MSLF-Net: A Multi-Scale and Multi-Level Feature Fusion Net for Diabetic Retinopathy Segmentation
Abstract
1. Introduction
- 1.
- This paper established effective pyramid feature extraction blocks, which can generate multi-scale features efficiently and enhance the segmentation of small lesions significantly.
- 2.
- This paper proposed the cross-layer and multi-scale class response feature fusion method, which makes the multi-scale feature fusion more effective, can adequately meet the challenges of small or complex features, avoids the complicated preprocessing work, achieves a high-performance end-to-end segmentation network, and reaches an advanced level in DR lesion segmentation.
- 3.
- This paper improved the image quality by preprocessing the field of view (FOV) mask images for the IDRID and e_ophtha dataset, so that the model can be trained and worked on small datasets, and to facilitate the future work of DR lesion segmentation.
2. Related Work
2.1. Convolutional Neural Networks in DR Lesion Segmentations
2.2. Feature Fusion in Medical Image
3. Materials and Methods
3.1. Datasets
3.2. Image Preprocessing and Augmentation
3.3. Model Architecture
3.3.1. Model Overall
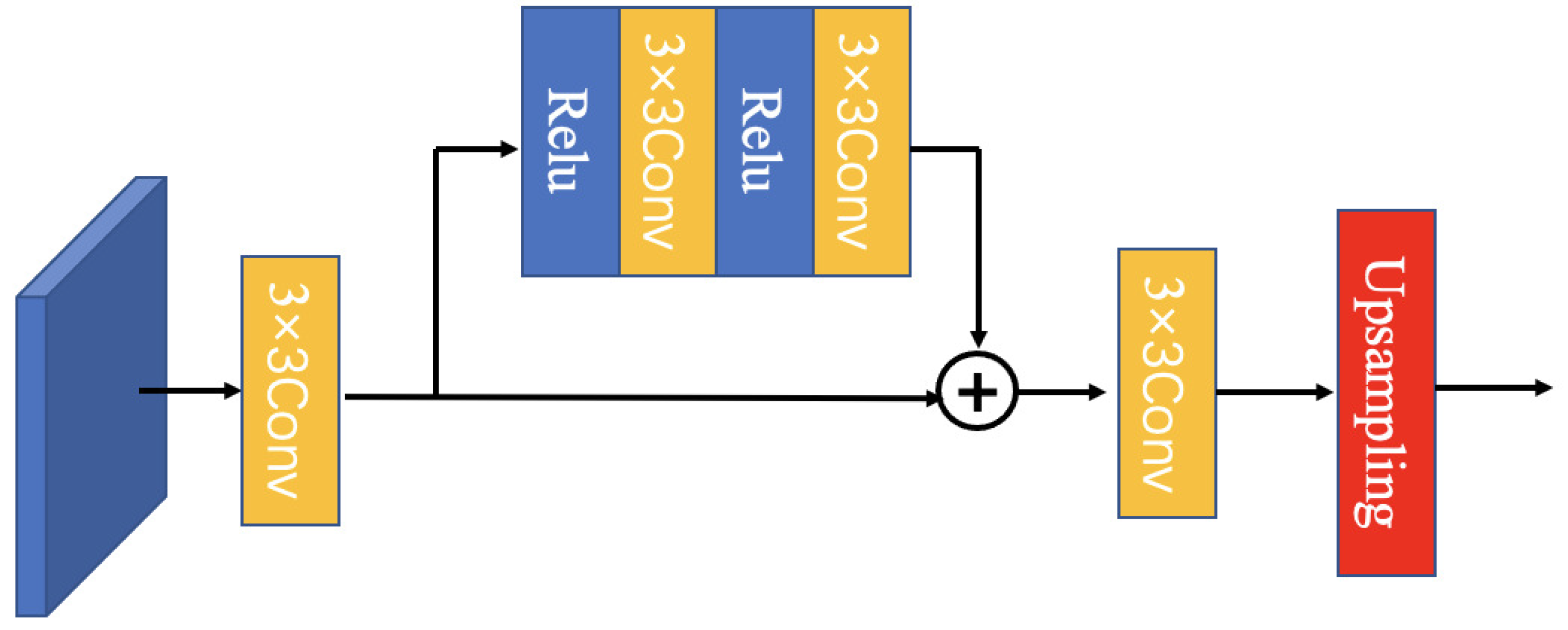
3.3.2. Multiscale Feature Module (MFEF)
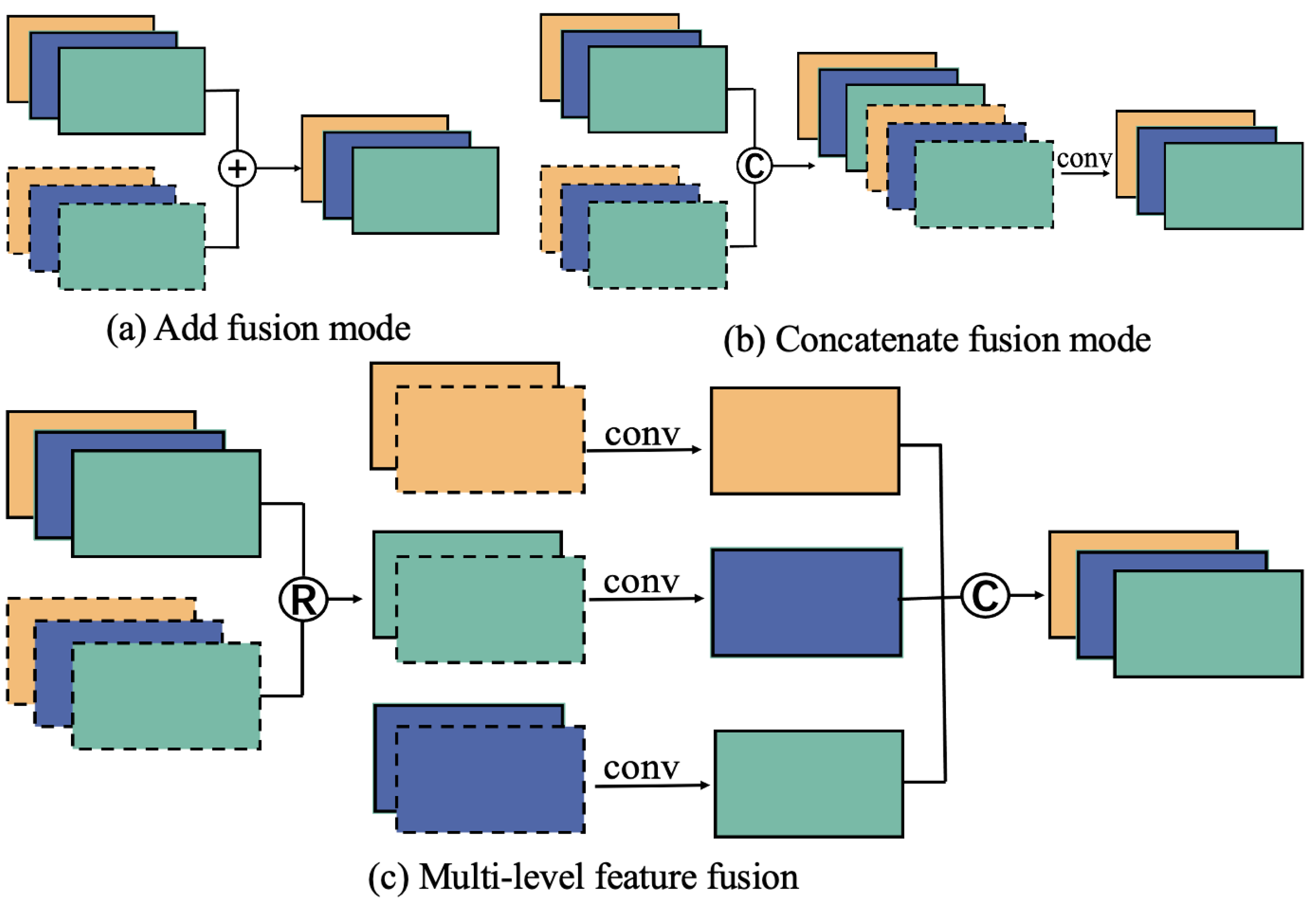
3.3.3. Feature Fusion Module (MLFF)
3.4. Implementation Details and Experiment Settings
3.5. Model Performance Evaluation
4. Results and Discussion
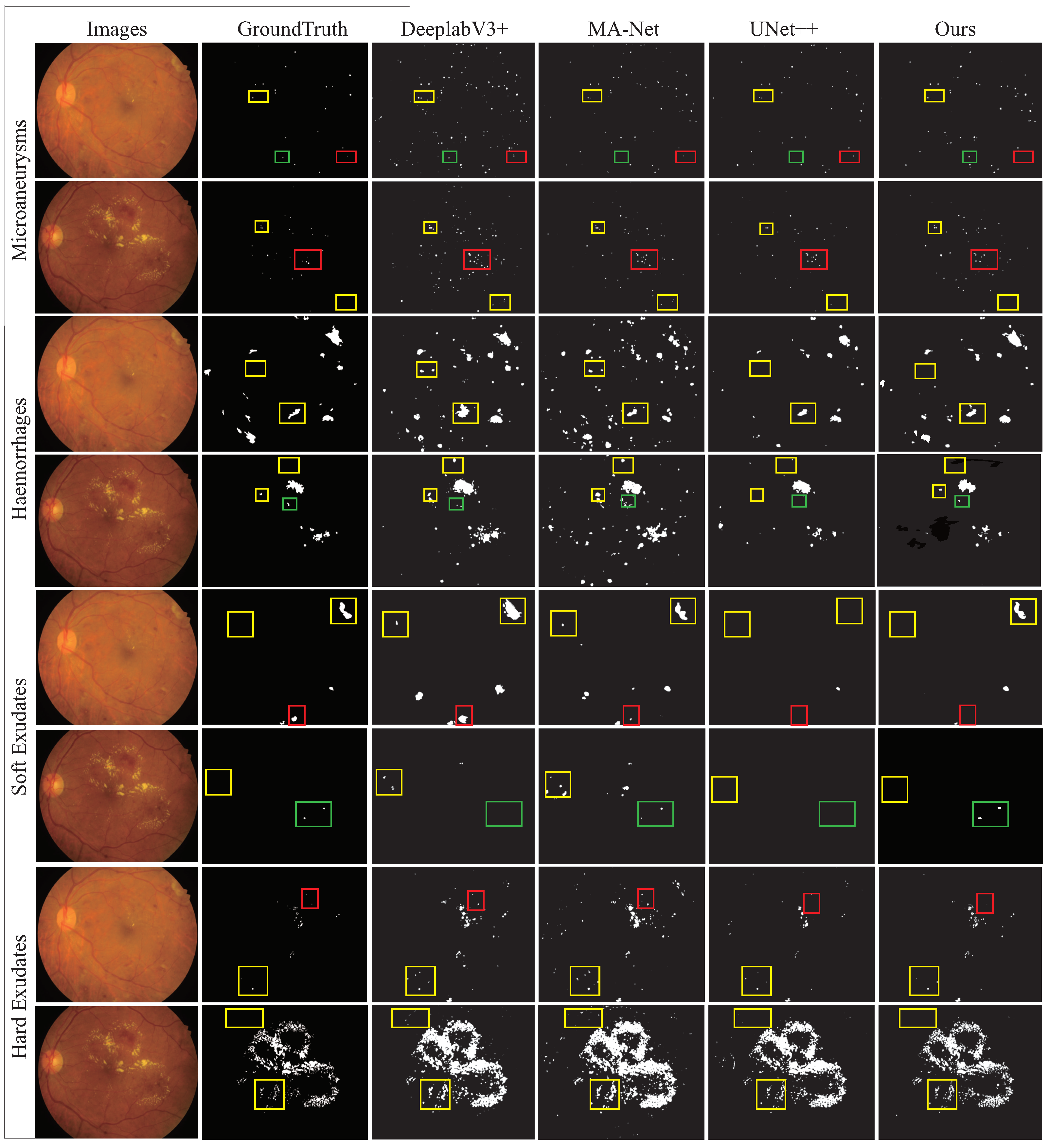
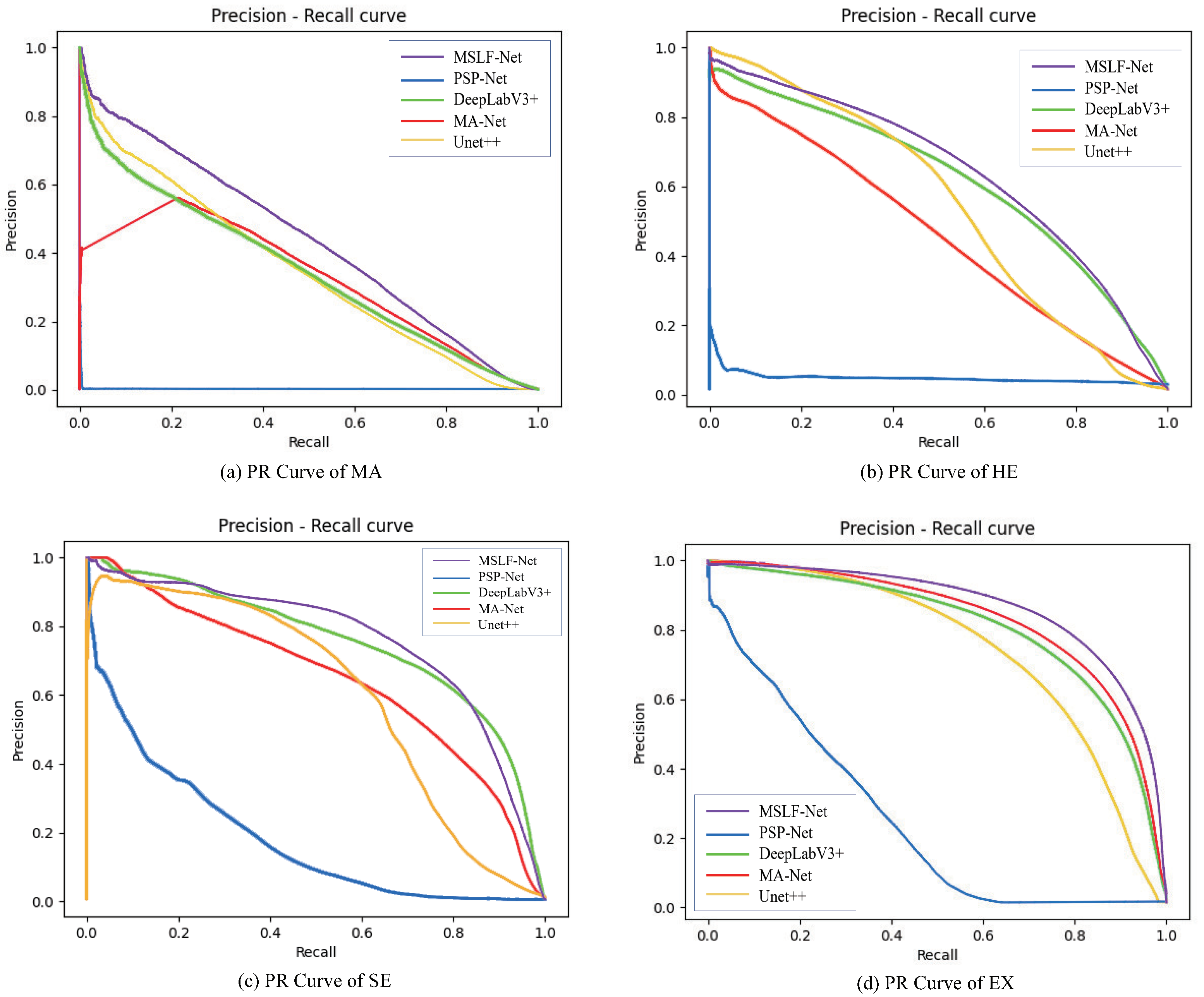
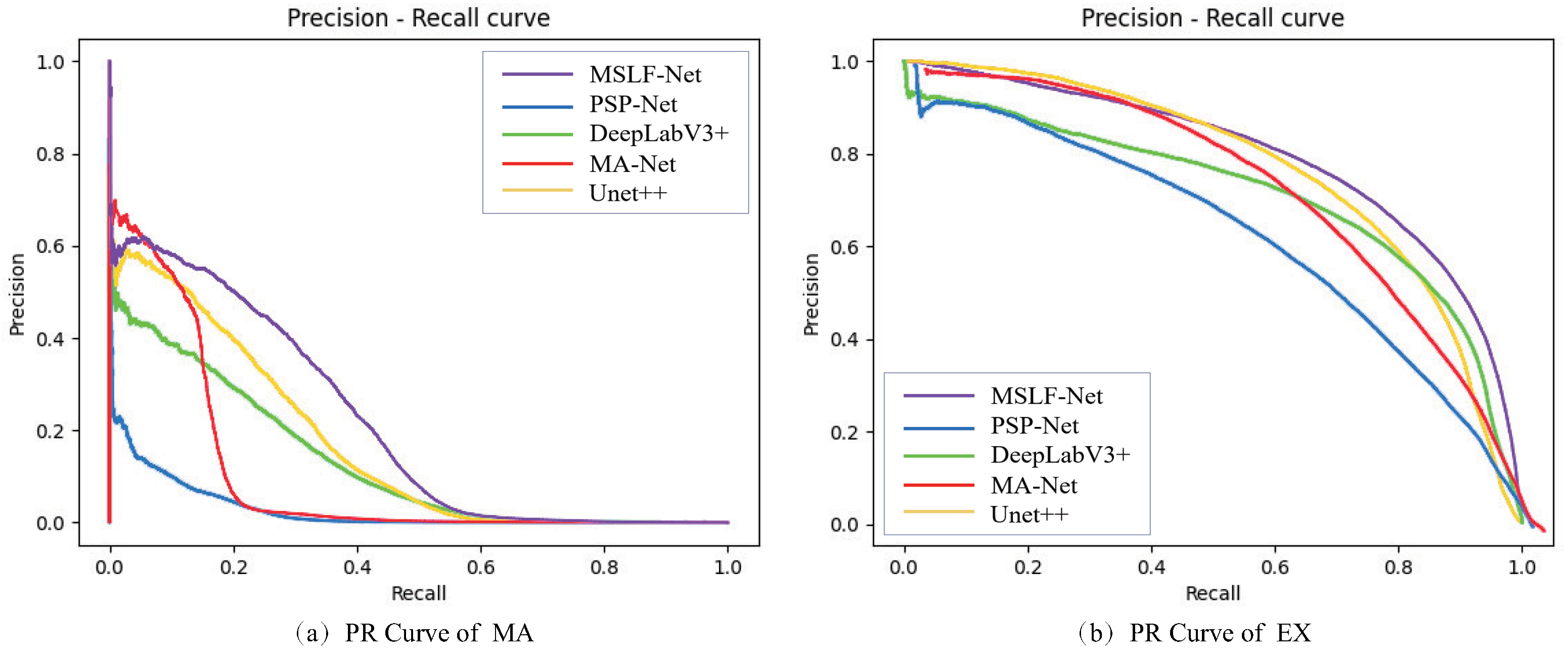
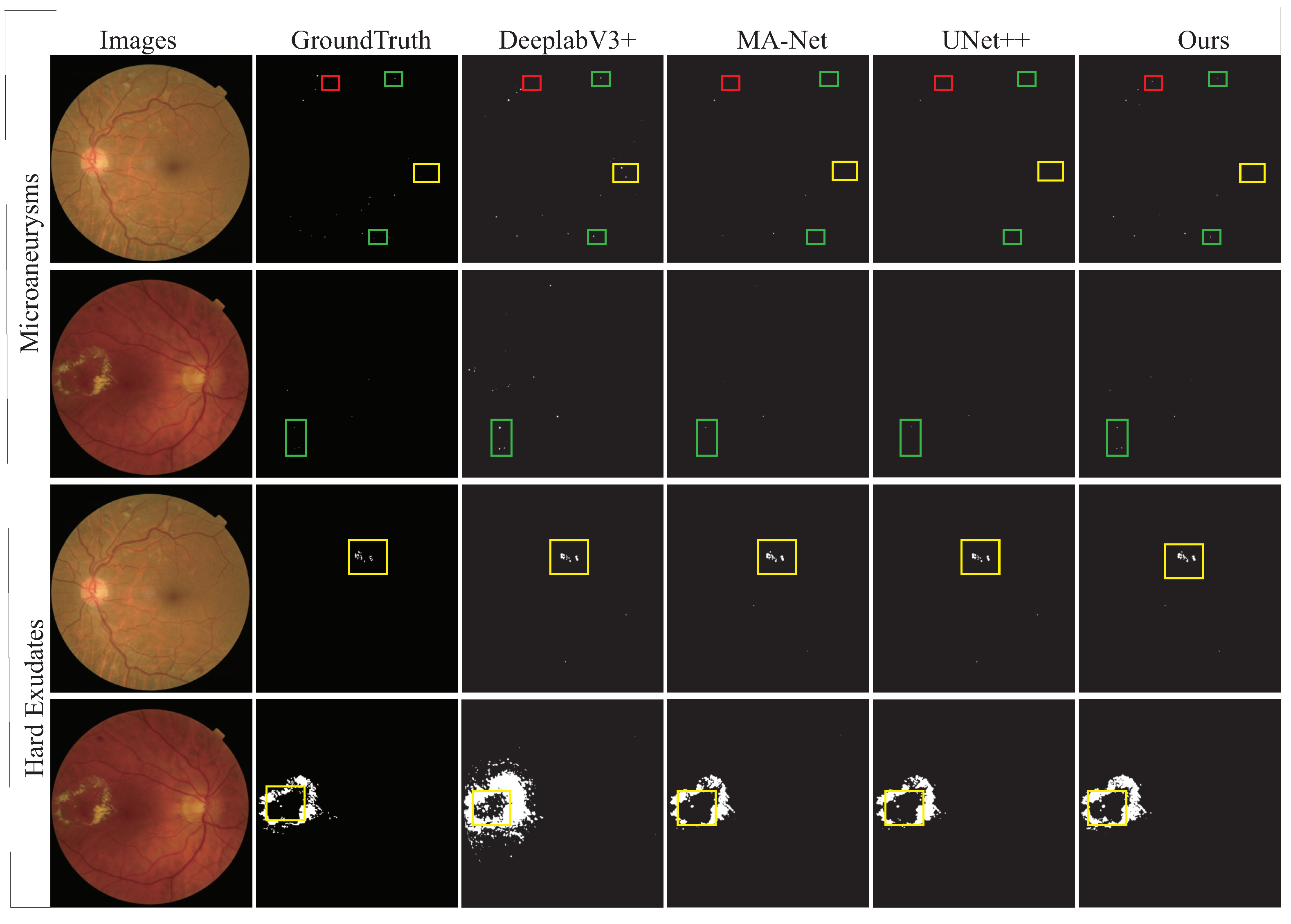
4.1. Comparative Experiments
4.2. Ablation Studies on IDRID Dataset
4.2.1. Analysis of the Model Components
4.2.2. Analysis of the Hybrid Loss Function
4.2.3. Analysis of the Image Augmentation
4.3. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Danaei, G.; Finucane, M.M.; Lu, Y.; Singh, G.M.; Cowan, M.J.; Paciorek, C.J.; Lin, J.K.; Farzadfar, F.; Khang, Y.H.; Stevens, G.A.; et al. National, regional, and global trends in fasting plasma glucose and diabetes prevalence since 1980: Systematic analysis of health examination surveys and epidemiological studies with 370 country-years and 2·7 million participants. Lancet 2011, 378, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Chen, H.; Chen, W.; Zhang, M. Prevalence and risk factors for diabetic retinopathy in China: A multi-hospital-based cross-sectional study. Br. J. Ophthalmol. 2017, 101, 1591–1595. [Google Scholar] [CrossRef] [PubMed]
- Atlas, I.D. A review of studies utilising retinal photography on the global prevalence of diabetes related retinopathy between 2015 and 2018. Thomas RL, Halim S, Gurudas S, Sivaprasad S, Owens DR. Diabetes Res. Clin. Pract. 2019, 157, 107840. [Google Scholar]
- Wilkinson, C.P.; Ferris, F.L., III; Klein, R.E.; Lee, P.P.; Agardh, C.D.; Davis, M.; Dills, D.; Kampik, A.; Pararajasegaram, R.; Verdaguer, J.T.; et al. Proposed international clinical diabetic retinopathy and diabetic macular edema disease severity scales. Ophthalmology 2003, 110, 1677–1682. [Google Scholar] [CrossRef]
- Bellemo, V.; Lim, Z.W.; Lim, G.; Nguyen, Q.D.; Xie, Y.; Yip, M.Y.; Hamzah, H.; Ho, J.; Lee, X.Q.; Hsu, W.; et al. Artificial intelligence using deep learning to screen for referable and vision-threatening diabetic retinopathy in Africa: A clinical validation study. Lancet Digit. Health 2019, 1, e35–e44. [Google Scholar] [CrossRef]
- Voets, M.; Mllersen, K.; Bongo, L.A. Replication study: Development and validation of deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. arXiv 2018, arXiv:1803.04337. [Google Scholar]
- Patton, N.; Aslam, T.M.; MacGillivray, T.; Deary, I.J.; Dhillon, B.; Eikelboom, R.H.; Yogesan, K.; Constable, I.J. Retinal image analysis: Concepts, applications and potential. Prog. Retin. Eye Res. 2006, 25, 99–127. [Google Scholar] [CrossRef]
- Shivaram, J.M.; Patil, R.; Aravind, H. Automated detection and quantification of haemorrhages in diabetic retinopathy images using image arithmetic and mathematical morphology methods. Int. J. Recent Trends Eng. 2009, 2, 174. [Google Scholar]
- Weiwei, G.; Jianxin, S.; Yuliang, W.; Chun, L.; Jing, Z. Application of local adaptive region growth method based on multi template matching in automatic detection of retinal hemorrhage. Spectrosc. Spectr. Anal. 2013, 33, 448–453. [Google Scholar]
- Zivkovic, Z. Improved adaptive Gaussian mixture model for background subtraction. In Proceedings of the 17th International Conference on Pattern Recognition, Cambridge, UK, 23–26 August 2004. [Google Scholar]
- Huang, Y.; Hu, J. Residual Neural Network Based Classification of Macular Edema in OCT. In Proceedings of the 2019 IEEE 31st International Conference on Tools with Artificial Intelligence, Portland, OR, USA, 4–6 November 2019. [Google Scholar]
- Xue, J.; Yan, S.; Qu, J.; Qi, F.; Qiu, C.; Zhang, H.; Chen, M.; Liu, T.; Li, D.; Liu, X. Deep membrane systems for multitask segmentation in diabetic retinopathy. Knowl.-Based Syst. 2019, 183, 104887. [Google Scholar] [CrossRef]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015. [Google Scholar]
- Eftekhari, N.; Pourreza, H.R.; Masoudi, M.; Ghiasi-Shirazi, K.; Saeedi, E. Microaneurysm detection in fundus images using a two-step convolutional neural network. Biomed. Eng. Online 2019, 18, 67. [Google Scholar] [CrossRef] [PubMed]
- Mo, J.; Zhang, L.; Feng, Y. Exudate-based diabetic macular edema recognition in retinal images using cascaded deep residual networks. Neurocomputing 2018, 290, 161–171. [Google Scholar] [CrossRef]
- Li, T.; Gao, Y.; Wang, K.; Guo, S.; Liu, H.; Kang, H. Diagnostic assessment of deep learning algorithms for diabetic retinopathy screening. Inf. Sci. 2019, 501, 511–522. [Google Scholar] [CrossRef]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-decoder with atrous separable convolution for semantic image segmentation. In Proceedings of the European Conference on Computer Vision, Munich, Germany, 8–14 September 2018. [Google Scholar]
- Guo, S.; Li, T.; Kang, H.; Li, N.; Zhang, Y.; Wang, K. L-Seg: An end-to-end unified framework for multi-lesion segmentation of fundus images. Neurocomputing 2019, 349, 52–63. [Google Scholar] [CrossRef]
- Tan, J.H.; Fujita, H.; Sivaprasad, S.; Bhandary, S.V.; Rao, A.K.; Chua, K.C.; Acharya, U.R. Automated segmentation of exudates, haemorrhages, microaneurysms using single convolutional neural network. Inf. Sci. 2017, 420, 66–76. [Google Scholar] [CrossRef]
- Kou, C.; Li, W.; Liang, W.; Yu, Z.; Hao, J. Microaneurysms segmentation with a U-Net based on recurrent residual convolutional neural network. J. Med. Imaging 2019, 6, 025008. [Google Scholar] [CrossRef]
- Huang, S.; Li, J.; Xiao, Y.; Shen, N.; Xu, T. RTNet: Relation transformer network for diabetic retinopathy multi-lesion segmentation. IEEE Trans. Med. Imaging 2022, 41, 1596–1607. [Google Scholar] [CrossRef]
- Zhang, L.; Feng, S.; Duan, G.; Li, Y.; Liu, G. Detection of microaneurysms in fundus images based on an attention mechanism. Genes 2019, 10, 817. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, X.; Peng, C.; Xue, X.; Sun, J. Exfuse: Enhancing feature fusion for semantic segmentation. In Proceedings of the European Conference on Computer Vision, Munich, Germany, 8–14 September 2018. [Google Scholar]
- Song, T.; Zhang, X.; Ding, M.; Rodriguez-Paton, A.; Wang, S.; Wang, G. DeepFusion: A deep learning based multi-scale feature fusion method for predicting drug-target interactions. Methods 2022, 204, 269–277. [Google Scholar] [CrossRef]
- Ramamurthy, K.; George, T.T.; Shah, Y.; Sasidhar, P. A Novel Multi-Feature Fusion Method for Classification of Gastrointestinal Diseases Using Endoscopy Images. Diagnostics 2022, 12, 2316. [Google Scholar] [CrossRef]
- Porwal, P.; Pachade, S.; Kokare, M.; Deshmukh, G.; Mériaudeau, F. IDRiD: Diabetic Retinopathy—Segmentation and Grading Challenge. Med. Image Anal. 2019, 59, 101561. [Google Scholar] [CrossRef]
- Decenciere, E.; Cazuguel, G.; Zhang, X.; Thibault, G.; Klein, J.C.; Meyer, F.; Marcotegui, B.; Quellec, G.; Lamard, M.; Danno, R.; et al. TeleOphta: Machine learning and image processing methods for teleophthalmology. Irbm 2013, 34, 196–203. [Google Scholar] [CrossRef]
- Staal, J.; Abràmoff, M.D.; Niemeijer, M.; Viergever, M.A.; Van Ginneken, B. Ridge-based vessel segmentation in color images of the retina. IEEE Trans. Med. Imaging 2004, 23, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Lee, C.Y.; Xie, S.; Gallagher, P.; Zhang, Z.; Tu, Z. Deeply-supervised nets. In Proceedings of the Artificial Intelligence and Statistics, San Diego, CA, USA, 9–12 May 2015. [Google Scholar]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016. [Google Scholar]
- Liu, W.; Anguelov, D.; Erhan, D.; Szegedy, C.; Reed, S.; Fu, C.Y.; Berg, A.C. Ssd: Single shot multibox detector. In Proceedings of the European Conference on Computer Vision, Amsterdam, The Netherlands, 8–16 October 2016. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2017. [Google Scholar]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–21 June 2009. [Google Scholar]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 2019, 39, 1856–1867. [Google Scholar] [CrossRef]
- Fan, T.; Wang, G.; Li, Y.; Wang, H. Ma-net: A multi-scale attention network for liver and tumor segmentation. IEEE Access 2020, 8, 179656–179665. [Google Scholar] [CrossRef]
- Sinthanayothin, C.; Boyce, J.F.; Williamson, T.H.; Cook, H.L.; Mensah, E.; Lal, S.; Usher, D. Automated detection of diabetic retinopathy on digital fundus images. Diabet. Med. 2002, 19, 105–112. [Google Scholar] [CrossRef]
- Furtado, P.; Baptista, C.; Paiva, I. Segmentation of Diabetic Retinopathy Lesions by Deep Learning:Achievements and Limitations. In Proceedings of the 13th International Joint Conference on Biomedical Engineering Systems and Technologies, Valletta, Malta, 24–26 February 2020. [Google Scholar]
- Antal, B.; Hajdu, A. An ensemble-based system for automatic screening of diabetic retinopathy. Knowl.-Based Syst. 2014, 60, 20–27. [Google Scholar] [CrossRef]




| Stage | Layers | Input | Input Size | Operations | Output Size |
|---|---|---|---|---|---|
| Encoder | Block1 | Input image | 1440 × 960 × 3 | Conv Maxpool | 720 × 480 × 64 |
| Block2 | Block1 | 720 × 480 × 64 | Conv Maxpool | 360 × 240 × 128 | |
| Block3 | Block2 | 360 × 240 × 128 | Conv Maxpool | 180 × 120 × 256 | |
| Block4 | Block3 | 180 × 120 × 256 | Conv Maxpool | 90 × 60 × 512 | |
| Block5 | Block4 | 90 × 60 × 512 | Conv Maxpool | 90 × 60 × 512 | |
| Decoder | UpBlock5 | Block5 | 90 × 60 × 512 | Conv Upsample Concat | 180 × 120 × 256 |
| UpBlock4 | Block5 Block4 | 180 × 120 × 512 | Concat Conv UpSample | 360 × 240 × 128 | |
| UpBlock3 | Block4 Block3 | 360 × 240 × 256 | Concat Conv UpSample | 720 × 480 × 64 | |
| UpBlock2 | Block3 Block2 | 720 × 480 × 128 | Concat Conv UpSample | 1440 × 960 × 64 | |
| UpBlock1 | Block2 Block1 | 1440 × 960 × 64 | Concat Conv UpSample | 1440 × 960 × 5 | |
| MSFE | Layer5 | UpBolck5 | 90 × 60 × 512 | Conv UpSample | 1440 × 960 × 5 |
| Layer4 | UpBolck4 | 180 × 120 × 512 | Conv UpSample | 1440 × 960 × 5 | |
| Layer3 | UpBolck3 | 360 × 240 × 256 | Conv UpSample | 1440 × 960 × 5 | |
| Layer2 | UpBolck2 | 720 × 480 × 128 | Conv UpSample | 1440 × 960 × 5 | |
| Layer1 | UpBolck1 | 1440 × 960 × 64 | Conv | 1440 × 960 × 5 | |
| MLFF | Fusion | Layer 1-5 | 1440 × 960 × 25 | Fusion | 1440 × 960 × 5 |
| Method | MA | HE | SE | EX | mAUC_PR |
|---|---|---|---|---|---|
| FRCN | 0.3383 | 0.4200 | 0.5152 | 0.5472 | 0.4552 |
| UNet++ | 0.3535 | 0.5472 | 0.6052 | 0.7415 | 0.5619 |
| MANet | 0.3220 | 0.4607 | 0.6466 | 0.8294 | 0.5647 |
| PSPNet | 0.0024 | 0.0340 | 0.1825 | 0.2483 | 0.1168 |
| DeeplabV3+ | 0.3500 | 0.6142 | 0.7416 | 0.8042 | 0.6275 |
| Paper [27] | 0.3499 | 0.5273 | 0.6449 | 0.7229 | 0.5613 |
| L-Seg | 0.4627 | 0.6374 | 0.7113 | 0.7945 | 0.6515 |
| RTNet-base | 0.4279 | 0.6570 | 0.5968 | 0.8659 | 0.6369 |
| MSLF-Net | 0.4393 | 0.6411 | 0.7597 | 0.8644 | 0.6761 |
| Method | MA | HE | SE | EX | mAUC_PR |
|---|---|---|---|---|---|
| FRCN | 0.0269 | - | - | 0.1675 | 0.0972 |
| UNet++ | 0.1636 | - | - | 0.7649 | 0.4643 |
| MANet | 0.1098 | - | - | 0.7216 | 0.4157 |
| PSPNet | 0.0237 | - | - | 0.6161 | 0.3199 |
| DeeplabV3+ | 0.1281 | - | - | 0.7105 | 0.4193 |
| Paper [27] | 0.1367 | - | - | 0.6578 | 0.3973 |
| L-Seg | 0.1687 | - | - | 0.4171 | 0.2929 |
| MSLF-Net | 0.2179 | - | - | 0.8001 | 0.5090 |
| Method | MA | HE | SE | EX | mAUC_PR |
|---|---|---|---|---|---|
| Baseline | 0.4059 | 0.6348 | 0.7017 | 0.8460 | 0.6471 |
| +MSFE | 0.3959 | 0.6446 | 0.7160 | 0.8464 | 0.6507 |
| +MLFF | 0.4393 | 0.6411 | 0.7597 | 0.8644 | 0.6761 |
| MA | HE | SE | EX | mAUC_PR | |
|---|---|---|---|---|---|
| −1 | 0.3826 | 0.6227 | 0.6635 | 0.8576 | 0.6316 |
| 0 | 0.4017 | 0.6174 | 0.7533 | 0.8457 | 0.6545 |
| 0.1 | 0.4234 | 0.6154 | 0.7490 | 0.8561 | 0.6604 |
| 0.2 | 0.4187 | 0.6162 | 0.7116 | 0.8091 | 0.6389 |
| 0.3 | 0.4325 | 0.6360 | 0.7300 | 0.8410 | 0.6599 |
| 0.4 | 0.4094 | 0.6179 | 0.7623 | 0.8469 | 0.6591 |
| 0.5 | 0.4198 | 0.6489 | 0.7324 | 0.8540 | 0.6638 |
| 0.6 | 0.4339 | 0.6290 | 0.7451 | 0.8517 | 0.6649 |
| 0.7 | 0.4227 | 0.6311 | 0.7508 | 0.8278 | 0.6581 |
| 0.8 | 0.3889 | 0.6431 | 0.7498 | 0.8419 | 0.6559 |
| 0.9 | 0.4459 | 0.6472 | 0.7394 | 0.8513 | 0.6709 |
| 1.0 | 0.4393 | 0.6411 | 0.7597 | 0.8644 | 0.6761 |
| 1.1 | 0.4469 | 0.6354 | 0.7211 | 0.8511 | 0.6636 |
| 1.2 | 0.4328 | 0.6390 | 0.7327 | 0.8479 | 0.6631 |
| Operation | MA | HE | SE | EX | mAUC_PR |
|---|---|---|---|---|---|
| baseline | 0.3326 | 0.5568 | 0.6236 | 0.7594 | 0.5681 |
| +Augmentation | 0.3742 | 0.5754 | 0.6756 | 0.8394 | 0.6162 |
| +FOV | 0.4393 | 0.6411 | 0.7597 | 0.8644 | 0.6761 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yan, H.; Xie, J.; Zhu, D.; Jia, L.; Guo, S. MSLF-Net: A Multi-Scale and Multi-Level Feature Fusion Net for Diabetic Retinopathy Segmentation. Diagnostics 2022, 12, 2918. https://doi.org/10.3390/diagnostics12122918
Yan H, Xie J, Zhu D, Jia L, Guo S. MSLF-Net: A Multi-Scale and Multi-Level Feature Fusion Net for Diabetic Retinopathy Segmentation. Diagnostics. 2022; 12(12):2918. https://doi.org/10.3390/diagnostics12122918
Chicago/Turabian StyleYan, Haitao, Jiexin Xie, Deliang Zhu, Lukuan Jia, and Shijie Guo. 2022. "MSLF-Net: A Multi-Scale and Multi-Level Feature Fusion Net for Diabetic Retinopathy Segmentation" Diagnostics 12, no. 12: 2918. https://doi.org/10.3390/diagnostics12122918
APA StyleYan, H., Xie, J., Zhu, D., Jia, L., & Guo, S. (2022). MSLF-Net: A Multi-Scale and Multi-Level Feature Fusion Net for Diabetic Retinopathy Segmentation. Diagnostics, 12(12), 2918. https://doi.org/10.3390/diagnostics12122918






