Applicability of Precision Medicine Approaches to Managing Hypertension in Rural Populations
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Approach
2.1.1. Description of High Blood Pressure Study
2.1.2. Description of Lifestyle Study
2.1.3. Study Measures
2.2. Genomic and Experimental Analyses
2.2.1. DNA Isolation, Purification, and Quality Control
2.2.2. Genotyping on the Illumina Platform
2.2.3. SNP-Level Analysis of Admixture and Relatedness
2.2.4. Imputing Single Nucleotide Polymorphisms
2.2.5. Pre-Modeling Activities
2.2.6. Pre-Modeling Activities
2.2.7. Baseline Systolic Blood Pressure Association Testing
2.3. Human Studies
3. Results
3.1. Study Population
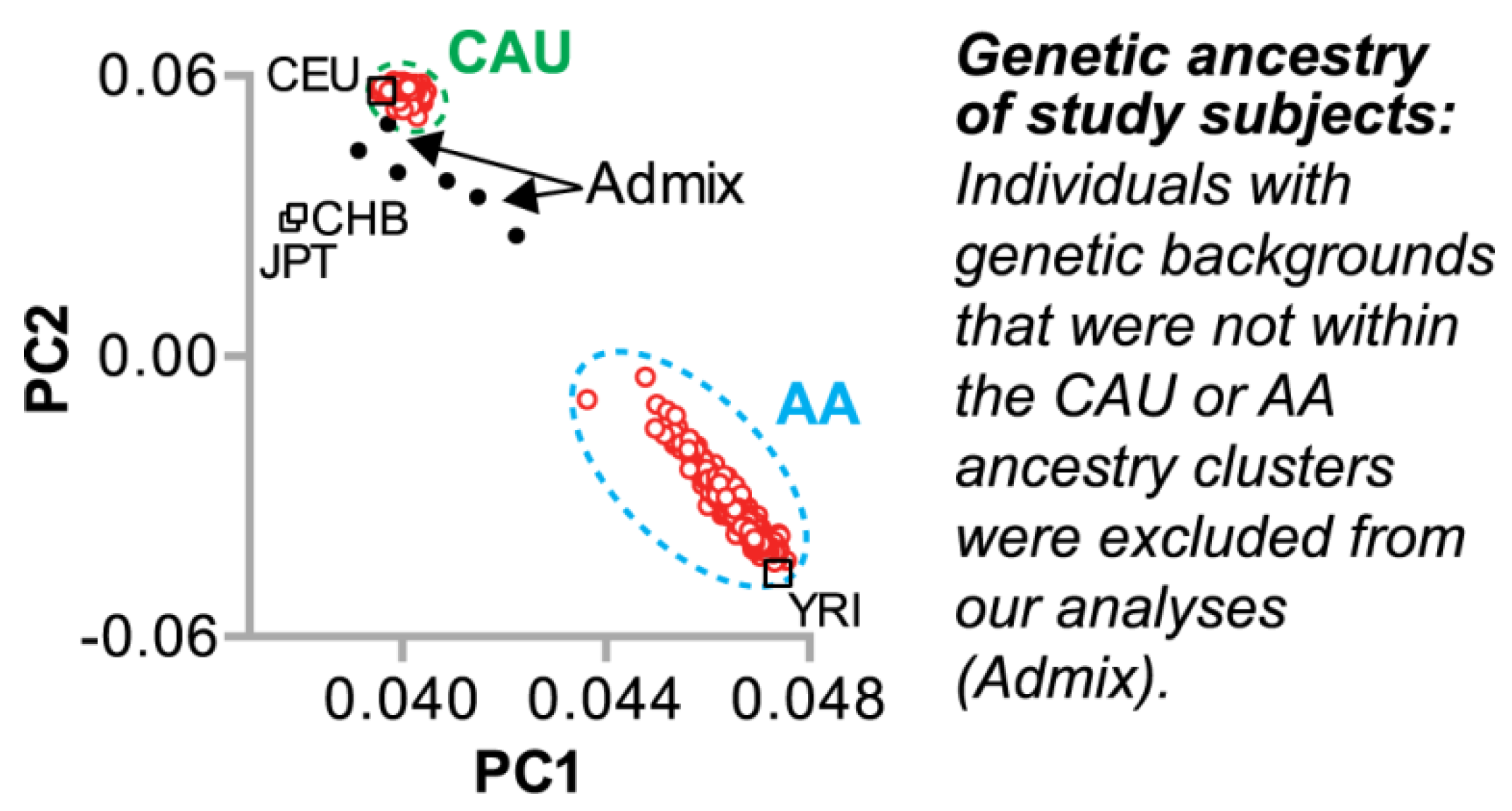
3.1.1. Genetic Ancestry of the Study Population
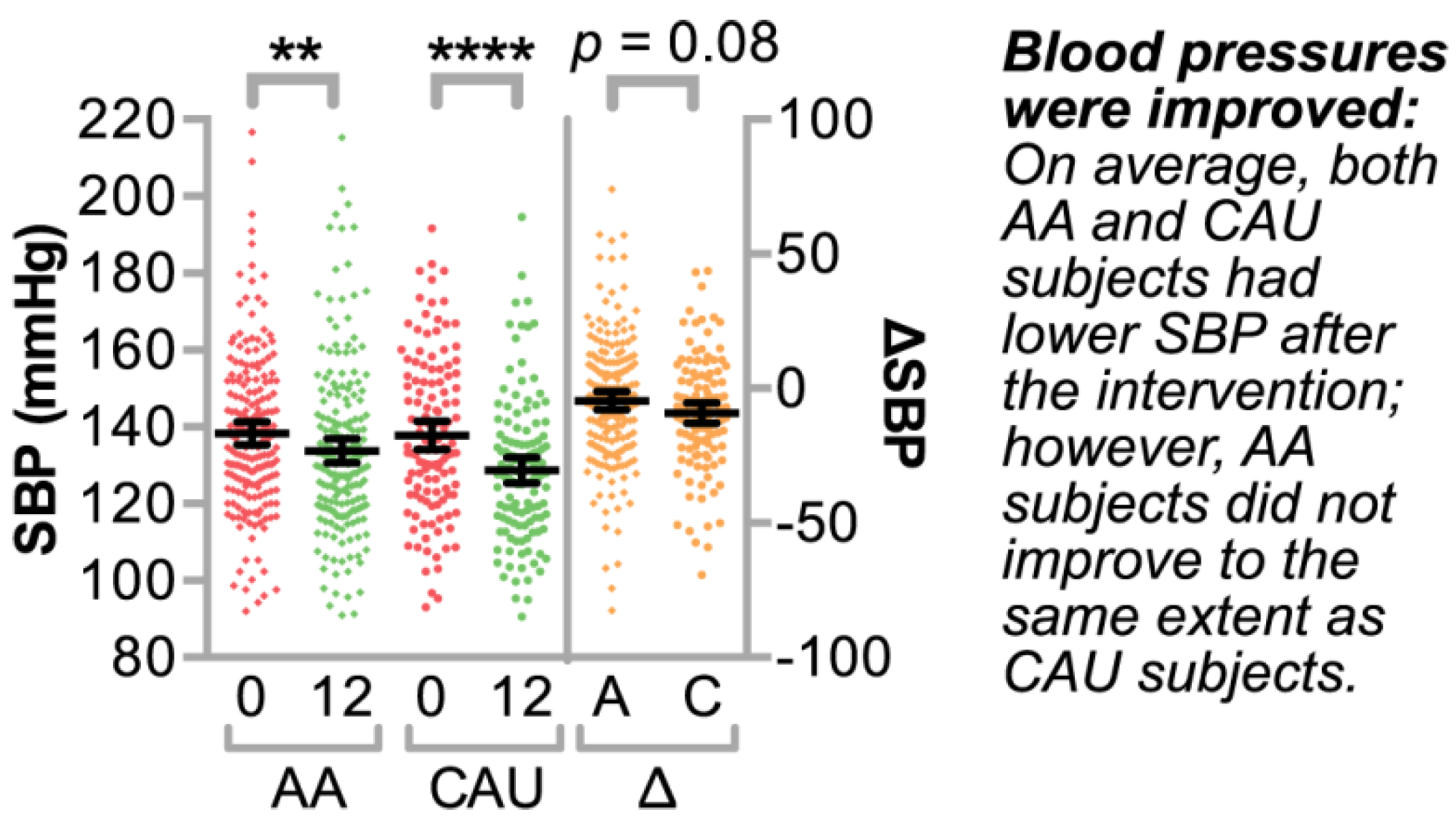
3.1.2. Clinical Characteristics of the Study Population
3.2. Identifying Clinically Relevant Variables that Associate with Systolic Blood Pressure
Bivariate and Multivariable Analyses of Clinical Variable Associations with ΔSBP over One Year
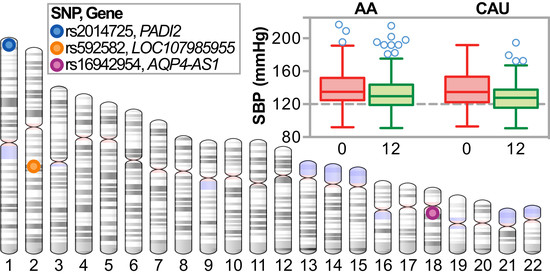
3.3. Association Analysis of Candidate Blood Pressure Polymorphisms in a Rural Population
3.4. Genome-Wide Association Analysis of Genetic Variation with ΔSBP
3.4.1. Single Nucleotide Polymorphism Associations with ΔSBP
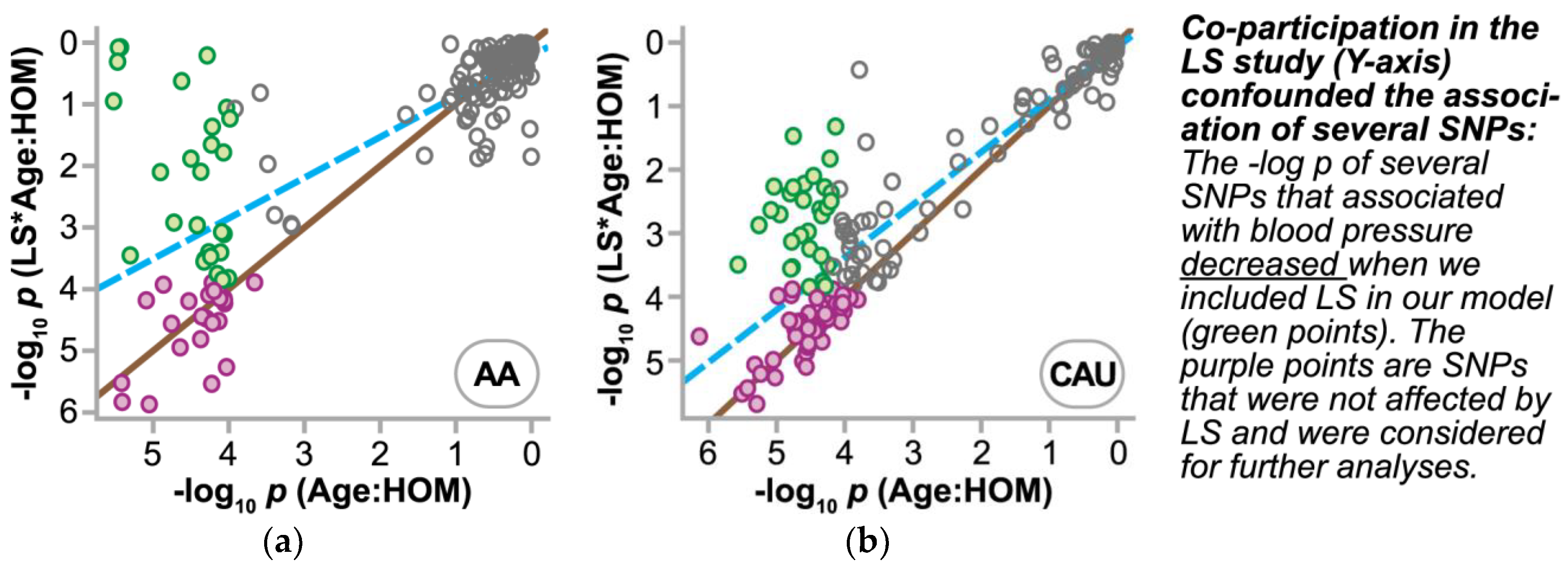
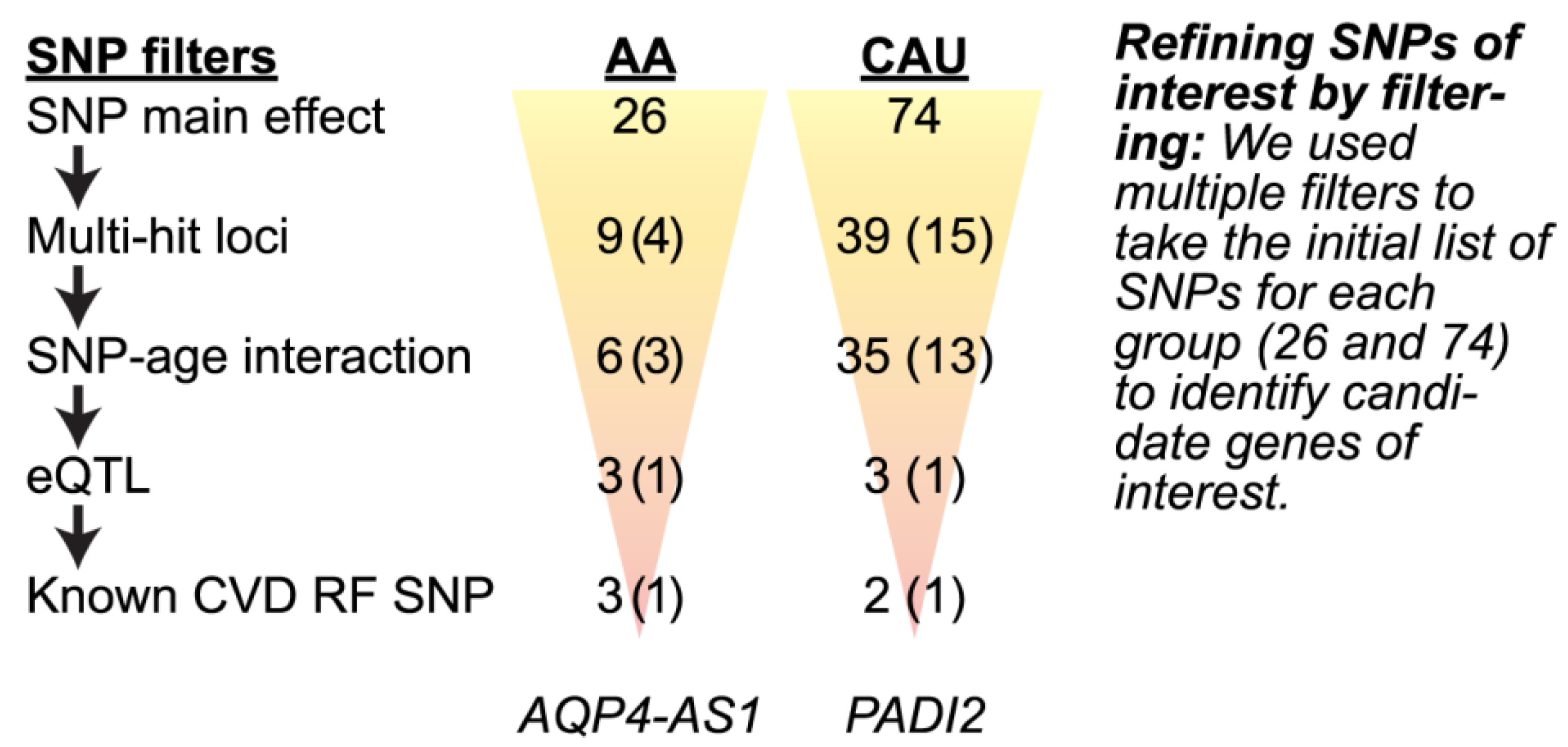
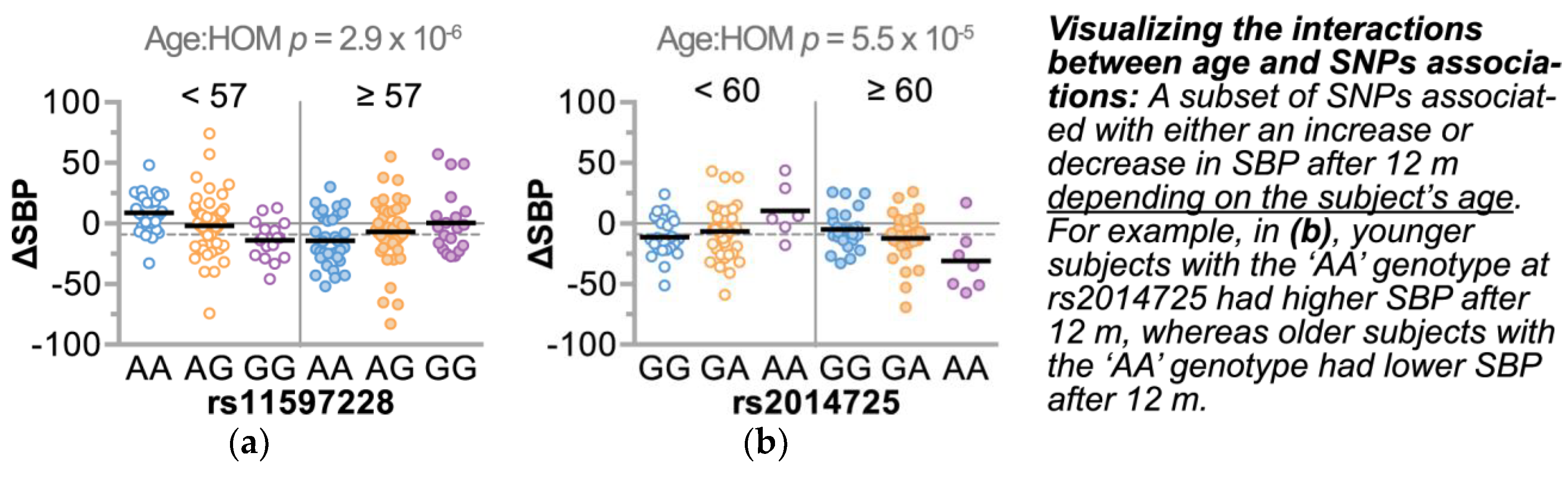
3.4.2. Refining SNPs of Interest and Identifying SNP-Age Interactions
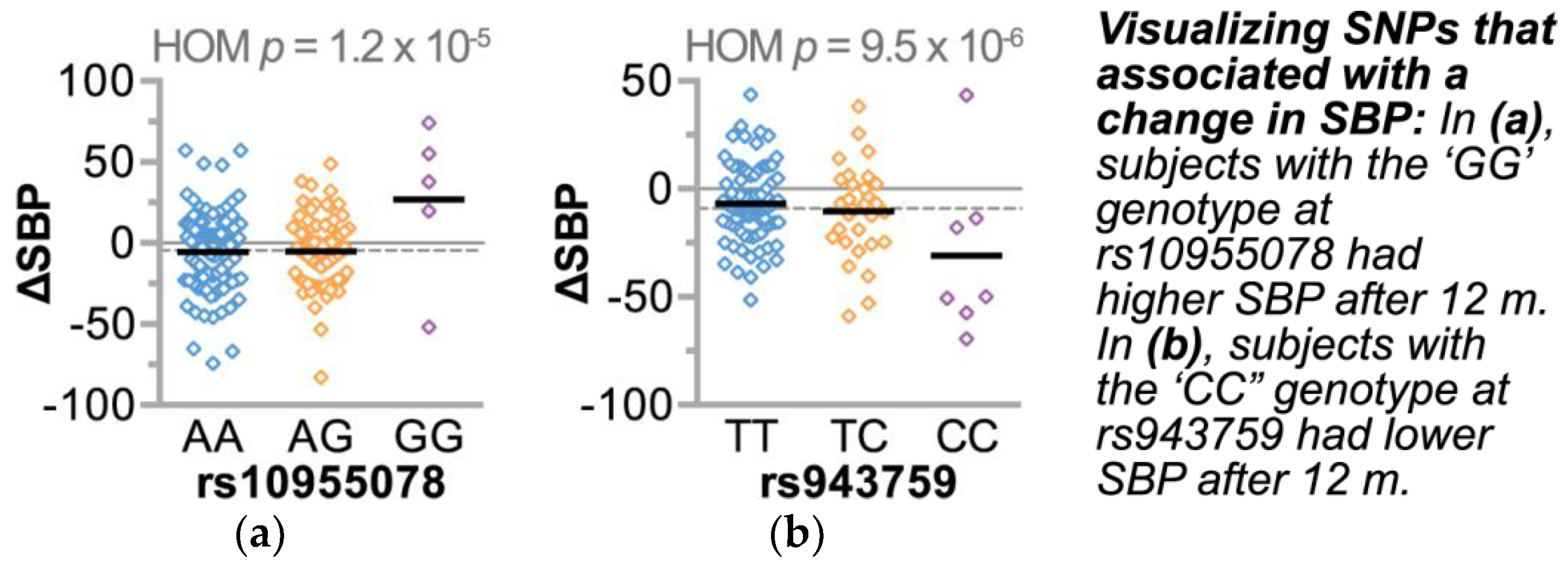
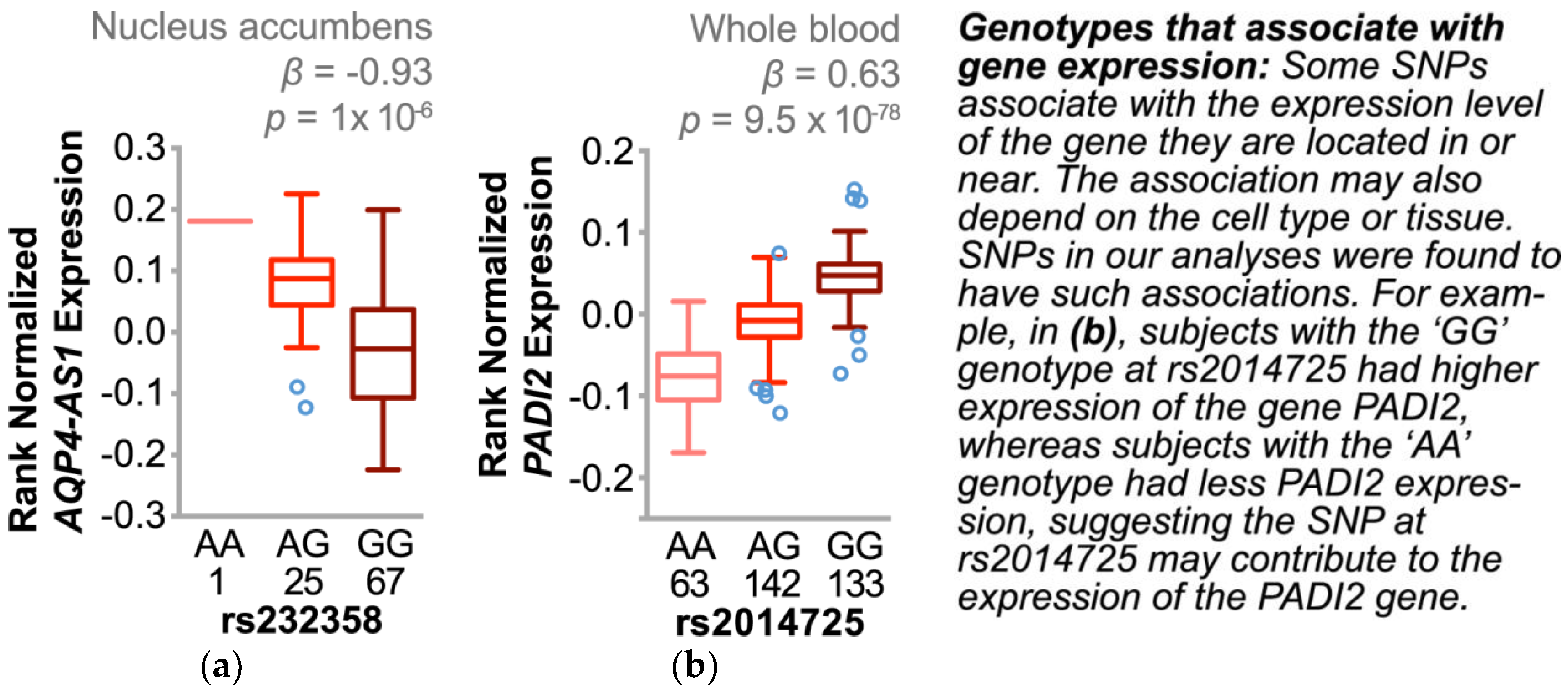
3.4.3. Single Nucleotide Polymorphisms Associate with Changes in Gene Expression and Other Cardiovascular Risk Factors
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chobanian, A.V.; Bakris, G.L.; Black, H.R.; Cushman, W.C.; Green, L.A.; Izzo, J.L.; Jones, D.W.; Materson, B.J.; Oparil, S.; Wright, J.T.; et al. Seventh report of the Joint National Committee on Prevention, Detection, Evaluation, and Treatment of High Blood Pressure. Hypertension 2003, 42, 1206–1252. [Google Scholar] [CrossRef] [PubMed]
- Mozaffarian, D.; Benjamin, E.J.; Go, A.S.; Arnett, D.K.; Blaha, M.J.; Cushman, M.; Das, S.R.; de Ferranti, S.; Després, J.P.; Fullerton, H.J.; et al. Heart Disease and Stroke Statistics—2016 Update. Circulation 2016, 133, e38–e360. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Jones, D.; Adams, R.; Carnethon, M.; De Simone, G.; Ferguson, T.B.; Flegal, K.; Ford, E.; Furie, K.; Go, A.; Greenlund, K.; et al. Heart disease and stroke statistics—2009 update: A report from the American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Circulation 2009, 119, 480–486. [Google Scholar] [CrossRef] [PubMed]
- Shea, S.; Misra, D.; Ehrlich, M.H.; Field, L.; Francis, C.K. Predisposing factors for severe, uncontrolled hypertension in an inner-city minority population. N. Engl. J. Med. 1992, 327, 776–781. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention (CDC). Racial/Ethnic disparities in the awareness, treatment, and control of hypertension—United States, 2003–2010. MMWR Morb. Mortal. Wkly. Rep. 2013, 62, 351–355. [Google Scholar]
- Purnell, T.S.; Calhoun, E.A.; Golden, S.H.; Halladay, J.R.; Krok-Schoen, J.L.; Appelhans, B.M.; Cooper, L.A. Achieving Health Equity: Closing The Gaps In Health Care Disparities, Interventions, and Research. Health Aff. (Millwood) 2016, 35, 1410–1415. [Google Scholar] [CrossRef] [PubMed]
- Hill, M.N.; Han, H.R.; Dennison, C.R.; Kim, M.T.; Roary, M.C.; Blumenthal, R.S.; Bone, L.R.; Levine, D.M.; Post, W.S. Hypertension care and control in underserved urban African American men: Behavioral and physiologic outcomes at 36 months. Am. J. Hypertens. 2003, 16, 906–913. [Google Scholar] [CrossRef]
- Rogers, M.A.; Small, D.; Buchan, D.A.; Butch, C.A.; Stewart, C.M.; Krenzer, B.E.; Husovsky, H.L. Home monitoring service improves mean arterial pressure in patients with essential hypertension. A randomized, controlled trial. Ann. Intern. Med. 2001, 134, 1024–1032. [Google Scholar] [CrossRef] [PubMed]
- Mueller, M.; Purnell, T.S.; Mensah, G.A.; Cooper, L.A. Reducing Racial and Ethnic Disparities in Hypertension Prevention and Control: What Will It Take to Translate Research into Practice and Policy? Am. J. Hypertens. 2015, 28, 699–716. [Google Scholar] [CrossRef] [PubMed]
- Safford, M.M.; Shewchuk, R.; Qu, H.; Williams, J.H.; Estrada, C.A.; Ovalle, F.; Allison, J.J. Reasons for Not Intensifying Medications: Differentiating “Clinical Inertia” from Appropriate Care. J. Gen. Intern. Med. 2007, 22, 1648–1655. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, P.J. Overcome clinical inertia to control systolic blood pressure. Arch. Intern. Med. 2003, 163, 2677–2678. [Google Scholar] [CrossRef] [PubMed]
- Simino, J.; Shi, G.; Bis, J.C.; Chasman, D.I.; Ehret, G.B.; Gu, X.; Guo, X.; Hwang, S.J.; Sijbrands, E.; Smith, A.V.; et al. Gene-Age Interactions in Blood Pressure Regulation: A Large-Scale Investigation with the CHARGE, Global BPgen, and ICBP Consortia. Am. J. Hum. Genet. 2014, 95, 24–38. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, F.; Isono, M.; Katsuya, T.; Yamamoto, K.; Yokota, M.; Sugiyama, T.; Nabika, T.; Fujioka, A.; Ohnaka, K.; Asano, H.; et al. Blood pressure and hypertension are associated with 7 loci in the Japanese population. Circulation 2010, 121, 2302–2309. [Google Scholar] [CrossRef] [PubMed]
- Newton-Cheh, C.; Johnson, T.; Gateva, V.; Tobin, M.D.; Bochud, M.; Coin, L.; Najjar, S.S.; Zhao, J.H.; Heath, S.C.; Eyheramendy, S.; et al. Eight blood pressure loci identified by genome-wide association study of 34,433 people of European ancestry. Nat. Genet. 2009, 41, 666–676. [Google Scholar] [CrossRef] [PubMed]
- Levy, D.; Ehret, G.B.; Rice, K.; Verwoert, G.C.; Launer, L.J.; Dehghan, A.; Glazer, N.L.; Morrison, A.C.; Johnson, A.D.; Aspelund, T.; et al. Genome-wide association study of blood pressure and hypertension. Nat. Genet. 2009, 41, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Kato, N.; Takeuchi, F.; Tabara, Y.; Kelly, T.N.; Go, M.J.; Sim, X.; Tay, W.T.; Chen, C.H.; Zhang, Y.; Yamamoto, K.; et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in East Asians. Nat. Genet. 2011, 43, 531–538. [Google Scholar] [CrossRef] [PubMed]
- Wain, L.V.; Verwoert, G.C.; O’Reilly, P.F.; Shi, G.; Johnson, T.; Johnson, A.D.; Bochud, M.; Rice, K.M.; Henneman, P.; Smith, A.V.; et al. Genome-wide association study identifies six new loci influencing pulse pressure and mean arterial pressure. Nat. Genet. 2011, 43, 1005–1011. [Google Scholar] [CrossRef] [PubMed]
- International Consortium for Blood Pressure Genome-Wide Association Studies; Ehret, G.B.; Munroe, P.B.; Rice, K.M.; Bochud, M.; Johnson, A.D.; Chasman, D.I.; Smith, A.V.; Tobin, M.D.; Verwoert, G.C.; et al. Genetic Variants in Novel Pathways Influence Blood Pressure and Cardiovascular Disease Risk. Nature 2011, 478, 103–109. [Google Scholar] [CrossRef] [PubMed]
- Fox, E.R.; Young, J.H.; Li, Y.; Dreisbach, A.W.; Keating, B.J.; Musani, S.K.; Liu, K.; Morrison, A.C.; Ganesh, S.; Kutlar, A.; et al. Association of genetic variation with systolic and diastolic blood pressure among African Americans: The Candidate Gene Association Resource study. Hum. Mol. Genet. 2011, 20, 2273–2284. [Google Scholar] [CrossRef] [PubMed]
- Montasser, M.E.; Shimmin, L.C.; Hanis, C.L.; Boerwinkle, E.; Hixson, J.E. Gene by Smoking Interaction in Hypertension: Identification of a Major QTL on Chromosome 15q for Systolic Blood Pressure in Mexican Americans. J. Hypertens. 2009, 27, 491–501. [Google Scholar] [CrossRef] [PubMed]
- Simino, J.; Rao, D.C.; Freedman, B.I. Novel findings and future directions on the genetics of hypertension. Curr. Opin. Nephrol. Hypertens. 2012, 21, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, D.B. Common genetic variation and human traits. N. Engl. J. Med. 2009, 360, 1696–1698. [Google Scholar] [CrossRef] [PubMed]
- Fuchsberger, C.; Flannick, J.; Teslovich, T.M.; Mahajan, A.; Agarwala, V.; Gaulton, K.J.; Ma, C.; Fontanillas, P.; Moutsianas, L.; McCarthy, D.J.; et al. The genetic architecture of type 2 diabetes. Nature 2016, 536, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Rappaport, S.M. Genetic Factors Are Not the Major Causes of Chronic Diseases. PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Marian, A.J. The personal genome and the practice of cardiovascular medicine. Methodist Debakey Cardiovasc. J. 2010, 6, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Marian, A.J.; Bolli, R. Circulation Research and Human Genetic Studies. Circ. Res. 2010, 107, 6–8. [Google Scholar] [CrossRef] [PubMed]
- Marian, A.J. Challenges in Medical Applications of Whole Exome/Genome Sequencing Discoveries. Trends Cardiovasc. Med. 2012, 22, 219–223. [Google Scholar] [CrossRef] [PubMed]
- Skinner, H.G.; Calancie, L.; Vu, M.B.; Garcia, B.; DeMarco, M.; Patterson, C.; Ammerman, A.; Schisler, J.C. Using Community-Based Participatory Research Principles to Develop More Understandable Recruitment and Informed Consent Documents in Genomic Research. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Halladay, J.R.; Donahue, K.E.; Hinderliter, A.L.; Cummings, D.M.; Cene, C.W.; Miller, C.L.; Garcia, B.A.; Tillman, J.; DeWalt, D. The heart healthy lenoir project-an intervention to reduce disparities in hypertension control: Study protocol. BMC Health Serv. Res. 2013, 13, 441. [Google Scholar] [CrossRef] [PubMed]
- Cené, C.W.; Halladay, J.R.; Gizlice, Z.; Donahue, K.E.; Cummings, D.M.; Hinderliter, A.; Miller, C.; Johnson, L.F.; Garcia, B.; Tillman, J.; et al. A multicomponent quality improvement intervention to improve blood pressure and reduce racial disparities in rural primary care practices. J. Clin. Hypertens. (Greenwich) 2017, 19, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Bosworth, H.B.; Olsen, M.K.; Dudley, T.; Orr, M.; Neary, A.; Harrelson, M.; Adams, M.; Svetkey, L.P.; Dolor, R.J.; Oddone, E.Z. The Take Control of Your Blood pressure (TCYB) study: Study design and methodology. Contemp. Clin. Trials 2007, 28, 33–47. [Google Scholar] [CrossRef] [PubMed]
- Keyserling, T.C.; Samuel-Hodge, C.D.; Pitts, S.J.; Garcia, B.A.; Johnston, L.F.; Gizlice, Z.; Miller, C.L.; Braxton, D.F.; Evenson, K.R.; Smith, J.C.; et al. A community-based lifestyle and weight loss intervention promoting a Mediterranean-style diet pattern evaluated in the stroke belt of North Carolina: The Heart Healthy Lenoir Project. BMC Public Health 2016, 16, 732. [Google Scholar] [CrossRef] [PubMed]
- Halladay, J.R.; Donahue, K.E.; Cené, C.W.; Li, Q.; Cummings, D.M.; Hinderliter, A.L.; Miller, C.L.; Garcia, B.A.; Little, E.; Rachide, M.; et al. The association of health literacy and blood pressure reduction in a cohort of patients with hypertension: The heart healthy lenoir trial. Patient Educ. Couns. 2017, 100, 542–549. [Google Scholar] [CrossRef] [PubMed]
- Donahue, K.E.; Vu, M.B.; Halladay, J.R.; Miller, C.; Garcia, B.A.; Cummings, D.M.; Cene, C.W.; Hinderliter, A.; Little, E.; Rachide, M.; et al. Patient and Practice Perspectives on Strategies for Controlling Blood Pressure, North Carolina, 2010–2012. Prev. Chronic Dis. 2014, 11. [Google Scholar] [CrossRef] [PubMed]
- Fuchsberger, C.; Abecasis, G.R.; Hinds, D.A. minimac2: faster genotype imputation. Bioinformatics 2015, 31, 782–784. [Google Scholar] [CrossRef] [PubMed]
- Browning, B.L.; Browning, S.R. Genotype Imputation with Millions of Reference Samples. Am. J. Hum. Genet. 2016, 98, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Gu, D.; Kelly, T.N.; Hixson, J.E.; Chen, J.; Liu, D.; Chen, J.; Rao, D.C.; Mu, J.; Ma, J.; Jaquish, C.E.; et al. Genetic Variants in the Renin-Angiotensin-Aldosterone System and Salt-Sensitivity of Blood Pressure. J. Hypertens. 2010, 28, 1210–1220. [Google Scholar] [CrossRef] [PubMed]
- Shetty, P.B.; Tang, H.; Tayo, B.O.; Morrison, A.C.; Hanis, C.L.; Rao, D.C.; Young, J.H.; Fox, E.R.; Boerwinkle, E.; Cooper, R.S.; et al. Variants in CXADR and F2RL1 are associated with blood pressure and obesity in African-Americans in regions identified through admixture mapping. J. Hypertens. 2012, 30, 1970–1976. [Google Scholar] [CrossRef] [PubMed]
- Franceschini, N.; Fox, E.; Zhang, Z.; Edwards, T.L.; Nalls, M.A.; Sung, Y.J.; Tayo, B.O.; Sun, Y.V.; Gottesman, O.; Adeyemo, A.; et al. Genome-wide Association Analysis of Blood-Pressure Traits in African-Ancestry Individuals Reveals Common Associated Genes in African and Non-African Populations. Am. J. Hum. Genet. 2013, 93, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Schisler, J.C.; Charles, P.C.; Parker, J.S.; Hilliard, E.G.; Mapara, S.; Meredith, D.; Lineberger, R.E.; Wu, S.S.; Alder, B.D.; Stouffer, G.A.; et al. Stable patterns of gene expression regulating carbohydrate metabolism determined by geographic ancestry. PLoS ONE 2009, 4, e8183. [Google Scholar] [CrossRef] [PubMed]
- Ravi, S.; Schuck, R.N.; Hilliard, E.; Lee, C.R.; Dai, X.; Lenhart, K.; Willis, M.S.; Jensen, B.C.; Stouffer, G.A.; Patterson, C.; et al. Clinical Evidence Supports a Protective Role for CXCL5 in Coronary Artery Disease. Am. J. Pathol. 2017, 187, 2895–2911. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Zhang, F.; Hu, H.; Bakshi, A.; Robinson, M.R.; Powell, J.E.; Montgomery, G.W.; Goddard, M.E.; Wray, N.R.; Visscher, P.M.; et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 2016, 48, 481–487. [Google Scholar] [CrossRef] [PubMed]
- GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 2013, 45, 580–585. [Google Scholar] [CrossRef]
- Zbrozyna, A.W.; Westwood, D.M. Stimulation in prefrontal cortex inhibits conditioned increase in blood pressure and avoidance bar pressing in rats. Physiol. Behav. 1991, 49, 705–708. [Google Scholar] [CrossRef]
- Flores, G.; Flores-Gómez, G.D.; de Jesús Gomez-Villalobos, M. Neuronal changes after chronic high blood pressure in animal models and its implication for vascular dementia. Synapse 2016, 70, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Clancy, K.W.; Weerapana, E.; Thompson, P.R. Detection and identification of protein citrullination in complex biological systems. Curr. Opin. Chem. Biol. 2016, 30, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Leslie, R.; O’Donnell, C.J.; Johnson, A.D. GRASP: Analysis of genotype–phenotype results from 1390 genome-wide association studies and corresponding open access database. Bioinformatics 2014, 30, i185–i194. [Google Scholar] [CrossRef] [PubMed]
- Speliotes, E.K.; Willer, C.J.; Berndt, S.I.; Monda, K.L.; Thorleifsson, G.; Jackson, A.U.; Allen, H.L.; Lindgren, C.M.; Luan, J.; Mägi, R.; et al. Association analyses of 249,796 individuals reveal eighteen new loci associated with body mass index. Nat. Genet. 2010, 42, 937–948. [Google Scholar] [CrossRef] [PubMed]
- Teslovich, T.M.; Musunuru, K.; Smith, A.V.; Edmondson, A.C.; Stylianou, I.M.; Koseki, M.; Pirruccello, J.P.; Ripatti, S.; Chasman, D.I.; Willer, C.J.; et al. Biological, Clinical, and Population Relevance of 95 Loci for Blood Lipids. Nature 2010, 466, 707–713. [Google Scholar] [CrossRef] [PubMed]
- Barber, M.J.; Mangravite, L.M.; Hyde, C.L.; Chasman, D.I.; Smith, J.D.; McCarty, C.A.; Li, X.; Wilke, R.A.; Rieder, M.J.; Williams, P.T.; et al. Genome-Wide Association of Lipid-Lowering Response to Statins in Combined Study Populations. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Winkler, T.W.; Justice, A.E.; Graff, M.; Barata, L.; Feitosa, M.F.; Chu, S.; Czajkowski, J.; Esko, T.; Fall, T.; Kilpeläinen, T.O.; et al. The Influence of Age and Sex on Genetic Associations with Adult Body Size and Shape: A Large-Scale Genome-Wide Interaction Study. PLoS Genet. 2015, 11. [Google Scholar] [CrossRef] [PubMed]
- Volders, P.J.; Verheggen, K.; Menschaert, G.; Vandepoele, K.; Martens, L.; Vandesompele, J.; Mestdagh, P. An update on LNCipedia: A database for annotated human lncRNA sequences. Nucleic Acids Res. 2015, 43, D174–D180. [Google Scholar] [CrossRef] [PubMed]
- Volders, P.J.; Helsens, K.; Wang, X.; Menten, B.; Martens, L.; Gevaert, K.; Vandesompele, J.; Mestdagh, P. LNCipedia: A database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res. 2013, 41, D246–D251. [Google Scholar] [CrossRef] [PubMed]
- Witalison, E.E.; Thompson, P.R.; Hofseth, L.J. Protein Arginine Deiminases and Associated Citrullination: Physiological Functions and Diseases Associated with Dysregulation. Curr. Drug Targets 2015, 16, 700–710. [Google Scholar] [CrossRef] [PubMed]
- Khajavi, M.; Zhou, Y.; Birsner, A.E.; Bazinet, L.; Rosa Di Sant, A.; Schiffer, A.J.; Rogers, M.S.; Krishnaji, S.T.; Hu, B.; Nguyen, V.; et al. Identification of Padi2 as a novel angiogenesis-regulating gene by genome association studies in mice. PLoS Genet. 2017, 13, e1006848. [Google Scholar] [CrossRef] [PubMed]
- Makrygiannakis, D.; Hermansson, M.; Ulfgren, A.K.; Nicholas, A.P.; Zendman, A.J.W.; Eklund, A.; Grunewald, J.; Skold, C.M.; Klareskog, L.; Catrina, A.I. Smoking increases peptidylarginine deiminase 2 enzyme expression in human lungs and increases citrullination in BAL cells. Ann. Rheum. Dis. 2008, 67, 1488–1492. [Google Scholar] [CrossRef] [PubMed]
- Damgaard, D.; Friberg Bruun Nielsen, M.; Quisgaard Gaunsbaek, M.; Palarasah, Y.; Svane-Knudsen, V.; Nielsen, C.H. Smoking is associated with increased levels of extracellular peptidylarginine deiminase 2 (PAD2) in the lungs. Clin. Exp. Rheumatol. 2015, 33, 405–408. [Google Scholar] [PubMed]
- Klareskog, L.; Stolt, P.; Lundberg, K.; Källberg, H.; Bengtsson, C.; Grunewald, J.; Rönnelid, J.; Harris, H.E.; Ulfgren, A.-K.; Rantapää-Dahlqvist, S.; et al. A new model for an etiology of rheumatoid arthritis: Smoking may trigger HLA-DR (shared epitope)-restricted immune reactions to autoantigens modified by citrullination. Arthritis Rheum. 2006, 54, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Kelly, T.N.; Hixson, J.E.; Rao, D.C.; Mei, H.; Rice, T.K.; Jaquish, C.E.; Shimmin, L.C.; Schwander, K.; Chen, C.-S.; Liu, D.; et al. Genome-wide Linkage and Positional Candidate Gene Study of Blood Pressure Response to Dietary Potassium Intervention: The GenSalt Study. Circ. Cardiovasc. Genet. 2010, 3, 539–547. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.T.; Schwartz, G.L. Gene markers and antihypertensive therapy. Curr. Hypertens. Rep. 2005, 7, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.T.; Boerwinkle, E.; O’Connell, J.R.; Bailey, K.R.; Gong, Y.; Chapman, A.B.; McDonough, C.W.; Beitelshees, A.L.; Schwartz, G.L.; Gums, J.G.; et al. Genomic Association Analysis of Common Variants Influencing Antihypertensive Response to Hydrochlorothiazide. Hypertension 2013, 62, 391–397. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.T.; Bailey, K.R.; Schwartz, G.L.; Chapman, A.B.; Chai, H.S.; Boerwinkle, E. Genomic association analysis identifies multiple loci influencing antihypertensive response to an angiotensin II receptor blocker. Hypertension 2012, 59, 1204–1211. [Google Scholar] [CrossRef] [PubMed]
- Sham, P.C.; Purcell, S.M. Statistical power and significance testing in large-scale genetic studies. Nat. Rev. Genet. 2014, 15, 335–346. [Google Scholar] [CrossRef] [PubMed]
- Shi, G.; Gu, C.C.; Kraja, A.T.; Arnett, D.K.; Myers, R.H.; Pankow, J.S.; Hunt, S.C.; Rao, D.C. Genetic Effect on Blood Pressure is Modulated by Age: The HyperGEN Study. Hypertension 2009, 53, 35–41. [Google Scholar] [CrossRef] [PubMed]

| AA | CAU | p | |
|---|---|---|---|
| Demographics | |||
| Number of genotyped participants | 193 | 123 | |
| * Age at enrollment, mean (range) | 57 (24–92) | 60 (25–91) | 0.0103 |
| Male sex, n (%) | 60 (31) | 41 (33) | 0.7113 |
| *** Education: HS or less, n (%) | 152 (79) | 74 (60) | 0.0005 |
| *** Low literacy†, n (%) | 52 (29) | 12 (10) | 0.0002 |
| Employed full or part time, n (%) | 71 (37) | 47 (38) | 0.8124 |
| *** Household income ≤ $40,000 n (%), (missing = 14%) | 148 (90) | 70 (65) | 0.0001 |
| * Currently have health insurance, n (%) | 135 (70) | 99 (80) | 0.0480 |
| Self-rated health good-excellent, n (%) | 118 (61) | 75 (61) | 1.0000 |
| Co-enrollment in lifestyle study, n (%) | 84 (44) | 43 (35) | 0.1579 |
| Cardiovascular Disease Risk Factors | |||
| Current cigarette smoker, n (%) | 44 (23) | 27 (22) | 0.8909 |
| * Diabetes (self-report or HbA1c ≥ 6.5), n (%) | 94 (49) | 42 (34) | 0.0143 |
| Total cholesterol (mg/dL), mean (SE) | 186 (3.0) | 194 (3.4) | 0.1715 |
| *** HDL-C (mg/dL), mean (SE) | 53 (1.0) | 47 (1.3) | 0.0003 |
| Systolic BP (mmHg), mean (SE) | 138 (1.5) | 138 (1.9) | 0.8523 |
| * Diastolic BP (mmHg), mean (SE) | 83 (1.0) | 80 (1.1) | 0.0310 |
| Systolic BP ≥140 mmHg, n (%) | 84 (44) | 54 (44) | 1.0000 |
| Physiologic Factors | |||
| * Weight (kg), mean (SE) | 101 (1.7) | 95 (2.4) | 0.0307 |
| * Body Mass Index, mean (SE) | 37 (0.7) | 35 (0.9) | 0.0214 |
| Number of comorbidities, mean (SE) | 3.4 (0.1) | 3.7 (0.2) | 0.2956 |
| ** Glomerular Filtration Rate (mg/dL), mean (SE) | 88 (1.7) | 81 (1.7) | 0.0045 |
| Medication and Adherence | |||
| *** Taking BP lowering medication, n (%) | 182 (94) | 100 (81) | 0.0006 |
| *** Number of BP medication classes, mean (SE) | 2.1 (0.1) | 1.6 (0.1) | 0.0010 |
| AA | CAU | |
|---|---|---|
| Trait | Estimate (SE) | Estimate (SE) |
| Age (years) | −0.43 (0.14) ** | −0.37 (0.18) * |
| Lifestyle participation (no) | 6.51 (3.42) ‡ | 1.77 (3.90) |
| BMI (per unit) | −0.17 (0.19) | 0.34 (0.18) ‡ |
| Smoking (some vs. none) | 13.68 (8.60) | −24.67 (11.87) * |
| Smoking (some vs. daily) | 15.30 (9.26) ‡ | −24.67 (12.40) * |
| Smoking (none vs. daily) | 1.61 (4.40) | 0.00 (4.62) |
| Diabetes (no) | −2.95 (3.42) | −5.77 (3.87) |
| Gender (male) | 2.31 (3.70) | 3.15 (3.92) |
| Weight loss (per percent) | −0.66 (0.33) * | −0.06 (0.29) |
| AA | CAU | |
|---|---|---|
| Trait | Estimate (SE) | Estimate (SE) |
| Age (years) | −0.59 (0.15) *** | −0.38 (0.19) * |
| Lifestyle participation (no) | 3.49 (1.73) * | 0.91 (1.84) |
| BMI (per unit) | −0.34 (0.21) | 0.19 (0.25) |
| Smoking history (ever) | −2.09 (2.14) | −3.89 (2.41) |
| Diabetes (no) | −2.52 (1.72) | −1.84 (2.11) |
| Gender (male) | 0.03 (1.90) | 2.11 (1.91) |
| Weight loss (per percent) | −0.39 (0.33) | −0.16 (0.29) |
| dbSNP ID | Gene(s) | Chr | GRCh37 | Cohort | β HET | β HOM | β Age: HET | β Age: HOM |
|---|---|---|---|---|---|---|---|---|
| rs592582 | XR_001739753 | 2 | 157773386 | AA | ↑ [↓] | ↓ [↑] | ||
| rs243601 | C21orf91-OT1 | 21 | 19159766 | AA | ↑ | ↓ | ||
| rs243603 | C21orf91-OT1 | 21 | 19160300 | AA | ↑ | ↓ | ||
| rs243605 | C21orf91-OT1, C21orf91 | 21 | 19161120 | AA | ↑ | ↓ | ||
| rs243607 | C21orf91-OT1, C21orf91 | 21 | 19161515 | AA | ↑ | ↓ | ||
| rs2220511 | C21orf91-OT1, C21orf91 | 21 | 19164911 | AA | ↑ | ↓ | ||
| rs2258119 | C21orf91 | 21 | 19167479 | AA | ↓ | ↑ | ||
| rs1799945 | HFE | 6 | 26091179 | AA † | ↓ | ↑ | ||
| rs381815 | PLEKHA7 | 11 | 16902268 | AA † | ↓ | ↑ | ↑ | ↓ |
| rs3184504 | SH2B3 | 12 | 111884608 | AA † | ↑ | ↓ | ||
| rs2521501 | FES | 15 | 91437388 | AA † | ↓ | ↑ | ||
| rs17477177 | CTB-30L5.1 | 7 | 106411858 | CAU | ↑ | ↓ | ||
| rs1378942 | CSK | 15 | 75077367 | CAU | ↓ | ↑ |
| dbSNP ID | Region | Gene(s) | Chr | GRCh37 | β HET | β HOM | β Age: HET | β Age: HOM |
|---|---|---|---|---|---|---|---|---|
| rs16942954 | intronic | AQP4-AS1, CHST9 | 18 | 24501350 | ↑ | ↓ | ||
| rs16942955 | intronic | AQP4-AS1, CHST9 | 18 | 24502493 | ↑ | ↓ | ||
| rs232358 | intronic, 3’ UTR | AQP4-AS1, CHST9 | 18 | 24492099 | ↓ | |||
| rs380625 | intronic, 3’ UTR | AQP4-AS1, CHST9 | 18 | 24493117 | ↓ | |||
| rs1181704 | intronic, 3’ UTR | AQP4-AS1, CHST9 | 18 | 24492641 | ↓ | |||
| rs11597228 | intergenic | CELF2 | 10 | 10660838 | ↓ | ↑ | ||
| rs4747873 | intergenic | CELF2 | 10 | 10686085 | ↓ | ↑ | ||
| rs7906433 | intergenic | KLF6 | 10 | 3888845 | ↓ | ↑ | ↑ | ↓ |
| rs12255472 | intergenic | KLF6 | 10 | 4466023 | ↑ | ↓ |
| dbSNP ID | Region | Gene(s) | Chr | GRCh37 | β HET | β HOM | β Age: HET | β Age: HOM |
|---|---|---|---|---|---|---|---|---|
| rs2014725 | intronic | PADI2 | 1 | 17417253 | ↓ | ↑ | ||
| rs2235910 | intronic | PADI2 | 1 | 17425829 | ↓ | ↑ | ||
| rs737428 | intronic | PADI2 | 1 | 17429185 | ↓ | ↑ | ||
| rs4949959 | intergenic | RWDD3 | 1 | 95766707 | ↓ | ↑ | ↑ | ↑ |
| rs4950044 | intergenic | RWDD3 | 1 | 95766797 | ↓ | ↑ | ↑ | ↓ |
| rs6683355 | intergenic | RWDD3 | 1 | 95773106 | ↓ | ↑ | ↑ | ↓ |
| rs7519220 | intergenic | SRP9, ENAH | 1 | 225863345 | ↓ | ↑ | ||
| rs7365361 | intergenic | SRP9, ENAH | 1 | 225864622 | ↓ | ↑ | ||
| rs943759 | intergenic | LOC102723834 | 1 | 225886318 | ↓ | ↑ | ||
| rs6576973 | 3’ UTR | ARID5A | 2 | 97218367 | ↓ | ↑ | ||
| rs7608325 | intergenic | KANSL3 | 2 | 97305080 | ↑ | |||
| rs7690085 | intronic | FSTL5 | 4 | 162709000 | ↑ | ↓ | ↓ | |
| rs13130537 | intronic | FSTL5 | 4 | 162718372 | ↓ | ↓ | ↑ | ↑ |
| rs10026821 | intronic | SORBS2 | 4 | 186540503 | ↑ | ↓ | ||
| rs10030246 | intronic | SORBS2 | 4 | 186541887 | ↑ | ↓ | ||
| rs37957 | intronic | LOC100505921, LOC105375139 | 7 | 8000971 | ↓ | |||
| rs37968 | intronic | LOC100505921, LOC105375139 | 7 | 8005973 | ↓ | |||
| rs1468594 | intronic | GLCCI1 | 7 | 8122313 | ↓ | ↑ | ↑ | ↓ |
| rs10966220 | intergenic | IZUMO3, ELAVL2 | 9 | 24113158 | ↓ | |||
| rs10812027 | intergenic | IZUMO3, ELAVL2 | 9 | 24113936 | ↓ | |||
| rs10886170 | intergenic | GHITM, NRG3 | 10 | 85091864 | ↓ | ↑ | ||
| rs10886214 | intergenic | GHITM, NRG3 | 10 | 85127739 | ↓ | ↑ | ↑ | ↓ |
| rs11244854 | intronic | ADAM12 | 10 | 127850629 | ↓ | |||
| rs1674927 | intronic | ADAM12 | 10 | 127852395 | ↓ | |||
| rs7337547 | intergenic | SPRYD7, KPNA3 | 13 | 50443527 | ↓ | |||
| rs11617754 | intronic | SPRYD7 | 13 | 50501980 | ↑ | |||
| rs9805613 | intergenic | SLITRK6 | 13 | 86982353 | ↑ | ↓ | ||
| rs9302073 | intergenic | SLITRK6 | 13 | 87002553 | ↑ | ↑ | ||
| rs8021103 | intronic | LOC105370510 | 14 | 56177363 | ↑ | ↓ | ↓ | ↑ |
| rs10498477 | intronic | LOC105370510 | 14 | 56180099 | ↑ | ↑ | ↓ | ↓ |
| rs2134919 | intergenic | EXOC5, OTX2 | 14 | 57412758 | ↑ | ↓ | ↓ | ↑ |
| rs6573129 | intergenic | EXOC5, OTX2 | 14 | 57648321 | ↓ | ↑ | ||
| rs7158266 | intergenic | EXOC5, OTX2 | 14 | 57648751 | ↓ | ↑ | ||
| rs10136042 | intergenic | EXOC5, OTX2 | 14 | 57665761 | ↑ | ↑ | ↓ | |
| rs10135064 | intergenic | EXOC5 | 14 | 57668859 | ↓ | ↑ | ||
| rs3742578 | missense, 3’ UTR | EXOC5 | 14 | 57672715 | ↓ | ↓ | ↓ | ↑ |
| rs7141911 | 3’ UTR | EXOC5 | 14 | 57672871 | ↓ | ↑ | ||
| rs198480 | intergenic | CTNNBL1, BLCAP | 20 | 36280827 | ↑ | ↓ | ||
| rs1928630 | intergenic | CTNNBL1, BLCAP | 20 | 36286035 | ↑ | ↑ | ↓ |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Halladay, J.R.; Lenhart, K.C.; Robasky, K.; Jones, W.; Homan, W.F.; Cummings, D.M.; Cené, C.W.; Hinderliter, A.L.; Miller, C.L.; Donahue, K.E.; et al. Applicability of Precision Medicine Approaches to Managing Hypertension in Rural Populations. J. Pers. Med. 2018, 8, 16. https://doi.org/10.3390/jpm8020016
Halladay JR, Lenhart KC, Robasky K, Jones W, Homan WF, Cummings DM, Cené CW, Hinderliter AL, Miller CL, Donahue KE, et al. Applicability of Precision Medicine Approaches to Managing Hypertension in Rural Populations. Journal of Personalized Medicine. 2018; 8(2):16. https://doi.org/10.3390/jpm8020016
Chicago/Turabian StyleHalladay, Jacqueline R., Kaitlin C. Lenhart, Kimberly Robasky, Wendell Jones, Wayne F. Homan, Doyle M. Cummings, Crystal W. Cené, Alan L. Hinderliter, Cassandra L. Miller, Katrina E. Donahue, and et al. 2018. "Applicability of Precision Medicine Approaches to Managing Hypertension in Rural Populations" Journal of Personalized Medicine 8, no. 2: 16. https://doi.org/10.3390/jpm8020016
APA StyleHalladay, J. R., Lenhart, K. C., Robasky, K., Jones, W., Homan, W. F., Cummings, D. M., Cené, C. W., Hinderliter, A. L., Miller, C. L., Donahue, K. E., Garcia, B. A., Keyserling, T. C., Ammerman, A. S., Patterson, C., DeWalt, D. A., Johnston, L. F., Willis, M. S., & Schisler, J. C. (2018). Applicability of Precision Medicine Approaches to Managing Hypertension in Rural Populations. Journal of Personalized Medicine, 8(2), 16. https://doi.org/10.3390/jpm8020016