Significance of Immune Status of SARS-CoV-2 Infected Patients in Determining the Efficacy of Therapeutic Interventions
Abstract
1. Introduction
2. Therapeutic Interventions
2.1. Pharmaceutical Interventions
2.2. Therapeutic Models for Pulmonary Phase
2.3. Anti-Inflammatory Therapeutics
3. Inflammatory Immunopathogenesis
4. Immune-Boosting Strategies
4.1. Interferon Mediated Interventions
4.2. Restoring Distressed T-Cells
5. Tracking Immune Status
6. Future Perspectives
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Guan, J.; Wei, X.; Qin, S.; Liu, X.; Jiang, Y.; Chen, Y.; Chen, Y.; Lu, H.; Qian, J.; Wang, Z.; et al. Continuous tracking of COVID-19 patients’ immune status. Int. Immunopharmacol. 2020, 89, 107034. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Mehta, O.P.; Bhandari, P.; Raut, A.; Kacimi, S.E.O.; Huy, N.T. Coronavirus Disease (COVID-19): Comprehensive Review of Clinical Presentation. Front. Public Health 2020, 8, 582932. [Google Scholar] [CrossRef] [PubMed]
- Brodin, P. Immune determinants of COVID-19 disease presentation and severity. Nat. Med. 2021, 27, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [PubMed]
- Moghadas, S.M.; Fitzpatrick, M.C.; Sah, P.; Pandey, A.; Shoukat, A.; Singer, B.H.; Galvani, A.P. The implications of silent transmission for the control of COVID-19 outbreaks. Proc. Natl. Acad. Sci. USA 2020, 117, 17513–17515. [Google Scholar] [CrossRef]
- Daher, A.; Balfanz, P.; Aetou, M.; Hartmann, B.; Müller-Wieland, D.; Müller, T.; Marx, N.; Dreher, M.; Cornelissen, C.G. Clinical course of COVID-19 patients needing supplemental oxygen outside the intensive care unit. Sci. Rep. 2021, 11, 2256. [Google Scholar] [CrossRef] [PubMed]
- Llitjos, J.-F.; Bredin, S.; Lascarrou, J.-B.; Soumagne, T.; Cojocaru, M.; Leclerc, M.; Lepetit, A.; Gouhier, A.; Charpentier, J.; Piton, G.; et al. Increased susceptibility to intensive care unit-acquired pneumonia in severe COVID-19 patients: A multicentre retrospective cohort study. Ann. Intensive Care 2021, 11, 20. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.-Y.; Shinde, S.K.; Lone, S.; Palem, R.R.; Ghodake, G.S. COVID-19 Pandemic: Public Health Risk Assessment and Risk Mitigation Strategies. J. Pers. Med. 2021, 11, 1243. [Google Scholar] [CrossRef]
- Hadjadj, J.; Yatim, N.; Barnabei, L.; Corneau, A.; Boussier, J.; Smith, N.; Péré, H.; Charbit, B.; Bondet, V.; Chenevier-Gobeaux, C.; et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 2020, 369, 718–724. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Olbei, M.; Hautefort, I.; Modos, D.; Treveil, A.; Poletti, M.; Gul, L.; Shannon-Lowe, C.D.; Korcsmaros, T. SARS-CoV-2 Causes a Different Cytokine Response Compared to Other Cytokine Storm-Causing Respiratory Viruses in Severely Ill Patients. Front. Immunol. 2021, 12, 629193. [Google Scholar] [CrossRef] [PubMed]
- Jafarzadeh, A.; Jafarzadeh, S.; Nozari, P.; Mokhtari, P.; Nemati, M. Lymphopenia an important immunological abnormality in patients with COVID-19: Possible mechanisms. Scand. J. Immunol. 2021, 93, e12967. [Google Scholar] [CrossRef] [PubMed]
- Winkler, E.S.; Bailey, A.L.; Kafai, N.M.; Nair, S.; McCune, B.T.; Yu, J.; Fox, J.M.; Chen, R.E.; Earnest, J.T.; Keeler, S.P.; et al. SARS-CoV-2 infection of human ACE2-transgenic mice causes severe lung inflammation and impaired function. Nat. Immunol. 2020, 21, 1327–1335. [Google Scholar] [CrossRef] [PubMed]
- Maccio, U.; Zinkernagel, A.S.; Shambat, S.M.; Zeng, X.; Cathomas, G.; Ruschitzka, F.; Schuepbach, R.A.; Moch, H.; Varga, Z. SARS-CoV-2 leads to a small vessel endotheliitis in the heart. EBioMedicine 2021, 63, 103182. [Google Scholar] [CrossRef] [PubMed]
- Bailey, A.L.; Dmytrenko, O.; Greenberg, L.; Bredemeyer, A.L.; Ma, P.; Liu, J.; Penna, V.; Winkler, E.S.; Sviben, S.; Brooks, E.; et al. SARS-CoV-2 Infects Human Engineered Heart Tissues and Models COVID-19 Myocarditis. JACC Basic Transl. Sci. 2021, 6, 331–345. [Google Scholar] [CrossRef]
- Taoufik, Y.; de Goër de Herve, M.-G.; Corgnac, S.; Durrbach, A.; Mami-Chouaib, F. When Immunity Kills: The Lessons of SARS-CoV-2 Outbreak. Front. Immunol. 2021, 12, 3833. [Google Scholar] [CrossRef]
- Diao, B.; Wang, C.; Wang, R.; Feng, Z.; Tan, Y.; Wang, H.; Wang, C.; Liu, L.; Liu, Y.; Liu, Y.; et al. Human Kidney is a Target for Novel Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection. medRxiv 2020. [Google Scholar] [CrossRef]
- Ghodake, G.S.; Shinde, S.K.; Kadam, A.A.; Saratale, R.G.; Saratale, G.D.; Syed, A.; Elgorban, A.M.; Marraiki, N.; Kim, D.-Y. Biological characteristics and biomarkers of novel SARS-CoV-2 facilitated rapid development and implementation of diagnostic tools and surveillance measures. Biosens. Bioelectron. 2021, 177, 112969. [Google Scholar] [CrossRef]
- Gelaye, B.; Foster, S.; Bhasin, M.; Tawakol, A.; Fricchione, G. SARS-CoV-2 morbidity and mortality in racial/ethnic minority populations: A window into the stress related inflammatory basis of health disparities? Brain Behav. Immun. Health 2020, 9, 100158. [Google Scholar] [CrossRef] [PubMed]
- Xiao, N.; Nie, M.; Pang, H.; Wang, B.; Hu, J.; Meng, X.; Li, K.; Ran, X.; Long, Q.; Deng, H.; et al. Integrated cytokine and metabolite analysis reveals immunometabolic reprogramming in COVID-19 patients with therapeutic implications. Nat. Commun. 2021, 12, 1618. [Google Scholar] [CrossRef] [PubMed]
- Panovska-Stavridis, I.; Ridova, N.; Stojanoska, T.; Demiri, I.; Stevanovic, M.; Stojanovska, S.; Ristevska, T.; Dimkovski, A.; Filipce, V.; Dimovski, A.; et al. Insight in the Current Progress in the Largest Clinical Trials for COVID-19 Drug Management (As of January 2021). Prilozi 2021, 42, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Dhupkar, P.; Mukherjee, S. Ethical dimensions in randomized trials and off-label use of investigational drugs for COVID-19 treatment. Clin. Ethics 2022, 17, 95–104. [Google Scholar] [CrossRef]
- Mulangu, S.; Dodd, L.E.; Davey, R.T.; Tshiani Mbaya, O.; Proschan, M.; Mukadi, D.; Lusakibanza Manzo, M.; Nzolo, D.; Tshomba Oloma, A.; Ibanda, A.; et al. A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics. N. Engl. J. Med. 2019, 381, 2293–2303. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of COVID-19—Final Report. N. Engl. J. Med. 2020, 383, 1813–1826. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Wang, X. COVID-19: A new challenge for human beings. Cell. Mol. Immunol. 2020, 17, 555–557. [Google Scholar] [CrossRef]
- Arora, G.; Shrivastava, R.; Kumar, P.; Bandichhor, R.; Krishnamurthy, D.; Sharma, R.K.; Matharu, A.S.; Pandey, J.; Rizwan, M. Recent advances made in the synthesis of small drug molecules for clinical applications: An insight. Curr. Res. Green Sustain. Chem. 2021, 4, 100097. [Google Scholar] [CrossRef]
- Prince, T.; Donovan-Banfield, I.a.; Goldswain, H.; Penrice-Randal, R.; Turtle, L.; Fletcher, T.; Khoo, S.; Hiscox, J.A. Antiviral activity of molnupiravir precursor NHC against SARS-CoV-2 Variants of Concern (VOCs) and its therapeutic window in a human lung cell model. bioRxiv 2021. [Google Scholar] [CrossRef]
- Dexamethasone in Hospitalized Patients with COVID-19. N. Engl. J. Med. 2020, 384, 693–704. [CrossRef]
- Ison, M.G.; Wolfe, C.; Boucher, H.W. Emergency Use Authorization of Remdesivir: The Need for a Transparent Distribution Process. JAMA 2020, 323, 2365–2366. [Google Scholar] [CrossRef] [PubMed]
- Gendrot, M.; Javelle, E.; Clerc, A.; Savini, H.; Pradines, B. Chloroquine as a prophylactic agent against COVID-19? Int. J. Antimicrob. Agents 2020, 55, 105980. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Singh, A.; Shaikh, A.; Singh, R.; Misra, A. Chloroquine and hydroxychloroquine in the treatment of COVID-19 with or without diabetes: A systematic search and a narrative review with a special reference to India and other developing countries. Diabetes Metab Syndr. 2020, 14, 241–246. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Cai, S.; Li, Y.; Li, Y.; Fan, Y.; Li, L.; Lei, C.; Tang, X.; Hu, F.; Li, F.; et al. Prognostic Factors for COVID-19 Pneumonia Progression to Severe Symptoms Based on Earlier Clinical Features: A Retrospective Analysis. Front. Med. 2020, 7, 643. [Google Scholar] [CrossRef]
- Horby, P.W.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Emberson, J.; Palfreeman, A.; Raw, J.; Elmahi, E.; Prudon, B.; et al. Lopinavir–ritonavir in patients admitted to hospital with COVID-19 (RECOVERY): A randomised, controlled, open-label, platform trial. Lancet 2020, 396, 1345–1352. [Google Scholar] [CrossRef]
- Fischer, W.; Eron, J.J.; Holman, W.; Cohen, M.S.; Fang, L.; Szewczyk, L.J.; Sheahan, T.P.; Baric, R.; Mollan, K.R.; Wolfe, C.R.; et al. Molnupiravir, an Oral Antiviral Treatment for COVID-19. medRxiv 2021. [Google Scholar] [CrossRef]
- Robinson, P.C.; Morand, E. Divergent effects of acute versus chronic glucocorticoids in COVID-19. Lancet Rheumatol. 2021, 3, e168–e170. [Google Scholar] [CrossRef]
- Zhuravel, S.V.; Khmelnitskiy, O.K.; Burlaka, O.O.; Gritsan, A.I.; Goloshchekin, B.M.; Kim, S.; Hong, K.Y. Nafamostat in hospitalized patients with moderate to severe COVID-19 pneumonia: A randomised Phase II clinical trial. eClinicalMedicine 2021, 41, 101169. [Google Scholar] [CrossRef]
- Axfors, C.; Schmitt, A.M.; Janiaud, P.; van’t Hooft, J.; Abd-Elsalam, S.; Abdo, E.F.; Abella, B.S.; Akram, J.; Amaravadi, R.K.; Angus, D.C.; et al. Mortality outcomes with hydroxychloroquine and chloroquine in COVID-19 from an international collaborative meta-analysis of randomized trials. Nat. Commun. 2021, 12, 2349. [Google Scholar] [CrossRef]
- Khalil, A.; Kamar, A.; Nemer, G. Thalidomide-Revisited: Are COVID-19 Patients Going to Be the Latest Victims of Yet Another Theoretical Drug-Repurposing? Front. Immunol. 2020, 11, 1248. [Google Scholar] [CrossRef]
- Loffredo, M.; Lucero, H.; Chen, D.-Y.; O’Connell, A.; Bergqvist, S.; Munawar, A.; Bandara, A.; De Graef, S.; Weeks, S.D.; Douam, F.; et al. The in-vitro effect of famotidine on SARS-CoV-2 proteases and virus replication. Sci. Rep. 2021, 11, 5433. [Google Scholar] [CrossRef]
- Wang, Q.; Guo, H.; Li, Y.; Jian, X.; Hou, X.; Zhong, N.; Fei, J.; Su, D.; Bian, Z.; Zhang, Y.; et al. Efficacy and Safety of Leflunomide for Refractory COVID-19: A Pilot Study. Front. Pharmacol. 2021, 12, 1217. [Google Scholar] [CrossRef] [PubMed]
- Durán-Méndez, A.; Aguilar-Arroyo, A.D.; Vivanco-Gómez, E.; Nieto-Ortega, E.; Pérez-Ortega, D.; Jiménez-Pérez, C.; Hernández-Skewes, K.Y.; Montiel-Bravo, G.; Roque-Reyes, O.J.; Romero-Lechuga, F.; et al. Tocilizumab reduces COVID-19 mortality and pathology in a dose and timing-dependent fashion: A multi-centric study. Sci. Rep. 2021, 11, 19728. [Google Scholar] [CrossRef] [PubMed]
- Kyriazopoulou, E.; Poulakou, G.; Milionis, H.; Metallidis, S.; Adamis, G.; Tsiakos, K.; Fragkou, A.; Rapti, A.; Damoulari, C.; Fantoni, M.; et al. Early treatment of COVID-19 with anakinra guided by soluble urokinase plasminogen receptor plasma levels: A double-blind, randomized controlled phase 3 trial. Nat. Med. 2021, 27, 1752–1760. [Google Scholar] [CrossRef]
- Pang, J.; Xu, F.; Aondio, G.; Li, Y.; Fumagalli, A.; Lu, M.; Valmadre, G.; Wei, J.; Bian, Y.; Canesi, M.; et al. Efficacy and tolerability of bevacizumab in patients with severe COVID-19. Nat. Commun. 2021, 12, 814. [Google Scholar] [CrossRef] [PubMed]
- Vankadari, N. Arbidol: A potential antiviral drug for the treatment of SARS-CoV-2 by blocking trimerization of the spike glycoprotein. Int. J. Antimicrob. Agents 2020, 56, 105998. [Google Scholar] [CrossRef] [PubMed]
- Marconi, V.C.; Ramanan, A.V.; de Bono, S.; Kartman, C.E.; Krishnan, V.; Liao, R.; Piruzeli, M.L.B.; Goldman, J.D.; Alatorre-Alexander, J.; de Cassia Pellegrini, R.; et al. Efficacy and safety of baricitinib for the treatment of hospitalised adults with COVID-19 (COV-BARRIER): A randomised, double-blind, parallel-group, placebo-controlled phase 3 trial. Lancet Respir. Med. 2021, 9, 1407–1418. [Google Scholar] [CrossRef]
- Nojomi, M.; Yassin, Z.; Keyvani, H.; Makiani, M.J.; Roham, M.; Laali, A.; Dehghan, N.; Navaei, M.; Ranjbar, M. Effect of Arbidol (Umifenovir) on COVID-19: A randomized controlled trial. BMC Infect. Dis. 2020, 20, 954. [Google Scholar] [CrossRef]
- Blum, V.F.; Cimerman, S.; Hunter, J.R.; Tierno, P.; Lacerda, A.; Soeiro, A.; Cardoso, F.; Bellei, N.C.; Maricato, J.; Mantovani, N.; et al. Nitazoxanide superiority to placebo to treat moderate COVID-19—A Pilot prove of concept randomized double-blind clinical trial. eClinicalMedicine 2021, 37, 100981. [Google Scholar] [CrossRef]
- Saleh, M.; Vaezi, A.A.; Aliannejad, R.; Sohrabpour, A.A.; Kiaei, S.Z.F.; Shadnoush, M.; Siavashi, V.; Aghaghazvini, L.; Khoundabi, B.; Abdoli, S.; et al. Cell therapy in patients with COVID-19 using Wharton’s jelly mesenchymal stem cells: A phase 1 clinical trial. Stem Cell Res. Ther. 2021, 12, 410. [Google Scholar] [CrossRef]
- Zanirati, G.; Provenzi, L.; Libermann, L.L.; Bizotto, S.C.; Ghilardi, I.M.; Marinowic, D.R.; Shetty, A.K.; Da Costa, J.C. Stem cell-based therapy for COVID-19 and ARDS: A systematic review. NPJ Regen. Med. 2021, 6, 73. [Google Scholar] [CrossRef]
- Bégin, P.; Callum, J.; Jamula, E.; Cook, R.; Heddle, N.M.; Tinmouth, A.; Zeller, M.P.; Beaudoin-Bussières, G.; Amorim, L.; Bazin, R.; et al. Convalescent plasma for hospitalized patients with COVID-19: An open-label, randomized controlled trial. Nat. Med. 2021, 27, 2012–2024. [Google Scholar] [CrossRef] [PubMed]
- Taylor, P.C.; Adams, A.C.; Hufford, M.M.; de la Torre, I.; Winthrop, K.; Gottlieb, R.L. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat. Rev. Immunol. 2021, 21, 382–393. [Google Scholar] [CrossRef] [PubMed]
- Yamin, R.; Jones, A.T.; Hoffmann, H.-H.; Schäfer, A.; Kao, K.S.; Francis, R.L.; Sheahan, T.P.; Baric, R.S.; Rice, C.M.; Ravetch, J.V.; et al. Fc-engineered antibody therapeutics with improved anti-SARS-CoV-2 efficacy. Nature 2021, 599, 465–470. [Google Scholar] [CrossRef] [PubMed]
- Mir, T.; Almas, T.; Kaur, J.; Faisaluddin, M.; Song, D.; Ullah, W.; Mamtani, S.; Rauf, H.; Yadav, S.; Latchana, S.; et al. Coronavirus disease 2019 (COVID-19): Multisystem review of pathophysiology. Ann. Med. Surg. 2021, 69, 102745. [Google Scholar] [CrossRef] [PubMed]
- Aleksova, A.; Ferro, F.; Gagno, G.; Cappelletto, C.; Santon, D.; Rossi, M.; Ippolito, G.; Zumla, A.; Beltrami, A.P.; Sinagra, G. COVID-19 and renin-angiotensin system inhibition: Role of angiotensin converting enzyme 2 (ACE2)—Is there any scientific evidence for controversy? J. Intern. Med. 2020, 288, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.J.; Edin, M.L.; Zeldin, D.C.; Li, C.; Wang, D.W.; Chen, C. Good or bad: Application of RAAS inhibitors in COVID-19 patients with cardiovascular comorbidities. Pharmacol. Ther. 2020, 215, 107628. [Google Scholar] [CrossRef] [PubMed]
- Rico-Mesa, J.S.; White, A.; Anderson, A.S. Outcomes in Patients with COVID-19 Infection Taking ACEI/ARB. Curr. Cardiol. Rep. 2020, 22, 31. [Google Scholar] [CrossRef]
- Vaduganathan, M.; Vardeny, O.; Michel, T.; McMurray, J.J.V.; Pfeffer, M.A.; Solomon, S.D. Renin–Angiotensin–Aldosterone System Inhibitors in Patients with COVID-19. N. Engl. J. Med. 2020, 382, 1653–1659. [Google Scholar] [CrossRef]
- Dean, A.Q.; Bozza, W.P.; Twomey, J.D.; Luo, S.; Nalli, A.; Zhang, B. The fight against COVID-19: Striking a balance in the renin–angiotensin system. Drug Discov. Today 2021, 26, 2214–2220. [Google Scholar] [CrossRef]
- Tang, T.; Bidon, M.; Jaimes, J.A.; Whittaker, G.R.; Daniel, S. Coronavirus membrane fusion mechanism offers a potential target for antiviral development. Antivir. Res. 2020, 178, 104792. [Google Scholar] [CrossRef]
- Verdecchia, P.; Cavallini, C.; Spanevello, A.; Angeli, F. The pivotal link between ACE2 deficiency and SARS-CoV-2 infection. Eur. J. Intern. Med. 2020, 76, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, T.; Hoshizaki, M.; Minato, T.; Nirasawa, S.; Asaka, M.N.; Niiyama, M.; Imai, M.; Uda, A.; Chan, J.F.-W.; Takahashi, S.; et al. ACE2-like carboxypeptidase B38-CAP protects from SARS-CoV-2-induced lung injury. Nat. Commun. 2021, 12, 6791. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Choi, W.J. Overview of COVID-19 inflammatory pathogenesis from the therapeutic perspective. Arch. Pharmacal. Res. 2021, 44, 99–116. [Google Scholar] [CrossRef]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e278. [Google Scholar] [CrossRef]
- Yamamoto, M.; Matsuyama, S.; Li, X.; Takeda, M.; Kawaguchi, Y.; Inoue, J.-i.; Matsuda, Z. Identification of Nafamostat as a Potent Inhibitor of Middle East Respiratory Syndrome Coronavirus S Protein-Mediated Membrane Fusion Using the Split-Protein-Based Cell-Cell Fusion Assay. Antimicrob. Agents Chemother. 2016, 60, 6532–6539. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Penninger, J.M.; Li, Y.; Zhong, N.; Slutsky, A.S. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: Molecular mechanisms and potential therapeutic target. Intensive Care Med. 2020, 46, 586–590. [Google Scholar] [CrossRef]
- Geng, J.; Chen, L.; Yuan, Y.; Wang, K.; Wang, Y.; Qin, C.; Wu, G.; Chen, R.; Zhang, Z.; Wei, D.; et al. CD147 antibody specifically and effectively inhibits infection and cytokine storm of SARS-CoV-2 and its variants delta, alpha, beta, and gamma. Signal Transduct. Target. Ther. 2021, 6, 347. [Google Scholar] [CrossRef]
- Bian, H.; Zheng, Z.-H.; Wei, D.; Wen, A.; Zhang, Z.; Lian, J.-Q.; Kang, W.-Z.; Hao, C.-Q.; Wang, J.; Xie, R.-H.; et al. Safety and efficacy of meplazumab in healthy volunteers and COVID-19 patients: A randomized phase 1 and an exploratory phase 2 trial. Signal Transduct. Target. Ther. 2021, 6, 194. [Google Scholar] [CrossRef] [PubMed]
- Gubernatorova, E.O.; Gorshkova, E.A.; Polinova, A.I.; Drutskaya, M.S. IL-6: Relevance for immunopathology of SARS-CoV-2. Cytokine Growth Factor Rev. 2020, 53, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, H.; Pillat, M.M. CD147 as a Target for COVID-19 Treatment: Suggested Effects of Azithromycin and Stem Cell Engagement. Stem Cell Rev. Rep. 2020, 16, 434–440. [Google Scholar] [CrossRef]
- Schijns, V.; Lavelle, E.C. Prevention and treatment of COVID-19 disease by controlled modulation of innate immunity. Eur. J. Immunol. 2020, 50, 932–938. [Google Scholar] [CrossRef]
- Reina, J.; Reina, N. Favipiravir, a new concept of antiviral drug against influenza viruses. Rev. Esp. Quimioter. Publ. Of. Soc. Esp. Quimioter. 2017, 30, 79–83. [Google Scholar]
- Tchesnokov, E.P.; Feng, J.Y.; Porter, D.P.; Götte, M. Mechanism of Inhibition of Ebola Virus RNA-Dependent RNA Polymerase by Remdesivir. Viruses 2019, 11, 326. [Google Scholar] [CrossRef] [PubMed]
- Buckland, M.S.; Galloway, J.B.; Fhogartaigh, C.N.; Meredith, L.; Provine, N.M.; Bloor, S.; Ogbe, A.; Zelek, W.M.; Smielewska, A.; Yakovleva, A.; et al. Treatment of COVID-19 with remdesivir in the absence of humoral immunity: A case report. Nat. Commun. 2020, 11, 6385. [Google Scholar] [CrossRef]
- Khan, F.I.; Kang, T.; Ali, H.; Lai, D. Remdesivir Strongly Binds to RNA-Dependent RNA Polymerase, Membrane Protein, and Main Protease of SARS-CoV-2: Indication From Molecular Modeling and Simulations. Front. Pharmacol. 2021, 12, 1616. [Google Scholar] [CrossRef]
- Doi, K.; Ikeda, M.; Hayase, N.; Moriya, K.; Morimura, N.; Maehara, H.; Tagami, S.; Fukushima, K.; Misawa, N.; Inoue, Y.; et al. Nafamostat mesylate treatment in combination with favipiravir for patients critically ill with COVID-19: A case series. Crit. Care 2020, 24, 392. [Google Scholar] [CrossRef]
- Hifumi, T.; Isokawa, S.; Otani, N.; Ishimatsu, S. Adverse events associated with nafamostat mesylate and favipiravir treatment in COVID-19 patients. Crit. Care 2020, 24, 497. [Google Scholar] [CrossRef]
- Osborne, V.; Davies, M.; Lane, S.; Evans, A.; Denyer, J.; Dhanda, S.; Roy, D.; Shakir, S. Lopinavir-Ritonavir in the Treatment of COVID-19: A Dynamic Systematic Benefit-Risk Assessment. Drug Saf. 2020, 43, 809–821. [Google Scholar] [CrossRef]
- Stower, H. Lopinavir–ritonavir in severe COVID-19. Nat. Med. 2020, 26, 465. [Google Scholar] [CrossRef] [PubMed]
- Bikdeli, B.; Madhavan, M.V.; Jimenez, D.; Chuich, T.; Dreyfus, I.; Driggin, E.; Nigoghossian, C.D.; Ageno, W.; Madjid, M.; Guo, Y.; et al. COVID-19 and Thrombotic or Thromboembolic Disease: Implications for Prevention, Antithrombotic Therapy, and Follow-Up: JACC State-of-the-Art Review. J. Am. Coll. Cardiol. 2020, 75, 2950–2973. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Chen, R.; Liu, C.; Liang, W.; Guan, W.; Tang, R.; Tang, C.; Zhang, N.; Zhong, N.; Li, S. Attention should be paid to venous thromboembolism prophylaxis in the management of COVID-19. Lancet Haematol. 2020, 7, e362–e363. [Google Scholar] [CrossRef]
- Paranjpe, I.; Fuster, V.; Lala, A.; Russak, A.J.; Glicksberg, B.S.; Levin, M.A.; Charney, A.W.; Narula, J.; Fayad, Z.A.; Bagiella, E.; et al. Association of Treatment Dose Anticoagulation With In-Hospital Survival Among Hospitalized Patients With COVID-19. J. Am. Coll. Cardiol. 2020, 76, 122–124. [Google Scholar] [CrossRef]
- Barnes, G.D.; Burnett, A.; Allen, A.; Blumenstein, M.; Clark, N.P.; Cuker, A.; Dager, W.E.; Deitelzweig, S.B.; Ellsworth, S.; Garcia, D.; et al. Thromboembolism and anticoagulant therapy during the COVID-19 pandemic: Interim clinical guidance from the anticoagulation forum. J. Thromb. Thrombolysis 2020, 50, 72–81. [Google Scholar] [CrossRef]
- Paar, V.; Wernly, B.; Zhou, Z.; Motloch, L.J.; Hoppe, U.C.; Egle, A.; Lichtenauer, M. Anti-coagulation for COVID-19 treatment: Both anti-thrombotic and anti-inflammatory? J. Thromb. Thrombolysis 2021, 51, 226–231. [Google Scholar] [CrossRef]
- Russo, V.; Cardillo, G.; Viggiano, G.V.; Mangiacapra, S.; Cavalli, A.; Fontanella, A.; Agrusta, F.; Bellizzi, A.; Amitrano, M.; Iannuzzo, M.; et al. Fondaparinux Use in Patients With COVID-19: A Preliminary Multicenter Real-World Experience. J. Cardiovasc. Pharmacol. 2020, 76, 369–371. [Google Scholar] [CrossRef]
- Flam, B.; Wintzell, V.; Ludvigsson, J.F.; Mårtensson, J.; Pasternak, B. Direct oral anticoagulant use and risk of severe COVID-19. J. Intern. Med. 2021, 289, 411–419. [Google Scholar] [CrossRef]
- Rossi, R.; Coppi, F.; Talarico, M.; Boriani, G. Protective role of chronic treatment with direct oral anticoagulants in elderly patients affected by interstitial pneumonia in COVID-19 era. Eur. J. Intern. Med. 2020, 77, 158–160. [Google Scholar] [CrossRef]
- Meini, S.; Zanichelli, A.; Sbrojavacca, R.; Iuri, F.; Roberts, A.T.; Suffritti, C.; Tascini, C. Understanding the Pathophysiology of COVID-19: Could the Contact System Be the Key? Front. Immunol. 2020, 11, 2014. [Google Scholar] [CrossRef]
- Mizurini, D.M.; Hottz, E.D.; Bozza, P.T.; Monteiro, R.Q. Fundamentals in COVID-19-Associated Thrombosis: Molecular and Cellular Aspects. Front. Cardiovasc. Med. 2021, 8, 785738. [Google Scholar] [CrossRef] [PubMed]
- Taus, F.; Salvagno, G.; Canè, S.; Fava, C.; Mazzaferri, F.; Carrara, E.; Petrova, V.; Barouni, R.M.; Dima, F.; Dalbeni, A.; et al. Platelets Promote Thromboinflammation in SARS-CoV-2 Pneumonia. Arterioscler. Thromb. Vasc. Biol. 2020, 40, 2975–2989. [Google Scholar] [CrossRef] [PubMed]
- Asakura, H.; Ogawa, H. Potential of heparin and nafamostat combination therapy for COVID-19. J. Thromb. Haemost. 2020, 18, 1521–1522. [Google Scholar] [CrossRef] [PubMed]
- Bonaventura, A.; Montecucco, F.; Dallegri, F. Update on the effects of treatment with recombinant tissue-type plasminogen activator (rt-PA) in acute ischemic stroke. Expert Opin. Biol. Ther. 2016, 16, 1323–1340. [Google Scholar] [CrossRef]
- Wang, J.; Hajizadeh, N.; Moore, E.E.; McIntyre, R.C.; Moore, P.K.; Veress, L.A.; Yaffe, M.B.; Moore, H.B.; Barrett, C.D. Tissue plasminogen activator (tPA) treatment for COVID-19 associated acute respiratory distress syndrome (ARDS): A case series. J. Thromb. Haemost. 2020, 18, 1752–1755. [Google Scholar] [CrossRef] [PubMed]
- Kanthi, Y.; Knight, J.S.; Zuo, Y.; Pinsky, D.J. New (re)purpose for an old drug: Purinergic modulation may extinguish the COVID-19 thromboinflammatory firestorm. JCI Insight 2020, 5, e140971. [Google Scholar] [CrossRef]
- Fata-Hartley, C.L.; Palmenberg, A.C. Dipyridamole Reversibly Inhibits Mengovirus RNA Replication. J. Virol. 2005, 79, 11062–11070. [Google Scholar] [CrossRef]
- Huang, B.; Chen, Z.; Geng, L.; Wang, J.; Liang, H.; Cao, Y.; Chen, H.; Huang, W.; Su, M.; Wang, H.; et al. Mucosal Profiling of Pediatric-Onset Colitis and IBD Reveals Common Pathogenics and Therapeutic Pathways. Cell 2019, 179, 1160–1176.e1124. [Google Scholar] [CrossRef]
- Insel, P.A.; Murray, F.; Yokoyama, U.; Romano, S.; Yun, H.; Brown, L.; Snead, A.; Lu, D.; Aroonsakool, N. cAMP and Epac in the regulation of tissue fibrosis. Br. J. Pharmacol. 2012, 166, 447–456. [Google Scholar] [CrossRef]
- Anderson, R.M.; Vegvari, C.; Hollingsworth, T.D.; Pi, L.; Maddren, R.; Ng, C.W.; Baggaley, R.F. The SARS-CoV-2 pandemic: Remaining uncertainties in our understanding of the epidemiology and transmission dynamics of the virus, and challenges to be overcome. Interface Focus 2021, 11, 20210008. [Google Scholar] [CrossRef]
- Liu, X.; Li, Z.; Liu, S.; Sun, J.; Chen, Z.; Jiang, M.; Zhang, Q.; Wei, Y.; Wang, X.; Huang, Y.-Y.; et al. Potential therapeutic effects of dipyridamole in the severely ill patients with COVID-19. Acta Pharm. Sin. B 2020, 10, 1205–1215. [Google Scholar] [CrossRef] [PubMed]
- Mitsios, A.; Chrysanthopoulou, A.; Arampatzioglou, A.; Angelidou, I.; Vidali, V.; Ritis, K.; Skendros, P.; Stakos, D. Ticagrelor Exerts Immune-Modulatory Effect by Attenuating Neutrophil Extracellular Traps. Int. J. Mol. Sci. 2020, 21, 3625. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Majumdar, S.; Singh, R.; Misra, A. Role of corticosteroid in the management of COVID-19: A systemic review and a Clinician’s perspective. Diabetes Metab. Syndr. Clin. Res. Rev. 2020, 14, 971–978. [Google Scholar] [CrossRef]
- Magro, G. SARS-CoV-2 and COVID-19: Is interleukin-6 (IL-6) the ‘culprit lesion’ of ARDS onset? What is there besides Tocilizumab? SGP130Fc. Cytokine X 2020, 2, 100029. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Han, M.; Li, T.; Sun, W.; Wang, D.; Fu, B.; Zhou, Y.; Zheng, X.; Yang, Y.; Li, X.; et al. Effective treatment of severe COVID-19 patients with tocilizumab. Proc. Natl. Acad. Sci. USA 2020, 117, 10970–10975. [Google Scholar] [CrossRef] [PubMed]
- Herold, T.; Jurinovic, V.; Arnreich, C.; Lipworth, B.J.; Hellmuth, J.C.; von Bergwelt-Baildon, M.; Klein, M.; Weinberger, T. Elevated levels of IL-6 and CRP predict the need for mechanical ventilation in COVID-19. J. Allergy Clin. Immunol. 2020, 146, 128–136.e124. [Google Scholar] [CrossRef]
- Liu, B.; Li, M.; Zhou, Z.; Guan, X.; Xiang, Y. Can we use interleukin-6 (IL-6) blockade for coronavirus disease 2019 (COVID-19)-induced cytokine release syndrome (CRS)? J. Autoimmun. 2020, 111, 102452. [Google Scholar] [CrossRef]
- Somers, E.C.; Eschenauer, G.A.; Troost, J.P.; Golob, J.L.; Gandhi, T.N.; Wang, L.; Zhou, N.; Petty, L.A.; Baang, J.H.; Dillman, N.O.; et al. Tocilizumab for Treatment of Mechanically Ventilated Patients With COVID-19. Clin. Infect. Dis. 2020, 73, e445–e454. [Google Scholar] [CrossRef]
- Richardson, P.; Griffin, I.; Tucker, C.; Smith, D.; Oechsle, O.; Phelan, A.; Rawling, M.; Savory, E.; Stebbing, J. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet 2020, 395, e30–e31. [Google Scholar] [CrossRef]
- Riegler, L.L.; Jones, G.P.; Lee, D.W. Current approaches in the grading and management of cytokine release syndrome after chimeric antigen receptor T-cell therapy. Clin. Risk Manag. 2019, 15, 323–335. [Google Scholar] [CrossRef]
- Shimabukuro-Vornhagen, A.; Gödel, P.; Subklewe, M.; Stemmler, H.J.; Schlößer, H.A.; Schlaak, M.; Kochanek, M.; Böll, B.; von Bergwelt-Baildon, M.S. Cytokine release syndrome. J. ImmunoTherapy Cancer 2018, 6, 56. [Google Scholar] [CrossRef] [PubMed]
- Synowiec, A.; Szczepański, A.; Barreto-Duran, E.; Lie, L.K.; Pyrc, K. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): A Systemic Infection. Clin. Microbiol. Rev. 2021, 34, e00133-00120. [Google Scholar] [CrossRef]
- Gorshkov, K.; Chen, C.Z.; Bostwick, R.; Rasmussen, L.; Tran, B.N.; Cheng, Y.-S.; Xu, M.; Pradhan, M.; Henderson, M.; Zhu, W.; et al. The SARS-CoV-2 Cytopathic Effect Is Blocked by Lysosome Alkalizing Small Molecules. ACS Infect. Dis. 2021, 7, 1389–1408. [Google Scholar] [CrossRef]
- Chen, I.-Y.; Moriyama, M.; Chang, M.-F.; Ichinohe, T. Severe Acute Respiratory Syndrome Coronavirus Viroporin 3a Activates the NLRP3 Inflammasome. Front. Microbiol. 2019, 10, 50. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Fu, B.; Zheng, X.; Wang, D.; Zhao, C.; Qi, Y.; Sun, R.; Tian, Z.; Xu, X.; Wei, H. Pathogenic T-cells and inflammatory monocytes incite inflammatory storms in severe COVID-19 patients. Natl. Sci. Rev. 2020, 7, 998–1002. [Google Scholar] [CrossRef] [PubMed]
- Schett, G.; Sticherling, M.; Neurath, M.F. COVID-19: Risk for cytokine targeting in chronic inflammatory diseases? Nat. Rev. Immunol. 2020, 20, 271–272. [Google Scholar] [CrossRef] [PubMed]
- Venerito, V.; Lopalco, G.; Iannone, F. COVID-19, rheumatic diseases and immunosuppressive drugs: An appeal for medication adherence. Rheumatol. Int. 2020, 40, 827–828. [Google Scholar] [CrossRef]
- Nile, S.H.; Nile, A.; Qiu, J.; Li, L.; Jia, X.; Kai, G. COVID-19: Pathogenesis, cytokine storm and therapeutic potential of interferons. Cytokine Growth Factor Rev. 2020, 53, 66–70. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Zhang, W.; Zhao, Y.; Zhang, F.; Wang, Q.; Li, T.; Liu, Z.; Wang, J.; Qin, Y.; Zhang, X.; Yan, X.; et al. The use of anti-inflammatory drugs in the treatment of people with severe coronavirus disease 2019 (COVID-19): The Perspectives of clinical immunologists from China. Clin. Immunol. 2020, 214, 108393. [Google Scholar] [CrossRef]
- Dariya, B.; Nagaraju, G.P. Understanding novel COVID-19: Its impact on organ failure and risk assessment for diabetic and cancer patients. Cytokine Growth Factor Rev. 2020, 53, 43–52. [Google Scholar] [CrossRef]
- Ronco, C.; Reis, T.; Husain-Syed, F. Management of acute kidney injury in patients with COVID-19. Lancet Respir. Med. 2020, 8, 738–742. [Google Scholar] [CrossRef]
- Moon, C. Fighting COVID-19 exhausts T cells. Nat. Rev. Immunol. 2020, 20, 277. [Google Scholar] [CrossRef] [PubMed]
- Côté, P.; Baril, J.-G.; Hébert, M.-N.; Klein, M.; Lalonde, R.; Poliquin, M.; Rouleau, D.; Therrien, R.; Vézina, S.; Willems, B.; et al. Management and Treatment of Hepatitis C Virus in Patients with HIV and Hepatitis C Virus Coinfection: A Practical Guide for Health Care Professionals. Can. J. Infect. Dis. Med. Microbiol. 2007, 18, 631054. [Google Scholar] [CrossRef]
- Taubenberger, J.K.; Morens, D.M. The Pathology of Influenza Virus Infections. Annu. Rev. Pathol. Mech. Dis. 2008, 3, 499–522. [Google Scholar] [CrossRef]
- Kim, Y.-I.; Kim, S.-G.; Kim, S.-M.; Kim, E.-H.; Park, S.-J.; Yu, K.-M.; Chang, J.-H.; Kim, E.J.; Lee, S.; Casel, M.A.B.; et al. Infection and Rapid Transmission of SARS-CoV-2 in Ferrets. Cell Host Microbe 2020, 27, 704–709.e702. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Philip, A.T.; Stoppacher, R. Pathologic Findings in Novel Influenza A (H1N1) Virus (“Swine Flu”) Infection: Contrasting Clinical Manifestations and Lung Pathology in Two Fatal Cases. Am. J. Clin. Pathol. 2010, 133, 380–387. [Google Scholar] [CrossRef]
- Pascal, L.; Hivert, B.; Trauet, J.; Deberranger, E.; Dessaint, J.-P.; Yakoub-Agha, I.; Labalette, M. A Low Effective Dose of Interleukin-7 Is Sufficient to Maintain Cord Blood T Cells Alive without Potentiating Allo-Immune Responses. Biol. Blood Marrow Transplant. 2015, 21, 625–631. [Google Scholar] [CrossRef]
- Zhong, J.; Tang, J.; Ye, C.; Dong, L. The immunology of COVID-19: Is immune modulation an option for treatment? Lancet Rheumatol. 2020, 2, e428–e436. [Google Scholar] [CrossRef]
- Rovere Querini, P.; De Lorenzo, R.; Conte, C.; Brioni, E.; Lanzani, C.; Yacoub, M.R.; Chionna, R.; Martinenghi, S.; Vitali, G.; Tresoldi, M.; et al. Post-COVID-19 follow-up clinic: Depicting chronicity of a new disease. Acta Biomed. Atenei Parm. 2020, 91, 22–28. [Google Scholar] [CrossRef]
- Yang, L.; Xie, X.; Tu, Z.; Fu, J.; Xu, D.; Zhou, Y. The signal pathways and treatment of cytokine storm in COVID-19. Signal Transduct. Target. Ther. 2021, 6, 255. [Google Scholar] [CrossRef] [PubMed]
- Awadasseid, A.; Yin, Q.; Wu, Y.; Zhang, W. Potential protective role of the anti-PD-1 blockade against SARS-CoV-2 infection. Biomed. Pharmacother. 2021, 142, 111957. [Google Scholar] [CrossRef] [PubMed]
- Bekele, Y.; Sui, Y.; Berzofsky, J.A. IL-7 in SARS-CoV-2 Infection and as a Potential Vaccine Adjuvant. Front. Immunol. 2021, 12, 3796. [Google Scholar] [CrossRef] [PubMed]
- El Baba, R.; Herbein, G. Management of epigenomic networks entailed in coronavirus infections and COVID-19. Clin. Epigenetics 2020, 12, 118. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.-C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.e1039. [Google Scholar] [CrossRef]
- Aevermann, B.D.; Pickett, B.E.; Kumar, S.; Klem, E.B.; Agnihothram, S.; Askovich, P.S.; Bankhead, A.; Bolles, M.; Carter, V.; Chang, J.; et al. A comprehensive collection of systems biology data characterizing the host response to viral infection. Sci. Data 2014, 1, 140033. [Google Scholar] [CrossRef]
- Menachery, V.D.; Eisfeld, A.J.; Schäfer, A.; Josset, L.; Sims, A.C.; Proll, S.; Fan, S.; Li, C.; Neumann, G.; Tilton, S.C.; et al. Pathogenic Influenza Viruses and Coronaviruses Utilize Similar and Contrasting Approaches To Control Interferon-Stimulated Gene Responses. mBio 2014, 5, e01174-01114. [Google Scholar] [CrossRef]
- Akira, S. Pathogen recognition by innate immunity and its signaling. Proc. Jpn. Acad. Ser. B 2009, 85, 143–156. [Google Scholar] [CrossRef]
- Kawai, T.; Akira, S. The role of pattern-recognition receptors in innate immunity: Update on Toll-like receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef]
- Bruening, J.; Weigel, B.; Gerold, G. The Role of Type III Interferons in Hepatitis C Virus Infection and Therapy. J. Immunol. Res. 2017, 2017, 7232361. [Google Scholar] [CrossRef]
- Shin, D.; Mukherjee, R.; Grewe, D.; Bojkova, D.; Baek, K.; Bhattacharya, A.; Schulz, L.; Widera, M.; Mehdipour, A.R.; Tascher, G.; et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 2020, 587, 657–662. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-Y.; Liao, C.-H.; Wang, Q.; Tan, Y.-J.; Luo, R.; Qiu, Y.; Ge, X.-Y. The ORF6, ORF8 and nucleocapsid proteins of SARS-CoV-2 inhibit type I interferon signaling pathway. Virus Res. 2020, 286, 198074. [Google Scholar] [CrossRef]
- Liu, X.; Cao, H.; Li, J.; Wang, B.; Zhang, P.; Dong Zhang, X.; Liu, Z.; Yuan, H.; Zhan, Z. Autophagy induced by DAMPs facilitates the inflammation response in lungs undergoing ischemia-reperfusion injury through promoting TRAF6 ubiquitination. Cell Death Differ. 2017, 24, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Schöllhorn, A.; Schuhmacher, J.; Besedovsky, L.; Fendel, R.; Jensen, A.T.R.; Stevanović, S.; Lange, T.; Rammensee, H.-G.; Born, J.; Gouttefangeas, C.; et al. Integrin Activation Enables Sensitive Detection of Functional CD4+ and CD8+ T Cells: Application to Characterize SARS-CoV-2 Immunity. Front. Immunol. 2021, 12, 522. [Google Scholar] [CrossRef] [PubMed]
- Mazzoni, A.; Maggi, L.; Capone, M.; Spinicci, M.; Salvati, L.; Colao, M.G.; Vanni, A.; Kiros, S.T.; Mencarini, J.; Zammarchi, L.; et al. Cell-mediated and humoral adaptive immune responses to SARS-CoV-2 are lower in asymptomatic than symptomatic COVID-19 patients. Eur. J. Immunol. 2020, 50, 2013–2024. [Google Scholar] [CrossRef] [PubMed]
- Kroemer, M.; Spehner, L.; Vettoretti, L.; Bouard, A.; Eberst, G.; Pili Floury, S.; Capellier, G.; Lepiller, Q.; Orillard, E.; Mansi, L.; et al. COVID-19 patients display distinct SARS-CoV-2 specific T-cell responses according to disease severity. J. Infect. 2021, 82, 282–327. [Google Scholar] [CrossRef] [PubMed]
- Hung, I.F.-N.; Lung, K.-C.; Tso, E.Y.-K.; Liu, R.; Chung, T.W.-H.; Chu, M.-Y.; Ng, Y.-Y.; Lo, J.; Chan, J.; Tam, A.R.; et al. Triple combination of interferon beta-1b, lopinavir–ritonavir, and ribavirin in the treatment of patients admitted to hospital with COVID-19: An open-label, randomised, phase 2 trial. Lancet 2020, 395, 1695–1704. [Google Scholar] [CrossRef]
- Mantlo, E.; Bukreyeva, N.; Maruyama, J.; Paessler, S.; Huang, C. Antiviral activities of type I interferons to SARS-CoV-2 infection. Antivir. Res. 2020, 179, 104811. [Google Scholar] [CrossRef]
- Choi, H.; Shin, E.-C. Roles of Type I and III Interferons in COVID-19. Yonsei Med. J. 2021, 62, 381–390. [Google Scholar] [CrossRef]
- Ramos, I.; Fernandez-Sesma, A. Modulating the Innate Immune Response to Influenza A Virus: Potential Therapeutic Use of Anti-Inflammatory Drugs. Front. Immunol. 2015, 6, 361. [Google Scholar] [CrossRef]
- Al-Tawfiq, J.A.; Momattin, H.; Dib, J.; Memish, Z.A. Ribavirin and interferon therapy in patients infected with the Middle East respiratory syndrome coronavirus: An observational study. Int. J. Infect. Dis. 2014, 20, 42–46. [Google Scholar] [CrossRef] [PubMed]
- Arabi, Y.M.; Shalhoub, S.; Mandourah, Y.; Al-Hameed, F.; Al-Omari, A.; Al Qasim, E.; Jose, J.; Alraddadi, B.; Almotairi, A.; Al Khatib, K.; et al. Ribavirin and Interferon Therapy for Critically Ill Patients With Middle East Respiratory Syndrome: A Multicenter Observational Study. Clin. Infect. Dis. 2019, 70, 1837–1844. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.M.; Cheng, V.C.C.; Hung, I.F.N.; Wong, M.M.L.; Chan, K.H.; Chan, K.S.; Kao, R.Y.T.; Poon, L.L.M.; Wong, C.L.P.; Guan, Y.; et al. Role of lopinavir/ritonavir in the treatment of SARS: Initial virological and clinical findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef] [PubMed]
- Tao, J.-H.; Barbi, J.; Pan, F. Hypoxia-inducible factors in T lymphocyte differentiation and function. A Review in the Theme: Cellular Responses to Hypoxia. Am. J. Physiol. Cell Physiol. 2015, 309, C580–C589. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.; Blum, H.E.; Thimme, R. T-cell responses in hepatitis B and C virus infection: Similarities and differences. Emerg. Microbes Infect. 2013, 2, e15. [Google Scholar] [CrossRef] [PubMed]
- Ostrand-Rosenberg, S.; Horn, L.A.; Haile, S.T. The Programmed Death-1 Immune-Suppressive Pathway: Barrier to Antitumor Immunity. J. Immunol. 2014, 193, 3835–3841. [Google Scholar] [CrossRef]
- Mishra, V.; Agas, A.; Schuetz, H.; Kalluru, J.; Haorah, J. Alcohol induces programmed death receptor-1 and programmed death-ligand-1 differentially in neuroimmune cells. Alcohol 2020, 86, 65–74. [Google Scholar] [CrossRef]
- Porichis, F.; Kaufmann, D.E. Role of PD-1 in HIV Pathogenesis and as Target for Therapy. Curr. HIV/AIDS Rep. 2012, 9, 81–90. [Google Scholar] [CrossRef]
- Bersanelli, M. Controversies about COVID-19 and anticancer treatment with immune checkpoint inhibitors. Immunotherapy 2020, 12, 269–273. [Google Scholar] [CrossRef]
- Poe, F.L.; Corn, J. N-Acetylcysteine: A potential therapeutic agent for SARS-CoV-2. Med. Hypotheses 2020, 143, 109862. [Google Scholar] [CrossRef]
- Shah, V.K.; Firmal, P.; Alam, A.; Ganguly, D.; Chattopadhyay, S. Overview of Immune Response During SARS-CoV-2 Infection: Lessons From the Past. Front. Immunol. 2020, 11, 1949. [Google Scholar] [CrossRef] [PubMed]
- Fogel, D.B. Factors associated with clinical trials that fail and opportunities for improving the likelihood of success: A review. Contemp. Clin. Trials Commun. 2018, 11, 156–164. [Google Scholar] [CrossRef] [PubMed]
- Gnjatic, S.; Bronte, V.; Brunet, L.R.; Butler, M.O.; Disis, M.L.; Galon, J.; Hakansson, L.G.; Hanks, B.A.; Karanikas, V.; Khleif, S.N.; et al. Identifying baseline immune-related biomarkers to predict clinical outcome of immunotherapy. J. ImmunoTherapy Cancer 2017, 5, 44. [Google Scholar] [CrossRef] [PubMed]
- Lei, H. A single transcript for the prognosis of disease severity in COVID-19 patients. Sci. Rep. 2021, 11, 12174. [Google Scholar] [CrossRef] [PubMed]
- Ganji, A.; Farahani, I.; Khansarinejad, B.; Ghazavi, A.; Mosayebi, G. Increased expression of CD8 marker on T-cells in COVID-19 patients. Blood Cells Mol. Dis. 2020, 83, 102437. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Yi, X.; Sun, Y.; Bi, X.; Du, J.; Zhang, C.; Quan, S.; Zhang, F.; Sun, R.; Qian, L.; et al. Proteomic and Metabolomic Characterization of COVID-19 Patient Sera. Cell 2020, 182, 59–72.e15. [Google Scholar] [CrossRef]
- Messner, C.B.; Demichev, V.; Wendisch, D.; Michalick, L.; White, M.; Freiwald, A.; Textoris-Taube, K.; Vernardis, S.I.; Egger, A.-S.; Kreidl, M.; et al. Ultra-High-Throughput Clinical Proteomics Reveals Classifiers of COVID-19 Infection. Cell Syst. 2020, 11, 11–24.e14. [Google Scholar] [CrossRef]
- Long, Q.-X.; Tang, X.-J.; Shi, Q.-L.; Li, Q.; Deng, H.-J.; Yuan, J.; Hu, J.-L.; Xu, W.; Zhang, Y.; Lv, F.-J.; et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 2020, 26, 1200–1204. [Google Scholar] [CrossRef]
- Pollard, A.J.; Bijker, E.M. A guide to vaccinology: From basic principles to new developments. Nat. Rev. Immunol. 2021, 21, 83–100. [Google Scholar] [CrossRef]
- Tong, X.; Cheng, A.; Yuan, X.; Zhong, X.; Wang, H.; Zhou, W.; Xu, X.; Li, Y. Characteristics of peripheral white blood cells in COVID-19 patients revealed by a retrospective cohort study. BMC Infect. Dis. 2021, 21, 1236. [Google Scholar] [CrossRef]
- Lu, C.; Liu, Y.; Chen, B.; Yang, H.; Hu, H.; Liu, Y.; Zhao, Y. Prognostic value of lymphocyte count in severe COVID-19 patients with corticosteroid treatment. Signal Transduct Target 2021, 6, 106. [Google Scholar] [CrossRef]
- deKay, J.T.; Emery, I.F.; Rud, J.; Eldridge, A.; Lord, C.; Gagnon, D.J.; May, T.L.; Herrera, V.L.M.; Ruiz-Opazo, N.; Riker, R.R.; et al. DEspRhigh neutrophils are associated with critical illness in COVID-19. Sci. Rep. 2021, 11, 22463. [Google Scholar] [CrossRef] [PubMed]
- Dávila-Collado, R.; Jarquín-Durán, O.; Solís-Vallejo, A.; Nguyen, M.A.; Espinoza, J.L. Elevated Monocyte to Lymphocyte Ratio and Increased Mortality among Patients with Chronic Kidney Disease Hospitalized for COVID-19. J. Pers. Med. 2021, 11, 224. [Google Scholar] [CrossRef] [PubMed]
- Prozan, L.; Shusterman, E.; Ablin, J.; Mitelpunkt, A.; Weiss-Meilik, A.; Adler, A.; Choshen, G.; Kehat, O. Prognostic value of neutrophil-to-lymphocyte ratio in COVID-19 compared with Influenza and respiratory syncytial virus infection. Sci. Rep. 2021, 11, 21519. [Google Scholar] [CrossRef]
- Gumus, H.; Demir, A.; Yükkaldıran, A. Is mean platelet volume a predictive marker for the diagnosis of COVID-19 in children? Int. J. Clin. Pract. 2021, 75, e13892. [Google Scholar] [CrossRef] [PubMed]
- Alsuwaidi, L.; Al Heialy, S.; Shaikh, N.; Al Najjar, F.; Seliem, R.; Han, A.; Hachim, M. Monocyte distribution width as a novel sepsis indicator in COVID-19 patients. BMC Infect. Dis. 2022, 22, 27. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudi, S.; Yaghmaei, B.; Sharifzadeh Ekbatani, M.; Pourakbari, B.; Navaeian, A.; Parvaneh, N.; Haghi Ashtiani, M.T.; Mamishi, S. Effects of Coronavirus Disease 2019 (COVID-19) on Peripheral Blood Lymphocytes and Their Subsets in Children: Imbalanced CD4+/CD8+ T Cell Ratio and Disease Severity. Front. Pediatrics 2021, 9, 267. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Rong, L.; Cui, R.; Feng, J.; Jin, Y.; Chen, X.; Xu, R. Dynamic changes in serum IL-6, IL-8, and IL-10 predict the outcome of ICU patients with severe COVID-19. Ann. Palliat. Med. 2021, 10, 3706–3714. [Google Scholar] [CrossRef]
- Hu, R.; Han, C.; Pei, S.; Yin, M.; Chen, X. Procalcitonin levels in COVID-19 patients. Int. J. Antimicrob. Agents 2020, 56, 106051. [Google Scholar] [CrossRef]
- Zinellu, A.; Paliogiannis, P.; Carru, C.; Mangoni, A.A. Serum amyloid A concentrations, COVID-19 severity and mortality: An updated systematic review and meta-analysis. Int. J. Infect. Dis. 2021, 105, 668–674. [Google Scholar] [CrossRef]
- Mardani, R.; Namavar, M.; Ghorbi, E.; Shoja, Z.; Zali, F.; Kaghazian, H.; Aghasadeghi, M.R.; Sadeghi, S.A.; Sabeti, S.; Darazam, I.A.; et al. Association between serum inflammatory parameters and the disease severity in COVID-19 patients. J. Clin. Lab. Anal. 2022, 36, e24162. [Google Scholar] [CrossRef] [PubMed]
- Luan, Y.-Y.; Yin, C.-H.; Yao, Y.-M. Update Advances on C-Reactive Protein in COVID-19 and Other Viral Infections. Front. Immunol. 2021, 12, 3153. [Google Scholar] [CrossRef] [PubMed]
- Tural Onur, S.; Altın, S.; Sokucu, S.N.; Fikri, B.; Barça, T.; Bolat, E.; Toptaş, M. Could ferritin level be an indicator of COVID-19 disease mortality? J. Med. Virol. 2021, 93, 1672–1677. [Google Scholar] [CrossRef]
- Marfia, G.; Navone, S.; Guarnaccia, L.; Campanella, R.; Mondoni, M.; Locatelli, M.; Barassi, A.; Fontana, L.; Palumbo, F.; Garzia, E.; et al. Decreased serum level of sphingosine-1-phosphate: A novel predictor of clinical severity in COVID-19. EMBO Mol. Med. 2021, 13, e13424. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.-C.; You, C.-Y.; Lu, S.-W.; Fu, Y.-Q. Characteristics of coagulation alteration in patients with COVID-19. Ann. Hematol. 2021, 100, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yan, X.; Fan, Q.; Liu, H.; Liu, X.; Liu, Z.; Zhang, Z. D-dimer levels on admission to predict in-hospital mortality in patients with COVID-19. J. Thromb. Haemost. JTH 2020, 18, 1324–1329. [Google Scholar] [CrossRef]
- Long, H.; Nie, L.; Xiang, X.; Li, H.; Zhang, X.; Fu, X.; Ren, H.; Liu, W.; Wang, Q.; Wu, Q. D-Dimer and Prothrombin Time Are the Significant Indicators of Severe COVID-19 and Poor Prognosis. BioMed Res. Int. 2020, 2020, 6159720. [Google Scholar] [CrossRef]
- Han, H.; Yang, L.; Liu, R.; Liu, F.; Wu, K.L.; Li, J.; Liu, X.H.; Zhu, C.L. Prominent changes in blood coagulation of patients with SARS-CoV-2 infection. Clin. Chem. Lab. Med. 2020, 58, 1116–1120. [Google Scholar] [CrossRef]
- Shimura, T.; Kurano, M.; Kanno, Y.; Ikeda, M.; Okamoto, K.; Jubishi, D.; Harada, S.; Okugawa, S.; Moriya, K.; Yatomi, Y. Clot waveform of APTT has abnormal patterns in subjects with COVID-19. Sci. Rep. 2021, 11, 5190. [Google Scholar] [CrossRef]
- Araya, S.; Mamo, M.A.; Tsegay, Y.G.; Atlaw, A.; Aytenew, A.; Hordofa, A.; Negeso, A.E.; Wordofa, M.; Niguse, T.; Cheru, M.; et al. Blood coagulation parameter abnormalities in hospitalized patients with confirmed COVID-19 in Ethiopia. PLoS ONE 2021, 16, e0252939. [Google Scholar] [CrossRef]
- Liaquat, H.; Shupp, B.; Rollins, S.; Schneider, Y.; Matin, A. Comparison of the impact of chronic corticosteroid therapy on critical care outcomes of COVID-19 patients with and without history of chronic liver disease. Sci. Rep. 2021, 11, 19245. [Google Scholar] [CrossRef] [PubMed]
- Szarpak, L.; Ruetzler, K.; Safiejko, K.; Hampel, M.; Pruc, M.; Kanczuga-Koda, L.; Filipiak, K.J.; Jaguszewski, M.J. Lactate dehydrogenase level as a COVID-19 severity marker. Am. J. Emerg. Med. 2021, 45, 638–639. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Wei, Y.; Lyu, X.; Zhao, B.; Feng, Y.; Li, T.; Cao, H.; Yang, X.; Zhou, X.; Wang, W.; et al. High aspartate aminotransferase to alanine aminotransferase ratio on admission as risk factor for poor prognosis in COVID-19 patients. Sci. Rep. 2020, 10, 16496. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Li, J.; Long, W.; Zeng, W.; Gao, R.; Zeng, G.; Chen, D.; Wang, S.; Li, Q.; Hu, D.; et al. Bilirubin Levels as Potential Indicators of Disease Severity in Coronavirus Disease Patients: A Retrospective Cohort Study. Front. Med. 2020, 7, 799. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Yu, C.; Yang, Q.; Yuan, X.; Yang, F.; Li, P.; Chen, G.; Liang, W.; Yang, Y. The clinical implication of gamma-glutamyl transpeptidase in COVID-19. Liver Res. 2021, 5, 209–216. [Google Scholar] [CrossRef]
- Takeshita, Y.; Terada, J.; Hirasawa, Y.; Kinoshita, T.; Tajima, H.; Koshikawa, K.; Kinouchi, T.; Isaka, Y.; Shionoya, Y.; Fujikawa, A.; et al. Elevated TAT in COVID-19 Patients with Normal D-Dimer as a Predictor of Severe Respiratory Failure: A Retrospective Analysis of 797 Patients. J. Clin. Med. 2022, 11, 134. [Google Scholar] [CrossRef]
- Ponti, G.; Ruini, C.; Tomasi, A. Homocysteine as a potential predictor of cardiovascular risk in patients with COVID-19. Med. Hypotheses 2020, 143, 109859. [Google Scholar] [CrossRef]
- Pranata, R.; Huang, I.; Lukito, A.A.; Raharjo, S.B. Elevated N-terminal pro-brain natriuretic peptide is associated with increased mortality in patients with COVID-19: Systematic review and meta-analysis. Postgrad. Med. J. 2020, 96, 387–391. [Google Scholar] [CrossRef]
- Kim, C.W.; Aronow, W.S. COVID-19, cardiovascular diseases and cardiac troponins. Future Cardiol. 2022, 18, 135–142. [Google Scholar] [CrossRef]
- Akbar, M.R.; Pranata, R.; Wibowo, A.; Lim, M.A.; Sihite, T.A.; Martha, J.W. The prognostic value of elevated creatine kinase to predict poor outcome in patients with COVID-19—A systematic review and meta-analysis. Diabetes Metab. Syndr. 2021, 15, 529–534. [Google Scholar] [CrossRef]
- Zhu, F.; Li, W.; Lin, Q.; Xu, M.; Du, J.; Li, H. Myoglobin and troponin as prognostic factors in patients with COVID-19 pneumonia. Med. Clin. 2021, 157, 164–171. [Google Scholar] [CrossRef] [PubMed]
- Sozio, E.; Tascini, C.; Fabris, M.; D’Aurizio, F.; De Carlo, C.; Graziano, E.; Bassi, F.; Sbrana, F.; Ripoli, A.; Pagotto, A.; et al. MR-proADM as prognostic factor of outcome in COVID-19 patients. Sci. Rep. 2021, 11, 5121. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Cai, X.; Wang, H.; He, G.; Lin, Y.; Lu, B.; Chen, C.; Pan, Y.; Hu, X. Abnormalities of peripheral blood system in patients with COVID-19 in Wenzhou, China. Clin. Chim. Acta 2020, 507, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Merad, M.; Martin, J.C. Pathological inflammation in patients with COVID-19: A key role for monocytes and macrophages. Nat. Rev. Immunol. 2020, 20, 355–362. [Google Scholar] [CrossRef]
- Tomar, B.; Anders, H.J.; Desai, J.; Mulay, S.R. Neutrophils and Neutrophil Extracellular Traps Drive Necroinflammation in COVID-19. Cells 2020, 9, 1383. [Google Scholar] [CrossRef]
- Lindsley, A.W.; Schwartz, J.T.; Rothenberg, M.E. Eosinophil responses during COVID-19 infections and coronavirus vaccination. J. Allergy Clin. Immunol. 2020, 146, 122–129. [Google Scholar] [CrossRef]
- Li, G.; De Clercq, E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat. Rev. Drug Discov. 2020, 19, 149–150. [Google Scholar] [CrossRef]
- Tu, Y.-F.; Chien, C.-S.; Yarmishyn, A.A.; Lin, Y.-Y.; Luo, Y.-H.; Lin, Y.-T.; Lai, W.-Y.; Yang, D.-M.; Chou, S.-J.; Yang, Y.-P.; et al. A Review of SARS-CoV-2 and the Ongoing Clinical Trials. Int. J. Mol. Sci. 2020, 21, 2657. [Google Scholar] [CrossRef]
- Acharya, D.; Liu, G.; Gack, M.U. Dysregulation of type I interferon responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 397–398. [Google Scholar] [CrossRef]
- Hernandez, A.V.; Roman, Y.M.; Pasupuleti, V.; Barboza, J.J.; White, C.M. Update Alert: Hydroxychloroquine or Chloroquine for the Treatment or Prophylaxis of COVID-19. Ann. Intern. Med. 2020, 173, W78–W79. [Google Scholar] [CrossRef]
- Kupferschmidt, K. Big studies dim hopes for hydroxychloroquine. Science 2020, 368, 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Saitz, R.; Schwitzer, G. Communicating Science in the Time of a Pandemic. JAMA 2020, 324, 443–444. [Google Scholar] [CrossRef] [PubMed]
- Diao, B.; Wang, C.; Tan, Y.; Chen, X.; Liu, Y.; Ning, L.; Chen, L.; Li, M.; Liu, Y.; Wang, G.; et al. Reduction and Functional Exhaustion of T Cells in Patients With Coronavirus Disease 2019 (COVID-19). Front. Immunol. 2020, 11, 827. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.-Y.; Zhang, M.; Yang, C.-X.; Zhang, N.; Wang, X.-C.; Yang, X.-P.; Dong, X.-Q.; Zheng, Y.-T. Elevated exhaustion levels and reduced functional diversity of T cells in peripheral blood may predict severe progression in COVID-19 patients. Cell. Mol. Immunol. 2020, 17, 541–543. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Gao, Y.; Wang, G.; Song, G.; Liu, S.; Sun, D.; Xu, Y.; Tian, Z. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell. Mol. Immunol. 2020, 17, 533–535. [Google Scholar] [CrossRef]
- Braun, J.; Loyal, L.; Frentsch, M.; Wendisch, D.; Georg, P.; Kurth, F.; Hippenstiel, S.; Dingeldey, M.; Kruse, B.; Fauchere, F.; et al. Presence of SARS-CoV-2-reactive T cells in COVID-19 patients and healthy donors. medRxiv 2020. [Google Scholar] [CrossRef]
- Chen, L.; Xiong, J.; Bao, L.; Shi, Y. Convalescent plasma as a potential therapy for COVID-19. Lancet Infect. Dis. 2020, 20, 398–400. [Google Scholar] [CrossRef]
- Shen, C.; Wang, Z.; Zhao, F.; Yang, Y.; Li, J.; Yuan, J.; Wang, F.; Li, D.; Yang, M.; Xing, L.; et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 2020, 323, 1582–1589. [Google Scholar] [CrossRef]
| Antiviral Agents | Mechanism of Action | Reference |
|---|---|---|
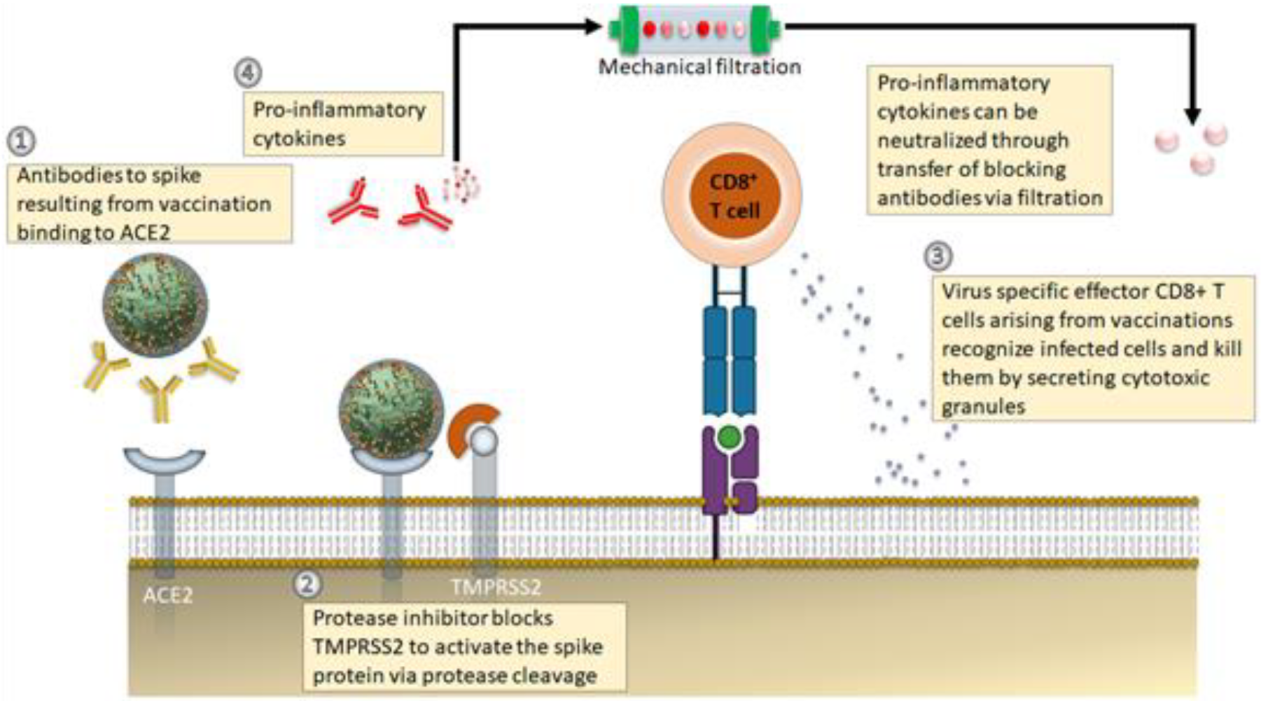
| Lopinavir–ritonavir | It is used as a protease inhibitor, inhibition of 3-chymotrypsin-like protease. | [34] |
| Molnupiravir | It is the first oral antiviral drug, highly effective in the treatment of SARS-CoV-2 infected patients. | [35] |
| Glucocorticoids | It is used for the suppression of immune and inflammatory responses. | [36] |
| Nafamostat | It acts as a serine protease inhibitor that helps in blocking the entry of SARS-CoV viruses. | [37] |
| Chloroquine or hydroxychloroquine | This drug creates interference with ACE2 and blocks virus invasion. It does not allow to increase of endosomal pH, which is required for virus fusion and helps in mild immune suppression. | [38] |
| Thalidomide | This agent has multiple benefits, including reducing inflammatory cell infiltration, relieving cytokine storm, reducing lung damage, and avoiding pulmonary interstitial fibrosis. | [39] |
| Famotidine | It acts as H2 receptor antagonist. | [40] |
| Leflunomide | It helps in the inhibition of virus replication. | [41] |
| Tocilizumab | It acts as a blockade for IL-1 receptors. It also involves downstream signaling pathways. It can reduce mortality and pathology. | [42] |
| Anakinra | It acts as a blockade for IL-1 receptors and involves downstream signaling pathways. | [43] |
| Bevacizumab | It helps in reduces vascular permeability and decreases the amount of fluid that enters the lung tissues. | [44] |
| Umifenovir | It is a small indole-derivative, which has antiviral properties. It can effectively block trimerization of S glycoprotein of SARS-CoV-2 virus. | [45] |
| Baricitinib | It acts as a Janus kinase (JAK) inhibitor; it can blockade viral entry by inhibiting AAK1. It also helps in immune suppression. | [46] |
| Arbidol (Ruxolitinib) | It acts as JAK inhibitor and helps in immune suppression. | [47] |
| Nitazoxanide | It involves blocking the maturation of the nucleocapsid N protein. | [48] |
| Stem cell therapy | It helps in suppressing inflammation and proviral silencing. Decreases inflammation indicators (lowers IL-6 and CRP levels) | [49,50] |
| Convalescent plasma therapy | It promotes the elimination of viruses through specific antibodies. | [51] |
| Neutralizing monoclonal antibodies | Pathogen-specific antibodies involve in antibody effector activity and allow the neutralization of pathogens. | [52] |
| Fc-engineered antibody | Application of monoclonal antibodies optimized Fc domains with the optimal Fc-effector function and better clinical effectiveness against the SARS-CoV-2 infection. | [53] |
| Pathway | Biomarkers | Comment | Reference |
|---|---|---|---|
| Hematological | White blood cells (WBC) count | Elevated levels. WBC count is significantly correlated with severity in SARS-CoV-2 infected patients. Higher levels of WBC suggest the need for more attention during the treatment of SARS-CoV-2 infected patients. | [170] |
| Lymphocyte count (LC) | Decreased levels. Recent findings show that absolute lymphocyte count remains lower in severe SARS-CoV-2 infected patients. | [171] | |
| Neutrophil count (NC) | Elevated levels. Increased levels of circulating neutrophils are characteristically realized in SARS-CoV-2 infected patients. | [172] | |
| Monocyte-to-lymphocyte ratio (MLR) | Elevated levels. Elevated MLR and higher mortality among SARS-CoV-2 infected patients. | [173] | |
| Neutrophils-to-lymphocyte ratio (NLR) | Elevated levels. NLR at the initial stage is lower and has a significant prognostic value in SARS-CoV-2 infected patients as compared with influenza and syncytial virus. | [174] | |
| Mean platelet volume (MPV) | Elevated levels. MPV found significantly higher levels in severe patients compared to mild patients. | [175] | |
| Red cell distribution width (RDW) | Elevated levels. RDW levels can be a potent predictor for the severity of SARS-CoV-2 infected patients and the probability of mortality. | ||
| Monocyte distribution width (MDW) | Elevated levels. MDW can be performed using cell count analyzers and can be reliable marker of sepsis for early-stage diagnosis. | [176] | |
| Lymphocyte subsets (neutrophilia, lymphopenia, T-helper (CD4+) and T-cytotoxic (CD8+)) | Lymphocyte subsets have a correlation with the severity and outcome of the disease in SARS-CoV-2 infected patients. | [177] | |
| Inflammation | Elevated IL-2, IL-6, IL-8, IL-10 levels | Elevated interleukin levels are highly associated with severe SARS-CoV-2 infected patients. | [178] |
| Procalcitonin (PCT) level | Elevated levels. Serial measurement of PCT levels help to predict the prognosis of SARS-CoV-2 infected patients. | [179] | |
| Serum amyloid A (SAA) | Elevated levels. Significantly higher SAA levels in SARS-CoV-2 infected patients with non-survivors and severe disease. | [180] | |
| Erythrocyte sedimentation rate (ESR) | Elevated levels. Significantly higher ESR levels were found in severe patients than non-severe patients. | [181] | |
| C-reactive protein (CRP) | Elevated levels. High serum levels of CRP contribute to the SARS-CoV-2 infection progression. | [182] | |
| Ferritin | Elevated levels. Ferritin is iron-containing an intracellular blood protein. | [183] | |
| Sphingosine-1-phosphate | Decreased levels. A decrease in serum sphingosine-1-phosphate levels can be a predictor of severity in SARS-CoV-2 infected patients. | [184] | |
| Coagulation | Elevated fibrin/fibrinogen (FIB) degradation factor | Elevated fibrinogen (FIB) can be one of the important indicator as fibrinolysis abnormalities and coagulation among SARS-CoV-2 infected patients. | [185] |
| D-dimer levels | Elevated level of D-dimer is one of the important measure can be used to detect thrombosis among SARS-CoV-2 infected patients. | [186] | |
| Prothrombin time (PT) | The admission prothrombin time (PT) significantly higher among non-survivor SARS-CoV-2 infected patients. | [187] | |
| Prothrombin time activity (PT-act) | Prothrombin time activity (PT-act) higher among non-survivor SARS-CoV-2 infected patients. | [185] | |
| Partial thromboplastin time (aPTT) | Anticoagulants based therapy required in SARS-CoV-2 infected patients for aPTT prolongation. | [188] | |
| PT-international normalized ratio (INR) | INR in SARS-CoV-2 infected patients were found higher than healthy individuals and were also found higher in the patients with thrombotic disease than that of without this disease. | [189] | |
| Liver fibrosis index (FIB-4) | This is one of the important test widely used to detect a stage of fibrosis and to monitor chronic liver diseases, such as chronic viral hepatitis, HCV/HIV co-infection, as well as metabolic-associated fatty liver disease. | [190] | |
| Necrosis | Lactate dehydrogenase (LDH) | Elevated levels. LDH is a marker of various inflammatory states, e.g., infections including SARS-CoV-2 infection, myocardial infractions, and malignancies. | [191] |
| Liver injury | Alanine transferase (ALT) | Elevated levels. Evidence of liver injury in SARS-CoV-2 infected patients with abnormal ALT levels is based on the severity of the disease. | [192] |
| Aspartate transaminase (AST) | A high AST/ALT ratio can indicate a worse state and mortality among SARS-CoV-2 infected patients. | [193] | |
| Serum total bilirubin (STB) | Elevated levels. SARS-CoV-2 infected patients with elevated STB levels had higher mortality. | [194] | |
| Unconjugated bilirubin (UCB) | Elevated UCB levels SARS-CoV-2 infected patients can predict the severity of disease and probability of mortality. | [194] | |
| Gamma-glutamyl transferase (GGT) | Elevated GGT levels indicate the severity and are positively correlated with a prolonged hospital stay. | [195] | |
| Cardiac injury | D-dimer | Early-stage elevated levels of D-dimer levels among SARS-CoV-2 infected patients indicate pulmonary intravascular coagulation. | [195] |
| Homocysteine | Elevated homocysteine levels are a potential predictor of cardiovascular risk in SARS-CoV-2 infected patients. | [196] | |
| N-terminal pro-brain natriuretic peptide (NT-pro-BNP) | Elevated NT-proBNP levels is associated with high mortality risk in SARS-CoV-2 infected patients even with no history of heart failure. | [197] | |
| Serum cardiac troponin (cTn) | Elevated cTn is a strong indication of myocardial injury in SARS-CoV-2 infected patients. | [198] | |
| Muscular injury | Creatine kinase | Elevated CK levels are a potential biomarker of muscle damage, possibly correlated with more severe SARS-CoV-2 infection. | [199] |
| Myoglobin | Myoglobin is oxygen and iron-binding protein that enables oxygen storage in cardiac and skeletal muscles. Significantly elevated levels of myoglobin were observed in SARS-CoV-2 infection with severe and death groups. | [200] | |
| Renal | Serum creatinine | Serum creatinine levels were significantly higher in the severe and mortality groups than recovery groups. | [201] |
| Organ failure | MR-pro-ADM | Elevated MR-proADM can be an emergent prognostic factor in evaluates deteriorating state SARS-CoV-2 infected patients. | [202] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saratale, G.D.; Shin, H.-S.; Shinde, S.K.; Kim, D.-Y.; Saratale, R.G.; Kadam, A.A.; Kumar, M.; Bahkali, A.H.; Syed, A.; Ghodake, G.S. Significance of Immune Status of SARS-CoV-2 Infected Patients in Determining the Efficacy of Therapeutic Interventions. J. Pers. Med. 2022, 12, 349. https://doi.org/10.3390/jpm12030349
Saratale GD, Shin H-S, Shinde SK, Kim D-Y, Saratale RG, Kadam AA, Kumar M, Bahkali AH, Syed A, Ghodake GS. Significance of Immune Status of SARS-CoV-2 Infected Patients in Determining the Efficacy of Therapeutic Interventions. Journal of Personalized Medicine. 2022; 12(3):349. https://doi.org/10.3390/jpm12030349
Chicago/Turabian StyleSaratale, Ganesh Dattatraya, Han-Seung Shin, Surendra Krushna Shinde, Dae-Young Kim, Rijuta Ganesh Saratale, Avinash Ashok Kadam, Manu Kumar, Ali Hassan Bahkali, Asad Syed, and Gajanan Sampatrao Ghodake. 2022. "Significance of Immune Status of SARS-CoV-2 Infected Patients in Determining the Efficacy of Therapeutic Interventions" Journal of Personalized Medicine 12, no. 3: 349. https://doi.org/10.3390/jpm12030349
APA StyleSaratale, G. D., Shin, H.-S., Shinde, S. K., Kim, D.-Y., Saratale, R. G., Kadam, A. A., Kumar, M., Bahkali, A. H., Syed, A., & Ghodake, G. S. (2022). Significance of Immune Status of SARS-CoV-2 Infected Patients in Determining the Efficacy of Therapeutic Interventions. Journal of Personalized Medicine, 12(3), 349. https://doi.org/10.3390/jpm12030349









