Temperature-Specific and Sex-Specific Fitness Effects of Sympatric Mitochondrial and Mito-Nuclear Variation in Drosophila obscura
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Desiccation Resistance
2.2. Egg-to-Pupa-to-Adult Viability, Developmental Time and Sex Ratio Experiment
2.3. Statistical Analysis
3. Results
3.1. Desiccation Resistance
3.2. Developmental Time
3.3. Viability
Egg-to-Adult
3.4. Proportion of Males
4. Discussion
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wolff, J.; Pichaud, N.; Camus, M.; Côté, G.; Blier, P.; Dowling, D. Evolutionary implications of mitochondrial genetic variation: Mitochondrial genetic effects on OXPHOS respiration and mitochondrial quantity change with age and sex in fruit flies. J. Evol. Biol. 2016, 29, 736–747. [Google Scholar] [CrossRef]
- Meiklejohn, C.D.; Holmbeck, M.A.; Siddiq, M.A.; Abt, D.N.; Rand, D.M.; Montooth, K.L. An incompatibility between a mitochondrial tRNA and its nuclear-encoded tRNA synthetase compromises development and fitness in Drosophila. PLoS Genet. 2013, 9, e1003238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ryan, M.T.; Hoogenraad, N.J. Mitochondrial-nuclear communications. Annu. Rev. Biochem. 2007, 76, 701–722. [Google Scholar] [CrossRef] [PubMed]
- Ellison, C.; Burton, R. Cytonuclear conflict in interpopulation hybrids: The role of RNA polymerase in mtDNA transcription and replication. J. Evol. Biol. 2010, 23, 528–538. [Google Scholar] [CrossRef] [PubMed]
- Gaspari, M.; Falkenberg, M.; Larsson, N.G.; Gustafsson, C.M. The mitochondrial RNA polymerase contributes critically to promoter specificity in mammalian cells. EMBO J. 2004, 23, 4606–4614. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chou, J.-Y.; Hung, Y.-S.; Lin, K.-H.; Lee, H.-Y.; Leu, J.-Y. Multiple molecular mechanisms cause reproductive isolation between three yeast species. PLoS Biol. 2010, 8, e1000432. [Google Scholar] [CrossRef] [Green Version]
- Ellison, C.K.; Burton, R.S. Genotype-dependent variation of mitochondrial transcriptional profiles in interpopulation hybrids. Proc. Natl. Acad. Sci. USA 2008, 105, 15831–15836. [Google Scholar] [CrossRef] [Green Version]
- Clayton, D.A. Transcription and replication of mitochondrial DNA. Hum. Reprod. 2000, 15, 11–17. [Google Scholar] [CrossRef]
- Falkenberg, M.; Larsson, N.-G.; Gustafsson, C.M. DNA replication and transcription in mammalian mitochondria. Annu. Rev. Biochem. 2007, 76, 679–699. [Google Scholar] [CrossRef]
- Burton, R.S.; Barreto, F.S. A disproportionate role for mt DNA in D obzhansky–M uller incompatibilities? Mol. Ecol. 2012, 21, 4942–4957. [Google Scholar] [CrossRef]
- Ballard, J.W.O.; Kreitman, M. Is mitochondrial DNA a strictly neutral marker? Trends Ecol. Evol. 1995, 10, 485–488. [Google Scholar] [CrossRef]
- Avise, J.C.; Arnold, J.; Ball, R.M.; Bermingham, E.; Lamb, T.; Neigel, J.E.; Reeb, C.A.; Saunders, N.C. Intraspecific phylogeography: The mitochondrial DNA bridge between population genetics and systematics. Annu. Rev. Ecol. Syst. 1987, 18, 489–522. [Google Scholar] [CrossRef]
- Charlesworth, B. The maintenance of genetic variation in life-history traits. Evol. Genet. Mol. Morphol. 2000, 369–392. [Google Scholar]
- Wolff, J.N.; Ladoukakis, E.D.; Enríquez, J.A.; Dowling, D.K. Mitonuclear interactions: Evolutionary consequences over multiple biological scales. Philos. Trans. R. Soc. B Biol. Sci. 2014, 369, 20130443. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rand, D.M.; Kann, L.M. Excess amino acid polymorphism in mitochondrial DNA: Contrasts among genes from Drosophila, mice, and humans. Mol. Biol. Evol. 1996, 13, 735–748. [Google Scholar] [CrossRef] [Green Version]
- Nachman, M.W.; Brown, W.M.; Stoneking, M.; Aquadro, C.F. Nonneutral mitochondrial DNA variation in humans and chimpanzees. Genetics 1996, 142, 953–963. [Google Scholar] [CrossRef] [PubMed]
- Pesole, G.; Gissi, C.; De Chirico, A.; Saccone, C. Nucleotide substitution rate of mammalian mitochondrial genomes. J. Mol. Evol. 1999, 48, 427–434. [Google Scholar] [CrossRef]
- Ruiz-Pesini, E.; Mishmar, D.; Brandon, M.; Procaccio, V.; Wallace, D.C. Effects of purifying and adaptive selection on regional variation in human mtDNA. Science 2004, 303, 223–226. [Google Scholar] [CrossRef] [Green Version]
- Bazin, E.; Glémin, S.; Galtier, N. Population size does not influence mitochondrial genetic diversity in animals. Science 2006, 312, 570–572. [Google Scholar] [CrossRef]
- Mishmar, D.; Ruiz-Pesini, E.; Golik, P.; Macaulay, V.; Clark, A.G.; Hosseini, S.; Brandon, M.; Easley, K.; Chen, E.; Brown, M.D. Natural selection shaped regional mtDNA variation in humans. Proc. Natl. Acad. Sci. USA 2003, 100, 171–176. [Google Scholar] [CrossRef] [Green Version]
- Grant, W.S.; Spies, I.B.; Canino, M.F. Biogeographic evidence for selection on mitochondrial DNA in North Pacific walleye pollock Theragra chalcogramma. J. Hered. 2006, 97, 571–580. [Google Scholar] [CrossRef] [PubMed]
- Dowling, D.K.; Abiega, K.C.; Arnqvist, G. Temperature-specific outcomes of cytoplasmic-nuclear interactions on egg-to-adult development time in seed beetles. Evolution 2007, 61, 194–201. [Google Scholar] [CrossRef]
- James, A.C.; Ballard, J.W.O. Mitochondrial genotype affects fitness in Drosophila simulans. Genetics 2003, 164, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Ballard, J.; Melvin, R. Linking the mitochondrial genotype to the organismal phenotype. Mol. Ecol. 2010, 19, 1523–1539. [Google Scholar] [CrossRef] [PubMed]
- Kurbalija Novičić, Z.; Immonen, E.; Jelić, M.; AnÐelković, M.; Stamenković-Radak, M.; Arnqvist, G. Within-population genetic effects of mt DNA on metabolic rate in D rosophila subobscura. J. Evol. Biol. 2015, 28, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Dowling, D.K.; Friberg, U.; Lindell, J. Evolutionary implications of non-neutral mitochondrial genetic variation. Trends Ecol. Evol. 2008, 23, 546–554. [Google Scholar] [CrossRef]
- Osada, N.; Akashi, H. Mitochondrial–nuclear interactions and accelerated compensatory evolution: Evidence from the primate cytochrome c oxidase complex. Mol. Biol. Evol. 2012, 29, 337–346. [Google Scholar] [CrossRef] [Green Version]
- Barreto, F.S.; Burton, R.S. Evidence for compensatory evolution of ribosomal proteins in response to rapid divergence of mitochondrial rRNA. Mol. Biol. Evol. 2012, 30, 310–314. [Google Scholar] [CrossRef] [Green Version]
- Howe, D.K.; Denver, D.R. Muller’s Ratchet and compensatory mutation in Caenorhabditis briggsae mitochondrial genome evolution. BMC Evol. Biol. 2008, 8, 62. [Google Scholar] [CrossRef] [Green Version]
- Filteau, M.; Hamel, V.; Pouliot, M.C.; Gagnon-Arsenault, I.; Dubé, A.K.; Landry, C.R. Evolutionary rescue by compensatory mutations is constrained by genomic and environmental backgrounds. Mol. Syst. Biol. 2015, 11, 832. [Google Scholar] [CrossRef]
- Hill, G.E. Mitonuclear compensatory coevolution. Trends Genet. 2020, 36, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.S.; Burton, R.S. Tracing hybrid incompatibilities to single amino acid substitutions. Mol. Biol. Evol. 2006, 23, 559–564. [Google Scholar] [CrossRef] [PubMed]
- Rand, D.M.; Haney, R.A.; Fry, A.J. Cytonuclear coevolution: The genomics of cooperation. Trends Ecol. Evol. 2004, 19, 645–653. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.G.; Lyckegaard, E. Natural selection with nuclear and cytoplasmic transmission. III. Joint analysis of segregation and mtDNA in Drosophila melanogaster. Genetics 1988, 118, 471–481. [Google Scholar] [CrossRef]
- Rand, D.M.; Clark, A.G.; Kann, L.M. Sexually antagonistic cytonuclear fitness interactions in Drosophila melanogaster. Genetics 2001, 159, 173–187. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.G. Natural selection with nuclear and cytoplasmic transmission. I. A deterministic model. Genetics 1984, 107, 679–701. [Google Scholar] [CrossRef]
- Clark, A.G. Natural selection with nuclear and cytoplasmic transmission. II. Tests with Drosophila from diverse populations. Genetics 1985, 111, 97–112. [Google Scholar] [CrossRef]
- Gregorius, H.; Ross, M. Selection with gene-cytoplasm interactions. I. Maintenance of cytoplasm polymorphisms. Genetics 1984, 107, 165–178. [Google Scholar] [CrossRef]
- Kojima, K.-I.; Tobari, Y.N. Selective modes associated with karyotypes in Drosophila ananassae. II. Heterosis and frequency-dependent selection. Genetics 1969, 63, 639. [Google Scholar] [CrossRef]
- Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2006, 2, e64. [Google Scholar] [CrossRef] [Green Version]
- Arnqvist, G.; Dowling, D.K.; Eady, P.; Gay, L.; Tregenza, T.; Tuda, M.; Hosken, D.J. Genetic architecture of metabolic rate: Environment specific epistasis between mitochondrial and nuclear genes in an insect. Evol. Int. J. Org. Evol. 2010, 64, 3354–3363. [Google Scholar] [CrossRef]
- Kazancıoğlu, E.; Arnqvist, G. The maintenance of mitochondrial genetic variation by negative frequency-dependent selection. Ecol. Lett. 2014, 17, 22–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novičić, Z.K.; Sayadi, A.; Jelić, M.; Arnqvist, G. Negative frequency dependent selection contributes to the maintenance of a global polymorphism in mitochondrial DNA. BMC Evol. Biol. 2020, 20, 20. [Google Scholar]
- Arnqvist, G.; Novičić, Z.K.; Castro, J.A.; Sayadi, A. Negative frequency dependent selection on sympatric mtDNA haplotypes in Drosophila subobscura. Hereditas 2016, 153, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babcock, C.S.; Asmussen, M.A. Effects of differential selection in the sexes on cytonuclear polymorphism and disequilibria. Genetics 1996, 144, 839–853. [Google Scholar] [CrossRef]
- Babcock, C.S.; Asmussen, M.A. Effects of differential selection in the sexes on cytonuclear dynamics: Life stages with sex differences. Genetics 1998, 149, 2063–2077. [Google Scholar] [CrossRef]
- Gemmell, N.J.; Metcalf, V.J.; Allendorf, F.W. Mother’s curse: The effect of mtDNA on individual fitness and population viability. Trends Ecol. Evol. 2004, 19, 238–244. [Google Scholar] [CrossRef]
- Dowling, D.K.; Adrian, R.E. Challenges and prospects for testing the mother’s curse hypothesis. Integr. Comp. Biol. 2019, 59, 875–889. [Google Scholar] [CrossRef]
- Maklakov, A.A.; Lummaa, V. Evolution of sex differences in lifespan and aging: Causes and constraints. BioEssays 2013, 35, 717–724. [Google Scholar] [CrossRef]
- Dowling, D.K.; Friberg, U.; Hailer, F.; Arnqvist, G.r. Intergenomic epistasis for fitness: Within-population interactions between cytoplasmic and nuclear genes in Drosophila melanogaster. Genetics 2007, 175, 235–244. [Google Scholar] [CrossRef] [Green Version]
- Maklakov, A.A.; Friberg, U.; Dowling, D.K.; Arnqvist, G. Within-population variation in cytoplasmic genes affects female life span and aging in Drosophila melanogaster. Evolution 2006, 60, 2081–2086. [Google Scholar] [CrossRef] [PubMed]
- Jelić, M.; Arnqvist, G.; Novičić, Z.K.; Kenig, B.; Tanasković, M.; Anđelković, M.; Stamenković-Radak, M. Sex-specific effects of sympatric mitonuclear variation on fitness in Drosophila subobscura. BMC Evol. Biol. 2015, 15, 135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Willett, C.S.; Burton, R.S. Environmental influences on epistatic interactions: Viabilities of cytochrome c genotypes in interpopulation crosses. Evolution 2003, 57, 2286–2292. [Google Scholar] [CrossRef] [PubMed]
- Immonen, E.; Berger, D.; Sayadi, A.; Liljestrand-Rönn, J.; Arnqvist, G. An experimental test of temperature-dependent selection on mitochondrial haplotypes in Callosobruchus maculatus seed beetles. Ecol. Evol. 2020, 10, 11387–11398. [Google Scholar] [CrossRef]
- Đorđević, M.; Stojković, B.; Savković, U.; Immonen, E.; Tucić, N.; Lazarević, J.; Arnqvist, G. Sex-specific mitonuclear epistasis and the evolution of mitochondrial bioenergetics, ageing, and life history in seed beetles. Evolution 2017, 71, 274–288. [Google Scholar] [CrossRef]
- Erić, P.; Stamenković-Radak, M.; Dragićević, M.; Kankare, M.; Wallace, M.A.; Veselinović, M.S.; Jelić, M. Mitochondrial DNA variation of Drosophila obscura (Diptera: Drosophilidae) across Europe. Eur. J. Entomol. 2022; 119, in press. [Google Scholar]
- Erić, P.; Aleksandra, P.; Erić, K.; Tanasković, M.; Davidović, S.; Rakić, M.; Savić Veselinović, M.; Stamenković-Radak, M.; Jelić, M. Unraveling the adaptive significance of mitochondrial genome variability of Drosophila obscura. In Proceedings of the 1st International Electronic Conference on Entomology, online, 1–15 July 2021. [Google Scholar]
- A Language and Environment for Statistical Computing, Version 4.0.3; R Foundation for Statistical Computing: Vienna, Austria, 2020.
- Wickham, H. Elegant graphics for data analysis. Media 2009, 35, 10.1007. [Google Scholar]
- Cox, D.R. Regression models and life-tables. J. R. Stat. Soc. Ser. B (Methodol.) 1972, 34, 187–202. [Google Scholar] [CrossRef]
- Therneau, T. A Package for Survival Analysis in R. Available online: https://cran.r-project.org/web/packages/survival/vignettes/survival.pdf (accessed on 22 December 2021).
- Bates, D.; Kliegl, R.; Vasishth, S.; Baayen, H. Parsimonious mixed models. arXiv 2015, arXiv:1506.04967. [Google Scholar]
- Kuznetsova, A.; Brockhoff, P.B.; Christensen, R.H. lmerTest package: Tests in linear mixed effects models. J. Stat. Softw. 2017, 82, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Bailey, S.F.; Alonso Morales, L.A.; Kassen, R. Effects of synonymous mutations beyond codon bias: The evidence for adaptive synonymous substitutions from microbial evolution experiments. Genome Biol. Evol. 2021, 13, evab141. [Google Scholar] [CrossRef]
- Courret, C.; Chang, C.-H.; Wei, K.H.-C.; Montchamp-Moreau, C.; Larracuente, A.M. Meiotic drive mechanisms: Lessons from Drosophila. Proc. R. Soc. B 2019, 286, 20191430. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gershenson, S. A new sex-ratio abnormality in Drosophila obscura. Genetics 1928, 13, 488. [Google Scholar] [CrossRef] [PubMed]
- Cazemajor, M.; Joly, D.; Montchamp-Moreau, C. Sex-ratio meiotic drive in Drosophila simulans is related to equational nondisjunction of the Y chromosome. Genetics 2000, 154, 229–236. [Google Scholar] [CrossRef] [PubMed]
- McKee, B.D.; Wilhelm, K.; Merrill, C.; Ren, X.-j. Male sterility and meiotic drive associated with sex chromosome rearrangements in Drosophila: Role of XY pairing. Genetics 1998, 149, 143–155. [Google Scholar] [CrossRef]
- Hiraizumi, Y. Temperature sensitivity of negative segregation distortion in Drosophila melanogaster. Genetics 1993, 135, 831–841. [Google Scholar] [CrossRef]
- Tao, Y.; Masly, J.P.; Araripe, L.; Ke, Y.; Hartl, D.L. A sex-ratio meiotic drive system in Drosophila simulans. I: An autosomal suppressor. PLoS Biol. 2007, 5, e292. [Google Scholar] [CrossRef] [Green Version]
- Ballard, J.W.O.; Pichaud, N. Mitochondrial DNA: More than an evolutionary bystander. Funct. Ecol. 2014, 28, 218–231. [Google Scholar] [CrossRef]
- Dowling, D.K. Evolutionary perspectives on the links between mitochondrial genotype and disease phenotype. Biochim. Et Biophys. Acta (BBA)-Gen. Subj. 2014, 1840, 1393–1403. [Google Scholar] [CrossRef]
- Katewa, S.D.; Ballard, J.W.O. Sympatric Drosophila simulans flies with distinct mtDNA show difference in mitochondrial respiration and electron transport. Insect Biochem. Mol. Biol. 2007, 37, 213–222. [Google Scholar] [CrossRef]
- Pichaud, N.; Ballard, J.W.O.; Tanguay, R.M.; Blier, P.U. Naturally occurring mitochondrial DNA haplotypes exhibit metabolic differences: Insight into functional properties of mitochondria. Evol. Int. J. Org. Evol. 2012, 66, 3189–3197. [Google Scholar] [CrossRef]
- Ekström, A.; Sandblom, E.; Blier, P.U.; Dupont Cyr, B.-A.; Brijs, J.; Pichaud, N. Thermal sensitivity and phenotypic plasticity of cardiac mitochondrial metabolism in European perch, Perca fluviatilis. J. Exp. Biol. 2017, 220, 386–396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lemieux, H.; Tardif, J.-C.; Blier, P.U. Thermal sensitivity of oxidative phosphorylation in rat heart mitochondria: Does pyruvate dehydrogenase dictate the response to temperature? J. Therm. Biol. 2010, 35, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Pichaud, N.; Ballard, J.W.O.; Tanguay, R.M.; Blier, P.U. Mitochondrial haplotype divergences affect specific temperature sensitivity of mitochondrial respiration. J. Bioenerg. Biomembr. 2013, 45, 25–35. [Google Scholar] [CrossRef]
- Camus, M.F.; Wolff, J.N.; Sgrò, C.M.; Dowling, D.K. Experimental support that natural selection has shaped the latitudinal distribution of mitochondrial haplotypes in Australian Drosophila melanogaster. Mol. Biol. Evol. 2017, 34, 2600–2612. [Google Scholar] [CrossRef] [PubMed]
- Consuegra, S.; John, E.; Verspoor, E.; de Leaniz, C.G. Patterns of natural selection acting on the mitochondrial genome of a locally adapted fish species. Genet. Sel. Evol. 2015, 47, 58. [Google Scholar] [CrossRef] [Green Version]
- Silva, G.; Lima, F.P.; Martel, P.; Castilho, R. Thermal adaptation and clinal mitochondrial DNA variation of European anchovy. Proc. R. Soc. B Biol. Sci. 2014, 281, 20141093. [Google Scholar] [CrossRef] [Green Version]
- Cheviron, Z.A.; Brumfield, R.T. Migration-selection balance and local adaptation of mitochondrial haplotypes in rufous-collared sparrows (Zonotrichia capensis) along an elevational gradient. Evol. Int. J. Org. Evol. 2009, 63, 1593–1605. [Google Scholar] [CrossRef]
- Fontanillas, P.; Depraz, A.; Giorgi, M.S.; Perrin, N. Nonshivering thermogenesis capacity associated to mitochondrial DNA haplotypes and gender in the greater white-toothed shrew, Crocidura russula. Mol. Ecol. 2005, 14, 661–670. [Google Scholar] [CrossRef]
- Rand, D.M.; Mossman, J.A.; Zhu, L.; Biancani, L.M.; Ge, J.Y. Mitonuclear epistasis, genotype-by-environment interactions, and personalized genomics of complex traits in Drosophila. IUBMB Life 2018, 70, 1275–1288. [Google Scholar] [CrossRef] [Green Version]
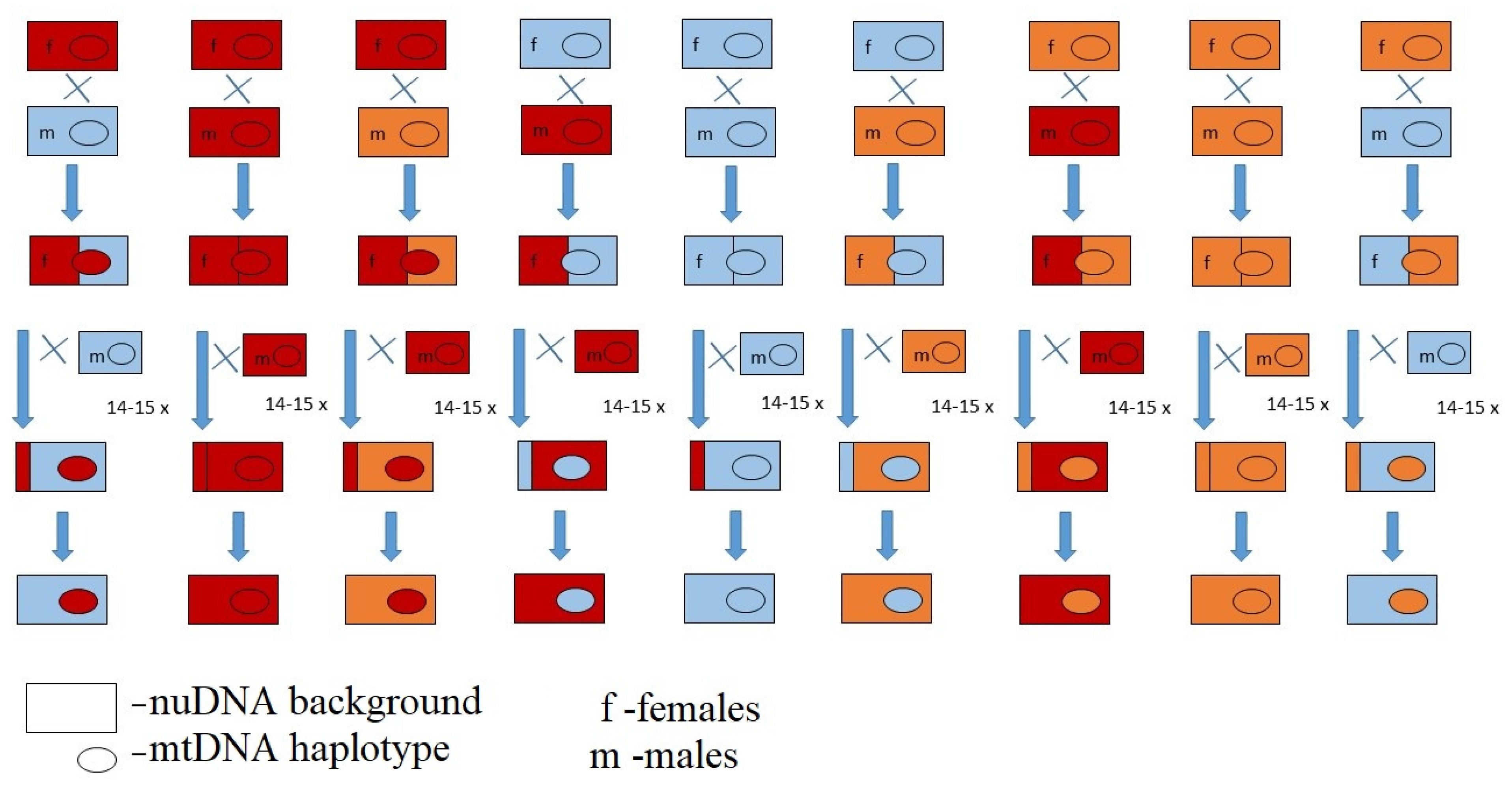
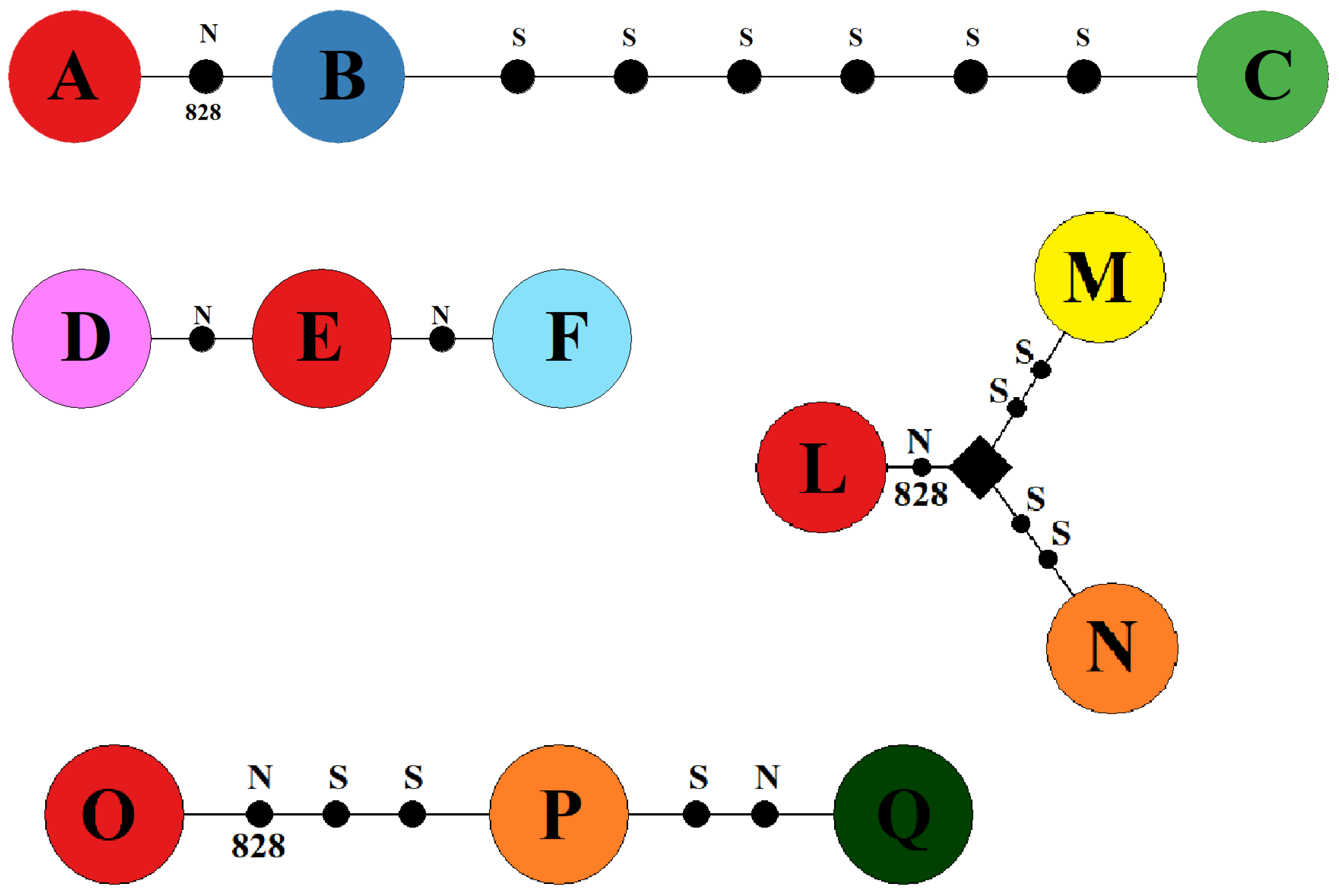
| I EB Population ST Site1 | II EB Population STSite2 | III EB Population SS | IV EB Population SG | ||||
|---|---|---|---|---|---|---|---|
| O2 | A | O9 | D | O2 | L | O2 | O |
| O1 | B | O2 | E | O4 | M | O3 | P |
| O8 | C | O10 | F | O3 | N | O18 | Q |
| EB I | AB | AC | BC | ABC | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | |
| MT | −2531.4 | 9.240 | 1 | 0.0024 | −2966.2 | 13.56 | 1 | 0.00023 | −2994.8 | 2.284 | 1 | 0.13072 | −6307.2 | 5.74 | 2 | 0.05684 |
| NU | strata | −2955.7 | 21.16 | 1 | 4.2 × 10−6 | strata | strata | |||||||||
| sex | −2521.6 | 19.570 | 1 | 9.7 × 10−6 | −2949.9 | 11.54 | 1 | 0.00068 | −2989.2 | 11.348 | 1 | 0.00076 | strata | |||
| temp(T) | strata | strata | −2952.2 | 73.989 | 1 | 2.2 × 10−16 | −6294.2 | 26.03 | 1 | 3.4 × 10−7 | ||||||
| MT:NU | −2506.3 | 30.534 | 1 | 3.3 × 10−8 | −2941.9 | 16.07 | 1 | 6.1 × 10−5 | −2949.7 | 4.846 | 1 | 0.02771 | −6264.9 | 58.51 | 4 | 6.0 × 10−12 |
| MT:sex | −2503.6 | 1.352 | 1 | 0.2450 | −2936.6 | 10.55 | 1 | 0.00116 | −2943.6 | 12.341 | 1 | 0.00044 | −6264 | 1.79 | 2 | 0.40841 |
| NU:sex | −2503.5 | 0.104 | 1 | 0.7473 | −2936.6 | 0.00 | 1 | 0.95096 | −2942.6 | 1.997 | 1 | 0.15766 | strata | |||
| MT:T | −2504.2 | 4.209 | 1 | 0.0402 | −2934 | 5.24 | 1 | 0.02209 | −2938.1 | 8.856 | 1 | 0.00292 | −6255.5 | 17.14 | 2 | 0.00019 |
| NU:T | strata | −2934 | 0.01 | 1 | 0.93708 | −2930.1 | 16.104 | 1 | 6.0 × 10−5 | −6254.9 | 1.09 | 2 | 0.57892 | |||
| T:sex | −2498.3 | 10.356 | 1 | 0.0013 | −2927.2 | 13.43 | 1 | 0.00025 | −2916.3 | 27.490 | 1 | 1.6 × 10−7 | −6233 | 43.86 | 1 | 3.5 × 10−11 |
| MT:NU:sex | −2477.9 | 0.056 | 1 | 0.8122 | −2926.4 | 1.61 | 1 | 0.20438 | −2916.3 | 0.004 | 1 | 0.94981 | −6202.2 | 61.54 | 4 | 1.4 × 10−12 |
| MT:NU:T | −2477.9 | 40.789 | 1 | 1.7 × 10−10 | −2925.7 | 1.43 | 1 | 0.23108 | −2913.7 | 5.255 | 1 | 0.02188 | −6200.9 | 2.60 | 4 | 0.62690 |
| MT:T:sex | −2476.7 | 2.378 | 1 | 0.1230 | −2925.4 | 0.59 | 1 | 0.44138 | −2912.2 | 3.115 | 1 | 0.07760 | −6200.6 | 0.56 | 2 | 0.75397 |
| NU:T:sex | −2476.6 | 0.151 | 1 | 0.6972 | −2925.1 | 0.62 | 1 | 0.43070 | −2911.7 | 0.856 | 1 | 0.35477 | −6200.1 | 0.99 | 2 | 0.61096 |
| MT:NU:T:sex | −2476.6 | 0.163 | 1 | 0.6866 | −2925 | 0.16 | 1 | 0.69248 | −2907.9 | 7.542 | 1 | 0.00603 | −6194.1 | 12.01 | 4 | 0.01725 |
| EB II | DE | DF | EF | DEF | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | |
| MT | −2993.6 | 4.84 | 1 | 0.02777 | −2601.3 | 0.61 | 1 | 0.43587 | −2964.5 | 28.42 | 1 | 9.7 × 10−8 | −7897.5 | 53.47 | 2 | 2.5 × 10−12 |
| NU | −2993.6 | 0.00 | 1 | 0.94573 | strata | −2945.6 | 37.76 | 1 | 8.0 × 10−10 | −7869.6 | 55.87 | 2 | 7.4 × 10−13 | |||
| sex | −2992 | 3.24 | 1 | 0.07200 | −2591.6 | 19.53 | 1 | 9.9 × 10−6 | −2944.5 | 2.16 | 1 | 0.14148 | strata | |||
| temp(T) | strata | strata | −7860.8 | 17.61 | 1 | 2.7 × 10−5 | ||||||||||
| MT:NU | −2991.1 | 1.74 | 1 | 0.18733 | −2591 | 1.27 | 1 | 0.26037 | −2926.3 | 36.45 | 1 | 1.6 × 10−9 | −7840.7 | 40.11 | 4 | 4.1 × 10−8 |
| MT:sex | −2988.3 | 5.54 | 1 | 0.01859 | −2587.5 | 7.00 | 1 | 0.00816 | −2926.3 | 0.01 | 1 | 0.90468 | −7832.5 | 16.34 | 2 | 0.00028 |
| NU:sex | −2973.7 | 29.23 | 1 | 6.4 × 10−8 | −2584.9 | 5.02 | 1 | 0.02511 | −2919.9 | 12.83 | 1 | 0.00034 | -7818.8 | 27.50 | 2 | 1.1 × 10−6 |
| MT:T | −2973.4 | 0.61 | 1 | 0.43353 | −2580.8 | 8.23 | 1 | 0.00412 | −2916.2 | 7.44 | 1 | 0.00638 | −7812.6 | 12.39 | 2 | 0.00204 |
| NU:T | −2963.6 | 19.60 | 1 | 9.6 × 10−6 | strata | −2910 | 12.40 | 1 | 0.00043 | −7790 | 45.24 | 2 | 1.5 × 10−10 | |||
| T:sex | −2961.2 | 4.74 | 1 | 0.02946 | −2578.8 | 3.96 | 1 | 0.04666 | −2904.3 | 11.41 | 1 | 0.00073 | −7781.2 | 17.49 | 1 | 0.00003 |
| MT:NU:sex | −2961 | 0.45 | 1 | 0.50107 | −2578.8 | 0.02 | 1 | 0.87763 | −2904.3 | 0.01 | 1 | 0.92003 | −7762.1 | 38.16 | 4 | 1.0 × 10−7 |
| MT:NU:T | −2961 | 0.06 | 1 | 0.81429 | −2577.1 | 3.57 | 1 | 0.05874 | −2882.4 | 43.78 | 1 | 3.7 × 10−11 | −7760.9 | 2.44 | 4 | 0.65468 |
| MT:T:sex | −2960.6 | 0.84 | 1 | 0.35998 | −2576.6 | 0.98 | 1 | 0.32177 | −2882.1 | 0.53 | 1 | 0.46709 | −7755.2 | 11.46 | 2 | 0.00325 |
| NU:T:sex | −2954.1 | 12.92 | 1 | 0.00032 | −2575.9 | 1.35 | 1 | 0.24562 | −2878.8 | 6.58 | 1 | 0.01032 | −7749.9 | 10.63 | 2 | 0.00491 |
| MT:NU:T:sex | −2952.2 | 3.76 | 1 | 0.05246 | −2574.6 | 2.65 | 1 | 0.10334 | −2872.9 | 11.73 | 1 | 0.00061 | −7738.2 | 23.39 | 4 | 0.00011 |
| EB III | LM | LN | MN | LMN | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | |
| MT | −2873.1 | 2.08 | 1 | 0.14912 | −3278.7 | 4.07 | 1 | 0.04362 | −2798.6 | 0.09 | 1 | 0.76424 | −6391.6 | 5.60 | 2 | 0.06090 |
| NU | −2853.4 | 39.27 | 1 | 3.7 × 10−10 | −3219 | 119.33 | 1 | 2.2 × 10−16 | −2780.6 | 36.17 | 1 | 1.8 × 10−9 | strata | |||
| sex | −2851.2 | 4.48 | 1 | 0.03431 | −3218.3 | 1.44 | 1 | 0.23044 | −2779.1 | 2.94 | 1 | 0.08623 | −6295.7 | 191.83 | 1 | 2.2 × 10−16 |
| temp(T) | −2769 | 164.44 | 1 | 2.2 × 10−16 | −3149.4 | 137.81 | 1 | 2.2 × 10−16 | −2761.2 | 35.71 | 1 | 2.3 × 10−9 | −6295.6 | 0.15 | 1 | 0.70212 |
| MT:NU | −2764.4 | 9.08 | 1 | 0.00259 | −3144.6 | 9.59 | 1 | 0.00195 | −2756.8 | 8.74 | 1 | 0.00312 | −6271.9 | 47.28 | 4 | 1.3 × 10−9 |
| MT:sex | −2764.4 | 0.03 | 1 | 0.86929 | −3142.3 | 4.55 | 1 | 0.03293 | −2756.8 | 0.06 | 1 | 0.80429 | −6263.5 | 16.84 | 2 | 0.00022 |
| NU:sex | −2761.3 | 6.26 | 1 | 0.01236 | −3137.7 | 9.26 | 1 | 0.00234 | −2755 | 3.65 | 1 | 0.05622 | −6246.6 | 33.81 | 2 | 4.6 × 10−8 |
| MT:T | −2754.5 | 13.65 | 1 | 0.00022 | −3134.7 | 6.01 | 1 | 0.01423 | −2752 | 5.98 | 1 | 0.01446 | −6245.2 | 2.87 | 2 | 0.23812 |
| NU:T | −2752.1 | 4.69 | 1 | 0.03030 | −3131.8 | 5.67 | 1 | 0.01723 | −2751.9 | 0.23 | 1 | 0.63140 | −6237.3 | 15.86 | 2 | 0.00036 |
| T:sex | −2752.1 | 0.03 | 1 | 0.85993 | −3129.3 | 5.11 | 1 | 0.02385 | −2750.3 | 3.26 | 1 | 0.07112 | −6235.9 | 2.68 | 1 | 0.10149 |
| MT:NU:sex | −2751.9 | 0.45 | 1 | 0.50464 | −3127.3 | 3.92 | 1 | 0.04782 | −2750.1 | 0.42 | 1 | 0.51560 | −6209.7 | 52.40 | 4 | 1.1 × 10−10 |
| MT:NU:T | −2749.1 | 5.48 | 1 | 0.01923 | −3123.6 | 7.48 | 1 | 0.00625 | −2737.8 | 24.51 | 1 | 7.4 × 10−7 | −6205.7 | 7.96 | 4 | 0.09311 |
| MT:T:sex | −2748.8 | 0.69 | 1 | 0.40653 | −3117 | 13.18 | 1 | 0.00028 | −2731.5 | 12.62 | 1 | 0.00038 | −6196.4 | 18.62 | 2 | 9.1 × 10−5 |
| NU:T:sex | −2747.1 | 3.43 | 1 | 0.06399 | −3116.9 | 0.17 | 1 | 0.67903 | −2730.3 | 2.45 | 1 | 0.11786 | −6195.3 | 2.18 | 2 | 0.33614 |
| MT:NU:T:sex | −2746.7 | 0.72 | 1 | 0.39545 | −3114.6 | 4.54 | 1 | 0.03320 | −2729.8 | 0.98 | 1 | 0.32162 | −6192.6 | 5.49 | 4 | 0.24058 |
| EB IV | OP | OQ | PQ | OPQ | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | LogLik | Chisq | Df | p | |
| MT | −2915.2 | 0.86 | 1 | 0.35422 | −2983.5 | 2.02 | 1 | 0.15542 | −2976.2 | 5.10 | 1 | 0.02397 | −6261.5 | 10.45 | 2 | 0.00537 |
| NU | −2915.2 | 0.02 | 1 | 0.88632 | strata | strata | strata | |||||||||
| sex | strata | −2972.6 | 21.85 | 1 | 2.9 × 10−6 | −2976.1 | 0.07 | 1 | 0.79310 | −6050.6 | 421.85 | 1 | 2.2 × 10−16 | |||
| temp(T) | −2810.4 | 209.51 | 1 | 2.2 × 10−16 | −2824.3 | 296.56 | 1 | 2.2 × 10−16 | −2888.9 | 174.40 | 1 | 2.2 × 10−16 | strata | |||
| MT:NU | −2809.9 | 0.99 | 1 | 0.32078 | −2824.3 | 0.00 | 1 | 0.96923 | −2882.8 | 12.31 | 1 | 0.00045 | −6042.6 | 16.07 | 4 | 0.00293 |
| MT:sex | −2801.8 | 16.30 | 1 | 5.4 × 10−5 | −2822.7 | 3.21 | 1 | 0.07312 | −2879.1 | 7.36 | 1 | 0.00667 | −6034.5 | 16.05 | 2 | 0.00033 |
| NU:sex | −2801.8 | 0.01 | 1 | 0.93663 | −2818.3 | 8.78 | 1 | 0.00304 | −2879.1 | 0.07 | 1 | 0.79796 | −6032.6 | 3.96 | 2 | 0.13834 |
| MT:T | −2800.4 | 2.72 | 1 | 0.09938 | −2816.2 | 4.26 | 1 | 0.03900 | −2879 | 0.06 | 1 | 0.80007 | −6017.3 | 30.56 | 2 | 2.3 × 10−7 |
| NU:T | −2800.4 | 0.01 | 1 | 0.91035 | −2815 | 2.31 | 1 | 0.12894 | −2878.2 | 1.75 | 1 | 0.18603 | strata | |||
| T:sex | −2797.4 | 5.96 | 1 | 0.01462 | −2814.6 | 0.80 | 1 | 0.36981 | −2878 | 0.35 | 1 | 0.55487 | −6015.4 | 3.78 | 1 | 0.05190 |
| MT:NU:sex | −2796.4 | 2.03 | 1 | 0.15397 | −2814.6 | 0.07 | 1 | 0.79290 | −2874.8 | 6.38 | 1 | 0.01152 | −6005.6 | 19.63 | 4 | 0.00059 |
| MT:NU:T | −2795.5 | 1.87 | 1 | 0.17163 | −2810.6 | 8.00 | 1 | 0.00469 | −2864.6 | 20.40 | 1 | 6.3 × 10−6 | −5998.3 | 14.43 | 4 | 0.00603 |
| MT:T:sex | −2788.6 | 13.76 | 1 | 0.00021 | −2805.3 | 10.55 | 1 | 0.00116 | −2864.6 | 0.06 | 1 | 0.81162 | −5990.1 | 16.49 | 2 | 0.00026 |
| NU:T:sex | −2785.2 | 6.81 | 1 | 0.00907 | −2798.7 | 13.27 | 1 | 0.00027 | −2862.5 | 4.16 | 1 | 0.04134 | −5969.5 | 41.23 | 2 | 1.1 × 10−9 |
| MT:NU:T:sex | −2781.5 | 7.34 | 1 | 0.00674 | −2794.8 | 7.64 | 1 | 0.00570 | −2861.4 | 2.21 | 1 | 0.13668 | −5962.8 | 13.45 | 4 | 0.00929 |
| EB I | AB | AC | BC | ABC | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | |
| MT | 0.450 | 70.0 | 0.62 | 0.4320 | 5.2 | 72.3 | 6.46 | 0.0132 | 0.1 | 69.6 | 0.10 | 0.7585 | 0.5 | 158.7 | 0.31 | 0.7374 |
| NU | 0.350 | 70.0 | 0.49 | 0.4870 | 7.8 | 72.3 | 9.61 | 0.0028 | 18.3 | 69.6 | 27.55 | 1.6 × 10−6 | 33.6 | 158.6 | 21.55 | 5.3 × 10−9 |
| sex | 5.400 | 931.4 | 7.43 | 0.0065 | 3.8 | 1042.2 | 4.67 | 0.0309 | 1.4 | 1028.8 | 2.18 | 0.1405 | 6.3 | 2214.3 | 8.12 | 0.0044 |
| temp(T) | 1726 | 70.0 | 2374 | 2.2 × 10−16 | 1675 | 72.3 | 2073 | 2.2 × 10−16 | 1353 | 69.6 | 2041 | 2.2 × 10−16 | 3835 | 158.7 | 4913 | 2.2 × 10−16 |
| MT:NU | 3.120 | 70.0 | 4.30 | 0.0419 | 5.3 | 72.3 | 6.50 | 0.0129 | 1.5 | 69.6 | 2.20 | 0.1428 | 14.4 | 158.5 | 4.63 | 0.0015 |
| MT:sex | 0.210 | 931.4 | 0.29 | 0.5928 | 1.3 | 1042.2 | 1.56 | 0.2123 | 1.1 | 1028.8 | 1.73 | 0.1892 | 1 | 2214.2 | 0.65 | 0.5217 |
| NU:sex | 1.670 | 931.4 | 2.30 | 0.1298 | 2.1 | 1042.2 | 2.56 | 0.1100 | 0.5 | 1028.8 | 0.68 | 0.4113 | 2.5 | 2213.9 | 1.60 | 0.2019 |
| MT:T | 1.740 | 70.0 | 2.39 | 0.1263 | 37.4 | 72.3 | 46.29 | 2.5 × 10−9 | 0.1 | 69.6 | 0.13 | 0.7176 | 5.1 | 158.7 | 3.25 | 0.0416 |
| NU:T | 0.950 | 70.0 | 1.31 | 0.2571 | 21.0 | 72.3 | 26.04 | 2.6 × 10−6 | 7.4 | 69.6 | 11.12 | 0.0014 | 34.9 | 158.6 | 22.35 | 2.8 × 10−9 |
| T:sex | 0.490 | 931.4 | 0.67 | 0.4131 | 3.7 | 1042.2 | 4.53 | 0.0336 | 5.6 | 1028.8 | 8.42 | 0.0038 | 4.6 | 2214.3 | 5.95 | 0.0148 |
| MT:NU:sex | 1.550 | 931.4 | 2.14 | 0.1441 | 0.0 | 1042.2 | 0.01 | 0.9115 | 0.0 | 1028.8 | 0.01 | 0.9201 | 1.9 | 2213.6 | 0.60 | 0.6591 |
| MT:NU:T | 14.96 | 70.0 | 20.58 | 2.3 × 10−5 | 0.2 | 72.3 | 0.24 | 0.6277 | 3.9 | 69.6 | 5.84 | 0.0183 | 55.4 | 158.5 | 17.75 | 4.6 × 10−12 |
| MT:T:sex | 2.310 | 931.4 | 3.18 | 0.0749 | 0.0 | 1042.2 | 0.00 | 0.9446 | 1.2 | 1028.8 | 1.75 | 0.1857 | 0.3 | 2214.2 | 0.20 | 0.8185 |
| NU:T:sex | 1.810 | 931.4 | 2.49 | 0.1146 | 0.0 | 1042.2 | 0.01 | 0.9263 | 1.9 | 1028.8 | 2.91 | 0.0885 | 3 | 2213.9 | 1.93 | 0.1452 |
| MT:NU:T:sex | 1.990 | 931.4 | 2.74 | 0.0984 | 0.2 | 1042.2 | 0.25 | 0.6154 | 0.0 | 1028.8 | 0.05 | 0.8181 | 7.6 | 2213.6 | 2.44 | 0.0446 |
| EB II | DE | DF | EF | DEF | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | |
| MT | 5.54 | 70.66 | 5.63 | 0.020 | 9.28 | 70.7 | 8.72 | 0.0043 | 32.1 | 71.14 | 33.26 | 1.9 × 10−7 | 45.3 | 160.0 | 22.66 | 2.2 × 10−9 |
| NU | 3.93 | 70.66 | 4.00 | 0.049 | 20.78 | 70.7 | 19.52 | 3.5 × 10−5 | 3.9 | 71.14 | 4.07 | 0.0474 | 12.7 | 160.0 | 6.33 | 0.0023 |
| sex | 6.36 | 954.08 | 6.46 | 0.011 | 0.25 | 996.9 | 0.24 | 0.6271 | 0.9 | 996 | 0.91 | 0.3396 | 3.7 | 2233.5 | 3.72 | 0.0538 |
| temp(T) | 2474 | 70.66 | 2514 | 2 × 10−16 | 3028 | 72.0 | 2845 | 2.2 × 10−16 | 3265 | 71.14 | 3378 | 2.2 × 10−16 | 6008 | 160.1 | 6008 | 2.2 × 10−16 |
| MT:NU | 0 | 70.66 | 0.00 | 0.982 | 0.08 | 70.7 | 0.08 | 0.7843 | 8.3 | 71.14 | 8.64 | 0.0044 | 8.2 | 160.0 | 2.06 | 0.0889 |
| MT:sex | 1.72 | 954.08 | 1.75 | 0.186 | 3.69 | 996.9 | 3.46 | 0.0631 | 6.5 | 996 | 6.71 | 0.0097 | 5.8 | 2233.1 | 2.90 | 0.0554 |
| NU:sex | 3.57 | 954.08 | 3.63 | 0.057 | 2.36 | 996.9 | 2.21 | 0.1371 | 1.6 | 996 | 1.67 | 0.1960 | 14.6 | 2233.0 | 7.28 | 0.0007 |
| MT:T | 0.96 | 70.66 | 0.98 | 0.326 | 6.81 | 72.0 | 6.40 | 0.0136 | 14.6 | 71.14 | 15.15 | 0.0002 | 19.6 | 160.0 | 9.81 | 9.5 × 10−5 |
| NU:T | 1.77 | 70.66 | 1.80 | 0.184 | 26.3 | 72.0 | 24.71 | 4.4 × 10−6 | 15.4 | 71.14 | 15.95 | 0.0002 | 38.4 | 160.0 | 19.20 | 3.4 × 10−8 |
| T:sex | 1.6 | 954.08 | 1.63 | 0.202 | 0.51 | 999.3 | 0.48 | 0.4875 | 0 | 996 | 0.02 | 0.8837 | 2.2 | 2233.5 | 2.20 | 0.1382 |
| MT:NU:sex | 1.38 | 954.08 | 1.41 | 0.236 | 0.05 | 996.9 | 0.04 | 0.8339 | 0.2 | 996 | 0.21 | 0.6484 | 5.2 | 2231.9 | 1.31 | 0.2643 |
| MT:NU:T | 0.05 | 70.66 | 0.05 | 0.826 | 0.09 | 72.0 | 0.08 | 0.7771 | 0.3 | 71.14 | 0.32 | 0.5719 | 0.5 | 160.0 | 0.14 | 0.9685 |
| MT:T:sex | 1.52 | 954.08 | 1.54 | 0.215 | 3.48 | 999.3 | 3.27 | 0.0710 | 4.9 | 996 | 5.03 | 0.0251 | 3.4 | 2233.1 | 1.69 | 0.1850 |
| NU:T:sex | 3.33 | 954.08 | 3.38 | 0.066 | 3.42 | 999.3 | 3.21 | 0.0735 | 7.4 | 996 | 7.66 | 0.0057 | 7.2 | 2233.0 | 3.62 | 0.0269 |
| MT:NU:T:sex | 0.01 | 954.08 | 0.01 | 0.943 | 0 | 999.3 | 0.00 | 0.9865 | 6.5 | 996 | 6.77 | 0.0094 | 12.2 | 2231.9 | 3.05 | 0.0162 |
| EB III | LM | LN | MN | LMN | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | |
| MT | 0.08 | 352.01 | 0.08 | 0.774 | 2.56 | 79.85 | 1.64 | 0.2040 | 10.17 | 263.43 | 10.84 | 0.0011 | 1.13 | 508.12 | 0.48 | 0.6201 |
| NU | 0.08 | 352.01 | 0.08 | 0.772 | 0.77 | 79.85 | 0.50 | 0.4832 | 19.82 | 263.43 | 21.13 | 6.7× 10−6 | 20.16 | 350.54 | 8.55 | 0.0002 |
| sex | 0.68 | 684.68 | 0.75 | 0.388 | 4.85 | 623.7 | 3.11 | 0.0783 | 0.35 | 782.68 | 0.37 | 0.5435 | 5.25 | 1563.83 | 4.45 | 0.0350 |
| temp(T) | 840 | 352.01 | 917 | 2× 10−16 | 2147 | 79.85 | 1376 | 2× 10−16 | 1447 | 263.43 | 1543 | 2.2× 10−16 | 2672 | 550.98 | 2267 | 2.2× 10−16 |
| MT:NU | 2.89 | 352.01 | 3.15 | 0.077 | 0.84 | 79.85 | 0.54 | 0.4657 | 9.71 | 263.43 | 10.35 | 0.0015 | 23.37 | 334.23 | 4.96 | 0.0007 |
| MT:sex | 0.41 | 684.68 | 0.44 | 0.506 | 0.03 | 623.7 | 0.02 | 0.8810 | 1.43 | 782.68 | 1.52 | 0.2174 | 0.09 | 1566.61 | 0.04 | 0.9619 |
| NU:sex | 2.61 | 684.68 | 2.84 | 0.092 | 0 | 623.7 | 0.00 | 0.9913 | 2.93 | 782.68 | 3.12 | 0.0776 | 1.56 | 1582.77 | 0.66 | 0.5158 |
| MT:T | 0.11 | 352.01 | 0.12 | 0.731 | 5.7 | 79.85 | 3.65 | 0.0595 | 2.61 | 263.43 | 2.78 | 0.0968 | 1.05 | 508.12 | 0.45 | 0.6408 |
| NU:T | 0.19 | 352.01 | 0.20 | 0.653 | 6.99 | 79.85 | 4.48 | 0.0375 | 11.09 | 263.43 | 11.82 | 0.0007 | 5.67 | 350.54 | 2.41 | 0.0917 |
| T:sex | 0.31 | 684.68 | 0.34 | 0.558 | 0.77 | 623.7 | 0.49 | 0.4830 | 2.09 | 782.68 | 2.23 | 0.1356 | 0.14 | 1563.83 | 0.11 | 0.7348 |
| MT:NU:sex | 4.55 | 684.68 | 4.97 | 0.026 | 0.06 | 623.7 | 0.04 | 0.8425 | 5.77 | 782.68 | 6.15 | 0.0134 | 8.51 | 1580.6 | 1.81 | 0.1252 |
| MT:NU:T | 8.14 | 352.01 | 8.89 | 0.003 | 0.6 | 79.85 | 0.39 | 0.5365 | 0 | 263.43 | 0.00 | 0.9458 | 10.77 | 334.23 | 2.29 | 0.0600 |
| MT:T:sex | 2.8 | 684.68 | 3.06 | 0.081 | 1.96 | 623.7 | 1.26 | 0.2626 | 0.07 | 782.68 | 0.07 | 0.7913 | 3.37 | 1566.61 | 1.43 | 0.2395 |
| NU:T:sex | 0.08 | 684.68 | 0.09 | 0.770 | 2.56 | 623.7 | 1.64 | 0.2009 | 0.31 | 782.68 | 0.33 | 0.5684 | 1.94 | 1582.77 | 0.82 | 0.4396 |
| MT:NU:T:sex | 1.14 | 684.68 | 1.25 | 0.265 | 0.39 | 623.7 | 0.25 | 0.6165 | 0.96 | 782.68 | 1.02 | 0.3117 | 2.41 | 1580.6 | 0.51 | 0.7280 |
| EB IV | OP | OQ | PQ | OPQ | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | SSq | ddf | F | p | |
| MT | 4.32 | 74.1 | 5.23 | 0.02510 | 3.07 | 82.0 | 2.99 | 0.0877 | 28.92 | 66.4 | 45.15 | 5.0× 10−9 | 22.88 | 172.1 | 14.38 | 1.7× 10−6 |
| NU | 25.84 | 74.1 | 31.28 | 3.6× 10−7 | 2.46 | 82.0 | 2.39 | 0.1263 | 0.73 | 66.4 | 1.15 | 0.2881 | 19.83 | 171.9 | 12.47 | 8.8× 10−6 |
| sex | 9.05 | 1146 | 10.95 | 0.00097 | 20.83 | 884.2 | 20.23 | 7.8× 10−6 | 9.46 | 1014 | 14.76 | 0.0001 | 29.81 | 2318 | 37.47 | 1.1× 10−9 |
| temp(T) | 1690 | 74.1 | 2045 | 2.2× 10−16 | 1235 | 82.0 | 1199 | 2.2× 10−16 | 1251 | 66.4 | 1953 | 2.2× 10−16 | 3075 | 172.6 | 3865 | 2.2× 10−16 |
| MT:NU | 3.18 | 74.1 | 3.85 | 0.05362 | 23.28 | 82.0 | 22.61 | 8.4× 10−6 | 9.90 | 66.4 | 15.46 | 0.0002 | 28.23 | 171.0 | 8.87 | 1.6× 10−6 |
| MT:sex | 0.01 | 1146 | 0.01 | 0.91750 | 0.08 | 884.2 | 0.07 | 0.7870 | 2.70 | 1014 | 4.21 | 0.0404 | 2.04 | 2315 | 1.28 | 0.2779 |
| NU:sex | 6.47 | 1146 | 7.82 | 0.00524 | 0.10 | 884.2 | 0.10 | 0.7572 | 0.89 | 1014 | 1.39 | 0.2392 | 6.15 | 2314 | 3.87 | 0.0211 |
| MT:T | 2.19 | 74.1 | 2.64 | 0.10813 | 0.00 | 82.0 | 0.00 | 0.9813 | 0.17 | 66.4 | 0.27 | 0.6049 | 2.55 | 172.1 | 1.60 | 0.2045 |
| NU:T | 5.55 | 74.1 | 6.72 | 0.01149 | 0.08 | 82.0 | 0.08 | 0.7833 | 8.87 | 66.4 | 13.84 | 0.0004 | 18.35 | 171.9 | 11.53 | 2.0× 10−5 |
| T:sex | 0.15 | 1146 | 0.18 | 0.66836 | 0.20 | 884.2 | 0.20 | 0.6581 | 0.12 | 1014 | 0.18 | 0.6711 | 0.38 | 2318 | 0.48 | 0.4905 |
| MT:NU:sex | 0.03 | 1146 | 0.04 | 0.83715 | 0.85 | 884.2 | 0.82 | 0.3643 | 0.10 | 1014 | 0.16 | 0.6907 | 2.65 | 2311 | 0.83 | 0.5049 |
| MT:NU:T | 2.1 | 74.1 | 2.55 | 0.11480 | 0.00 | 82.0 | 0.00 | 0.9927 | 0.23 | 66.4 | 0.36 | 0.5500 | 2.12 | 171.0 | 0.67 | 0.6167 |
| MT:T:sex | 1.21 | 1146 | 1.47 | 0.22586 | 0.41 | 884.2 | 0.40 | 0.5279 | 0.71 | 1014 | 1.11 | 0.2914 | 0.52 | 2315 | 0.33 | 0.7202 |
| NU:T:sex | 0.15 | 1146 | 0.18 | 0.66802 | 2.99 | 884.2 | 2.90 | 0.0888 | 0.03 | 1014 | 0.05 | 0.8156 | 1.03 | 2314 | 0.65 | 0.5230 |
| MT:NU:T:sex | 6.44 | 1146 | 7.79 | 0.00534 | 0.02 | 884.2 | 0.02 | 0.8987 | 0.11 | 1014 | 0.18 | 0.6740 | 7.18 | 2311 | 2.26 | 0.0607 |
| EB I | AB | AC | BC | ABC | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 1.11 | 0.2932 | 1 | 7.24 | 0.0071 | 1 | 1.874 | 0.1711 | 2 | 7.041 | 0.0296 |
| NU | 1 | 40.66 | 1.8× 10−10 | 1 | 132.6 | 2.2× 10−16 | 1 | 171.0 | 2.2× 10−16 | 2 | 288.7 | 2.2× 10−16 |
| temp(T) | 1 | 0.18 | 0.6732 | 1 | 20.15 | 7.2× 10−6 | 1 | 57.03 | 4.3× 10−14 | 1 | 48.41 | 3.5× 10−12 |
| MT:NU | 1 | 17.38 | 3.1× 10−5 | 1 | 78.50 | 2.2× 10−16 | 1 | 2.71 | 0.0997 | 4 | 101.2 | 2.2× 10−16 |
| MT:T | 1 | 18.39 | 1.8× 10−5 | 1 | 1.45 | 0.2285 | 1 | 15.77 | 7.1× 10−5 | 2 | 16.51 | 2.6× 10−4 |
| NU:T | 1 | 3.23 | 0.0723 | 1 | 70.44 | 2.2× 10−16 | 1 | 15.24 | 9.5× 10−5 | 2 | 62.91 | 2.2× 10−14 |
| MT:NU:T | 1 | 11.82 | 0.0006 | 1 | 1.839 | 0.1751 | 1 | 2.761 | 0.0966 | 4 | 40.30 | 3.7× 10−8 |
| EB II | DE | DF | EF | DEF | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 15.51 | 8.2× 10−5 | 1 | 10.55 | 0.0012 | 1 | 1.458 | 0.2273 | 2 | 11.51 | 0.0032 |
| NU | 1 | 66.97 | 2.8× 10−16 | 1 | 34.21 | 4.9× 10−9 | 1 | 35.74 | 2.3× 10−9 | 2 | 130.5 | 2.2× 10−16 |
| temp(T) | 1 | 29.32 | 6.1× 10−8 | 1 | 87.20 | 2.2× 10−16 | 1 | 40.59 | 1.9× 10−10 | 1 | 200.5 | 2.2× 10−16 |
| MT:NU | 1 | 2.596 | 0.1071 | 1 | 23.32 | 1.4× 10−6 | 1 | 13.11 | 0.0003 | 4 | 76.41 | 1.0× 10−15 |
| MT:T | 1 | 31.34 | 2.2× 10−8 | 1 | 9.333 | 0.0023 | 1 | 19.69 | 9.1× 10−6 | 2 | 23.56 | 7.7× 10−6 |
| NU:T | 1 | 0.811 | 0.3678 | 1 | 19.71 | 9.0× 10−6 | 1 | 62.83 | 2.3× 10−15 | 2 | 31.95 | 1.2× 10−7 |
| MT:NU:T | 1 | 149.9 | 2.2× 10−16 | 1 | 15.07 | 0.0001 | 1 | 36.52 | 1.5× 10−9 | 4 | 173.4 | 2.2× 10−16 |
| EB III | LM | LN | MN | LMN | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 1.085 | 0.2976 | 1 | 28.27 | 1.1× 10−7 | 1 | 6.41 | 0.0114 | 2 | 18.83 | 8.1× 10−5 |
| NU | 1 | 238.3 | 2.2× 10−16 | 1 | 56.36 | 6.0× 10−14 | 1 | 94.99 | 2.2× 10−16 | 2 | 522.3 | 2.2× 10−16 |
| temp(T) | 1 | 313.8 | 2.2× 10−16 | 1 | 18.26 | 1.9× 10−5 | 1 | 225.9 | 2.2× 10−16 | 1 | 266.4 | 2.2× 10−16 |
| MT:NU | 1 | 8.779 | 0.0030 | 1 | 17.70 | 2.6× 10−5 | 1 | 16.53 | 4.8× 10−5 | 4 | 55.97 | 2.0× 10−11 |
| MT:T | 1 | 32.16 | 1.4× 10−8 | 1 | 27.78 | 1.4× 10−7 | 1 | 1.581 | 0.2087 | 2 | 37.28 | 8.0× 10−9 |
| NU:T | 1 | 0.115 | 0.7341 | 1 | 2.159 | 0.1418 | 1 | 125.2 | 2.2× 10−16 | 2 | 106.3 | 2.2× 10−16 |
| MT:NU:T | 1 | 28.06 | 1.2× 10−7 | 1 | 50.24 | 1.4× 10−12 | 1 | 1.9 | 0.1681 | 4 | 126.0 | 2.2× 10−16 |
| EB IV | OP | OQ | PQ | OPQ | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 20.38 | 6.3× 10−6 | 1 | 78.53 | 2.2× 10−16 | 1 | 87.32 | 2.0× 10−16 | 2 | 200.8 | 2.2× 10−16 |
| NU | 1 | 9.33 | 0.0023 | 1 | 103.0 | 2.2× 10−16 | 1 | 327.1 | 2.0× 10−16 | 2 | 447.3 | 2.2× 10−16 |
| temp(T) | 1 | 45.95 | 1.2× 10−11 | 1 | 2.752 | 0.0971 | 1 | 0 | 0.9581 | 1 | 17.14 | 3.5× 10−5 |
| MT:NU | 1 | 7.112 | 0.0077 | 1 | 0.01 | 0.9197 | 1 | 128.7 | 2.0× 10−16 | 4 | 148.3 | 2.2× 10−16 |
| MT:T | 1 | 0.061 | 0.8051 | 1 | 57.04 | 4.3× 10−14 | 1 | 1.42 | 0.2339 | 2 | 61.24 | 5.0× 10−14 |
| NU:T | 1 | 0.299 | 0.5844 | 1 | 6.415 | 0.0113 | 1 | 5.57 | 0.0183 | 2 | 2.32 | 0.3130 |
| MT:NU:T | 1 | 2.017 | 0.1555 | 1 | 0.134 | 0.7144 | 1 | 0.48 | 0.4900 | 4 | 23.16 | 0.0001 |
| EB I | AB | AC | BC | ABC | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 0.02 | 0.8808 | 1 | 0.04739 | 0.8277 | 1 | 1.33724 | 0.24750 | 2 | 0.36 | 0.834 |
| NU | 1 | 0.00 | 0.9872 | 1 | 2.74073 | 0.0978 | 1 | 0.00149 | 0.96920 | 2 | 1.12 | 0.571 |
| temp(T) | 1 | 0.30 | 0.5828 | 1 | 0.09581 | 0.7569 | 1 | 0.00376 | 0.95110 | 1 | 0.24 | 0.626 |
| MT:NU | 1 | 0.01 | 0.9396 | 1 | 0.48097 | 0.4880 | 1 | 0.30348 | 0.58170 | 4 | 2.47 | 0.651 |
| MT:T | 1 | 0.09 | 0.7636 | 1 | 1.0408 | 0.3076 | 1 | 0.0083 | 0.92740 | 2 | 0.16 | 0.922 |
| NU:T | 1 | 1.78 | 0.1825 | 1 | 0.48283 | 0.4871 | 1 | 0.67503 | 0.41130 | 2 | 0.35 | 0.841 |
| MT:NU:T | 1 | 1.92 | 0.1659 | 1 | 0.06717 | 0.7955 | 1 | 1.32005 | 0.25060 | 4 | 5.53 | 0.237 |
| EB II | DE | DF | EF | DEF | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 4.51 | 0.03379 | 1 | 1.6082 | 0.20470 | 1 | 0.1452 | 0.70314 | 2 | 12.25 | 0.002 |
| NU | 1 | 0.62 | 0.43183 | 1 | 0.1475 | 0.70100 | 1 | 2.5504 | 0.11027 | 2 | 28.76 | 6× 10−7 |
| temp(T) | 1 | 7.31 | 0.00686 | 1 | 1.6117 | 0.20430 | 1 | 15.36 | 8.9× 10−5 | 1 | 0.31 | 0.578 |
| MT:NU | 1 | 1.09 | 0.29647 | 1 | 0.0122 | 0.91210 | 1 | 4.9746 | 0.02572 | 4 | 19.09 | 0.001 |
| MT:T | 1 | 4.97 | 0.02586 | 1 | 15.2255 | 9.5× 10−5 | 1 | 0.5121 | 0.47425 | 2 | 0.74 | 0.691 |
| NU:T | 1 | 8.00 | 0.00468 | 1 | 0.2847 | 0.59360 | 1 | 10.6933 | 0.00108 | 2 | 5.59 | 0.061 |
| MT:NU:T | 1 | 1.10 | 0.29332 | 1 | 0.5458 | 0.46000 | 1 | 0.3765 | 0.53949 | 4 | 6.17 | 0.187 |
| EB III | LM | LN | MN | LMN | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 0.00 | 0.97074 | 1 | 0.3288 | 0.56638 | 1 | 12.92 | 0.00032 | 2 | 11.7 | 0.003 |
| NU | 1 | 472.89 | 2.2× 10−16 | 1 | 0.0007 | 0.97957 | 1 | 633.78 | 2.2× 10−16 | 2 | 1125.3 | 2× 10−16 |
| temp(T) | 1 | 8.40 | 0.00375 | 1 | 2.7945 | 0.09459 | 1 | 4.59 | 0.03210 | 1 | 5.5 | 0.019 |
| MT:NU | 1 | 15.68 | 7.5× 10−5 | 1 | 3.479 | 0.06215 | 1 | 23.35 | 1.4× 10−6 | 4 | 33.1 | 1× 10−6 |
| MT:T | 1 | 0.35 | 0.55523 | 1 | 0.8008 | 0.37085 | 1 | 3.57 | 0.05889 | 2 | 1.0 | 0.610 |
| NU:T | 1 | 8.54 | 0.00348 | 1 | 1.3167 | 0.25119 | 1 | 13.88 | 0.00019 | 2 | 9.3 | 0.010 |
| MT:NU:T | 1 | 14.50 | 0.00014 | 1 | 0.0018 | 0.96654 | 1 | 5.64 | 0.01751 | 4 | 16.9 | 0.002 |
| EB IV | OP | OQ | PQ | OPQ | ||||||||
| Df | Dev | p | Df | Dev | p | Df | Dev | p | Df | Dev | p | |
| MT | 1 | 35.30 | 2.8× 10−9 | 1 | 3.409 | 0.06486 | 1 | 0.0482 | 0.82617 | 2 | 2.20 | 0.334 |
| NU | 1 | 90.58 | 2.2× 10−16 | 1 | 7.96 | 0.00478 | 1 | 0.4858 | 0.48582 | 2 | 188.33 | 2× 10−16 |
| temp(T) | 1 | 1.13 | 0.28824 | 1 | 2.405 | 0.12093 | 1 | 0.5486 | 0.45890 | 1 | 0.68 | 0.409 |
| MT:NU | 1 | 66.80 | 3.0× 10−16 | 1 | 102.83 | 2.2× 10−16 | 1 | 0.9611 | 0.32692 | 4 | 168.34 | 2× 10−16 |
| MT:T | 1 | 39.46 | 3.3× 10−10 | 1 | 44.23 | 2.9× 10−11 | 1 | 0.1021 | 0.74928 | 2 | 27.32 | 1× 10−6 |
| NU:T | 1 | 0.48 | 0.49005 | 1 | 140.156 | 2.2× 10−16 | 1 | 4.3152 | 0.03777 | 2 | 31.88 | 1× 10−7 |
| MT:NU:T | 1 | 11.23 | 0.00081 | 1 | 0.674 | 0.41178 | 1 | 3.7143 | 0.05395 | 4 | 169.68 | 2× 10−16 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Erić, P.; Patenković, A.; Erić, K.; Tanasković, M.; Davidović, S.; Rakić, M.; Savić Veselinović, M.; Stamenković-Radak, M.; Jelić, M. Temperature-Specific and Sex-Specific Fitness Effects of Sympatric Mitochondrial and Mito-Nuclear Variation in Drosophila obscura. Insects 2022, 13, 139. https://doi.org/10.3390/insects13020139
Erić P, Patenković A, Erić K, Tanasković M, Davidović S, Rakić M, Savić Veselinović M, Stamenković-Radak M, Jelić M. Temperature-Specific and Sex-Specific Fitness Effects of Sympatric Mitochondrial and Mito-Nuclear Variation in Drosophila obscura. Insects. 2022; 13(2):139. https://doi.org/10.3390/insects13020139
Chicago/Turabian StyleErić, Pavle, Aleksandra Patenković, Katarina Erić, Marija Tanasković, Slobodan Davidović, Mina Rakić, Marija Savić Veselinović, Marina Stamenković-Radak, and Mihailo Jelić. 2022. "Temperature-Specific and Sex-Specific Fitness Effects of Sympatric Mitochondrial and Mito-Nuclear Variation in Drosophila obscura" Insects 13, no. 2: 139. https://doi.org/10.3390/insects13020139
APA StyleErić, P., Patenković, A., Erić, K., Tanasković, M., Davidović, S., Rakić, M., Savić Veselinović, M., Stamenković-Radak, M., & Jelić, M. (2022). Temperature-Specific and Sex-Specific Fitness Effects of Sympatric Mitochondrial and Mito-Nuclear Variation in Drosophila obscura. Insects, 13(2), 139. https://doi.org/10.3390/insects13020139








