Development of a Highly Sensitive Technique for Capturing Renal Cell Cancer Circulating Tumor Cells
Abstract
1. Introduction
2. Materials and Methods
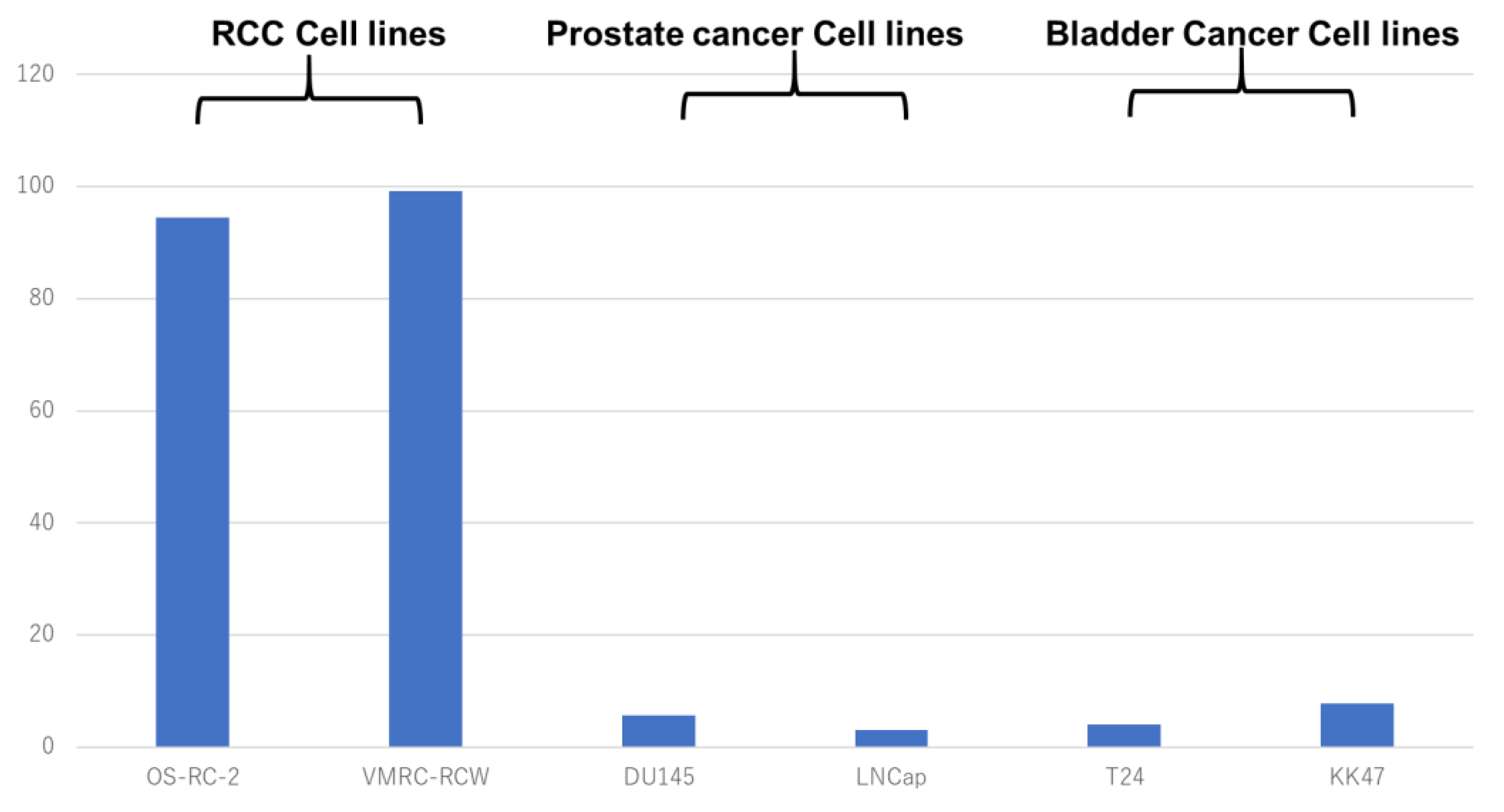
2.1. Stainability of Anti-G250 Antibody in Various Cancer Cell Lines
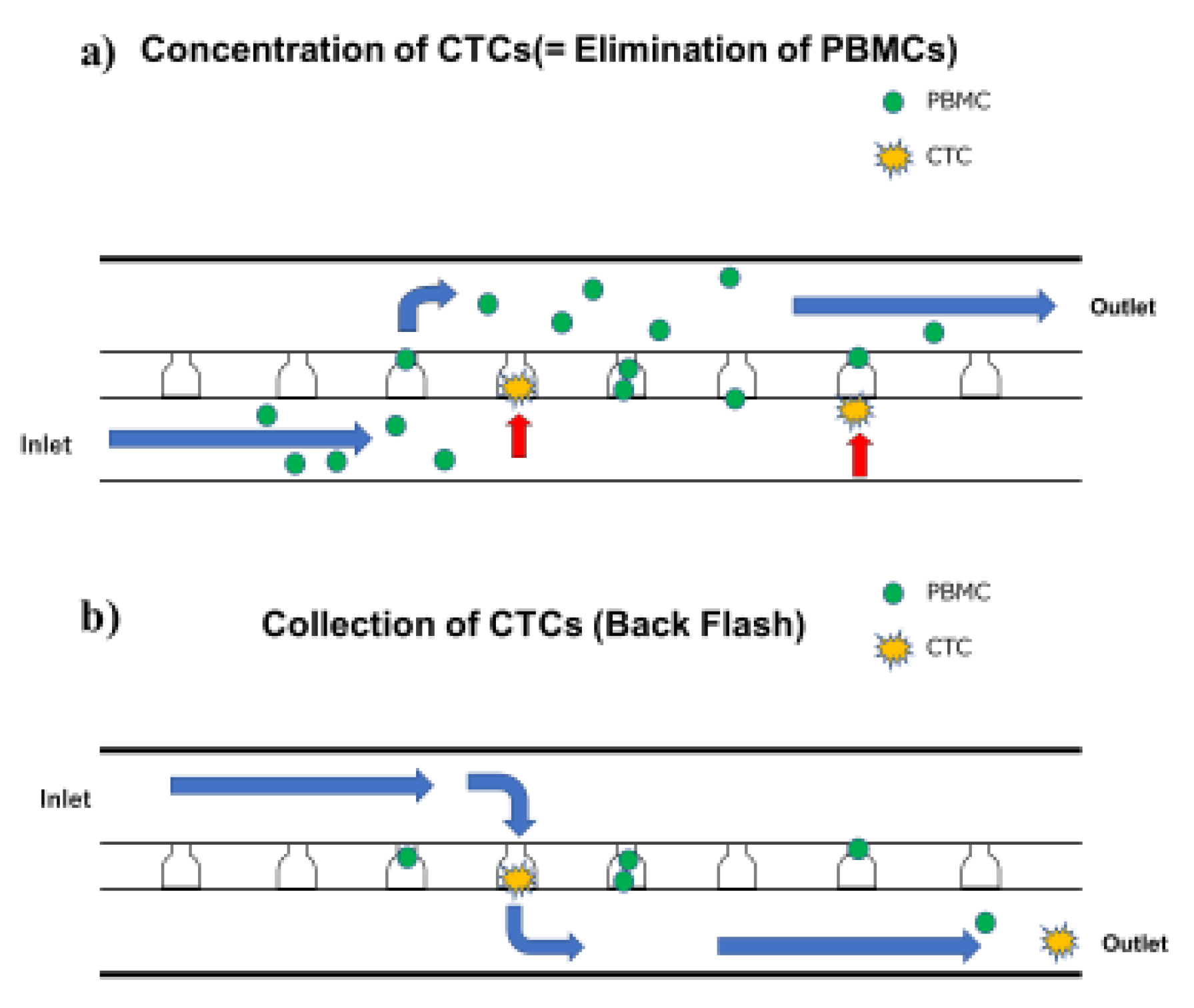
2.2. Concentration and Analysis of CTC (Using Celsee® Combined with On-chip Sort®)
2.3. Cell Retrieval Using Celsee PREP100® Combined with On-Chip Sort®
2.4. Patient Samples
3. Results
3.1. Anti-G250 Antibody Staining Pattern in Various Cancer Cell Lines
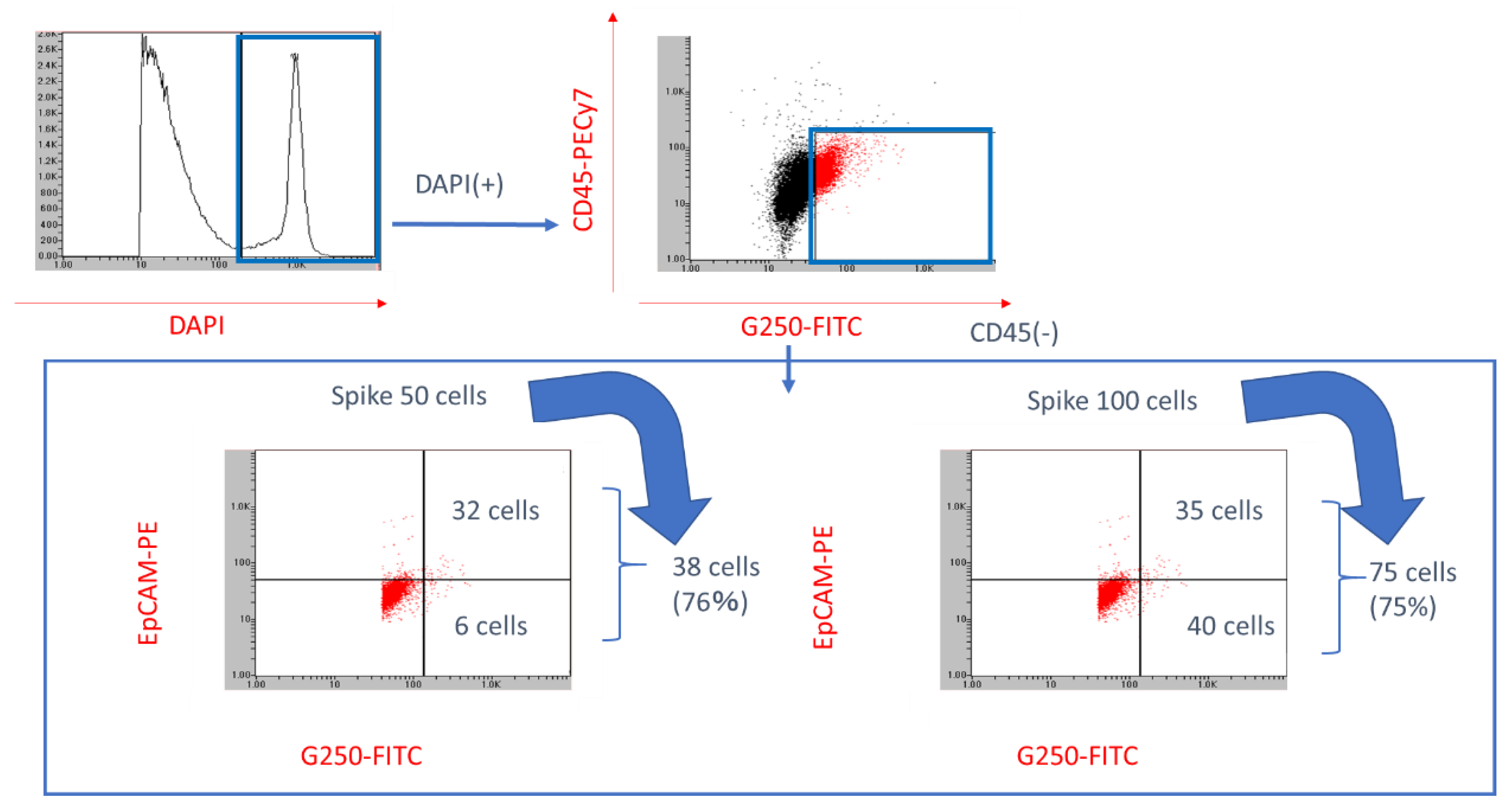
3.2. Identification Rate of RCC CTC Using On-Chip Sort® with G250 Antibody
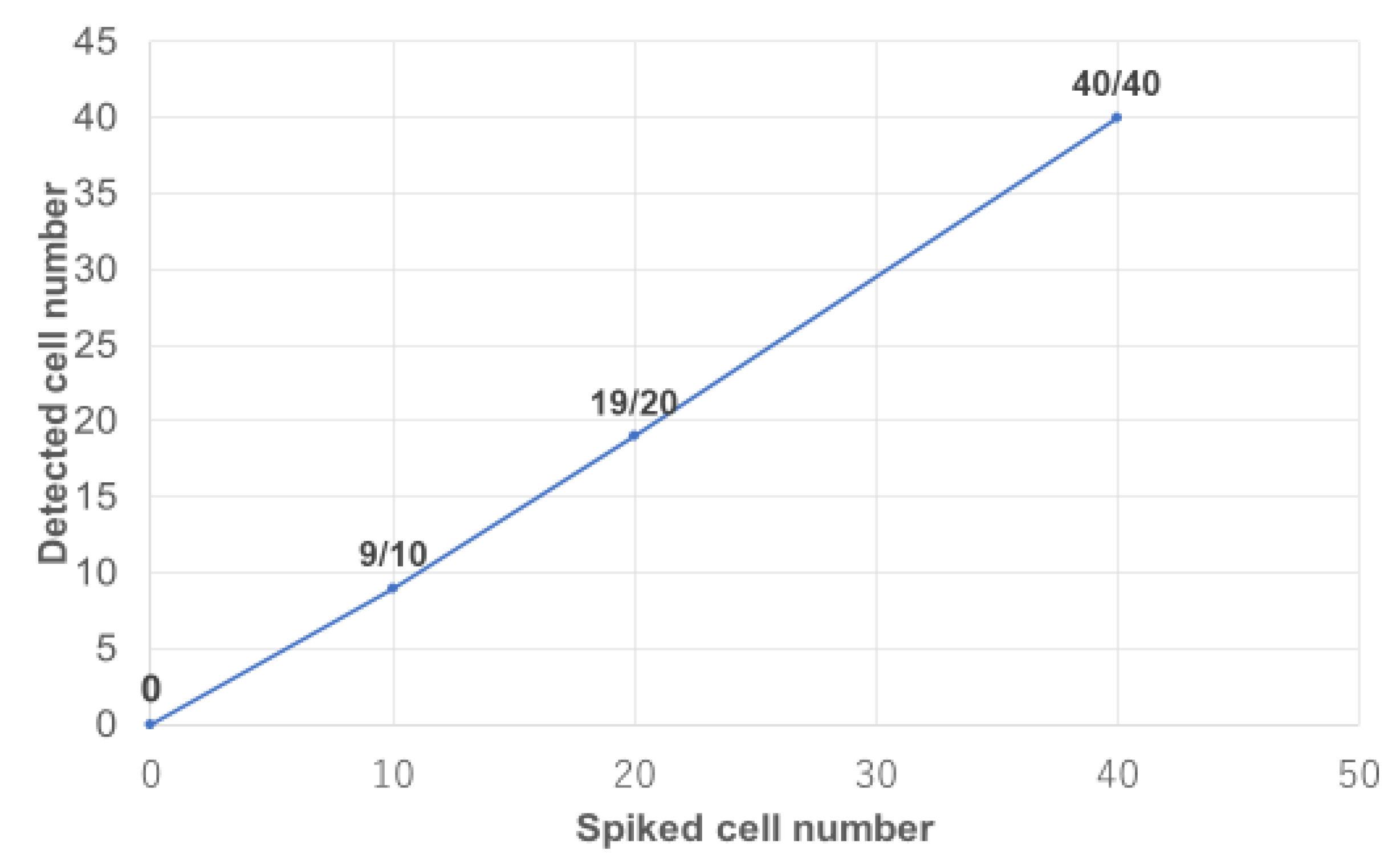
3.3. RCC Cells Concentration Using CelSee® and Spiked RCC Cells Counting Using On-Chip Sort® with Anti-G250 Antibody
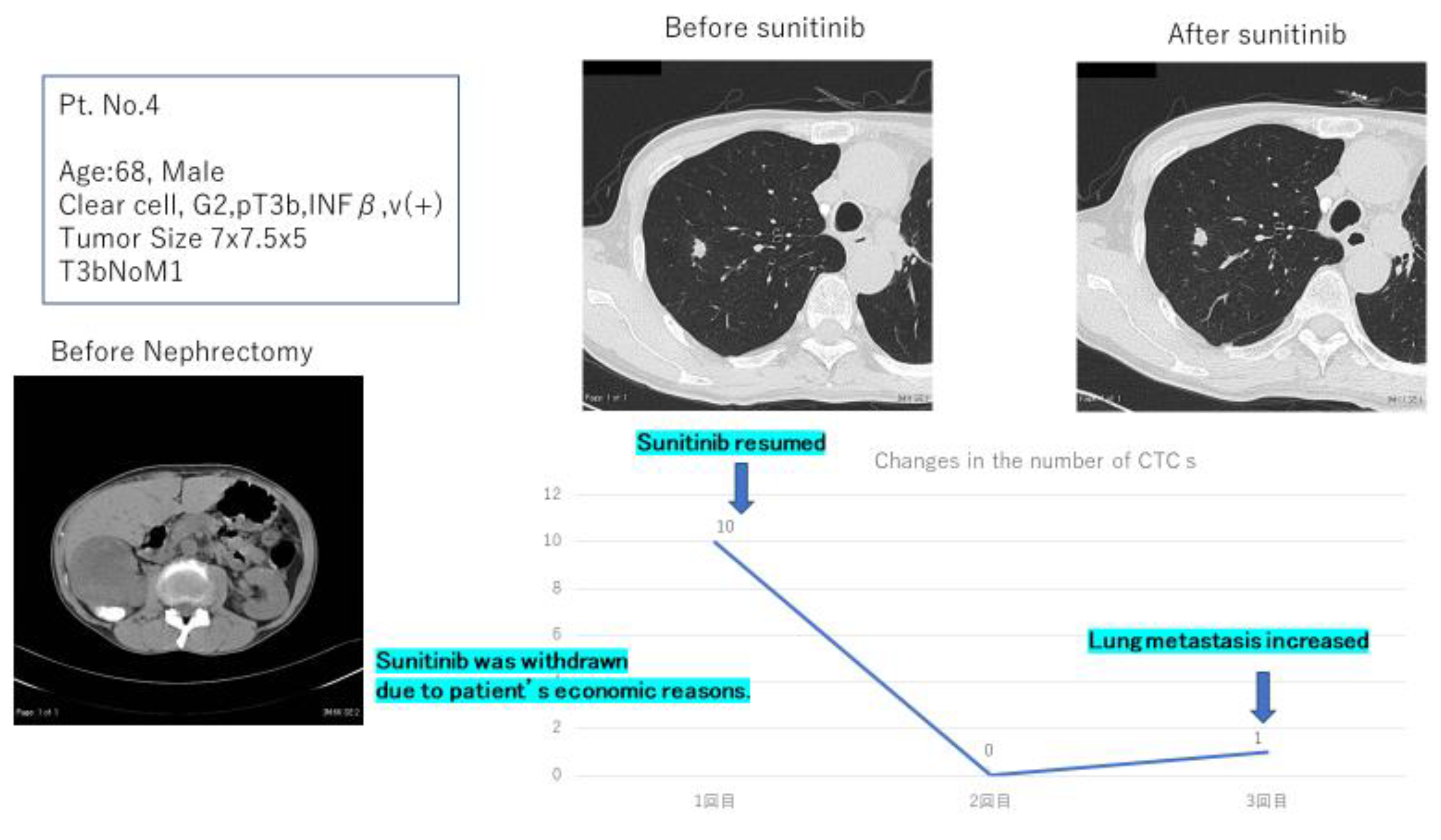
3.4. Pilot CTC Examination in Patients with Metastatic RCC
4. Discussion
4.1. History of CTC Identification
4.2. Regarding the Weak Point of CellSearch System®
4.3. Microfluidic Chip of CelSee®
4.4. On-chip Sort®
4.5. G250 antigen
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Andree, K.C.; van Dalum, G.; Terstappen, L.W. Challenges in circulating tumor cell detection by the CellSearch system. Mol. Oncol. 2016, 10, 395–407. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Hayes, D.F.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Reuben, J.M.; Doyle, G.V.; Matera, J.; Allard, W.J. Circulating tumor cells: A novel prognostic factor for newly diagnosed metastatic breast cancer. J. Clin. Oncol. 2005, 23, 1420–1430. [Google Scholar] [CrossRef] [PubMed]
- Spizzo, G.; Fong, D.; Wurm, M.; Ensinger, C.; Obrist, P.; Hofer, C.; Mazzoleni, G.; Gastl, G.; Went, P. EpCAM expression in primary tumour tissues and metastases: An immunohistochemical analysis. J. Clin. Pathol. 2011, 64, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Baeuerle, P.A.; Gires, O. EpCAM (CD326) finding its role in cancer. Br. J. Cancer 2007, 96, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Van der Gun, B.T.; Melchers, L.J.; Ruiters, M.H.; de Leij, L.F.; McLaughlin, P.M.; Rots, M.G. EpCAM in carcinogenesis: The good, the bad or the ugly. Carcinogenesis 2010, 31, 1913–1921. [Google Scholar] [CrossRef] [PubMed]
- Riahi, R.; Gogoi, P.; Sepehri, S.; Zhou, Y.; Handique, K.; Godsey, J.; Wang, Y. A novel microchannel-based device to capture and analyze circulating tumor cells (CTCs) of breast cancer. Int. J. Oncol. 2014, 44, 1870–1878. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Uehara, Y.; Yamashita, N.; Fujimura, Y.; Nishio, K.; Sawada, T.; Takeda, K.; Koizumi, F.; Koh, Y. Multicolor detection of rare tumor cells in blood using a novel flow cytometry-based system. Cytom. Part A 2014, 85, 206–213. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.; Gumbleton, M.; Griffiths, D.F. Caveolin-1 overexpression predicts poor disease-free survival of patients with clinically confined renal cell carcinoma. Br. J. Cancer 2003, 89, 1909–1913. [Google Scholar] [CrossRef] [PubMed]
- Lam, J.S.; Klatte, T.; Kim, H.L.; Patard, J.J.; Breda, A.; Zisman, A.; Pantuck, A.J.; Figlin, R.A. Prognostic factors and selection for clinical studies of patients with kidney cancer. Crit. Rev. Oncol. Hematol. 2008, 65, 235–262. [Google Scholar] [CrossRef] [PubMed]
- Oosterwijk, E.; Ruiter, D.J.; Hoedemaeker, P.J.; Pauwels, E.K.; Jonas, U.; Zwartendijk, J.; Warnaar, S.O. Monoclonal antibody G 250 recognizes a determinant present in renal-cell carcinoma and absent from normal kidney. Int. J. Cancer 1986, 38, 489–494. [Google Scholar] [CrossRef] [PubMed]
- Oosterwijk, E.; Ruiter, D.J.; Wakka, J.C.; Huiskens-van der Meij, J.W.; Jonas, U.; Fleuren, G.J.; Zwartendijk, J.; Hoedemaeker, P.; Warnaar, S.O. Immunohistochemical analysis of monoclonal antibodies to renal antigens. Application in the diagnosis of renal cell carcinoma. Am. J. Pathol. 1986, 123, 301–309. [Google Scholar] [PubMed]
- Uemura, H.; Nakagawa, Y.; Yoshida, K.; Saga, S.; Yoshikawa, K.; Hirao, Y.; Oosterwijk, E. MN/CA IX/G250 as a potential target for immunotherapy of renal cell carcinomas. Br. J. Cancer. 1999, 81, 741–746. [Google Scholar] [CrossRef] [PubMed]
- Cairns, P. Renal cell carcinoma. Cancer Biomark. 2010, 9, 461–473. [Google Scholar] [CrossRef] [PubMed]
- Takacova, M.; Bartosova, M.; Skvarkova, L.; Zatovicova, M.; Vidlickova, I.; Csaderova, L.; Barathova, M.; Breza, J., Jr.; Bujdak, P.; Pastorek, J.; et al. Carbonic anhydrase IX is a clinically significant tissue and serum biomarker associated with renal cell carcinoma. Oncol. Lett. 2013, 5, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Bluemke, K.; Bilkenroth, U.; Meye, A.; Fuessel, S.; Lautenschlaeger, C.; Goebel, S.; Melchior, A.; Heynemann, H.; Fornara, P.; Taubert, H. Detection of circulating tumor cells in peripheral blood of patients with renal cell carcinoma correlates with prognosis. Cancer. Epidemiol. Biomarkers. Prev. 2009, 18, 2190–2194. [Google Scholar] [CrossRef] [PubMed]
- Small, A.C.; Gong, Y.; Oh, W.K.; Hall, S.J.; van Rijn, C.J.; Galsky, M.D. The emerging role of circulating tumor cell detection in genitourinary cancer. J. Urol. 2012, 188, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Myung, J.H.; Tam, K.A.; Park, S.J.; Cha, A.; Hong, S. Recent advances in nanotechnology-based detection and separation of circulating tumor cells. Wiley. Interdiscip. Rev. Nanomed. Nanobiotechnol. 2016, 8, 223–239. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabières, C.; Pantel, K. Technologies for detection of circulating tumor cells: Facts and vision. Lab Chip 2014, 14, 57–62. [Google Scholar] [CrossRef] [PubMed]
| Pt.No | Sex | Age | Operation | Pathological Result | Tumor Size (cm) | TNM Classification | Number of CTC/4 mL |
|---|---|---|---|---|---|---|---|
| 1 | F | 65 | Total | clear cell, G2, INFα, pT1b | 4.5 × 4 × 3 | T1bN0M0 | 1 |
| 2 | M | 68 | Partial | papillary, G1, pT1a | 1.4 × 0.8 | T1aN0M0 | 2 |
| 3 | M | 57 | Total | clear cell, G2, INFβ, pT3a | 9 × 6.9 | T3aN0M0 | 3 |
| 4 | M | 68 | Total | clear cell, G2, pT3b, INFβ, v (+) | 7 × 7.5 × 5 | T3bN0M1 | 10 |
| 5 | M | 61 | Total | clear cell, G2>G3, pT3a | 3.7 × 5.7 | T3aN0M0 | 1 |
| 6 | M | 70 | Nil | unknown | 9.5 × 8.1 | T3cN0M1 | 0 |
| 7 | M | 64 | Nil | unknown | 4.6 | T3bN2M1 | 1 |
| 8 | M | 59 | Total | clear cell, G2>G3, pT3a, v (−) | 9.5 × 9.5 | T1bN0M0 | 3 |
| 9 | M | 71 | Nil | unknown | 2.0 | T1aN0M0 | 2 |
| 10 | M | 74 | Nil | unknown | 17 × 13 × 12 | T2bN0M0 | 0 |
| 11 | M | 74 | Total | clear cell, G2, pT1a, v (+) | 6.1 × 4.4 | T1aN0M0 | 0 |
| 12 | M | 54 | Total | clear cell, G2, pT1a, v (+) | 15 × 10 | T1aN0M1 | 3 |
| 13 | M | 58 | Nil | unknown | 20 × 17 × 15 | T2bN0M1 | 0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naoe, M.; Kusaka, C.; Ohta, M.; Hasebe, Y.; Unoki, T.; Shimoyama, H.; Nakasato, T.; Oshinomi, K.; Morita, J.; Fuji, K.; et al. Development of a Highly Sensitive Technique for Capturing Renal Cell Cancer Circulating Tumor Cells. Diagnostics 2019, 9, 96. https://doi.org/10.3390/diagnostics9030096
Naoe M, Kusaka C, Ohta M, Hasebe Y, Unoki T, Shimoyama H, Nakasato T, Oshinomi K, Morita J, Fuji K, et al. Development of a Highly Sensitive Technique for Capturing Renal Cell Cancer Circulating Tumor Cells. Diagnostics. 2019; 9(3):96. https://doi.org/10.3390/diagnostics9030096
Chicago/Turabian StyleNaoe, Michio, Chiho Kusaka, Mika Ohta, Yuki Hasebe, Tsutomu Unoki, Hideaki Shimoyama, Takehiko Nakasato, Kazuhiko Oshinomi, Jun Morita, Kohzo Fuji, and et al. 2019. "Development of a Highly Sensitive Technique for Capturing Renal Cell Cancer Circulating Tumor Cells" Diagnostics 9, no. 3: 96. https://doi.org/10.3390/diagnostics9030096
APA StyleNaoe, M., Kusaka, C., Ohta, M., Hasebe, Y., Unoki, T., Shimoyama, H., Nakasato, T., Oshinomi, K., Morita, J., Fuji, K., Ogawa, Y., Tsukada, M., Sunagawa, M., & Ishii, H. (2019). Development of a Highly Sensitive Technique for Capturing Renal Cell Cancer Circulating Tumor Cells. Diagnostics, 9(3), 96. https://doi.org/10.3390/diagnostics9030096






