Analyzing Precision Medicine Utilization with Real-World Data: A Scoping Review
Abstract
:1. Introduction
2. Materials and Methods
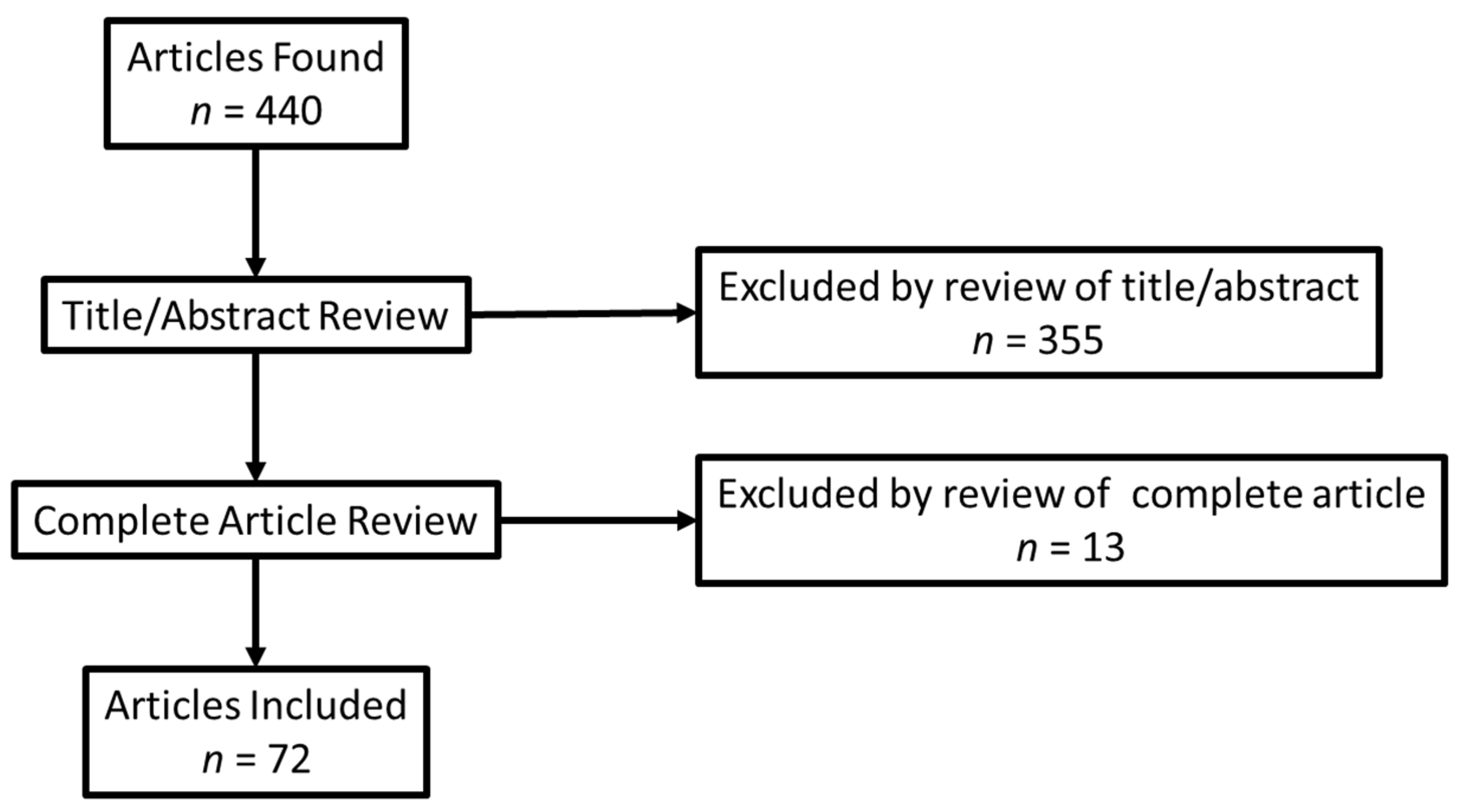
2.1. Inclusion/Exclusion
2.2. Data Abstraction
- Demographic (PubMed ID, first author, year);
- Research question (research question);
- Name of RWD sources and whether or not the study used an integrated or merged dataset from two or more sources (clinical data source, admin claims (commercial) source, admin claims (public) source, registry source);
- Laboratory name that performed the PM testing, if available (lab name, other);
- Condition or disease, including if it was cancer or non-cancer condition (condition);
- Test name and/or genes, including if the test was BRCA, gene expression profiling/Oncotype Dx, multigene pane, tumor sequencing, WES/WGS, or other (test name and/or genes).
2.3. Analysis
3. Results
4. Discussion
Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wetterstrand, K.A. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Available online: https://www.genome.gov/sequencingcostsdata (accessed on 18 February 2022).
- Grapov, D.; Fahrmann, J.; Wanichthanarak, K.; Khoomrung, S. Rise of Deep Learning for Genomic, Proteomic, and Metabolomic Data Integration in Precision Medicine. Omics 2018, 22, 630–636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phillips, K.A.; Douglas, M.P.; Marshall, D.A. Expanding Use of Clinical Genome Sequencing and the Need for More Data on Implementation. JAMA 2020, 324, 2029–2030. [Google Scholar] [CrossRef] [PubMed]
- Deverka, P.A.; Douglas, M.P.; Phillips, K.A. Use of Real-World Evidence in US Payer Coverage Decision-Making for Next-Generation Sequencing-Based Tests: Challenges, Opportunities, and Potential Solutions. Value Health 2020, 23, 540–550. [Google Scholar] [CrossRef] [PubMed]
- Rudrapatna, V.A.; Butte, A.J. Opportunities and challenges in using real-world data for health care. J. Clin. Investig. 2020, 130, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Crown, W.H. Real-World Evidence, Causal Inference, and Machine Learning. Value Health 2019, 22, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Teutsch, S.M.; Bradley, L.A.; Palomaki, G.E.; Haddow, J.E.; Piper, M.; Calonge, N.; Dotson, W.D.; Douglas, M.P.; Berg, A.O. The Evaluation of Genomic Applications in Practice and Prevention (EGAPP) Initiative: Methods of the EGAPP Working Group. Genet. Med. 2009, 11, 3–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kolata, G. Breaking Ranks, Lab Offers Test to Assess Risk of Breast Cancer. New York Times, 1 April 1996. Available online: https://www.nytimes.com/1996/04/01/us/breaking-ranks-lab-offers-test-to-assess-risk-of-breast-cancer.html (accessed on 11 December 2021).
- Phillips, K.A.; Douglas, M.P.; Wordsworth, S.; Buchanan, J.; Marshall, D.A. Availability and funding of clinical genomic sequencing globally. BMJ Glob. Health 2021, 6, e004415. [Google Scholar] [CrossRef] [PubMed]
- Garrison, L.P., Jr.; Neumann, P.J.; Erickson, P.; Marshall, D.; Mullins, C.D. Using real-world data for coverage and payment decisions: The ISPOR Real-World Data Task Force report. Value Health 2007, 10, 326–335. [Google Scholar] [CrossRef] [Green Version]
- Roberts, M.C.; Kurian, A.W.; Petkov, V.I. Uptake of the 21-Gene Assay among Women with Node-Positive, Hormone Receptor-Positive Breast Cancer. J. Natl. Compr. Cancer Netw. 2019, 17, 662–668. [Google Scholar] [CrossRef] [PubMed]
- Oncotype DX Database (2004–2016) with or without SES/Rurality: National Cancer Institute Surveillance, Epidemiology, and End Results Program. 2022. Available online: https://seer.cancer.gov/seerstat/databases/oncotype-dx/index.html (accessed on 18 February 2022).
- Dimmock, D.; Caylor, S.; Waldman, B.; Benson, W.; Ashburner, C.; Carmichael, J.L.; Carroll, J.; Cham, E.; Chowdhury, S.; Cleary, J.; et al. Project Baby Bear: Rapid precision care incorporating rWGS in 5 California children’s hospitals demonstrates improved clinical outcomes and reduced costs of care. Am. J. Hum. Genet. 2021, 108, 1231–1238. [Google Scholar] [CrossRef] [PubMed]
- Carter, A.B.; Abruzzo, L.V.; Hirschhorn, J.W.; Jones, D.; Jordan, D.C.; Nassiri, M.; Ogino, S.; Patel, N.R.; Suciu, C.G.; Temple-Smolkin, R.L.; et al. Electronic Health Records and Genomics: Perspectives from the Association for Molecular Pathology Electronic Health Record (EHR) Interoperability for Clinical Genomics Data Working Group. J. Mol. Diagn. 2022, 24, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Beckmann, J.S.; Lew, D. Reconciling evidence-based medicine and precision medicine in the era of big data: Challenges and opportunities. Genome Med. 2016, 8, 134. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- RWE Registry Developed From the Real-World Evidence Transparency Initiative: The Professional Society for Health Economics and Outcomes Research (ISPOR). 2021. Available online: https://www.ispor.org/heor-resources/news/2021/10/26/new-real-world-evidence-registry-launches (accessed on 11 December 2021).
- Abul-Husn, N.S.; Manickam, K.; Jones, L.K.; Wright, E.A.; Hartzel, D.N.; Gonzaga-Jauregui, C.; O’Dushlaine, C.; Leader, J.B.; Kirchner, H.L.; Lindbuchler, D.M.; et al. Genetic identification of familial hypercholesterolemia within a single U.S. health care system. Science 2016, 354, aaf7000. [Google Scholar] [CrossRef] [PubMed]
- Altman, A.M.; Marmor, S.; Tuttle, T.M.; Hui, J.Y.C. 21-Gene Recurrence Score Testing in HER2-positive Patients. Clin. Breast Cancer 2019, 19, 126–130. [Google Scholar] [CrossRef]
- Anderson, H.D.; Crooks, K.R.; Kao, D.P.; Aquilante, C.L. The landscape of pharmacogenetic testing in a US managed care population. Genet. Med. 2020, 22, 1247–1253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Armstrong, J.; Toscano, M.; Kotchko, N.; Friedman, S.; Schwartz, M.D.; Virgo, K.S.; Lynch, K.; Andrews, J.E.; Loi, C.X.A.; Bauer, J.E.; et al. Utilization and Outcomes of BRCAGenetic Testing and Counseling in a National Commercially Insured Population. JAMA Oncol. 2015, 1, 1251–1260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bardakjian, T.M.; Helbig, I.; Quinn, C.; Elman, L.B.; McCluskey, L.F.; Scherer, S.S.; Gonzalez-Alegre, P. Genetic test utilization and diagnostic yield in adult patients with neurological disorders. Neurogenetics 2018, 19, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Benitez, J.; Cool, C.L.; Scotti, D.J. Use of combinatorial pharmacogenomic guidance in treating psychiatric disorders. Pers. Med. 2018, 15, 481–494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhutiani, N.; Vuong, B.; Egger, M.E.; Eldredge-Hindy, H.; McMasters, K.M.; Ajkay, N. Evaluating patterns of utilization of gene signature panels and impact on treatment patterns in patients with ductal carcinoma in situ of the breast. Surgery 2019, 166, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Blaes, A.H.; Jewett, P.I.; McKay, K.; Riley, D.; Jatoi, I.; Trentham-Dietz, A.; Chrischilles, E.; Klemp, J.R. Factors associated with genetic testing in a cohort of breast cancer survivors. Breast J. 2019, 25, 1241–1244. [Google Scholar] [CrossRef]
- Blagec, K.; Kuch, W.; Samwald, M. The Importance of Gene-Drug-Drug-Interactions in Pharmacogenomics Decision Support: An Analysis Based on Austrian Claims Data. In Health Informatics Meets eHealth; Schreier, G., Ammenwerth, E., Hörbst, A., Hayn, D., Eds.; Studies in Health Technology and Informatics; IOS Press: Amsterdam, The Netherlands, 2017; Volume 236, pp. 121–127. Available online: https://ebooks.iospress.nl/publication/46468 (accessed on 18 February 2022).
- Byfield, S.D.; Wei, H.; DuCharme, M.; Lancaster, J.M. Economic impact of multigene panel testing for hereditary breast and ovarian cancer. J. Comp. Eff. Res. 2021, 10, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Caplan, E.O.; Wong, W.B.; Ferries, E.; Hulinsky, R.; Brown, V.T.; Bordenave, K.; Suehs, B.T. Novel Approach Using Administrative Claims to Evaluate Trends in Oncology Multigene Panel Testing for Patients Enrolled in Medicare Advantage Health Plans. JCO Precis. Oncol. 2021, 5, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Kolor, K.; Grosse, S.D.; Rodriguez, J.L.; Lynch, J.A.; Green, R.F.; Dotson, W.D.; Bowen, M.S.; Khoury, M.J. Trends in utilization and costs of BRCA testing among women aged 18–64 years in the United States, 2003–2014. Genet. Med. 2018, 20, 428–434. [Google Scholar] [CrossRef] [PubMed]
- Childers, K.K.; Maggard-Gibbons, M.; Macinko, J.; Childers, C.P. National Distribution of Cancer Genetic Testing in the United States. JAMA Oncol. 2018, 4, 876. [Google Scholar] [CrossRef] [PubMed]
- Chitty, L.S.; Wright, D.; Hill, M.; Verhoef, T.I.; Daley, R.; Lewis, C.; Mason, S.; McKay, F.; Jenkins, L.; Howarth, A.; et al. Uptake, outcomes, and costs of implementing non-invasive prenatal testing for Down’s syndrome into NHS maternity care: Prospective cohort study in eight diverse maternity units. BMJ 2016, 354, i3426. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cress, R.D.; Chen, Y.S.; Morris, C.R.; Chew, H.; Kizer, K.W. Underutilization of gene expression profiling for early-stage breast cancer in California. Cancer Causes Control 2016, 27, 721–727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dalal, A.A.; Guerin, A.; Mutebi, A.; Culver, K.W. Economic analysis of BRAF gene mutation testing in real world practice using claims data: Costs of single gene versus panel tests in patients with lung cancer. J. Med. Econ. 2018, 21, 649–655. [Google Scholar] [CrossRef] [PubMed]
- DeFrancesco, M.S.; Waldman, R.N.; Pearlstone, M.M.; Karanik, D.; Bernhisel, R.; Logan, J.; Alico, L.; Adkins, R.T. Hereditary Cancer Risk Assessment and Genetic Testing in the Community-Practice Setting. Obstet. Gynecol. 2018, 132, 1121–1129. [Google Scholar] [CrossRef] [PubMed]
- Desai, S.; Jena, A.B. Do celebrity endorsements matter? Observational study of BRCA gene testing and mastectomy rates after Angelina Jolie’s New York Times editorial. BMJ 2016, 355, i6357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinan, M.A.; Mi, X.; Reed, S.D.; Hirsch, B.R.; Lyman, G.H.; Curtis, L.H. Initial Trends in the Use of the 21-Gene Recurrence Score Assay for Patients With Breast Cancer in the Medicare Population, 2005–2009. JAMA Oncol. 2015, 1, 158–166. [Google Scholar] [CrossRef] [PubMed]
- Dinan, M.A.; Mi, X.; Reed, S.D.; Lyman, G.H.; Curtis, L.H. Association Between Use of the 21-Gene Recurrence Score Assay and Receipt of Chemotherapy Among Medicare Beneficiaries With Early-Stage Breast Cancer, 2005–2009. JAMA Oncol. 2015, 1, 1098–1109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinan, M.A.; Wilson, L.; Reed, S.D. Chemotherapy Costs and 21-Gene Recurrence Score Genomic Testing Among Medicare Beneficiaries With Early-Stage Breast Cancer, 2005 to 2011. J. Natl. Compr. Cancer Netw. 2019, 17, 245–254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinan, M.A.; Wilson, L.E.; Reed, S.D.; Griggs, J.J.; Norton, E.C. Association of 21-Gene Assay (OncotypeDX) Testing and Receipt of Chemotherapy in the Medicare Breast Cancer Patient Population Following Initial Adoption. Clin. Breast Cancer 2020, 20, 487–494.e1. [Google Scholar] [CrossRef] [PubMed]
- Dubrovsky, E.; Raymond, S.; Chun, J.; Fong, A.; Patel, N.; Guth, A.; Schnabel, F. Genomic testing in early stage invasive male breast cancer: An NCDB analysis from 2008 to 2014. Breast J. 2019, 25, 425–433. [Google Scholar] [CrossRef]
- El Rouby, N.; Alrwisan, A.; Langaee, T.; Lipori, G.; Angiolillo, D.J.; Franchi, F.; Riva, A.; Elsey, A.; Johnson, J.A.; Cavallari, L.H.; et al. Clinical Utility of Pharmacogene Panel-Based Testing in Patients Undergoing Percutaneous Coronary Intervention. Clin. Transl. Sci. 2019, 13, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Fohner, A.E.; Ranatunga, D.K.; Thai, K.K.; Lawson, B.L.; Risch, N.; Oni-Orisan, A.; Jelalian, A.T.; Rettie, A.E.; Liu, V.X.; Schaefer, C.A. Assessing the clinical impact of CYP2C9 pharmacogenetic variation on phenytoin prescribing practice and patient response in an integrated health system. Pharmacogenet. Genom. 2019, 29, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Franc, B.L.; Copeland, T.P.; Thombley, R.; Park, M.; Marafino, B.; Dean, M.; Boscardin, W.J.; Rugo, H.S.; Dudley, R.A. Geographic and Patient Characteristics Associated With Election of Prophylactic Mastectomy in Young Breast Cancer Patients With Early Disease. Am. J. Clin. Oncol. 2018, 41, 1037–1042. [Google Scholar] [CrossRef] [PubMed]
- Geddes, G.C.; Basel, D.; Frommelt, P.; Kinney, A.; Earing, M. Genetic Testing Protocol Reduces Costs and Increases Rate of Genetic Diagnosis in Infants with Congenital Heart Disease. Pediatr. Cardiol. 2017, 38, 1465–1470. [Google Scholar] [CrossRef] [Green Version]
- Guo, F.; Bs, M.S.; Fuchs, E.L.; Berenson, A.B.; Kuo, Y. BRCA testing in unaffected young women in the United States, 2006–2017. Cancer 2020, 126, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Scholl, M.; Fuchs, E.L.; Wong, R.; Kuo, Y.-F.; Berenson, A.B. Trends in Positive BRCA Test Results Among Older Women in the United States, 2008–2018. JAMA Netw. Open 2020, 3, e2024358. [Google Scholar] [CrossRef] [PubMed]
- Hefti, E.; Jacobs, D.M.; Rana, K.; Blanco, J.G. Analysis of outpatient HER2 testing in New York state using the statewide planning and research cooperative system. Pharmacogenomics 2018, 19, 1395–1401. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Kamath, P.; Schlumbrecht, M.; Miao, F.; Driscoll, D.; Oldak, S.; Slomovitz, B.; Koru-Sengul, T.; George, S. Identifying disparities in germline and somatic testing for ovarian cancer. Gynecol. Oncol. 2019, 153, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Hui, L.; Hutchinson, B.; Poulton, A.; Halliday, J. Population-based impact of noninvasive prenatal screening on screening and diagnostic testing for fetal aneuploidy. Genet. Med. 2017, 19, 1338–1345. [Google Scholar] [CrossRef] [PubMed]
- Hull, L.E.; Lynch, J.A.; Berse, B.B.; DuVall, S.L.; Chun, D.S.; Venne, V.L.; Efimova, O.V.; Icardi, M.S.; Kelley, M.J. Clinical Impact of 21-Gene Recurrence Score Test Within the Veterans Health Administration: Utilization and Receipt of Guideline-Concordant Care. Clin. Breast Cancer 2017, 18, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Katz, S.J.; Bondarenko, I.; Ward, K.C.; Hamilton, A.S.; Morrow, M.; Kurian, A.W.; Hofer, T.P. Association of Attending Surgeon With Variation in the Receipt of Genetic Testing After Diagnosis of Breast Cancer. JAMA Surg. 2018, 153, 909. [Google Scholar] [CrossRef] [PubMed]
- Nzale, S.K.; Weeks, W.B.; Ouafik, L.; Rouquette, I.; Beau-Faller, M.; Lemoine, A.; Bringuier, P.-P.; Soriano, A.-G.L.C.; Barlesi, F.; Ventelou, B. Inequity in access to personalized medicine in France: Evidences from analysis of geo variations in the access to molecular profiling among advanced non-small-cell lung cancer patients: Results from the IFCT Biomarkers France Study. PLoS ONE 2020, 15, e0234387. [Google Scholar] [CrossRef]
- Kolor, K.; Chen, Z.; Grosse, S.D.; Rodriguez, J.L.; Green, R.F.; Dotson, W.D.; Bowen, M.S.; Lynch, J.A.; Khoury, M.J. BRCA Genetic Testing and Receipt of Preventive Interventions among Women Aged 18–64 Years with Employer-Sponsored Health Insurance in Nonmetropolitan and Metropolitan Areas-United States, 2009–2014, 2017. Centers for Diseases Control and Prevention Website. Available online: https://www.cdc.gov/mmwr/volumes/66/ss/ss6615a1.htm (accessed on 18 February 2022).
- Larson, K.L.; Huang, B.; Chen, Q.; Tucker, T.; Schuh, M.; Arnold, S.M.; Kolesar, J.M. EGFR testing and erlotinib use in non-small cell lung cancer patients in Kentucky. PLoS ONE 2020, 15, e0237790. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Fan, J.; Vitali, F.; Berghout, J.; Aberasturi, D.; Li, J.; Wilson, L.; Chiu, W.; Pumarejo, M.; Han, J.; et al. Novel disease syndromes unveiled by integrative multiscale network analysis of diseases sharing molecular effectors and comorbidities. BMC Med. Genom. 2018, 11, 112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liede, A.; Cai, M.; Crouter, T.F.; Niepel, D.; Callaghan, F.; Evans, D.G. Risk-reducing mastectomy rates in the US: A closer examination of the Angelina Jolie effect. Breast Cancer Res. Treat. 2018, 171, 435–442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lynch, J.A.; Berse, B.; Coomer, N.; Kautter, J. 21-Gene recurrence score testing among Medicare beneficiaries with breast cancer in 2010–2013. Genet. Med. 2017, 19, 1134–1143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lynch, J.A.; Berse, B.; Dotson, W.D.; Khoury, M.J.; Coomer, N.; Kautter, J. Utilization of genetic tests: Analysis of gene-specific billing in Medicare claims data. Genet. Med. 2017, 19, 890–899. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, S.J.; Lin, C.C.; Todd, P.K.; Burke, J.F.; Callaghan, B.C. Genetic testing utilization for patients with neurologic disease and the limitations of claims data. Neurol. Genet. 2020, 6, e405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mathias, P.C.; Hendrix, N.; Wang, W.-J.; Keyloun, K.; Khelifi, M.; Tarczy-Hornoch, P.; Devine, B. Characterizing Pharmacogenomic-Guided Medication Use With a Clinical Data Repository. Clin. Pharmacol. Ther. 2017, 102, 340–348. [Google Scholar] [CrossRef] [PubMed]
- McCuaig, J.M.; Care, M.; Ferguson, S.E.; Kim, R.H.; Stockley, T.L.; Metcalfe, K.A. Year 1: Experiences of a tertiary cancer centre following implementation of reflex BRCA1 and BRCA2 tumor testing for all high-grade serous ovarian cancers in a universal healthcare system. Gynecol. Oncol. 2020, 158, 747–753. [Google Scholar] [CrossRef]
- Mittmann, N.; Earle, C.C.; Cheng, S.Y.; Julian, J.A.; Rahman, F.; Seung, S.J.; Levine, M.N. Population-Based Study to Determine the Health System Costs of Using the 21-Gene Assay. J. Clin. Oncol. 2018, 36, 238–243. [Google Scholar] [CrossRef]
- Muller, C.; Lee, S.M.; Barge, W.; Siddique, S.M.; Berera, S.; Wideroff, G.; Tondon, R.; Chang, J.; Peterson, M.; Stoll, J.; et al. Low Referral Rate for Genetic Testing in Racially and Ethnically Diverse Patients Despite Universal Colorectal Cancer Screening. Clin. Gastroenterol. Hepatol. 2018, 16, 1911–1918. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, S.C.; Isaacs, C.; Chao, C.; Tsai, H.-T.; Liu, C.; Ekezue, B.F.; Selvam, N.; Kessler, L.G.; Schwartz, M.D.; Lobo, T.; et al. Adoption of Gene Expression Profiling for Breast Cancer in US Oncology Practice for Women Younger Than 65 Years. J. Natl. Compr. Cancer Netw. 2015, 13, 1216–1224. [Google Scholar] [CrossRef] [PubMed]
- O’neill, S.C.; Isaacs, C.; Lynce, F.; Graham, D.M.A.; Chao, C.; Sheppard, V.B.; Zhou, Y.; Liu, C.; Selvam, N.; Schwartz, M.D.; et al. Endocrine therapy initiation, discontinuation and adherence and breast imaging among 21-gene recurrence score assay-eligible women under age 65. Breast Cancer Res. 2017, 19, 45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Orucevic, A.; Heidel, R.E.; Bell, J.L. Utilization and impact of 21-gene recurrence score assay for breast cancer in clinical practice across the United States: Lessons learned from the 2010 to 2012 National Cancer Data Base analysis. Breast Cancer Res. Treat. 2016, 157, 427–435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pace, L.E.; Baum, C.F.; Horvath, K.; Raja, S.; Cohen, J.; Hawkins, S.S. BRCA1/2 Testing in Massachusetts Among Women With Private Insurance or Medicaid, 2011–2015. Med. Care 2020, 58, 963–967. [Google Scholar] [CrossRef] [PubMed]
- Palazzo, L.L.; Sheehan, D.F.; Tramontano, A.C.; Kong, C.Y. Disparities and Trends in Genetic Testing and Erlotinib Treatment among Metastatic Non–Small Cell Lung Cancer Patients. Cancer Epidemiol. Biomarkers Prev. 2019, 28, 926–934. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parkhurst, E.; Calonico, E.; Abboy, S. Utilization of Genetic Testing for RET Mutations in Patients with Medullary Thyroid Carcinoma: A Single-Center Experience. J. Genet. Couns. 2018, 27, 1411–1416. [Google Scholar] [CrossRef] [PubMed]
- Pavey, A.R.; Bodian, D.L.; Vilboux, T.; Khromykh, A.; Hauser, N.S.; Huddleston, K.; Klein, E.; Black, A.; Kane, M.S.; Iyer, R.K.; et al. Utilization of genomic sequencing for population screening of immunodeficiencies in the newborn. Genet. Med. 2017, 19, 1367–1375. [Google Scholar] [CrossRef] [PubMed]
- Perlis, R.H.; Mehta, R.; Edwards, A.M.; Tiwari, A.; Imbens, G.W. Pharmacogenetic testing among patients with mood and anxiety disorders is associated with decreased utilization and cost: A propensity-score matched study. Depress. Anxiety 2018, 35, 946–952. [Google Scholar] [CrossRef] [PubMed]
- Petelin, L.; James, P.A.; Trainer, A.H. Changing landscape of hereditary breast and ovarian cancer germline genetic testing in Australia. Intern. Med. J. 2018, 48, 1269–1272. [Google Scholar] [CrossRef] [PubMed]
- Pokharel, H.P.; Hacker, N.F.; Andrews, L. Changing patterns of referrals and outcomes of genetic participation in gynaecological-oncology multidisciplinary care. Aust. N. Z. J. Obstet. Gynaecol. 2016, 56, 633–638. [Google Scholar] [CrossRef]
- Potosky, A.L.; O’Neill, S.C.; Isaacs, C.; Tsai, H.-T.; Chao, C.; Liu, C.; Ekezue, B.F.; Selvam, N.; Kessler, L.G.; Zhou, Y.; et al. Population-based study of the effect of gene expression profiling on adjuvant chemotherapy use in breast cancer patients under the age of 65 years. Cancer 2015, 121, 4062–4070. [Google Scholar] [CrossRef] [Green Version]
- Poulton, A.; Lewis, S.; Hui, L.; Halliday, J.L. Prenatal and preimplantation genetic diagnosis for single gene disorders: A population-based study from 1977 to 2016. Prenat. Diagn. 2018, 38, 904–910. [Google Scholar] [CrossRef]
- Ray, G.T.; Mandelblatt, J.; Habel, L.A.; Ramsey, S.; Kushi, L.H.; Li, Y.; Lieu, T.A. Breast cancer multigene testing trends and impact on chemotherapy use. Am. J. Manag. Care 2016, 22, e153. [Google Scholar]
- Ritter, A.; Bedoukian, E.; Berger, J.H.; Copenheaver, D.; Gray, C.; Krantz, I.; Izumi, K.; Juusola, J.; Leonard, J.; Lin, K.; et al. Clinical utility of exome sequencing in infantile heart failure. Genet. Med. 2019, 22, 423–426. [Google Scholar] [CrossRef]
- Roberts, M.C.; Dusetzina, S.B. Use and Costs for Tumor Gene Expression Profiling Panels in the Management of Breast Cancer From 2006 to 2012: Implications for Genomic Test Adoption Among Private Payers. J. Oncol. Pract. 2015, 11, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.C.; Dusetzina, S. The effect of a celebrity health disclosure on demand for health care: Trends in BRCA testing and subsequent health services use. J. Community Genet. 2017, 8, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.C.; Weinberger, M.; Dusetzina, S.; Dinan, M.A.; Reeder-Hayes, K.E.; Carey, L.A.; Troester, M.A.; Wheeler, S.B. Racial Variation in the Uptake of Oncotype DX Testing for Early-Stage Breast Cancer. J. Clin. Oncol. 2016, 34, 130–138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schoen, C.; Santolaya-Forgas, J.; Genc, M.; Ashkinadze, E. Differential utilization of expanded genetic screening tests in patients of reproductive ages from private and academic practices. J. Périnat. Med. 2015, 43, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, T.; Handorf, E.A.; Meyer, J.E.; Hall, M.J.; Esnaola, N.F. Mismatch Repair Deficiency Testing in Patients With Colorectal Cancer and Nonadherence to Testing Guidelines in Young Adults. JAMA Oncol. 2018, 4, e173580. [Google Scholar] [CrossRef] [PubMed]
- Singer, J.; Hanna, J.W.; Visaria, J.; Gu, T.; McCoy, M.; Kloos, R.T. Impact of a gene expression classifier on the long-term management of patients with cytologically indeterminate thyroid nodules. Curr. Med. Res. Opin. 2016, 32, 1225–1232. [Google Scholar] [CrossRef] [PubMed]
- Sivapiragasam, A.; Kumar, P.A.; Sokol, E.S.; Albacker, L.A.; Killian, J.K.; Ramkissoon, S.H.; Huang, R.S.P.; Severson, E.A.; Brown, C.A.; Danziger, N.; et al. Predictive Biomarkers for Immune Checkpoint Inhibitors in Metastatic Breast Cancer. Cancer Med. 2021, 10, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Stein, Q.P.; Vockley, C.W.; Edick, M.J.; Zhai, S.; Hiner, S.J.; Loman, R.S.; Davis-Keppen, L.; Zuck, T.A.; Cameron, C.A.; Berry, S.A.; et al. An Exploration of Genetic Test Utilization, Genetic Counseling, and Consanguinity within the Inborn Errors of Metabolism Collaborative (IBEMC). J. Genet. Couns. 2017, 26, 1238–1243. [Google Scholar] [CrossRef] [PubMed]
- Tanner, J.-A.; Brown, L.C.; Yu, K.; Li, J.; Dechairo, B.M. Canadian Medication Cost Savings Associated with Combinatorial Pharmacogenomic Guidance for Psychiatric Medications. Clin. Outcomes Res. 2019, ume 11, 779–787. [Google Scholar] [CrossRef] [Green Version]
- Tiller, G.E.; Kershberg, H.B.; Goff, J.; Coffeen, C.; Liao, W.; Sehnert, A.J. Women’s views and the impact of noninvasive prenatal testing on procedures in a managed care setting. Prenat. Diagn. 2015, 35, 428–433. [Google Scholar] [CrossRef] [PubMed]
- Vaeth, S.; Christensen, R.; Dunø, M.; Lildballe, D.L.; Thorsen, K.; Vissing, J.; Svenstrup, K.; Hertz, J.M.; Andersen, H.; Jensen, U.B. Genetic analysis of Charcot-Marie-Tooth disease in Denmark and the implementation of a next generation sequencing platform. Eur. J. Med Genet. 2018, 62, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Real-World Data Type * | Integrated Dataset | ||||
|---|---|---|---|---|---|
| Administrative Claims (Commercial) | Administrative Claims (Public) | Clinical Databases | Lab Databases | Registries | Used Integrated Dataset for Analyses |
| 33% 24/72 | 25% 18/72 | 29% 21/72 | 31% 22/72 | 55% 40/72 | 38% 27/72 |
| Test | Percentage (n) * | Test Examples |
|---|---|---|
| BRCA1 and BRCA2 | 16% (14) | BRCA1 and BRCA2 |
| GEP/Oncotype Dx® | 22% (19) | Oncotype Dx® |
| Multi-gene panel | 10% (9) | Hereditary Cancer Genetic Testing, Myriad MyRisk® Panel |
| Tumor sequencing | 2% (2) | FoundationOne® |
| WES or WGS | 2% (2) | Whole Exome or Genome Sequencing |
| Other | 48% (42) | GeneSight® Psychotropic Test, Afirma® CYP450, PGx (undefined) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Douglas, M.P.; Kumar, A. Analyzing Precision Medicine Utilization with Real-World Data: A Scoping Review. J. Pers. Med. 2022, 12, 557. https://doi.org/10.3390/jpm12040557
Douglas MP, Kumar A. Analyzing Precision Medicine Utilization with Real-World Data: A Scoping Review. Journal of Personalized Medicine. 2022; 12(4):557. https://doi.org/10.3390/jpm12040557
Chicago/Turabian StyleDouglas, Michael P., and Anika Kumar. 2022. "Analyzing Precision Medicine Utilization with Real-World Data: A Scoping Review" Journal of Personalized Medicine 12, no. 4: 557. https://doi.org/10.3390/jpm12040557
APA StyleDouglas, M. P., & Kumar, A. (2022). Analyzing Precision Medicine Utilization with Real-World Data: A Scoping Review. Journal of Personalized Medicine, 12(4), 557. https://doi.org/10.3390/jpm12040557







