Pap Smear Images Classification Using Machine Learning: A Literature Matrix
Abstract
:1. Introduction
2. Review Method
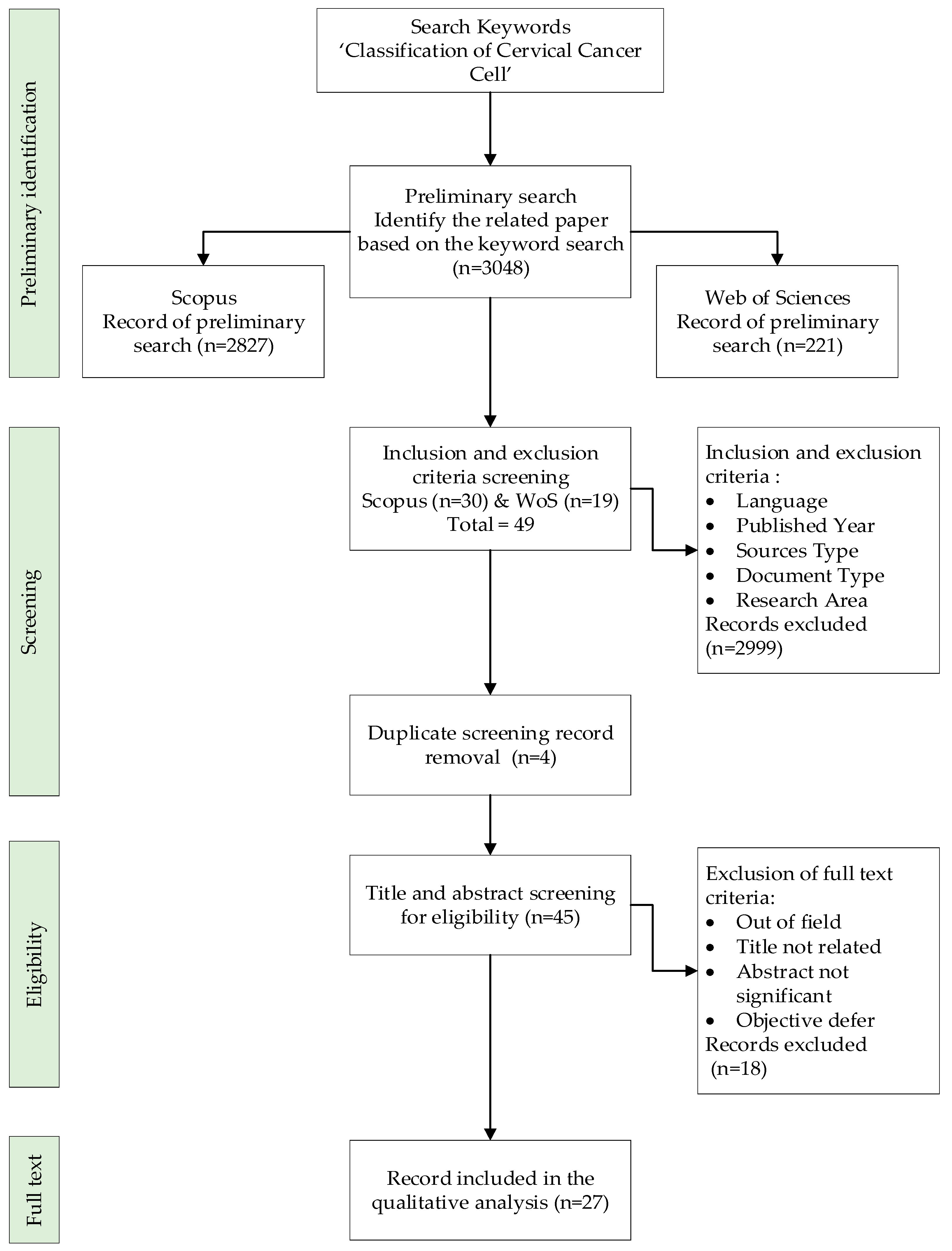
2.1. Preliminary Identification
2.2. Screening
2.3. Eligibility
2.4. Data Abstraction and Analysis
3. Results and Findings
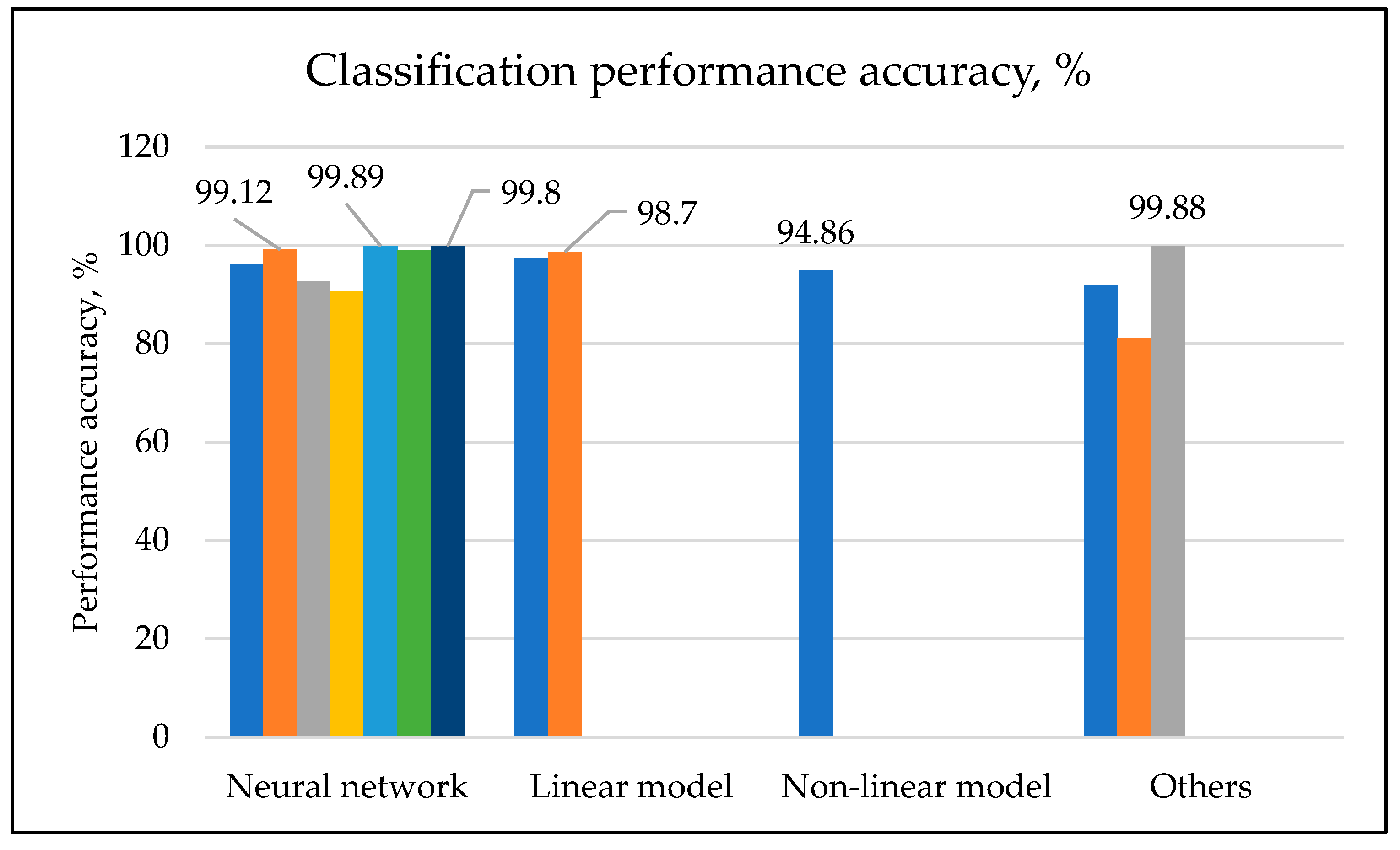
Classification of Cells Based on Machine Learning Approach
4. Discussion and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jia, D.; He, Z.; Zhang, C.; Yin, W.; Wu, N.; Li, Z. Detection of cervical cancer cells in complex situation based on improved YOLOv3 network. Multimed. Tools Appl. 2022, 81, 8939–8961. [Google Scholar] [CrossRef]
- Balaji, G.N.; Suryanarayana, S.V.; Sengathir, J. Enhanced boykov’s graph cuts based segmentation for cervical cancer detection. EAI Endorsed Trans. Pervasive Health Technol. 2021, 7, e3. [Google Scholar] [CrossRef]
- Li, J.; Dou, Q.; Yang, H.; Liu, J.; Fu, L.; Zhang, Y.; Zheng, L.; Zhang, D. Cervical cell multi-classification algorithm using global context information and attention mechanism. Tissue Cell 2022, 74, 101677. [Google Scholar] [CrossRef] [PubMed]
- Kumawat, G.; Vishwakarma, S.K.; Chakrabarti, P.; Chittora, P.; Chakrabarti, T.; Lin, J. Prognosis of Cervical Cancer Disease by Applying Machine Learning Techniques. J. Circuits Syst. Comput. 2022, 2350019. [Google Scholar] [CrossRef]
- Elakkiya, R.; Teja, K.S.S.; Jegatha Deborah, L.; Bisogni, C.; Medaglia, C. Imaging based cervical cancer diagnostics using small object detection—Generative adversarial networks. Multimed. Tools Appl. 2022, 81, 191–207. [Google Scholar] [CrossRef]
- Lia, M.; Horn, L.C.; Sodeikat, P.; Höckel, M.; Aktas, B.; Wolf, B. The diagnostic value of core needle biopsy in cervical cancer: A retrospective analysis. PLoS ONE 2022, 17, e0262257. [Google Scholar] [CrossRef]
- Lavanya Devi, N.; Thirumurugan, P. Cervical Cancer Classification from Pap Smear Images Using Modified Fuzzy C Means, PCA, and KNN. IETE J. Res. 2021, 68, 1591–1598. [Google Scholar] [CrossRef]
- Waly, M.I.; Sikkandar, M.Y.; Aboamer, M.A.; Kadry, S.; Thinnukool, O. Optimal deep convolution neural network for cervical cancer diagnosis model. Comput. Mater. Contin. 2022, 70, 3297–3309. [Google Scholar] [CrossRef]
- Vaiyapuri, T.; Alaskar, H.; Syed, L.; Aljohani, E.; Alkhayyat, A.; Shankar, K.; Kumar, S. Modified metaheuristics with stacked sparse denoising autoencoder model for cervical cancer classification. Comput. Electr. Eng. 2022, 103, 108292. [Google Scholar] [CrossRef]
- Lilhore, U.K.; Poongodi, M.; Kaur, A.; Simaiya, S.; Algarni, A.D.; Elmannai, H.; Vijayakumar, V.; Tunze, G.B.; Hamdi, M. Hybrid Model for Detection of Cervical Cancer Using Causal Analysis and Machine Learning Techniques. Comput. Math. Methods Med. 2022, 2022, 4688327. [Google Scholar] [CrossRef]
- Chitra, B.; Kumar, S. Early cervical cancer diagnosis using Sooty tern-optimized CNN-LSTM classifier. Int. J. Imaging Syst. Technol. 2022, 32, 1846–1860. [Google Scholar] [CrossRef]
- Kaur, H.; Sharma, R.; Kaur, L. Automated Cervical Cancer Image Analysis using Deep Learning Techniques from Pap-Smear Images: A Literature Review. In Proceedings of the 2021 9th International Conference on Reliability, Infocom Technologies and Optimization (Trends and Future Directions) (ICRITO), Noida, India, 3–4 September 2021; pp. 1–7. [Google Scholar] [CrossRef]
- Diniz, D.N.; Rezende, M.T.; Bianchi, A.G.C.; Carneiro, C.M.; Luz, E.J.S.; Moreira, G.J.P.; Ushizima, D.M.; de Medeiros, F.N.S.; Souza, M.J.F. A deep learning ensemble method to assist cytopathologists in pap test image classification. J. Imaging 2021, 7, 111. [Google Scholar] [CrossRef]
- Jahan, S.; Islam, M.D.S.; Islam, L.; Rashme, T.Y.; Prova, A.A.; Paul, B.K.; Islam, M.D.M.; Mosharof, M.K. Automated invasive cervical cancer disease detection at early stage through suitable machine learning model. SN Appl. Sci. 2021, 3, 806. [Google Scholar] [CrossRef]
- Diniz, D.N.; Rezende, M.T.; Bianchi, A.G.C.; Carneiro, C.M.; Ushizima, D.M.; de Medeiros, F.N.S.; Souza, M.J.F. A hierarchical feature-based methodology to perform cervical cancer classification. Appl. Sci. 2021, 11, 4091. [Google Scholar] [CrossRef]
- Ali, M.M.; Ahmed, K.; Bui, F.M.; Paul, B.K.; Ibrahim, S.M.; Quinn, J.M.W.; Moni, M.A. Machine learning-based statistical analysis for early stage detection of cervical cancer. Comput. Biol. Med. 2021, 139, 104985. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Jia, D.; Li, Z.; Wu, N. Auxiliary classification of cervical cells based on multi-domain hybrid deep learning framework. Biomed. Signal Process. Control 2022, 77, 103739. [Google Scholar] [CrossRef]
- Liu, W.; Li, C.; Xu, N.; Jiang, T.; Rahaman, M.M.; Sun, H.; Wu, X.; Hu, W.; Chen, H.; Sun, C.; et al. CVM-Cervix: A hybrid cervical Pap-smear image classification framework using CNN, visual transformer and multilayer perceptron. Pattern Recognit. 2022, 130, 108829. [Google Scholar] [CrossRef]
- Zhao, C.; Shuai, R.; Ma, L.; Liu, W.; Wu, M. Improving cervical cancer classification with imbalanced datasets combining taming transformers with T2T-ViT. Multimed. Tools Appl. 2022, 81, 24265–24300. [Google Scholar] [CrossRef]
- Jia, D.; Zhou, J.; Zhang, C. Detection of cervical cells based on improved SSD network. Multimed. Tools Appl. 2022, 81, 13371–13387. [Google Scholar] [CrossRef]
- Janiesch, C.; Zschech, P.; Heinrich, K. Machine learning and deep learning. Electron. Mark. 2021, 31, 685–695. [Google Scholar] [CrossRef]
- Huang, J.; Wang, T.; Zheng, D.; He, Y. Nucleus segmentation of cervical cytology images based on multi-scale fuzzy clustering algorithm. Bioengineered 2020, 11, 484–501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, X.; Che, J.; Chen, Y. Weed identification using deep learning and image processing in vegetable plantation. IEEE Access 2021, 9, 10940–10950. [Google Scholar] [CrossRef]
- Maier, A.; Syben, C.; Lasser, T.; Riess, C. A gentle introduction to deep learning in medical image processing. Z. Med. Phys. 2019, 29, 86–101. [Google Scholar] [CrossRef]
- Tekchandani, H.; Verma, S.; Londhe, N.D.; Jain, R.R.; Tiwari, A. Computer aided diagnosis system for cervical lymph nodes in CT images using deep learning. Biomed. Signal Process. Control 2022, 71, 103158. [Google Scholar] [CrossRef]
- Yaman, O.; Tuncer, T. Exemplar pyramid deep feature extraction based cervical cancer image classification model using pap-smear images. Biomed. Signal Process. Control 2022, 73, 103428. [Google Scholar] [CrossRef]
- Sellamuthu Palanisamy, V.; Athiappan, R.K.; Nagalingam, T. Pap smear based cervical cancer detection using residual neural networks deep learning architecture. Concurr. Comput. Pract. Exp. 2022, 34, e6608. [Google Scholar] [CrossRef]
- Zhao, S.; He, Y.; Qin, J.; Wang, Z.; Urzua, U. A Semi-supervised Deep Learning Method for Cervical Cell Classification. Anal. Cell. Pathol. 2022, 2022, 4376178. [Google Scholar] [CrossRef]
- Mulmule, P.V.; Kanphade, R.D. Classification of Cervical Cytology Overlapping Cell Images with Transfer Learning Architectures. Biomed. Pharmacol. J. 2022, 15, 277–284. [Google Scholar] [CrossRef]
- Chen, W.; Gao, L.; Li, X.; Shen, W. Lightweight convolutional neural network with knowledge distillation for cervical cells classification. Biomed. Signal Process. Control 2022, 71, 103177. [Google Scholar] [CrossRef]
- Chen, M.; Li, W.; Hao, Y.; Qian, Y.; Humar, I. Edge cognitive computing based smart healthcare system. Futur. Gener. Comput. Syst. 2018, 86, 403–411. [Google Scholar] [CrossRef]
- Shamim Hossain, M.; Amin, S.U.; Alsulaiman, M.; Muhammad, G. Applying deep learning for epilepsy seizure detection and brain mapping visualization. ACM Trans. Multimed. Comput. Commun. Appl. 2019, 15, 1–17. [Google Scholar] [CrossRef]
- Lu, J.; Song, E.; Ghoneim, A.; Alrashoud, M. Machine learning for assisting cervical cancer diagnosis: An ensemble approach. Futur. Gener. Comput. Syst. 2020, 106, 199–205. [Google Scholar] [CrossRef]
- Elakkiya, R.; Subramaniyaswamy, V.; Vijayakumar, V.; Mahanti, A. Cervical Cancer Diagnostics Healthcare System Using Hybrid Object Detection Adversarial Networks. IEEE J. Biomed. Health Inform. 2022, 26, 1464–1471. [Google Scholar] [CrossRef] [PubMed]
- Hiebl, M.R.W. Sample Selection in Systematic Literature Reviews of Management Research. Organ. Res. Methods 2021. [Google Scholar] [CrossRef]
- Kalbhor, M.; Shinde, S.V.; Jude, H. Cervical cancer diagnosis based on cytology pap smear image classification using fractional coefficient and machine learning classifiers. TELKOMNIKA Telecommun. Comput. Electron. Control 2022, 20, 1091–1102. [Google Scholar] [CrossRef]
- Karasu Benyes, Y.; Welch, E.C.; Singhal, A.; Ou, J.; Tripathi, A. A Comparative Analysis of Deep Learning Models for Automated Cross-Preparation Diagnosis of Multi-Cell Liquid Pap Smear Images. Diagnostics 2022, 12, 1838. [Google Scholar] [CrossRef]
- Karri, M.; Annavarapu, C.S.R.; Mallik, S.; Zhao, Z.; Acharya, U.R. Multi-class nucleus detection and classification using deep convolutional neural network with enhanced high dimensional dissimilarity translation model on cervical cells. Biocybern. Biomed. Eng. 2022, 42, 797–814. [Google Scholar] [CrossRef]
- Chen, W.; Shen, W.; Gao, L.; Li, X. Hybrid Loss-Constrained Lightweight Convolutional Neural Networks for Cervical Cell Classification. Sensors 2022, 22, 3272. [Google Scholar] [CrossRef]
- Shinde, S.; Kalbhor, M.; Wajire, P. DeepCyto: A hybrid framework for cervical cancer classification by using deep feature fusion of cytology images. Math. Biosci. Eng. 2022, 19, 6415–6434. [Google Scholar] [CrossRef]
- Fekri-Ershad, S.; Ramakrishnan, S. Cervical cancer diagnosis based on modified uniform local ternary patterns and feed forward multilayer network optimized by genetic algorithm. Comput. Biol. Med. 2022, 144, 105392. [Google Scholar] [CrossRef]
- Alquran, H.; Mustafa, W.A.; Qasmieh, I.A.; Yacob, Y.M.; Alsalatie, M.; Al-Issa, Y.; Alqudah, A.M. Cervical Cancer Classification Using Combined Machine Learning and Deep Learning Approach. Comput. Mater. Contin. 2022, 72, 5117–5134. [Google Scholar] [CrossRef]
- Maylaa, T.; Windal, F.; Benhabiles, H.; Maubon, G.; Maubon, N.; Vandenhaute, E.; Collard, D. An Evaluation of Computational Learning-based Methods for the Segmentation of Nuclei in Cervical Cancer Cells from Microscopic Images. Curr. Comput. Aided Drug Des. 2022, 18, 81–94. [Google Scholar] [CrossRef]
- Liu, W.; Li, C.; Rahaman, M.M.; Jiang, T.; Sun, H.; Wu, X.; Hu, W.; Chen, H.; Sun, C.; Yao, Y.; et al. Is the aspect ratio of cells important in deep learning? A robust comparison of deep learning methods for multi-scale cytopathology cell image classification: From convolutional neural networks to visual transformers. Comput. Biol. Med. 2022, 141. [Google Scholar] [CrossRef]
- Mahmoud, H.A.H.; Alarfaj, A.A.; Hafez, A.M. A Fast Hybrid Classification Algorithm with Feature Reduction for Medical Images. Appl. Bionics Biomech. 2022, 2022, 1367366. [Google Scholar] [CrossRef]
- Baghialaxmi, R.L.P.S.; Kirubagari, B. Ensemble feature extraction model with optimal kernelized clustering algorithm for identifying the cancer from cervical histopathology images. J. Theor. Appl. Inf. Technol. 2022, 100, 4866–4877. [Google Scholar]
| Scopus | Web of Science |
|---|---|
| TITLE-ABS-KEY (classification AND cervical AND cancer AND cell) AND (LIMIT-TO (SRCTYPE, “j”)) AND (LIMIT-TO (PUBSTAGE, “final”)) AND (LIMIT-TO (DOCTYPE, “ar”)) AND (EXCLUDE (SUBJAREA, “MEDI”)) AND (LIMIT-TO (PUBYEAR, 2022)) AND (LIMIT-TO (LANGUAGE, “English”)) Date of access: October 2022 | Classification of cervical cancer cell (Abstract) or classification of cervical cancer cell (Title) or classification of cervical cancer cell (Author Key-words) and 2022 (Publication Years) and Article (Document Types) and English (Languages) and Oncology or Engineering Biomedical or Engineering Electrical Electronic (Web of Science Categories). Date of access: October 2022 |
| Criterion | Inclusion | Exclusion |
|---|---|---|
| Language | English | Non-English |
| Published Year | 2022 | <2022 |
| Sources type | Journal (only research articles) | Conference proceeding |
| Document Type | Article | Letter, Review, Conference, Note |
| Research Area | Computer Science and Engineering | Besides Computer Science and Engineering |
| Title | Class and Database | Methodology | Results |
|---|---|---|---|
| CVM-Cervix: A hybrid cervical Pap smear image classification framework using CNN, visual transformer, and multilayer perceptron [18] | Database: CRIC dataset, SIPaKMeD dataset and combination of CRIC and SIPaKMeD Class: CRIC-6 class, SIPaKMeD 5 Class | Framework CVM-Cervix based on deep learning. Type: Machine learning—Neural network | Effective and potential of the proposed CVM-Cervix proved. |
| A Comparative Analysis of Deep Learning Models for Automated Cross-Preparation Diagnosis of Multi-Cell Liquid Pap Smear Images [37] | Database: Thin Prep Pap dataset Class: 4 Bethesda class | An ensemble novel convolutional neural network (CNN) and a CNN with autoencoder (AE). Type: Machine learning—Neural network | All models’ accuracy >905. The individual transfer model had high variability in performance, while CNN and AE CNN did not. ResNet101 accuracy is 92.65%. |
| Multi-class nucleus detection and classification using a deep convolutional neural network with enhanced high dimensional dissimilarity translation model on cervical cells [38] | Database: Herlev dataset, SIPaKMeD dataset, CRIC dataset Class: Herlev-7 class, SIPaKMeD-5 Class, CRIC-6 class | Segmentation—hybrid system that incorporates two binary image patches obtained by a 19-layered convolutional neural network (ConvNet) model with an enhanced deep high dimensional dissimilarity translation (HDDT). A Pre trained Resnet-50 model. T-distribution stochastic neighbour embedding (t-SNE) for down-sampled. Classification using a multi-class weighted kernel extreme learning machine (WKELM) classifier via a sparse multicanonical correlation (SMCCA) method. Type: Machine learning—Neural network | Accuracy 99.12% Specificity 99.45% Sensitivity 99.25% Execution time 99.6248 s The proposed model is more effective compared to existing approaches. |
| Hybrid Loss-Constrained Lightweight Convolutional Neural Networks for Cervical Cell Classification [39] | Database: Herlev dataset Class: Herlev-7 class | Classification using hybrid loss function with label smoothing to improve the distinguishing power of lightweight convolutional neural networks (CNN) Type: Machine learning—Neural network | ShufflenetV2 results:- Accuracy 96.18% Precision 96.30% Recall 96.23% Specificity 99.08% GhostnetV2 results:- Accuracy 96.39% Precision 96.42% Recall 96.39% Specificity 99.09% |
| Detection of cervical cells based on improved SSD network [20] | Database: Herlev dataset Class: Herlev-7 class | Integration of Single Shot MultiBox Detector with the positive and negative features to address the problem of insufficient sensitivity for small objects. Type: Machine learning—Neural network | Accuracy 90.80% Mean average precision (mAP) is 81.53%, which is 7.54% and 4.92% higher than YOLO and classical SSD. |
| Detection of cervical cancer cells in a complex situation based on improved YOLOv3 network [1] | Database: Herlev dataset Class: Herlev-7 class | Detection using the YOLO algorithm. Feature extraction generalization by adding the dense block and S3Pool algorithm on the basis of the feature extraction network DarkNet-53. Clustering algorithm of improved algorithm k-means++. Type: Machine learning—Neural network | Mean average precision (mAP) is 78.87%, which is 7.54% and 4.92% higher than YOLO (You Only Look Once) and classical SSD |
| Pap smear-based cervical cancer detection using residual neural networks deep learning architecture [27] | Database: Mendeley LBC SIPaKMeD dataset Class: Mendeley LBC SIPaKMeD—4 class | Data augmentation module of DTWCT module and convolutional neural networks (CNN). Classification using ResNet 18, which defines four classes of sources for Pap smear cell images. Type: Machine learning—Neural network | Average Pap smear detection index (PDI) is 99%. |
| Cervical cell multi-classification algorithm using global context information and attention mechanism [3] | Database: SIPaKMeD dataset Class: SIPaKMeD—5 class | Convolutional neural network (L-PCNN) that integrates global context information and attention mechanism. Improved ResNet-50 backbone network for feature extraction. Type: Machine learning—Neural network | Accuracy 98.89%. Sensitivity 99.9%. Specificity 99.8%. F-measure 99.89%. |
| DeepCyto: a hybrid framework for cervical cancer classification using deep feature fusion of cytology images [40] | Database: Herlev dataset, SIPaKMeD dataset, LBC dataset Class: Herlev-7 class, SIPaKMeD-5 class, LBC-4 class | Novel classification using DeepCyto. Principal component analysis and machine learning ensemble for classification of Pap smear images. Artificial neural network with feature fusion vectors as an input for classification. Type: Machine learning—Neural network | DeepCyto is a powerful tool for precise feature extraction and Pap smear image classification. |
| Classification of Cervical Cytology Overlapping Cell Images with Transfer Learning Architectures [29] | Database: Cervix93 cervical cytology image Class: 3 class | Transfer learning using deep learning convolutional neural network. Cutting edge pretrained networks: AlexNet, ImageNet, and Places 365. Type: Machine learning—Neural network | Accuracy 99.03%. Kappa coefficient showing perfect agreement. AlexNet proved a successful assistive tool for cervical cancer detection. |
| Optimal deep convolution neural network for cervical cancer diagnosis model [8] | Database: Herlev dataset Class: Herlev-7 class | Detection using intelligent deep convolutional neural network. Classification (IDCNN-CDC) model using biomedical Pap smear images. Noise removal using Gaussian Filter. Segmentation using the Tsallis entropy technique with the dragonfly optimization. Deep learned feature using SqueezeNet. Classification using weighted extreme learning machine (ELM). Type: Machine learning—Neural network | Higher performance of the proposed technique in terms of sensitivity, specificity, accuracy, and F-Score. |
| Modified metaheuristics with stacked sparse denoising autoencoder model for cervical cancer classification [9]. | Database: Herlev dataset Class: Herlev-7 class | Novel Modified Firefly Optimization Algorithm with Deep Learning-enabled cervical cancer classification (MFFOA-DL3) model. Noise removal using Bilateral Filtering (BF)-based. Segmentation technique of Kapur’s entropy-based image to define affected area. Generate feature vectors using EfficientNet. Classification of the cell using MFFOA with Stacked Sparse Denoising Autoencoder (SSDA) model. Type: Machine learning—Neural network | The findings of a comprehensive comparison investigation revealed that the MFFOA-DL3 model outperformed other recent approaches. |
| Imaging based cervical cancer diagnostics using small object detection—generative adversarial networks [5] | Database: Herlev dataset, Colposcopy images, Clinical references Class: not applicable | An effective hybrid deep learning technique using Small-Object Detection-Generative Adversarial Networks (SOD-GAN) with Fine-tuned Stacked Autoencoder (F-SAE). Generation and discrimination of the cervical cell using Region-based Convolutional Neural Network (RCNN). Type: Machine learning—Neural network | The proposed method identifies and classifies cervical premalignant and malignant diseases based on deep characteristics without the necessity for initial classification and segmentation. |
| Cervical cancer diagnosis based on modified uniform local ternary patterns and feed forward multilayer network optimized by genetic algorithm [41] | Database: Herlev dataset Class: Herlev-7 class | Segmentation of the image using a thresholding approach. Feature extraction by applying a texture descriptor titled modified uniform local ternary patterns (MULTP). Classification of the cell using an optimized multilayer feed-forward neural network. Type: Machine learning—Neural network | MULTP, the proposed texture descriptor, is a generic operator that may be used to characterise texture features of images in numerous computer vision issues. In addition, the suggested optimization approach may be utilised to increase performance in deep networks. |
| Early cervical cancer diagnosis using Sooty tern-optimized CNN-LSTM classifier [11] | Database: Herlev dataset Class: Herlev-7 class | Augmentation process of image enhancement, image flipping, and image rotating to reduce the number of parameters necessary. Segmentation of the cancer-affected regions with the help of kernel weighted fuzzy local information c-means clustering (KWFLICM) model. Classification using the Sooty Tern Optimization (STO) algorithm with CNN-based long short-term memory classifier (CNN-LSTM). Type: Machine learning—Neural network | Accuracy 99.80%. Specificity 99%. Sensitivity 98.83%. F-Score 97.8. Improvement of 28.5% better than Random Forest and 19.46% better than ensemble classifier. |
| Hybrid Model for Detection of Cervical Cancer Using Causal Analysis and Machine Learning Techniques [10] | Database: Private Class: not applicable | Boruta analysis and SVM method for an efficient feature selection and prediction of the model for the cervical cell dataset. Type: Machine learning—Linear model | Boruta analysis shows a better performance approach compared to the existing techniques available. |
| Cervical Cancer Classification Using Combined Machine Learning and Deep Learning Approach [42] | Database: Herlev dataset Class: Herlev-2 class | Feature extraction using ResNet-101. Classification using Support vector Machine (SVM). Type: Machine learning—Linear model | Accuracy 97.30%. |
| Auxiliary classification of cervical cells based on the multi-domain hybrid deep learning framework [17] | Database: Herlev dataset, SIPaKMeD dataset, BJTU dataset Class: Herlev-2&7 class, SIPaKMeD-5 Class, BJTU-7 class | Deep features extraction using deep Convolutional Neural Network of pretrained Visual Geometry Group-19 (VGG-19). Hand-crafted images undergo the process of feature selection, clustering, and dimensionality reduction. Classification using a Support Vector Machine (SVM) classifier. Type: Machine learning-Linear model | Accuracy 98.70%. Sensitivity 98.20%. Specificity 98.90%. The suggested novel screening methodology is promising for early cervical cancer detection, with multi-domain and hybrid characteristics proving realistic in clinical practise. |
| An Evaluation of Computational Learning-based Methods for the Segmentation of Nuclei in Cervical Cancer Cells from Microscopic Images [43] | Database: Z-Stack cellular microscopy proliferation images provided by the HCS Pharma Class: not applicable | Machine learning architecture of Random Forest, Ada Boost, and MLP algorithm. Type: Machine learning—Nonlinear model | All machine learning architectures gave outstanding nuclei segmentation in cervical cancer cells but did not solve the overlapping nuclei and Z-stack segmentation problems. |
| Prognosis of Cervical Cancer Disease by Applying Machine Learning Techniques [4] | Database: Dataset of 858 cervical cancer patients with 36 risk factors and one outcome variable Class: not applicable | Analysis of the different supervised machine learning techniques. The classification algorithm used Artificial Neural Network, Bayesian Network, SVM, Random Tree, Logistic Tree and XG-Boost Tree. Selection algorithm for feature selection: relief rank, wrapper method, and LASSO regression. Type: Machine learning—Nonlinear model | Maximum accuracy achieved using XG-Boost with complete features 94.94%. This approach offers much potential for clinical use and cervical cancer cell detection. |
| Is the aspect ratio of cells important in deep learning? A robust comparison of deep learning methods for multi-scale cytopathology cell image classification: From convolutional neural networks to visual transformers [44] | Database-SIPaKMeD dataset Class: not applicable | Twenty-two deep learning models were used to classify the cervical cancer cells into two categories of standard and scaled datasets. Type: Machine learning—Nonlinear model | Deep learning models are robust to changes in the aspect ratio of cervical cells in cervical cytopathological images. |
| A Fast Hybrid Classification Algorithm with Feature Reduction for Medical Images [45] | Database: Herlev dataset Class: Herlev-7 class | A novel fast hybrid fuzzy classification algorithm with feature reduction for medical images. Integration of quantum-based grasshopper computing algorithm (QGH) with a fuzzy clustering technique for feature extraction. The second integration of the fusion technique utilises QGH with the fuzzy c-means algorithm to determine the best features. Type: Machine learning—Nonlinear model | Established the importance of the feature selection on the accuracy of the proposed classifier |
| Cervical Cancer Classification from Pap Smear Images Using Modified Fuzzy C Means, PCA, and KNN [7] | Database: Herlev dataset Class: Herlev-7 class | Geometrical and feature extraction using a novel approach of modified fuzzy c-means. Augmentation of the images using Principal Component Analysis (PCA) to maintain the uncorrelated features and thus reduce the algorithm processing time. Classification of the Pap smear image into normal and abnormal cells using K Nearest Neighbour (KNN). Type: Machine learning—Nonlinear model | Minimum accuracy 94.15%. Maximum accuracy 96.28%. Average accuracy 94.86%. Sensitivity 97.96%. Specificity 83.65%. F1-Score 96.87%. Precision 96.31%. |
| A Semi-supervised Deep Learning Method for Cervical Cell Classification [28] | Database: Herlev dataset, SIPaKMeD dataset Class: Herlev-7 class, SIPaKMeD-5 Class | A novel manual features and voting mechanism to achieve data expansion in semi-supervised learning. Clarity function to filter out higher-quality images, annotating a small amount of the high-quality images, and voting mechanism for balancing and training data. Type: Machine learning—Classifier | Accuracy 91.94%. |
| Ensemble feature extraction model with optimal kernelized clustering algorithm for identifying cancer from cervical histopathology images [46] | Database: 962 histopathological cervical images. Class: Not applicable | Provide improvement in the efficiency of the clustering network algorithm of Optimal Kernelized Fuzzy C-Means (OKFCM). Accurate histopathological image ensemble-based feature extraction model. Type: Machine learning—Clustering | Higher value of performance achieved. Precision Specificity Recall AUC Accuracy FPR FNR |
| Cervical cancer diagnosis based on cytology Pap smear image classification using fractional coefficient and machine learning classifiers [36] | Database: SIPaKMeD dataset Class: SIPaKMeD—5 class | Feature extraction using the discrete coefficient transform (DCT) and Haar transform. Classification using seven different machine learning algorithms for normal and abnormal Pap smear images. Optimization of feature extraction using fractional coefficient to form the five different sizes of feature vectors. | Accuracy 81.11% |
| Improving cervical cancer classification with imbalanced datasets combining taming transformers with T2T-ViT [19] | Database: Liquid-based cytology, Herlev dataset, SIPaKMeD dataset, Class: Liquid-based cytology—4 class, Herlev dataset—7 class, SIPaKMeD dataset—5 class | Taming transformers (CCG-taming transformers). Improve the encoder structure by introducing SE-block and MultiRes-block. Layer Normalization to standardize the data. SMOTE-Tomek Links to balance the source data set and the number of samples and weights. Classification using Tokens-to-Token Vision Transformers (T2T-ViT) combing transfer learning. | Accuracy:- 4-class liquid-based cytology Pap smear dataset 98.79%. 5-Class SIPAKMeD 99.58%. 7-Class Herlev 99.88%. Inception score (IS) 3.75. Frechet inception distance (FID) 0.71. Recall 0.32. Precision 0.65. Novel approach that applies the transformer to the generation and recognition of cervical cancer cell images. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alias, N.A.; Mustafa, W.A.; Jamlos, M.A.; Alquran, H.; Hanafi, H.F.; Ismail, S.; Rahman, K.S.A. Pap Smear Images Classification Using Machine Learning: A Literature Matrix. Diagnostics 2022, 12, 2900. https://doi.org/10.3390/diagnostics12122900
Alias NA, Mustafa WA, Jamlos MA, Alquran H, Hanafi HF, Ismail S, Rahman KSA. Pap Smear Images Classification Using Machine Learning: A Literature Matrix. Diagnostics. 2022; 12(12):2900. https://doi.org/10.3390/diagnostics12122900
Chicago/Turabian StyleAlias, Nur Ain, Wan Azani Mustafa, Mohd Aminudin Jamlos, Hiam Alquran, Hafizul Fahri Hanafi, Shahrina Ismail, and Khairul Shakir Ab Rahman. 2022. "Pap Smear Images Classification Using Machine Learning: A Literature Matrix" Diagnostics 12, no. 12: 2900. https://doi.org/10.3390/diagnostics12122900






