Artificial Intelligence for Fibrosis Diagnosis in Metabolic-Dysfunction-Associated Steatotic Liver Disease: A Systematic Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Selection
2.2. Risk of Bias and Quality of Studies
2.3. Data Extraction and Synthesis Methods
3. Results
3.1. Risk of Bias
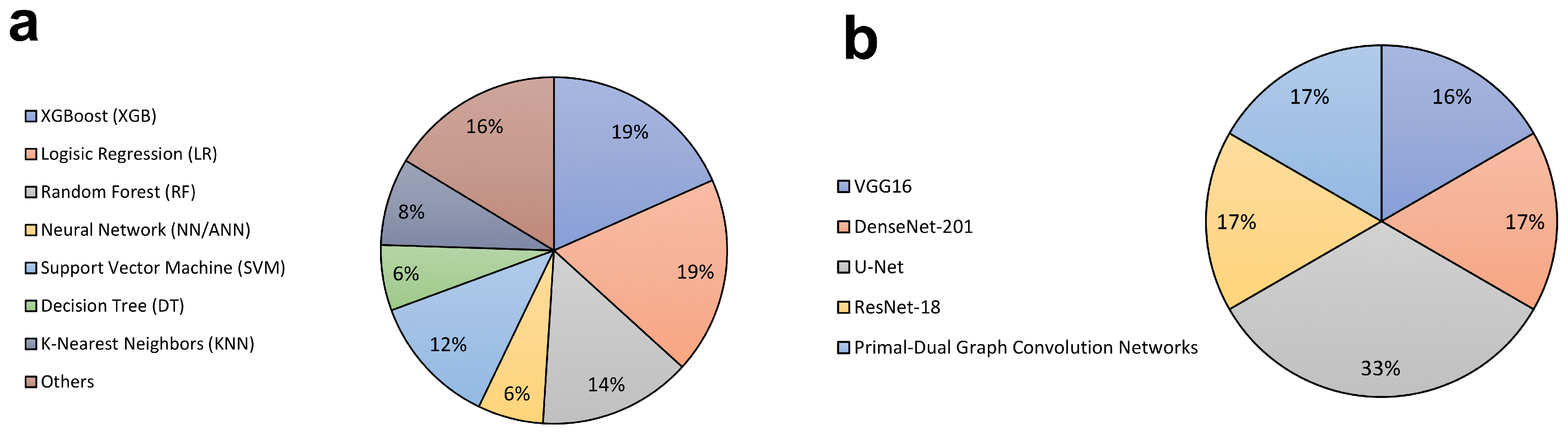
3.2. Model Architecture
3.3. Model Performance
3.4. Model Evaluation
3.5. Most Frequently Used Features Across AI Models
4. Discussion
4.1. AI-Assisted Imaging and Radiomics for Liver Fibrosis: A Complementary Perspective
4.2. Study Transparency
4.3. Limitations of This Study
4.4. Implementation Feasibility of AI Models in Clinical Practice
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| ALT | Alanine Aminotransferase |
| ANN | Artificial Neural Network |
| AST | Aspartate Aminotransferase |
| AUROC (AUC) | Area Under the Receiver Operating Characteristic Curve |
| BLDA | Bayesian Linear Discriminant Analysis |
| BMI | Body Mass Index |
| CNN | Convolutional Neural Network |
| DT | Decision Tree |
| FF | Feed-Forward Neural Network |
| FIB-4 | Fibrosis-4 Index |
| GBM | Gradient Boosting Machine |
| GGT | Gamma-Glutamyl Transferase |
| GNB | Gaussian Naive Bayes |
| HDL | High-Density Lipoprotein |
| KNN | K-Nearest Neighbors |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| LR | Logistic Regression |
| LSN | Liver Surface Nodularity |
| MASH | Metabolic-Dysfunction-Associated Steatohepatitis |
| MASLD | Metabolic-Dysfunction-Associated Steatotic Liver Disease |
| MINIMAR | Minimum Information for Medical AI Reporting |
| MRE | Magnetic Resonance Elastography |
| NAIF | NAFLD Artificial Intelligence Fibrosis Model |
| NB | Naive Bayes |
| NFS | NAFLD Fibrosis Score |
| NITs | Non-Invasive Tests |
| NN | Neural Network |
| PDGCN | Primal-Dual Graph Convolution Networks |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses |
| QUADAS-2 | Quality Assessment of Diagnostic Accuracy Studies, version 2 |
| RF | Random Forest |
| SAFE | Steatosis-Associated Fibrosis Estimator |
| SR | Systematic Review |
| SVM | Support Vector Machine |
| T2DM | Type 2 Diabetes Mellitus |
| VCTE | Vibration-Controlled Transient Elastography |
| VGG16 | Visual Geometry Group 16 |
| XGBoost (XGB) | Extreme Gradient Boosting |
References
- Chan, W.-K.; Chuah, K.-H.; Rajaram, R.B.; Lim, L.-L.; Ratnasingam, J.; Vethakkan, S.R. Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD): A State-of-the-Art Review. J. Obes. Metab. Syndr. 2023, 32, 197–213. [Google Scholar] [CrossRef]
- Younossi, Z.M.; Kalligeros, M.; Henry, L. Epidemiology of Metabolic Dysfunction-Associated Steatotic Liver Disease. Clin. Mol. Hepatol. 2025, 31, S32–S50. [Google Scholar] [CrossRef]
- Li, Y.; Yang, P.; Ye, J.; Xu, Q.; Wu, J.; Wang, Y. Updated Mechanisms of MASLD Pathogenesis. Lipids Health Dis. 2024, 23, 117. [Google Scholar] [CrossRef] [PubMed]
- Goodman, Z.D. Grading and Staging Systems for Inflammation and Fibrosis in Chronic Liver Diseases. J. Hepatol. 2007, 47, 598–607. [Google Scholar] [CrossRef] [PubMed]
- Sumida, Y.; Nakajima, A.; Itoh, Y. Limitations of Liver Biopsy and Non-Invasive Diagnostic Tests for the Diagnosis of Nonalcoholic Fatty Liver Disease/Nonalcoholic Steatohepatitis. World J. Gastroenterol. 2014, 20, 475–485. [Google Scholar] [CrossRef]
- Patel, K.; Sebastiani, G. Limitations of Non-Invasive Tests for Assessment of Liver Fibrosis. JHEP Rep. 2020, 2, 100067. [Google Scholar] [CrossRef] [PubMed]
- Al Kuwaiti, A.; Nazer, K.; Al-Reedy, A.; Al-Shehri, S.; Al-Muhanna, A.; Subbarayalu, A.V.; Al Muhanna, D.; Al-Muhanna, F.A. A Review of the Role of Artificial Intelligence in Healthcare. J. Pers. Med. 2023, 13, 951. [Google Scholar] [CrossRef]
- Liu, X.; Faes, L.; Kale, A.U.; Wagner, S.K.; Fu, D.J.; Bruynseels, A.; Mahendiran, T.; Moraes, G.; Shamdas, M.; Kern, C.; et al. A Comparison of Deep Learning Performance against Health-Care Professionals in Detecting Diseases from Medical Imaging: A Systematic Review and Meta-Analysis. Lancet Digit. Health 2019, 1, e271–e297. [Google Scholar] [CrossRef]
- Pugliese, N.; Bertazzoni, A.; Hassan, C.; Schattenberg, J.M.; Aghemo, A. Revolutionizing MASLD: How Artificial Intelligence Is Shaping the Future of Liver Care. Cancers 2025, 17, 722. [Google Scholar] [CrossRef]
- Njei, B.; Osta, E.; Njei, N.; Al-Ajlouni, Y.A.; Lim, J.K. An Explainable Machine Learning Model for Prediction of High-Risk Nonalcoholic Steatohepatitis. Sci. Rep. 2024, 14, 8589. [Google Scholar] [CrossRef]
- Yin, C.; Zhang, H.; Du, J.; Zhu, Y.; Zhu, H.; Yue, H. Artificial Intelligence in Imaging for Liver Disease Diagnosis. Front. Med. 2025, 12, 1591523. [Google Scholar] [CrossRef]
- Wakabayashi, S.; Kimura, T.; Tamaki, N.; Iwadare, T.; Okumura, T.; Kobayashi, H.; Yamashita, Y.; Tanaka, N.; Kurosaki, M.; Umemura, T. AI-Based Platelet-Independent Noninvasive Test for Liver Fibrosis in MASLD Patients. JGH Open 2025, 9, e70150. [Google Scholar] [CrossRef]
- Hernandez-Boussard, T.; Bozkurt, S.; Ioannidis, J.P.A.; Shah, N.H. MINIMAR (MINimum Information for Medical AI Reporting): Developing Reporting Standards for Artificial Intelligence in Health Care. J. Am. Med. Inform. Assoc. 2020, 27, 2011–2015. [Google Scholar] [CrossRef]
- Alkhouri, N.; Cheuk-Fung Yip, T.; Castera, L.; Takawy, M.; Adams, L.A.; Verma, N.; Arab, J.P.; Jafri, S.-M.; Zhong, B.; Dubourg, J.; et al. ALADDIN: A Machine Learning Approach to Enhance the Prediction of Significant Fibrosis or Higher in Metabolic Dysfunction-Associated Steatotic Liver Disease. Am. J. Gastroenterol. 2022. [Google Scholar] [CrossRef] [PubMed]
- Dabbah, S.; Mishani, I.; Davidov, Y.; Ben Ari, Z. Implementation of Machine Learning Algorithms to Screen for Advanced Liver Fibrosis in Metabolic Dysfunction-Associated Steatotic Liver Disease: An In-Depth Explanatory Analysis. Digestion 2025, 106, 189–202. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.; Yu, N.; Li, G.; Arshad, T.; Liu, W.-Y.; Wong, G.L.-H.; Liang, X.; Chen, Y.; Jin, X.-Z.; Leung, H.H.-W.; et al. Machine-Learning Model Comprising Five Clinical Indices and Liver Stiffness Measurement Can Accurately Identify MASLD-Related Liver Fibrosis. Liver Int. 2024, 44, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Zheng, K.I.; Li, Y.-Y.; Rios, R.S.; Zhu, P.-W.; Pan, X.-Y.; Li, G.; Ma, H.-L.; Tang, L.-J.; Byrne, C.D.; et al. Machine Learning Algorithm Outperforms Fibrosis Markers in Predicting Significant Fibrosis in Biopsy-Confirmed NAFLD. J. Hepatobiliary Pancreat. Sci. 2021, 28, 593–603. [Google Scholar] [CrossRef]
- Ginter-Matuszewska, B.; Adamek, A.; Majchrzak, M.; Rozplochowski, B.; Zientarska, A.; Kowala-Piaskowska, A.; Lukasiak, P. FibrAIm—The Machine Learning Approach to Identify the Early Stage of Liver Fibrosis and Steatosis. Int. J. Med. Inform. 2025, 197, 105837. [Google Scholar] [CrossRef]
- Hassoun, S.; Bruckmann, C.; Ciardullo, S.; Perseghin, G.; Marra, F.; Curto, A.; Arena, U.; Broccolo, F.; Di Gaudio, F. NAIF: A Novel Artificial Intelligence-Based Tool for Accurate Diagnosis of Stage F3/F4 Liver Fibrosis in the General Adult Population, Validated with Three External Datasets. Int. J. Med. Inform. 2024, 185, 105373. [Google Scholar] [CrossRef]
- Lu, C.-H.; Wang, W.; Li, Y.-C.J.; Chang, I.-W.; Chen, C.-L.; Su, C.-W.; Chang, C.-C.; Kao, W.-Y. Machine Learning Models for Predicting Significant Liver Fibrosis in Patients with Severe Obesity and Nonalcoholic Fatty Liver Disease. Obes. Surg. 2024, 34, 4393–4404. [Google Scholar] [CrossRef]
- Mamandipoor, B.; Wernly, S.; Semmler, G.; Flamm, M.; Jung, C.; Aigner, E.; Datz, C.; Wernly, B.; Osmani, V. Machine Learning Models Predict Liver Steatosis but Not Liver Fibrosis in a Prospective Cohort Study. Clin. Res. Hepatol. Gastroenterol. 2023, 47, 102181. [Google Scholar] [CrossRef]
- Okanoue, T.; Shima, T.; Mitsumoto, Y.; Umemura, A.; Yamaguchi, K.; Itoh, Y.; Yoneda, M.; Nakajima, A.; Mizukoshi, E.; Kaneko, S.; et al. Artificial Intelligence/Neural Network System for the Screening of Nonalcoholic Fatty Liver Disease and Nonalcoholic Steatohepatitis. Hepatol. Res. 2021, 51, 554–569. [Google Scholar] [CrossRef]
- Sang, C.; Yan, H.; Chan, W.K.; Zhu, X.; Sun, T.; Chang, X.; Xia, M.; Sun, X.; Hu, X.; Gao, X.; et al. Diagnosis of Fibrosis Using Blood Markers and Logistic Regression in Southeast Asian Patients With Non-Alcoholic Fatty Liver Disease. Front. Med. 2021, 8, 637652. [Google Scholar] [CrossRef] [PubMed]
- Suárez, M.; Martínez, R.; Torres, A.M.; Torres, B.; Mateo, J. A Machine Learning Method to Identify the Risk Factors for Liver Fibrosis Progression in Nonalcoholic Steatohepatitis. Dig. Dis. Sci. 2023, 68, 3801–3809. [Google Scholar] [CrossRef]
- Suárez, M.; Martínez, R.; Torres, A.M.; Ramón, A.; Blasco, P.; Mateo, J. A Machine Learning-Based Method for Detecting Liver Fibrosis. Diagnostics 2023, 13, 2952. [Google Scholar] [CrossRef] [PubMed]
- Verma, N.; Duseja, A.; Mehta, M.; De, A.; Lin, H.; Wong, V.W.-S.; Wong, G.L.-H.; Rajaram, R.B.; Chan, W.-K.; Mahadeva, S.; et al. Machine Learning Improves the Prediction of Significant Fibrosis in Asian Patients with Metabolic Dysfunction-Associated Steatotic Liver Disease—The Gut and Obesity in Asia (GO-ASIA) Study. Aliment. Pharmacol. Ther. 2024, 59, 774–788. [Google Scholar] [CrossRef] [PubMed]
- Xiong, F.-X.; Sun, L.; Zhang, X.-J.; Chen, J.-L.; Zhou, Y.; Ji, X.-M.; Meng, P.-P.; Wu, T.; Wang, X.-B.; Hou, Y.-X. Machine Learning-Based Models for Advanced Fibrosis in Non-Alcoholic Steatohepatitis Patients: A Cohort Study. World J. Gastroenterol. 2025, 31, 101383. [Google Scholar] [CrossRef]
- Yamaguchi, K.; Shima, T.; Mitsumoto, Y.; Seko, Y.; Umemura, A.; Itoh, Y.; Nakajima, A.; Kaneko, S.; Harada, K.; Watkins, T.; et al. Fibro-Scope V1.0.1: An Artificial Intelligence/Neural Network System for Staging of Nonalcoholic Steatohepatitis. Hepatol. Int. 2023, 17, 573–583. [Google Scholar] [CrossRef]
- Cunha, G.M.; Delgado, T.I.; Middleton, M.S.; Liew, S.; Henderson, W.C.; Batakis, D.; Wang, K.; Loomba, R.; Huss, R.S.; Myers, R.P.; et al. Automated CNN-Based Analysis Versus Manual Analysis for MR Elastography in Nonalcoholic Fatty Liver Disease: Intermethod Agreement and Fibrosis Stage Discriminative Performance. Am. J. Roentgenol. 2022, 219, 224–232. [Google Scholar] [CrossRef]
- Lu, X.-Z.; Hu, H.-T.; Li, W.; Deng, J.-F.; Chen, L.; Cheng, M.-Q.; Huang, H.; Ke, W.-P.; Wang, W.; Sun, B.-G. Exploring Hepatic Fibrosis Screening via Deep Learning Analysis of Tongue Images. J. Tradit. Complement. Med. 2024, 14, 544–549. [Google Scholar] [CrossRef]
- Naik, S.N.; Forlano, R.; Manousou, P.; Goldin, R.; Angelini, E.D. Fibrosis Severity Scoring on Sirius Red Histology with Multiple-Instance Deep Learning. Biol. Imaging 2023, 3, e17. [Google Scholar] [CrossRef]
- Preechathammawong, N.; Charoenpitakchai, M.; Wongsason, N.; Karuehardsuwan, J.; Prasoppokakorn, T.; Pitisuttithum, P.; Sanpavat, A.; Yongsiriwit, K.; Aribarg, T.; Chaisiriprasert, P.; et al. Development of a Diagnostic Support System for the Fibrosis of Nonalcoholic Fatty Liver Disease Using Artificial Intelligence and Deep Learning. Kaohsiung J. Med. Sci. 2024, 40, 757–765. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.; Liu, S.; Lyu, F.; Lu, J.; Darkner, S.; Wong, V.W.-S.; Yuen, P.C. XFibrosis: Explicit Vessel-Fiber Modeling for Fibrosis Staging from Liver Pathology Images. In Proceedings of the 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Seattle, WA, USA, 17–21 June 2024; IEEE: Seattle, WA, USA, 2024; pp. 11282–11291. [Google Scholar]
- Zhan, H.; Chen, S.; Gao, F.; Wang, G.; Chen, S.-D.; Xi, G.; Yuan, H.-Y.; Li, X.; Liu, W.-Y.; Byrne, C.D.; et al. AutoFibroNet: A Deep Learning and Multi-Photon Microscopy-Derived Automated Network for Liver Fibrosis Quantification in MAFLD. Aliment. Pharmacol. Ther. 2023, 58, 573–584. [Google Scholar] [CrossRef]
- Bates, S.; Hastie, T.; Tibshirani, R. Cross-Validation: What Does It Estimate and How Well Does It Do It? J. Am. Stat. Assoc. 2024, 119, 1434–1445. [Google Scholar] [CrossRef] [PubMed]
- Karagoz, M.A.; Akay, B.; Basturk, A.; Karaboga, D.; Nalbantoglu, O.U. An Unsupervised Transfer Learning Model Based on Convolutional Auto Encoder for Non-Alcoholic Steatohepatitis Activity Scoring and Fibrosis Staging of Liver Histopathological Images. Neural Comput. Appl. 2023, 35, 10605–10619. [Google Scholar] [CrossRef]
- Jana, A.; Qu, H.; Rattan, P.; Minacapelli, C.D.; Rustgi, V.; Metaxas, D. Deep Learning Based NAS Score and Fibrosis Stage Prediction from CT and Pathology Data. arXiv 2020, arXiv:2009.10687. [Google Scholar] [CrossRef]
- Kayaaltı, Ö.; Aksebzeci, B.H.; Karahan, İ.Ö.; Deniz, K.; Öztürk, M.; Yılmaz, B.; Kara, S.; Asyalı, M.H. Liver Fibrosis Staging Using CT Image Texture Analysis and Soft Computing. Appl. Soft Comput. 2014, 25, 399–413. [Google Scholar] [CrossRef]
- Natarajan, R.; Swathika, R.; V, R.; Antony, S.S. Predictive Modeling for Non Alcoholic Fatty Liver Disease Detection. In Proceedings of the 2025 11th International Conference on Communication and Signal Processing (ICCSP), Melmaruvathur, India, 5–7 June 2025; pp. 1026–1031. [Google Scholar]
- Tang, M.; Wu, Y.; Hu, N.; Lin, C.; He, J.; Xia, X.; Yang, M.; Lei, P.; Luo, P. A Combination Model of CT-Based Radiomics and Clinical Biomarkers for Staging Liver Fibrosis in the Patients with Chronic Liver Disease. Sci. Rep. 2024, 14, 20230. [Google Scholar] [CrossRef]
- Pickhardt, P.J.; Lubner, M.G. Noninvasive Quantitative CT for Diffuse Liver Diseases: Steatosis, Iron Overload, and Fibrosis. Radiographics 2025, 45, e240176. [Google Scholar] [CrossRef]
- Heo, S.; Kim, D.W.; Choi, S.H.; Kim, S.W.; Jang, J.K. Diagnostic Performance of Liver Fibrosis Assessment by Quantification of Liver Surface Nodularity on Computed Tomography and Magnetic Resonance Imaging: Systematic Review and Meta-Analysis. Eur. Radiol. 2022, 32, 3377–3387. [Google Scholar] [CrossRef]
- Roe, K.D.; Jawa, V.; Zhang, X.; Chute, C.G.; Epstein, J.A.; Matelsky, J.; Shpitser, I.; Taylor, C.O. Feature Engineering with Clinical Expert Knowledge: A Case Study Assessment of Machine Learning Model Complexity and Performance. PLoS ONE 2020, 15, e0231300. [Google Scholar] [CrossRef]
- Kapoor, S.; Narayanan, A. Leakage and the Reproducibility Crisis in Machine-Learning-Based Science. Patterns 2023, 4, 100804. [Google Scholar] [CrossRef] [PubMed]
- Berisha, V.; Krantsevich, C.; Hahn, P.R.; Hahn, S.; Dasarathy, G.; Turaga, P.; Liss, J. Digital Medicine and the Curse of Dimensionality. npj Digit. Med. 2021, 4, 153. [Google Scholar] [CrossRef]
- Clusmann, J.; Balaguer-Montero, M.; Bassegoda, O.; Schneider, C.V.; Seraphin, T.; Paintsil, E.; Luedde, T.; Lopez, R.P.; Calderaro, J.; Gilbert, S.; et al. The Barriers to Uptake of Artificial Intelligence in Hepatology and How to Overcome Them. J. Hepatol. 2025, 83, 1410–1426. [Google Scholar] [CrossRef]
- Decharatanachart, P.; Chaiteerakij, R.; Tiyarattanachai, T.; Treeprasertsuk, S. Application of Artificial Intelligence in Chronic Liver Diseases: A Systematic Review and Meta-Analysis. BMC Gastroenterol. 2021, 21, 10. [Google Scholar] [CrossRef] [PubMed]
- Abbas, Q.; Jeong, W.; Lee, S.W. Explainable AI in Clinical Decision Support Systems: A Meta-Analysis of Methods, Applications, and Usability Challenges. Healthcare 2025, 13, 2154. [Google Scholar] [CrossRef]
- Balsano, C.; Burra, P.; Duvoux, C.; Alisi, A.; Piscaglia, F.; Gerussi, A.; Special Interest Group (SIG) Artificial Intelligence and Liver Disease; Italian Association for the Study of Liver (AISF). Artificial Intelligence and Liver: Opportunities and Barriers. Dig. Liver Dis. 2023, 55, 1455–1461. [Google Scholar] [CrossRef] [PubMed]
- Popa, S.L.; Ismaiel, A.; Abenavoli, L.; Padureanu, A.M.; Dita, M.O.; Bolchis, R.; Munteanu, M.A.; Brata, V.D.; Pop, C.; Bosneag, A.; et al. Diagnosis of Liver Fibrosis Using Artificial Intelligence: A Systematic Review. Medicina 2023, 59, 992. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef]


| Parameters | Inclusion Criteria | Exclusion Criteria |
|---|---|---|
| Population | Patients ≥ 18 years old diagnosed with MASLD | Studies in animal models, in vitro studies, pediatric populations, or other liver diseases |
| Intervention | Applications of artificial intelligence (AI), such as machine learning, deep learning, or neural networks, for the assessment or prediction of liver fibrosis | Studies that do not use AI tools as a central part of the analysis |
| Comparator | Studies with or without a comparator group. When present, comparisons with traditional methods (e.g., elastography, biopsy, clinical scores) | Studies that only describe imaging methods or laboratory tests without a link to AI |
| Outcome | Performance of AI in identifying, classifying, or predicting the degree of liver fibrosis; diagnostic accuracy; sensitivity/specificity; AUROC | Studies without analysis of clinical or predictive outcomes, or that do not report AI performance metrics |
| Study Type | Original articles from retrospective or prospective studies, cohorts, validation studies, or cross-sectional studies with real clinical data (medical records, imaging, histology, laboratory tests) used in AI models | Abstracts, systematic reviews, editorials, letters to the editor, study protocols, commentaries, or opinion articles based exclusively on theoretical simulations |
| Performance Assessment | Presence of quantitative metrics such as AUROC, sensitivity, specificity, and accuracy | Studies that do not present any objective performance metrics |
| Study Language | English | Languages other than English |
| Data Splitting | Studies that applied data splitting into training/testing/validation or train/validation sets | Studies without a clear description of the model validation methodology |
| Gold Standard | Studies that used accepted methods for fibrosis diagnosis as a reference (e.g., liver biopsy, elastography) | Studies without a clear definition of the gold standard to validate the model’s results |
| Features | Presented the features used (e.g., clinical, laboratory, histological, radiological, or combined data) as input variables in the AI models | Studies that did not clearly describe the data used. Lack of transparency in the feature selection method, excluding studies that used the test set in the feature selection process, eliminating the risk of the curse of dimensionality with a minimum ~10:1 ratio |
| Reference | Sample Size (n) Patients | Data Splitting * | Validation | Gold Standard | Fibrosis Classification | AI ** Architecture | AI Performance (AUROC) | Traditional NIT Performance (AUROC) |
|---|---|---|---|---|---|---|---|---|
| Alkhouri N et al. (2022) [14] | 3630 | 827/1504/1299 | External | Biopsy | ≥F2 | RF, GBM, XGB | 0.717 (Ensemble); 0.683 (XGB); 0.678 (GBM); 0.688 (RF) | 0.655 (FIB-4); 0.632 (Liver Risk) |
| Dabbah S et al. (2025) [15] | 1158 | 618/540 | Internal | Elastography or Biopsy | ≥F3 or ≥9.3 kPa | XGB, LR, ANN, SVM, RF | 0.91 (XGB); 0.89 (LR); 0.89 (ANN); 0.89 (SVM); 0.90 (RF) | 0.78 (FIB-4); 0.81 (NFS) |
| Fan R et al. (2024) [16] | 828 | 703/125 | External | Biopsy | F3 or F4 | RF, LR, XGB, NB, KNN, SVM, Bagging | 0.808–0.964 (F3); 0.718–0.985 (F4) | 0.795 (FIB-4 for F3); 0.857 (FIB-4 for F4) |
| Feng G et al. (2021) [17] | 553 | 278/275 | Internal | Biopsy | ≥F2 | RF, LR | 0.893 (RF); 0.786 (LR) | 0.578 (FIB-4) |
| Ginter-Matuszewska B et al. (2025) [18] | 178 | 153/25 | Internal | Elastography | ≥F2 or >7.0 kPa | LR, DT | 0.656 (LR); 0.622 (DT) | N |
| Hassoun S et al. (2024) [19] | 6082 | 5962/120 | External | Elastography or Biopsy | ≥F3 | NAIF | 0.83 | N |
| Lu CH et al. (2024) [20] | 194 | 135/59 | Internal | Biopsy | ≥F2 | SVM, RF, KNN, XGB, LR | 0.770 (XGB); 0.768 (SVM); 0.748 (LR); 0.712 (KNN); 0.738 (RF) | 0.710 (FIB-4) |
| Mamandipoor B et al. (2023) [21] | 1151 | 808/343 | Internal | Elastography | >8 kPa | XGB, FF, LR | 0.71 (XGB); 0.70 (LR); 0.74 (FF) | 0.61 (FIB-4) |
| Okanoue T et al. (2021) [22] | 434 | 324/110 | Internal | Biopsy | ≥F1, ≥F2 or ≥F3 | NN | 0.922 (≥F1); 0.901 (≥F2); 0.911 (≥F3) | 0.766 (FIB-4 for ≥F1); 0.809 (FIB-4 for ≥F2); 0.771 (FIB-4 for ≥F3) |
| Sang C et al. (2021) [23] | 784 | 540/244 | Internal | Biopsy | ≥F3 | LR | 0.89 | 0.85 (FIB-4) |
| Suárez M et al. (2023) [24] | 215 | 150/65 | Internal | Biopsy | ≥F3 | XGB, SVM, DT, GNB, KNN | 0.95 (XGB); 0.91 (KNN); 0.84 (GNB); 0.88 (DT); 0.87 (SVM) | N |
| Suárez M et al. (2023) [25] | 211 | 148/63 | Internal | Biopsy or Elastography | ≥F2 | XGB, SVM, BLDA, LR, DT, KNN | 0.92 (XBG); 0.82 (SVM); 0.79 (BLDA); 0.75 (LR); 0.83 (DT); 0.84 (KNN) | N |
| Verma N et al. (2024) [26] | 1656 | 1153/283/220 | Internal | Biopsy | ≥F2 | RF, XGB | 0.714 (RF); 0.764 (XBG) | 0.699 (FIB-4) |
| Xiong FX et al. (2025) [27] | 746 | 522/224 | Internal | Biopsy | ≥F3 | XGB, RF, SVM, LR, NB | 0.917 (XGB); 0.840 (RF); 0.740 (SVM); 0.790 (LR); 0.503 (NB) | 0.752 (FIB-4) |
| Yamaguchi K et al. (2023) [28] | 1198 | 898/300 | Internal | Biopsy | ≥F3 | NN | 0.976 | N |
| Reference | Sample Size (n) | Data Splitting * | Validation | Gold Standard | Fibrosis Classification | AI Architecture | Image Type | AI Performance (AUROC) |
|---|---|---|---|---|---|---|---|---|
| Cunha GM et al. (2022) [29] | 2319 images | 1761/558 | External | Biopsy | ≥F1, ≥F2, ≥F3, =F4 | U-Net | Magnetic Resonance Elastography (MRE) | 0.89 (≥F1); 0.92 (≥F2); 0.92 (≥F3); 0.93 (=F4) |
| Lu X et al. (2024) [30] | 1083 images | 707/209/167 | Internal | Elastography | ≥7 kPa | DenseNet-201 | Tongue photographs | 0.893 |
| Naik SN et al. (2023) [31] | 152 images | 5-fold cross-validation (70-20-10%) | Internal | Biopsy | >F2 | ResNet-18 | Histological | 0.87 |
| Preechathammawong N et al. (2024) [32] | 176 images | (**)/146/30 | Internal | Biopsy | F0, F1, F2, F3, F4 | U-Net | Histological | *** 80.82% (F0–F1 and F2–F4); 89.73% (F0–F2 and F3–F4) |
| Yin C et al. (2024) [33] | 132 images | 3-fold-cross-validation | Internal | Biopsy | ≥F1, ≥F2, ≥F3, ≥F4 | PDGCN | Histological | 0.83 (≥F1); 0.78 (≥F2); 0.86 (≥F3); 0.88 (≥F4) |
| Zhan H et al. (2023) [34] | 1375 images and 203 patients | 143/60 | External | Biopsy | F0, F1, F2, F3–F4 | VGG16 | Histological | 0.99 (F0); 0.83 (F1); 0.80 (F2); 0.90 (F3–F4) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Souza, N.S.d.; Vitório, T.C.V.; Souza, R.A.d.; Machado, M.A.D.; Cotrim, H.P. Artificial Intelligence for Fibrosis Diagnosis in Metabolic-Dysfunction-Associated Steatotic Liver Disease: A Systematic Review. Diagnostics 2026, 16, 261. https://doi.org/10.3390/diagnostics16020261
Souza NSd, Vitório TCV, Souza RAd, Machado MAD, Cotrim HP. Artificial Intelligence for Fibrosis Diagnosis in Metabolic-Dysfunction-Associated Steatotic Liver Disease: A Systematic Review. Diagnostics. 2026; 16(2):261. https://doi.org/10.3390/diagnostics16020261
Chicago/Turabian StyleSouza, Neilson Silveira de, Théo Cordeiro Veiga Vitório, Raphael Augusto de Souza, Marcos Antônio Dórea Machado, and Helma Pinchemel Cotrim. 2026. "Artificial Intelligence for Fibrosis Diagnosis in Metabolic-Dysfunction-Associated Steatotic Liver Disease: A Systematic Review" Diagnostics 16, no. 2: 261. https://doi.org/10.3390/diagnostics16020261
APA StyleSouza, N. S. d., Vitório, T. C. V., Souza, R. A. d., Machado, M. A. D., & Cotrim, H. P. (2026). Artificial Intelligence for Fibrosis Diagnosis in Metabolic-Dysfunction-Associated Steatotic Liver Disease: A Systematic Review. Diagnostics, 16(2), 261. https://doi.org/10.3390/diagnostics16020261








