A Real-Time Dynamic Warning Method for MODS in Trauma Sepsis Patients Based on a Pre-Trained Transfer Learning Algorithm
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Definitions
2.3. Overall Flow Chart for MODS Prediction
- 1.
- Pre-train Model: Data of all ICU patients except trauma sepsis patients were collected from the MIMIC-IV and eICU databases, totaling 250,394 cases for model pre-training. The original data are divided into high-frequency time-series data and low-frequency data. These are input into the LSTM and the multilayer perceptron (MLP) models, respectively, to predict the risk of death within the next 30 days (“Pre-train Model” section of Figure 2);
- 2.
- MODS Model: Based on the pre-trained model, 80% of the MIMIC-IV trauma sepsis patient data were used for fine-tuning, resulting in the final real-time MODS prediction model (“MODS Model” section of Figure 2). This model employs a 4-h observation window and predicts the occurrence of MODS within the next 6, 12, and 24 h (“Time window” section of Figure 2);
- 3.
- Validation: The remaining 20% of the MIMIC-IV data and all eICU trauma sepsis patient data were used for internal and multicenter external validation of the model (“Validation” section of Figure 2);
- 4.
- Interpretability: The SHAP method is used to perform interpretability analysis on the model to reveal the impact of key features (“Interpretability” section of Figure 2).
2.4. Data Pre-Processing
2.5. Feature Selection
2.6. Model Development
2.6.1. MODS Prediction Model
2.6.2. Five Derivative Models
2.7. Time Windows and Labeling
2.8. Performance Evaluation
2.9. Model Interpretation
2.10. Model Parameters
3. Results
3.1. Development Cohort Analysis
3.2. Ablation Experiment
3.2.1. Model Performance Under Different Prediction Windows
3.2.2. Comparative Analysis of Models
3.2.3. Comparison with SOFA
3.3. Relationship Between Model Performance and Dataset Size
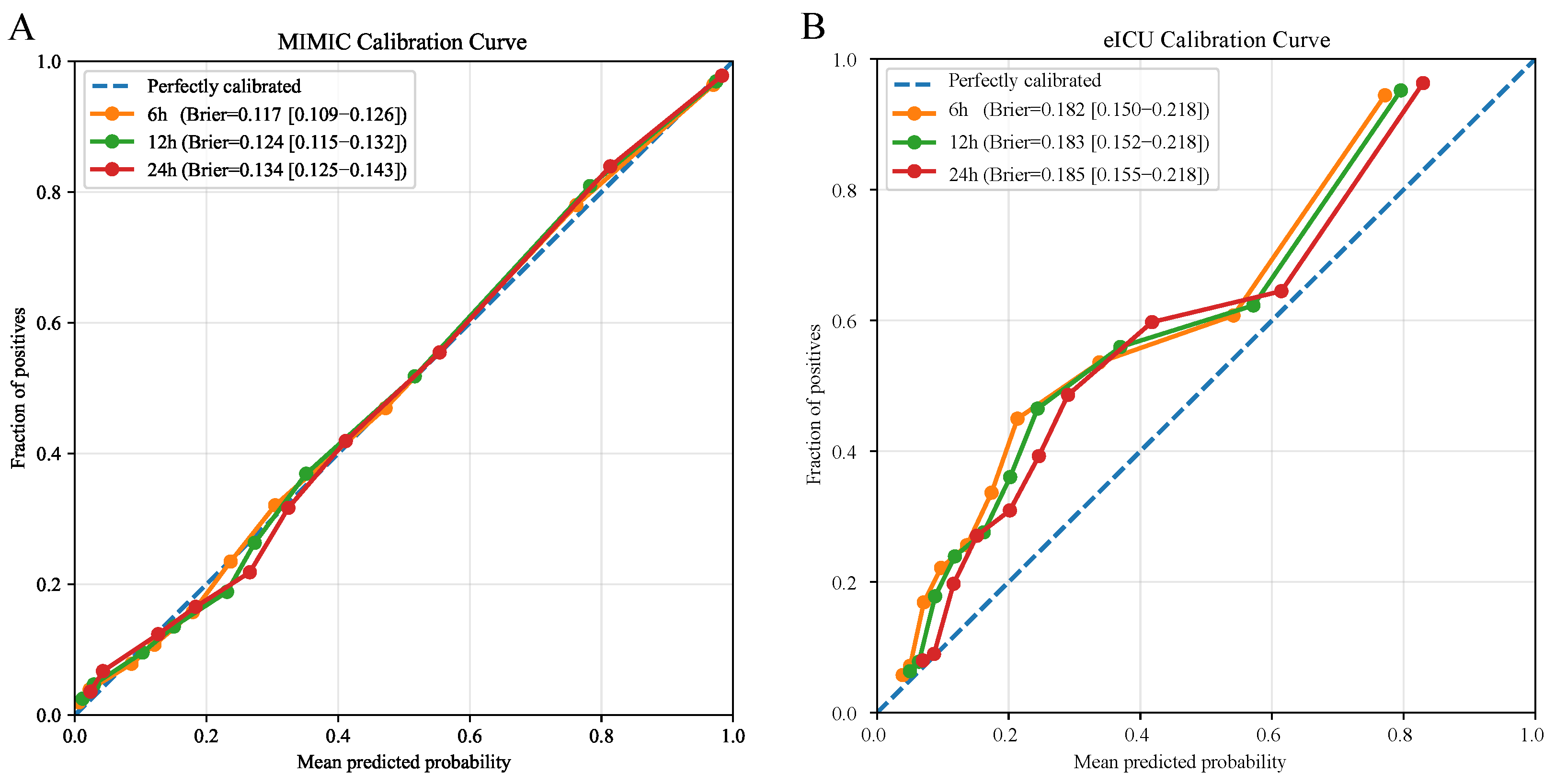
3.4. Model Calibration
3.5. Clinical Threshold Performance
3.6. Model Interpretation
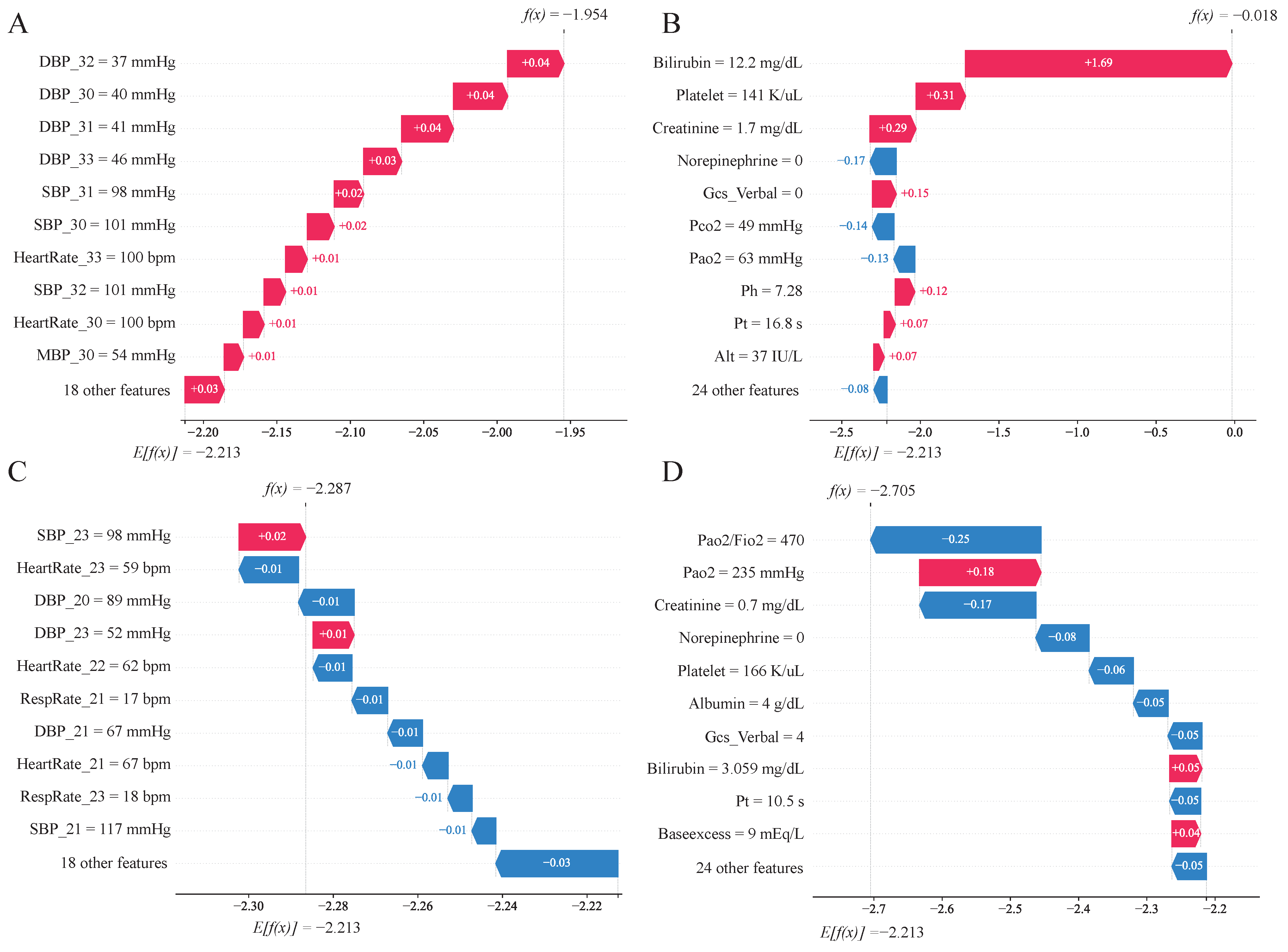
3.6.1. Feature Contributions
3.6.2. Case Analysis
4. Discussion
Study Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MODS | Multiple Organ Dysfunction Syndrome |
| AUC | Area Under the Curve |
| ACC | Accuracy |
| SHAP | SHapley Additive exPlanations |
| CI | Confidence Interval |
| PaO2 | Partial pressure of arterial oxygen |
| PaCO2 | Partial pressure of arterial carbon dioxide |
| FiO2 | Fraction of inspired oxygen |
| PaO2/FiO2 | Oxygenation index |
| GCS | Glasgow Coma Scale |
| SBP | Systolic blood pressure |
| DBP | Diastolic blood pressure |
| ICU LOS | ICU length of stay |
| WBC | White blood cell count |
| PT | Prothrombin time |
| ALT | Alanine aminotransferase |
| AST | Aspartate aminotransferase |
References
- Singer, M.; Deutschman, C.S.; Seymour, C.W.; Shankar-Hari, M.; Annane, D.; Bauer, M.; Bellomo, R.; Bernard, G.R.; Chiche, J.D.; Coopersmith, C.M.; et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016, 315, 801–810. [Google Scholar] [CrossRef]
- Rudd, K.E.; Johnson, S.C.; Agesa, K.M.; Shackelford, K.A.; Tsoi, D.; Kievlan, D.R.; Colombara, D.V.; Ikuta, K.S.; Kissoon, N.; Finfer, S.; et al. Global, Regional, and National Sepsis Incidence and Mortality, 1990–2017: Analysis for the Global Burden of Disease Study. Lancet 2020, 395, 200–211. [Google Scholar] [CrossRef]
- Injuries and Violence. Available online: https://www.who.int/news-room/fact-sheets/detail/injuries-and-violence (accessed on 5 January 2026).
- Raju, R. Immune and Metabolic Alterations Following Trauma and Sepsis—An Overview. Biochim. Biophys. Acta. Mol. Basis Dis. 2017, 1863, 2523–2525. [Google Scholar] [CrossRef]
- Mayr, F.B.; Yende, S.; Angus, D.C. Epidemiology of Severe Sepsis. Virulence 2014, 5, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Mao, Q.; Liu, Y.; Zhang, J.; Li, W.; Zhang, W.; Zhou, C. Blood Virome of Patients with Traumatic Sepsis. Virol. J. 2023, 20, 198. [Google Scholar] [CrossRef] [PubMed]
- Bone, R.C. Immunologic Dissonance: A Continuing Evolution in Our Understanding of the Systemic Inflammatory Response Syndrome (SIRS) and the Multiple Organ Dysfunction Syndrome (MODS). Ann. Intern. Med. 1996, 125, 680–687. [Google Scholar] [CrossRef]
- Sun, G.D.; Zhang, Y.; Mo, S.S.; Zhao, M.Y. Multiple Organ Dysfunction Syndrome Caused by Sepsis: Risk Factor Analysis. Int. J. Gen. Med. 2021, 14, 7159–7164. [Google Scholar] [CrossRef]
- Afessa, B.; Green, B.; Delke, I.; Koch, K. Systemic Inflammatory Response Syndrome, Organ Failure, and Outcome in Critically Ill Obstetric Patients Treated in an ICU. Chest 2001, 120, 1271–1277. [Google Scholar] [CrossRef] [PubMed]
- Vincent, J.L.; Moreno, R.; Takala, J.; Willatts, S.; De Mendonça, A.; Bruining, H.; Reinhart, C.K.; Suter, P.M.; Thijs, L.G. The SOFA (Sepsis-related Organ Failure Assessment) Score to Describe Organ Dysfunction/Failure. On Behalf of the Working Group on Sepsis-Related Problems of the European Society of Intensive Care Medicine. Intensive Care Med. 1996, 22, 707–710. [Google Scholar] [CrossRef]
- Liu, G.; Xu, J.; Wang, C.; Yu, M.; Yuan, J.; Tian, F.; Zhang, G. A Machine Learning Method for Predicting the Probability of MODS Using Only Non-Invasive Parameters. Comput. Methods Programs Biomed. 2022, 227, 107236. [Google Scholar] [CrossRef]
- Liu, X.; DuMontier, C.; Hu, P.; Liu, C.; Yeung, W.; Mao, Z.; Ho, V.; Thoral, P.J.; Kuo, P.C.; Hu, J.; et al. Clinically Interpretable Machine Learning Models for Early Prediction of Mortality in Older Patients with Multiple Organ Dysfunction Syndrome: An International Multicenter Retrospective Study. J. Gerontol. Ser. A Biol. Sci. Med Sci. 2023, 78, 718–726. [Google Scholar] [CrossRef]
- Fan, B.; Klatt, J.; Moor, M.M.; Daniels, L.A.; Swiss Pediatric Sepsis Study; Sanchez-Pinto, L.N.; Agyeman, P.K.A.; Schlapbach, L.J.; Borgwardt, K.M. Prediction of Recovery from Multiple Organ Dysfunction Syndrome in Pediatric Sepsis Patients. Bioinformatics 2022, 38, i101–i108. [Google Scholar] [CrossRef]
- Devlin, J.; Chang, M.W.; Lee, K.; Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. arXiv 2019. [Google Scholar] [CrossRef]
- Howard, J.; Ruder, S. Universal Language Model Fine-tuning for Text Classification. arXiv 2018. [Google Scholar] [CrossRef]
- Li, Y.; Mamouei, M.; Salimi-Khorshidi, G.; Rao, S.; Hassaine, A.; Canoy, D.; Lukasiewicz, T.; Rahimi, K. Hi-BEHRT: Hierarchical Transformer-Based Model for Accurate Prediction of Clinical Events Using Multimodal Longitudinal Electronic Health Records. IEEE J. Biomed. Health Inform. 2023, 27, 1106–1117. [Google Scholar] [CrossRef]
- Huang, D.; Cogill, S.; Hsia, R.Y.; Yang, S.; Kim, D. Development and External Validation of a Pretrained Deep Learning Model for the Prediction of Non-Accidental Trauma. NPJ Digit. Med. 2023, 6, 131. [Google Scholar] [CrossRef] [PubMed]
- Johnson, A.E.; Bulgarelli, L.; Shen, L.; Gayles, A.; Shammout, A.; Horng, S.; Pollard, T.J.; Hao, S.; Moody, B.; Gow, B. MIMIC-IV, a Freely Accessible Electronic Health Record Dataset. Sci. Data 2023, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Pollard, T.J.; Johnson, A.E.; Raffa, J.D.; Celi, L.A.; Mark, R.G.; Badawi, O. The eICU Collaborative Research Database, a Freely Available Multi-Center Database for Critical Care Research. Sci. Data 2018, 5, 180178. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Xu, J.; Wang, C.; Zhang, G.; Wang, H. Method of Non-Invasive Parameters for Predicting the Probability of Early in-Hospital Death of Patients in Intensive Care Unit. Biomed. Signal Process. Control 2022, 73, 103405. [Google Scholar] [CrossRef]
- Seymour, C.W.; Liu, V.X.; Iwashyna, T.J.; Brunkhorst, F.M.; Rea, T.D.; Scherag, A.; Rubenfeld, G.; Kahn, J.M.; Shankar-Hari, M.; Singer, M.; et al. Assessment of Clinical Criteria for Sepsis: For the Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016, 315, 762–774. [Google Scholar] [CrossRef]
- Liu, C.; Yao, Z.; Liu, P.; Tu, Y.; Chen, H.; Cheng, H.; Xie, L.; Xiao, K. Early Prediction of MODS Interventions in the Intensive Care Unit Using Machine Learning. J. Big Data 2023, 10, 55. [Google Scholar] [CrossRef]
- Zhang, G.; Yuan, J.; Yu, M.; Wu, T.; Luo, X.; Chen, F. A Machine Learning Method for Acute Hypotensive Episodes Prediction Using Only Non-Invasive Parameters. Comput. Methods Programs Biomed. 2021, 200, 105845. [Google Scholar] [CrossRef]
- Jakobsen, J.C.; Gluud, C.; Wetterslev, J.; Winkel, P. When and How Should Multiple Imputation Be Used for Handling Missing Data in Randomised Clinical Trials—A Practical Guide with Flowcharts. BMC Med. Res. Methodol. 2017, 17, 162. [Google Scholar] [CrossRef]
- Andrade, C. Z Scores, Standard Scores, and Composite Test Scores Explained. Indian J. Psychol. Med. 2021, 43, 555–557. [Google Scholar] [CrossRef]
- Poslavskaya, E.; Korolev, A. Encoding Categorical Data: Is There yet Anything ’hotter’ than One-Hot Encoding? arXiv 2023. [Google Scholar] [CrossRef]
- Ye, J.; Zhao, J.; Ye, K.; Xu, C. How to Build a Graph-Based Deep Learning Architecture in Traffic Domain: A Survey. IEEE Trans. Intell. Transp. Syst. 2020, 23, 3904–3924. [Google Scholar] [CrossRef]
- Choromanski, K.; Likhosherstov, V.; Dohan, D.; Song, X.; Gane, A.; Sarlos, T.; Hawkins, P.; Davis, J.; Mohiuddin, A.; Kaiser, L.; et al. Rethinking Attention with Performers. arXiv 2022. [Google Scholar] [CrossRef]
- Zeng, A.; Chen, M.; Zhang, L.; Xu, Q. Are Transformers Effective for Time Series Forecasting? In Proceedings of the AAAI Conference on Artificial Intelligence, Washington, DC, USA, 7 February 2023; Volume 37, pp. 11121–11128. [Google Scholar]
- Hochreiter, S.; Schmidhuber, J. Long Short-Term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef]
- Hornik, K.; Stinchcombe, M.; White, H. Multilayer Feedforward Networks Are Universal Approximators. Neural Netw. 1989, 2, 359–366. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.I. A Unified Approach to Interpreting Model Predictions. Adv. Neural Inf. Process. Syst. 2017, 30, 4768–4777. [Google Scholar]
- Peng, J.; Li, Y.; Liu, C.; Mao, Z.; Kang, H.; Zhou, F. Predicting Multiple Organ Dysfunction Syndrome in Trauma-Induced Sepsis: Nomogram and Machine Learning Approaches. J. Intensive Med. 2025, 5, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Vanderschueren, S.; De Weerdt, A.; Malbrain, M.; Vankersschaever, D.; Frans, E.; Wilmer, A.; Bobbaers, H. Thrombocytopenia and Prognosis in Intensive Care. Crit. Care Med. 2000, 28, 1871–1876. [Google Scholar] [CrossRef] [PubMed]
- Delano, M.J.; Ward, P.A. The Immune System’s Role in Sepsis Progression, Resolution, and Long-term Outcome. Immunol. Rev. 2016, 274, 330–353. [Google Scholar] [CrossRef] [PubMed]
- Semple, J.W.; Italiano, J.E.; Freedman, J. Platelets and the Immune Continuum. Nat. Rev. Immunol. 2011, 11, 264–274. [Google Scholar] [CrossRef]
- Zarychanski, R.; Houston, D.S. Assessing Thrombocytopenia in the Intensive Care Unit: The Past, Present, and Future. Hematol. Am. Soc. Hematol. Educ. Program 2017, 2017, 660–666. [Google Scholar] [CrossRef]
- Li, X.; Yuan, F.; Zhou, L. Organ Crosstalk in Acute Kidney Injury: Evidence and Mechanisms. J. Clin. Med. 2022, 11, 6637. [Google Scholar] [CrossRef]
- Goncalves, J.A.; Hydo, L.J.; Barie, P.S. Factors Influencing Outcome of Prolonged Norepinephrine Therapy for Shock in Critical Surgical Illness. Shock 1998, 10, 231–236. [Google Scholar] [CrossRef]
- Xiang, H.; Zhao, Y.; Ma, S.; Li, Q.; Kashani, K.B.; Peng, Z.; Li, J.; Hu, B. Dose-Related Effects of Norepinephrine on Early-Stage Endotoxemic Shock in a Swine Model. J. Intensive Med. 2023, 3, 335–344. [Google Scholar] [CrossRef]
- Xu, X.; Yang, T.; An, J.; Li, B.; Dou, Z. Liver Injury in Sepsis: Manifestations, Mechanisms and Emerging Therapeutic Strategies. Front. Immunol. 2025, 16, 1575554. [Google Scholar] [CrossRef]
- Borges, A.; Bento, L. Organ Crosstalk and Dysfunction in Sepsis. Ann. Intensive Care 2024, 14, 147. [Google Scholar] [CrossRef]
- Hotchkiss, R.S.; Monneret, G.; Payen, D. Immunosuppression in Sepsis: A Novel Understanding of the Disorder and a New Therapeutic Approach. Lancet Infect. Dis. 2013, 13, 260–268. [Google Scholar] [CrossRef] [PubMed]
- Bakker, J.; Gris, P.; Coffernils, M.; Kahn, R.J.; Vincent, J.L. Serial Blood Lactate Levels Can Predict the Development of Multiple Organ Failure Following Septic Shock. Am. J. Surg. 1996, 171, 221–226. [Google Scholar] [CrossRef] [PubMed]





| Features | Details |
|---|---|
| Demographics | Age, Gender, ICU LOS |
| Vital Signs | Heart rate *, Mean Blood Pressure (MBP) *, Systolic Blood Pressure (SBP) *, Diastolic Blood Pressure (DBP) *, Temperature *, SpO2 *, Respiratory Rate *, GCS, GCS Motor, GCS Verbal, GCS Eyes |
| Biochemical Markers | PaO2, Hematocrit, WBC, Creatinine, BUN, Sodium, Albumin, Bilirubin, Glucose, pH, PaCO2, PaO2/FiO2, Platelet, PT, Potassium, ALT, AST, Base Excess, Chloride, Total CO2, Lactate, Free Calcium, FiO2 |
| Medication Records | Epinephrine, Norepinephrine, Dopamine, Dobutamine |
| Variables | Overall (3502) | MODS (1854) | Non-MODS (1648) | p Value |
|---|---|---|---|---|
| Age (mean ± std in years) | 64.79 ± 19.73 | 64.97 ± 18.51 | 64.58 ± 21.02 | 0.557 |
| Gender (males) | 2237 (63.88%) | 1251 (67.48%) | 986 (59.83%) | <0.001 |
| ICU length of stay (days) | 6.04 ± 5.95 | 8.16 ± 6.68 | 3.65 ± 3.79 | <0.001 |
| Hospital length of stay (days) | 15.61 ± 18.66 | 19.21 ± 22.73 | 11.55 ± 11.29 | <0.001 |
| ICU type (%) | <0.001 | |||
| CCU | 230 (6.57%) | 155 (8.36%) | 75 (4.55%) | |
| SICU | 532 (15.19%) | 250 (13.48%) | 282 (17.11%) | |
| Neuro Stepdown | 36 (1.03%) | 7 (0.38%) | 29 (1.76%) | |
| Neuro Intermediate | 55 (1.57%) | 7 (0.38%) | 48 (2.91%) | |
| CVICU | 150 (4.28%) | 96 (5.18%) | 54 (3.28%) | |
| Neuro SICU | 126 (3.6%) | 59 (3.18%) | 67 (4.07%) | |
| TSICU | 1389 (39.66%) | 718 (38.73%) | 671 (40.72%) | |
| MICU | 686 (19.59%) | 400 (21.57%) | 286 (17.35%) | |
| MICU/SICU | 298 (8.51%) | 162 (8.74%) | 136 (8.25%) | |
| Ethnicity (%) | 0.005 | |||
| Asian | 73 (2.08%) | 42 (2.27%) | 31 (1.88%) | |
| Black | 217 (6.2%) | 105 (5.66%) | 112 (6.8%) | |
| Hispanic | 119 (3.4%) | 64 (3.45%) | 55 (3.34%) | |
| Other/Unknown | 800 (22.84%) | 468 (25.24%) | 332 (20.15%) | |
| White | 2293 (65.48%) | 1175 (63.38%) | 1118 (67.84%) | |
| Mortality | 711 (20.30%) | 541 (29.18%) | 170 (10.32%) | <0.001 |
| Model | Window | AUC (95%CI) | ACC (95%CI) | TPR (95%CI) | TNR (95%CI) |
|---|---|---|---|---|---|
| PT-MLP·LSTM-eICU | 6 h | 0.913 (0.907, 0.920) | 0.831 (0.818, 0.842) | 0.829 (0.814, 0.845) | 0.831 (0.810, 0.850) |
| 12 h | 0.907 (0.901, 0.914) | 0.823 (0.811, 0.836) | 0.829 (0.806, 0.839) | 0.820 (0.803, 0.844) | |
| 24 h | 0.899 (0.891, 0.906) | 0.816 (0.807, 0.825) | 0.817 (0.799, 0.828) | 0.815 (0.800, 0.830) | |
| MLP·LSTM | 6 h | 0.884 (0.876, 0.892) | 0.810 (0.803, 0.818) | 0.812 (0.797, 0.825) | 0.809 (0.797, 0.816) |
| 12 h | 0.879 (0.870, 0.887) | 0.803 (0.794, 0.809) | 0.806 (0.795, 0.817) | 0.801 (0.790, 0.809) | |
| 24 h | 0.870 (0.860, 0.879) | 0.795 (0.785, 0.802) | 0.794 (0.783, 0.801) | 0.796 (0.784, 0.807) | |
| PT-LSTM-eICU | 6 h | 0.904 (0.897, 0.911) | 0.824 (0.812, 0.830) | 0.815 (0.804, 0.824) | 0.829 (0.820, 0.839) |
| 12 h | 0.898 (0.890, 0.905) | 0.813 (0.804, 0.822) | 0.813 (0.803, 0.824) | 0.814 (0.803, 0.823) | |
| 24 h | 0.889 (0.879, 0.897) | 0.804 (0.792, 0.81) | 0.800 (0.785, 0.816) | 0.807 (0.794, 0.817) | |
| PT-MLP-eICU | 6 h | 0.894 (0.886, 0.901) | 0.811 (0.804, 0.823) | 0.819 (0.802, 0.824) | 0.808 (0.802, 0.825) |
| 12 h | 0.888 (0.879, 0.895) | 0.810 (0.798, 0.817) | 0.803 (0.796, 0.818) | 0.813 (0.796, 0.819) | |
| 24 h | 0.879 (0.870, 0.887) | 0.801 (0.788, 0.808) | 0.793 (0.786, 0.810) | 0.805 (0.786, 0.811) | |
| PT-MLP·LSTM | 6 h | 0.891 (0.883, 0.898) | 0.809 (0.786, 0.826) | 0.806 (0.776, 0.819) | 0.811 (0.767, 0.844) |
| 12 h | 0.885 (0.876, 0.893) | 0.802 (0.783, 0.818) | 0.802 (0.776, 0.823) | 0.802 (0.765, 0.821) | |
| 24 h | 0.876 (0.867, 0.885) | 0.794 (0.779, 0.804) | 0.792 (0.774, 0.837) | 0.795 (0.766, 0.836) |
| Model | Pre-Train | MLP | LSTM | eICU | MIMIC-IV AUC (Mean) | eICU AUC (Mean) |
|---|---|---|---|---|---|---|
| PT-MLP·LSTM-eICU | ✓ | ✓ | ✓ | ✓ | 0.906 | 0.809 |
| MLP·LSTM | ✓ | ✓ | 0.878 | 0.735 | ||
| PT-LSTM-eICU | ✓ | ✓ | ✓ | 0.897 | 0.800 | |
| PT-MLP-eICU | ✓ | ✓ | ✓ | 0.887 | 0.788 | |
| PT-MLP·LSTM | ✓ | ✓ | ✓ | 0.884 | 0.722 |
| 6-h Prediction Window | 12-h Prediction Window | 24-h Prediction Window | |||||
|---|---|---|---|---|---|---|---|
| Dataset | Method | AUC | ACC | AUC | ACC | AUC | ACC |
| MIMIC-IV | SOFA-only | 0.748 (0.700, 0.789) | 0.734 (0.675, 0.787) | 0.736 (0.685, 0.783) | 0.725 (0.670, 0.776) | 0.727 (0.675, 0.777) | 0.711 (0.660, 0.758) |
| SOFA-var | 0.880 (0.863, 0.895) | 0.806 (0.788, 0.823) | 0.869 (0.851, 0.886) | 0.795 (0.776, 0.813) | 0.854 (0.834, 0.873) | 0.777 (0.760, 0.799) | |
| noSOFA-var | 0.809 (0.782, 0.833) | 0.730 (0.705, 0.755) | 0.805 (0.778, 0.829) | 0.728 (0.701, 0.748) | 0.800 (0.772, 0.824) | 0.715 (0.692, 0.738) | |
| ALL-var | 0.913 (0.907, 0.920) | 0.831 (0.818, 0.842) | 0.907 (0.901, 0.914) | 0.823 (0.811, 0.836) | 0.899 (0.891, 0.906) | 0.816 (0.807, 0.825) | |
| eICU | SOFA-only | 0.730 (0.652, 0.816) | 0.730 (0.646, 0.810) | 0.722 (0.628, 0.820) | 0.732 (0.653, 0.811) | 0.681 (0.559, 0.797) | 0.722 (0.636, 0.805) |
| SOFA-var | 0.771 (0.716, 0.822) | 0.680 (0.619, 0.735) | 0.764 (0.711, 0.819) | 0.666 (0.605, 0.723) | 0.759 (0.700, 0.813) | 0.642 (0.572, 0.703) | |
| noSOFA-var | 0.756 (0.702, 0.809) | 0.642 (0.582, 0.699) | 0.749 (0.695, 0.803) | 0.633 (0.574, 0.69) | 0.742 (0.688, 0.794) | 0.630 (0.574, 0.685) | |
| ALL-var | 0.812 (0.757, 0.864) | 0.735 (0.681, 0.784) | 0.810 (0.754, 0.862) | 0.734 (0.679, 0.782) | 0.805 (0.748, 0.857) | 0.724 (0.673, 0.777) | |
| 6-h Prediction Window | 12-h Prediction Window | 24-h Prediction Window | |||||
|---|---|---|---|---|---|---|---|
| Dataset | Threshold | Sensitivity | Specificity | Sensitivity | Specificity | Sensitivity | Specificity |
| MIMIC-IV | 10% | 0.909 (0.899, 0.919) | 0.676 (0.660, 0.691) | 0.948 (0.941, 0.955) | 0.523 (0.506, 0.540) | 0.967 (0.962, 0.972) | 0.376 (0.358, 0.393) |
| 20% | 0.630 (0.605, 0.656) | 0.956 (0.950, 0.961) | 0.726 (0.705, 0.748) | 0.898 (0.890, 0.906) | 0.816 (0.799, 0.832) | 0.791 (0.779, 0.803) | |
| 30% | 0.411 (0.380, 0.442) | 0.991 (0.989, 0.993) | 0.431 (0.401, 0.461) | 0.990 (0.988, 0.992) | 0.478 (0.449, 0.507) | 0.985 (0.982, 0.997) | |
| eICU | 10% | 0.881 (0.825, 0.931) | 0.494 (0.390, 0.596) | 0.913 (0.862, 0.958) | 0.439 (0.335, 0.546) | 0.958 (0.923, 0.987) | 0.304 (0.204, 0.415) |
| 20% | 0.666 (0.564, 0.764) | 0.785 (0.705, 0.856) | 0.748 (0.665, 0.830) | 0.714 (0.618, 0.796) | 0.807 (0.738, 0.871) | 0.614 (0.511, 0.706) | |
| 30% | 0.533 (0.421, 0.644) | 0.879 (0.810, 0.932) | 0.550 (0.443, 0.656) | 0.970 (0.800, 0.925) | 0.579 (0.475, 0.683) | 0.844 (0.768, 0.906) | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Wen, J.; Liu, G.; Chang, P.; Hu, P.; Liu, B.; Jiang, C.; Xu, X.; Ma, J.; Zhang, G. A Real-Time Dynamic Warning Method for MODS in Trauma Sepsis Patients Based on a Pre-Trained Transfer Learning Algorithm. Diagnostics 2026, 16, 270. https://doi.org/10.3390/diagnostics16020270
Wen J, Liu G, Chang P, Hu P, Liu B, Jiang C, Xu X, Ma J, Zhang G. A Real-Time Dynamic Warning Method for MODS in Trauma Sepsis Patients Based on a Pre-Trained Transfer Learning Algorithm. Diagnostics. 2026; 16(2):270. https://doi.org/10.3390/diagnostics16020270
Chicago/Turabian StyleWen, Jiahe, Guanjun Liu, Panpan Chang, Pan Hu, Bin Liu, Chunliang Jiang, Xiaoyun Xu, Jun Ma, and Guang Zhang. 2026. "A Real-Time Dynamic Warning Method for MODS in Trauma Sepsis Patients Based on a Pre-Trained Transfer Learning Algorithm" Diagnostics 16, no. 2: 270. https://doi.org/10.3390/diagnostics16020270
APA StyleWen, J., Liu, G., Chang, P., Hu, P., Liu, B., Jiang, C., Xu, X., Ma, J., & Zhang, G. (2026). A Real-Time Dynamic Warning Method for MODS in Trauma Sepsis Patients Based on a Pre-Trained Transfer Learning Algorithm. Diagnostics, 16(2), 270. https://doi.org/10.3390/diagnostics16020270







