Homologous Recombination Deficiency in Ovarian Cancer: from the Biological Rationale to Current Diagnostic Approaches
Abstract
1. Homologous Recombination Deficiency (HRD)
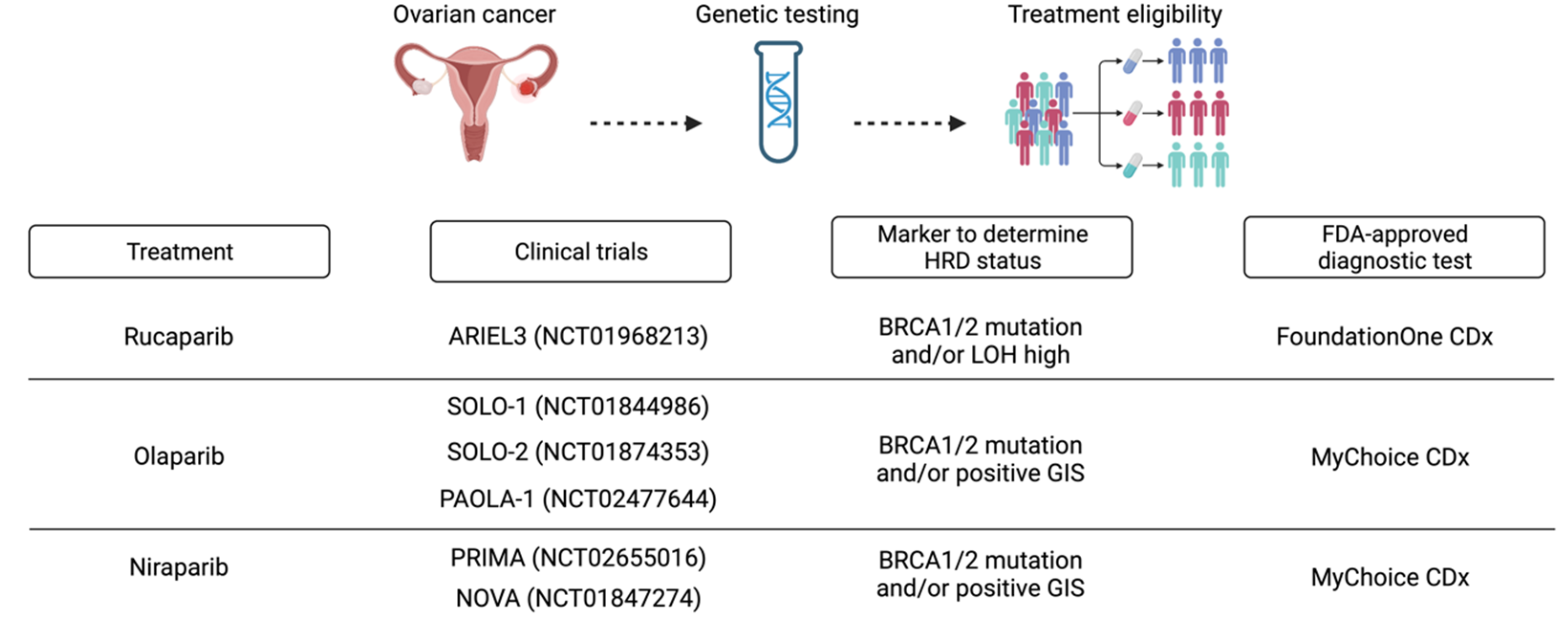
2. HRD in Ovarian Cancer
| Trial | Drug | Efficacy Data | Ref. |
|---|---|---|---|
| ARIEL2 | Rucaparib | Response rates: 69% for BRCA-mutated tumors, 39% for BRCA wild-type and high LOH tumors, and 11% for BRCA wild-type and low LOH tumors (p < 0.0001). | [42,43] |
| ARIEL3 | Rucaparib | Improved PFS compared with placebo in both patients with mutated BRCA (16.6 versus 5 months, HR = 0.23, 95% CI = 0.16–0.34) and those with HRD (13.6 versus 5.4 months, HR = 0.32, 95% CI = 0.24–0.42), and in the whole population (10.8 versus 5.4 months, HR = 0.36, 95% CI = 0.30–0.45) including LOH negative patients. | [44] |
| NOVA | Niraparib | Niraparib was associated with better median PFS in all subgroups, with a more substantial benefit in the gBRCA cohort (21.0 versus 5.5 months, HR = 0.27; 95% CI = 0.17–0.41) than in the non-gBRCA cohort with HRD (12.9 versus 3.8 months, HR = 0.38; 95% CI = 0.24–0.59) and the entire non-gBRCA cohort (9.3 versus 3.9 months, HR = 0.45; 95% CI = 0.34). | [45] |
| Study19 | Olaparib | Improvement in OS in patients with BRCA-mutated recurrent ovarian cancer who received olaparib maintenance therapy following platinum-based chemotherapy (29.8 versus 27.8 months, HR = 0.73; 95% CI = 0.55–0.95). This occurrence represents a secondary endpoint of the trial. The results support previously reported benefits of olaparib in PFS compared to placebo, which is the primary trial endpoint (8.4 versus 4.8 months, HR = 0.35; 95% CI = 0.25–0.49). | [46,47] |
| SOLO-1 | Olaparib | The SOLO-1 trial demonstrated a long-term PFS advantage for olaparib versus placebo in the first-line maintenance treatment of patients with HGSOC with a germline or somatic mutation in BRCA1/2 who had a complete or partial response after platinum-based chemotherapy. Data revealed olaparib reduced the risk of disease progression or death by 67% (based on an HR of 0.33; 95% CI = 0.25–0.43) and increased PFS to a median of 56.0 months compared with 13.8 months of placebo. | [48,49] |
| SOLO-2 | Olaparib | Results from the phase III SOLO-2 trial demonstrate an improvement in PFS in patients with platinum-sensitive relapsed ovarian cancer and gBRCA mutations treated with olaparib compared to placebo in the maintenance setting: PFS (19.1 versus 5.5 months, HR = 0.30; 95% CI = 0.22–0.41). Although there was no statistically significant difference in OS, the results supported the use of olaparib for maintenance in these patients. | [50,51,52] |
| PRIMA | Niraparib | In the HRD population, maintenance therapy with niraparib led to a reduction in the risk of progression or death by 57% (HR = 0.43; 95% CI 0.31–0.59), whereas in the intention-to-treat population, the risk reduction was 38% (HR = 0.62; 95% CI 0.5–0.76). In the HR-proficient subgroup maintenance therapy with niraparib led to 32% reduction in the risk of progression or death (HR = 0.68; 95% CI, 0.49–0.94). | [20,49] |
| VELIA | Veliparib | Across all patient subgroups, PFS for the combined induction and maintenance phases in the veliparib arm was 23.5 months versus 17.3 months in the placebo arm (HR = 0.68; 95% CI 0.56–0.83). The benefit was most evident for those with BRCA mutations. In this group, the median PFS was 34.7 months, compared with 22.0 months for veliparib and placebo, respectively (HR 0.44; 95% CI 0.28–0.68), whereas in the HRD cohort, PFS was 31.9 versus. 20.5 months (HR = 0.57; 95% CI: 0.43–0.76). Unfortunately, veliparib is not FDA-approved. | [53] |
| PAOLA-1 | Olaparib | Results showed that olaparib in combination with bevacizumab reduced the risk of disease progression or death by 41% and improved PFS in the intention-to-treat population with a median of 22.1 months compared with 16.6 months in patients treated with bevacizumab monotherapy (HR = 0.59; 95% CI 0.49–0.72). Subgroup analysis also highlighted an essential synergistic effect of the combination in patients with BRCA mutation and HRD, with a mean PFS of 37.2 months and an HR of 0.31 (95% CI 0.20–0.47) and 0.33 (95% CI 0.25–0.45), respectively. However, adding olaparib to bevacizumab showed almost no effect in the HRD-negative or unknown HRD status subgroup, where the median PFS was 16.9 months in the experimental arm versus 16 months in the control arm. These data support the hypothesis of the synergistic effect of olaparib and bevacizumab in BRCA mutant and HRD patients, underscoring the importance of HRD status as a novel prognostic factor and predictor of response to PARP inhibitors. | [49,54] |
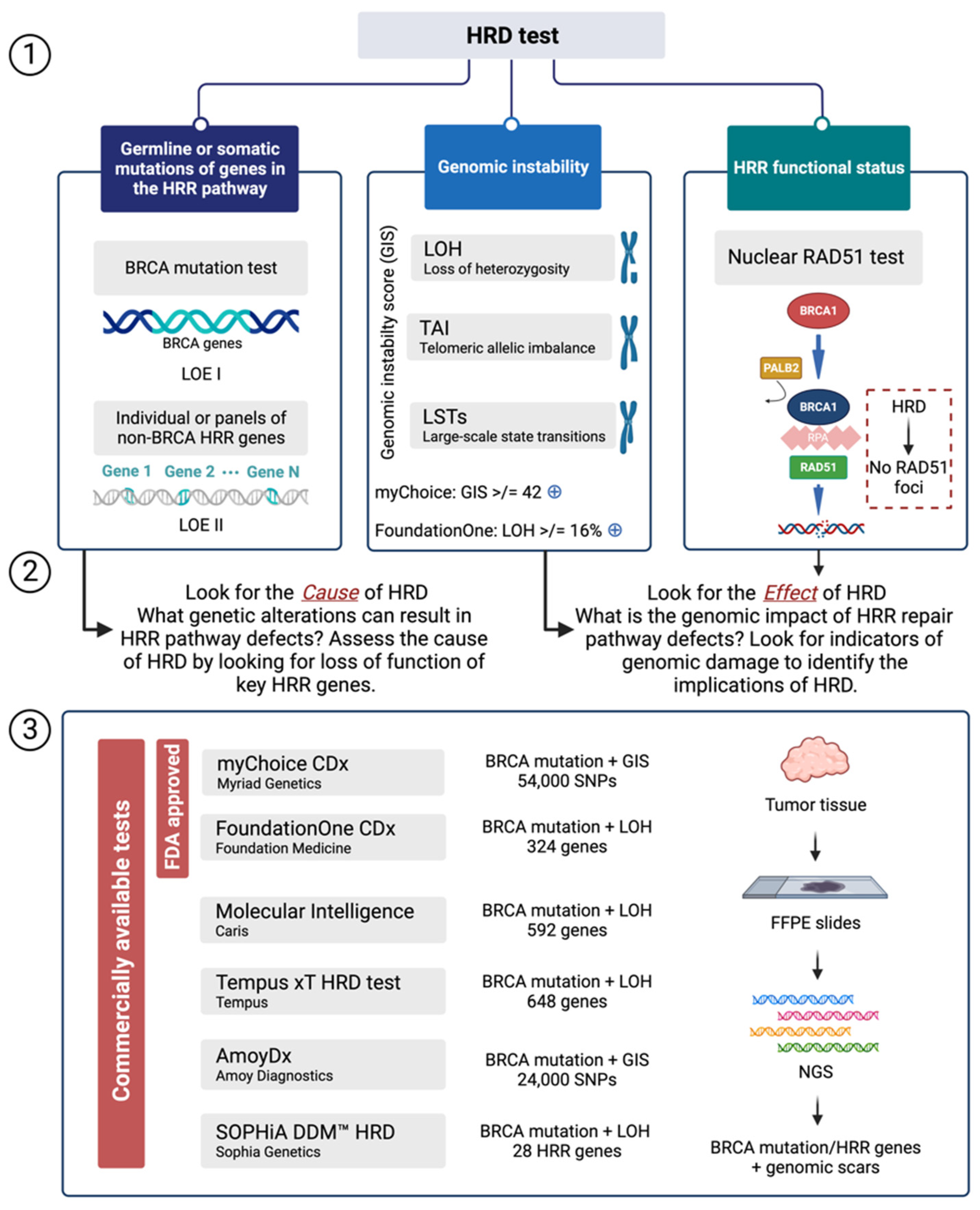
3. HRD Assays
3.1. Mutations of Genes in the HRR Pathway
3.2. Genomic Scars or Mutational Markers Showing Profiles of Genomic Instability
3.3. Functional Assays of HRD
4. Outsourcing of HRD Analysis
5. HRD in a Laboratory Routine
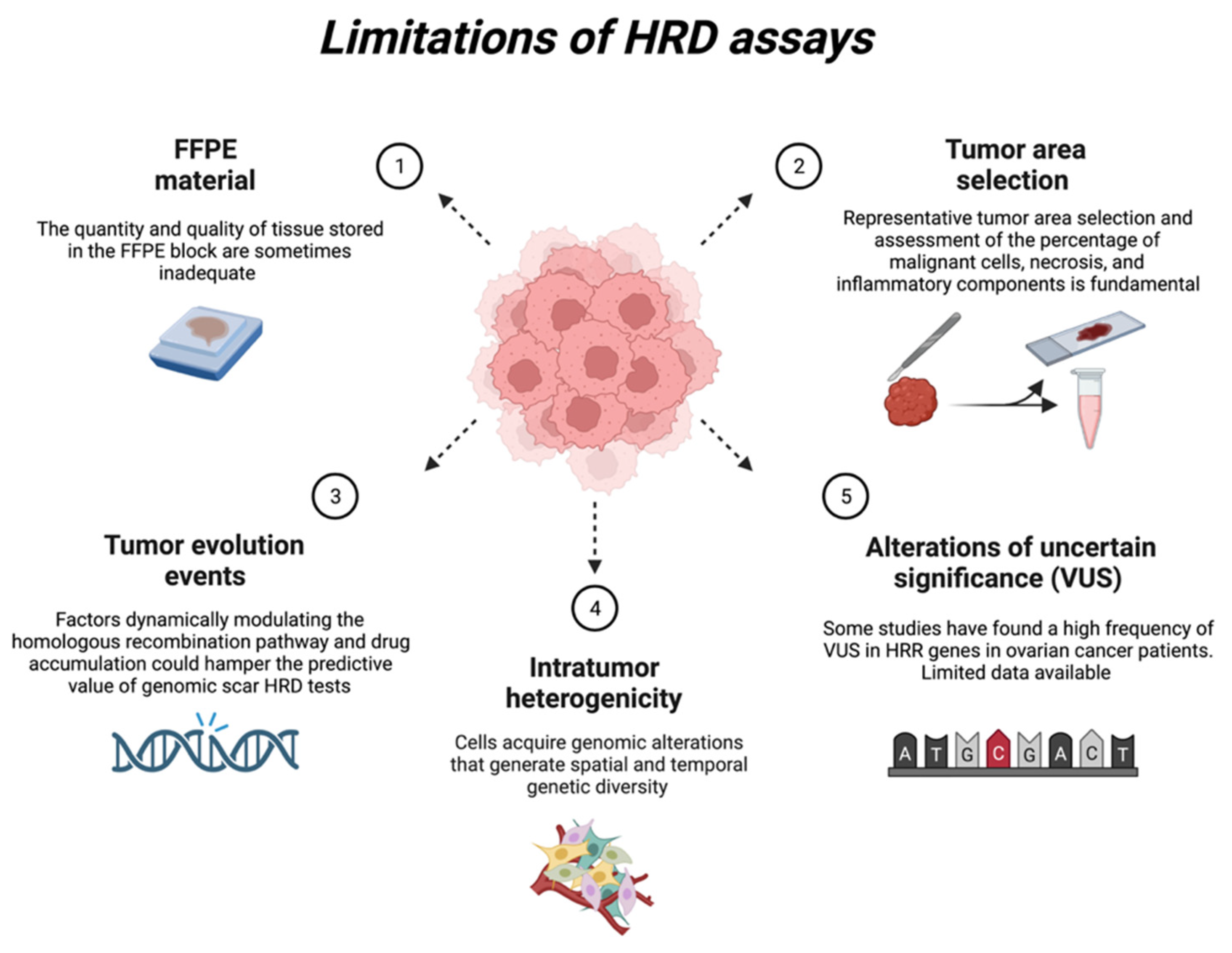
6. Current HRD Assay Limitations
6.1. FFPE Material
6.2. Representative Tumour Area Selection
6.3. Tumour Evolution Events
6.4. Intratumor Heterogenicity
6.5. Alterations of Uncertain Significance (VUS)
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yamamoto, H.; Hirasawa, A. Homologous Recombination Deficiencies and Hereditary Tumors. Int. J. Mol. Sci. 2021, 23, 348. [Google Scholar] [CrossRef] [PubMed]
- Konstantinopoulos, P.A.; Ceccaldi, R.; Shapiro, G.I.; D’Andrea, A.D. Homologous Recombination Deficiency: Exploiting the Fundamental Vulnerability of Ovarian Cancer. Cancer Discov. 2015, 5, 1137–1154. [Google Scholar] [CrossRef] [PubMed]
- Prakash, R.; Zhang, Y.; Feng, W.; Jasin, M. Homologous Recombination and Human Health: The Roles of BRCA1, BRCA2, and Associated Proteins. Cold Spring Harb. Perspect. Biol. 2015, 7, a016600. [Google Scholar] [CrossRef] [PubMed]
- Moynahan, M.E.; Jasin, M. Mitotic Homologous Recombination Maintains Genomic Stability and Suppresses Tumorigenesis. Nat. Rev. Mol. Cell Biol. 2010, 11, 196–207. [Google Scholar] [CrossRef]
- King, M.-C. “The Race” to Clone BRCA1. Science 2014, 343, 1462–1465. [Google Scholar] [CrossRef]
- da Cunha Colombo Bonadio, R.R.; Fogace, R.N.; Miranda, V.C.; Diz, M.D.P.E. Homologous Recombination Deficiency in Ovarian Cancer: A Review of Its Epidemiology and Management. Clin. Sao Paulo Braz. 2018, 73, e450s. [Google Scholar] [CrossRef]
- O’Connor, M.J. Targeting the DNA Damage Response in Cancer. Mol. Cell 2015, 60, 547–560. [Google Scholar] [CrossRef]
- Abkevich, V.; Timms, K.M.; Hennessy, B.T.; Potter, J.; Carey, M.S.; Meyer, L.A.; Smith-McCune, K.; Broaddus, R.; Lu, K.H.; Chen, J.; et al. Patterns of Genomic Loss of Heterozygosity Predict Homologous Recombination Repair Defects in Epithelial Ovarian Cancer. Br. J. Cancer 2012, 107, 1776–1782. [Google Scholar] [CrossRef]
- Birkbak, N.J.; Wang, Z.C.; Kim, J.-Y.; Eklund, A.C.; Li, Q.; Tian, R.; Bowman-Colin, C.; Li, Y.; Greene-Colozzi, A.; Iglehart, J.D.; et al. Telomeric Allelic Imbalance Indicates Defective DNA Repair and Sensitivity to DNA-Damaging Agents. Cancer Discov. 2012, 2, 366–375. [Google Scholar] [CrossRef]
- Popova, T.; Manié, E.; Rieunier, G.; Caux-Moncoutier, V.; Tirapo, C.; Dubois, T.; Delattre, O.; Sigal-Zafrani, B.; Bollet, M.; Longy, M.; et al. Ploidy and Large-Scale Genomic Instability Consistently Identify Basal-like Breast Carcinomas with BRCA1/2 Inactivation. Cancer Res. 2012, 72, 5454–5462. [Google Scholar] [CrossRef]
- Stewart, M.D.; Merino Vega, D.; Arend, R.C.; Baden, J.F.; Barbash, O.; Beaubier, N.; Collins, G.; French, T.; Ghahramani, N.; Hinson, P.; et al. Homologous Recombination Deficiency: Concepts, Definitions, and Assays. Oncologist 2022, 27, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Timms, K.M.; Abkevich, V.; Hughes, E.; Neff, C.; Reid, J.; Morris, B.; Kalva, S.; Potter, J.; Tran, T.V.; Chen, J.; et al. Association of BRCA1/2 Defects with Genomic Scores Predictive of DNA Damage Repair Deficiency among Breast Cancer Subtypes. Breast Cancer Res. BCR 2014, 16, 475. [Google Scholar] [CrossRef] [PubMed]
- Marquard, A.M.; Eklund, A.C.; Joshi, T.; Krzystanek, M.; Favero, F.; Wang, Z.C.; Richardson, A.L.; Silver, D.P.; Szallasi, Z.; Birkbak, N.J. Pan-Cancer Analysis of Genomic Scar Signatures Associated with Homologous Recombination Deficiency Suggests Novel Indications for Existing Cancer Drugs. Biomark. Res. 2015, 3, 9. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, T.; Zhang, Z.; Payne, S.H.; Zhang, B.; McDermott, J.E.; Zhou, J.-Y.; Petyuk, V.A.; Chen, L.; Ray, D.; et al. Integrated Proteogenomic Characterization of Human High-Grade Serous Ovarian Cancer. Cell 2016, 166, 755–765. [Google Scholar] [CrossRef]
- Ngoi, N.Y.L.; Tan, D.S.P. The Role of Homologous Recombination Deficiency Testing in Ovarian Cancer and Its Clinical Implications: Do We Need It? ESMO Open 2021, 6, 100144. [Google Scholar] [CrossRef]
- Weaver, A.N.; Yang, E.S. Beyond DNA Repair: Additional Functions of PARP-1 in Cancer. Front. Oncol. 2013, 3, 290. [Google Scholar] [CrossRef] [PubMed]
- Helleday, T.; Petermann, E.; Lundin, C.; Hodgson, B.; Sharma, R.A. DNA Repair Pathways as Targets for Cancer Therapy. Nat. Rev. Cancer 2008, 8, 193–204. [Google Scholar] [CrossRef]
- Denkert, C.; Romey, M.; Swedlund, B.; Hattesohl, A.; Teply-Szymanski, J.; Kommoss, S.; Kaiser, K.; Staebler, A.; du Bois, A.; Grass, A.; et al. Homologous Recombination Deficiency as an Ovarian Cancer Biomarker in a Real-World Cohort: Validation of Decentralized Genomic Profiling. J. Mol. Diagn. JMD 2022, 24, 1254–1263. [Google Scholar] [CrossRef] [PubMed]
- Robson, M.E.; Tung, N.; Conte, P.; Im, S.-A.; Senkus, E.; Xu, B.; Masuda, N.; Delaloge, S.; Li, W.; Armstrong, A.; et al. OlympiAD Final Overall Survival and Tolerability Results: Olaparib versus Chemotherapy Treatment of Physician’s Choice in Patients with a Germline BRCA Mutation and HER2-Negative Metastatic Breast Cancer. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2019, 30, 558–566. [Google Scholar] [CrossRef]
- González-Martín, A.; Pothuri, B.; Vergote, I.; DePont Christensen, R.; Graybill, W.; Mirza, M.R.; McCormick, C.; Lorusso, D.; Hoskins, P.; Freyer, G.; et al. Niraparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2391–2402. [Google Scholar] [CrossRef]
- de Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Incorvaia, L.; Capoluongo, E.; Tagliaferri, P.; Gori, S.; Cortesi, L.; Genuardi, M.; Turchetti, D.; De Giorgi, U.; Di Maio, M.; et al. Implementation of Preventive and Predictive BRCA Testing in Patients with Breast, Ovarian, Pancreatic, and Prostate Cancer: A Position Paper of Italian Scientific Societies. ESMO Open 2022, 7, 100459. [Google Scholar] [CrossRef]
- Stegel, V.; Blatnik, A.; Škof, E.; Dragoš, V.Š.; Krajc, M.; Gregorič, B.; Škerl, P.; Strojnik, K.; Klančar, G.; Banjac, M.; et al. Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing. Cancers 2022, 14, 1434. [Google Scholar] [CrossRef] [PubMed]
- Weren, R.D.A.; Mensenkamp, A.R.; Simons, M.; Eijkelenboom, A.; Sie, A.S.; Ouchene, H.; van Asseldonk, M.; Gomez-Garcia, E.B.; Blok, M.J.; de Hullu, J.A.; et al. Novel BRCA1 and BRCA2 Tumor Test as Basis for Treatment Decisions and Referral for Genetic Counselling of Patients with Ovarian Carcinomas. Hum. Mutat. 2017, 38, 226–235. [Google Scholar] [CrossRef] [PubMed]
- Venkitaraman, A.R. A Growing Network of Cancer-Susceptibility Genes. N. Engl. J. Med. 2003, 348, 1917–1919. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network Integrated Genomic Analyses of Ovarian Carcinoma. Nature 2011, 474, 609–615. [CrossRef]
- Buisson, R.; Dion-Côté, A.-M.; Coulombe, Y.; Launay, H.; Cai, H.; Stasiak, A.Z.; Stasiak, A.; Xia, B.; Masson, J.-Y. Cooperation of Breast Cancer Proteins PALB2 and Piccolo BRCA2 in Stimulating Homologous Recombination. Nat. Struct. Mol. Biol. 2010, 17, 1247–1254. [Google Scholar] [CrossRef]
- Taniguchi, T.; Tischkowitz, M.; Ameziane, N.; Hodgson, S.V.; Mathew, C.G.; Joenje, H.; Mok, S.C.; D’Andrea, A.D. Disruption of the Fanconi Anemia-BRCA Pathway in Cisplatin-Sensitive Ovarian Tumors. Nat. Med. 2003, 9, 568–574. [Google Scholar] [CrossRef]
- Wilkerson, P.M.; Dedes, K.J.; Wetterskog, D.; Mackay, A.; Lambros, M.B.; Mansour, M.; Frankum, J.; Lord, C.J.; Natrajan, R.; Ashworth, A.; et al. Functional Characterization of EMSY Gene Amplification in Human Cancers. J. Pathol. 2011, 225, 29–42. [Google Scholar] [CrossRef]
- Baer, R.; Ludwig, T. The BRCA1/BARD1 Heterodimer, a Tumor Suppressor Complex with Ubiquitin E3 Ligase Activity. Curr. Opin. Genet. Dev. 2002, 12, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Alsop, K.; Fereday, S.; Meldrum, C.; deFazio, A.; Emmanuel, C.; George, J.; Dobrovic, A.; Birrer, M.J.; Webb, P.M.; Stewart, C.; et al. BRCA Mutation Frequency and Patterns of Treatment Response in BRCA Mutation-Positive Women with Ovarian Cancer: A Report from the Australian Ovarian Cancer Study Group. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2012, 30, 2654–2663. [Google Scholar] [CrossRef]
- Turner, N.; Tutt, A.; Ashworth, A. Hallmarks of “BRCAness” in Sporadic Cancers. Nat. Rev. Cancer 2004, 4, 814–819. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.S.P.; Rothermundt, C.; Thomas, K.; Bancroft, E.; Eeles, R.; Shanley, S.; Ardern-Jones, A.; Norman, A.; Kaye, S.B.; Gore, M.E. “BRCAness” Syndrome in Ovarian Cancer: A Case-Control Study Describing the Clinical Features and Outcome of Patients with Epithelial Ovarian Cancer Associated with BRCA1 and BRCA2 Mutations. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2008, 26, 5530–5536. [Google Scholar] [CrossRef] [PubMed]
- Pennington, K.P.; Walsh, T.; Harrell, M.I.; Lee, M.K.; Pennil, C.C.; Rendi, M.H.; Thornton, A.; Norquist, B.M.; Casadei, S.; Nord, A.S.; et al. Germline and Somatic Mutations in Homologous Recombination Genes Predict Platinum Response and Survival in Ovarian, Fallopian Tube, and Peritoneal Carcinomas. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2014, 20, 764–775. [Google Scholar] [CrossRef]
- Konstantinopoulos, P.A.; Spentzos, D.; Karlan, B.Y.; Taniguchi, T.; Fountzilas, E.; Francoeur, N.; Levine, D.A.; Cannistra, S.A. Gene Expression Profile of BRCAness That Correlates with Responsiveness to Chemotherapy and with Outcome in Patients with Epithelial Ovarian Cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2010, 28, 3555–3561. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.E.; Leary, A.; Scott, C.L.; Serra, V.; Lord, C.J.; Bowtell, D.; Chang, D.K.; Garsed, D.W.; Jonkers, J.; Ledermann, J.A.; et al. ESMO Recommendations on Predictive Biomarker Testing for Homologous Recombination Deficiency and PARP Inhibitor Benefit in Ovarian Cancer. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2020, 31, 1606–1622. [Google Scholar] [CrossRef]
- Armstrong, D.K.; Alvarez, R.D.; Bakkum-Gamez, J.N.; Barroilhet, L.; Behbakht, K.; Berchuck, A.; Chen, L.-M.; Cristea, M.; DeRosa, M.; Eisenhauer, E.L.; et al. Ovarian Cancer, Version 2.2020, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. JNCCN 2021, 19, 191–226. [Google Scholar] [CrossRef]
- AstraZeneca Shares Progress on LYNPARZATM (Olaparib) Tablets in the US. Available online: https://www.astrazeneca-us.com/media/press-releases/2017/astrazeneca-shares-progress-on-lynparza-olaparib-tablets-in-the-us-03282017.html (accessed on 11 January 2023).
- FDA Label for Niraparib. Research, C. for D.E. and Niraparib (ZEJULA). FDA 2019. Available online: https://www.fda.gov/drugs/resources-information-approved-drugs/niraparib-zejula (accessed on 11 January 2023).
- FDA Label for Rucaparib. Research, C. for D.E. and Rucaparib. FDA 2019. Available online: https://www.fda.gov/drugs/resources-information-approved-drugs/rucaparib (accessed on 11 January 2023).
- Banerjee, S.; Gonzalez-Martin, A.; Harter, P.; Lorusso, D.; Moore, K.N.; Oaknin, A.; Ray-Coquard, I. First-Line PARP Inhibitors in Ovarian Cancer: Summary of an ESMO Open-Cancer Horizons Round-Table Discussion. ESMO Open 2020, 5, e001110. [Google Scholar] [CrossRef]
- Pfisterer, J.; Weber, B.; Reuss, A.; Kimmig, R.; du Bois, A.; Wagner, U.; Bourgeois, H.; Meier, W.; Costa, S.; Blohmer, J.-U.; et al. Randomized Phase III Trial of Topotecan Following Carboplatin and Paclitaxel in First-Line Treatment of Advanced Ovarian Cancer: A Gynecologic Cancer Intergroup Trial of the AGO-OVAR and GINECO. J. Natl. Cancer Inst. 2006, 98, 1036–1045. [Google Scholar] [CrossRef]
- Jenner, Z.B.; Sood, A.K.; Coleman, R.L. Evaluation of Rucaparib and Companion Diagnostics in the PARP Inhibitor Landscape for Recurrent Ovarian Cancer Therapy. Future Oncol. Lond. Engl. 2016, 12, 1439–1456. [Google Scholar] [CrossRef]
- Coleman, R.L.; Oza, A.M.; Lorusso, D.; Aghajanian, C.; Oaknin, A.; Dean, A.; Colombo, N.; Weberpals, J.I.; Clamp, A.; Scambia, G.; et al. Rucaparib Maintenance Treatment for Recurrent Ovarian Carcinoma after Response to Platinum Therapy (ARIEL3): A Randomised, Double-Blind, Placebo-Controlled, Phase 3 Trial. Lancet Lond. Engl. 2017, 390, 1949–1961. [Google Scholar] [CrossRef] [PubMed]
- Mirza, M.R.; Monk, B.J.; Herrstedt, J.; Oza, A.M.; Mahner, S.; Redondo, A.; Fabbro, M.; Ledermann, J.A.; Lorusso, D.; Vergote, I.; et al. Niraparib Maintenance Therapy in Platinum-Sensitive, Recurrent Ovarian Cancer. N. Engl. J. Med. 2016, 375, 2154–2164. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, M.; Matulonis, U.; Gourley, C.; du Bois, A.; Vergote, I.; Rustin, G.; Scott, C.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Long-Term Efficacy, Tolerability and Overall Survival in Patients with Platinum-Sensitive, Recurrent High-Grade Serous Ovarian Cancer Treated with Maintenance Olaparib Capsules Following Response to Chemotherapy. Br. J. Cancer 2018, 119, 1075–1085. [Google Scholar] [CrossRef] [PubMed]
- Ledermann, J.; Harter, P.; Gourley, C.; Friedlander, M.; Vergote, I.; Rustin, G.; Scott, C.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Olaparib Maintenance Therapy in Platinum-Sensitive Relapsed Ovarian Cancer. N. Engl. J. Med. 2012, 366, 1382–1392. [Google Scholar] [CrossRef]
- Moore, K.; Colombo, N.; Scambia, G.; Kim, B.-G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; Sonke, G.S.; et al. Maintenance Olaparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2018, 379, 2495–2505. [Google Scholar] [CrossRef]
- Franzese, E.; Diana, A.; Centonze, S.; Pignata, S.; De Vita, F.; Ciardiello, F.; Orditura, M. PARP Inhibitors in First-Line Therapy of Ovarian Cancer: Are There Any Doubts? Front. Oncol. 2020, 10, 782. [Google Scholar] [CrossRef]
- Poveda, A.; Floquet, A.; Ledermann, J.A.; Asher, R.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Pignata, S.; Friedlander, M.; et al. Final Overall Survival (OS) Results from SOLO2/ENGOT-Ov21: A Phase III Trial Assessing Maintenance Olaparib in Patients (Pts) with Platinum-Sensitive, Relapsed Ovarian Cancer and a BRCA Mutation. J. Clin. Oncol. 2020, 38, 6002. [Google Scholar] [CrossRef]
- Poveda, A.; Floquet, A.; Ledermann, J.A.; Asher, R.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Pignata, S.; Friedlander, M.; et al. Olaparib Tablets as Maintenance Therapy in Patients with Platinum-Sensitive Relapsed Ovarian Cancer and a BRCA1/2 Mutation (SOLO2/ENGOT-Ov21): A Final Analysis of a Double-Blind, Randomised, Placebo-Controlled, Phase 3 Trial. Lancet Oncol. 2021, 22, 620–631. [Google Scholar] [CrossRef]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib Tablets as Maintenance Therapy in Patients with Platinum-Sensitive, Relapsed Ovarian Cancer and a BRCA1/2 Mutation (SOLO2/ENGOT-Ov21): A Double-Blind, Randomised, Placebo-Controlled, Phase 3 Trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef]
- Coleman, R.L.; Fleming, G.F.; Brady, M.F.; Swisher, E.M.; Steffensen, K.D.; Friedlander, M.; Okamoto, A.; Moore, K.N.; Efrat Ben-Baruch, N.; Werner, T.L.; et al. Veliparib with First-Line Chemotherapy and as Maintenance Therapy in Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2403–2415. [Google Scholar] [CrossRef]
- Ray-Coquard, I.; Pautier, P.; Pignata, S.; Pérol, D.; González-Martín, A.; Berger, R.; Fujiwara, K.; Vergote, I.; Colombo, N.; Mäenpää, J.; et al. Olaparib plus Bevacizumab as First-Line Maintenance in Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2416–2428. [Google Scholar] [CrossRef] [PubMed]
- McAlpine, J.N.; Porter, H.; Köbel, M.; Nelson, B.H.; Prentice, L.M.; Kalloger, S.E.; Senz, J.; Milne, K.; Ding, J.; Shah, S.P.; et al. BRCA1 and BRCA2 Mutations Correlate with TP53 Abnormalities and Presence of Immune Cell Infiltrates in Ovarian High-Grade Serous Carcinoma. Mod. Pathol. 2012, 25, 740–750. [Google Scholar] [CrossRef]
- Strickland, K.C.; Howitt, B.E.; Shukla, S.A.; Rodig, S.; Ritterhouse, L.L.; Liu, J.F.; Garber, J.E.; Chowdhury, D.; Wu, C.J.; D’Andrea, A.D.; et al. Association and Prognostic Significance of BRCA1/2-Mutation Status with Neoantigen Load, Number of Tumor-Infiltrating Lymphocytes and Expression of PD-1/PD-L1 in High Grade Serous Ovarian Cancer. Oncotarget 2016, 7, 13587–13598. [Google Scholar] [CrossRef] [PubMed]
- Gadducci, A.; Guerrieri, M.E. PARP Inhibitors Alone and in Combination with Other Biological Agents in Homologous Recombination Deficient Epithelial Ovarian Cancer: From the Basic Research to the Clinic. Crit. Rev. Oncol. Hematol. 2017, 114, 153–165. [Google Scholar] [CrossRef]
- Franzese, O.; Graziani, G. Role of PARP Inhibitors in Cancer Immunotherapy: Potential Friends to Immune Activating Molecules and Foes to Immune Checkpoints. Cancers 2022, 14, 5633. [Google Scholar] [CrossRef] [PubMed]
- Stover, E.H.; Fuh, K.; Konstantinopoulos, P.A.; Matulonis, U.A.; Liu, J.F. Clinical Assays for Assessment of Homologous Recombination DNA Repair Deficiency. Gynecol. Oncol. 2020, 159, 887–898. [Google Scholar] [CrossRef]
- Paulet, L.; Trecourt, A.; Leary, A.; Peron, J.; Descotes, F.; Devouassoux-Shisheboran, M.; Leroy, K.; You, B.; Lopez, J. Cracking the Homologous Recombination Deficiency Code: How to Identify Responders to PARP Inhibitors. Eur. J. Cancer 2022, 166, 87–99. [Google Scholar] [CrossRef]
- Chiang, Y.-C.; Lin, P.-H.; Cheng, W.-F. Homologous Recombination Deficiency Assays in Epithelial Ovarian Cancer: Current Status and Future Direction. Front. Oncol. 2021, 11, 675972. [Google Scholar] [CrossRef]
- Pennington, K.P.; Swisher, E.M. Hereditary Ovarian Cancer: Beyond the Usual Suspects. Gynecol. Oncol. 2012, 124, 347–353. [Google Scholar] [CrossRef]
- Roy, R.; Chun, J.; Powell, S.N. BRCA1 and BRCA2: Different Roles in a Common Pathway of Genome Protection. Nat. Rev. Cancer 2011, 12, 68–78. [Google Scholar] [CrossRef]
- Loveday, C.; Turnbull, C.; Ramsay, E.; Hughes, D.; Ruark, E.; Frankum, J.R.; Bowden, G.; Kalmyrzaev, B.; Warren-Perry, M.; Snape, K.; et al. Germline Mutations in RAD51D Confer Susceptibility to Ovarian Cancer. Nat. Genet. 2011, 43, 879–882. [Google Scholar] [CrossRef] [PubMed]
- Rafnar, T.; Gudbjartsson, D.F.; Sulem, P.; Jonasdottir, A.; Sigurdsson, A.; Jonasdottir, A.; Besenbacher, S.; Lundin, P.; Stacey, S.N.; Gudmundsson, J.; et al. Mutations in BRIP1 Confer High Risk of Ovarian Cancer. Nat. Genet. 2011, 43, 1104–1107. [Google Scholar] [CrossRef]
- Bajrami, I.; Frankum, J.R.; Konde, A.; Miller, R.E.; Rehman, F.L.; Brough, R.; Campbell, J.; Sims, D.; Rafiq, R.; Hooper, S.; et al. Genome-Wide Profiling of Genetic Synthetic Lethality Identifies CDK12 as a Novel Determinant of PARP1/2 Inhibitor Sensitivity. Cancer Res. 2014, 74, 287–297. [Google Scholar] [CrossRef]
- Stronach, E.A.; Paul, J.; Timms, K.M.; Hughes, E.; Brown, K.; Neff, C.; Perry, M.; Gutin, A.; El-Bahrawy, M.; Steel, J.H.; et al. Biomarker Assessment of HR Deficiency, Tumor BRCA1/2 Mutations, and CCNE1 Copy Number in Ovarian Cancer: Associations with Clinical Outcome Following Platinum Monotherapy. Mol. Cancer Res. MCR 2018, 16, 1103–1111. [Google Scholar] [CrossRef] [PubMed]
- Hill, S.J.; Decker, B.; Roberts, E.A.; Horowitz, N.S.; Muto, M.G.; Worley, M.J.; Feltmate, C.M.; Nucci, M.R.; Swisher, E.M.; Nguyen, H.; et al. Prediction of DNA Repair Inhibitor Response in Short-Term Patient-Derived Ovarian Cancer Organoids. Cancer Discov. 2018, 8, 1404–1421. [Google Scholar] [CrossRef] [PubMed]
- Castroviejo-Bermejo, M.; Cruz, C.; Llop-Guevara, A.; Gutiérrez-Enríquez, S.; Ducy, M.; Ibrahim, Y.H.; Gris-Oliver, A.; Pellegrino, B.; Bruna, A.; Guzmán, M.; et al. A RAD51 Assay Feasible in Routine Tumor Samples Calls PARP Inhibitor Response beyond BRCA Mutation. EMBO Mol. Med. 2018, 10, e9172. [Google Scholar] [CrossRef]
- Blanc-Durand, F.; Yaniz, E.; Genestie, C.; Rouleau, E.; Berton, D.; Lortholary, A.; Dohollou, N.; Desauw, C.; Fabbro, M.; Malaurie, E.; et al. Evaluation of a RAD51 Functional Assay in Advanced Ovarian Cancer, a GINECO/GINEGEPS Study. J. Clin. Oncol. 2021, 39, 5513. [Google Scholar] [CrossRef]
- Telli, M.L.; Timms, K.M.; Reid, J.; Hennessy, B.; Mills, G.B.; Jensen, K.C.; Szallasi, Z.; Barry, W.T.; Winer, E.P.; Tung, N.M.; et al. Homologous Recombination Deficiency (HRD) Score Predicts Response to Platinum-Containing Neoadjuvant Chemotherapy in Patients with Triple-Negative Breast Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2016, 22, 3764–3773. [Google Scholar] [CrossRef]
- Capoluongo, E.D.; Pellegrino, B.; Arenare, L.; Califano, D.; Scambia, G.; Beltrame, L.; Serra, V.; Scaglione, G.L.; Spina, A.; Cecere, S.C.; et al. Alternative Academic Approaches for Testing Homologous Recombination Deficiency in Ovarian Cancer in the MITO16A/MaNGO-OV2 Trial. ESMO Open 2022, 7, 100585. [Google Scholar] [CrossRef]
- Loverix, L.; Vergote, I.; Busschaert, P.; Vanderstichele, A.; Boeckx, B.; Venken, T.; Harter, P.; Brems, H.; Nieuwenhuysen, E.V.; Pignata, S.; et al. Predictive Value of the Leuven HRD Test Compared with Myriad MyChoice PLUS on 468 Ovarian Cancer Samples from the PAOLA-1/ENGOT-Ov25 Trial (LBA 6). Gynecol. Oncol. 2022, 166, S51–S52. [Google Scholar] [CrossRef]
- Wagener-Ryczek, S.; Merkelbach-Bruse, S.; Siemanowski, J. Biomarkers for Homologous Recombination Deficiency in Cancer. J. Pers. Med. 2021, 11, 612. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, C.; Betella, I.; Ranghiero, A.; Guerini-Rocco, E.; Bonaldo, G.; Rappa, A.; Vacirca, D.; Colombo, N.; Barberis, M. In-House Testing for Homologous Recombination Repair Deficiency (HRD) Testing in Ovarian Carcinoma: A Feasibility Study Comparing AmoyDx HRD Focus Panel with Myriad MyChoiceCDx Assay. Pathologica 2022, 114, 288–294. [Google Scholar] [CrossRef] [PubMed]
- Magliacane, G.; Brunetto, E.; Calzavara, S.; Bergamini, A.; Pipitone, G.B.; Marra, G.; Redegalli, M.; Grassini, G.; Rabaiotti, E.; Taccagni, G.; et al. Locally Performed HRD Testing for Ovarian Cancer? Yes, We Can! Cancers 2023, 15, 43. [Google Scholar] [CrossRef]
- Pellegrino, B.; Mateo, J.; Serra, V.; Balmaña, J. Controversies in Oncology: Are Genomic Tests Quantifying Homologous Recombination Repair Deficiency (HRD) Useful for Treatment Decision Making? ESMO Open 2019, 4, e000480. [Google Scholar] [CrossRef]
- Edwards, S.L.; Brough, R.; Lord, C.J.; Natrajan, R.; Vatcheva, R.; Levine, D.A.; Boyd, J.; Reis-Filho, J.S.; Ashworth, A. Resistance to Therapy Caused by Intragenic Deletion in BRCA2. Nature 2008, 451, 1111–1115. [Google Scholar] [CrossRef]
- Norquist, B.; Wurz, K.A.; Pennil, C.C.; Garcia, R.; Gross, J.; Sakai, W.; Karlan, B.Y.; Taniguchi, T.; Swisher, E.M. Secondary Somatic Mutations Restoring BRCA1/2 Predict Chemotherapy Resistance in Hereditary Ovarian Carcinomas. J. Clin. Oncol. 2011, 29, 3008–3015. [Google Scholar] [CrossRef]
- Swisher, E.M.; Sakai, W.; Karlan, B.Y.; Wurz, K.; Urban, N.; Taniguchi, T. Secondary BRCA1 Mutations in BRCA1-Mutated Ovarian Carcinomas with Platinum Resistance. Cancer Res. 2008, 68, 2581–2586. [Google Scholar] [CrossRef]
- Burger, H.; Loos, W.J.; Eechoute, K.; Verweij, J.; Mathijssen, R.H.J.; Wiemer, E.A.C. Drug Transporters of Platinum-Based Anticancer Agents and Their Clinical Significance. Drug Resist. Updat. Rev. Comment. Antimicrob. Anticancer Chemother. 2011, 14, 22–34. [Google Scholar] [CrossRef]
- von Wahlde, M.-K.; Timms, K.M.; Chagpar, A.; Wali, V.B.; Jiang, T.; Bossuyt, V.; Saglam, O.; Reid, J.; Gutin, A.; Neff, C.; et al. Intratumor Heterogeneity of Homologous Recombination Deficiency in Primary Breast Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2017, 23, 1193–1199. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, Z.; Chen, L.; Pang, J.; Wu, H.; Liang, Z. Next-Generation Sequencing Reveals a Very Low Prevalence of Deleterious Mutations of Homologous Recombination Repair Genes and Homologous Recombination Deficiency in Ovarian Clear Cell Carcinoma. Front. Oncol. 2022, 11, 5641. [Google Scholar] [CrossRef] [PubMed]

| ARID1A | EMSY | MSH2 |
| ATM | FANCA | NBN |
| ATR | FANCC | PALB2 |
| BRCA1/2 | FANCE | PTEN |
| BARD1 | FANCF | RAD50 |
| BAP1 | FANCD2 | RAD51 |
| BRIP1 | FANCG | RAD51B |
| BLM | FANCI | RAD51C |
| CDK12 | FANCL | RAD51D |
| CHEK1 | H2AX | RAD54L |
| CHEK2 | MRE11 | TP53 |
| Genomic scar | Description |
|---|---|
| Loss of heterozygosity (LOH) | One of the two alleles for a gene is lost, resulting in a homozygous cell. Failure of the remaining allele could result in the growth of malignant cells. |
| Telomeric allelic imbalance (TAI) | The proportion of alleles at the end of the chromosome (telomere) in a pair does not correspond, indicating that one chromosome has more alleles than the other. |
| Large-scale transitions (LSTs) | Breakpoints between regions of the chromosome cause discrepancies within the chromosome pair. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mangogna, A.; Munari, G.; Pepe, F.; Maffii, E.; Giampaolino, P.; Ricci, G.; Fassan, M.; Malapelle, U.; Biffi, S. Homologous Recombination Deficiency in Ovarian Cancer: from the Biological Rationale to Current Diagnostic Approaches. J. Pers. Med. 2023, 13, 284. https://doi.org/10.3390/jpm13020284
Mangogna A, Munari G, Pepe F, Maffii E, Giampaolino P, Ricci G, Fassan M, Malapelle U, Biffi S. Homologous Recombination Deficiency in Ovarian Cancer: from the Biological Rationale to Current Diagnostic Approaches. Journal of Personalized Medicine. 2023; 13(2):284. https://doi.org/10.3390/jpm13020284
Chicago/Turabian StyleMangogna, Alessandro, Giada Munari, Francesco Pepe, Edoardo Maffii, Pierluigi Giampaolino, Giuseppe Ricci, Matteo Fassan, Umberto Malapelle, and Stefania Biffi. 2023. "Homologous Recombination Deficiency in Ovarian Cancer: from the Biological Rationale to Current Diagnostic Approaches" Journal of Personalized Medicine 13, no. 2: 284. https://doi.org/10.3390/jpm13020284
APA StyleMangogna, A., Munari, G., Pepe, F., Maffii, E., Giampaolino, P., Ricci, G., Fassan, M., Malapelle, U., & Biffi, S. (2023). Homologous Recombination Deficiency in Ovarian Cancer: from the Biological Rationale to Current Diagnostic Approaches. Journal of Personalized Medicine, 13(2), 284. https://doi.org/10.3390/jpm13020284









