HAMP Downregulation Contributes to Aggressive Hepatocellular Carcinoma via Mechanism Mediated by Cyclin4-Dependent Kinase-1/STAT3 Pathway
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Samples
2.2. Cell Culture
2.3. Western Blotting
2.4. HAMP shRNA Transfection
2.5. Quantitative RT-PCR
2.6. Tumor Colony Forming Assay
2.7. 5-Ethynyl-20-deoxyuridine (EdU) Incorporation Assay
2.8. Wound-Healing Assay
2.9. Cell Cycle Analysis
2.10. Animal Treatment
2.11. Statistical Analysis
3. Results
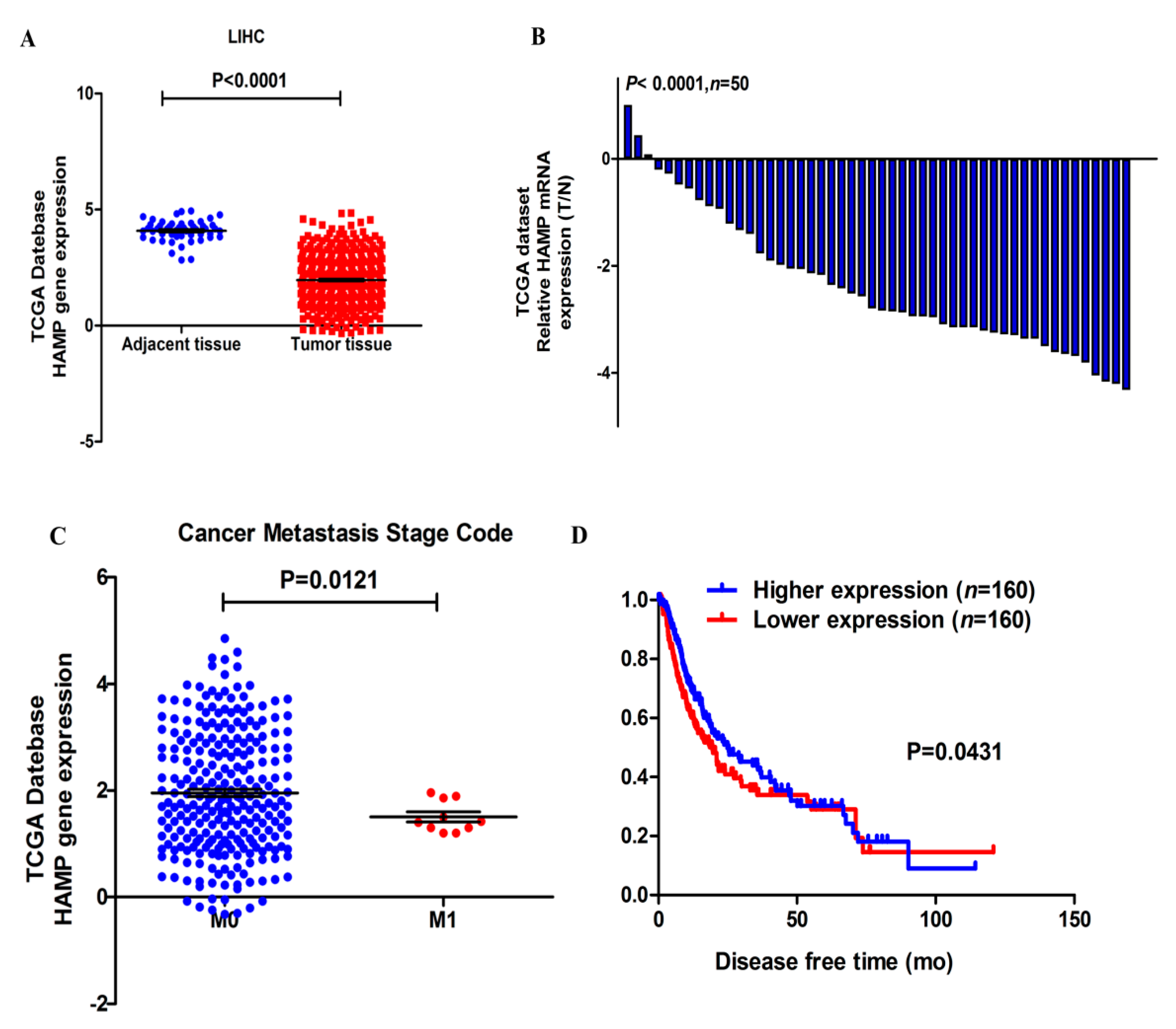
3.1. HAMP Expression Is Significantly Suppressed in Hepatocellular Carcinoma
3.2. Lower HAMP Expression Is Associated with Higher Cancer Metastasis Rate and Recurrence
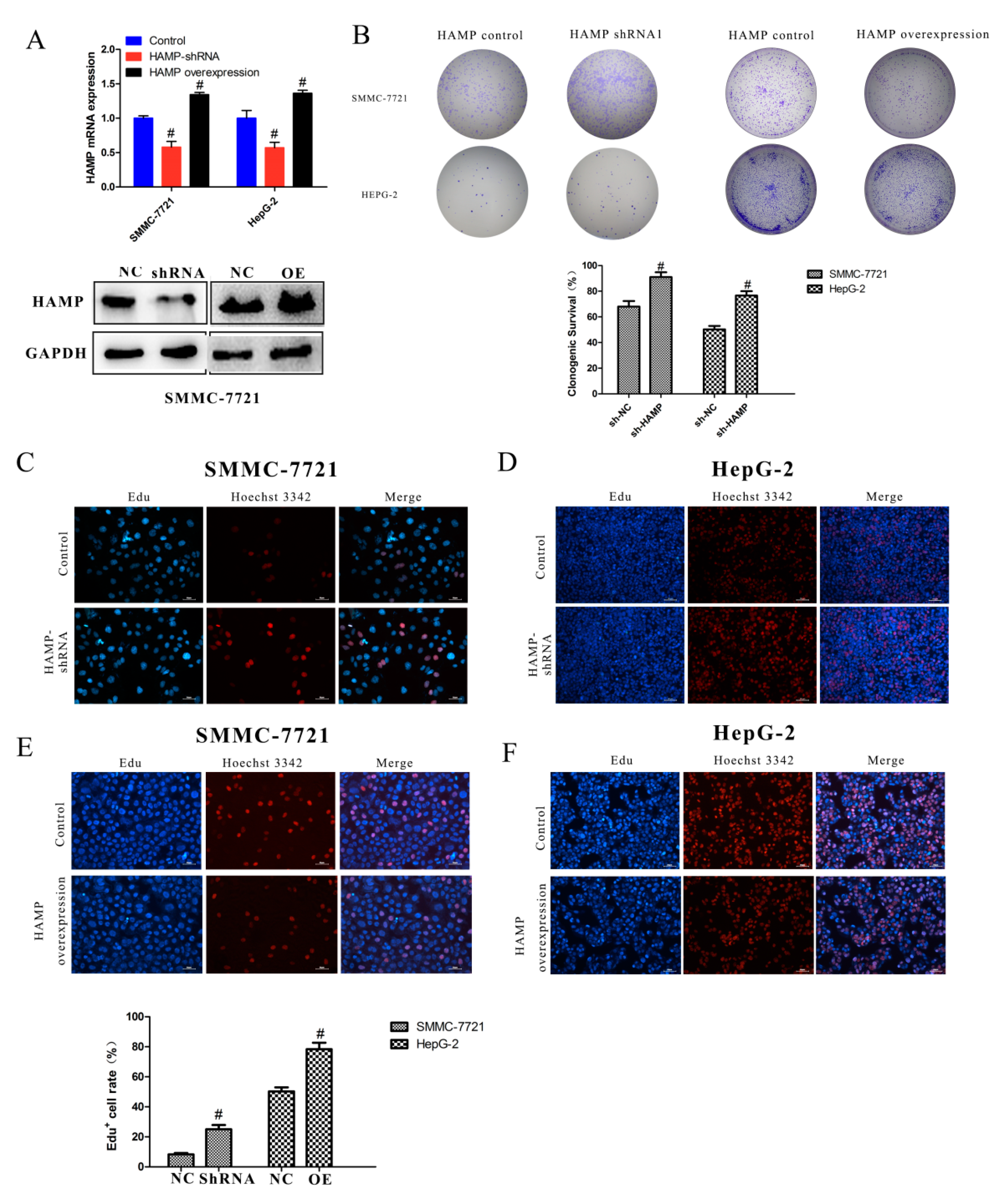
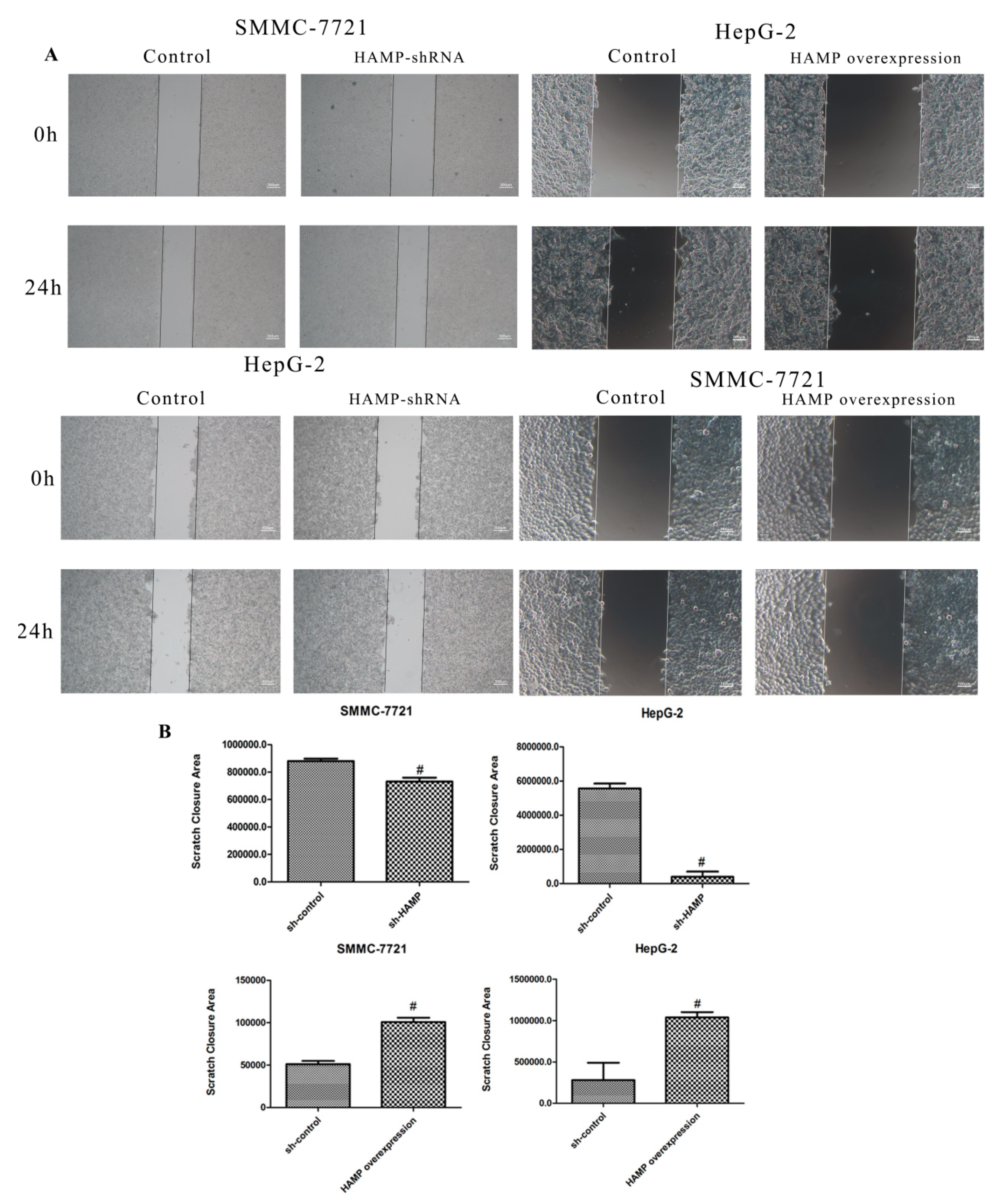
3.3. Low Expression of HAMP Results in Increased Proliferation and Migration in SMMC-7721 and HepG-2 Cells
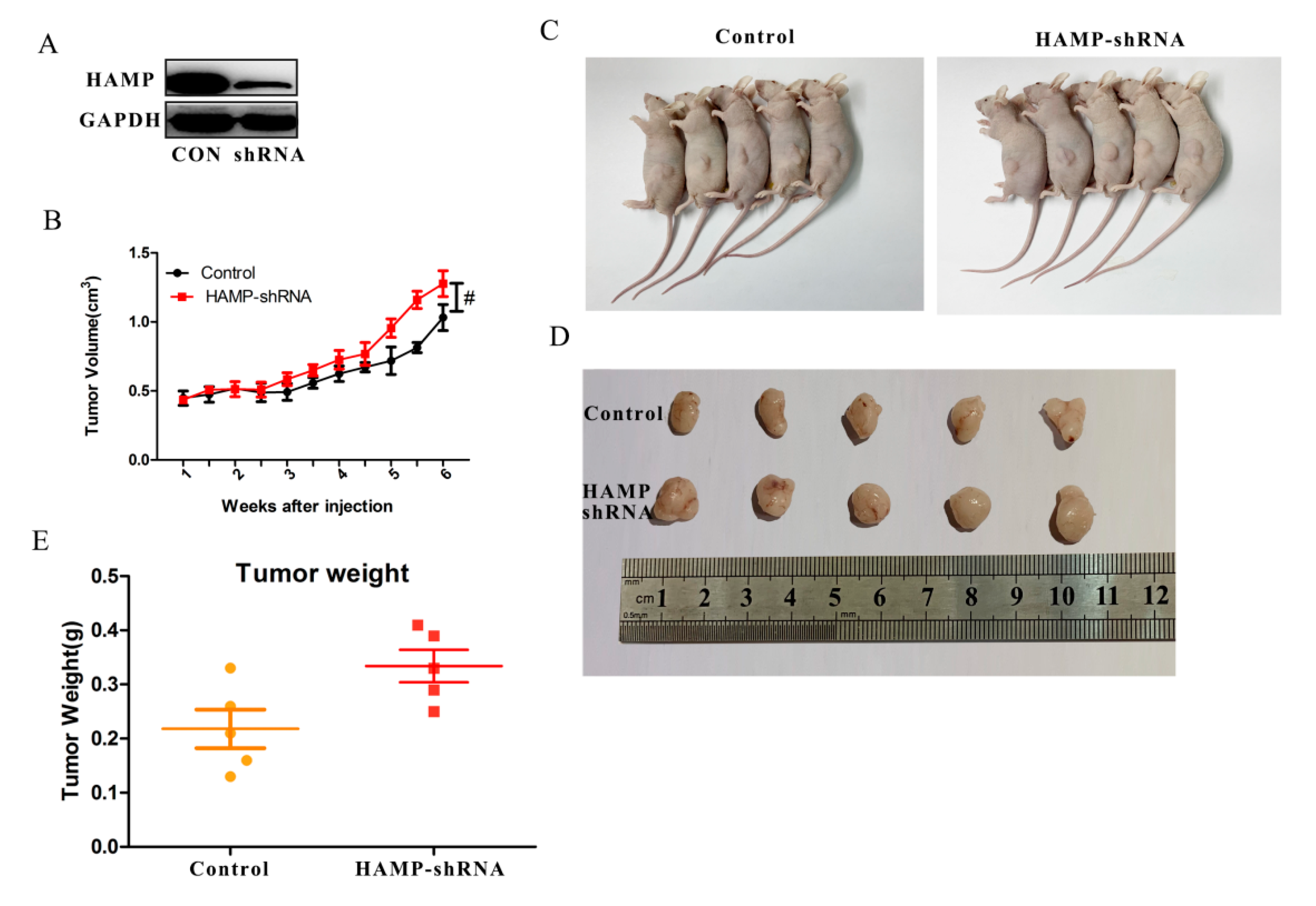
3.4. Low HAMP Expression Promotes Hepatocellular Carcinoma Proliferation In Vivo
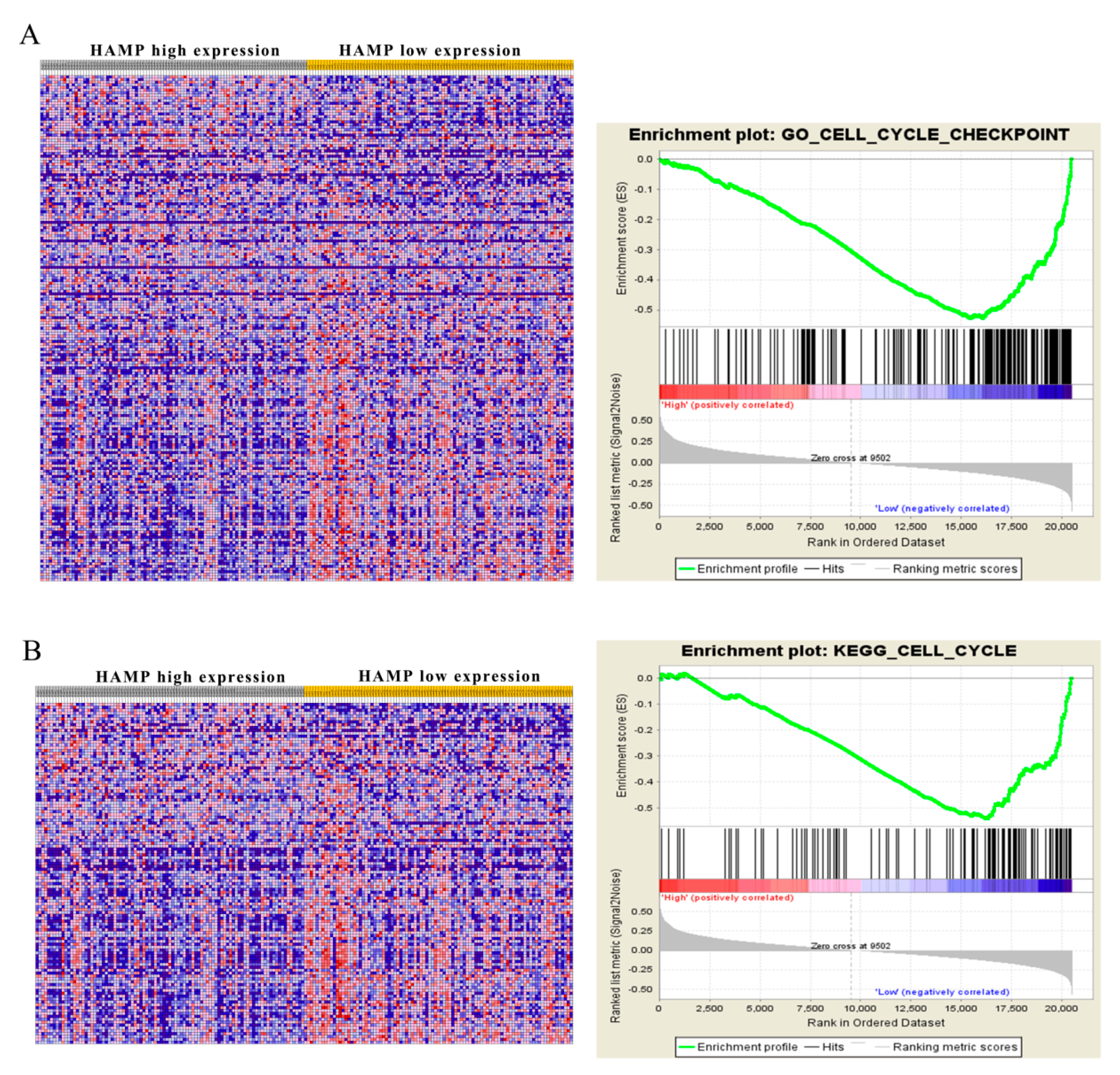
3.5. Low HAMP Expression Is Associated with Cell Cycle Checkpoint in Gene Set Enrichment Analysis (GSEA) of the TCGA
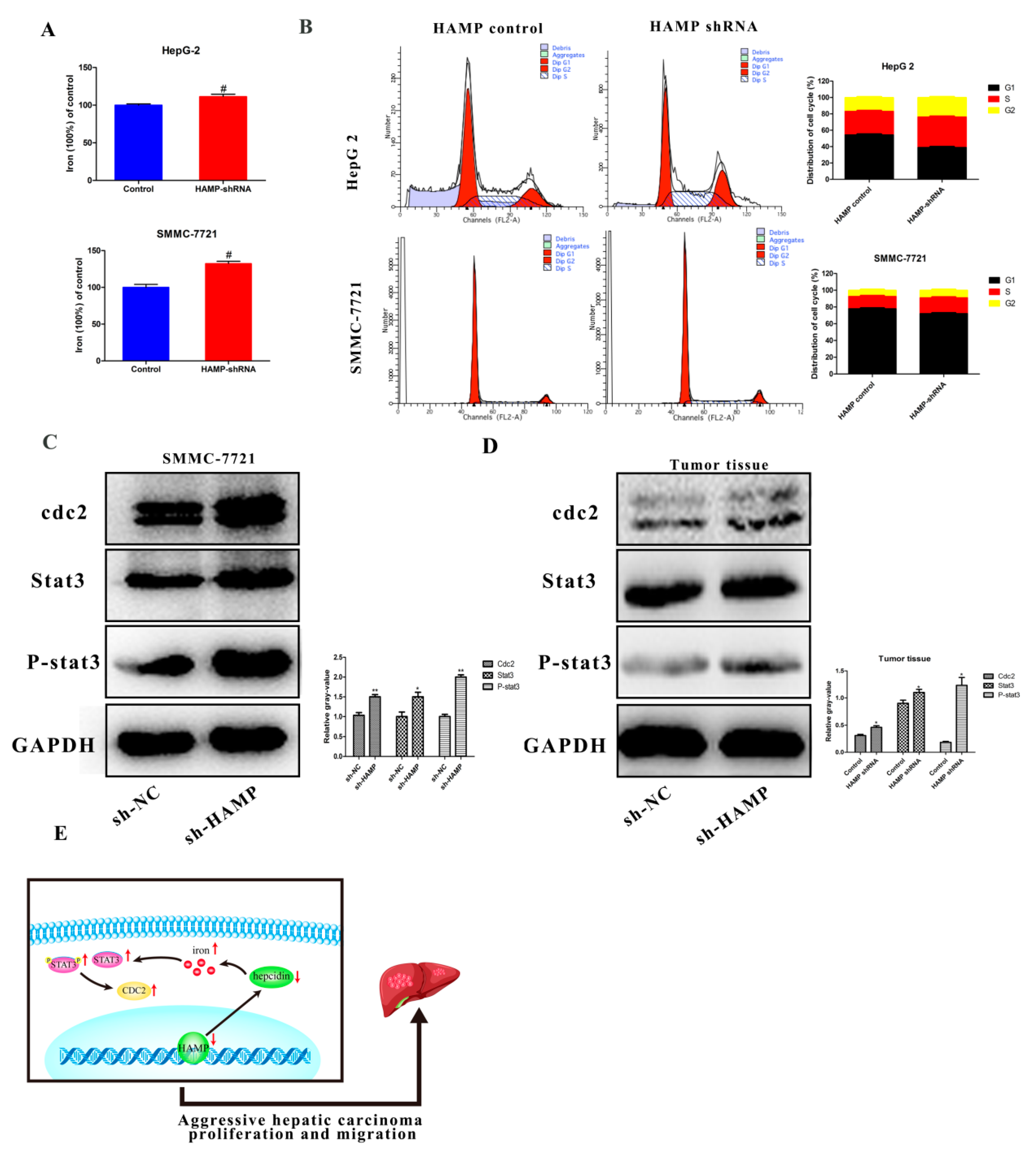
3.6. HAMP Suppression Results in High Cellular Iron Concentration in Cells and Activated CDK1/STAT3 Pathway
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CDK1 | Cyclin-dependent kinases 1 |
| FPN | Ferroportin |
| GSEA | Gene Set Enrichment Analysis |
| HAMP | Hepcidin |
| HCC | Hepatocellular carcinoma |
| SLC40A1 | Solute carrier family 40 member 1 |
| STAT3 | Signal transducer and activator of transcription 3 |
| TCGA | The Cancer Genome Atlas |
References
- Sia, D.; Villanueva, A.; Friedman, S.L.; Llovet, J.M. Liver Cancer Cell of Origin, Molecular Class, and Effects on Patient Prognosis. Gastroenterology 2017, 152, 745–761. [Google Scholar] [CrossRef]
- Lin, M.V.; King, L.Y.; Chung, R.T. Hepatitis C virus-associated cancer. Annu. Rev. Pathol. 2015, 10, 345–370. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Villanueva, A.; Lachenmayer, A.; Finn, R.S. Advances in targeted therapies for hepatocellular carcinoma in the genomic era. Nat. Rev. Clin. Oncol. 2015, 12, 408–424. [Google Scholar] [CrossRef]
- Talwalkar, J.A.; Gores, G.J. Diagnosis and staging of hepatocellular carcinoma. Gastroenterology 2004, 127, S126–S132. [Google Scholar] [CrossRef]
- Kim, C.K.; He, P.; Bialkowska, A.B.; Yang, V.W. SP and KLF Transcription Factors in Digestive Physiology and Diseases. Gastroenterology 2017, 152, 1845–1875. [Google Scholar] [CrossRef]
- Shimano, H.; Sato, R. SREBP-regulated lipid metabolism: Convergent physiology—Divergent pathophysiology. Nat. Rev. Endocrinol. 2017, 13, 710–730. [Google Scholar] [CrossRef] [PubMed]
- Siebel, C.; Lendahl, U. Notch Signaling in Development, Tissue Homeostasis, and Disease. Physiol. Rev. 2017, 97, 1235–1294. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, J.; Yuan, L.; Lay, Y.; Wong, Y.K.; Lim, T.K.; Ong, C.S.; Lin, Q.; Wang, J.; Hua, Z. Andrographolide Suppresses MV4-11 Cell Proliferation through the Inhibition of FLT3 Signaling, Fatty Acid Synthesis and Cellular Iron Uptake. Molecules 2017, 22, 1444. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Manz, D.H.; Torti, S.V.; Torti, F.M. Effects of Ferroportin-Mediated Iron Depletion in Cells Representative of Different Histological Subtypes of Prostate Cancer. Antioxid. Redox Signal. 2019, 30, 1043–1061. [Google Scholar] [CrossRef]
- Harjes, U. Cancer: Ironing it out. Nat. Rev. Drug Discov. 2017, 16, 602. [Google Scholar] [CrossRef]
- Regenboog, M.; Bohte, A.E.; Akkerman, E.M.; Stoker, J.; Hollak, C. Iron storage in liver, bone marrow and splenic Gaucheroma reflects residual disease in type 1 Gaucher disease patients on treatment. Br. J. Haematol. 2017, 179, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Funauchi, Y.; Tanikawa, C.; Yi Lo, P.H.; Mori, J.; Daigo, Y.; Takano, A.; Miyagi, Y.; Okawa, A.; Nakamura, Y.; Matsuda, K. Regulation of iron homeostasis by the p53-ISCU pathway. Sci. Rep. 2015, 5, 16497. [Google Scholar] [CrossRef] [PubMed]
- Aleksandrova, K.; Bamia, C.; Drogan, D.; Lagiou, P.; Trichopoulou, A.; Jenab, M.; Fedirko, V.; Romieu, I.; Bueno-de-Mesquita, H.B.; Pischon, T.; et al. The association of coffee intake with liver cancer risk is mediated by biomarkers of inflammation and hepatocellular injury: Data from the European Prospective Investigation into Cancer and Nutrition. Am. J. Clin. Nutr. 2015, 102, 1498–1508. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Guo, P.; Xie, X.; Wang, Y.; Chen, G. Ferroptosis, a new form of cell death, and its relationships with tumourous diseases. J. Cell. Mol. Med. 2017, 21, 648–657. [Google Scholar] [CrossRef]
- Crielaard, B.J.; Lammers, T.; Rivella, S. Targeting iron metabolism in drug discovery and delivery. Nat. Rev. Drug Discov. 2017, 16, 400–423. [Google Scholar] [CrossRef]
- Abid, Z.; Cross, A.J.; Sinha, R. Meat, dairy, and cancer. Am. J. Clin. Nutr. 2014, 100 (Suppl. 1), 386S–393S. [Google Scholar] [CrossRef] [PubMed]
- Mesías, M.; Seiquer, I.; Navarro, M.P. Iron nutrition in adolescence. Crit. Rev. Food Sci. Nutr. 2013, 53, 1226–1237. [Google Scholar] [CrossRef]
- Aschemeyer, S.; Qiao, B.; Stefanova, D.; Valore, E.V.; Sek, A.C.; Ruwe, T.A.; Vieth, K.R.; Jung, G.; Casu, C.; Rivella, S.; et al. Structure-function analysis of ferroportin defines the binding site and an alternative mechanism of action of hepcidin. Blood 2017, 131, 899–910. [Google Scholar] [CrossRef]
- Park, C.H.; Valore, E.V.; Waring, A.J.; Ganz, T. Hepcidin, a urinary antimicrobial peptide synthesized in the liver. J. Biol. Chem. 2001, 276, 7806–7810. [Google Scholar] [CrossRef] [PubMed]
- Bao, W.D.; Fan, Y.; Deng, Y.Z.; Long, L.Y.; Wang, J.J.; Guan, D.X.; Qian, Z.Y.; An, P.; Feng, Y.Y.; He, Z.Y.; et al. Iron overload in hereditary tyrosinemia type 1 induces liver injury through the Sp1/Tfr2/hepcidin axis. J. Hepatol. 2016, 65, 137–145. [Google Scholar] [CrossRef]
- Nahon, P.; Nuraldeen, R.; Rufat, P.; Sutton, A.; Trautwein, C.; Strnad, P. In alcoholic cirrhosis, low-serum hepcidin levels associate with poor long-term survival. Liver Int. 2016, 36, 185–188. [Google Scholar] [CrossRef]
- Xue, X.; Ramakrishnan, S.K.; Weisz, K.; Triner, D.; Xie, L.; Attili, D.; Pant, A.; Győrffy, B.; Zhan, M.; Carter-Su, C.; et al. Iron Uptake via DMT1 Integrates Cell Cycle with JAK-STAT3 Signaling to Promote Colorectal Tumorigenesis. Cell Metab. 2016, 24, 447–461. [Google Scholar] [CrossRef]
- Horniblow, R.D.; Bedford, M.; Hollingworth, R.; Evans, S.; Sutton, E.; Lal, N.; Beggs, A.; Iqbal, T.H.; Tselepis, C. BRAF mutations are associated with increased iron regulatory protein-2 expression in colorectal tumorigenesis. Cancer Sci. 2017, 108, 1135–1143. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, J.; Ding, C.; Dong, D.; Shang, P. Regulation of Osteoblast Differentiation and Iron Content in MC3T3-E1 Cells by Static Magnetic Field with Different Intensities. Biol. Trace Elem. Res. 2017, 184, 214–225. [Google Scholar] [CrossRef]
- De Falco, L.; Tortora, R.; Imperatore, N.; Bruno, M.; Capasso, M.; Girelli, D.; Castagna, A.; Caporaso, N.; Iolascon, A.; Rispo, A. The role of TMPRSS6 and HFE variants in iron deficiency anemia in celiac disease. Am. J. Hematol. 2018, 93, 383–393. [Google Scholar] [CrossRef]
- Lefebvre, T.; Reihani, N.; Daher, R.; de Villemeur, T.B.; Belmatoug, N.; Rose, C.; Colin-Aronovicz, Y.; Puy, H.; Le Van Kim, C.; Franco, M.; et al. Involvement of hepcidin in iron metabolism dysregulation in Gaucher disease. Haematologica 2018, 103, 587–596. [Google Scholar] [CrossRef] [PubMed]
- You, L.H.; Yan, C.Z.; Zheng, B.J.; Ci, Y.Z.; Chang, S.Y.; Yu, P.; Gao, G.F.; Li, H.Y.; Dong, T.Y.; Chang, Y.Z. Astrocyte hepcidin is a key factor in LPS-induced neuronal apoptosis. Cell Death Dis 2017, 8, e2676. [Google Scholar] [CrossRef]
- Pinnix, Z.K.; Miller, L.D.; Wang, W.; D’Agostino, R.; Kute, T.; Willingham, M.C.; Hatcher, H.; Tesfay, L.; Sui, G.; Di, X.; et al. Ferroportin and iron regulation in breast cancer progression and prognosis. Sci. Transl. Med. 2010, 2, 43ra56. [Google Scholar] [CrossRef] [PubMed]
- Ruchala, P.; Nemeth, E. The pathophysiology and pharmacology of hepcidin. Trends Pharmacol. Sci. 2014, 35, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Pusatcioglu, C.K.; Nemeth, E.; Fantuzzi, G.; Llor, X.; Freels, S.; Tussing-Humphreys, L.; Cabay, R.J.; Linzmeier, R.; Ng, D.; Clark, J.; et al. Systemic and tumor level iron regulation in men with colorectal cancer: A case control study. Nutr. Metab. (Lond.) 2014, 11, 21. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Liu, A.; Bai, R.; Zhang, B.; Jin, Y.; Guo, W.; Li, Y.; Gao, J.; Liu, L. Hepcidin and iron metabolism in the pathogenesis of prostate cancer. J. BUON 2017, 22, 1328–1332. [Google Scholar] [PubMed]
- Zhang, S.; Chen, Y.; Guo, W.; Yuan, L.; Zhang, D.; Xu, Y.; Nemeth, E.; Ganz, T.; Liu, S. Disordered hepcidin-ferroportin signaling promotes breast cancer growth. Cell. Signal. 2014, 26, 2539–2550. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Wang, L.; Ma, Y.; Wu, X.; Jin, L.; Yu, F. Increased hepcidin expression in non-small cell lung cancer tissue and serum is associated with clinical stage. Thorac. Cancer 2014, 5, 14–24. [Google Scholar] [CrossRef]
- Tesfay, L.; Clausen, K.A.; Kim, J.W.; Hegde, P.; Wang, X.; Miller, L.D.; Deng, Z.; Blanchette, N.; Arvedson, T.; Miranti, C.K.; et al. Hepcidin regulation in prostate and its disruption in prostate cancer. Cancer Res. 2015, 75, 2254–2263. [Google Scholar] [CrossRef]
- Vela, D.; Vela-Gaxha, Z. Differential regulation of hepcidin in cancer and non-cancer tissues and its clinical implications. Exp. Mol. Med. 2018, 50, e436. [Google Scholar] [CrossRef]
- Kessler, S.M.; Barghash, A.; Laggai, S.; Helms, V.; Kiemer, A.K. Hepatic hepcidin expression is decreased in cirrhosis and HCC. J. Hepatol. 2015, 62, 977–979. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Cui, Y.; Xu, B.; Gu, J.; Wang, W.; Luo, X. Hypermethylation leads to bone morphogenetic protein 6 downregulation in hepatocellular carcinoma. PLoS ONE 2014, 9, e87994. [Google Scholar] [CrossRef] [PubMed]
- Maegdefrau, U.; Arndt, S.; Kivorski, G.; Hellerbrand, C.; Bosserhoff, A.K. Downregulation of hemojuvelin prevents inhibitory effects of bone morphogenetic proteins on iron metabolism in hepatocellular carcinoma. Lab. Invest. 2011, 91, 1615–1623. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Pathology Character | n | HAMP Expression | p Value | |
|---|---|---|---|---|
| Low | High | |||
| Adjacent inflammation | 0.3829 | |||
| None | 117 | 63 | 54 | |
| Mild | 100 | 46 | 54 | |
| Severe | 17 | 6 | 11 | |
| M stage | 0.0121 * | |||
| M0 | 267 | 135 | 132 | |
| M1 | 4 | 0 | 4 | |
| T stage | 0.0361 * | |||
| T1 | 181 | 82 | 99 | |
| T2 | 94 | 48 | 46 | |
| T3 | 80 | 47 | 33 | |
| T4 | 13 | 7 | 6 | |
| Age (year) | 0.1581 | |||
| ≤60 | 177 | 81 | 96 | |
| >60 | 195 | 104 | 91 | |
| Postoperative radiotherapy | 0.1339 | |||
| No | 241 | 121 | 120 | |
| Yes | 4 | 2 | 2 | |
| Family history | 0.8110 | |||
| No | 208 | 100 | 108 | |
| Yes | 112 | 60 | 52 | |
| History risk factor | 0.1123 | |||
| Alcohol consumption | 68 | 37 | 31 | |
| Hemochromatosis | 6 | 4 | 2 | |
| Hepatitis B | 76 | 37 | 39 | |
| Hepatitis C | 32 | 13 | 19 | |
| Liver fibrosis Ishak score | 0.6397 | |||
| 0—No Fibrosis | 74 | 40 | 34 | |
| 1,2—Portal Fibrosis | 31 | 16 | 15 | |
| 3,4—Fibrous Speta | 28 | 14 | 14 | |
| 5—Nodular Formation | 9 | 3 | 6 | |
| 6—Established Cirrhosis | 70 | 35 | 35 | |
| Lymph node stage | 0.4924 | |||
| N0 | 253 | 125 | 128 | |
| N1 | 4 | 3 | 1 | |
| Neoplasm stage | 0.9142 | |||
| Stage I | 171 | 78 | 93 | |
| Stage II | 86 | 43 | 43 | |
| Stage III | 85 | 52 | 33 | |
| Stage IV | 5 | 1 | 4 | |
| Neoplasm histologic grade | 0.0129 * | |||
| G1 | 55 | 19 | 36 | |
| G2 | 178 | 71 | 107 | |
| G3 | 122 | 40 | 82 | |
| G4 | 12 | 3 | 9 | |
| Sex | 0.4953 | |||
| Female | 121 | 91 | 30 | |
| Male | 250 | 192 | 58 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, Y.; Li, X.; Su, Y.; Badshah, S.A.; Zhang, B.; Xue, Y.; Shang, P. HAMP Downregulation Contributes to Aggressive Hepatocellular Carcinoma via Mechanism Mediated by Cyclin4-Dependent Kinase-1/STAT3 Pathway. Diagnostics 2019, 9, 48. https://doi.org/10.3390/diagnostics9020048
Shen Y, Li X, Su Y, Badshah SA, Zhang B, Xue Y, Shang P. HAMP Downregulation Contributes to Aggressive Hepatocellular Carcinoma via Mechanism Mediated by Cyclin4-Dependent Kinase-1/STAT3 Pathway. Diagnostics. 2019; 9(2):48. https://doi.org/10.3390/diagnostics9020048
Chicago/Turabian StyleShen, Ying, Xin Li, Yanwei Su, Shaikh Atik Badshah, Bin Zhang, Yanru Xue, and Peng Shang. 2019. "HAMP Downregulation Contributes to Aggressive Hepatocellular Carcinoma via Mechanism Mediated by Cyclin4-Dependent Kinase-1/STAT3 Pathway" Diagnostics 9, no. 2: 48. https://doi.org/10.3390/diagnostics9020048
APA StyleShen, Y., Li, X., Su, Y., Badshah, S. A., Zhang, B., Xue, Y., & Shang, P. (2019). HAMP Downregulation Contributes to Aggressive Hepatocellular Carcinoma via Mechanism Mediated by Cyclin4-Dependent Kinase-1/STAT3 Pathway. Diagnostics, 9(2), 48. https://doi.org/10.3390/diagnostics9020048




