Identification and Expression Analysis of Four Small Heat Shock Protein Genes in Cigarette Beetle, Lasioderma serricorne (Fabricius)
Abstract
1. Introduction
2. Materials and Methods
2.1. Insect and Sample Preparation
2.2. 20E and Temperature Treatment
2.3. Immune Challenge and Controlled Atmosphere Treatment
2.4. RNA Extraction and cDNA Synthesis
2.5. Identification and Sequencing of LsHsp cDNAs
2.6. Bioinformatics and Phylogenetic Analyses
2.7. Quantitative Real-Time PCR
2.8. Data Analyses
3. Results
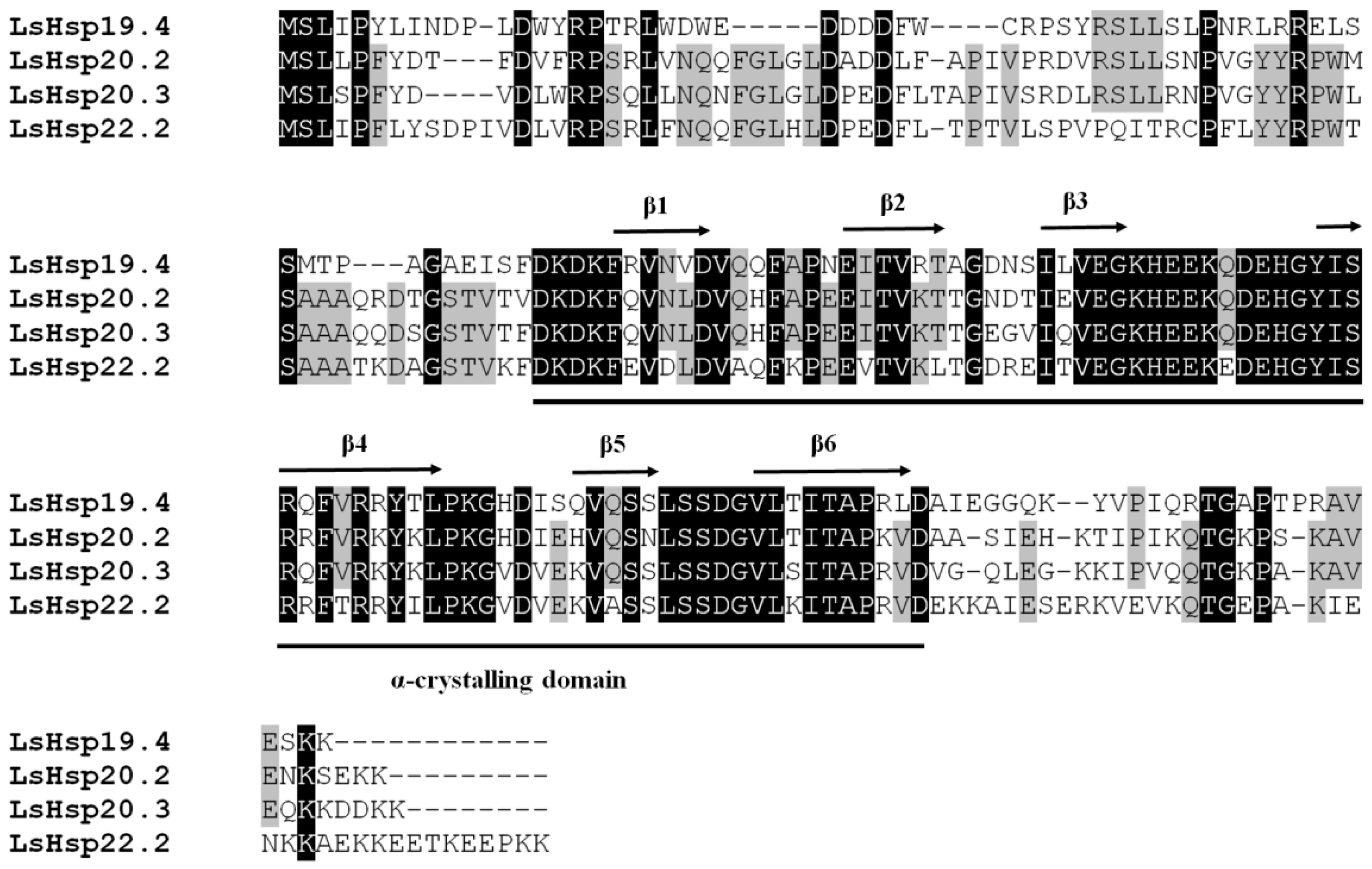
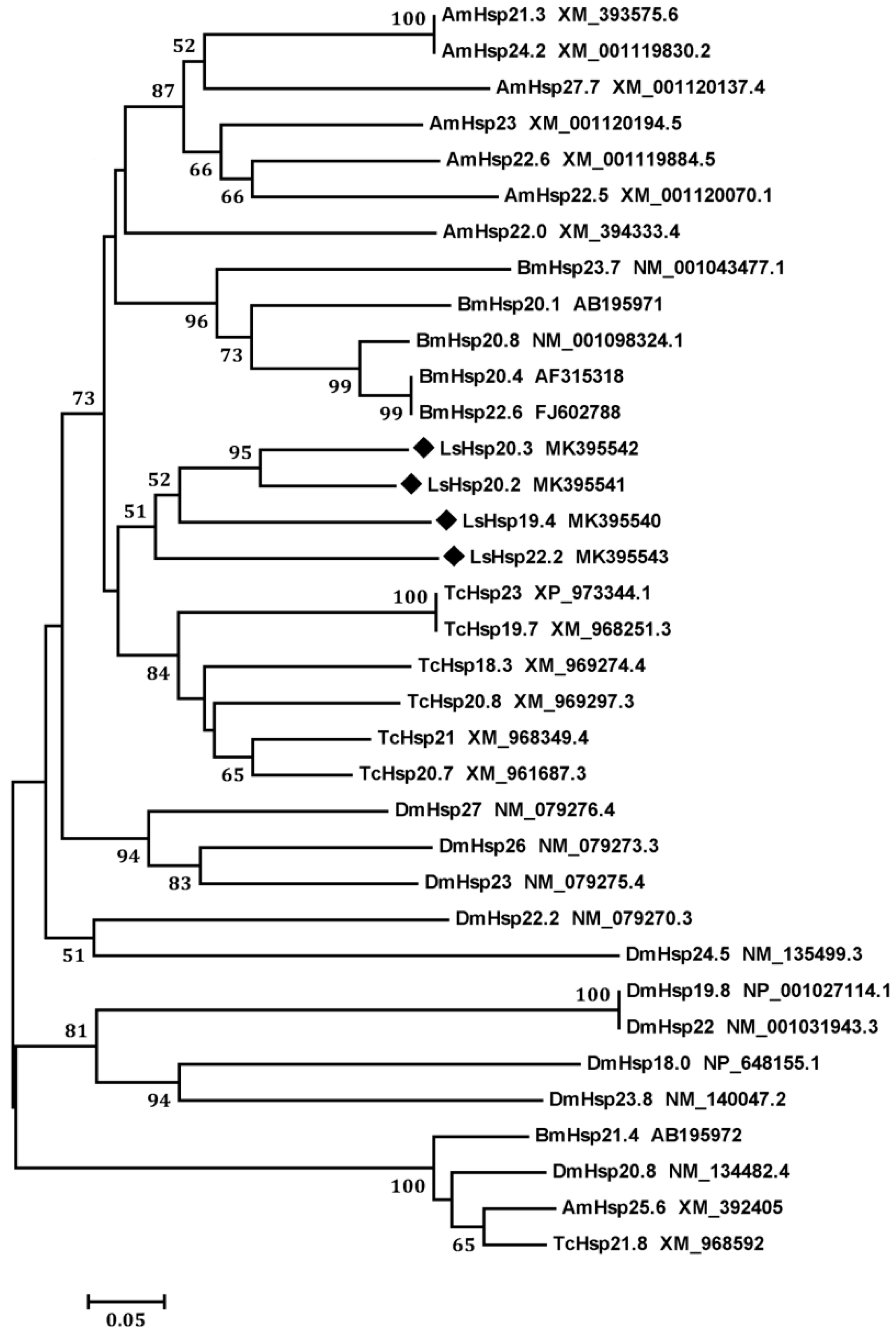
3.1. Identification and Characterization of Four LsHsp Genes
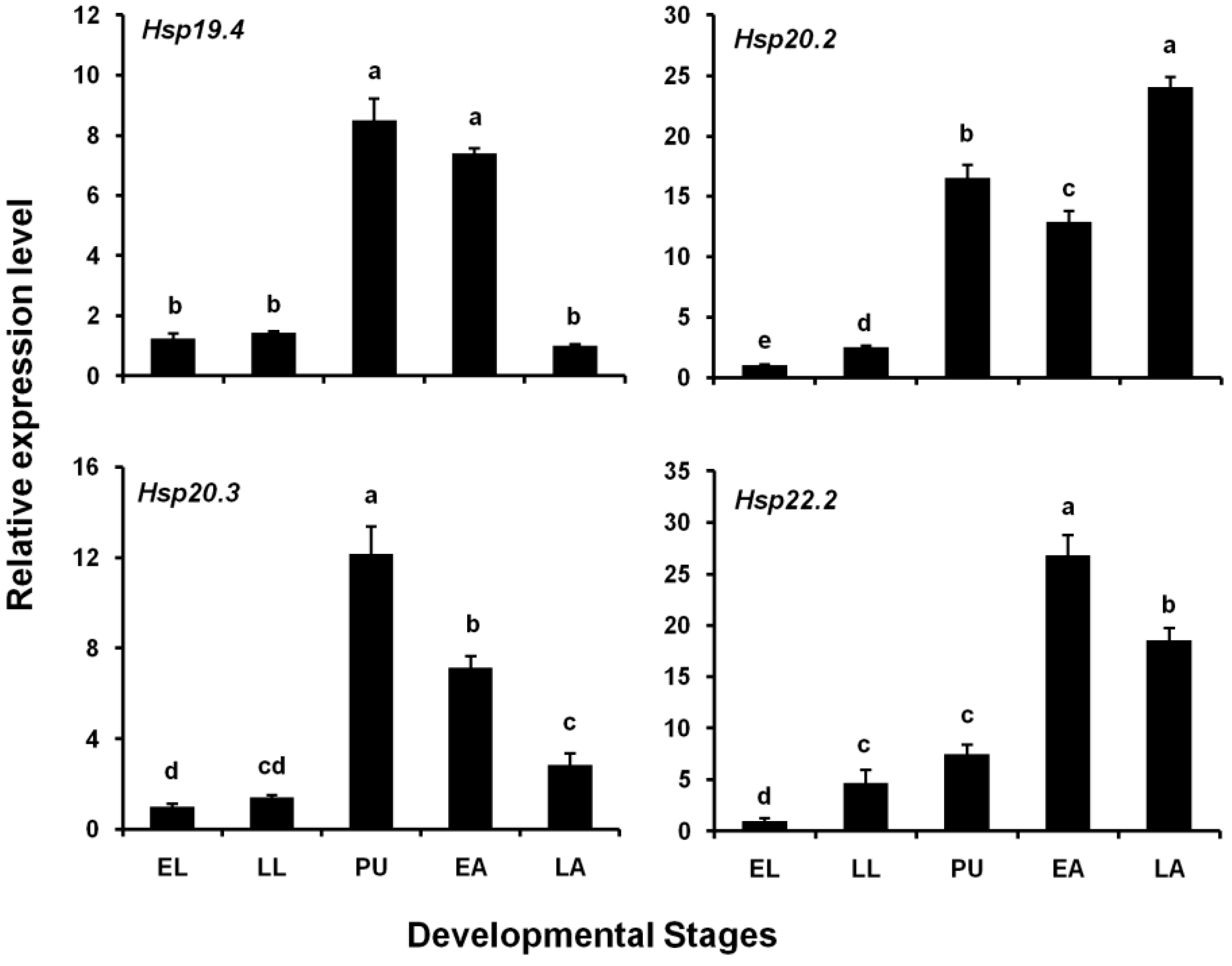
3.2. Expression Levels of Four LsHsp Genes at Different Developmental Stages
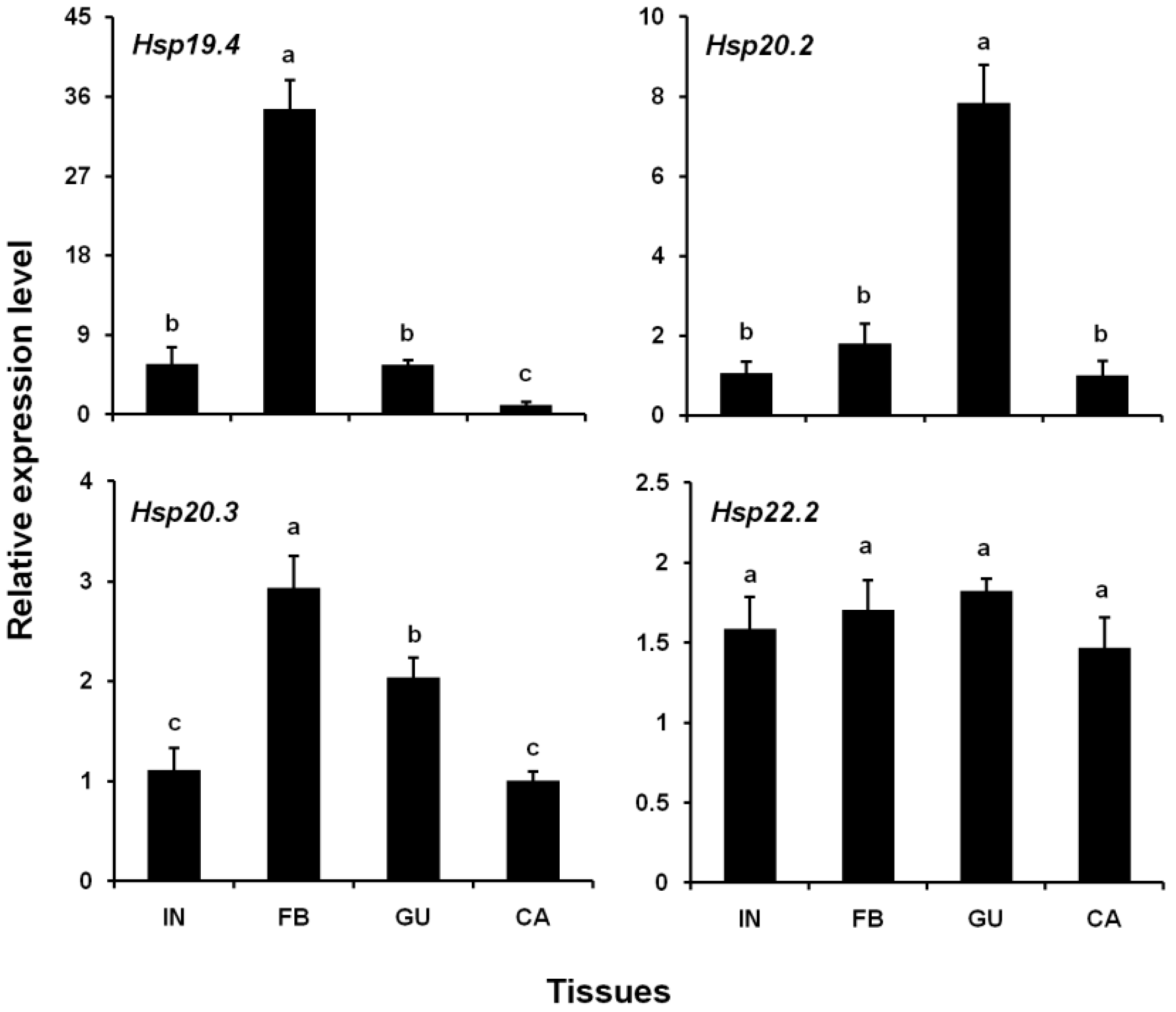
3.3. Expression Levels of Four LsHsp Genes in Different Tissues
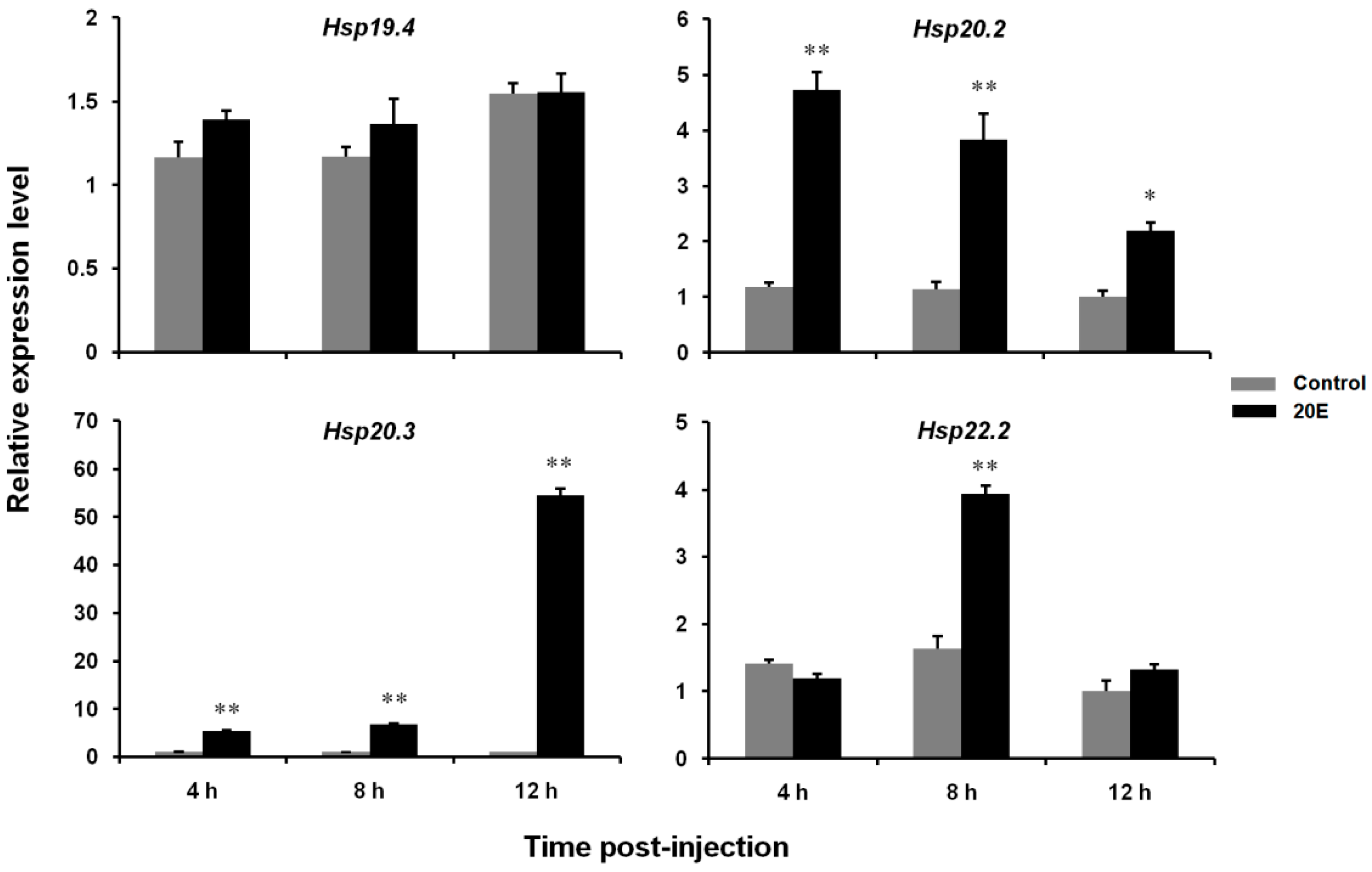
3.4. Effects of 20E Treatment on the Expression Levels of Four LsHsp Genes
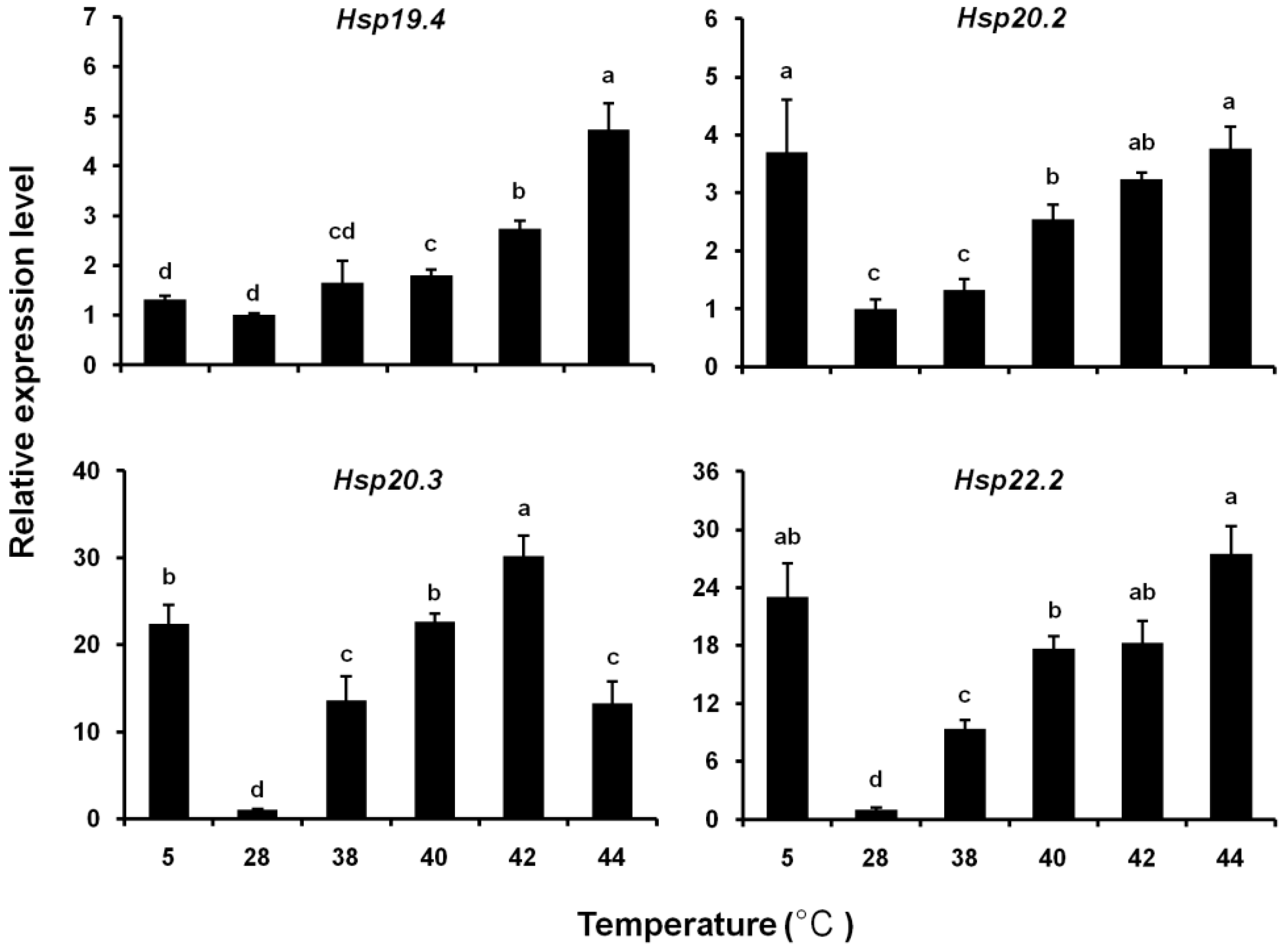
3.5. Expression Levels of Four LsHsp Genes in Response to Thermal Stress
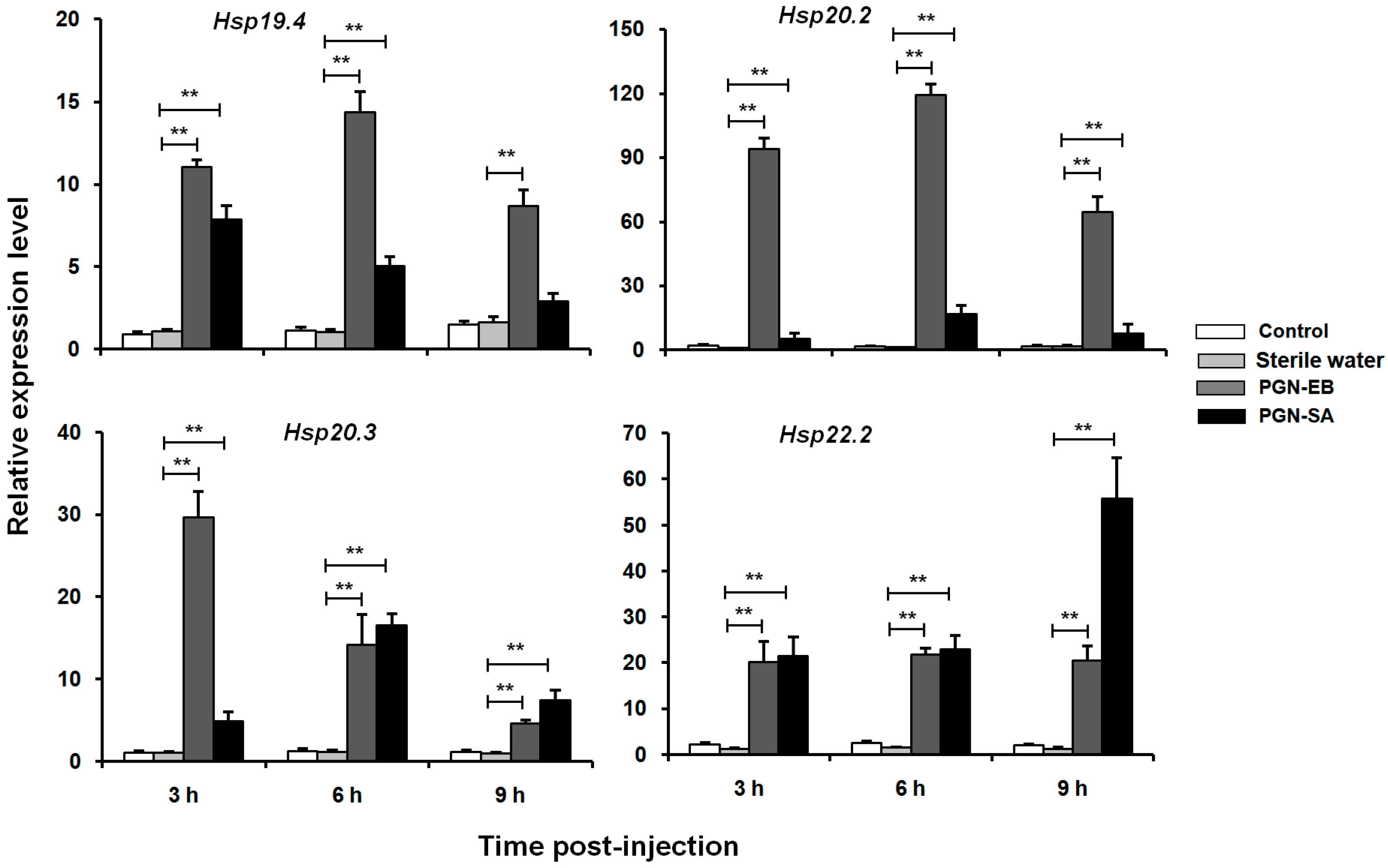
3.6. Expression Levels of Four LsHsp Genes in Response to Immune Challenges
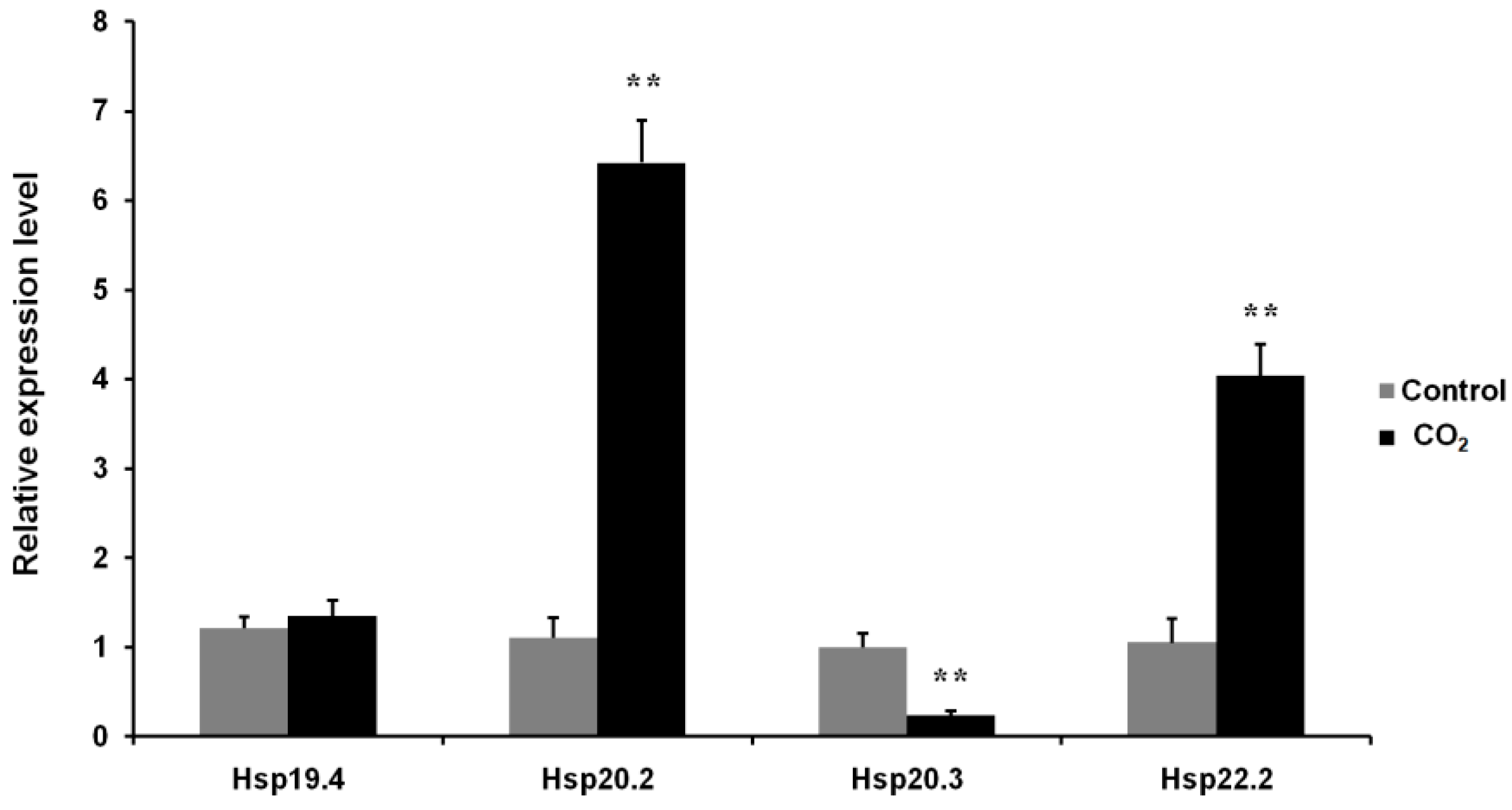
3.7. Effects of CO2 Exposure on the Expression Levels of Four LsHsp Genes
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Johnston, J.A.; Ward, C.L.; Kopito, R.R. Aggresomes: A cellular response to misfolded proteins. J. Cell Biol. 1998, 143, 1883–1898. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Ye, K.; Tu, X.F.; Lu, Y.J.; Thakur, K.; Wei, Z.J. Identification and expression analysis of four heat shock protein genes associated with thermal stress in rice weevil, Sitophilus oryzae. J. Asia-Pac. Entomol. 2018, 21, 872–879. [Google Scholar] [CrossRef]
- Xu, Q.; Zou, Q.; Zheng, H.; Zhang, F.; Tang, B.; Wang, S.G. Three heat shock proteins from Spodoptera exigua: Gene cloning, characterization and comparative stress response during heat and cold shocks. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2011, 159, 92–102. [Google Scholar] [CrossRef]
- Sang, W.; Ma, W.H.; Qiu, L.; Zhu, Z.H.; Lei, C.L. The involvement of heat shock protein and cytochrome P450 genes in response to UV-A exposure in the beetle Tribolium castaneum. J. Insect Physiol. 2012, 58, 830–836. [Google Scholar] [CrossRef]
- Zhang, Y.P.; Liu, Y.M.; Zhang, J.Z.; Guo, Y.P.; Ma, E.B. Molecular cloning and mRNA expression of heat shock protein genes and their response to cadmium stress in the grasshopper Oxya chinensis. PLoS ONE 2015, 10, e0131244. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Li, K.; Zhu, J.Y.; Fang, Q.; Ye, G.Y. Cloning and expression pattern of heat shock protein genes from the endoparasitoid wasp, Pteromalus puparum in response to environmental stresses. Arch. Insect Biochem. Physiol. 2012, 79, 247–263. [Google Scholar] [CrossRef]
- Planelló, R.; Herrero, O.; Martínez-Guitarte, J.L.; Morcillo, G. Comparative effects of butyl benzyl phthalate (BBP) and di (2-ethylhexyl) phthalate (DEHP) on the aquatic larvae of Chironomus riparius based on gene expression assays related to the endocrine system, the stress response and ribosomes. Aquat. Toxicol. 2011, 105, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.M.; Zhang, S.; Timakov, B.; Zhang, P. The Hsp27 gene is not required for Drosophila development but its activity is associated with starvation resistance. Cell Stress Chaperones 2007, 12, 364–372. [Google Scholar] [CrossRef]
- Liu, Z.H.; Xi, D.M.; Kang, M.J.; Guo, X.Q.; Xu, B.H. Molecular cloning and characterization of Hsp27.6: The first reported small heat shock protein from Apis cerana cerana. Cell Stress Chaperones 2012, 17, 539–551. [Google Scholar] [CrossRef]
- Hendrick, J.P.; Hartl, F.U. Molecular chaperone functions of heat-shock proteins. Annu. Rev. Biochem. 1993, 62, 349–384. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.K.; Kim, R.; Kim, S.H. Crystal structure of a small heat-shock protein. Nature. 1998, 394, 595–599. [Google Scholar] [CrossRef] [PubMed]
- Garrido, C.; Paul, C.; Seigneuric, R.; Kampinga, H.H. The small heat shock proteins family: The long forgotten chaperones. Int. J. Biochem. Cell. Biol. 2012, 44, 1588–1592. [Google Scholar] [CrossRef]
- Franck, E.; Madsen, O.; van Rheede, T.; Ricard, G.; Huynen, M.A.; de Jong, W.W. Evolutionary diversity of vertebrate small heat shock proteins. J. Mol. Evol. 2004, 59, 792–805. [Google Scholar] [CrossRef] [PubMed]
- Basha, E.; Friedrich, K.L.; Vierling, E. The N-terminal arm of small heat shock proteins is important for both chaperone activity and substrate specificity. J. Biol. Chem. 2006, 281, 39943–39952. [Google Scholar] [CrossRef] [PubMed]
- Bagnéris, C.; Bateman, O.A.; Naylor, C.E.; Cronin, N.; Boelens, W.C.; Keep, N.H.; Slingsby, C. Crystal structures of alpha-crystallin domain dimers of alphaB-crystallin and Hsp20. J. Mol. Biol. 2009, 392, 1242–1252. [Google Scholar] [CrossRef]
- Kostenko, S.; Moens, U. Heat shock protein 27 phosphorylation: Kinases, phosphatases, functions and pathology. Cell. Mol. Life Sci. 2009, 66, 3289–3307. [Google Scholar] [CrossRef] [PubMed]
- MacRae, T.H. Structure and function of small heat shock/α-crystallin proteins: Established concepts and emerging ideas. Cell. Mol. Life Sci. 2000, 57, 899–913. [Google Scholar] [CrossRef] [PubMed]
- Reineke, A. Identification and expression of a small heat shock protein in two lines of the endoparasitic wasp Venturia canescens. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2005, 141, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Arrigo, A.P. sHsp as novel regulators of programmed cell death and tumorigenicity. Pathol. Biol. 2000, 48, 280–288. [Google Scholar]
- Tsvetkova, N.M.; Horvath, I.; Torok, Z.; Wolkers, W.F.; Balogi, Z.; Shigapova, N.; Crowe, L.M.; Tablin, F.; Vierling, E.; Crowe, J.H.; et al. Small heat-shock proteins regulate membrane lipid polymorphism. Proc. Natl. Acad. Sci. USA 2002, 99, 13504–13509. [Google Scholar] [CrossRef]
- Morrow, G.; Battistini, S.; Zhang, P.; Tanguay, R.M. Decreased lifespan in the absence of expression of the mitochondrial small heat shock protein Hsp22 in Drosophila. J. Biol. Chem. 2004, 279, 43382–43385. [Google Scholar] [CrossRef] [PubMed]
- Gkouvitsas, T.; Kontogiannatos, D.; Kourti, A. Differential expression of two small Hsps during diapause in the corn stalk borer Sesamia nonagrioides (Lef.). J. Insect Physiol. 2008, 54, 1503–1510. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.W.; Li, X.; Yu, Q.Y.; Xiang, Z.H.; Kishino, H.; Zhang, Z. The small heat shock protein (sHSP) genes in the silkworm, Bombyx mori, and comparative analysis with other insect sHSP genes. BMC Evol. Biol. 2009, 9, 215. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.E.; Zhang, Y.L. Identification of multiple small heat-shock protein genes in Plutella xylostella (L.) and their expression profiles in response to abiotic stresses. Cell Stress Chaperones 2015, 20, 23–35. [Google Scholar] [CrossRef] [PubMed]
- Quan, G.X.; Duan, J.; Ladd, T.; Krell, P.J. Identification and expression analysis of multiple small heat shock protein genes in spruce budworm, Choristoneura fumiferana (L.). Cell Stress Chaperones 2018, 23, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.R.; Wang, C.; Ban, F.X.; Zhu, D.T.; Liu, S.S.; Wang, X.W. Genome-wide identification and characterization of HSP gene superfamily in whitefly (Bemisia tabaci) and expression profiling analysis under temperature stress. Insect Sci. 2019, 26, 44–57. [Google Scholar] [CrossRef]
- Wang, L.H.; Zhang, Y.L.; Pan, L.; Wang, Q.; Han, Y.C.; Niu, H.T.; Shan, D.; Hoffmann, A.; Fang, J.C. Induced expression of small heat shock proteins is associated with thermotolerance in female Laodelphax striatellus planthoppers. Cell Stress Chaperones 2018. [Google Scholar] [CrossRef]
- Yocum, G.D.; Joplin, K.H.; Denlinger, D.L. Upregulation of a 23 kDa small heat shock protein transcript diapause in the flesh fly, Sarcophaga crassipalpis. Insect Biochem. Mol. Biol. 1998, 28, 677–682. [Google Scholar] [CrossRef]
- Morrow, G.; Samson, M.; Michaud, S.; Tanguay, R.M. Overexpression of the small mitochondrial Hsp22 extends Drosophila life span and increases resistance to oxidative stress. FASEB J. 2004, 18, 598–599. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Hu, X.X.; Zhai, M.F.; Yu, X.J.; Song, X.W.; Gao, S.S.; Wu, W.; Li, B. Characterization and functional analysis of hsp18.3 gene in the red flour beetle, Tribolium castaneum. Insect Sci. 2017. [Google Scholar] [CrossRef]
- Zhang, Y.Y.; Liu, Y.L.; Guo, X.L.; Li, Y.L.; Gao, H.R.; Guo, X.Q.; Xu, B.H. sHsp22.6, an intronless small heat shock protein gene, is involved in stress defence and development in Apis cerana cerana. Insect Biochem. Mol. Biol. 2014, 53, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, J.R. The biology of Lasioderma serricorne. J. Stored Prod. Res. 1993, 29, 291–303. [Google Scholar] [CrossRef]
- Mahroof, R.M.; Phillips, T.W. Life history parameters of Lasioderma serricorne (F.) as influenced by food sources. J. Stored Prod. Res. 2008, 44, 219–226. [Google Scholar] [CrossRef]
- Li, C.; Li, Z.Z.; Cao, Y.; Zhou, B.; Zheng, X.W. Partial characterization of stress-induced carboxylesterase from adults of Stegobium paniceum and Lasioderma serricorne (Coleoptera: Anobiidae) subjected to CO2-enriched atmosphere. J. Pest Sci. 2009, 82, 7–11. [Google Scholar] [CrossRef]
- Minor, M.F. Do adult cigarette beetle feed? Tob. Sci. 1979, 23, 61–64. [Google Scholar]
- Riudavets, J.; Salas, I.; Pons, M.J. Damage characteristics produced by insect pests in packaging film. J. Stored Prod. Res. 2007, 43, 564–570. [Google Scholar] [CrossRef]
- Allahvaisi, S. Controlling Lasioderma serricorne F. (Col.: Anobiidae) by fumigation and packaging. World Appl. Sci. J. 2013, 28, 1983–1988. [Google Scholar]
- Kim, M.; Yang, J.; Lee, H. Phototactic behavior: Repellent effects of cigarette beetle, Lasioderma serricorne (Coleoptera: Anobiidae), to light-emitting diodes. J. Korean Soc. Appl. Biol. Chem. 2013, 56, 331–333. [Google Scholar] [CrossRef]
- Rajendran, S.; Narasimhan, K.S. Phosphine resistance in the cigarette beetle Lasioderma serricorne (Coleoptera: Anobiidae) and overcoming control failures during fumigation of stored tobacco. Int. J. Pest Manag. 1994, 40, 207–210. [Google Scholar] [CrossRef]
- Sağlam, Ö.; Edde, P.A.; Phillips, T.W. Resistance of Lasioderma serricorne (Coleoptera: Anobiidae) to fumigation with phosphine. J. Econ. Entomol. 2015, 108, 2489–2495. [Google Scholar] [CrossRef] [PubMed]
- Collins, D.; Conyers, S. The effect of sub-zero temperatures on different lifestages of Lasioderma serricorne (F.) and Ephestia elutella (Hübner). J. Stored Prod. Res. 2010, 46, 234–241. [Google Scholar] [CrossRef]
- Yu, C.; Subramanyam, B.; Flinn, P.W.; Gwirtz, J.A. Susceptibility of Lasioderma serricorne (Coleoptera: Anobiidae) life stages to elevated temperatures used during structural heat treatments. J. Econ. Entomol. 2011, 104, 317–324. [Google Scholar] [CrossRef]
- Imai, T. The additive effect of carbon dioxide on mortality of the cigarette beetle Lasioderma serricorne (Coleoptera: Anobiidae) in low-oxygen atmospheres. Appl. Entomol. Zool. 2015, 50, 11–15. [Google Scholar] [CrossRef]
- Combet, C.; Blanchet, C.; Geourjon, C.; Deléage, G. NPS@: Network protein sequence analysis. Trends Biochem. Sci. 2000, 291, 147–150. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Gu, J.; Huang, L.H.; Zheng, S.C.; Liu, L.; Xu, W.H.; Feng, Q.L.; Kang, L. Cloning and expression analysis of six small heat shock protein genes in the common cutworm, Spodoptera litura. J. Insect Physiol. 2011, 57, 908–914. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Zheng, J.; Peng, Y.; Liu, X.; Hoffmann, A.A.; Ma, C.S. Stress responses of small heat shock protein genes in Lepidoptera point to limited conservation of function across phylogeny. PLoS ONE 2015, 10, e0132700. [Google Scholar] [CrossRef]
- Lu, M.X.; Hua, J.; Cui, Y.D.; Du, Y.Z. Five small heat shock protein genes from Chilo suppressalis: Characteristics of gene, genomic organization, structural analysis, and transcription profiles. Cell Stress Chaperones 2014, 19, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Kou, L.H.; Wu, H.H.; Liu, Y.M.; Zhang, Y.P.; Zhang, J.Z.; Guo, Y.P.; Ma, E.B. Molecular characterization of six small heat shock proteins and their responses under cadmium stress in Oxya chinensis (Orthoptera: Acridoidea). Environ. Entomol. 2016, 45, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Dou, W.; Tian, Y.; Liu, H.; Shi, Y.; Smagghe, G.; Wang, J.J. Characteristics of six small heat shock protein genes from Bactrocera dorsalis: Diverse expression under conditions of thermal stress and normal growth. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2017, 213, 8–16. [Google Scholar] [CrossRef] [PubMed]
- Mason, P.J.; Hall, L.M.C.; Gausz, J. The expression of heat shock genes during normal development in Drosophila melanogaster (heat shock/abundant transcripts/developmental regulation). Mol. Gen. Genet. 1984, 194, 73–78. [Google Scholar] [CrossRef]
- Sonoda, S.; Ashfaq, M.; Tsumuki, H. Cloning and nucleotide sequencing of three heat shock protein genes (hsp90, hsc70, and hsp19.5) from the diamondback moth, Plutella xylostella (L.) and their expression in relation to developmental stage and temperature. Arch. Insect Biochem. Physiol. 2006, 62, 80–90. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.H.; Wang, C.Z.; Kang, L. Cloning and expression of five heat shock protein genes in relation to cold hardening and development in the leafminer. Liriomyza sativa. J. Insect Physiol. 2009, 55, 279–285. [Google Scholar] [CrossRef]
- Gu, J.; Huang, L.X.; Shen, L.H.; Feng, Q.L. Hsp70 and small Hsps are the major heat shock protein members involved in midgut metamorphosis in the common cutworm, Spodoptera litura. Insect Mol. Biol. 2012, 21, 535–543. [Google Scholar] [CrossRef] [PubMed]
- Graczynski, S.F.; Unruh, T.R.; Guédot, C.; Neven, L.G. Characterization of three transcripts encoding small heat shock proteins expressed in the codling moth, Cydia pomonella (Lepidoptera: Tortricidae). Insect Sci. 2011, 18, 473–483. [Google Scholar] [CrossRef]
- Wang, H.J.; Shi, Z.K.; Shen, Q.D.; Xu, C.D.; Wang, B.; Meng, Z.J.; Wang, S.G.; Tang, B.; Wang, S. Molecular cloning and induced expression of six small heat shock proteins mediating cold-hardiness in Harmonia axyridis (Coleoptera: Coccinellidae). Front. Physiol. 2017, 8, 60. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Lu, M.X.; Tang, X.T.; Du, Y.Z. Characterization and expression of genes encoding three small heat shock proteins in Sesamia inferens (Lepidoptera: Noctuidae). Int. J. Mol. Sci. 2014, 15, 23196–23211. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Fang, Y.; Bao, Z.Z.; Jin, X.; Zhu, W.J.; Wang, L.P.; Liu, T.; Ji, H.P.; Wang, H.Y.; Xu, S.Q.; et al. Identification of a Bombyx mori gene encoding small heat shock protein BmHsp27.4 expressed in response to high-temperature stress. Gene 2014, 538, 56–62. [Google Scholar] [CrossRef]
- Lezzi, M. Chromosome puffing: Supramolecular aspects of ecdysone action. In Metamorphosis: Post-Embryonic Reprogramming of Gene Expression in Amphibian and Insect Cells; Gilbert, L.I., Tata, J.R., Atkinson, B.G., Eds.; Academic Press: New York, NY, USA, 1996; pp. 145–173. [Google Scholar]
- Antoniewski, C.; Laval, M.; Lepesant, J.A. Structural features critical to the activity of an ecdysone receptor binding site. Insect Biochem. Mol. Biol. 1993, 23, 105–114. [Google Scholar] [CrossRef]
- Kokolakis, G.; Tatari, M.; Zacharopoulou, A.; Mintzas, A.C. The hsp27 gene of the Mediterranean fruit fly, Ceratitis capitata: Structural characterization, regulation and developmental expression. Insect Mol. Biol. 2008, 17, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Dubrovsky, E.B.; Dubrovskaya, V.A.; Berger, E.M. Selective binding of Drosophila BR-C isoforms to a distal regulatory element in the hsp23 promoter. Insect Biochem. Mol. Biol. 2001, 3, 1231–1239. [Google Scholar] [CrossRef]
- Liu, Z.H.; Yao, P.B.; Guo, X.Q.; Xu, B.H. Two small heat shock protein genes in Apis cerana cerana: Characterization, regulation, and developmental expression. Gene 2014, 545, 205–214. [Google Scholar] [CrossRef]
- Wu, P.; Wang, X.; Qin, G.X.; Liu, T.; Jiang, Y.F.; Li, M.W.; Guo, X.J. Microarray analysis of the gene expression profile in the midgut of silkworm infected with cytoplasmic polyhedrosis virus. Mol. Biol. Rep. 2011, 38, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.F.; Dai, L.S.; Wang, L.; Qian, C.; Wei, G.Q.; Li, J.; Zhu, B.J.; Liu, C.L. Eicosanoids mediate sHSP 20.8 gene response to biotic stress in larvae of the Chinese oak silkworm Antheraea pernyi. Gene 2015, 562, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.F.; Xie, J.; Wei, L.T.; Zhang, S.S.; Song, X.W.; Gao, S.S.; Li, B. Transcriptome analysis of hsp18.3 functions and regulatory systems using RNA-sequencing in the red flour beetle, Tribolium castaneum. J. Integr. Agric. 2018, 17, 1040–1056. [Google Scholar] [CrossRef]
- Gunasekaran, N.; Rajendran, S. Toxicity of carbon dioxide to drugstore beetle Stegobium paniceum and cigarette beetle Lasioderma serricorne. J. Stored Prod. Res. 2005, 41, 283–294. [Google Scholar] [CrossRef]
- Kennington, G.S.; Cannel, S. Biochemical correlates of respiratory and developmental changes in anoxic Tribolium confusum pupae. Physiol. Zool. 1967, 40, 403–408. [Google Scholar] [CrossRef]
- Xu, K.K.; Ding, T.B.; Yan, Y.; Li, C.; Yang, W.J. Expression analysis of glutathione S-transferase genes in Lasioderma serricorne (Coleoptera: Anobiidae) subjected to CO2-enriched atmosphere. J. Zhejiang Univ. Agric. Life Sci. 2017, 43, 599–607. [Google Scholar]
| Application of Primers | Gene | Primer Sequence (5′ to 3′) |
|---|---|---|
| Full-length confirmation | LsHsp19.4 | F: TGATAAACGACCCTCTGGAT R: ATGTACCCGTGTTCGTCTTG |
| LsHsp20.2 | F: CATTCTACGACACCTTTGAC R: AATGGAAGCTGCATCAACTT | |
| LsHsp20.3 | F: TTGTCTCCATTCTACGACGT R: CCTTTTGGCAACTTATACTT | |
| LsHsp22.2 | F: TTCCTATATTCCGACCCCAT R: TGATCTTCAACACACCATCA | |
| qPCR analysis | LsHsp19.4 | F: CTGGGAGGATGATGACGACT R: ACATCGACGTTCACACGAAA |
| LsHsp20.2 | F: ATTTCCAGGCGATTTGTGAG R: CTTTGCTTGGCTTTCCAGTC | |
| LsHsp20.3 | F: AAACACGAGGAGAAGCAGGA R: CTGGTTTCCCAGTTTGCTGT | |
| LsHsp22.2 | F: AAAGACGCTGGCTCAACTGT R: TCCTTTCGGCAGAATGTACC | |
| 18S | F: GTTGATCACGTCGCAAGCTA R: AGGTTTCCCTCTGGCTTGTT |
| Gene | Sequence Length (bp) | Protein Length (aa) | Molecular Weight (kDa) | Isoelectric Point | GenBank Accession Number |
|---|---|---|---|---|---|
| LsHsp19.4 | 510 | 169 | 19.4 | 5.94 | MK395540 |
| LsHsp20.2 | 543 | 180 | 20.2 | 6.60 | MK395541 |
| LsHsp20.3 | 546 | 181 | 20.3 | 6.32 | MK395542 |
| LsHsp22.2 | 585 | 194 | 22.2 | 6.04 | MK395543 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, W.-J.; Xu, K.-K.; Cao, Y.; Meng, Y.-L.; Liu, Y.; Li, C. Identification and Expression Analysis of Four Small Heat Shock Protein Genes in Cigarette Beetle, Lasioderma serricorne (Fabricius). Insects 2019, 10, 139. https://doi.org/10.3390/insects10050139
Yang W-J, Xu K-K, Cao Y, Meng Y-L, Liu Y, Li C. Identification and Expression Analysis of Four Small Heat Shock Protein Genes in Cigarette Beetle, Lasioderma serricorne (Fabricius). Insects. 2019; 10(5):139. https://doi.org/10.3390/insects10050139
Chicago/Turabian StyleYang, Wen-Jia, Kang-Kang Xu, Yu Cao, Yong-Lu Meng, Yan Liu, and Can Li. 2019. "Identification and Expression Analysis of Four Small Heat Shock Protein Genes in Cigarette Beetle, Lasioderma serricorne (Fabricius)" Insects 10, no. 5: 139. https://doi.org/10.3390/insects10050139
APA StyleYang, W.-J., Xu, K.-K., Cao, Y., Meng, Y.-L., Liu, Y., & Li, C. (2019). Identification and Expression Analysis of Four Small Heat Shock Protein Genes in Cigarette Beetle, Lasioderma serricorne (Fabricius). Insects, 10(5), 139. https://doi.org/10.3390/insects10050139






