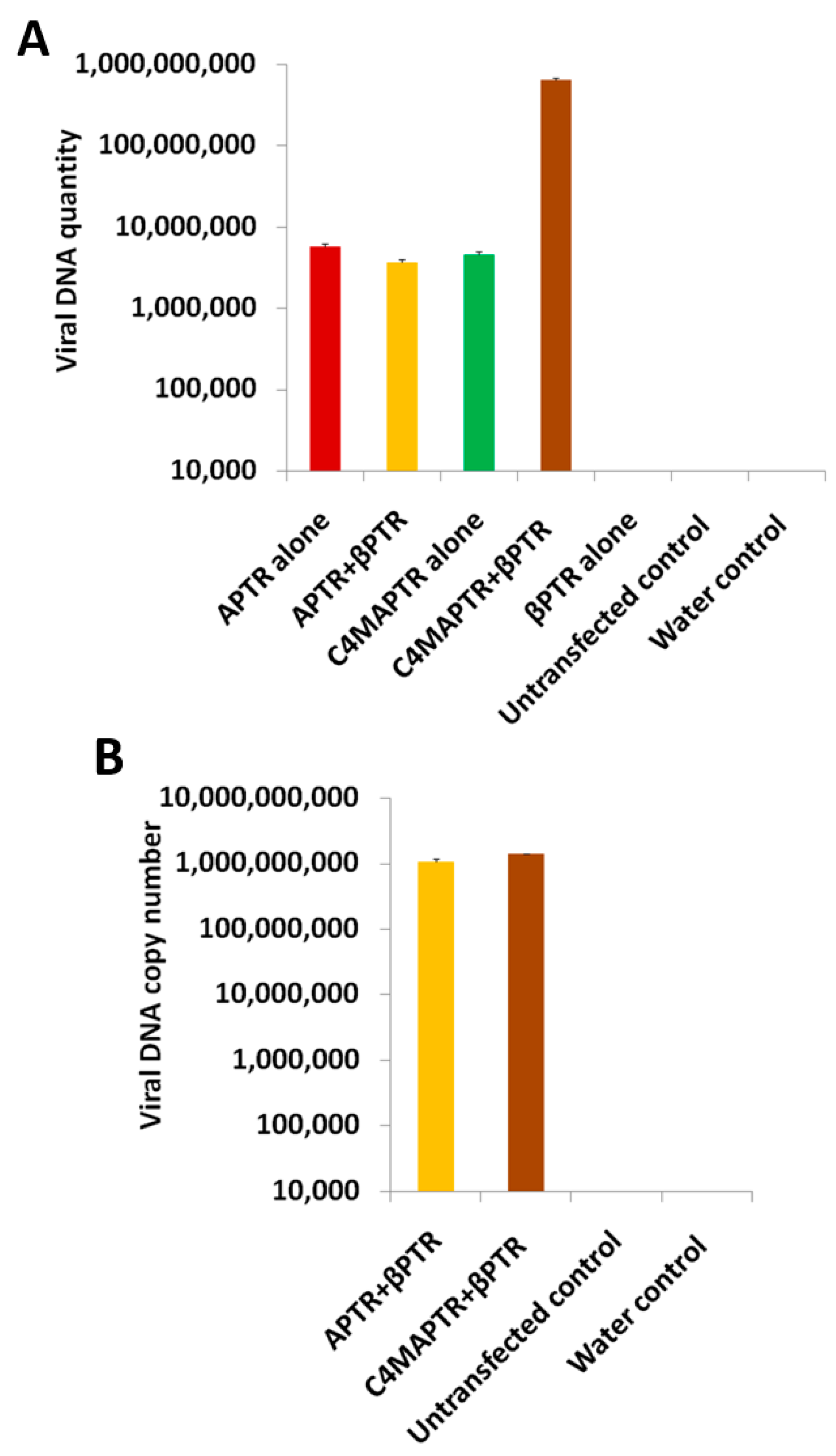
2.3. BYVMV C4 and C2 Hinders Transcriptional Gene Silencing
2.4. C4 Mutation Impairs Viral Movement
3. Discussion
4. Materials and Methods
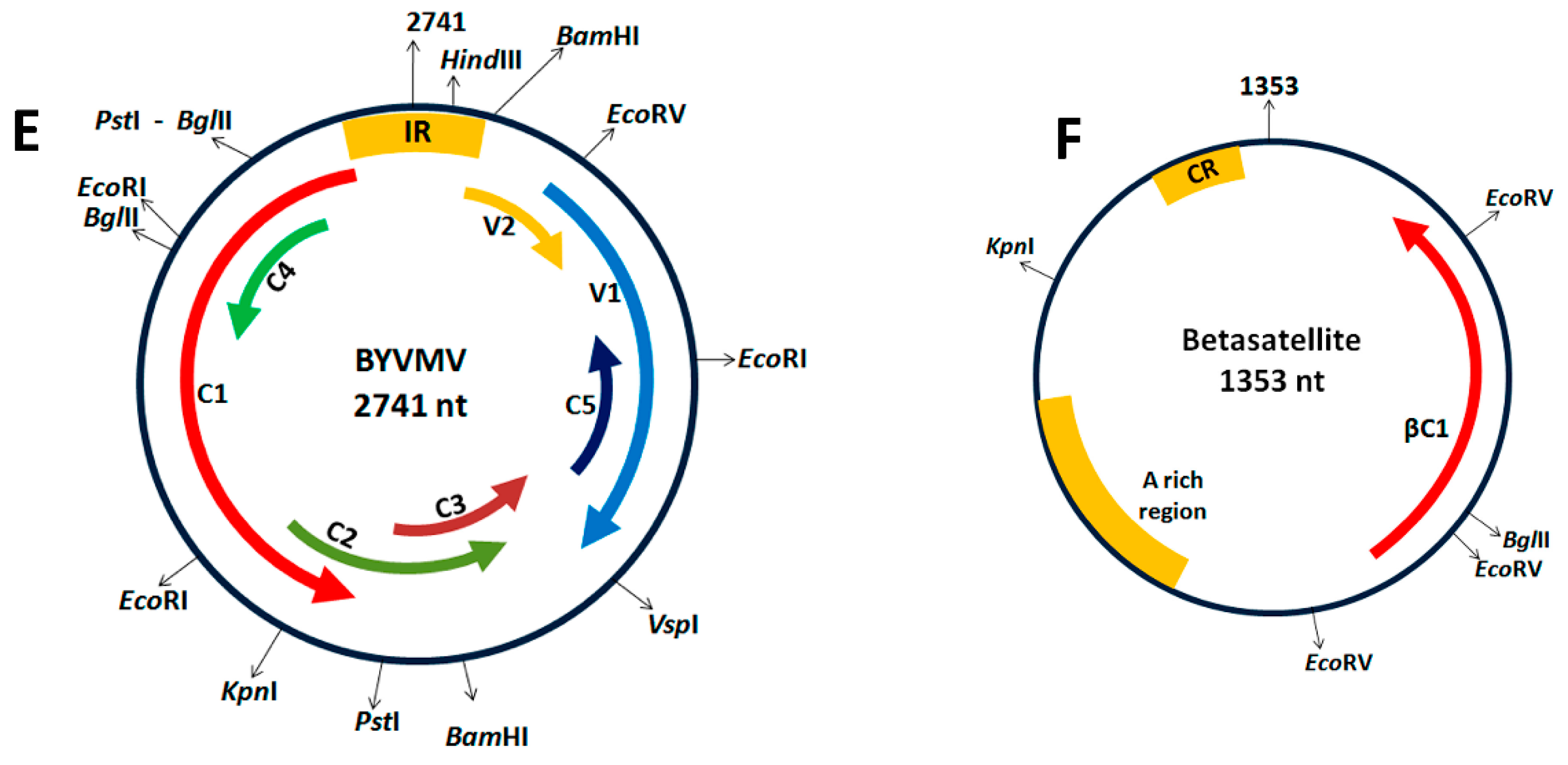
4.1. Cloning
4.2. Agroinfiltration
4.3. Viral DNA Isolation
4.4. Chop-PCR
4.5. Southern Hybridization
4.6. Quantitative PCR (qPCR)
4.7. Protoplast Isolation and Transfection
4.8. Bisulfite Conversion
4.9. RCA
4.10. DNA Processing for Illumina Sequencing and Library Preparation
4.11. Mapping Cytosine Methylation Using MethGo
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Varsani, A.; Roumagnac, P.; Fuchs, M.; Navas-Castillo, J.; Moriones, E.; Idris, A.; Briddon, R.W.; Rivera-Bustamante, R.; Murilo Zerbini, F.; Martin, D.P. Capulavirus and Grablovirus: Two new genera in the family Geminiviridae. Arch. Virol. 2017, 162, 1819–1831. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.K.; Zerbini, F.M.; Navas-Castillo, J.; Moriones, E.; Ramos-Sobrinho, R.; Silva, J.C.F.; Fiallo-Olivé, E.; Briddon, R.W.; Hernández-Zepeda, C.; Idris, A.; et al. Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch. Virol. 2015, 160, 1593–1619. [Google Scholar] [CrossRef] [PubMed]
- Hanley-Bowdoin, L.; Bejarano, E.R.; Robertson, D.; Mansoor, S. Geminiviruses: Masters at redirecting and reprogramming plant processes. Nat. Rev. Microbiol. 2013, 11, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Schneider, R.J.; Mohr, I. Translation initiation and viral tricks. Trends Biochem. Sci. 2003, 28, 130–136. [Google Scholar] [CrossRef]
- Pilartz, M.; Jeske, H. Mapping of abutilon mosaic geminivirus minichromosomes. J. Virol. 2003, 77, 10808–10818. [Google Scholar] [CrossRef] [PubMed]
- Velayudha Vimala Kumar, K.; Srikakulam, N.; Padbhanabhan, P.; Pandi, G. Deciphering microRNAs and Their Associated Hairpin Precursors in a Non-Model Plant, Abelmoschus esculentus. Non Coding RNA 2017, 3, 19. [Google Scholar] [CrossRef] [PubMed]
- Raja, P.; Sanville, B.C.; Buchmann, R.C.; Bisaro, D.M. Viral Genome Methylation as an Epigenetic Defense against Geminiviruses. J. Virol. 2008, 82, 8997–9007. [Google Scholar] [CrossRef] [PubMed]
- Ermak, G.; Paszkowski, U.; Wohlmuth, M.; Scheid, O.M.; Paszkowski, J. Cytosine methylation inhibits replication of African cassava mosaic virus by two distinct mechanisms. Nucleic Acids Res. 1993, 21, 3445–3450. [Google Scholar] [CrossRef] [PubMed]
- Raja, P.; Wolf, J.N.; Bisaro, D.M. RNA silencing directed against geminiviruses: Post-transcriptional and epigenetic components. Biochim. Biophys. Acta BBA Gene Regul. Mech. 2010, 1799, 337–351. [Google Scholar] [CrossRef] [PubMed]
- Ashapkin, V.V.; Kutueva, L.I.; Vanyushin, B.F. Plant DNA Methyltransferase Genes: Multiplicity, Expression, Methylation Patterns. Biochem. Biokhimiia 2016, 81, 141–151. [Google Scholar] [CrossRef] [PubMed]
- Mitsudome, T.; Mon, H.; Xu, J.; Li, Z.; Lee, J.M.; Patil, A.A.; Masuda, A.; Iiyama, K.; Morokuma, D.; Kusakabe, T. Biochemical characterization of maintenance DNA methyltransferase DNMT-1 from silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 2015, 58, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Pavlopoulou, A.; Kossida, S. Plant cytosine-5 DNA methyltransferases: Structure, function, and molecular evolution. Genomics 2007, 90, 530–541. [Google Scholar] [CrossRef] [PubMed]
- Hashimshony, T.; Zhang, J.; Keshet, I.; Bustin, M.; Cedar, H. The role of DNA methylation in setting up chromatin structure during development. Nat. Genet. 2003, 34, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Severin, P.M.D.; Zou, X.; Gaub, H.E.; Schulten, K. Cytosine methylation alters DNA mechanical properties. Nucleic Acids Res. 2011, 39, 8740–8751. [Google Scholar] [CrossRef] [PubMed]
- Bisaro, D.M. Silencing suppression by geminivirus proteins. Virology 2006, 344, 158–168. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Negrete, E.; Lozano-Durán, R.; Piedra-Aguilera, A.; Cruzado, L.; Bejarano, E.R.; Castillo, A.G. Geminivirus Rep protein interferes with the plant DNA methylation machinery and suppresses transcriptional gene silencing. New Phytol. 2013, 199, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Li, F.; Huang, C.; Yang, X.; Qian, Y.; Xie, Y.; Zhou, X. V2 of tomato yellow leaf curl virus can suppress methylation-mediated transcriptional gene silencing in plants. J. Gen. Virol. 2014, 95, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xie, Y.; Raja, P.; Li, S.; Wolf, J.N.; Shen, Q.; Bisaro, D.M.; Zhou, X. Suppression of methylation-mediated transcriptional gene silencing by βC1-SAHH protein interaction during geminivirus-β satellite infection. PLoS Pathog. 2011, 7, e1002329. [Google Scholar] [CrossRef] [PubMed]
- Jose, J.; Usha, R. Bhendi yellow vein mosaic disease in India is caused by association of a DNA β satellite with a begomovirus. Virology 2003, 305, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Priyavathi, P.; Kavitha, V.; Gopal, P. Complex Nature of Infection Associated with Yellow Vein Mosaic Disease in Bhendi (Abelmoschus esculentus). Curr. Sci. 2016, 111, 1511–1515. [Google Scholar] [CrossRef]
- Chandran, S.A.; Levy, Y.; Mett, A.; Belausov, E.; Ramakrishnan, U.; Gafni, Y. Mapping of functional region conferring nuclear localization and karyopherin α-binding activity of the C2 protein of bhendi yellow vein mosaic virus. J. Gen. Virol. 2012, 93, 1367–1374. [Google Scholar] [CrossRef] [PubMed]
- Gopal, P.; Pravin Kumar, P.; Sinilal, B.; Jose, J.; Kasin Yadunandam, A.; Usha, R. Differential roles of C4 and βC1 in mediating suppression of post-transcriptional gene silencing: Evidence for transactivation by the C2 of Bhendi yellow vein mosaic virus, a monopartite begomovirus. Virus Res. 2007, 123, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Chandran, S.A.; Jeyabharathy, C.; Usha, R. The C2 protein of Bhendi yellow vein mosaic virus plays an important role in symptom determination and virus replication. Virus Genes 2014, 48, 203–207. [Google Scholar] [CrossRef] [PubMed]
- Rigden, J.E.; Krake, L.R.; Rezaian, M.A.; Dry, I.B. ORF C4 of tomato leaf curl geminivirus is a determinant of symptom severity. Virology 1994, 204, 847–850. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Liu, Z.; Song, F.; Xie, Q.; Hanley-Bowdoin, L.; Zhou, X. Tomato SlSnRK1 protein interacts with and phosphorylates βC1, a pathogenesis protein encoded by a geminivirus β-satellite. Plant Physiol. 2011, 157, 1394–1406. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.; Behjatnia, S.A.A.; Mansoor, S.; Zafar, Y.; Hasnain, S.; Rezaian, M.A. A single complementary-sense transcript of a geminiviral DNA β satellite is determinant of pathogenicity. Mol. Plant Microbe Interact. MPMI 2005, 18, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.-Y.; Iwasaki, M.; Machida, C.; Machida, Y.; Zhou, X.; Chua, N.-H. βC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev. 2008, 22, 2564–2577. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.-W.; Yen, M.-R.; Ju, E.; Hsu, F.-M.; Lam, L.; Chen, P.-Y. MethGo: A comprehensive tool for analyzing whole-genome bisulfite sequencing data. BMC Genom. 2015, 16 (Suppl. 12), S11. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Zhao, L.; Jiao, X.; Jiang, T.; Gong, H.; Wang, B.; Briddon, R.W.; Zhou, X. A recombinant begomovirus resulting from exchange of the C4 gene. J. Gen. Virol. 2013, 94, 1896–1907. [Google Scholar] [CrossRef] [PubMed]
- Vanitharani, R.; Chellappan, P.; Pita, J.S.; Fauquet, C.M. Differential roles of AC2 and AC4 of cassava geminiviruses in mediating synergism and suppression of posttranscriptional gene silencing. J. Virol. 2004, 78, 9487–9498. [Google Scholar] [CrossRef] [PubMed]
- Llave, C.; Kasschau, K.D.; Carrington, J.C. Virus-encoded suppressor of posttranscriptional gene silencing targets a maintenance step in the silencing pathway. Proc. Natl. Acad. Sci. USA 2000, 97, 13401–13406. [Google Scholar] [CrossRef] [PubMed]
- Vanitharani, R.; Chellappan, P.; Fauquet, C.M. Geminiviruses and RNA silencing. Trends Plant Sci. 2005, 10, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.-B.; Wu, Q.; Ito, T.; Cillo, F.; Li, W.-X.; Chen, X.; Yu, J.-L.; Ding, S.-W. RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hao, L.; Shung, C.-Y.; Sunter, G.; Bisaro, D.M. Adenosine Kinase Is Inactivated by Geminivirus AL2 and L2 Proteins. Plant Cell 2003, 15, 3020–3032. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chen, H.; Huang, X.; Xia, R.; Zhao, Q.; Lai, J.; Teng, K.; Li, Y.; Liang, L.; Du, Q.; et al. BSCTV C2 Attenuates the Degradation of SAMDC1 to Suppress DNA Methylation-Mediated Gene Silencing in Arabidopsis. Plant Cell Online 2011, 23, 273–288. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Zhou, X. AC2 and AC4 proteins of Tomato yellow leaf curl China virus and Tobacco curly shoot virus mediate suppression of RNA silencing. Chin. Sci. Bull. 2004, 49, 2607–2612. [Google Scholar] [CrossRef]
- Stanley, J.; Latham, J.R.; Pinner, M.S.; Bedford, I.; Markham, P.G. Mutational analysis of the monopartite geminivirus beet curly top virus. Virology 1992, 191, 396–405. [Google Scholar] [CrossRef]
- Jupin, I.; De Kouchkovsky, F.; Jouanneau, F.; Gronenborn, B. Movement of tomato yellow leaf curl geminivirus (TYLCV): Involvement of the protein encoded by ORF C4. Virology 1994, 204, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Krake, L.R.; Rezaian, M.A.; Dry, I.B. Expression of the Tomato Leaf Curl Geminivirus C4 Gene Produces Viruslike Symptoms in Transgenic Plants. Mol. Plant. Microbe Interact. 1998, 11, 413–417. [Google Scholar] [CrossRef]
- Latham, J.R.; Saunders, K.; Pinner, M.S.; Stanley, J. Induction of plant cell division by beet curly top virus gene C4. Plant J. 1997, 11, 1273–1283. [Google Scholar] [CrossRef]
- Krichevsky, A.; Kozlovsky, S.V.; Gafni, Y.; Citovsky, V. Nuclear import and export of plant virus proteins and genomes. Mol. Plant Pathol. 2006, 7, 131–146. [Google Scholar] [CrossRef] [PubMed]
- Kunik, T.; Palanichelvam, K.; Czosnek, H.; Citovsky, V.; Gafni, Y. Nuclear import of the capsid protein of tomato yellow leaf curl virus (TYLCV) in plant and insect cells. Plant J. Cell Mol. Biol. 1998, 13, 393–399. [Google Scholar] [CrossRef]
- Palanichelvam, K.; Kunik, T.; Citovsky, V.; Gafni, Y. The capsid protein of tomato yellow leaf curl virus binds cooperatively to single-stranded DNA. J. Gen. Virol. 1998, 79 Pt 11, 2829–2833. [Google Scholar] [CrossRef] [PubMed]
- Usha, R.; Zrachya, A.; Levy, Y.; Spanov, H.; Gafni, Y. Protein–protein interactions and nuclear trafficking of coat protein and βC1 protein associated with Bhendi yellow vein mosaic disease. Virus Res. 2006, 122, 127–136. [Google Scholar] [CrossRef]
- Guha, D.; Poornima Priyadarshini, C.G.; Purakayastha, A.; Thippeswamy, R.; Lakshmikanth, M.; Savithri, H.S. Biochemical characterization of C4 protein of Cotton leaf curl Kokhran Virus-Dabawali. Biochim. Biophys. Acta 2013, 1830, 3734–3744. [Google Scholar] [CrossRef] [PubMed]
- Rojas, M.R.; Jiang, H.; Salati, R.; Xoconostle-Cázares, B.; Sudarshana, M.R.; Lucas, W.J.; Gilbertson, R.L. Functional analysis of proteins involved in movement of the monopartite begomovirus, Tomato yellow leaf curl virus. Virology 2001, 291, 110–125. [Google Scholar] [CrossRef] [PubMed]
- Buchmann, R.C.; Asad, S.; Wolf, J.N.; Mohannath, G.; Bisaro, D.M. Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. J. Virol. 2009, 83, 5005–5013. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Buckley, K.J.; Yang, X.; Buchmann, R.C.; Bisaro, D.M. Adenosine kinase inhibition and suppression of RNA silencing by geminivirus AL2 and L2 proteins. J. Virol. 2005, 79, 7410–7418. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Huang, C.; Li, Z.; Zhou, X. Suppression of RNA Silencing by a Plant DNA Virus Satellite Requires a Host Calmodulin-Like Protein to Repress RDR6 Expression. PLoS Pathog. 2014, 10, e1003921. [Google Scholar] [CrossRef] [PubMed]
- Vieira, J.; Messing, J. New pUC-derived cloning vectors with different selectable markers and DNA replication origins. Gene 1991, 100, 189–194. [Google Scholar] [PubMed]
- Ditta, G.; Stanfield, S.; Corbin, D.; Helinski, D.R. Broad host range DNA cloning system for gram-negative bacteria: Construction of a gene bank of Rhizobium meliloti. Proc. Natl. Acad. Sci. USA 1980, 77, 7347–7351. [Google Scholar] [CrossRef] [PubMed]
- Jose, J.; Usha, R. Extraction of geminiviral DNA from a highly mucilaginous plant (Abelmoschus esculentus). Plant Mol. Biol. Rep. 2000, 18, 349–355. [Google Scholar] [CrossRef]
- Hong, Y. Virus Resistance in Nicotiana benthamiana Conferred by African Cassava Mosaic Virus Replication-Associated Protein (ACI) Transgene. Mol. Plant. Microbe Interact. 1996, 9, 219–255. [Google Scholar] [CrossRef]
- Karthikeyan, C.; Patil, B.L.; Borah, B.K.; Resmi, T.R.; Turco, S.; Pooggin, M.M.; Hohn, T.; Veluthambi, K. Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus. Viruses 2016, 8, 264. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.-D.; Cho, Y.-H.; Sheen, J. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat. Protoc. 2007, 2, 1565–1572. [Google Scholar] [CrossRef] [PubMed]
- Paprotka, T.; Deuschle, K.; Metzler, V.; Jeske, H. Conformation-Selective Methylation of Geminivirus DNA. J. Virol. 2011, 85, 12001–12012. [Google Scholar] [CrossRef] [PubMed]
- Deuschle, K.; Kepp, G.; Jeske, H. Differential methylation of the circular DNA in geminiviral minichromosomes. Virology 2016, 499, 243–258. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Fiziev, P.; Yan, W.; Cokus, S.; Sun, X.; Zhang, M.Q.; Chen, P.-Y.; Pellegrini, M. BS-Seeker2: A versatile aligning pipeline for bisulfite sequencing data. BMC Genom. 2013, 14, 774. [Google Scholar] [CrossRef] [PubMed]



| S. No | Primers | Sequences | Target DNA & Nucleotide Coordinate | Details of Experimental Use |
|---|---|---|---|---|
| 1 | BYVMV DNA A IR Forward | 5′ GTTGACTATTTTTGAGACCCGATT 3′ | BYVMV DNA (1–24) | Chop-PCR |
| 2 | BYVMV DNA A IR Reverse | 5′ GGATCCCACATTTTTTGAATTG 3′ | BYVMV DNA A (285–264) | Chop-PCR |
| 3 | BglII BYVMV AV2 Forward | 5′ AGATCTATGTGGGATCCACTATTAAACG 3′ | BYVMV DNA A (119–131) | Chop-PCR |
| 4 | HindIII BYVMV AV2 Reverse | 5′ AAGCTTTCACATCCCCTTGGAACATCC 3′ | BYVMV DNA A (485–465) | Chop-PCR |
| 5 | HindIII BYVMV C2 Forward | 5′ AAGCTTATGCAGCATTCGTCTTTC 3′ | BYVMV DNA A (1584–1601) | Chop-PCR |
| 6 | BglII BYVMV C2 Reverse | 5′ AGATCTTTAGAGATATTTGAGGAC 3′ | BYVMV DNA A (1170–1190) | Chop-PCR |
| 7 | BamHI BYVMV AC4 Forward | 5′ GGATCCTTATCTTTAAGAACTCTAAGAGC 3′ | BYVMV DNA A (2130–2154) | Chop-PCR |
| 8 | HindIII BYVMV AC4 Reverse | 5′ AGCTTATGAAAATGGGGAACCTCATCTTCACG 3′ | BYVMV DNA A (2438–2412) | Chop-PCR |
| 9 | BglII BYVMV βC1 Forward | 5′ AGATCTAATTATTATCTTATTATCAATAGTAC 3′ | BYVMV DNA β (180–207) | Chop-PCR |
| 10 | HindIII BYVMV βC1 Reverse | 5′ AAGCTTATGAAAATATCTATACATTTCATC 3′ | BYVMV DNA β (602–578) | Chop-PCR |
| 11 | βC1 Promoter Forward | 5′ ATAATGTGTGGATGAAATTTATAGA 3′ | BYVMV DNA β 603–627 | Chop-PCR |
| 12 | βC1 Promoter Reverse | 5′ CTCATTTACCTATCGGTGTCTG 3′ | BYVMV DNA β 1080–1059 | Chop-PCR |
| 13 | BYVMV V2 RT Forward | 5′ GCAACTTTTGTCGCAGGATT 3′ | BYVMV DNA A (194–213) | Quantitative PCR |
| 14 | BYVMV V2 RT Reverse | 5′ ATAGGCCTGTTTGTCCATGC 3′ | BYVMV DNA A (406–425) | Quantitative PCR |
| 15 | BYVMV β RT Forward | 5′ CGCGCGGTAAATGGTAAATA 3′ | BYVMV DNA β (1017–1036) | Quantitative PCR |
| 16 | BYVMV β RT Reverse | 5′ CTACGACGCGCGATATAACT 3′ | BYVMV DNA β (1158–1177) | Quantitative PCR |
| 17 | C4 stop 1 Forward | 5′ ATT TTCCTTCGAACTGGATTAGCACGTGAAGATGAGGTT 3′ | BYVMV DNA A (2386–2426) | Site-specific mutagenesis |
| 18 | C4 stop 1 Reverse | 5′ AACCTCATCTTCACGTGCTAATCCAGTTCGAAGGAAAAT 3′ | BYVMV DNA A (2387–2426) | Site-specific mutagenesis |
| 19 | C2 stop 1 Forward | 5′ CCGAGTACAACGGTTGGGTTAGAAAGACGAATGCTGCAT 3′ | BYVMV DNA A 1563–1600 | Site-specific mutagenesis |
| 20 | C2 stop 1 Reverse | 5′ ATGCAGCATTCGTCTTTCTAACCCAACCGTTGTACTCGG 3′ | BYVMV DNA A 1600–1563 | Site-specific mutagenesis |
| 21 | C2 stop 2 Forward | 5′CGCTTCCTTGTGTTGAACTTAGATTGGTACCCGAGTACA 3′ | BYVMV DNA A 1533–1571 | Site-specific mutagenesis |
| 22 | C2 stop 2 Reverse | 5′ TGTACTCGGGTACCAATCTAAGTTCAACACAAGGAAGCG 3′ | BYVMV DNA A 1571–1533 | Site-specific mutagenesis |
| S.No | Type of Infiltration | Total No. of Plants Infiltrated | Severe Symptomatic Plants | Mild Symptomatic Plants | Asymptomatic |
|---|---|---|---|---|---|
| 1 | APTR alone | 22 | 0 | 13 | 9 |
| 2 | APTR + βPTR | 27 | 13 | 4 | 10 |
| 3 | C4MAPTR alone | 10 | 0 | 0 | 10 |
| 4 | C4MAPTR + βPTR | 10 | 0 | 0 | 10 |
| 5 | C2MAPTR alone | 10 | 0 | 0 | 10 |
| 6 | C2MAPTR + βPTR | 10 | 0 | 0 | 10 |
| 7 | βPTR alone | 14 | 0 | 0 | 14 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Babu, K.S.D.; Guria, A.; Karanthamalai, J.; Srikakulam, N.; Kumari, K.; Sharma, P.; Chandran, S.A.; Barnabas, A.D.; Tennyson, J.; Pandi, G. DNA Methylation Suppression by Bhendi Yellow Vein Mosaic Virus. Epigenomes 2018, 2, 7. https://doi.org/10.3390/epigenomes2020007
Babu KSD, Guria A, Karanthamalai J, Srikakulam N, Kumari K, Sharma P, Chandran SA, Barnabas AD, Tennyson J, Pandi G. DNA Methylation Suppression by Bhendi Yellow Vein Mosaic Virus. Epigenomes. 2018; 2(2):7. https://doi.org/10.3390/epigenomes2020007
Chicago/Turabian StyleBabu, Kandhalu Sagadevan Dinesh, Ashirbad Guria, Jeyalakshmi Karanthamalai, Nagesh Srikakulam, Kamlesh Kumari, Priyanka Sharma, Sam Aldrin Chandran, Anburaj Daniel Barnabas, Jebasingh Tennyson, and Gopal Pandi. 2018. "DNA Methylation Suppression by Bhendi Yellow Vein Mosaic Virus" Epigenomes 2, no. 2: 7. https://doi.org/10.3390/epigenomes2020007
APA StyleBabu, K. S. D., Guria, A., Karanthamalai, J., Srikakulam, N., Kumari, K., Sharma, P., Chandran, S. A., Barnabas, A. D., Tennyson, J., & Pandi, G. (2018). DNA Methylation Suppression by Bhendi Yellow Vein Mosaic Virus. Epigenomes, 2(2), 7. https://doi.org/10.3390/epigenomes2020007





