FGF2 as a Potential Tumor Suppressor in Lung Adenocarcinoma
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection from Public GEO Datasets
2.2. Identification of DEGs
2.3. GO and KEGG Pathway Enrichment of DEGs
2.4. Protein–Protein Interaction (PPI) Network Analysis and Hub Gene Selection
2.5. Prognostic Significance and Expression Validation of Hub Genes
2.6. Cell Lines and Culture
2.7. Transwell Migration and Wound Healing Assays
2.8. Statistical Analysis
3. Results
3.1. Identification of Common Differentially Expressed Genes in NSCLC
3.2. Functional Enrichment Analyses Highlight Metastasis-Associated Hallmarks and Signaling Pathways in LUAD
3.3. Identification and Expression Profiling of Hub Genes in NSCLC
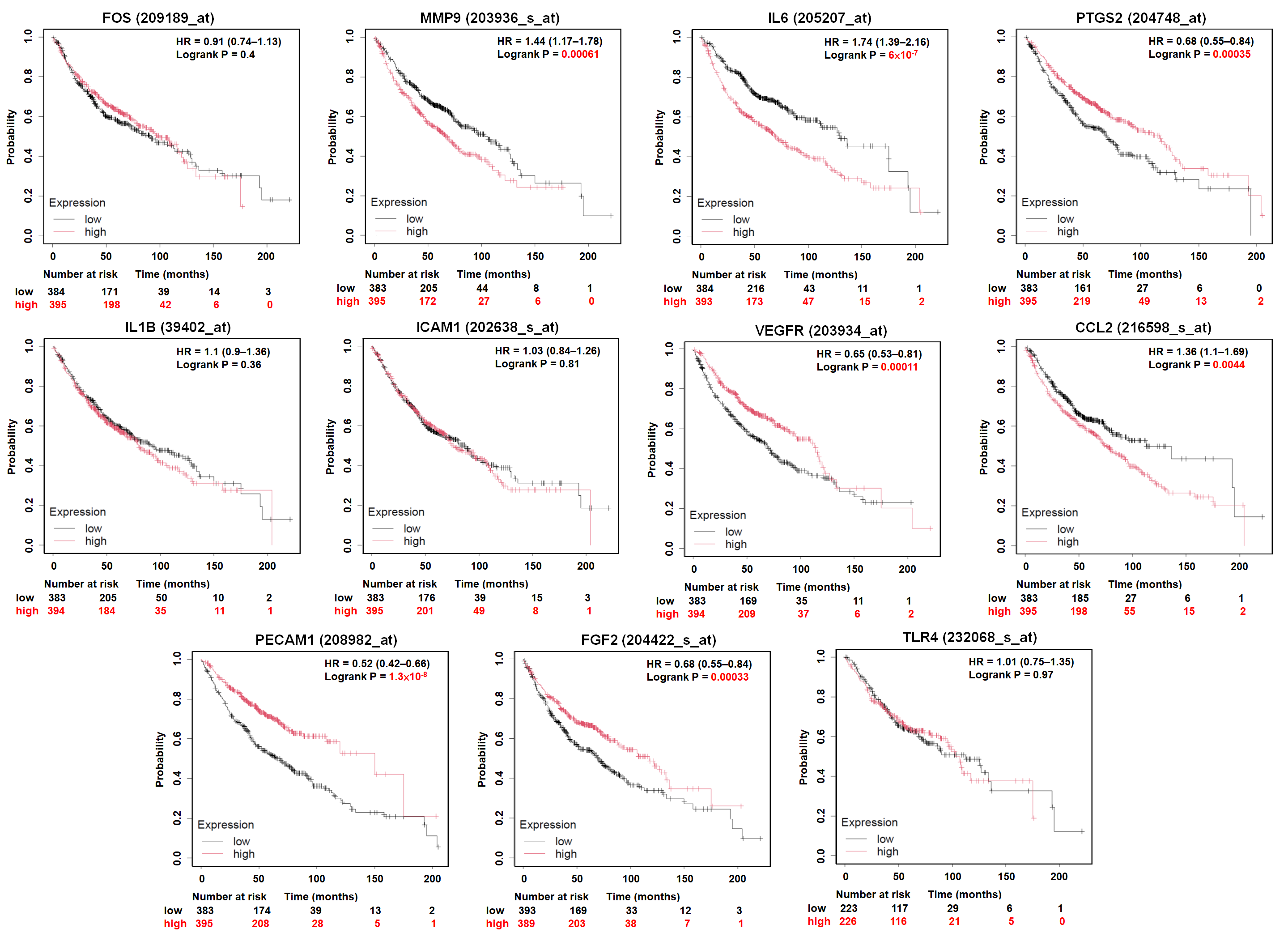
3.4. Prognostic Value of Hub Genes in LUAD Patients
3.5. FGF2 Downregulation Is Associated with Tumor Progression and Reduced Cell Motility in NSCLC
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| FGF2 | Fibroblast growth factor 2 |
| bFGF | Basic fibroblast growth factor |
| NSCLC | Non-small cell lung cancer |
| LUAD | Lung adenocarcinoma |
| LUSC | Lung squamous cell carcinoma |
| SCLC | Small cell lung cancer |
| DEGs | Differentially expressed genes |
| GEO | Gene expression omnibus |
| GO | Gene ontology |
| KEGG | Kyoto encyclopedia of genes and genomes |
| PPI | Protein–protein interaction |
| BP | Biological process |
| MF | Molecular function |
| CC | Cellular component |
| FDR | False discovery rate |
| OS | Overall survival |
| FP | First progression |
| TCGA | The cancer genome atlas |
| CPTAC | Clinical proteomic tumor analysis consortium |
| LCE | Lung cancer explorer |
| MCODE | Molecular complex detection |
| MNC | Maximum neighborhood component |
| VEGFR | Vascular endothelial growth factor receptor |
| FGFR | Fibroblast growth factor receptor |
| TNF | Tumor necrosis factor |
| IL | Interleukin |
| ICAM1 | Intercellular adhesion molecule 1 |
| KDR | Kinase insert domain receptor |
| PECAM1 | Platelet and endothelial cell adhesion molecule 1 |
| CCL2 | C-C motif chemokine ligand 2 |
| PTGS2 | Prostaglandin-endoperoxide synthase 2 |
| TLR4 | Toll-like receptor 4 |
| MAPK | Mitogen-activated protein kinase |
| PI3K/Akt | Phosphatidylinositol 3-kinase–protein kinase B pathway |
| GSEA | Gene set enrichment analysis |
| UALCAN | University of Alabama at Birmingham cancer data portal |
| TNMplot | Tumor–node–metastasis plotting platform |
References
- Abu Rous, F.; Singhi, E.K.; Sridhar, A.; Faisal, M.S.; Desai, A. Lung Cancer Treatment Advances in 2022. Cancer Investig. 2023, 41, 12–24. [Google Scholar] [CrossRef]
- Chan, A.W.; Chau, S.L.; Tong, J.H.; Chow, C.; Kwan, J.S.H.; Chung, L.Y.; Lung, R.W.; Tong, C.Y.; Tin, E.K.; Law, P.P.; et al. The Landscape of Actionable Molecular Alterations in Immunomarker-Defined Large-Cell Carcinoma of the Lung. J. Thorac. Oncol. 2019, 14, 1213–1222. [Google Scholar] [CrossRef]
- Wu, Y.L.; Dziadziuszko, R.; Ahn, J.S.; Barlesi, F.; Nishio, M.; Lee, D.H.; Lee, J.S.; Zhong, W.; Horinouchi, H.; Mao, W.; et al. Alectinib in Resected ALK-Positive Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2024, 390, 1265–1276. [Google Scholar] [CrossRef]
- Nicholson, A.G.; Tsao, M.S.; Beasley, M.B.; Borczuk, A.C.; Brambilla, E.; Cooper, W.A.; Dacic, S.; Jain, D.; Kerr, K.M.; Lantuejoul, S.; et al. The 2021 WHO Classification of Lung Tumors: Impact of Advances Since 2015. J. Thorac. Oncol. 2022, 17, 362–387. [Google Scholar] [CrossRef]
- Gale, D.; Heider, K.; Ruiz-Valdepenas, A.; Hackinger, S.; Perry, M.; Marsico, G.; Rundell, V.; Wulff, J.; Sharma, G.; Knock, H.; et al. Residual ctDNA after treatment predicts early relapse in patients with early-stage non-small cell lung cancer. Ann. Oncol. 2022, 33, 500–510. [Google Scholar] [CrossRef]
- Lee, J.H.; Machtay, M.; Kaiser, L.R.; Friedberg, J.S.; Hahn, S.M.; McKenna, M.G.; McKenna, W.G. Non-small cell lung cancer: Prognostic factors in patients treated with surgery and postoperative radiation therapy. Radiology 1999, 213, 845–852. [Google Scholar] [CrossRef]
- Jiang, C.; Zhang, N.; Hu, X.; Wang, H. Tumor-associated exosomes promote lung cancer metastasis through multiple mechanisms. Mol. Cancer 2021, 20, 117. [Google Scholar] [CrossRef]
- Yousefi, M.; Bahrami, T.; Salmaninejad, A.; Nosrati, R.; Ghaffari, P.; Ghaffari, S.H. Lung cancer-associated brain metastasis: Molecular mechanisms and therapeutic options. Cell. Oncol. 2017, 40, 419–441. [Google Scholar] [CrossRef]
- He, J.; Yang, L.; Zhou, N.; Zu, L.; Xu, S. The role and underlying mechanisms of tumour-derived exosomes in lung cancer metastasis. Curr. Opin. Oncol. 2023, 35, 46–53. [Google Scholar] [CrossRef]
- Tanabe, S. EMT Mechanism, Lung Cancer Metastasis, and microRNA. Front. Mol. Biosci. 2021, 8, 731788. [Google Scholar] [CrossRef]
- Mamon, H.J.; Yeap, B.Y.; Janne, P.A.; Reblando, J.; Shrager, S.; Jaklitsch, M.T.; Mentzer, S.; Lukanich, J.M.; Sugarbaker, D.J.; Baldini, E.H.; et al. High risk of brain metastases in surgically staged IIIA non-small-cell lung cancer patients treated with surgery, chemotherapy, and radiation. J. Clin. Oncol. 2005, 23, 1530–1537. [Google Scholar] [CrossRef]
- Khadela, A.; Postwala, H.; Rana, D.; Dave, H.; Ranch, K.; Boddu, S.H.S. A review of recent advances in the novel therapeutic targets and immunotherapy for lung cancer. Med. Oncol. 2023, 40, 152. [Google Scholar] [CrossRef] [PubMed]
- Cook, S.; Samuel, V.; Meyers, D.E.; Stukalin, I.; Litt, I.; Sangha, R.; Morris, D.G.; Heng, D.Y.C.; Pabani, A.; Dean, M.; et al. Immune-Related Adverse Events and Survival Among Patients With Metastatic NSCLC Treated With Immune Checkpoint Inhibitors. JAMA Netw. Open 2024, 7, e2352302. [Google Scholar] [CrossRef]
- Mori, S.; Hatori, N.; Kawaguchi, N.; Hamada, Y.; Shih, T.C.; Wu, C.Y.; Lam, K.S.; Matsuura, N.; Yamamoto, H.; Takada, Y.K.; et al. The integrin-binding defective FGF2 mutants potently suppress FGF2 signalling and angiogenesis. Biosci. Rep. 2017, 37, BSR20170173. [Google Scholar] [CrossRef]
- Akl, M.R.; Nagpal, P.; Ayoub, N.M.; Tai, B.; Prabhu, S.A.; Capac, C.M.; Gliksman, M.; Goy, A.; Suh, K.S. Molecular and clinical significance of fibroblast growth factor 2 (FGF2 /bFGF) in malignancies of solid and hematological cancers for personalized therapies. Oncotarget 2016, 7, 44735–44762. [Google Scholar] [CrossRef] [PubMed]
- Lund, P.J.; Lopes, M.; Sidoli, S.; Coradin, M.; Vitorino, F.N.L.; da Cunha, J.P.C.; Garcia, B.A. FGF-2 induces a failure of cell cycle progression in cells harboring amplified K-Ras, revealing new insights into oncogene-induced senescence. Mol. Omics 2021, 17, 725–739. [Google Scholar] [CrossRef]
- Chi, F.; Griffiths, J.I.; Nath, A.; Bild, A.H. Paradoxical cancer cell proliferation after FGFR inhibition through decreased p21 signaling in FGFR1-amplified breast cancer cells. Breast Cancer Res. 2024, 26, 54. [Google Scholar] [CrossRef]
- Chen, Z.F.; Wang, J.; Yu, Y.; Wei, W. MicroRNA-936 promotes proliferation and invasion of gastric cancer cells by down-regulating FGF2 expression and activating P13K/Akt signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 6707–6715. [Google Scholar] [CrossRef] [PubMed]
- Im, J.H.; Buzzelli, J.N.; Jones, K.; Franchini, F.; Gordon-Weeks, A.; Markelc, B.; Chen, J.; Kim, J.; Cao, Y.; Muschel, R.J. FGF2 alters macrophage polarization, tumour immunity and growth and can be targeted during radiotherapy. Nat. Commun. 2020, 11, 4064. [Google Scholar] [CrossRef]
- Ardizzone, A.; Bova, V.; Casili, G.; Repici, A.; Lanza, M.; Giuffrida, R.; Colarossi, C.; Mare, M.; Cuzzocrea, S.; Esposito, E.; et al. Role of Basic Fibroblast Growth Factor in Cancer: Biological Activity, Targeted Therapies, and Prognostic Value. Cells 2023, 12, 1002. [Google Scholar] [CrossRef]
- Pacini, L.; Jenks, A.D.; Lima, N.C.; Huang, P.H. Targeting the Fibroblast Growth Factor Receptor (FGFR) Family in Lung Cancer. Cells 2021, 10, 1154. [Google Scholar] [CrossRef]
- Landi, M.T.; Dracheva, T.; Rotunno, M.; Figueroa, J.D.; Liu, H.; Dasgupta, A.; Mann, F.E.; Fukuoka, J.; Hames, M.; Bergen, A.W.; et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS ONE 2008, 3, e1651. [Google Scholar] [CrossRef]
- Lu, T.P.; Hsiao, C.K.; Lai, L.C.; Tsai, M.H.; Hsu, C.P.; Lee, J.M.; Chuang, E.Y. Identification of regulatory SNPs associated with genetic modifications in lung adenocarcinoma. BMC Res. Notes 2015, 8, 92. [Google Scholar] [CrossRef]
- Sanchez-Palencia, A.; Gomez-Morales, M.; Gomez-Capilla, J.A.; Pedraza, V.; Boyero, L.; Rosell, R.; Farez-Vidal, M.E. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int. J. Cancer 2011, 129, 355–364. [Google Scholar] [CrossRef]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets--update. Nucleic Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef] [PubMed]
- Heberle, H.; Meirelles, G.V.; da Silva, F.R.; Telles, G.P.; Minghim, R. InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015, 16, 169. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Ishiguro-Watanabe, M.; Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 2021, 49, D545–D551. [Google Scholar] [CrossRef]
- Luo, W.; Brouwer, C. Pathview: An R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 2013, 29, 1830–1831. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Kuhn, M.; Simonovic, M.; Roth, A.; Minguez, P.; Doerks, T.; Stark, M.; Muller, J.; Bork, P.; et al. The STRING database in 2011: Functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 2011, 39, D561–D568. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: Protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Chin, C.H.; Chen, S.H.; Wu, H.H.; Ho, C.W.; Ko, M.T.; Lin, C.Y. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 2014, 8, S11. [Google Scholar] [CrossRef]
- Hou, G.X.; Liu, P.; Yang, J.; Wen, S. Mining expression and prognosis of topoisomerase isoforms in non-small-cell lung cancer by using Oncomine and Kaplan-Meier plotter. PLoS ONE 2017, 12, e0174515. [Google Scholar] [CrossRef]
- Gyorffy, B. Transcriptome-level discovery of survival-associated biomarkers and therapy targets in non-small-cell lung cancer. Br. J. Pharmacol. 2024, 181, 362–374. [Google Scholar] [CrossRef]
- Li, T.; Fan, J.; Wang, B.; Traugh, N.; Chen, Q.; Liu, J.S.; Li, B.; Liu, X.S. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017, 77, e108–e110. [Google Scholar] [CrossRef]
- Li, T.; Fu, J.; Zeng, Z.; Cohen, D.; Li, J.; Chen, Q.; Li, B.; Liu, X.S. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020, 48, W509–W514. [Google Scholar] [CrossRef] [PubMed]
- Uhlen, M.; Zhang, C.; Lee, S.; Sjostedt, E.; Fagerberg, L.; Bidkhori, G.; Benfeitas, R.; Arif, M.; Liu, Z.; Edfors, F.; et al. A pathology atlas of the human cancer transcriptome. Science 2017, 357, eaan2507. [Google Scholar] [CrossRef] [PubMed]
- Hung, S.Y.; Shih, Y.P.; Chen, M.; Lo, S.H. Up-regulated cten by FGF2 contributes to FGF2-mediated cell migration. Mol. Carcinog. 2014, 53, 787–792. [Google Scholar] [CrossRef][Green Version]
- Awan, B.; Turkov, D.; Schumacher, C.; Jacobo, A.; McEnerney, A.; Ramsey, A.; Xu, G.; Park, D.; Kalomoiris, S.; Yao, W.; et al. FGF2 Induces Migration of Human Bone Marrow Stromal Cells by Increasing Core Fucosylations on N-Glycans of Integrins. Stem Cell Rep. 2018, 11, 325–333. [Google Scholar] [CrossRef]
- Nowwarote, N.; Sukarawan, W.; Pavasant, P.; Foster, B.L.; Osathanon, T. Basic fibroblast growth factor regulates phosphate/pyrophosphate regulatory genes in stem cells isolated from human exfoliated deciduous teeth. Stem Cell Res. Ther. 2018, 9, 345. [Google Scholar] [CrossRef]
- Yaoita, E.; Nameta, M.; Yoshida, Y.; Fujinaka, H. Dynamic changes of podocytes caused by fibroblast growth factor 2 in culture. Cell Tissue Res. 2021, 386, 117–126. [Google Scholar] [CrossRef]
- Tsugeno, Y.; Furuhashi, M.; Sato, T.; Watanabe, M.; Umetsu, A.; Suzuki, S.; Ida, Y.; Hikage, F.; Ohguro, H. FGF-2 enhances fibrogenetic changes in TGF-beta2 treated human conjunctival fibroblasts. Sci. Rep. 2022, 12, 16006. [Google Scholar] [CrossRef] [PubMed]
- Araghi, M.; Mannani, R.; Heidarnejad Maleki, A.; Hamidi, A.; Rostami, S.; Safa, S.H.; Faramarzi, F.; Khorasani, S.; Alimohammadi, M.; Tahmasebi, S.; et al. Recent advances in non-small cell lung cancer targeted therapy; an update review. Cancer Cell Int. 2023, 23, 162. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.M.; Zheng, M.M.; Pan, Y.; Liu, S.Y.; Li, Y.; Wu, Y.L. Emerging evidence and treatment paradigm of non-small cell lung cancer. J. Hematol. Oncol. 2023, 16, 40. [Google Scholar] [CrossRef]
- Zhang, P.; Yue, L.; Leng, Q.; Chang, C.; Gan, C.; Ye, T.; Cao, D. Targeting FGFR for cancer therapy. J. Hematol. Oncol. 2024, 17, 39. [Google Scholar] [CrossRef]
- DiGiacomo, J.W.; Godet, I.; Trautmann-Rodriguez, M.; Gilkes, D.M. Extracellular Matrix-Bound FGF2 Mediates Estrogen Receptor Signaling and Therapeutic Response in Breast Cancer. Mol. Cancer Res. MCR 2021, 19, 136–149. [Google Scholar] [CrossRef]
- Ye, Y.; Yang, F.; Gu, Z.; Li, W.; Yuan, Y.; Liu, S.; Zhou, L.; Han, B.; Zheng, R.; Cao, Z. Fibroblast growth factor pathway promotes glycolysis by activating LDHA and suppressing LDHB in a STAT1-dependent manner in prostate cancer. J. Transl. Med. 2024, 22, 474. [Google Scholar] [CrossRef] [PubMed]
- Otte, J.; Dizdar, L.; Behrens, B.; Goering, W.; Knoefel, W.T.; Wruck, W.; Stoecklein, N.H.; Adjaye, J. FGF Signalling in the Self-Renewal of Colon Cancer Organoids. Sci. Rep. 2019, 9, 17365. [Google Scholar] [CrossRef]
- Carter, E.P.; Coetzee, A.S.; Tomas Bort, E.; Wang, Q.; Kocher, H.M.; Grose, R.P. Dissecting FGF Signalling to Target Cellular Crosstalk in Pancreatic Cancer. Cells 2021, 10, 847. [Google Scholar] [CrossRef]
- de Luna Vitorino, F.N.; Levy, M.J.; Mansano Wailemann, R.A.; Lopes, M.; Silva, M.L.; Sardiu, M.E.; Garcia, B.A.; Machado Motta, M.C.; Oliveira, C.C.; Armelin, H.A.; et al. The antiproliferative effect of FGF2 in K-Ras-driven tumor cells involves modulation of rRNA and the nucleolus. J. Cell Sci. 2023, 136, jcs260989. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, X.; Peng, S.; Hu, H.; Wang, Y.; Shao, M.; Feng, G.; Liu, Y.; Bai, Y. Single-Cell Analysis Reveals Spatial Heterogeneity of Immune Cells in Lung Adenocarcinoma. Front. Cell Dev. Biol. 2021, 9, 638374. [Google Scholar] [CrossRef] [PubMed]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Lin, S.-S.; Lu, H.-Y.; Chang, T.-M.; Sun, Y.-S.; Liu, J.-F. FGF2 as a Potential Tumor Suppressor in Lung Adenocarcinoma. Diagnostics 2026, 16, 250. https://doi.org/10.3390/diagnostics16020250
Lin S-S, Lu H-Y, Chang T-M, Sun Y-S, Liu J-F. FGF2 as a Potential Tumor Suppressor in Lung Adenocarcinoma. Diagnostics. 2026; 16(2):250. https://doi.org/10.3390/diagnostics16020250
Chicago/Turabian StyleLin, Shih-Sen, Hsin-Ying Lu, Tsung-Ming Chang, Ying-Sui Sun, and Ju-Fang Liu. 2026. "FGF2 as a Potential Tumor Suppressor in Lung Adenocarcinoma" Diagnostics 16, no. 2: 250. https://doi.org/10.3390/diagnostics16020250
APA StyleLin, S.-S., Lu, H.-Y., Chang, T.-M., Sun, Y.-S., & Liu, J.-F. (2026). FGF2 as a Potential Tumor Suppressor in Lung Adenocarcinoma. Diagnostics, 16(2), 250. https://doi.org/10.3390/diagnostics16020250





