Recapping the Features of SARS-CoV-2 and Its Main Variants: Status and Future Paths
Abstract
:1. Introduction
2. A General Perspective for SARS-CoV-2
2.1. Molecular Biology and Taxonomy
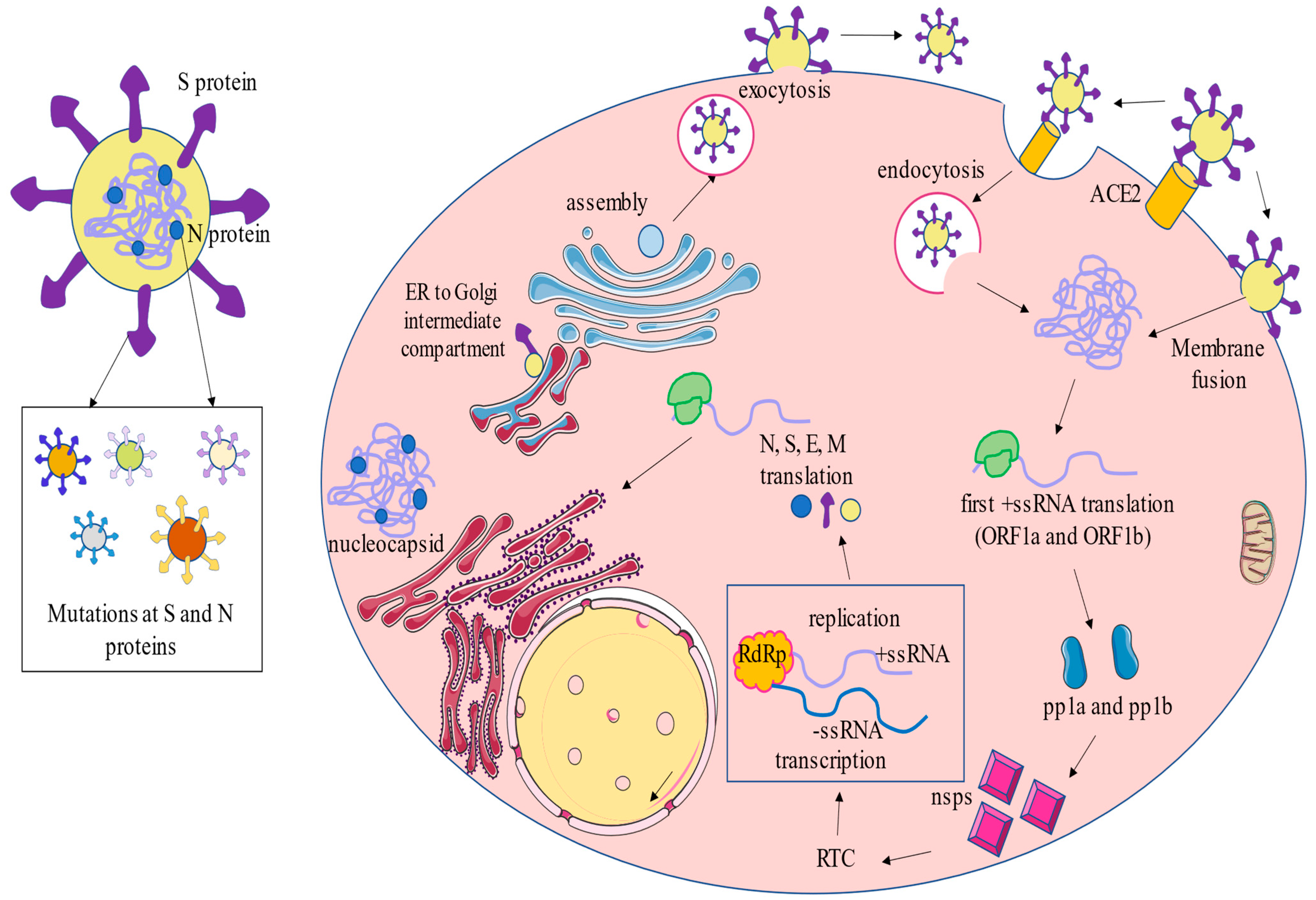
2.2. Infection Cycle
2.3. Immune Response and Vaccinology
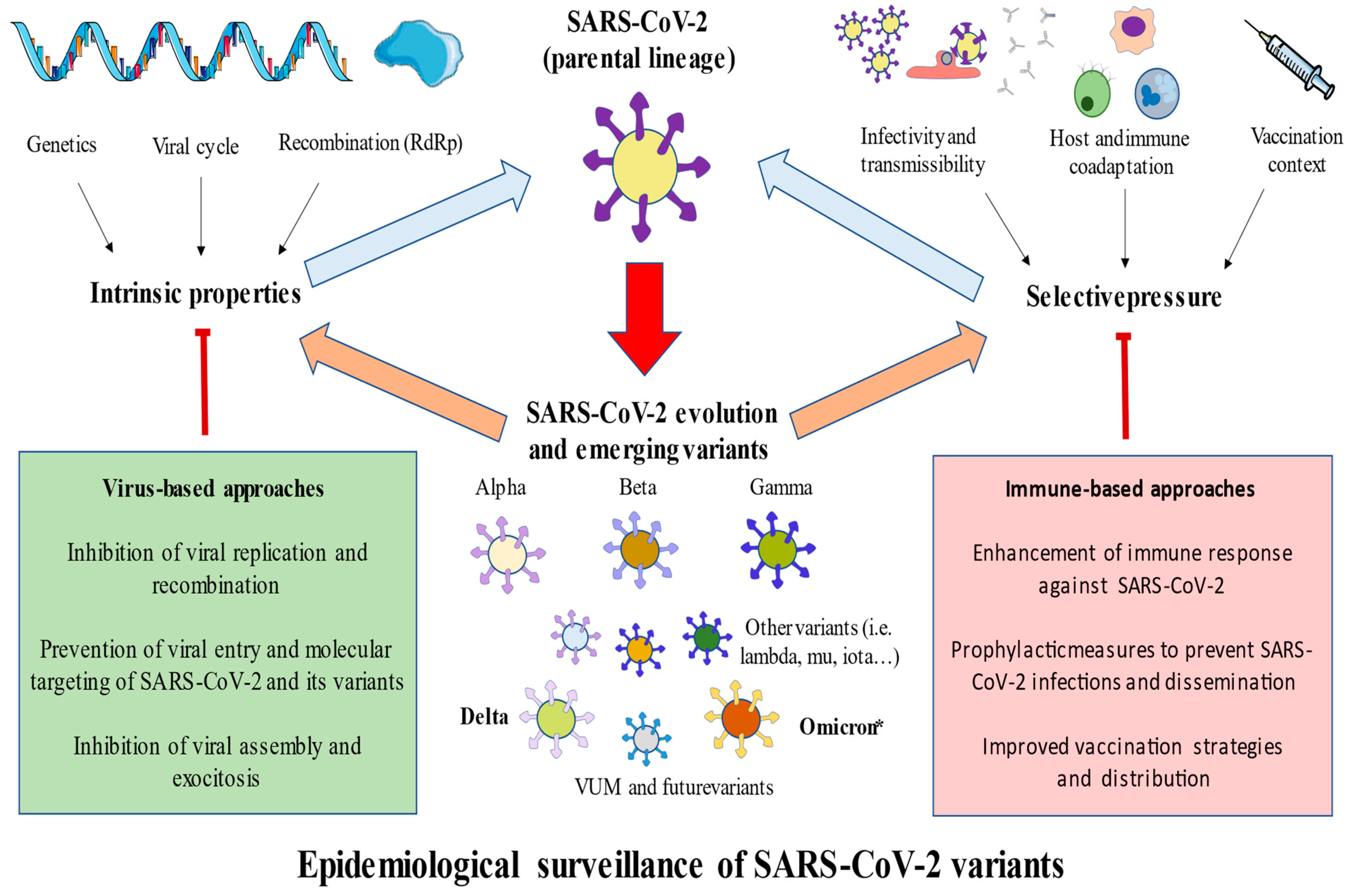
3. Main SARS-CoV-2 Variants
3.1. Alpha
3.2. Beta
3.3. Gamma
3.4. Delta
3.5. Omicron
3.6. Omicron Sublineages
3.7. Omicron-Hybrid Variants
4. Conclusions and Future Directions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vijayanand, P.; Wilkins, E.; Woodhead, M. Severe acute respiratory syndrome (SARS): A review. Clin. Med. 2004, 4, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kasuga, Y.; Zhu, B.; Jang, K.J.; Yoo, J.S. Innate immune sensing of coronavirus and viral evasion strategies. Exp. Mol. Med. 2021, 53, 723–736. [Google Scholar] [CrossRef] [PubMed]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Wu, D.; Wu, T.; Liu, Q.; Yang, Z. The SARS-CoV-2 outbreak: What we know. Int. J. Infect. Dis. 2020, 94, 44–48. [Google Scholar] [CrossRef]
- Abdelrahman, Z.; Li, M.; Wang, X. Comparative Review of SARS-CoV-2, SARS-CoV, MERS-CoV, and Influenza A Respiratory Viruses. Front. Immunol. 2020, 11, 552909. [Google Scholar] [CrossRef]
- Aimrane, A.; Laaradia, M.A.; Sereno, D.; Perrin, P.; Draoui, A.; Bougadir, B.; Hadach, M.; Zahir, M.; Fdil, N.; El Hiba, O.; et al. Insight into COVID-19’s epidemiology, pathology, and treatment. Heliyon 2022, 8, e08799. [Google Scholar] [CrossRef]
- Alsobaie, S. Understanding the Molecular Biology of SARS-CoV-2 and the COVID-19 Pandemic: A Review. Infect. Drug Resist. 2021, 14, 2259–2268. [Google Scholar] [CrossRef] [PubMed]
- Berman, J.J. Group IV Viruses: Single-Stranded (+)Sense RNA. In Taxonomic Guide to Infectious Diseases; Elsevier: Amsterdam, The Netherlands, 2012; pp. 237–246. [Google Scholar] [CrossRef]
- Zhu, C.; He, G.; Yin, Q.; Zeng, L.; Ye, X.; Shi, Y.; Xu, W. Molecular biology of the SARs-CoV-2 spike protein: A review of current knowledge. J. Med. Virol. 2021, 93, 5729. [Google Scholar] [CrossRef]
- Severe Acute Respiratory Syndrome Coronavirus 2 Isolate Wuhan-Hu-1, co-Nucleotide-NCBI. Available online: https://www.ncbi.nlm.nih.gov/nuccore/MN908947.3 (accessed on 18 March 2022).
- Chan, J.F.W.; Kok, K.H.; Zhu, Z.; Chu, H.; To, K.K.W.; Yuan, S.; Yuen, K.Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes. Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shang, J.; Du, L.; Han, N.; Lv, D.; Wang, J.; Yang, H.; Bai, L.; Tang, H. Severe acute respiratory syndrome coronavirus 2 for physicians: Molecular characteristics and host immunity (Review). Mol. Med. Rep. 2021, 23, 262. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yang, C.; Xu, X.F.; Xu, W.; Liu, S.W. Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–1149. [Google Scholar] [CrossRef] [PubMed]
- Jochheim, F.A.; Tegunov, D.; Hillen, H.S.; Schmitzová, J.; Kokic, G.; Dienemann, C.; Cramer, P. The structure of a dimeric form of SARS-CoV-2 polymerase. Commun. Biol. 2021, 4, 999. [Google Scholar] [CrossRef]
- Hillen, H.S.; Kokic, G.; Farnung, L.; Dienemann, C.; Tegunov, D.; Cramer, P. Structure of replicating SARS-CoV-2 polymerase. Nature 2020, 584, 154–156. [Google Scholar] [CrossRef] [PubMed]
- Naydenova, K.; Muir, K.W.; Wu, L.F.; Zhang, Z.; Coscia, F.; Peet, M.J.; Castro-Hartmann, P.; Qian, P.; Sader, K.; Dent, K.; et al. Structure of the SARS-CoV-2 RNA-dependent RNA polymerase in the presence of favipiravir-RTP. Proc. Natl. Acad. Sci. USA 2021, 118, e2021946118. [Google Scholar] [CrossRef]
- V’kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2020, 19, 155–170. [Google Scholar] [CrossRef]
- Wang, S.; Xu, X.; Wei, C.; Li, S.; Zhao, J.; Zheng, Y.; Liu, X.; Zeng, X.; Yuan, W.; Peng, S. Molecular evolutionary characteristics of SARS-CoV-2 emerging in the United States. J. Med. Virol. 2022, 94, 310–317. [Google Scholar] [CrossRef]
- Phan, T. Genetic diversity and evolution of SARS-CoV-2. Infect. Genet. Evol. 2020, 81, 104260. [Google Scholar] [CrossRef]
- Payne, S. Family Coronaviridae. In Viruses; Academic Press: Cambridge, MA, USA, 2017; p. 149. [Google Scholar] [CrossRef]
- Malune, P.; Piras, G.; Monne, M.; Fiamma, M.; Asproni, R.; Fancello, T.; Manai, A.; Carta, F.; Pira, G.; Fancello, P.; et al. Molecular Characterization of Severe Acute Respiratory Syndrome Coronavirus 2 Isolates From Central Inner Sardinia. Front. Microbiol. 2022, 12, 4335. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Thao, T.T.N.; Hoffmann, D.; Taddeo, A.; Ebert, N.; Labroussaa, F.; Pohlmann, A.; King, J.; Steiner, S.; Kelly, J.N.; et al. SARS-CoV-2 spike D614G change enhances replication and transmission. Nature 2021, 592, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Peng, F.; Wang, R.; Guan, K.; Jiang, T.; Xu, G.; Sun, J.; Chang, C. The deadly coronaviruses: The 2003 SARS pandemic and the 2020 novel coronavirus epidemic in China. J. Autoimmun. 2020, 109, 102434. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2021, 23, 3–20. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Li, W.; Feng, J.; da Silva, S.R.; Ju, E.; Zhang, H.; Chang, Y.; Moore, P.S.; Guo, H.; Gao, S.J. SARS-CoV-2 pseudovirus infectivity and expression of viral entry-related factors ACE2, TMPRSS2, Kim-1, and NRP-1 in human cells from the respiratory, urinary, digestive, reproductive, and immune systems. J. Med. Virol. 2021, 93, 6671–6685. [Google Scholar] [CrossRef]
- Glowacka, I.; Bertram, S.; Müller, M.A.; Allen, P.; Soilleux, E.; Pfefferle, S.; Steffen, I.; Tsegaye, T.S.; He, Y.; Gnirss, K.; et al. Evidence that TMPRSS2 activates the severe acute respiratory syndrome coronavirus spike protein for membrane fusion and reduces viral control by the humoral immune response. J. Virol. 2011, 85, 4122–4134. [Google Scholar] [CrossRef] [Green Version]
- Roe, M.K.; Junod, N.A.; Young, A.R.; Beachboard, D.C.; Stobart, C.C. Targeting novel structural and functional features of coronavirus protease nsp5 (3CLpro, Mpro) in the age of COVID-19. J. Gen. Virol. 2021, 102, 1558. [Google Scholar] [CrossRef]
- Das, A.; Ahmed, R.; Akhtar, S.; Begum, K.; Banu, S. An overview of basic molecular biology of SARS-CoV-2 and current COVID-19 prevention strategies. Gene Rep. 2021, 23, 101122. [Google Scholar] [CrossRef]
- Pizzato, M.; Baraldi, C.; Boscato Sopetto, G.; Finozzi, D.; Gentile, C.; Gentile, M.D.; Marconi, R.; Paladino, D.; Raoss, A.; Riedmiller, I.; et al. SARS-CoV-2 and the Host Cell: A Tale of Interactions. Front. Virol. 2022, 1, 46. [Google Scholar] [CrossRef]
- Espinoza, C.; Alarcón, M. The Immune Response to SARS-CoV-2: Mechanisms, Aging, Sequelae and Vaccines. Mini-Rev. Med. Chem. 2022; Epub ahead of print. [Google Scholar] [CrossRef]
- Chen, X.; Huang, Z.; Wang, J.; Zhao, S.; Wong, M.C.; Chong, K.C.; He, D.; Li, J. Ratio of asymptomatic COVID-19 cases among ascertained SARS-CoV-2 infections in different regions and population groups in 2020: A systematic review and meta-analysis including 130 123 infections from 241 studies. BMJ Open 2021, 11, e049752. [Google Scholar] [CrossRef]
- Wu, Z.; McGoogan, J.M. Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72314 Cases From the Chinese Center for Disease Control and Prevention. JAMA 2020, 323, 1239–1242. [Google Scholar] [CrossRef]
- Ortega, M.A.; Fraile-Martínez, O.; García-Montero, C.; García-Gallego, S.; Sánchez-Trujillo, L.; Torres-Carranza, D.; Álvarez-Mon, M.Á.; Pekarek, L.; García-Honduvilla, N.; Bujan, J.; et al. An integrative look at SARS-CoV-2 (Review). Int. J. Mol. Med. 2021, 47, 415–434. [Google Scholar] [CrossRef] [PubMed]
- Siddiqi, H.K.; Mehra, M.R. COVID-19 illness in native and immunosuppressed states: A clinical–therapeutic staging proposal. J. Hear. Lung Transplant. 2020, 39, 405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Azkur, A.K.; Akdis, M.; Azkur, D.; Sokolowska, M.; van de Veen, W.; Brüggen, M.C.; O’Mahony, L.; Gao, Y.; Nadeau, K.; Akdis, C.A. Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy 2020, 75, 1564–1581. [Google Scholar] [CrossRef] [PubMed]
- Rabaan, A.A.; Al-Ahmed, S.H.; Muhammad, J.; Khan, A.; Sule, A.A.; Tirupathi, R.; Al Mutair, A.; Alhumaid, S.; Al-Omari, A.; Dhawan, M.; et al. Role of Inflammatory Cytokines in COVID-19 Patients: A Review on Molecular Mechanisms, Immune Functions, Immunopathology and Immunomodulatory Drugs to Counter Cytokine Storm. Vaccines 2021, 9, 436. [Google Scholar] [CrossRef] [PubMed]
- Monserrat, J.; Asunsolo, A.; Gómez-Lahoz, A.; Ortega, M.A.; Gasalla, J.M.; Gasulla, Ó.; Fortuny-Profitós, J.; Mazaira-Font, F.A.; Román, M.T.; Arranz, A.; et al. Impact of the Innate Inflammatory Response on ICU Admission and Death in Hospitalized Patients with COVID-19. Biomedicines 2021, 9, 1675. [Google Scholar] [CrossRef] [PubMed]
- Costela-Ruiz, V.J.; Illescas-Montes, R.; Puerta-Puerta, J.M.; Ruiz, C.; Melguizo-Rodríguez, L. SARS-CoV-2 infection: The role of cytokines in COVID-19 disease. Cytokine Growth Factor Rev. 2020, 54, 62–75. [Google Scholar] [CrossRef]
- Álvarez-Mon, M.; Ortega, M.A.; Gasulla, Ó.; Fortuny-Profitós, J.; Mazaira-Font, F.A.; Saurina, P.; Monserrat, J.; Plana, M.N.; Troncoso, D.; Moreno, J.S.; et al. A predictive model and risk factors for case fatality of covid-19. J. Pers. Med. 2021, 11, 32. [Google Scholar] [CrossRef]
- Ruetalo, N.; Flehmig, B.; Schindler, M.; Pridzun, L.; Haage, A.; Reichenbächer, M.; Kirchner, T.; Kirchner, T.; Klingel, K.; Ranke, M.B.; et al. Long-Term Humoral Immune Response against SARS-CoV-2 after Natural Infection and Subsequent Vaccination According to WHO International Binding Antibody Units (BAU/mL). Viruses 2021, 13, 2336. [Google Scholar] [CrossRef]
- Krashias, G.; Deeba, E.; Constantinou, A.; Hadjiagapiou, M.; Koptides, D.; Richter, J.; Tryfonos, C.; Bashiardes, S.; Lambrianides, A.; Loizidou, M.A.; et al. Characterization of IgG Antibody Response against SARS-CoV-2 (COVID-19) in the Cypriot Population. Microorganisms 2021, 10, 85. [Google Scholar] [CrossRef]
- Misra, A.; Theel, E.S. Immunity to SARS-CoV-2: What Do We Know and Should We Be Testing for It? J. Clin. Microbiol. 2022, 60, e00482-21. [Google Scholar] [CrossRef]
- García-Montero, C.; Fraile-Martínez, O.; Bravo, C.; Torres-Carranza, D.; Sanchez-Trujillo, L.; Gómez-Lahoz, A.M.; Guijarro, L.G.; García-Honduvilla, N.; Asúnsolo, A.; Bujan, J.; et al. An Updated Review of SARS-CoV-2 Vaccines and the Importance of Effective Vaccination Programs in Pandemic Times. Vaccines 2021, 9, 433. [Google Scholar] [CrossRef] [PubMed]
- Giurgea, L.T.; Morens, D.M. Great Expectations of COVID-19 Herd Immunity. MBio 2022, 13, e03495-21. [Google Scholar] [CrossRef]
- Alexandridi, M.; Mazej, J.; Palermo, E.; Hiscott, J. The Coronavirus pandemic—2022: Viruses, variants & vaccines. Cytokine Growth Factor Rev. 2022, 63, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Moss, P. The T cell immune response against SARS-CoV-2. Nat. Immunol. 2022, 23, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Hall, V.; Foulkes, S.; Insalata, F.; Kirwan, P.; Saei, A.; Atti, A.; Wellington, E.; Khawam, J.; Munro, K.; Cole, M.; et al. Protection against SARS-CoV-2 after COVID-19 Vaccination and Previous Infection. N. Engl. J. Med. 2022, 386, 1207–1220. [Google Scholar] [CrossRef] [PubMed]
- Gazit, S.; Shlezinger, R.; Perez, G.; Lotan, R.; Peretz, A.; Ben-Tov, A.; Herzel, E.; Alapi, H.; Cohen, D.; Muhsen, K.; et al. SARS-CoV-2 Naturally Acquired Immunity vs. Vaccine-induced Immunity, Reinfections versus Breakthrough Infections: A Retrospective Cohort Study. Clin. Infect. Dis. 2022, ciac262. [Google Scholar] [CrossRef]
- Kang, C.K.; Kim, M.; Hong, J.; Kim, G.; Lee, S.; Chang, E.; Choe, P.G.; Kim, N.J.; Kim, I.S.; Seo, J.-Y.; et al. Distinct Immune Response at 1 Year Post-COVID-19 According to Disease Severity. Front. Immunol. 2022, 13, 1085. [Google Scholar] [CrossRef] [PubMed]
- Accorsi, E.K.; Britton, A.; Fleming-Dutra, K.E.; Smith, Z.R.; Shang, N.; Derado, G.; Miller, J.; Schrag, S.J.; Verani, J.R. Association Between 3 Doses of mRNA COVID-19 Vaccine and Symptomatic Infection Caused by the SARS-CoV-2 Omicron and Delta Variants. JAMA 2022, 327, 639–651. [Google Scholar] [CrossRef]
- Alefishat, E.; Jelinek, H.F.; Mousa, M.; Tay, G.K.; Alsafar, H.S. Immune response to SARS-CoV-2 variants: A focus on severity, susceptibility, and preexisting immunity. J. Infect. Public Health 2022, 15, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Röltgen, K.; Nielsen, S.C.A.; Silva, O.; Younes, S.F.; Zaslavsky, M.; Costales, C.; Yang, F.; Wirz, O.F.; Solis, D.; Hoh, R.A.; et al. Immune imprinting, breadth of variant recognition, and germinal center response in human SARS-CoV-2 infection and vaccination. Cell 2022, 185, 1025–1040.e14. [Google Scholar] [CrossRef]
- Andreano, E.; Paciello, I.; Piccini, G.; Manganaro, N.; Pileri, P.; Hyseni, I.; Leonardi, M.; Pantano, E.; Abbiento, V.; Benincasa, L.; et al. Hybrid immunity improves B cells and antibodies against SARS-CoV-2 variants. Nature 2021, 600, 530–535. [Google Scholar] [CrossRef] [PubMed]
- Pilz, S.; Theiler-Schwetz, V.; Trummer, C.; Krause, R.; Ioannidis, J.P.A. SARS-CoV-2 reinfections: Overview of efficacy and duration of natural and hybrid immunity. Environ. Res. 2022, 209, 112911. [Google Scholar] [CrossRef] [PubMed]
- European Centre for Disease Prevention and Control. Changes to List of SARS-CoV-2 Variants of Concern, Variants of Interest, and Variants under Monitoring; ECDC: Solna, Sweden, 2022. [Google Scholar]
- SARS-CoV-2 Variants of Concern as of 17 March 2022. Available online: https://www.ecdc.europa.eu/en/covid-19/variants-concern (accessed on 5 June 2022).
- Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/ (accessed on 5 June 2022).
- Walker, A.S.; Vihta, K.-D.; Gethings, O.; Pritchard, E.; Jones, J.; House, T.; Bell, I.; Bell, J.I.; Newton, J.N.; Farrar, J.; et al. Tracking the Emergence of SARS-CoV-2 Alpha Variant in the United Kingdom. N. Engl. J. Med. 2021, 385, 2582–2585. [Google Scholar] [CrossRef]
- Hart, W.S.; Miller, E.; Andrews, N.J.; Waight, P.; Maini, P.K.; Funk, S.; Thompson, R.N. Generation time of the alpha and delta SARS-CoV-2 variants: An epidemiological analysis. Lancet Infect. Dis. 2022, 22, 603–610. [Google Scholar] [CrossRef]
- SARS-CoV-2 Alpha Variant Cases Worldwide 2022|Statista. Available online: https://www.statista.com/statistics/1246219/number-alpha-variant-worldwide-by-country/ (accessed on 5 June 2022).
- Parra-Lucares, A.; Segura, P.; Rojas, V.; Pumarino, C.; Saint-Pierre, G.; Toro, L. Emergence of SARS-CoV-2 Variants in the World: How Could This Happen? Life 2022, 12, 194. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.; Mishra, S.; Chand, M.; Barrett, J.C.; Johnson, R.; Geidelberg, L.; Hinsley, W.R.; Laydon, D.J.; Dabrera, G.; O’Toole, Á.; et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature 2021, 593, 266–269. [Google Scholar] [CrossRef]
- Garcia Borrega, J.; Naendrup, J.H.; Heindel, K.; Hamacher, L.; Heger, E.; Di Cristanziano, V.; Deppe, A.C.; Dusse, F.; Wetsch, W.A.; Eichenauer, D.A.; et al. Clinical Course and Outcome of Patients with SARS-CoV-2 Alpha Variant Infection Compared to Patients with SARS-CoV-2 Wild-Type Infection Admitted to the ICU. Microorganisms 2021, 9, 1944. [Google Scholar] [CrossRef]
- Graham, M.S.; Sudre, C.H.; May, A.; Antonelli, M.; Murray, B.; Varsavsky, T.; Kläser, K.; Canas, L.S.; Molteni, E.; Modat, M.; et al. Changes in symptomatology, reinfection, and transmissibility associated with the SARS-CoV-2 variant B.1.1.7: An ecological study. Lancet Public Health 2021, 6, e335–e345. [Google Scholar] [CrossRef]
- Davies, N.G.; Jarvis, C.I.; van Zandvoort, K.; Clifford, S.; Sun, F.Y.; Funk, S.; Medley, G.; Jafari, Y.; Meakin, S.R.; Lowe, R.; et al. Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7. Nature 2021, 593, 270–274. [Google Scholar] [CrossRef]
- Eyre, D.W.; Taylor, D.; Purver, M.; Chapman, D.; Fowler, T.; Pouwels, K.B.; Walker, A.S.; Peto, T.E.A. Effect of COVID-19 Vaccination on Transmission of Alpha and Delta Variants. N. Engl. J. Med. 2022, 386, 744–756. [Google Scholar] [CrossRef]
- Lauring, A.S.; Tenforde, M.W.; Chappell, J.D.; Gaglani, M.; Ginde, A.A.; McNeal, T.; Ghamande, S.; Douin, D.J.; Talbot, H.K.; Casey, J.D.; et al. Clinical severity of, and effectiveness of mRNA vaccines against, covid-19 from omicron, delta, and alpha SARS-CoV-2 variants in the United States: Prospective observational study. BMJ 2022, 376, e069761. [Google Scholar] [CrossRef]
- Cevik, M.; Grubaugh, N.D.; Iwasaki, A.; Openshaw, P. COVID-19 vaccines: Keeping pace with SARS-CoV-2 variants. Cell 2021, 184, 5077–5081. [Google Scholar] [CrossRef]
- Duerr, R.; Dimartino, D.; Marier, C.; Zappile, P.; Wang, G.; Lighter, J.; Elbel, B.; Troxel, A.B.; Heguy, A. Dominance of Alpha and Iota variants in SARS-CoV-2 vaccine breakthrough infections in New York City. J. Clin. Investig. 2021, 131, e152702. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, G.R.; Yadav, P.D.; Abraham, P.; Nyayanit, D.A.; Sapkal, G.N.; Shete, A.M.; Gupta, N.; Vadrevu, K.M.; Ella, R.; Panda, S.; et al. Booster dose of the inactivated COVID-19 vaccine BBV152 (Covaxin) enhances the neutralizing antibody response against Alpha, Beta, Delta and Omicron variants of concern. J. Travel. Med. 2022, 29, taac039. [Google Scholar] [CrossRef] [PubMed]
- Heath, P.T.; Galiza, E.P.; Baxter, D.N.; Boffito, M.; Browne, D.; Burns, F.; Chadwick, D.R.; Clark, R.; Cosgrove, C.; Galloway, J.; et al. Safety and Efficacy of NVX-CoV2373 COVID-19 Vaccine. N. Engl. J. Med. 2021, 385, 1172–1183. [Google Scholar] [CrossRef]
- Puranik, A.; Lenehan, P.J.; Silvert, E.; Niesen, M.J.M.; Corchado-Garcia, J.; O’Horo, J.C.; Virk, A.; Swift, M.D.; Halamka, J.; Badley, A.D.; et al. Comparison of two highly effective mRNA vaccines for COVID-19 during periods of Alpha and Delta variant prevalence. medRxiv 2021. [Google Scholar] [CrossRef]
- Park, H.-S.; Shapiro, J.R.; Sitaras, I.; Woldemeskel, B.A.; Garliss, C.C.; Dziedzic, A.; Sachithanandham, J.; Jedlicka, A.E.; Caputo, C.A.; Rousseau, K.E.; et al. Adaptive immune responses in vaccinated patients with symptomatic SARS-CoV-2 Alpha infection. JCI Insight 2022, 7. [Google Scholar] [CrossRef]
- Domingo, P.; de Benito, N. Alpha variant SARS-CoV-2 infection: How it all starts. EBioMedicine 2021, 74, 103703. [Google Scholar] [CrossRef]
- Thorne, L.G.; Bouhaddou, M.; Reuschl, A.K.; Zuliani-Alvarez, L.; Polacco, B.; Pelin, A.; Batra, J.; Whelan, M.V.X.; Hosmillo, M.; Fossati, A.; et al. Evolution of enhanced innate immune evasion by SARS-CoV-2. Nature 2021, 602, 487–495. [Google Scholar] [CrossRef]
- Paniskaki, K.; Anft, M.; Meister, T.L.; Marheinecke, C.; Pfaender, S.; Skrzypczyk, S.; Seibert, F.S.; Thieme, C.J.; Konik, M.J.; Dolff, S.; et al. Immune Response in Moderate to Critical Breakthrough COVID-19 Infection After mRNA Vaccination. Front. Immunol. 2022, 13, 74. [Google Scholar] [CrossRef] [PubMed]
- Yadav, P.D.; Sarkale, P.; Razdan, A.; Gupta, N.; Nyayanit, D.A.; Sahay, R.R.; Potdar, V.; Patil, D.Y.; Baradkar, S.; Kumar, A.; et al. Isolation and characterization of SARS-CoV-2 Beta variant from UAE travelers. J. Infect. Public Health 2022, 15, 182–186. [Google Scholar] [CrossRef] [PubMed]
- Lippi, G.; Mattiuzzi, C.; Henry, B.M. Updated picture of SARS-CoV-2 variants and mutations. Diagnosis 2022, 9, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Zia, T.; Suleman, M.; Khan, T.; Ali, S.S.; Abbasi, A.A.; Mohammad, A.; Wei, D.Q. Higher infectivity of the SARS-CoV-2 new variants is associated with K417N/T, E484K, and N501Y mutants: An insight from structural data. J. Cell. Physiol. 2021, 236, 7045–7057. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Liu, Y.; Tang, X.; He, D. The Disease Severity and Clinical Outcomes of the SARS-CoV-2 Variants of Concern. Front. Public Health 2021, 9. [Google Scholar] [CrossRef]
- Butt, A.A.; Dargham, S.R.; Chemaitelly, H.; Al Khal, A.; Tang, P.; Hasan, M.R.; Coyle, P.V.; Thomas, A.G.; Borham, A.M.; Concepcion, E.G.; et al. Severity of Illness in Persons Infected With the SARS-CoV-2 Delta Variant vs Beta Variant in Qatar. JAMA Intern. Med. 2022, 182, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Riou, C.; Keeton, R.; Moyo-Gwete, T.; Hermanus, T.; Kgagudi, P.; Baguma, R.; Valley-Omar, Z.; Smith, M.; Tegally, H.; Doolabh, D.; et al. Escape from recognition of SARS-CoV-2 variant spike epitopes but overall preservation of T cell immunity. Sci. Transl. Med. 2022, 14, eabj6824. [Google Scholar] [CrossRef] [PubMed]
- Chemaitelly, H.; Bertollini, R.; Abu-Raddad, L.J. Efficacy of Natural Immunity against SARS-CoV-2 Reinfection with the Beta Variant. N. Engl. J. Med. 2021, 385, 2585–2586. [Google Scholar] [CrossRef]
- Becker, M.; Dulovic, A.; Junker, D.; Ruetalo, N.; Kaiser, P.D.; Pinilla, Y.T.; Heinzel, C.; Haering, J.; Traenkle, B.; Wagner, T.R.; et al. Immune response to SARS-CoV-2 variants of concern in vaccinated individuals. Nat. Commun. 2021, 12, 3109. [Google Scholar] [CrossRef]
- Duong, D. Alpha, Beta, Delta, Gamma: What’s important to know about SARS-CoV-2 variants of concern? CMAJ 2021, 193, E1059–E1060. [Google Scholar] [CrossRef]
- Mahase, E. COVID-19: Novavax vaccine efficacy is 86% against UK variant and 60% against South African variant. BMJ 2021, 372, n296. [Google Scholar] [CrossRef]
- Mor, O.; Zuckerman, N.S.; Hazan, I.; Fluss, R.; Ash, N.; Ginish, N.; Mendelson, E.; Alroy-Preis, S.; Freedman, L.; Huppert, A. BNT162b2 vaccine effectiveness was marginally affected by the SARS-CoV-2 beta variant in fully vaccinated individuals. J. Clin. Epidemiol. 2022, 142, 38–44. [Google Scholar] [CrossRef]
- Singer, S.R.; Angulo, F.J.; Swerdlow, D.L.; McLaughlin, J.M.; Hazan, I.; Ginish, N.; Anis, E.; Mendelson, E.; Mor, O.; Zuckerman, N.S.; et al. Effectiveness of BNT162b2 mRNA COVID-19 vaccine against SARS-CoV-2 variant Beta (B.1.351) among persons identified through contact tracing in Israel: A prospective cohort study. EClinicalMedicine 2021, 42, 101190. [Google Scholar] [CrossRef] [PubMed]
- Naveca, F.G.; Nascimento, V.; de Souza, V.C.; Corado, A.D.L.; Nascimento, F.; Silva, G.; Costa, Á.; Duarte, D.; Pessoa, K.; Mejía, M.; et al. COVID-19 in Amazonas, Brazil, was driven by the persistence of endemic lineages and P.1 emergence. Nat. Med. 2021, 27, 1230–1238. [Google Scholar] [CrossRef]
- Naveca, F.G.; Nascimento, V.; Souza, V.; Corado, A.D.L.; Nascimento, F.; Silva, G.; Mejía, M.C.; Brandão, M.J.; Costa, Á.; Duarte, D.; et al. Spread of Gamma (P.1) Sub-Lineages Carrying Spike Mutations Close to the Furin Cleavage Site and Deletions in the N-Terminal Domain Drives Ongoing Transmission of SARS-CoV-2 in Amazonas, Brazil. Microbiol. Spectr. 2022, 10. [Google Scholar] [CrossRef] [PubMed]
- Padilha, D.A.; Benetti-Filho, V.; Moreira, R.S.; Soratto, T.A.; Maia, A.; Christoff, A.P.; Barazzetti, F.H.; Schörner, M.A.; Ferrari, F.L.; Martins, C.L.; et al. Emergence of two distinct SARS-CoV-2 Gamma variants and the rapid spread of P.1-like-II SARS-CoV-2 during the second wave of COVID-19 in Santa Catarina, Southern Brazil. medRxiv 2022. [Google Scholar] [CrossRef]
- Tagliamonte, M.S.; Mavian, C.; Zainabadi, K.; Cash, M.N.; Lednicky, J.A.; Magalis, B.R.; Riva, A.; Deschamps, M.M.; Liautaud, B.; Rouzier, V.; et al. Rapid Emergence and Spread of Severe Acute Respiratory Syndrome Coronavirus 2 Gamma (P.1) Variant in Haiti. Clin. Infect. Dis. 2021, 74, 2057–2060. [Google Scholar] [CrossRef] [PubMed]
- Nicolete, V.C.; Rodrigues, P.T.; Fernandes, A.R.J.; Corder, R.M.; Tonini, J.; Buss, L.F.; Sales, F.C.; Faria, N.R.; Sabino, E.C.; Castro, M.C.; et al. Epidemiology of COVID-19 after Emergence of SARS-CoV-2 Gamma Variant, Brazilian Amazon, 2020–2021. Emerg. Infect. Dis. 2022, 28, 709–712. [Google Scholar] [CrossRef]
- Fiolet, T.; Kherabi, Y.; MacDonald, C.J.; Ghosn, J.; Peiffer-Smadja, N. Comparing COVID-19 vaccines for their characteristics, efficacy and effectiveness against SARS-CoV-2 and variants of concern: A narrative review. Clin. Microbiol. Infect. 2022, 28, 202–221. [Google Scholar] [CrossRef]
- Chmielewska, A.M.; Czarnota, A.; Bieńkowska-Szewczyk, K.; Grzyb, K. Immune response against SARS-CoV-2 variants: The role of neutralization assays. NPJ Vaccines 2021, 6, 142. [Google Scholar] [CrossRef]
- Ferreira, I.A.T.M.; Kemp, S.A.; Datir, R.; Saito, A.; Meng, B.; Rakshit, P.; Takaori-Kondo, A.; Kosugi, Y.; Uriu, K.; Kimura, I.; et al. SARS-CoV-2 B.1.617 Mutations L452R and E484Q Are Not Synergistic for Antibody Evasion. J. Infect. Dis. 2021, 224, 989–994. [Google Scholar] [CrossRef] [PubMed]
- Shiehzadegan, S.; Alaghemand, N.; Fox, M.; Venketaraman, V. Analysis of the Delta Variant, B.1.617.2 COVID-19. Clin. Pract. 2021, 11, 778–784. [Google Scholar] [CrossRef]
- Gunadi; Hakim, M. S.; Wibawa, H.; Marcellus; Setiawaty, V.; Slamet; Trisnawati, I.; Supriyati, E.; El Khair, R.; Iskandar, K.; et al. Is the infection of the SARS-CoV-2 Delta variant associated with the outcomes of COVID-19 patients? medRxiv 2021, 10. [Google Scholar] [CrossRef]
- Lee, B.U. Why Does the SARS-CoV-2 Delta VOC Spread So Rapidly? Universal Conditions for the Rapid Spread of Respiratory Viruses, Minimum Viral Loads for Viral Aerosol Generation, Effects of Vaccination on Viral Aerosol Generation, and Viral Aerosol Clouds. Int. J. Environ. Res. Public Health 2021, 18, 9804. [Google Scholar] [CrossRef] [PubMed]
- Riediker, M.; Briceno-Ayala, L.; Ichihara, G.; Albani, D.; Poffet, D.; Tsai, D.H.; Iff, S.; Monn, C. Higher viral load and infectivity increase risk of aerosol transmission for Delta and Omicron variants of SARS-CoV-2. Swiss Med. Wkly. 2022, 152, w30133. [Google Scholar] [CrossRef] [PubMed]
- Tani-Sassa, C.; Iwasaki, Y.; Ichimura, N.; Nagano, K.; Takatsuki, Y.; Yuasa, S.; Takahashi, Y.; Nakajima, J.; Sonobe, K.; Nukui, Y.; et al. Viral loads and profile of the patients infected with SARS-CoV-2 Delta, Alpha, or R.1 variants in Tokyo. J. Med. Virol. 2022, 94, 1707–1710. [Google Scholar] [CrossRef] [PubMed]
- Yazdanpanah, F.; Yazdanpanah, S.; Rezaei, N. COVID-19 Delta variation; more contagious or more pernicious? Acta Biomed. 2022, 92, e2021454. [Google Scholar] [CrossRef]
- Aleem, A.; Samad, A.B.A.; Slenker, A.K. Emerging Variants of SARS-CoV-2 and Novel Therapeutics against Coronavirus (COVID-19); StatPearls: Treasure Island, FL, USA, 2022. [Google Scholar]
- Kannan, S.R.; Spratt, A.N.; Cohen, A.R.; Naqvi, S.H.; Chand, H.S.; Quinn, T.P.; Lorson, C.L.; Byrareddy, S.N.; Singh, K. Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. J. Autoimmun. 2021, 124, 102715. [Google Scholar] [CrossRef]
- Thakur, N.; Gallo, G.; Newman, J.; Peacock, T.P.; Biasetti, L.; Hall, C.N.; Wright, E.; Barclay, W.; Bailey, D. SARS-CoV-2 variants of concern alpha, beta, gamma and delta have extended ACE2 receptor host ranges. J. Gen. Virol. 2022, 103, 001735. [Google Scholar] [CrossRef]
- Rahman, F.I.; Ether, S.A.; Islam, M.R. The “Delta Plus” COVID-19 variant has evolved to become the next potential variant of concern: Mutation history and measures of prevention. J. Basic Clin. Physiol. Pharmacol. 2021, 33, 109–112. [Google Scholar] [CrossRef]
- Ong, S.W.X.; Chiew, C.J.; Ang, L.W.; Mak, T.-M.; Cui, L.; Toh, M.P.H.S.; Lim, Y.D.; Lee, P.H.; Lee, T.H.; Chia, P.Y.; et al. Clinical and virological features of SARS-CoV-2 variants of concern: A retrospective cohort study comparing B.1.1.7 (Alpha), B.1.315 (Beta), and B.1.617.2 (Delta). Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef] [PubMed]
- Twohig, K.A.; Nyberg, T.; Zaidi, A.; Thelwall, S.; Sinnathamby, M.A.; Aliabadi, S.; Seaman, S.R.; Harris, R.J.; Hope, R.; Lopez-Bernal, J.; et al. Hospital admission and emergency care attendance risk for SARS-CoV-2 delta (B.1.617.2) compared with alpha (B.1.1.7) variants of concern: A cohort study. Lancet Infect. Dis. 2022, 22, 35–42. [Google Scholar] [CrossRef]
- Hu, Z.; Huang, X.; Zhang, J.; Fu, S.; Ding, D.; Tao, Z. Differences in Clinical Characteristics Between Delta Variant and Wild-Type SARS-CoV-2 Infected Patients. Front. Med. 2022, 8, 2526. [Google Scholar] [CrossRef] [PubMed]
- Chohan, F.; Ishak, A.; Alderette, T.; Rad, P.; Michel, G. Clinical Presentation of a COVID-19 Delta Variant Patient: Case Report and Literature Review. Cureus 2021, 13, e18603. [Google Scholar] [CrossRef] [PubMed]
- Risk, M.; Shen, C.; Hayek, S.S.; Holevinski, L.; Schiopu, E.; Freed, G.; Akin, C.; Zhao, L. Comparative Effectiveness of COVID-19 Vaccines against the Delta Variant. Clin. Infect. Dis. 2022. [Google Scholar] [CrossRef]
- Zhang, M.; Gong, Y.; Jiao, S. Neutralization heterogeneity of circulating SARS-CoV-2 variants to sera elicited by a vaccinee or convalescent. Future Virol. 2022, 17, 403–413. [Google Scholar] [CrossRef]
- McKeigue, P.M.; McAllister, D.A.; Hutchinson, S.J.; Robertson, C.; Stockton, D.; Colhoun, H.M. Vaccine efficacy against severe COVID-19 in relation to delta variant (B.1.617.2) and time since second dose in patients in Scotland (REACT-SCOT): A case-control study. Lancet Respir. Med. 2022, 10, 566–572. [Google Scholar] [CrossRef]
- Mlcochova, P.; Kemp, S.; Dhar, M.S.; Papa, G.; Meng, B.; Ferreira, I.A.T.M.; Datir, R.; Collier, D.A.; Albecka, A.; Singh, S.; et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 2021, 599, 114–119. [Google Scholar] [CrossRef]
- Planas, D.; Veyer, D.; Baidaliuk, A.; Staropoli, I.; Guivel-Benhassine, F.; Rajah, M.M.; Planchais, C.; Porrot, F.; Robillard, N.; Puech, J.; et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 2021, 596, 276–280. [Google Scholar] [CrossRef]
- Gao, S.J.; Guo, H.; Luo, G. Omicron variant (B.1.1.529) of SARS-CoV-2, a global urgent public health alert! J. Med. Virol. 2022, 94, 1255–1256. [Google Scholar] [CrossRef]
- Ferré, V.M.; Peiffer-Smadja, N.; Visseaux, B.; Descamps, D.; Ghosn, J.; Charpentier, C. Omicron SARS-CoV-2 variant: What we know and what we don’t. Anaesth. Crit. Care Pain Med. 2022, 41, 100998. [Google Scholar] [CrossRef] [PubMed]
- Meo, S.A.; Meo, A.S.; Al-Jassir, F.F.; Klonoff, D.C. Omicron SARS-CoV-2 new variant: Global prevalence and biological and clinical characteristics. Eur. Rev. Med. Pharmacol. Sci. 2021, 25, 8012–8018. [Google Scholar] [CrossRef] [PubMed]
- Rahmani, S.; Rezaei, N. Omicron (B.1.1.529) variant: Development, dissemination, and dominance. J. Med. Virol. 2022, 94, 1787–1788. [Google Scholar] [CrossRef] [PubMed]
- Lennerstrand, J.; Svensson, L.; Lundkvist, A. How did Omicron evolve and why does this SARS-CoV-2 variant spread so fast? Lakartidningen 2022, 119, 21242. [Google Scholar]
- Wei, C.; Shan, K.J.; Wang, W.; Zhang, S.; Huan, Q.; Qian, W. Evidence for a mouse origin of the SARS-CoV-2 Omicron variant. J. Genet. Genom. 2021, 48, 1111–1121. [Google Scholar] [CrossRef]
- Thakur, V.; Ratho, R.K. OMICRON (B.1.1.529): A new SARS-CoV-2 variant of concern mounting worldwide fear. J. Med. Virol. 2022, 94, 1821–1824. [Google Scholar] [CrossRef]
- Bhattacharyya, R.P.; Hanage, W.P. Challenges in Inferring Intrinsic Severity of the SARS-CoV-2 Omicron Variant. N. Engl. J. Med. 2022, 386, e14. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Huo, J.; Zhou, D.; Zahradník, J.; Supasa, P.; Liu, C.; Duyvesteyn, H.M.E.; Ginn, H.M.; Mentzer, A.J.; Tuekprakhon, A.; et al. SARS-CoV-2 Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell 2022, 185, 467–484.e15. [Google Scholar] [CrossRef]
- Andrews, N.; Stowe, J.; Kirsebom, F.; Toffa, S.; Rickeard, T.; Gallagher, E.; Gower, C.; Kall, M.; Groves, N.; O’Connell, A.-M.; et al. COVID-19 Vaccine Effectiveness against the Omicron (B.1.1.529) Variant. N. Engl. J. Med. 2022, 386, 1532–1546. [Google Scholar] [CrossRef]
- Minka, S.O.; Minka, F.H. A tabulated summary of the evidence on humoral and cellular responses to the SARS-CoV-2 Omicron VOC, as well as vaccine efficacy against this variant. Immunol. Lett. 2022, 243, 38–43. [Google Scholar] [CrossRef]
- Yadav, P.D.; Sapkal, G.N.; Sahay, R.R.; Potdar, V.A.; Deshpande, G.R.; Patil, D.Y.; Nyayanit, D.A.; Shete, A.M.; Shastri, J.; Awate, P.; et al. Substantial immune response in Omicron infected breakthrough and unvaccinated individuals against SARS-CoV-2 variants of concern. J. Infect. 2022, 84, e80–e81. [Google Scholar] [CrossRef] [PubMed]
- Adam, D. Will Omicron end the pandemic? Here’s what experts say. Nature 2022, 602, 20–21. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Chen, L.; Shi, Y. Booster COVID-19 vaccination against the SARS-CoV-2 Omicron variant: A systematic review. Hum. Vaccines Immunother. 2022, 1–19. [Google Scholar] [CrossRef]
- Regev-Yochay, G.; Gonen, T.; Gilboa, M.; Mandelboim, M.; Indenbaum, V.; Amit, S.; Meltzer, L.; Asraf, K.; Cohen, C.; Fluss, R.; et al. Efficacy of a Fourth Dose of COVID-19 mRNA Vaccine against Omicron. N. Engl. J. Med. 2022, 386, 1377–1380. [Google Scholar] [CrossRef] [PubMed]
- Magen, O.; Waxman, J.G.; Makov-Assif, M.; Vered, R.; Dicker, D.; Hernán, M.A.; Lipsitch, M.; Reis, B.Y.; Balicer, R.D.; Dagan, N. Fourth Dose of BNT162b2 mRNA COVID-19 Vaccine in a Nationwide Setting. N. Engl. J. Med. 2022, 386, 1603–1614. [Google Scholar] [CrossRef]
- Mungmunpuntipantip, R.; Wiwanitkit, V. Fourth dose of COVID-19 vaccine: Does it still contribute any additional immunoprotection? J. Taibah Univ. Med. Sci. 2022, 17, 533–536. [Google Scholar] [CrossRef]
- Mallapaty, S. Fourth dose of COVID vaccine offers only slight boost against Omicron infection. Nature 2022. [Google Scholar] [CrossRef]
- Li, X. Omicron: Call for updated vaccines. J. Med. Virol. 2022, 94, 1261–1263. [Google Scholar] [CrossRef]
- Po, A.L.W. Omicron variant as nature’s solution to the COVID-19 pandemic. J. Clin. Pharm. Ther. 2022, 47, 3–5. [Google Scholar] [CrossRef]
- Vauhkonen, H.; Nguyen, P.T.; Kant, R.; Plyusnin, I.; Erdin, M.; Kurkela, S.; Liimatainen, H.; Ikonen, N.; Blomqvist, S.; Liitsola, K.; et al. Introduction and Rapid Spread of SARS-CoV-2 Omicron Variant and Dynamics of BA.1 and BA.1.1 Sublineages, Finland, December 2021. Emerg. Infect. Dis. 2022, 28. [Google Scholar] [CrossRef]
- Chen, J.; Wei, G.-W. Omicron BA.2 (B.1.1.529.2): High potential to becoming the next dominating variant. arXiv 2022, arXiv:2202.05031v1. [Google Scholar]
- Hirotsu, Y.; Maejima, M.; Shibusawa, M.; Natori, Y.; Nagakubo, Y.; Hosaka, K.; Sueki, H.; Mochizuki, H.; Tsutsui, T.; Kakizaki, Y.; et al. SARS-CoV-2 Omicron sublineage BA.2 replaces BA.1.1: Genomic surveillance in Japan from September 2021 to March 2022. J. Infect. 2022. [Google Scholar] [CrossRef] [PubMed]
- Bowen, J.E.; Sprouse, K.R.; Walls, A.C.; Mazzitelli, I.G.; Logue, J.K.; Franko, N.M.; Ahmed, K.; Shariq, A.; Cameroni, E.; Gori, A.; et al. Omicron BA.1 and BA.2 neutralizing activity elicited by a comprehensive panel of human vaccines. bioRxiv Prepr. Serv. Biol. 2022. [Google Scholar] [CrossRef]
- Yu, J.; Collier, A.Y.; Rowe, M.; Mardas, F.; Ventura, J.D.; Wan, H.; Miller, J.; Powers, O.; Chung, B.; Siamatu, M.; et al. Comparable Neutralization of the SARS-CoV-2 Omicron BA.1 and BA.2 Variants. medRxiv 2022. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; Chu, H. Pathogenicity of SARS-CoV-2 Omicron BA.1.1 in hamsters. eBioMedicine 2022, 80, 104035. [Google Scholar] [CrossRef]
- Maxmen, A. Are new Omicron subvariants a threat? Here’s how scientists are keeping watch. Nature 2022, 604, 605–606. [Google Scholar] [CrossRef]
- Wang, L.; Gao, G.F. The “Wolf” Is Indeed Coming: Recombinant “Deltacron” SARS-CoV-2 Detected. China CDC Wkly. 2022, 4, 285–287. [Google Scholar] [CrossRef]
- SARS-CoV-2 Variants of Concern and Variants Under Investigation in England, Technical Briefing 32. 17 December 2021. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/1042688/RA_Technical_Briefing_32_DRAFT_17_December_2021_2021_12_17.pdf (accessed on 20 May 2022).
- Nicola, M.; Alsafi, Z.; Sohrabi, C.; Kerwan, A.; Al-Jabir, A.; Iosifidis, C.; Agha, M.; Agha, R. The socio-economic implications of the coronavirus pandemic (COVID-19): A review. Int. J. Surg. 2020, 78, 185–193. [Google Scholar] [CrossRef]
- Xiong, J.; Lipsitz, O.; Nasri, F.; Lui, L.M.W.; Gill, H.; Phan, L.; Chen-Li, D.; Iacobucci, M.; Ho, R.; Majeed, A.; et al. Impact of COVID-19 pandemic on mental health in the general population: A systematic review. J. Affect. Disord. 2020, 277, 55–64. [Google Scholar] [CrossRef]
| Variant | Current Epidemiological Classification (ECDC) | First Report | Mutations in the Spike Protein of Interest | Clinical Manifestations | Immune Evasion | Vaccination Cocerns | References |
|---|---|---|---|---|---|---|---|
| Alpha (B.1.1.7) | De-escalated variant | United Kingdom, September 2020. | N501Y, D614G, P681H | Similar presentation to wild-type SARS-CoV-2, although some studies observed an increased mortality and severity | Possible influence in the innate immune system due to some mutations in the N protein, Orf9b, and Orf6 | Two doses of vaccine, especially mRNA-1273, appears to exert protective effects against this variant. | [8,9,45,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78] |
| Beta (B.1.351) | De-escalated variant | South Africa, May 2020. | K417N, E484K, N501Y, D614G, A701V | Increased risk of hospitalization, ICU admission, and mortality in comparison to Alpha and Gamma variants, but less than Delta | Spike mutations of Beta variant are not recognized by T-cells, but the immunity is equally preserved even 1 year after primary infection. E484K mutation may lead to escape from immune response | Some vaccines have presented a reduced efficacy against this variant regarding its transmission and mild-to-moderate presentations, although they exert a protective role against severe or fatal disease | [8,9,80,91] |
| Gamma (P.1) | De-escalated variant | Brazil, November 2020. | K417T, E484K, N501Y, D614G, H655Y | No significant association between Gamma variant infections and the severity of COVID-19 symptoms has been reported | E484K mutation may lead to escape from immune response | Little information available, but it seems that mRNA-based vaccines, AZD1222, and CoronaVac are effective in preventing symptomatic COVID-19 and severe infections against this variant | [8,9,81,92,93,94,95,96,97,98] |
| Delta (B.1.617.2) | Variant of concern (VOC) | India, October 2020. | L452R, T478K, D614G, P681R. | Delta variant presents a lessened time interval between disease onset to hospitalization in comparison to the wild-type variant, while also showing significant changes in hematological profile | 6-fold less sensitive to serum neutralizing antibodies from recovered individuals, and 8-fold less sensitive to vaccine-elicited antibodies, compared with wild-type SARS-CoV-2. | While the overall efficacy of the vaccines are diminished with this variant, mRNA-based vaccines (Pfizer and Moderna) provide greater protection against this variant | [8,9,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118] |
| Omicron (B.1.1.529) | Variant of concern (VOC) | Identified in multiple regions in November 2021. | A67V, del69-70, T95I, del142-144, Y145D, del211, L212I, ins214EPE, G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y, Y505H, T547K, D614G, H655Y, N679K, P681H, N764K, D796Y, N856K, Q954H, N969K, L981F | Despite some evidence suggesting that this variant leads to milder clinical presentations than Delta, it is difficult to establish the intrinsic severity of this variant due to different factors (i.e., the global percentage of vaccinated people) | Mutations in the ACE-2 binding site boost the immune escape of this variant, especially from neutralizing antibody responses | Three vaccination doses are needed to elicit a more appropriate immune response against symptomatic or non-symptomatic infections, transmission, and serious manifestations. Being infected by Omicron seems to confer a greater protection against this and other variants. | [8,9,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137] |
| Omicron sublineage BA.1 and BA.1.1 | Variant of concern (VOC) | First recognized in South Africa and Botswana in November 2021; they were predominant variants until the onset of BA.2 | A67V, del69-70, T95I, G142D, del143-145, N211I, del212, ins215EPE, G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y, Y505H, T547K, D614G, H655Y, N679K, P681H, N764K, D796Y, N856K, Q954H, N969K, L981F BA.1.1 = BA.1 with the spike R346K substitution | Similar to B.1.1.529 | Boosted immune escape, similar to BA.2 | Similar to B.1.1.529 | [138,139,140,141,142,143] |
| BA.2 | Variant of concern (VOC) | Dominant Omicron variant | G142D, N211I, del212, V213G, G339D, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, S477N, T478K, E484A, Q493R, Q498R, N501Y, Y505H, D614G, H655Y, N679K, P681H, N764K, D796Y, Q954H, N969K | Similar to B.1.1.529 | Boosted immune escape, similar to BA.1, but increased transmissibility. | Similar to B.1.1.529 | [138,139,140,141,142,143] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ortega, M.A.; García-Montero, C.; Fraile-Martinez, O.; Colet, P.; Baizhaxynova, A.; Mukhtarova, K.; Alvarez-Mon, M.; Kanatova, K.; Asúnsolo, A.; Sarría-Santamera, A. Recapping the Features of SARS-CoV-2 and Its Main Variants: Status and Future Paths. J. Pers. Med. 2022, 12, 995. https://doi.org/10.3390/jpm12060995
Ortega MA, García-Montero C, Fraile-Martinez O, Colet P, Baizhaxynova A, Mukhtarova K, Alvarez-Mon M, Kanatova K, Asúnsolo A, Sarría-Santamera A. Recapping the Features of SARS-CoV-2 and Its Main Variants: Status and Future Paths. Journal of Personalized Medicine. 2022; 12(6):995. https://doi.org/10.3390/jpm12060995
Chicago/Turabian StyleOrtega, Miguel A., Cielo García-Montero, Oscar Fraile-Martinez, Paolo Colet, Ardak Baizhaxynova, Kymbat Mukhtarova, Melchor Alvarez-Mon, Kaznagul Kanatova, Angel Asúnsolo, and Antonio Sarría-Santamera. 2022. "Recapping the Features of SARS-CoV-2 and Its Main Variants: Status and Future Paths" Journal of Personalized Medicine 12, no. 6: 995. https://doi.org/10.3390/jpm12060995
APA StyleOrtega, M. A., García-Montero, C., Fraile-Martinez, O., Colet, P., Baizhaxynova, A., Mukhtarova, K., Alvarez-Mon, M., Kanatova, K., Asúnsolo, A., & Sarría-Santamera, A. (2022). Recapping the Features of SARS-CoV-2 and Its Main Variants: Status and Future Paths. Journal of Personalized Medicine, 12(6), 995. https://doi.org/10.3390/jpm12060995









