Addressing Class Imbalance in Fetal Health Classification: Rigorous Benchmarking of Multi-Class Resampling Methods on Cardiotocography Data
Abstract
1. Introduction
2. Related Work
3. Methodology
3.1. Dataset Overview
3.2. Workflow of the Model
- Data acquisition and labeling: The CTG dataset is obtained with three outcome classes: normal, suspect, and pathologic.
- Train–test split: The data is partitioned using a 70/30 stratified split. The test set is set aside as an independent validation cohort for unbiased final evaluation.
- Cross-validation for hyperparameter tuning: Within the training set, 5-fold cross-validation is performed. During each fold, four parts are used for training and one part serves as the validation set. This process is used exclusively for selecting the best hyperparameters. A fixed random seed (seed = 123) is used to ensure identical cross-validation folds across all models, enabling a fair performance comparison.
- Class imbalance handling: Five different resampling methods are applied: SMOTE, BSMOTE, ADASYN, NearMiss, and SCUT. Each method is applied exclusively to the training data to address class imbalance and prevent data leakage. The original unbalanced training set is also retained as a baseline.
- Balanced training data: Resampling produces balanced training sets with corrected class proportions. This reduces majority class bias during model training.
- Feature preprocessing: Both training and test sets are standardized. The mean and standard deviation are computed only on the training data and then these same parameters are applied to the test data.
- Model training and hyperparameter tuning: Seven machine learning algorithms are trained: NB, RF, LDA, KNN, SVM, MLR, and MLP. Hyperparameters are tuned via grid search within the cross-validation process. Probability outputs are generated for ROC analysis.
- Prediction generation: Each trained model is used to produce predictions and probabilities on the held-out validation fold, aggregating results across folds.
- Final evaluation: The retrained models are evaluated on the untouched test set. Performance is measured using BACC, Macro-F1, Macro-MCC, and Macro-Averaged ROC-AUC. Per-class ROC curves are generated using a one-vs-all approach, and individual class AUC values are averaged to compute the Macro-AUC.
3.3. Algorithms
3.3.1. NB
3.3.2. RF
3.3.3. KNN
3.3.4. Linear SVMs
3.3.5. LDA
3.3.6. MLR
3.3.7. MLP
3.4. Class Imbalance and Resampling Methods
3.4.1. SMOTE
3.4.2. BSMOTE
3.4.3. ADASYN
3.4.4. NearMiss
3.4.5. SCUT
3.5. Model Evaluation
- BACC: This metric mitigates the effect of class imbalance by averaging sensitivity () and specificity ().
- Macro-F1-Score: The F1-score is the harmonic mean of precision and recall, providing a balanced measure that accounts for both false positives and false negatives. In multi-class settings, the Macro-F1 is computed by calculating the F1-score separately for each class and then averaging the results. For each class i, precision (), recall (), and the corresponding F1-score () These are computed as follows:Finally, the Macro-F1-Score is obtained by taking the arithmetic mean across all k classes:This macro-averaging approach treats all classes equally, regardless of their size. This makes it particularly suitable for imbalanced datasets where minority class performance is critical. The Macro-F1 is widely applied across binary, multi-class, and multi-label classification problems, particularly in domains where class imbalance is prevalent [112,113].
- Macro-MCC: The Matthews Correlation Coefficient (MCC) evaluates the quality of binary classifications by incorporating all four entries of the confusion matrix. It ranges from (perfect agreement) to (complete disagreement), with 0 representing random prediction [114]. For multi-class problems, we employ a macro-averaging approach. MCC is computed separately for each class using a one-vs-rest strategy, where each class is treated as positive while all others are grouped as negative. The per-class MCC values are then averaged to obtain the Macro-Averaged MCC:where k represents the number of classes, and , , , and denote the true positives, true negatives, false positives, and false negatives for class i, respectively. This macro-averaging approach ensures that each class contributes equally to the evaluation, making the metric robust to class imbalance [115,116].
- Macro-Averaged ROC-AUC: Receiver Operating Characteristic (ROC) curves provide a visual and quantitative tool for evaluating classifier performance across different decision thresholds. Originally developed for medical decision-making, ROC analysis has become widely adopted in machine learning and data mining research. The area under the ROC curve (AUC) offers a single scalar measure of overall classification performance. AUC values range from 0 to 1, where 1 indicates perfect discrimination and corresponds to random guessing. In practice, any meaningful classifier should achieve an AUC above 0.5 [117,118].For the multi-class problems, we employed a one-vs-rest strategy to compute the ROC curves and AUC values for each class independently [119]. In this approach, each class is treated sequentially as the positive class, while all remaining classes are combined to form the negative class. For our three-class CTG dataset (normal, suspect, pathological), this procedure generated three separate ROC curves. Each curve plots sensitivity (true-positive rate) against 1-specificity (false-positive rate) across all possible classification thresholds. The Macro-Averaged ROC-AUC was then calculated as the arithmetic mean of the individual class AUC values:where k represents the number of classes and denotes the area under the ROC curve for class i. By averaging across classes rather than aggregating predictions, this macro-averaging approach ensures that each class contributes equally to the final metric, regardless of its frequency in the dataset. This property makes the Macro-AUC particularly well-suited for imbalanced classification tasks [120].
4. Results and Discussion
- NB (Figure 3): SMOTE achieves the highest performance (AUC = 0.942) compared to the base model (0.926), whereas BSMOTE and NearMiss reduce it to approximately 0.911–0.913. Despite these AUC improvements, BACC decreases under both SMOTE (0.8128 to 0.7671) and SCUT (0.8128 to 0.7013), demonstrating that enhanced ranking ability does not guarantee improved threshold-based classification accuracy.
- RF (Figure 4): This achieves near-perfect AUCs (approximately 0.986–0.989) across all resampling methods, confirming its robustness and superior predictive calibration. The base, SMOTE, and BSMOTE models achieve identical top performance (AUC = 0.989), indicating highly stable and accurate class separation. Minor declines under ADASYN, NearMiss, and SCUT suggest that resampling offers no additional benefit.
- LDA (Figure 5): This maintains high AUCs across methods (approximately 0.951–0.956), with a minor reduction under ADASYN (0.934). The base configuration remains optimal.
- KNN (Figure 6): Overall AUC improves from the base model (0.870) under SMOTE (0.899) and SCUT (0.905), while NearMiss achieves the highest value (0.949). However, BACC consistently decreases across all configurations despite these AUC gains. This divergence occurs because improving ranking ability (AUC) does not guarantee better performance at a fixed decision threshold; resampling shifts class priors and alters optimal threshold positions, resulting in improved discrimination without corresponding gains in classification accuracy.
- SVM (Figure 7): This records excellent performance with base and BSMOTE models (approximately 0.971–0.972), while ADASYN and NearMiss slightly lower AUCs to approximately 0.954–0.958.
- MLR (Figure 8): This shows consistently strong performance (AUC approximately 0.964–0.970), with the base model performing best (0.970).
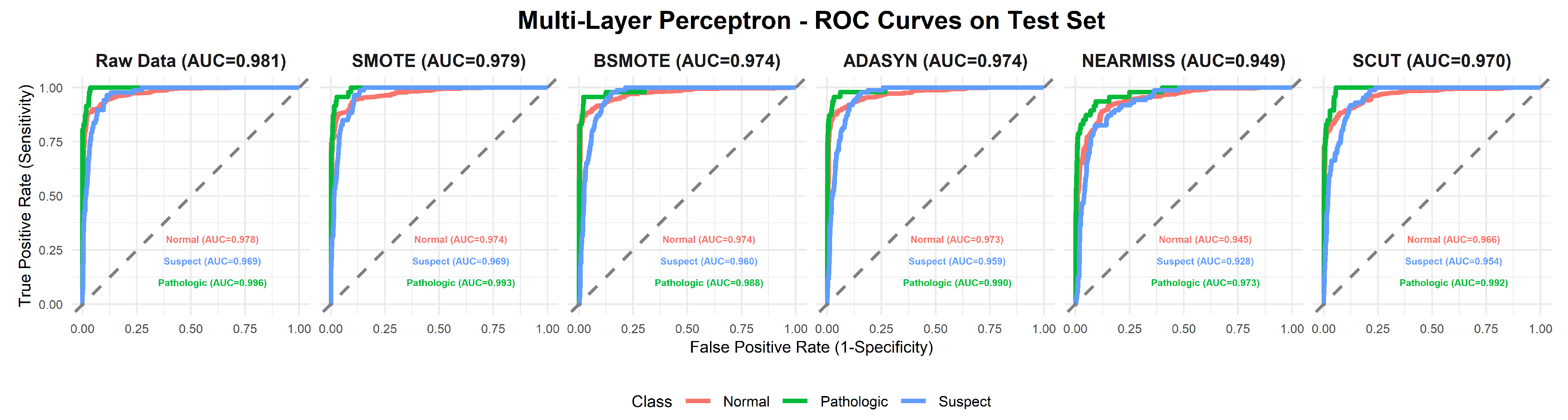
- MLP (Figure 9): The base model exhibits the highest AUC (0.981), while oversampling slightly decreases performance (SMOTE = 0.979, BSMOTE and ADASYN= 0.974).
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Study | Year | Methods | Classes | Performance | Imbalance Handling |
|---|---|---|---|---|---|
| Sahin et al. [25] | 2015 | ANN, SVM, k-NN, RF, CART, LR, C4.5, RBFN | Binary | Accuracy = 99.2%; F1-score = 99.2%; AUC = 99.9% | No |
| Piri et al. [29] | 2019 | DT, LR, SVM, KNN, GNB, RF, XGBoost | 3-class | Accuracy = 84% | No |
| Piri et al. [31] | 2020 | MOGA-CD + DT, SVM, GNB, RF, XGBoost | 3-class | Accuracy = 94% | No |
| Pradhan et al. [34] | 2021 | RF, LR, KNN, GBM | 3-class | Accuracy = 93% | No |
| Rahmayanti et al. [35] | 2022 | ANN, LSTM, XGBoost, SVM, KNN, LightGBM, RF | 3-class | Accuracy = 89–99%; F1-score = 98%; AUC = 99% | SMOTE |
| Kaliappan et al. [36] | 2023 | DT, RF, SVM, KNN, GNB, AdaBoost, Gradient Boosting, Voting Classifier, Neural Networks | 3-class | Accuracy = 99%; F1-score ≈ 97%; MCC = 96% | Random Oversampling |
| Salini et al. [38] | 2024 | RF, LR, DT, SVC, KNN, Voting Classifier | 3-class | Accuracy = 93% | No |
| Nazlı et al. [40] | 2025 | CatBoost, DT, ExtraTrees, GB, KNN, LightGBM, RF, SVM, ANN, DNN | 3-class | BACC = 91.34% | SMOTE |
| Ahmed et al. [13] | 2025 | ML (RF, SVM, KNN, DT, LR, XGBoost, ET); DL (ANN, CNN, RNN, LSTM, GRU) + Stacking Meta-Classifier | 3-class | Accuracy = 98.9%; F1-score = 99.3%; AUC = 99.8% | SMOTE |
| Bhukya et al. [41] | 2025 | ML: KNN, DT, SVM, RF, SGD, GB, AdaBoost, XGBoost; DL: CNN, LSTM, BiLSTM | 3-class | Accuracy ≈ 99.39%; F1-score = 99% | No |
| Proposed work | 2026 | NB, RF, LDA, KNN, Linear SVM, MLR, MLP | 3-class | BACC = 91.18% (RF + Base), 90.92% (RF + BSMOTE); Macro-F1 = 90.73%; Macro-MCC = 85.33%; Macro-ROC-AUC = 98.9% (RF + BSMOTE) | Base, SMOTE, BSMOTE, ADASYN, NearMiss, SCUT |
References
- Mehbodniya, A.; Lazar, A.J.P.; Webber, J.; Sharma, D.K.; Jayagopalan, S.; Singh, K.K.P.; Rajan, R.; Pandya, S.; Sengan, S. Fetal health classification from cardiotocographic data using machine learning. Expert Syst. 2022, 39, e12899. [Google Scholar] [CrossRef]
- Mendis, L.; Palaniswami, M.; Brownfoot, F.; Keenan, E. Computerised cardiotocography analysis for the automated detection of fetal compromise during labour: A review. Bioengineering 2023, 10, 1007. [Google Scholar] [CrossRef] [PubMed]
- Hantoushzadeh, S.; Gargari, O.K.; Jamali, M.; Farrokh, F.; Eshraghi, N.; Asadi, F.; Mirzamoradi, M.; Razavi, S.J.; Ghaemi, M.; Aski, S.K.; et al. The association between increased fetal movements in the third trimester and perinatal outcomes: A systematic review and meta-analysis. BMC Pregnancy Childbirth 2024, 24, 365. [Google Scholar] [CrossRef] [PubMed]
- Sundar, C.; Chitradevi, M.; Geetharamani, G. An analysis on the performance of K-means clustering algorithm for cardiotocogram data clustering. Int. J. Comput. Sci. Appl. 2012, 2, 11–20. [Google Scholar] [CrossRef]
- Namusoke, H.; Nannyonga, M.M.; Ssebunya, R.; Nakibuuka, V.K.; Mworozi, E. Incidence and short-term outcomes of neonates with hypoxic ischemic encephalopathy in a peri-urban teaching hospital, Uganda: A prospective cohort study. Matern. Health Neonatol. Perinatol. 2018, 4, 6. [Google Scholar] [CrossRef]
- Ayres-de-Campos, D.; Costa-Santos, C.; Bernardes, J. Prediction of neonatal state by computer analysis of fetal heart rate tracings: The antepartum arm of the SisPorto® multicentre validation study. Eur. J. Obstet. Gynecol. Reprod. Biol. 2005, 118, 52–60. [Google Scholar] [CrossRef]
- Pinas, A.; Chandraharan, E. Continuous cardiotocography during labour: Analysis, classification and management. Best Pract. Res. Clin. Obstet. Gynaecol. 2016, 30, 33–47. [Google Scholar] [CrossRef]
- Chittacharoen, A.; Chaitum, A.; Suthutvoravut, S.; Herabutya, Y. Fetal acoustic stimulation for early intrapartum assessment of fetal well-being. Int. J. Gynecol. Obstet. 2000, 69, 275–277. [Google Scholar] [CrossRef]
- Goonewardene, M.; Hanwellage, K. Fetal acoustic stimulation test for early intrapartum fetal monitoring. Ceylon Med. J. 2011, 56, 14–18. [Google Scholar] [CrossRef]
- Rahman, H.; Renjhen, P.; Dutta, S. Reliability of admission cardiotocography for intrapartum monitoring in low resource setting. Niger. Med. J. 2012, 53, 145–149. [Google Scholar] [CrossRef]
- David, B.; Saraswathi, K. Role of admission CTG as a screening test to predict fetal outcome and mode of delivery. Res. J. Pharm. Biol. Chem. Sci. 2014, 5, 295–299. [Google Scholar]
- Housseine, N.; Punt, M.C.; Browne, J.L.; Meguid, T.; Klipstein-Grobusch, K.; Kwast, B.E.; Franx, A.; Grobbee, D.E.; Rijken, M.J. Strategies for intrapartum foetal surveillance in low- and middle-income countries: A systematic review. PLoS ONE 2018, 13, e0206295. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.S.; Mahmoud, N.M. Early detection of fetal health status based on cardiotocography using artificial intelligence. Neural Comput. Appl. 2025, 37, 16753–16779. [Google Scholar] [CrossRef]
- Ignatov, P.N.; Lutomski, J.E. Quantitative cardiotocography to improve fetal assessment during labor: A preliminary randomized controlled trial. Eur. J. Obstet. Gynecol. Reprod. Biol. 2016, 205, 91–97. [Google Scholar] [CrossRef]
- Eenkhoorn, C.; van den Wildenberg, S.; Goos, T.G.; Dankelman, J.; Franx, A.; Eggink, A.J. A systematic catalog of studies on fetal heart rate pattern and neonatal outcome variables. J. Perinat. Med. 2025, 53, 94–109. [Google Scholar] [CrossRef]
- Lovers, A.; Daumer, M.; Frasch, M.G.; Ugwumadu, A.; Warrick, P.; Vullings, R.; Pini, N.; Tolladay, J.; Petersen, O.B.; Lederer, C.; et al. Advancements in fetal heart rate monitoring: A report on opportunities and strategic initiatives for better intrapartum care. BJOG Int. J. Obstet. Gynaecol. 2025, 132, 853–866. [Google Scholar] [CrossRef]
- Ocak, H. A medical decision support system based on support vector machines and the genetic algorithm for the evaluation of fetal well-being. J. Med. Syst. 2013, 37, 9913. [Google Scholar] [CrossRef]
- Hoodbhoy, Z.; Noman, M.; Shafique, A.; Nasim, A.; Chowdhury, D.; Hasan, B. Use of machine learning algorithms for prediction of fetal risk using cardiotocographic data. Int. J. Appl. Basic Med. Res. 2019, 9, 226–230. [Google Scholar] [CrossRef]
- Mooijman, P.; Catal, C.; Tekinerdogan, B.; Lommen, A.; Blokland, M. The effects of data balancing approaches: A case study. Appl. Soft Comput. 2023, 132, 109853. [Google Scholar] [CrossRef]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar] [CrossRef]
- Fernández, A.; García, S.; Galar, M.; Prati, R.C.; Krawczyk, B.; Herrera, F. Learning from Imbalanced Data Sets; Springer: Cham, Switzerland, 2018. [Google Scholar]
- Devoe, L.D.; Castillo, R.A.; Sherline, D.M. The nonstress test as a diagnostic test: A critical reappraisal. Am. J. Obstet. Gynecol. 1985, 152, 1047–1053. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Cömert, Z.; Kocamaz, A. Comparison of machine learning techniques for fetal heart rate classification. Acta Phys. Pol. A 2017, 132, 451–454. [Google Scholar] [CrossRef]
- Sahin, H.; Subasi, A. Classification of the cardiotocogram data for anticipation of fetal risks using machine learning techniques. Appl. Soft Comput. 2015, 33, 231–238. [Google Scholar] [CrossRef]
- Spilka, J.; Frecon, J.; Leonarduzzi, R.; Pustelnik, N.; Abry, P.; Doret, M. Intrapartum fetal heart rate classification from trajectory in sparse SVM feature space. In Proceedings of the IEEE Engineering in Medicine and Biology Conference (EMBC), Milan, Italy, 25–29 August 2015; IEEE: Piscataway, NJ, USA, 2015; pp. 2335–2338. [Google Scholar] [CrossRef]
- Ramla, M.; Sangeetha, S.; Nickolas, S. Fetal health state monitoring using decision tree classifier from cardiotocography measurements. In Proceedings of the International Conference on Intelligent Computing and Control Systems (ICICCS), Madurai, India, 14–15 June 2018; IEEE: Piscataway, NJ, USA, 2018; pp. 1799–1803. [Google Scholar] [CrossRef]
- Madiraju, R.; Upadhyay, U.; C, M.; Bharati, R. Fetal health analysis based on CTG. In Proceedings of the 11th International Conference on Communication and Signal Processing (ICCSP), Melmaruvathur, India, 5–7 June 2025; IEEE: Piscataway, NJ, USA, 2025; pp. 1706–1711. [Google Scholar] [CrossRef]
- Piri, J.; Mohapatra, P. Exploring fetal health status using an association-based classification approach. In Proceedings of the International Conference on Information Technology (ICIT), Bhubaneswar, India, 19–21 December 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 166–171. [Google Scholar] [CrossRef]
- Vani, R. Weighted deep neural network based clinical decision support system for the determination of fetal health. Int. J. Recent Technol. Eng. 2019, 8, 8564–8569. [Google Scholar] [CrossRef]
- Piri, J.; Mohapatra, P.; Dey, R. Fetal health status classification using MOGA-CD based feature selection approach. In Proceedings of the IEEE International Conference on Electronics, Computing and Communication Technologies (CONECCT), Bangalore, India, 2–4 July 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 1–6. [Google Scholar] [CrossRef]
- Zhou, D.; Wang, J.; Xu, X. Multi-channel signal analysis for fetal heart rate monitoring using cardiotocography. IEEE Trans. Instrum. Meas. 2020, 69, 1013–1023. [Google Scholar]
- Li, J.; Liu, X. Fetal health classification based on machine learning. In Proceedings of the IEEE 2nd International Conference on Big Data, Artificial Intelligence and Internet of Things Engineering (ICBAIE), Nanchang, China, 26–28 March 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 899–902. [Google Scholar] [CrossRef]
- Pradhan, A.K.; Rout, J.K.; Maharana, A.B.; Balabantaray, B.K.; Ray, N.K. A machine learning approach for the prediction of fetal health using CTG. In Proceedings of the International Conference on Optical and Intelligent Technologies (OCIT), Hubli, India, 25–27 June 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 239–244. [Google Scholar] [CrossRef]
- Rahmayanti, N.; Pradani, H.; Pahlawan, M.; Vinarti, R. Comparison of machine learning algorithms to classify fetal health using cardiotocogram data. Procedia Comput. Sci. 2022, 197, 162–171. [Google Scholar] [CrossRef]
- Kaliappan, J.; Bagepalli, A.R.; Almal, S.; Mishra, R.; Hu, Y.-C.; Srinivasan, K. Impact of cross-validation on machine learning models for early detection of intrauterine fetal demise. Diagnostics 2023, 13, 1692. [Google Scholar] [CrossRef]
- Regmi, B.; Shah, C. Classification methods based on machine learning for the analysis of fetal health data. arXiv 2023, arXiv:2311.10962. [Google Scholar] [CrossRef]
- Salini, Y.; Mohanty, S.N.; Ramesh, J.V.N.; Yang, M.; Mukkoti, M.V.C. Cardiotocography data analysis for fetal health classification using machine learning models. IEEE Access 2024, 12, 26005–26022. [Google Scholar] [CrossRef]
- Zeng, R.; Lu, Y.; Long, S.; Wang, C.; Bai, J. Cardiotocography signal abnormality classification using time-frequency features and ensemble cost-sensitive SVM classifier. Comput. Biol. Med. 2021, 130, 104218. [Google Scholar] [CrossRef] [PubMed]
- Nazli, I.; Korbeko, E.; Dogru, S.; Kugu, E.; Sahingoz, O.K. Early detection of fetal health conditions using machine learning for classifying imbalanced cardiotocographic data. Diagnostics 2025, 15, 1250. [Google Scholar] [CrossRef] [PubMed]
- Bhukya, R.K.; Kayande, D.D.; Jain, A.; Agrawal, S.; Chitravanshi, P.; Walia, S.V. Enhancing fetal health classification: A study on fetal cardiotocograms. Procedia Comput. Sci. 2025, 260, 217–225. [Google Scholar] [CrossRef]
- Ayres-de-Campos, D.; Spong, C.Y.; Chandraharan, E. FIGO consensus guidelines on intrapartum fetal monitoring: Cardiotocography. Int. J. Gynecol. Obstet. 2015, 131, 13–24. [Google Scholar] [CrossRef]
- Wong, W.K.; Juwono, F.H.; Apriono, C.; Fitri, I.R. Fetal health prediction from cardiotocography recordings using Kolmogorov–Arnold networks. IEEE Open J. Eng. Med. Biol. 2025, 6, 345–351. [Google Scholar] [CrossRef]
- Campos, D.; Bernardes, J. Cardiotocography. UCI Mach. Learn. Repos. 2000. [Google Scholar] [CrossRef]
- Ayres-de-Campos, D.; Bernardes, J.; Garrido, A.; Marques-de-Sa, J.; Pereira-Leite, L. SisPorto 2.0: A program for automated analysis of cardiotocograms. J. Matern.-Fetal Med. 2000, 9, 311–318. [Google Scholar] [CrossRef]
- Ilham, A.; Kindarto, A.; Fathurohman, A.; Khikmah, L.; Ramadhani, R.D.; Jawad, S.A.; Liana, D.A.; Amylia, A.; Oleiwi, A.K.; Mutiar, A.; et al. CFCM-SMOTE: A robust fetal health classification to improve precision modelling in multi-class scenarios. Int. J. Comput. Digit. Syst. 2024, 16, 471–486. [Google Scholar] [CrossRef]
- Yin, Y.; Bingi, Y. Using machine learning to classify human fetal health and analyze feature importance. BioMedInformatics 2023, 3, 280–298. [Google Scholar] [CrossRef]
- Solt, I.; Caspi, O.; Beloosesky, R.; Weiner, Z.; Avdor, E. Machine learning approach to fetal weight estimation. Am. J. Obstet. Gynecol. 2019, 220, S666–S667. [Google Scholar] [CrossRef]
- Alsaggaf, W.; Cömert, Z.; Nour, M.; Polat, K.; Brdesee, H.; Toğaçar, M. Predicting fetal hypoxia using common spatial pattern and machine learning from cardiotocography signals. Appl. Acoust. 2020, 167, 107429. [Google Scholar] [CrossRef]
- Ananth, C.V.; Brandt, J.S. Fetal growth and gestational age prediction by machine learning. Lancet Digit. Health 2020, 2, e336–e337. [Google Scholar] [CrossRef] [PubMed]
- Lindley, D.V. Fiducial distributions and Bayes’ theorem. J. R. Stat. Soc. Ser. B (Methodol.) 1958, 20, 102–107. [Google Scholar] [CrossRef]
- Rish, I. An empirical study of the naive Bayes classifier. In Proceedings of the IJCAI Workshop on Empirical Methods in Artificial Intelligence, Seattle, WA, USA, 4 August 2001; IJCAI: Seattle, WA, USA, 2001; pp. 41–46. [Google Scholar]
- Uddin, S.; Khan, A.; Hossain, M.E.; Moni, M.A. Comparing different supervised machine learning algorithms for disease prediction. BMC Med. Inform. Decis. Mak. 2019, 19, 281. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Batla, A.; Kumar, C.; Jain, G. Comparative analysis of machine learning models for fake news classification. J. Xi’An Shiyou Univ. Nat. Sci. Ed. 2023, 19, 1250–1254. [Google Scholar]
- Zhang, L. Features extraction based on Naive Bayes algorithm and TF-IDF for news classification. PLoS ONE 2025, 20, e0327347. [Google Scholar] [CrossRef]
- Montesinos-López, O.A.; Montesinos-López, A.; Mosqueda-Gonzalez, B.A.; Montesinos-López, J.C.; Crossa, J.; Ramirez, N.L.; Valladares-Anguiano, F.A. A zero altered Poisson random forest model for genomic-enabled prediction. G3 Genes Genomes Genet. 2021, 11, jkaa057. [Google Scholar] [CrossRef]
- Tyralis, H.; Papacharalampous, G.; Langousis, A. A brief review of random forests for water scientists and practitioners and their recent history in water resources. Water 2019, 11, 910. [Google Scholar] [CrossRef]
- Nhu, V.H.; Shirzadi, A.; Shahabi, H.; Singh, S.K.; Al-Ansari, N.; Clague, J.J.; Ahmad, B.B. Shallow landslide susceptibility mapping: A comparison between logistic model tree, logistic regression, naive Bayes tree, artificial neural networks, and support vector machine algorithms. Int. J. Environ. Res. Public Health 2020, 17, 2749. [Google Scholar] [CrossRef]
- Singh, M.S.; Thongam, K.; Choudhary, P.; Bhagat, P.K. An integrated machine learning approach for congestive heart failure prediction. Diagnostics 2024, 14, 736. [Google Scholar] [CrossRef]
- Joachims, T. Making large-scale SVM learning practical. In Advances in Kernel Methods: Support Vector Learning; MIT Press: Cambridge, MA, USA, 1999; pp. 169–184. Available online: http://svmlight.joachims.org/ (accessed on 22 October 2025).
- Hussain, M.; Wajid, S.K.; Elzaart, A.; Berbar, M. A comparison of SVM kernel functions for breast cancer detection. In Proceedings of the International Conference on Computer Graphics, Imaging and Visualization (CGIV), Singapore, 17–19 August 2011; IEEE: Piscataway, NJ, USA, 2011; pp. 145–150. [Google Scholar] [CrossRef]
- Pal, S.; Peng, Y.; Aselisewine, W.; Barui, S. A support vector machine-based cure rate model for interval censored data. Stat. Methods Med. Res. 2023, 32, 2405–2422. [Google Scholar] [CrossRef]
- Iftikhar, H.; Khan, M.; Khan, Z.; Khan, F.; Alshanbari, H.M.; Ahmad, Z. A comparative analysis of machine learning models: A case study in predicting chronic kidney disease. Sustainability 2023, 15, 2754. [Google Scholar] [CrossRef]
- Dogantekin, E.; Dogantekin, A.; Avci, D.; Avci, L. An intelligent diagnosis system for diabetes on linear discriminant analysis and adaptive network based fuzzy inference system: LDA-ANFIS. Digit. Signal Process. 2010, 20, 1248–1255. [Google Scholar] [CrossRef]
- Çalişir, D.; Doğantekin, E. An automatic diabetes diagnosis system based on LDA–Wavelet support vector machine classifier. Expert Syst. Appl. 2011, 38, 8311–8315. [Google Scholar] [CrossRef]
- Tharwat, A.; Gaber, T.; Ibrahim, A.; Hassanien, A.E. Linear discriminant analysis: A detailed tutorial. AI Commun. 2017, 30, 169–190. [Google Scholar] [CrossRef]
- Egwom, O.J.; Hassan, M.; Tanimu, J.J.; Hamada, M.; Ogar, O.M. An LDA–SVM machine learning model for breast cancer classification. BioMedInformatics 2022, 2, 345–358. [Google Scholar] [CrossRef]
- Choubey, D.K.; Kumar, M.; Shukla, V.; Tripathi, S.; Dhandhania, V.K. Comparative analysis of classification methods with PCA and LDA for diabetes. Curr. Diabetes Rev. 2020, 16, 833–850. [Google Scholar] [CrossRef]
- Friedman, J.H.; Hastie, T.; Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 2010, 33, 1–22. [Google Scholar] [CrossRef]
- Madhu, B.; Ashok, N.C.; Balasubramanian, S. Multinomial logistic regression predicted probability map to visualize the influence of socio-economic factors on breast cancer occurrence in Southern Karnataka. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2014, 40, 193–196. [Google Scholar] [CrossRef]
- Ueki, K.; Hino, H.; Kuwatani, T. Geochemical discrimination and characteristics of magmatic tectonic settings: A machine-learning-based approach. Geochem. Geophys. Geosystems 2018, 19, 1327–1347. [Google Scholar] [CrossRef]
- Itano, K.; Ueki, K.; Iizuka, T.; Kuwatani, T. Geochemical discrimination of monazite source rock based on machine learning techniques and multinomial logistic regression analysis. Geosciences 2020, 10, 63. [Google Scholar] [CrossRef]
- Jain, A.K.; Mao, J.; Mohiuddin, K.M. Artificial neural networks: A tutorial. Computer 1996, 29, 31–44. [Google Scholar] [CrossRef]
- Abdelbasset, W.K.; Elkholi, S.M.; Opulencia, M.J.C.; Diana, T.; Su, C.-H.; Alashwal, M.; Nguyen, H.C. Development of multiple machine-learning computational techniques for optimization of heterogeneous catalytic biodiesel production from waste vegetable oil. Arab. J. Chem. 2022, 15, 103843. [Google Scholar] [CrossRef]
- Ekinci, G.; Ozturk, H.K. Forecasting wind farm production in the short, medium, and long terms using various machine learning algorithms. Energies 2025, 18, 1125. [Google Scholar] [CrossRef]
- Sumayli, A. Development of advanced machine learning models for optimization of methyl ester biofuel production from papaya oil: Gaussian process regression (GPR), multilayer perceptron (MLP), and K-nearest neighbor (KNN) regression models. Arab. J. Chem. 2023, 16, 104833. [Google Scholar] [CrossRef]
- Suguna, R.; Prakash, J.S.; Pai, H.A.; Mahesh, T.R.; Kumar, V.V.; Yimer, T.E. Mitigating class imbalance in churn prediction with ensemble methods and SMOTE. Sci. Rep. 2025, 15, 16256. [Google Scholar] [CrossRef]
- Haykin, S. Neural Networks: A Comprehensive Foundation; Prentice Hall PTR: Englewood Cliffs, NJ, USA, 1994. [Google Scholar]
- Chen, S.M. Data Science and Big Data: An Environment of Computational Intelligence; Springer: Cham, Switzerland, 2017. [Google Scholar]
- Leevy, J.L.; Khoshgoftaar, T.M.; Bauder, R.A.; Seliya, N. A survey on addressing high-class imbalance in big data. J. Big Data 2018, 5, 42. [Google Scholar] [CrossRef]
- Sowjanya, A.M.; Mrudula, O. Effective treatment of imbalanced datasets in health care using modified SMOTE coupled with stacked deep learning algorithms. Appl. Nanosci. 2023, 13, 1829–1840. [Google Scholar] [CrossRef]
- Chowdhury, M.M.; Ayon, R.S.; Hossain, M.S. An investigation of machine learning algorithms and data augmentation techniques for diabetes diagnosis using class imbalanced BRFSS dataset. Healthc. Anal. 2024, 5, 100297. [Google Scholar] [CrossRef]
- Suresh, T.; Brijet, Z.; Subha, T.D. Imbalanced medical disease dataset classification using enhanced generative adversarial network. Comput. Methods Biomech. Biomed. Eng. 2023, 26, 1702–1718. [Google Scholar] [CrossRef]
- Altalhan, M.; Algarni, A.; Alouane, M.T.H. Imbalanced data problem in machine learning: A review. IEEE Access 2025, 13, 13686–13699. [Google Scholar] [CrossRef]
- Chakravarthy, A.D.; Bonthu, S.; Chen, Z.; Zhu, Q. Predictive models with resampling: A comparative study of machine learning algorithms and their performances on handling imbalanced datasets. In Proceedings of the IEEE International Conference on Machine Learning and Applications (ICMLA), Boca Raton, FL, USA, 16–19 December 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 1492–1495. [Google Scholar] [CrossRef]
- Almadi, M.; Alotaibi, F.; Almudawah, R.; Ali, A.; Nasser, Y.; Nasser, N. Data-driven machine learning models for enhanced fetal health classification and monitoring. In Proceedings of the International Conference on Computing, Data Management and Analytics (CDMA), Riyadh, Saudi Arabia, 16–17 February 2025; pp. 189–192. [Google Scholar] [CrossRef]
- Khan, M.; Ahmad, A.; Sarfraz, M. L1 regularization based fetal health analysis using ML techniques. In Proceedings of the International Conference on Computing, Communication and Networking Technologies (ICCCNT), Kamand, India, 24–28 June 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 1–6. [Google Scholar] [CrossRef]
- Piri, J.; Mohapatra, P. Imbalanced cardiotocography data classification using re-sampling techniques. In Machine Intelligence and Data Analytics for Sustainable Systems; Springer: Singapore, 2021; pp. 681–692. [Google Scholar] [CrossRef]
- Cicak, S.; Avci, U. Handling imbalanced data in predictive maintenance: A resampling-based approach. In Proceedings of the International Conference on Human-Oriented Robotics and Applications (HORA), Istanbul, Turkey, 8–10 June 2023; IEEE: Piscataway, NJ, USA, 2023; pp. 1–6. [Google Scholar] [CrossRef]
- Brownlee, J. Random Oversampling and Undersampling for Imbalanced Classification. 2021. Available online: https://machinelearningmastery.com/random-oversampling-and-undersampling-for-imbalanced-classification/ (accessed on 28 October 2025).
- Paula, B.; Torgo, L.; Ribeiro, R. A survey of predictive modelling under imbalanced distributions. arXiv 2015, arXiv:1505.01658. [Google Scholar] [CrossRef]
- Kraiem, M.S.; Sánchez-Hernández, F.; Moreno-García, M.N. Selecting the suitable resampling strategy for imbalanced data classification regarding dataset properties: An approach based on association models. Appl. Sci. 2021, 11, 8546. [Google Scholar] [CrossRef]
- Yang, Y.; Khorshidi, H.A.; Aickelin, U. A review on over-sampling techniques in classification of multi-class imbalanced datasets: Insights for medical problems. Front. Digit. Health 2024, 6, 1430245. [Google Scholar] [CrossRef] [PubMed]
- Kaur, P.; Gosain, A. Comparing the behavior of oversampling and undersampling approach of class imbalance learning by combining class imbalance problem with noise. In ICT Based Innovations; Springer: Singapore, 2017. [Google Scholar] [CrossRef]
- Ichihashi, H.; Honda, K.; Notsu, A.; Miyamoto, E. FCM classifier for high-dimensional data. In Proceedings of the IEEE World Congress on Computational Intelligence (FUZZ-IEEE), Hong Kong, China, 1–6 June 2008; IEEE: Piscataway, NJ, USA, 2008; pp. 200–206. [Google Scholar] [CrossRef][Green Version]
- Pezoulas, V.C.; Zaridis, D.I.; Mylona, E.; Androutsos, C.; Apostolidis, K.; Tachos, N.S.; Fotiadis, D.I. Synthetic data generation methods in healthcare: A review on open-source tools and methods. Comput. Struct. Biotechnol. J. 2024, 23, 2892–2910. [Google Scholar] [CrossRef] [PubMed]
- Turlapati, V.P.K.; Prusty, M.R. Outlier-SMOTE: A refined oversampling technique for improved detection of COVID-19. Intell.-Based Med. 2020, 3, 100023. [Google Scholar] [CrossRef]
- Han, H.; Wang, W.Y.; Mao, B.H. Borderline-SMOTE: A new over-sampling method in imbalanced data sets learning. In Intelligent Computing; Springer: Berlin/Heidelberg, Germany, 2005; pp. 878–887. [Google Scholar] [CrossRef]
- Zhuang, D.; Zhang, B.; Yang, Q.; Yan, J.; Chen, Z.; Chen, Y. Efficient text classification by weighted proximal SVM. In Proceedings of the IEEE International Conference on Data Mining (ICDM), Houston, TX, USA, 27–30 November 2005; IEEE: Piscataway, NJ, USA, 2005; p. 8. [Google Scholar] [CrossRef]
- Majzoub, H.A.; Elgedawy, I. AB-SMOTE: An affinitive borderline SMOTE approach for imbalanced data binary classification. Int. J. Mach. Learn. Comput. 2020, 10, 31–37. [Google Scholar] [CrossRef]
- Göcs, L.; Johanyák, Z.C. Feature selection with weighted ensemble ranking for improved classification performance on the CSE-CIC-IDS2018 dataset. Computers 2023, 12, 147. [Google Scholar] [CrossRef]
- Shoohi, L.M.; Saud, J.H. Adaptation proposed methods for handling imbalanced datasets based on over-sampling technique. Al-Mustansiriyah J. Sci. 2020, 31, 25–29. [Google Scholar] [CrossRef]
- Chen, Z.; Zhou, L.; Yu, W. ADASYN-Random forest based intrusion detection model. In Proceedings of the 2021 4th International Conference on Signal Processing and Machine Learning, Beijing, China, 18–20 August 2021; ACM: New York, NY, USA, 2021; pp. 152–159. [Google Scholar] [CrossRef]
- Mani, I.; Zhang, J. kNN Approach to Unbalanced Data Distributions: A Case Study involving Information Extraction. In Proceedings of the Workshop on Learning from Imbalanced Datasets (ICML), Washington, DC, USA, 21 August 2003; ICML: Washington, DC, USA, 2003; pp. 1–7. [Google Scholar]
- Wickramasinghe, I.; Kalutarage, H. Naive Bayes: Applications, variations and vulnerabilities: A review of literature with code snippets for implementation. Soft Comput. 2021, 25, 2277–2293. [Google Scholar] [CrossRef]
- Blanquero, R.; Carrizosa, E.; Ramírez-Cobo, P.; Sillero-Denamiel, M.R. Constrained Naïve Bayes with application to unbalanced data classification. Cent. Eur. J. Oper. Res. 2022, 30, 1403–1425. [Google Scholar] [CrossRef]
- Gunawan, W.; Devianto, Y.; Sari, A.P. Imbalanced data NearMiss for comparison of SVM and Naive Bayes algorithms. Comput. Eng. Appl. J. 2024, 13, 34–43. [Google Scholar] [CrossRef]
- Jafarnejad, A.; Rezasoltani, A.; Khani, A.M. Comparative analysis of machine learning algorithms in predicting jumps in stock closing price: Case study of Iran Khodro using NearMiss and SMOTE approaches. Iran. J. Financ. 2025, 9, 27–54. [Google Scholar]
- Dempster, A.P.; Laird, N.M.; Rubin, D.B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Ser. B (Methodol.) 1977, 39, 1–22. [Google Scholar] [CrossRef]
- Agrawal, A.; Viktor, H.L.; Paquet, E. SCUT: Multi-class imbalanced data classification using SMOTE and cluster-based undersampling. In Proceedings of the International Joint Conference on Knowledge Discovery, Knowledge Engineering and Knowledge Management (IC3K), Lisbon, Portugal, 12–14 November 2015; IEEE: Piscataway, NJ, USA, 2015; Volume 1, pp. 226–234. [Google Scholar]
- Burduk, R. Classification performance metric for imbalance data based on recall and selectivity normalized in class labels. arXiv 2020, arXiv:2006.13319. [Google Scholar] [CrossRef]
- Lipton, Z.C.; Elkan, C.; Narayanaswamy, B. Optimal thresholding of classifiers to maximize F1 measure. In Machine Learning and Knowledge Discovery in Databases; Springer: Berlin/Heidelberg, Germany, 2014; pp. 225–239. [Google Scholar] [CrossRef]
- Opitz, J.; Burst, S. Macro F1 and micro F1. arXiv 2019, arXiv:1911.03347. [Google Scholar] [CrossRef]
- Matthews, B.W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta—Protein Struct. 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Ahuja, S.; Panigrahi, B.K.; Dey, N.; Taneja, A.; Gandhi, T.K. McS-Net: Multi-class Siamese network for severity of COVID-19 infection classification from lung CT scan slices. Appl. Soft Comput. 2022, 131, 109683. [Google Scholar] [CrossRef]
- Tamura, J.; Itaya, Y.; Hayashi, K.; Yamamoto, K. Statistical inference of the Matthews correlation coefficient for multiclass classification. arXiv 2025, arXiv:2503.06450. [Google Scholar] [CrossRef]
- Bradley, A.P. The use of the area under the ROC curve in the evaluation of machine learning algorithms. Pattern Recognit. 1997, 30, 1145–1159. [Google Scholar] [CrossRef]
- Hanley, J.A.; McNeil, B.J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982, 143, 29–36. [Google Scholar] [CrossRef]
- Hand, D.J.; Till, R.J. A simple generalisation of the area under the ROC curve for multiple class classification problems. Mach. Learn. 2001, 45, 171–186. [Google Scholar] [CrossRef]
- Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]








| Symbol | Variable Description |
|---|---|
| Class | Fetal state class: normal; suspect; pathologic |
| LB | Fetal heart beats per minute |
| AC | Accelerations per second |
| FM | Fetal movements per second |
| UC | Number of uterine contractions per second |
| DL | Number of light decelerations per second |
| DS | Number of severe decelerations per second |
| DP | Number of prolonged decelerations per second |
| ASTV | Percentage of time with abnormal short-term variability |
| MSTV | Mean value of short-term variability |
| ALTV | Percentage of time with abnormal long-term variability |
| MLTV | Mean value of long-term variability |
| Width | Width of FHR histogram |
| Max | Maximum of FHR histogram |
| Min | Minimum of FHR histogram |
| Nmax | Number of histogram peaks |
| Nzeros | Number of histogram zeros |
| Mode | Histogram mode |
| Mean | Histogram mean |
| Median | Histogram median |
| Variance | Histogram variance |
| Tendency | Histogram tendency: = left asymmetric; 0 = symmetric; 1 = right asymmetric |
| Algorithm | Resampling Method | BACC | Macro-MCC | Macro-F1 |
|---|---|---|---|---|
| NB | base | 0.8128 | 0.6474 | 0.7550 |
| SMOTE | 0.7671 | 0.6511 | 0.7616 | |
| BSMOTE | 0.7914 | 0.6340 | 0.7474 | |
| ADASYN | 0.7787 | 0.6061 | 0.7256 | |
| NearMiss | 0.7083 | 0.6210 | 0.7288 | |
| SCUT | 0.7013 | 0.6182 | 0.7276 | |
| RF | base | 0.9118 | 0.8477 | 0.9003 |
| SMOTE | 0.8897 | 0.8428 | 0.8973 | |
| BSMOTE | 0.9092 | 0.8533 | 0.9073 | |
| ADASYN | 0.8815 | 0.8437 | 0.8957 | |
| NearMiss | 0.7880 | 0.7616 | 0.8357 | |
| SCUT | 0.8657 | 0.8283 | 0.8898 | |
| LDA | base | 0.7874 | 0.6540 | 0.7626 |
| SMOTE | 0.7041 | 0.6415 | 0.7352 | |
| BSMOTE | 0.7209 | 0.6574 | 0.7490 | |
| ADASYN | 0.6490 | 0.5880 | 0.6916 | |
| NearMiss | 0.7005 | 0.6315 | 0.7324 | |
| SCUT | 0.7018 | 0.6359 | 0.7313 | |
| KNN | base | 0.8473 | 0.7574 | 0.8346 |
| SMOTE | 0.8449 | 0.7820 | 0.8513 | |
| BSMOTE | 0.8375 | 0.7713 | 0.8413 | |
| ADASYN | 0.8312 | 0.7658 | 0.8401 | |
| NearMiss | 0.7479 | 0.6820 | 0.7850 | |
| SCUT | 0.8245 | 0.7679 | 0.8445 | |
| SVM | base | 0.8190 | 0.6946 | 0.7835 |
| SMOTE | 0.7575 | 0.7042 | 0.7855 | |
| BSMOTE | 0.7791 | 0.7150 | 0.7932 | |
| ADASYN | 0.7013 | 0.6556 | 0.7429 | |
| NearMiss | 0.7193 | 0.6654 | 0.7566 | |
| SCUT | 0.7546 | 0.6903 | 0.7793 | |
| MLR | base | 0.7742 | 0.6683 | 0.7699 |
| SMOTE | 0.7437 | 0.6924 | 0.7770 | |
| BSMOTE | 0.7533 | 0.6908 | 0.7801 | |
| ADASYN | 0.6914 | 0.6393 | 0.7340 | |
| NearMiss | 0.7073 | 0.6355 | 0.7443 | |
| SCUT | 0.7382 | 0.6753 | 0.7649 | |
| MLP | base | 0.8636 | 0.7849 | 0.8560 |
| SMOTE | 0.8326 | 0.7862 | 0.8564 | |
| BSMOTE | 0.8230 | 0.7610 | 0.8391 | |
| ADASYN | 0.8040 | 0.7509 | 0.8306 | |
| NearMiss | 0.7527 | 0.6779 | 0.7797 | |
| SCUT | 0.7852 | 0.7219 | 0.8126 |
| Algorithm | Resampling Method | Sensitivity/Recall | Specificity | Precision/PPV | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Normal | Suspect | Pathological | Normal | Suspect | Pathological | Normal | Suspect | Pathological | ||
| NB | base | 0.935 | 0.674 | 0.575 | 0.722 | 0.924 | 0.997 | 0.927 | 0.580 | 0.931 |
| SMOTE | 0.901 | 0.779 | 0.638 | 0.797 | 0.908 | 0.987 | 0.944 | 0.568 | 0.790 | |
| BSMOTE | 0.921 | 0.721 | 0.553 | 0.729 | 0.919 | 0.993 | 0.928 | 0.579 | 0.867 | |
| ADASYN | 0.897 | 0.802 | 0.468 | 0.722 | 0.904 | 0.993 | 0.925 | 0.566 | 0.846 | |
| NearMiss | 0.834 | 0.884 | 0.660 | 0.902 | 0.851 | 0.975 | 0.970 | 0.481 | 0.674 | |
| SCUT | 0.828 | 0.884 | 0.681 | 0.902 | 0.851 | 0.971 | 0.970 | 0.481 | 0.653 | |
| RF | base | 0.974 | 0.779 | 0.915 | 0.857 | 0.973 | 0.997 | 0.963 | 0.817 | 0.956 |
| SMOTE | 0.956 | 0.849 | 0.915 | 0.902 | 0.958 | 0.995 | 0.974 | 0.760 | 0.935 | |
| BSMOTE | 0.966 | 0.814 | 0.936 | 0.872 | 0.969 | 0.997 | 0.966 | 0.805 | 0.957 | |
| ADASYN | 0.953 | 0.872 | 0.915 | 0.925 | 0.955 | 0.993 | 0.980 | 0.750 | 0.915 | |
| NearMiss | 0.903 | 0.849 | 0.957 | 0.932 | 0.931 | 0.971 | 0.981 | 0.658 | 0.726 | |
| SCUT | 0.929 | 0.884 | 0.957 | 0.932 | 0.935 | 0.995 | 0.981 | 0.679 | 0.938 | |
| LDA | base | 0.953 | 0.570 | 0.702 | 0.699 | 0.953 | 0.985 | 0.923 | 0.653 | 0.786 |
| SMOTE | 0.812 | 0.919 | 0.745 | 0.977 | 0.832 | 0.970 | 0.993 | 0.459 | 0.660 | |
| BSMOTE | 0.844 | 0.919 | 0.702 | 0.955 | 0.859 | 0.973 | 0.986 | 0.503 | 0.674 | |
| ADASYN | 0.802 | 0.733 | 0.851 | 0.985 | 0.826 | 0.946 | 0.995 | 0.396 | 0.556 | |
| NearMiss | 0.814 | 0.872 | 0.766 | 0.955 | 0.833 | 0.970 | 0.986 | 0.449 | 0.667 | |
| SCUT | 0.822 | 0.895 | 0.723 | 0.970 | 0.837 | 0.970 | 0.991 | 0.461 | 0.654 | |
| KNN | base | 0.962 | 0.698 | 0.809 | 0.797 | 0.964 | 0.988 | 0.947 | 0.750 | 0.844 |
| SMOTE | 0.947 | 0.826 | 0.809 | 0.872 | 0.951 | 0.988 | 0.966 | 0.725 | 0.844 | |
| BSMOTE | 0.951 | 0.802 | 0.787 | 0.865 | 0.953 | 0.987 | 0.964 | 0.726 | 0.822 | |
| ADASYN | 0.943 | 0.802 | 0.809 | 0.865 | 0.948 | 0.987 | 0.964 | 0.704 | 0.826 | |
| NearMiss | 0.846 | 0.872 | 0.851 | 0.895 | 0.875 | 0.978 | 0.968 | 0.521 | 0.755 | |
| SCUT | 0.921 | 0.849 | 0.851 | 0.895 | 0.929 | 0.988 | 0.971 | 0.652 | 0.851 | |
| SVM | base | 0.951 | 0.698 | 0.638 | 0.782 | 0.937 | 0.993 | 0.943 | 0.632 | 0.882 |
| SMOTE | 0.879 | 0.861 | 0.787 | 0.962 | 0.880 | 0.980 | 0.989 | 0.529 | 0.755 | |
| BSMOTE | 0.899 | 0.872 | 0.723 | 0.940 | 0.895 | 0.985 | 0.983 | 0.564 | 0.791 | |
| ADASYN | 0.875 | 0.756 | 0.809 | 0.970 | 0.882 | 0.959 | 0.991 | 0.500 | 0.613 | |
| NearMiss | 0.846 | 0.872 | 0.787 | 0.962 | 0.862 | 0.970 | 0.988 | 0.497 | 0.673 | |
| SCUT | 0.873 | 0.837 | 0.787 | 0.940 | 0.875 | 0.981 | 0.982 | 0.511 | 0.771 | |
| MLR | base | 0.937 | 0.663 | 0.702 | 0.782 | 0.935 | 0.983 | 0.942 | 0.613 | 0.767 |
| SMOTE | 0.877 | 0.849 | 0.787 | 0.955 | 0.884 | 0.975 | 0.987 | 0.533 | 0.712 | |
| BSMOTE | 0.893 | 0.837 | 0.745 | 0.910 | 0.902 | 0.976 | 0.974 | 0.571 | 0.714 | |
| ADASYN | 0.867 | 0.744 | 0.809 | 0.947 | 0.882 | 0.956 | 0.984 | 0.496 | 0.594 | |
| NearMiss | 0.848 | 0.802 | 0.787 | 0.895 | 0.871 | 0.968 | 0.968 | 0.493 | 0.661 | |
| SCUT | 0.875 | 0.837 | 0.745 | 0.940 | 0.879 | 0.976 | 0.982 | 0.518 | 0.714 | |
| MLP | base | 0.962 | 0.733 | 0.851 | 0.820 | 0.966 | 0.990 | 0.953 | 0.768 | 0.870 |
| SMOTE | 0.935 | 0.826 | 0.894 | 0.895 | 0.946 | 0.985 | 0.971 | 0.703 | 0.824 | |
| BSMOTE | 0.929 | 0.802 | 0.851 | 0.880 | 0.933 | 0.988 | 0.967 | 0.651 | 0.851 | |
| ADASYN | 0.915 | 0.814 | 0.872 | 0.902 | 0.922 | 0.985 | 0.973 | 0.620 | 0.820 | |
| NearMiss | 0.873 | 0.826 | 0.787 | 0.880 | 0.890 | 0.980 | 0.965 | 0.538 | 0.755 | |
| SCUT | 0.907 | 0.791 | 0.851 | 0.872 | 0.920 | 0.981 | 0.964 | 0.607 | 0.784 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Hawrami, Z.S.M.; Cengiz, M.A.; Dünder, E. Addressing Class Imbalance in Fetal Health Classification: Rigorous Benchmarking of Multi-Class Resampling Methods on Cardiotocography Data. Diagnostics 2026, 16, 485. https://doi.org/10.3390/diagnostics16030485
Hawrami ZSM, Cengiz MA, Dünder E. Addressing Class Imbalance in Fetal Health Classification: Rigorous Benchmarking of Multi-Class Resampling Methods on Cardiotocography Data. Diagnostics. 2026; 16(3):485. https://doi.org/10.3390/diagnostics16030485
Chicago/Turabian StyleHawrami, Zainab Subhi Mahmood, Mehmet Ali Cengiz, and Emre Dünder. 2026. "Addressing Class Imbalance in Fetal Health Classification: Rigorous Benchmarking of Multi-Class Resampling Methods on Cardiotocography Data" Diagnostics 16, no. 3: 485. https://doi.org/10.3390/diagnostics16030485
APA StyleHawrami, Z. S. M., Cengiz, M. A., & Dünder, E. (2026). Addressing Class Imbalance in Fetal Health Classification: Rigorous Benchmarking of Multi-Class Resampling Methods on Cardiotocography Data. Diagnostics, 16(3), 485. https://doi.org/10.3390/diagnostics16030485






