Overexpression of CDC20 Confer a Poorer Prognosis in Bladder Cancer Identified by Gene Co-Expression Network Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Acquisition of Raw Data
2.2. Identifying a Co-Expression Module by WGCNA
2.3. Differential Expression Analysis and Interaction with Modules of Interest
2.4. Function Enrichment Analysis
2.5. PPI Network and Hub Gene Identification
2.6. Verification of the Prognostic Value of Hub Genes
2.7. Verification of the Expressions of Hub Genes with Prognostic Value and Gene Set Enrichment Analysis
2.8. Tumor-Infiltrating Immune Cells
2.9. Cell Culture
2.10. siRNA Transfection
2.11. RNA Extraction and Real-Time Quantitative Polymerase Chain Reaction
2.12. Cell Counting Kit-8 Assay
2.13. Transwell Assays
2.14. Statistical Analysis
3. Results
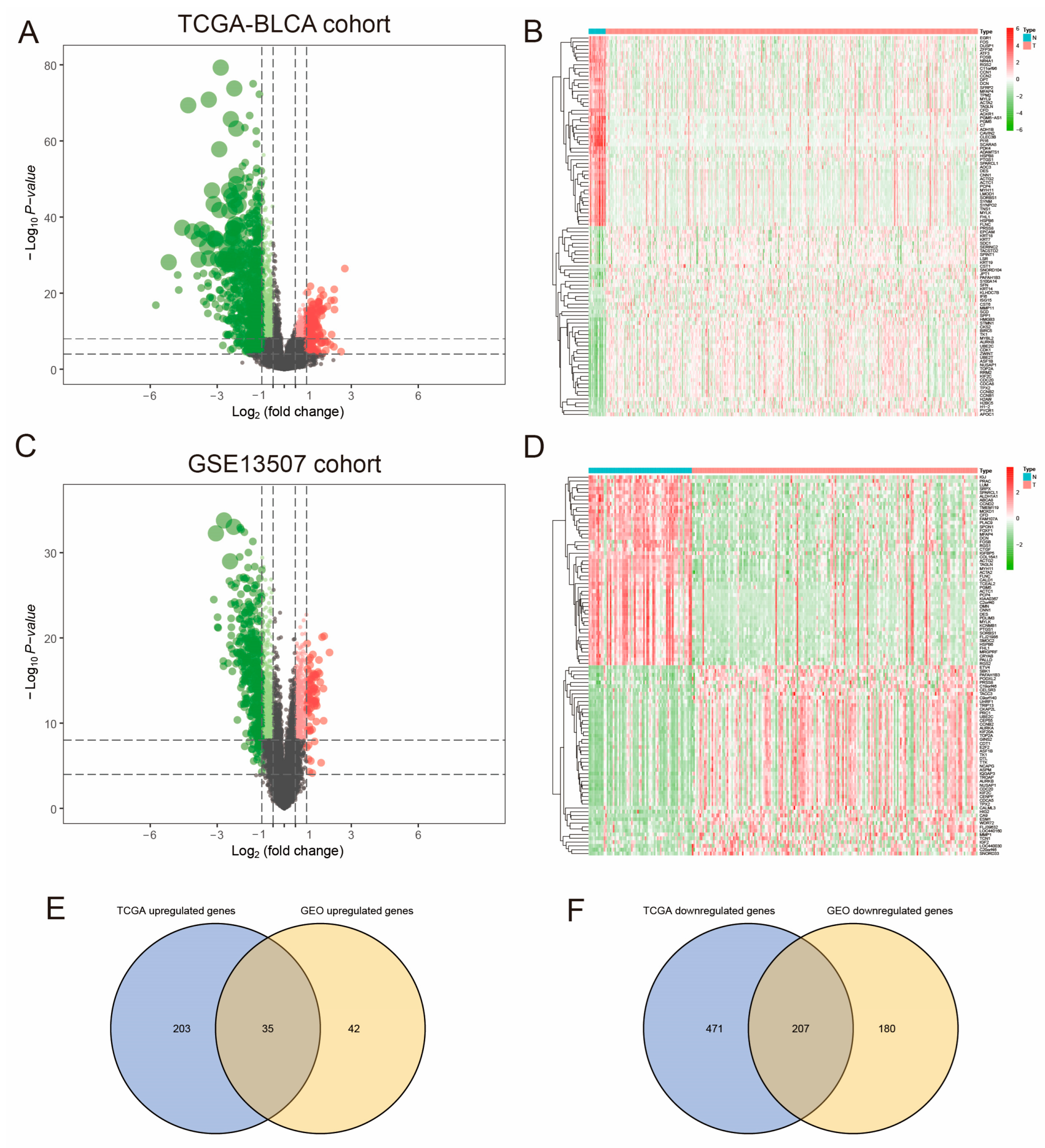
3.1. Gene Intersection Between Differentially Expressed Genes and Co-Expression Modules
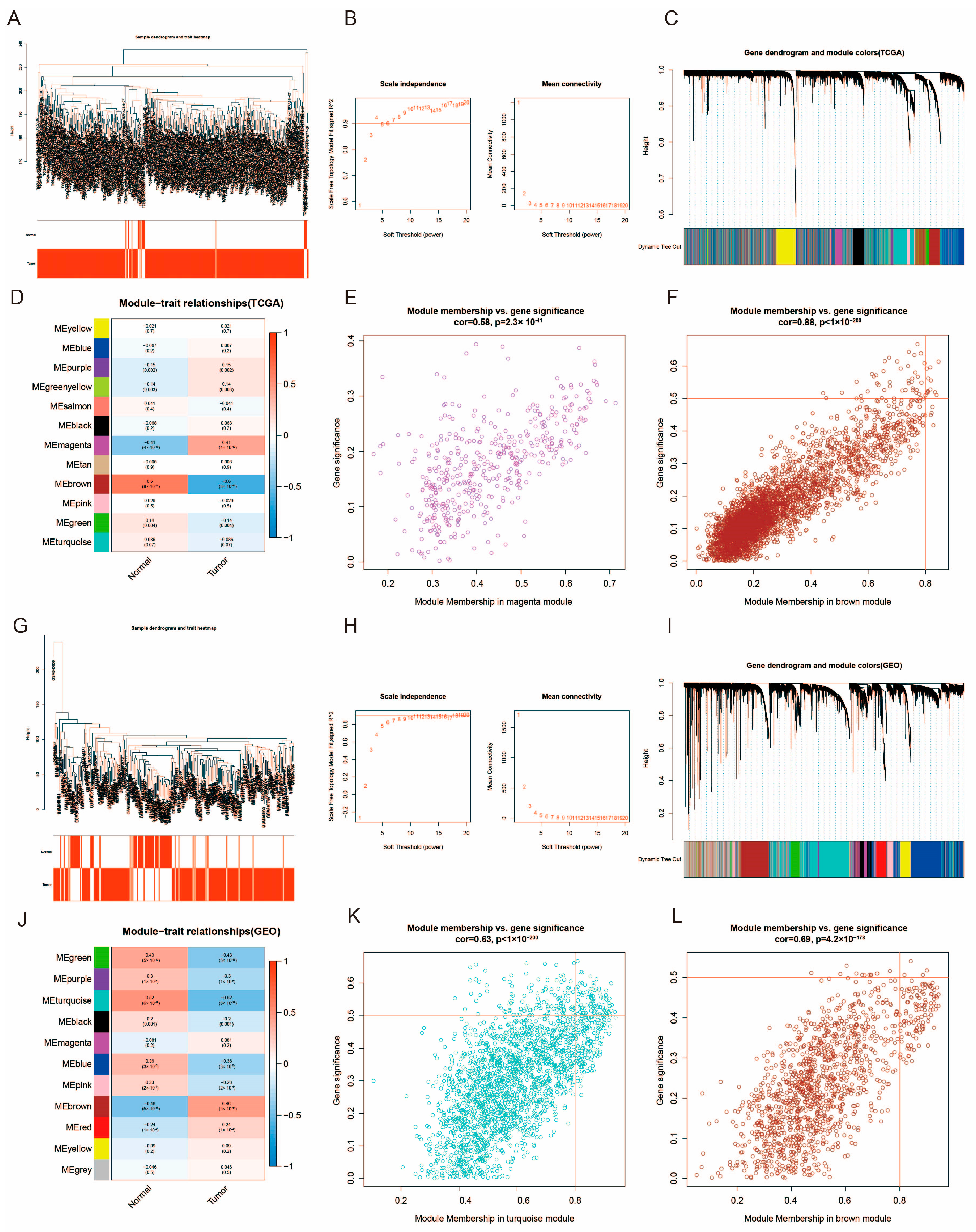
3.2. Construction of Weighted Co-Expression Network and Identification of Key Modules
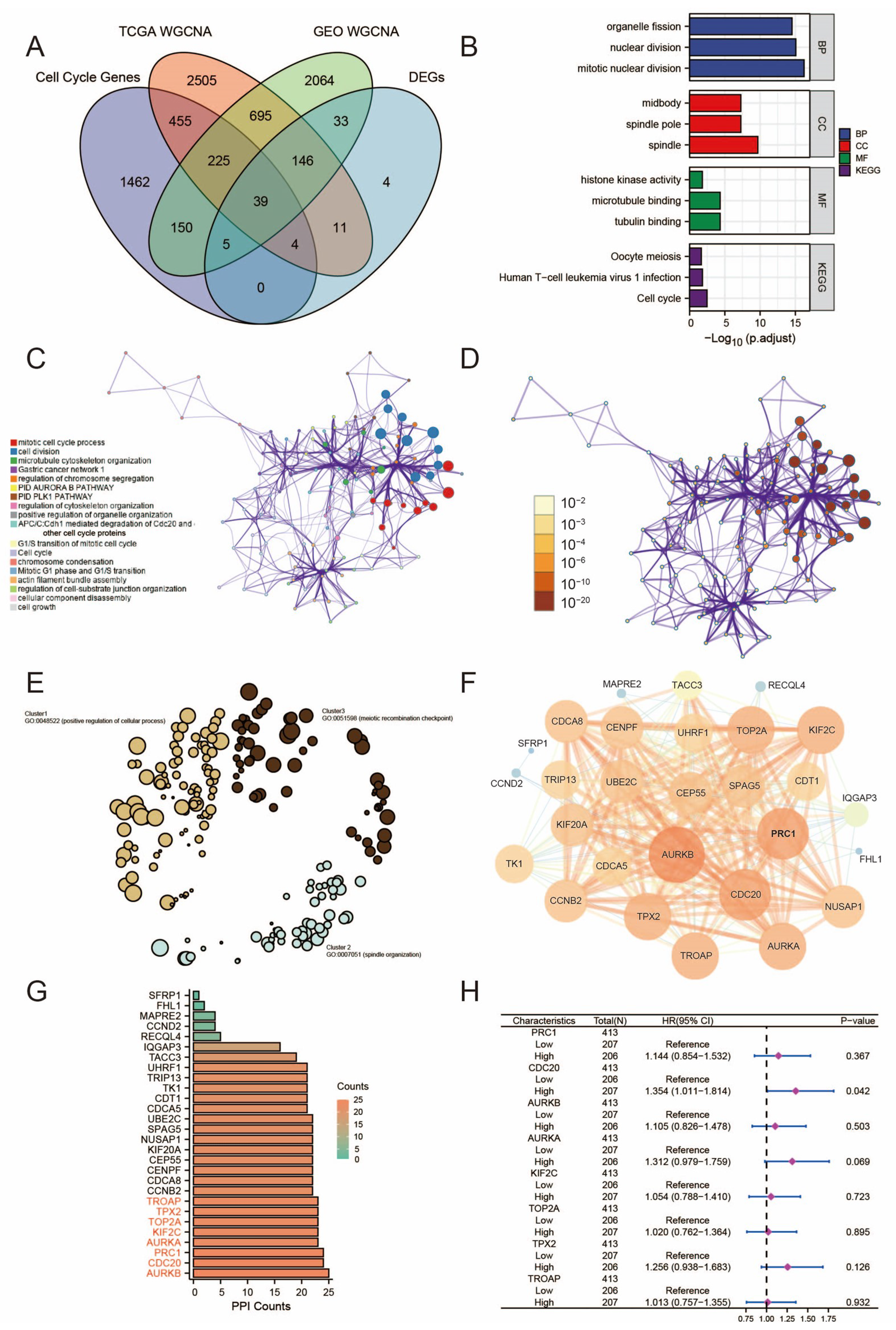
3.3. Gene Set Enrichment Analysis
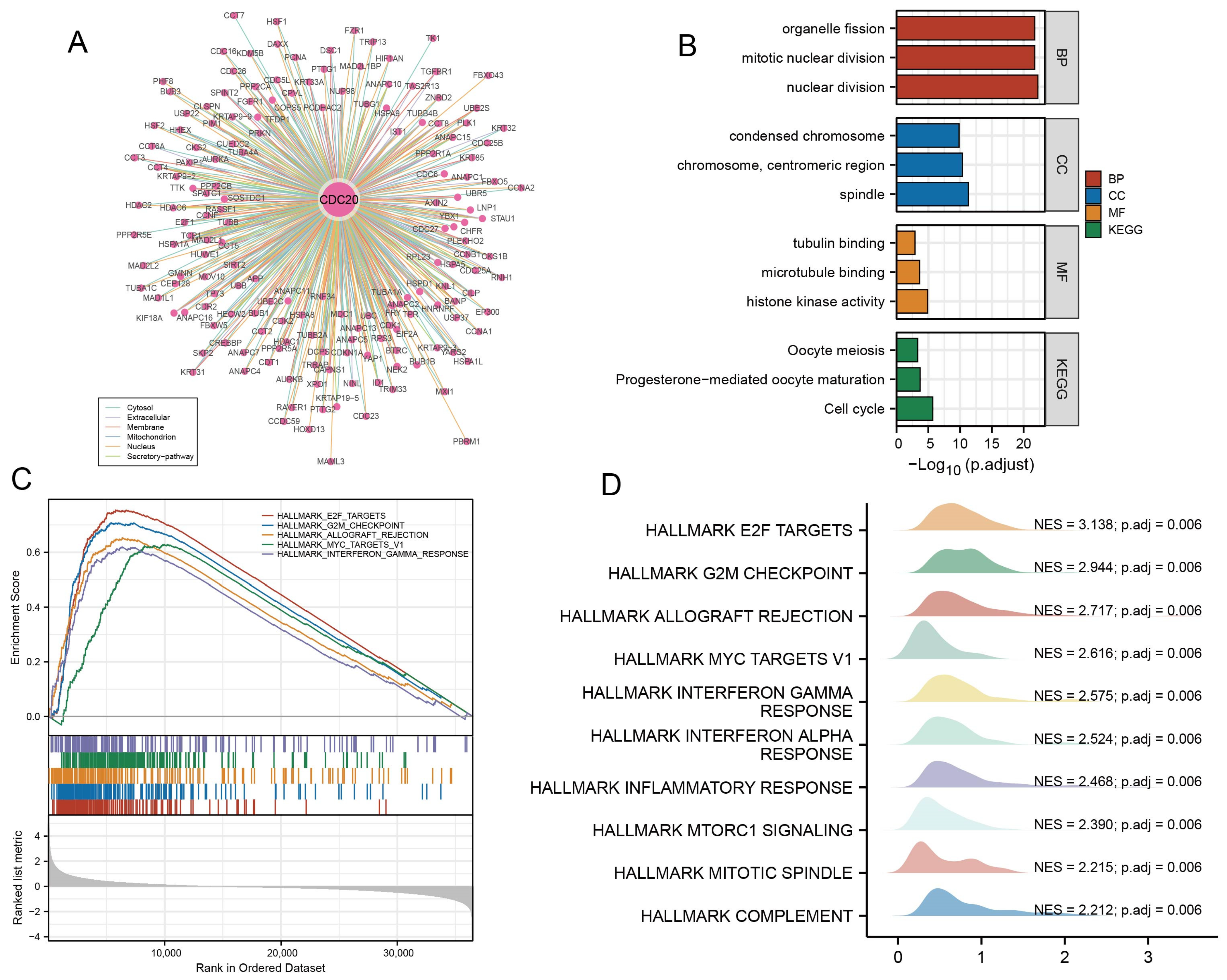
3.4. Construction of PPI Network and Validation of CDC20 Gene Identification
3.5. Functional Enrichment of CDC20 and Validation of GSEA
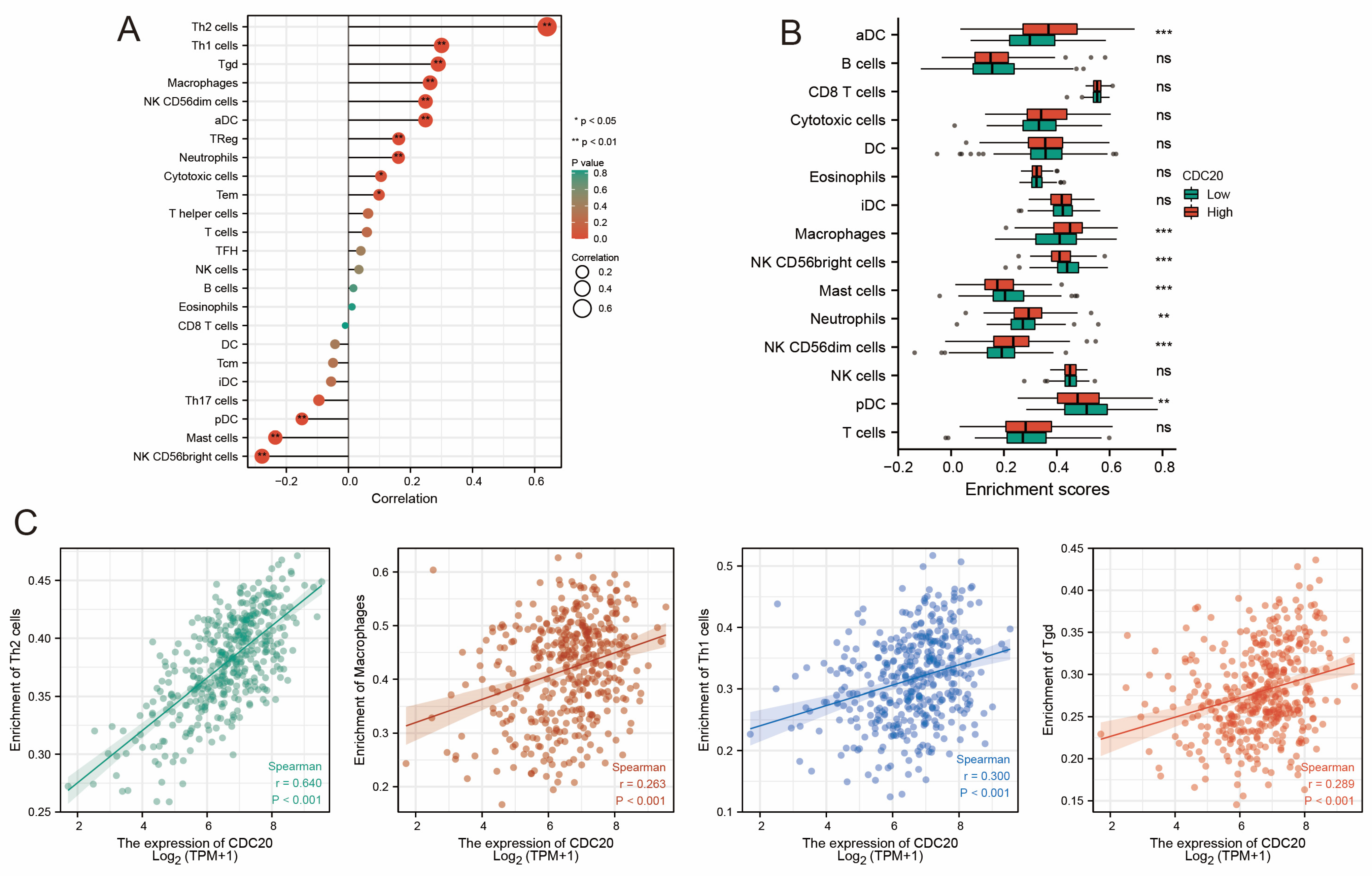
3.6. Correlation Between CDC20 and Tumor Immune Microenvironment
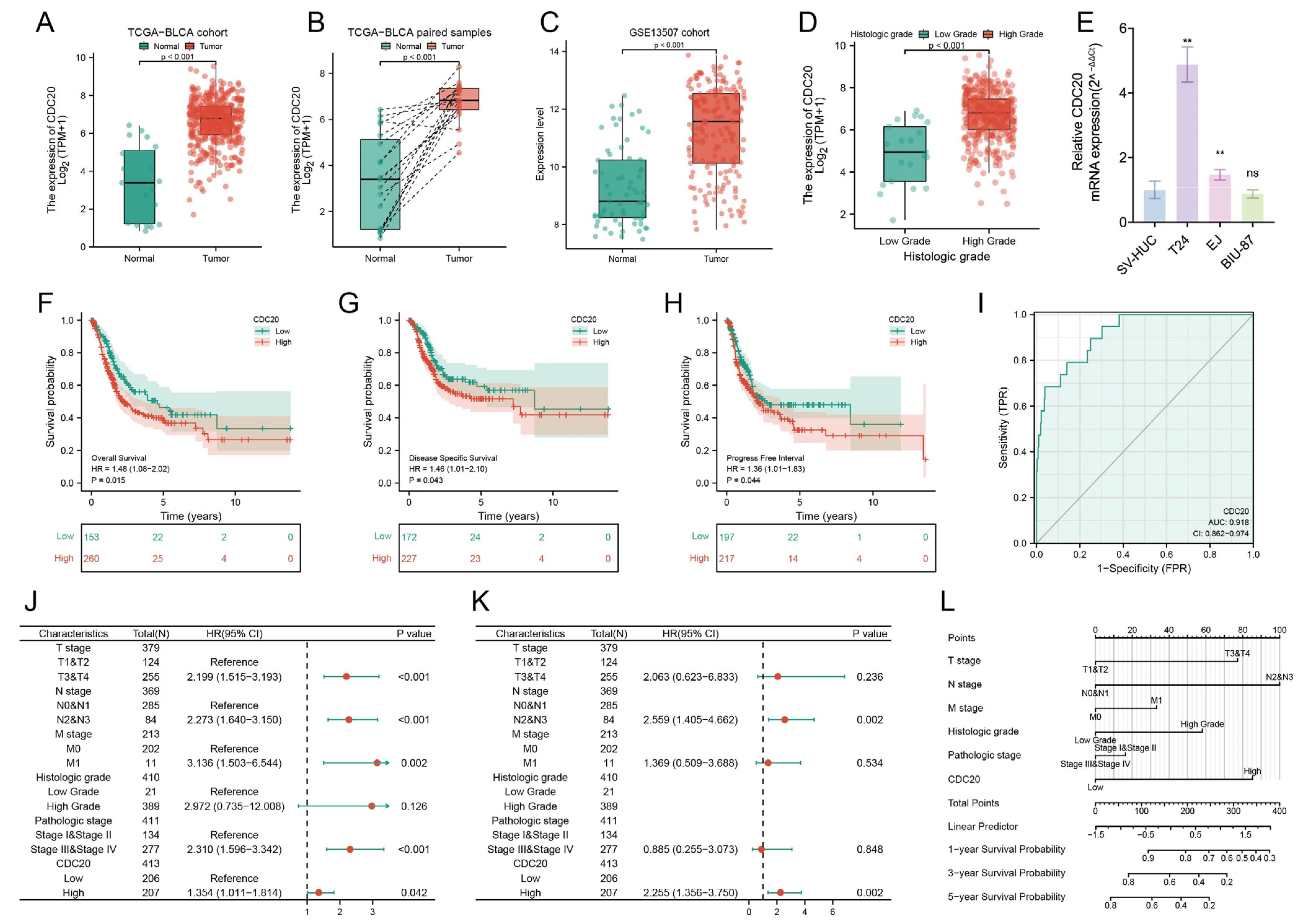
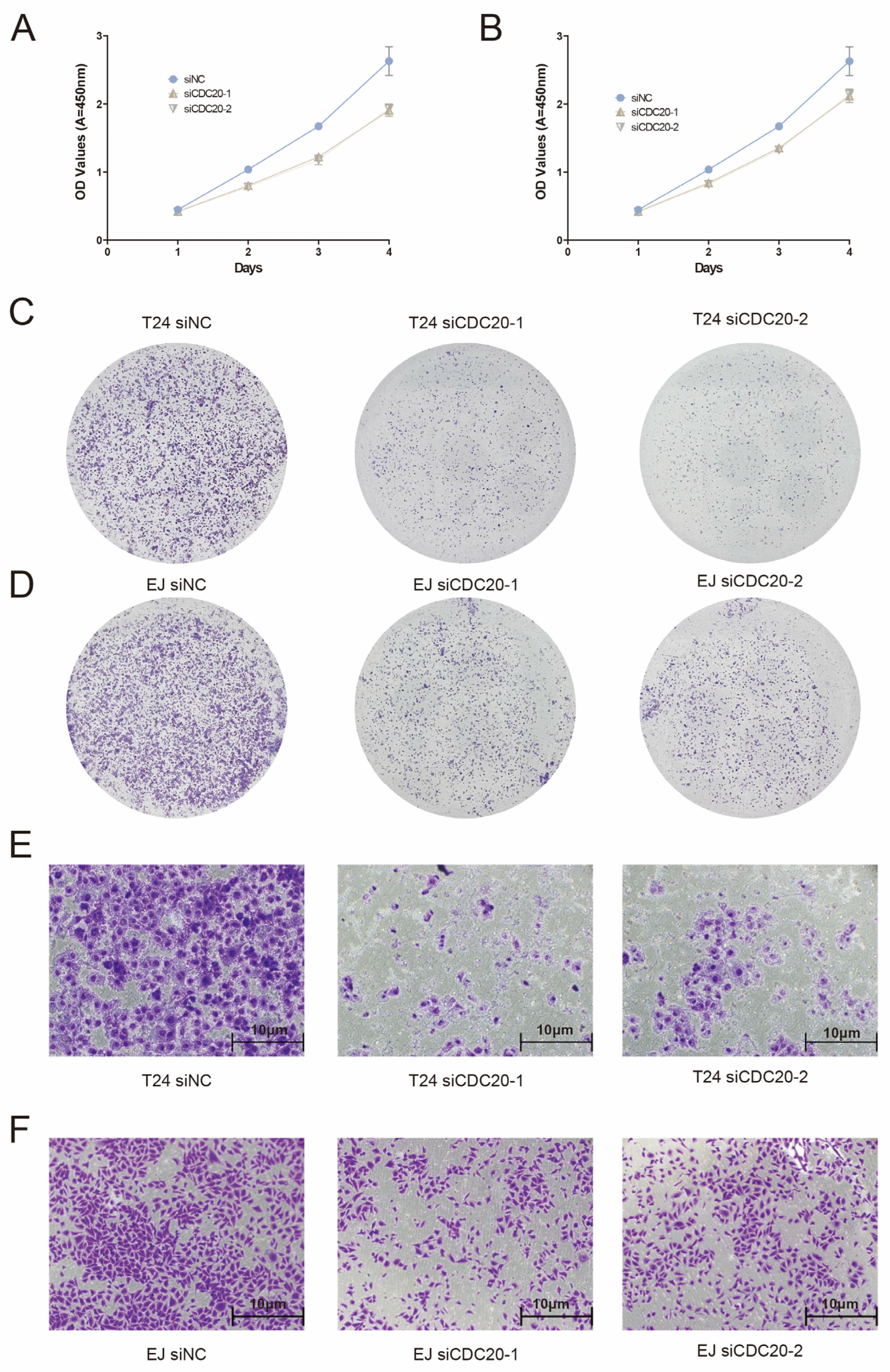
3.7. Expression and Functional Characterization of CDC20
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lobo, N.; Afferi, L.; Moschini, M.; Mostafid, H.; Porten, S.; Psutka, S.P.; Gupta, S.; Smith, A.B.; Williams, S.B.; Lotan, Y. Epidemiology, Screening, and Prevention of Bladder Cancer. Eur. Urol. Oncol. 2022, 5, 628–639. [Google Scholar] [CrossRef]
- Kayama, E.; Uemura, M.; Onagi, A.; Meguro, S.; Ogawa, S.; Yaginuma, K.; Matsuoka, K.; Hoshi, S.; Koguchi, T.; Hata, J.; et al. A Novel Gene Expression Scoring System Predicts Recurrence in Non-Muscle-Invasive Bladder Cancer Patients. Cancer Med. 2024, 13, e70349. [Google Scholar] [CrossRef]
- Li, H.; Wang, L.; Li, H.; Zhang, P.; Li, Z.; Xue, L.; Wang, Z.; Fu, D.; Chen, Q.; Luo, Q.; et al. Analysis of Risk Factors for Recurrence after Transurethral Resection of Bladder Tumor in Patients with Non-Muscle Invasive Bladder Cancer: 2-Year Follow-Up Outcomes. Oncology 2024, 102, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.P.; Cai, J.; Miranda, G.; Bhanvadia, S.; Quinn, D.I.; Schuckman, A.K.; Djaladat, H.; Daneshmand, S. Management Trends and Outcomes of Patients Undergoing Radical Cystectomy for Urothelial Carcinoma of the Bladder: Evolution of the University of Southern California Experience over 3,347 Cases. J. Urol. 2022, 207, 302–313. [Google Scholar] [CrossRef]
- Dobruch, J.; Oszczudłowski, M. Bladder Cancer: Current Challenges and Future Directions. Medicina 2021, 57, 749. [Google Scholar] [CrossRef] [PubMed]
- Englinger, B. Molecular research in bladder cancer. Curr. Opin. Urol. 2024, 34, 225–226. [Google Scholar] [CrossRef] [PubMed]
- Varchulová Nováková, Z.; Kuniaková, M.; Žiaran, S.; Harsányi, Š. Molecular Biomarkers of Bladder Cancer: A Mini-Review. Physiol. Res. 2023, 72, S247–s256. [Google Scholar] [CrossRef]
- Wondergem, N.E.; Nauta, I.H.; Muijlwijk, T.; Leemans, C.R.; van de Ven, R. The Immune Microenvironment in Head and Neck Squamous Cell Carcinoma: On Subsets and Subsites. Curr. Oncol. Rep. 2020, 22, 81. [Google Scholar] [CrossRef]
- Elia, I.; Haigis, M.C. Metabolites and the tumour microenvironment: From cellular mechanisms to systemic metabolism. Nat. Metab. 2021, 3, 21–32. [Google Scholar] [CrossRef]
- Zeng, X.; Lu, Z.; Dai, C.; Su, H.; Liu, Z.; Cheng, S. Establish TIIC signature score based the machine learning fusion in bladder cancer. Discov. Oncol. 2024, 15, 368. [Google Scholar] [CrossRef]
- Costa, A.C.; Santos, J.M.O.; da Costa, R.M.G.; Medeiros, R. Impact of immune cells on the hallmarks of cancer: A literature review. Crit. Rev. Oncol. Hematol. 2021, 168, 103541. [Google Scholar] [CrossRef]
- Hao, Y.; Li, D.; Xu, Y.; Ouyang, J.; Wang, Y.; Zhang, Y.; Li, B.; Xie, L.; Qin, G. Investigation of lipid metabolism dysregulation and the effects on immune microenvironments in pan-cancer using multiple omics data. BMC Bioinform. 2019, 20 (Suppl. 7), 195. [Google Scholar] [CrossRef] [PubMed]
- Siracusano, S.; Rizzetto, R.; Porcaro, A.B. Bladder cancer genomics. Urologia 2020, 87, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Danishuddin; Haque, M.A.; Khan, S.; Kim, J.J.; Ahmad, K. Molecular Landscape of Bladder Cancer: Key Genes, Transcription Factors, and Drug Interactions. Int. J. Mol. Sci. 2024, 25, 10997. [Google Scholar] [CrossRef]
- Lai, G.; Zhong, X.; Liu, H.; Deng, J.; Li, K.; Xie, B. A Novel m7G-Related Genes-Based Signature with Prognostic Value and Predictive Ability to Select Patients Responsive to Personalized Treatment Strategies in Bladder Cancer. Cancers 2022, 14, 5346. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Beltran, A.; Cookson, M.S.; Guercio, B.J.; Cheng, L. Advances in diagnosis and treatment of bladder cancer. BMJ 2024, 384, e076743. [Google Scholar] [CrossRef]
- Heard, J.R.; Ahdoot, M.; Theodorescu, D.; Mitra, A.P. Biomarkers of treatment response in bladder cancer. Expert Rev. Mol. Diagn. 2024, 24, 957–969. [Google Scholar] [CrossRef]
- He, W.; Meng, J. CDC20: A novel therapeutic target in cancer. Am. J. Transl. Res. 2023, 15, 678–693. [Google Scholar]
- Bruno, S.; Ghelli Luserna di Rorà, A.; Napolitano, R.; Soverini, S.; Martinelli, G.; Simonetti, G. CDC20 in and out of mitosis: A prognostic factor and therapeutic target in hematological malignancies. J. Exp. Clin. Cancer Res. 2022, 41, 159. [Google Scholar] [CrossRef]
- Greil, C.; Engelhardt, M.; Wäsch, R. The Role of the APC/C and Its Coactivators Cdh1 and Cdc20 in Cancer Development and Therapy. Front. Genet. 2022, 13, 941565. [Google Scholar] [CrossRef]
- Schrock, M.S.; Stromberg, B.R.; Scarberry, L.; Summers, M.K. APC/C ubiquitin ligase: Functions and mechanisms in tumorigenesis. Semin. Cancer Biol. 2020, 67, 80–91. [Google Scholar] [CrossRef]
- Ryan, C.E.; Fasaye, G.A.; Gallanis, A.F.; Gamble, L.A.; McClelland, P.H.; Duemler, A.; Samaranayake, S.G.; Blakely, A.M.; Drogan, C.M.; Kingham, K.; et al. Germline CDH1 Variants and Lifetime Cancer Risk. JAMA 2024, 332, 722–729. [Google Scholar] [CrossRef]
- Nobre, A.R.; Dalla, E.; Yang, J.; Huang, X.; Wullkopf, L.; Risson, E.; Razghandi, P.; Anton, M.L.; Zheng, W.; Seoane, J.A.; et al. ZFP281 drives a mesenchymal-like dormancy program in early disseminated breast cancer cells that prevents metastatic outgrowth in the lung. Nat. Cancer 2022, 3, 1165–1180. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Yang, Y.; Cao, Y.; Meng, M.; Wang, X.; Zhang, Z.; Fu, W.; Duan, S.; Tang, L. Histone deacetylase inhibitors inhibit lung adenocarcinoma metastasis via HDAC2/YY1 mediated downregulation of Cdh1. Sci. Rep. 2023, 13, 12069. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Ge, Y.; Zang, Y.; Xu, M.; Jin, L.; Wang, Y.; Xu, X.; Xue, B.; Wang, Z.; Wang, L. CDC20 promotes radioresistance of prostate cancer by activating Twist1 expression. Apoptosis 2023, 28, 1584–1595. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Kong, L.; Zhao, Y.; Zhai, F.; Zhan, Z.; Li, Y.; Jingfei, Z.; Chunhong, Y.; Jin, X. The oncogenic role of EIF4A3/CDC20 axis in the endometrial cancer. J. Mol. Med. 2024, 102, 1395–1410. [Google Scholar] [CrossRef]
- Pan, J.; Chen, R.H. Spindle checkpoint regulates Cdc20p stability in Saccharomyces cerevisiae. Genes Dev. 2004, 18, 1439–1451. [Google Scholar] [CrossRef] [PubMed]
- Manchado, E.; Guillamot, M.; de Carcer, G.; Eguren, M.; Trickey, M.; Garcia-Higuera, I.; Moreno, S.; Yamano, H.; Canamero, M.; Malumbres, M. Targeting mitotic exit leads to tumor regression in vivo: Modulation by Cdk1, Mastl, and the PP2A/B55alpha, delta phosphatase. Cancer Cell 2010, 18, 641–654. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, J.; Wan, L.; Zhou, X.; Wang, Z.; Wei, W. Targeting Cdc20 as a novel cancer therapeutic strategy. Pharmacol. Ther. 2015, 151, 141–151. [Google Scholar] [CrossRef]
- Li, M.; Fang, X.; Wei, Z.; York, J.P.; Zhang, P. Loss of spindle assembly checkpoint-mediated inhibition of Cdc20 promotes tumorigenesis in mice. J. Cell Biol. 2009, 185, 983–994. [Google Scholar] [CrossRef]
- Fu, C.; Jiang, L.; Hao, S.; Liu, Z.; Ding, S.; Zhang, W.; Yang, X.; Li, S. Activation of the IL-4/STAT6 Signaling Pathway Promotes Lung Cancer Progression by Increasing M2 Myeloid Cells. Front. Immunol. 2019, 10, 2638. [Google Scholar] [CrossRef]
- Zhang, Q.; Qin, J.; Zhong, L.; Gong, L.; Zhang, B.; Zhang, Y.; Gao, W.Q. CCL5-Mediated Th2 Immune Polarization Promotes Metastasis in Luminal Breast Cancer. Cancer Res. 2015, 75, 4312–4321. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, B.; Wang, Y.; Shang, G. Cdc20 inhibitor apcin inhibits the growth and invasion of osteosarcoma cells. Oncol. Rep. 2018, 40, 841–848. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wan, L.; Zhong, J.; Inuzuka, H.; Liu, P.; Sarkar, F.H.; Wei, W. Cdc20: A potential novel therapeutic target for cancer treatment. Curr. Pharm. Des. 2013, 19, 3210–3214. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Low Expression of CDC20 | High Expression of CDC20 | p | |
|---|---|---|---|---|
| Total, n | 207 | 207 | ||
| Age, n (%) | ≤70 | 124 (30%) | 110 (26.6%) | 0.197 |
| >70 | 83 (20%) | 97 (23.4%) | ||
| Gender, n (%) | Female | 52 (12.6%) | 57 (13.8%) | 0.655 |
| Male | 155 (37.4%) | 150 (36.2%) | ||
| Histologic grade, n (%) | High Grade | 186 (45.3%) | 204 (49.6%) | <0.001 |
| Low Grade | 20 (4.9%) | 1 (0.2%) | ||
| Pathologic stage, n (%) | Stage I | 3 (0.7%) | 1 (0.2%) | 0.570 |
| Stage II | 68 (16.5%) | 62 (15%) | ||
| Stage III | 66 (16%) | 76 (18.4%) | ||
| Stage IV | 69 (16.7%) | 67 (16.3%) | ||
| T stage, n (%) | T1 | 3 (0.8%) | 2 (0.5%) | 0.117 |
| T2 | 59 (15.5%) | 60 (15.8%) | ||
| T3 | 90 (23.7%) | 106 (27.9%) | ||
| T4 | 38 (10%) | 22 (5.8%) | ||
| N stage, n (%) | N0 | 117 (31.6%) | 122 (33%) | 0.153 |
| N1 | 18 (4.9%) | 28 (7.6%) | ||
| N2 | 46 (12.4%) | 31 (8.4%) | ||
| N3 | 4 (1.1%) | 4 (1.1%) | ||
| M stage, n (%) | M0 | 117 (54.9%) | 85 (39.9%) | 1.000 |
| M1 | 6 (2.8%) | 5 (2.3%) | ||
| Lymph vascular invasion, n (%) | No | 56 (19.8%) | 74 (26.1%) | 0.008 |
| Yes | 91 (32.2%) | 62 (21.9%) | ||
| Primary therapy outcome, n (%) | PD | 29 (8.1%) | 41 (11.5%) | 0.289 |
| SD | 15 (4.2%) | 16 (4.5%) | ||
| PR | 12 (3.4%) | 10 (2.8%) | ||
| CR | 127 (35.6%) | 107 (30%) | ||
| Characteristics | Total (N) | Odds Ratio (OR) | p Value |
|---|---|---|---|
| T stage (T3&T4 vs. T1&T2) | 380 | 1.000 (0.651–1.536) | 1.000 |
| N stage (N1&N2&N3 vs. N0) | 370 | 0.889 (0.579–1.361) | 0.587 |
| M stage (M1 vs. M0) | 213 | 1.147 (0.321–3.929) | 0.825 |
| Histologic grade (High Grade vs. Low Grade) | 411 | 21.935 (4.505–395.580) | 0.003 |
| Pathologic stage (Stage III&Stage IV vs. Stage I&Stage II) | 412 | 1.194 (0.790–1.806) | 0.400 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, X.; Guo, Y.; Wang, L.; Miao, Z.; Lu, X.; Ruan, J.; Tian, W. Overexpression of CDC20 Confer a Poorer Prognosis in Bladder Cancer Identified by Gene Co-Expression Network Analysis. Diagnostics 2025, 15, 3016. https://doi.org/10.3390/diagnostics15233016
Yang X, Guo Y, Wang L, Miao Z, Lu X, Ruan J, Tian W. Overexpression of CDC20 Confer a Poorer Prognosis in Bladder Cancer Identified by Gene Co-Expression Network Analysis. Diagnostics. 2025; 15(23):3016. https://doi.org/10.3390/diagnostics15233016
Chicago/Turabian StyleYang, Xuejian, Yunjie Guo, Lanyu Wang, Zengli Miao, Xiaojie Lu, Jun Ruan, and Wei Tian. 2025. "Overexpression of CDC20 Confer a Poorer Prognosis in Bladder Cancer Identified by Gene Co-Expression Network Analysis" Diagnostics 15, no. 23: 3016. https://doi.org/10.3390/diagnostics15233016
APA StyleYang, X., Guo, Y., Wang, L., Miao, Z., Lu, X., Ruan, J., & Tian, W. (2025). Overexpression of CDC20 Confer a Poorer Prognosis in Bladder Cancer Identified by Gene Co-Expression Network Analysis. Diagnostics, 15(23), 3016. https://doi.org/10.3390/diagnostics15233016






