Progress in Oral Microbiome Related to Oral and Systemic Diseases: An Update
Abstract
1. Introduction
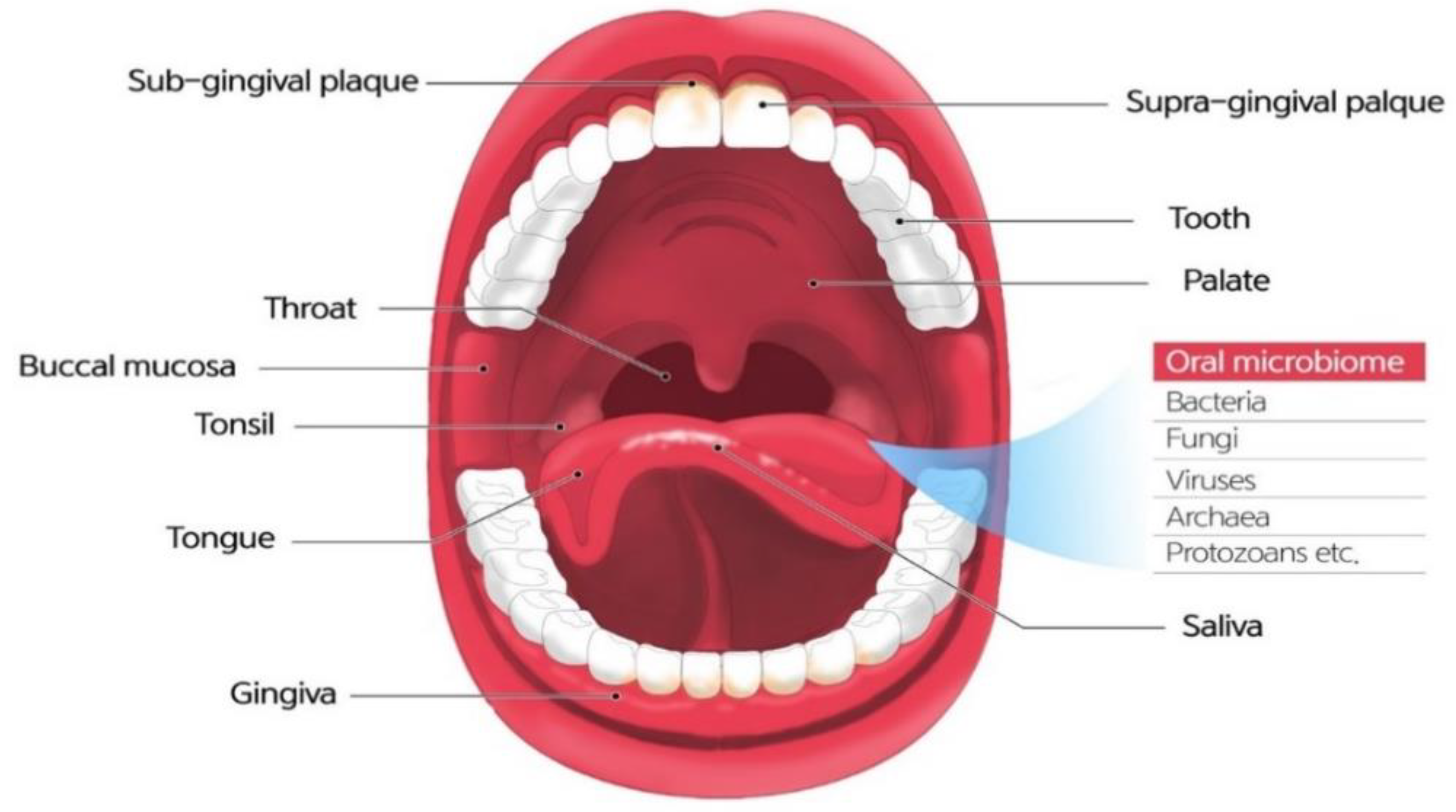
2. The Uniqueness of the Oral Cavity as a Microbial Niche
3. The Salivary Microbiome in Health
3.1. Oral Fungal and Protozoal Microbiota
3.2. Oral Viral Microbiota
4. Effective and Rapid Detection of Oral Microbiota Diversity
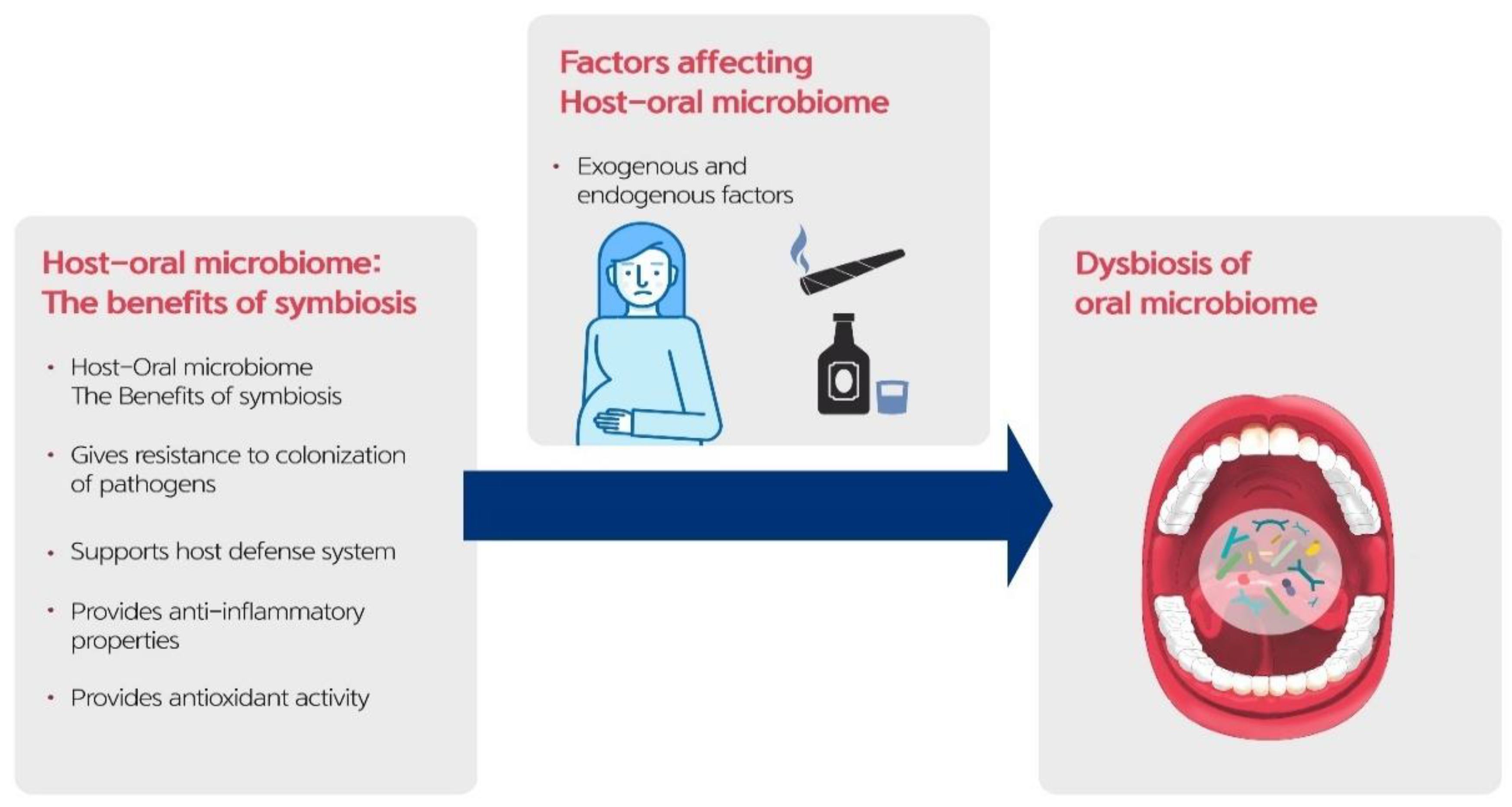
5. Factors Influencing the Oral Microbiome
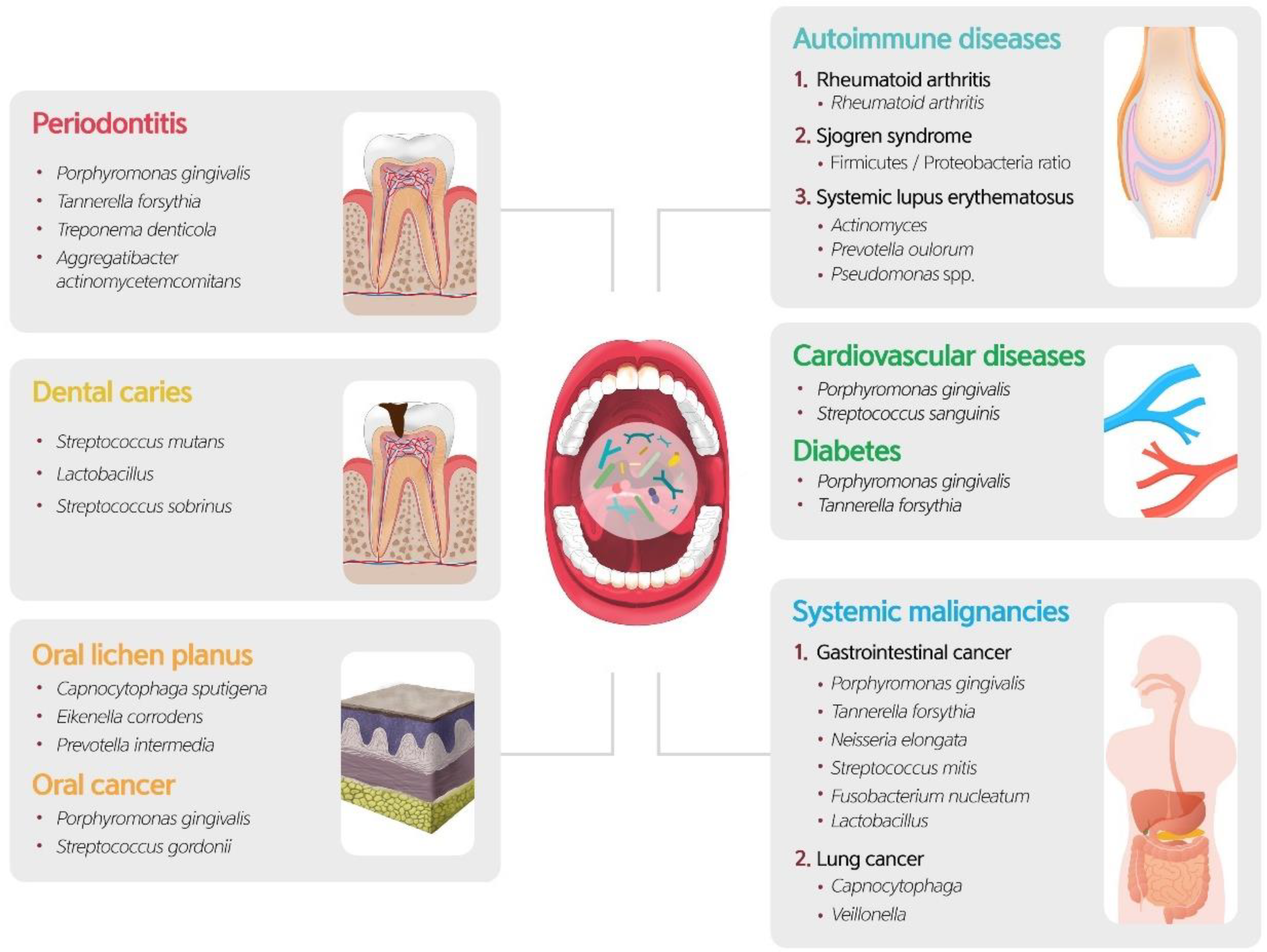
6. The Oral Microbiome within Common Oral Diseases
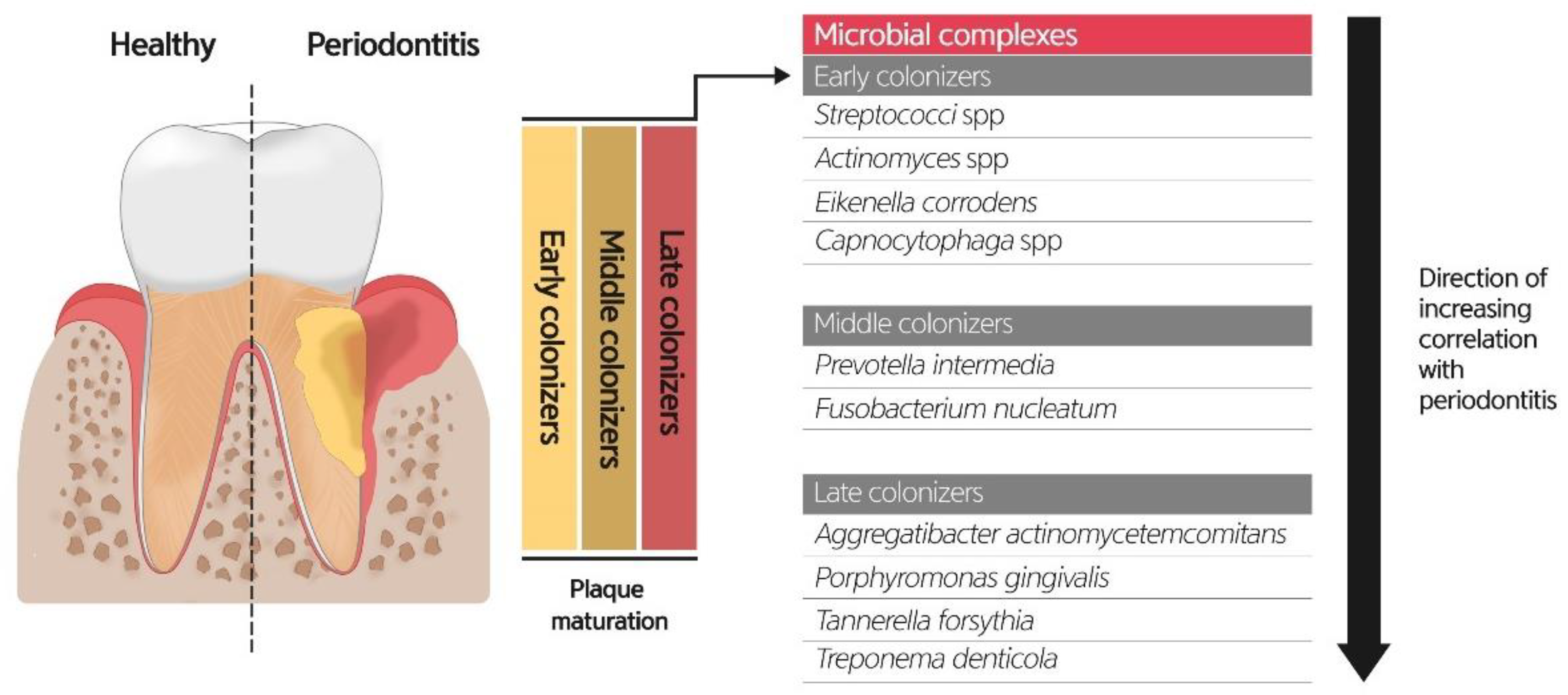
6.1. Role of Oral Microbiome on Periodontitis
6.2. Role of Oral Microbiome and Dental Caries
6.3. Role of Oral Microbiome on Oral Lichen Planus
6.4. Role of the Oral Microbiome on Pre-Malignancy and Oral Cancers
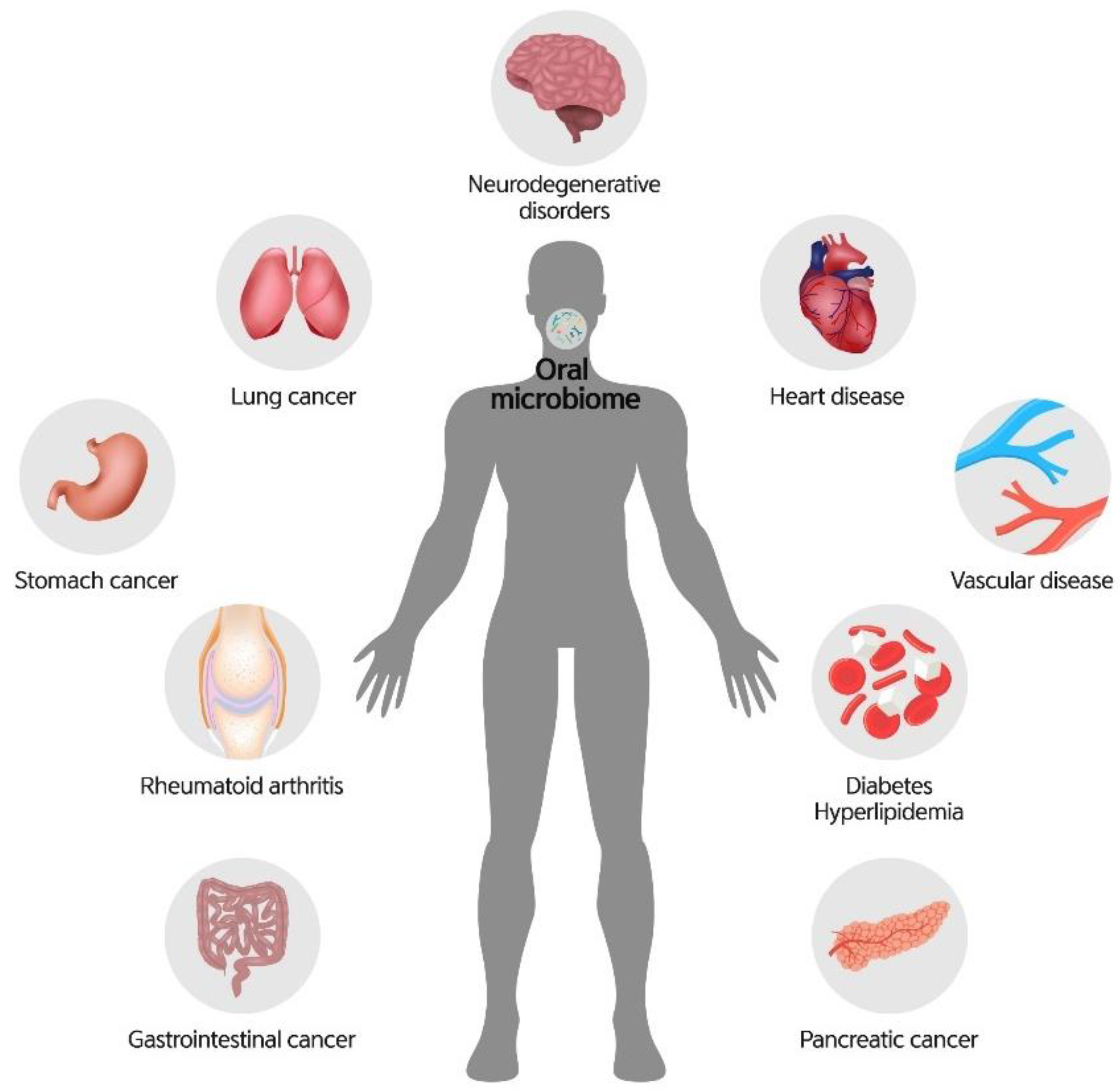
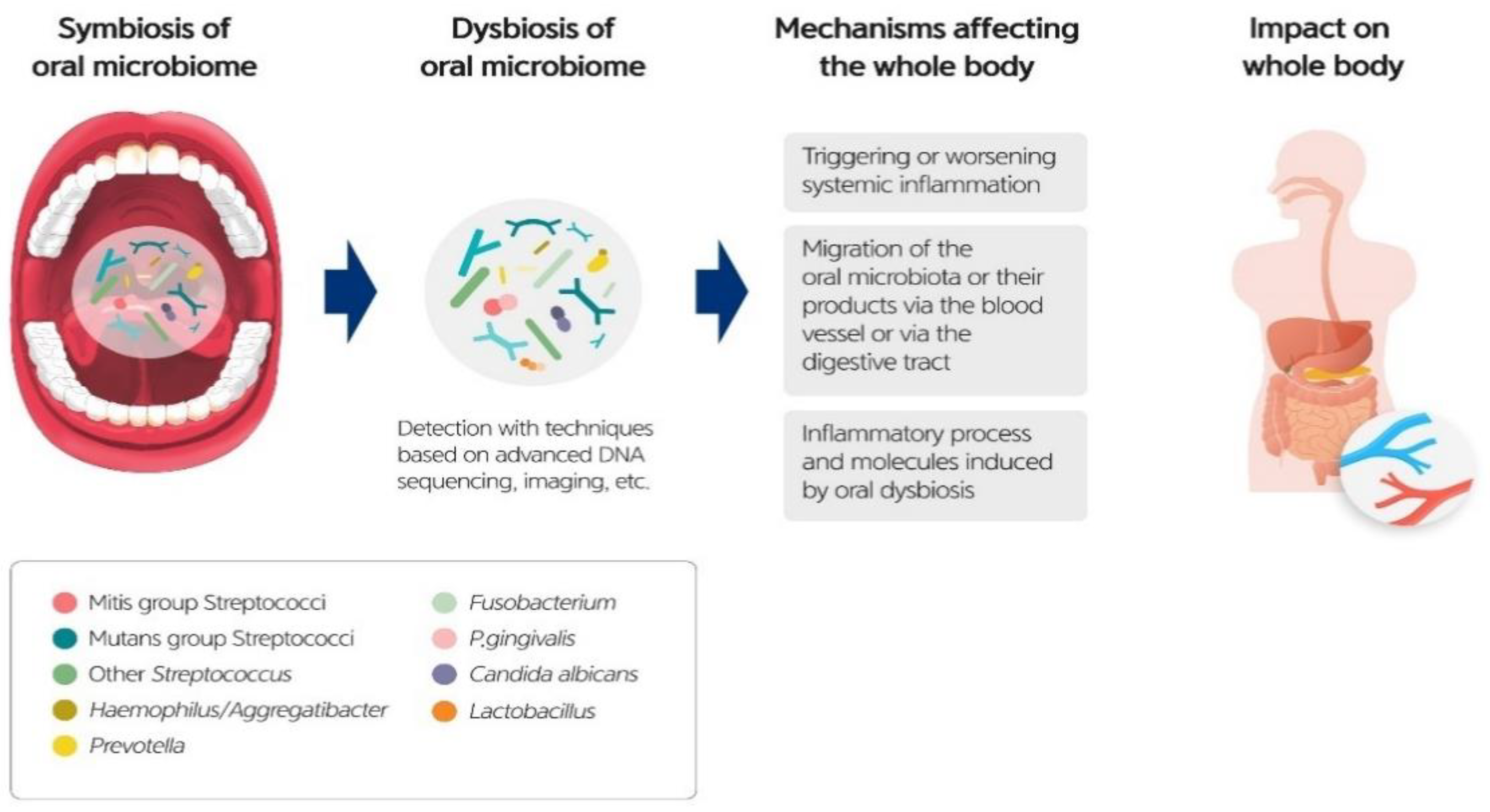
7. Oral Microbiome and Systemic Diseases
7.1. Autoimmune Disease
7.2. Systemic Malignancies
7.3. Pregnancy Outcomes
7.4. Other Systemic Diseases
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Sender, R.; Fuchs, S.; Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 2016, 164, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Delgado, S.; Cabrera-Rubio, R.; Mira, A.; Suárez, A.; Mayo, B. Microbiological survey of the human gastric ecosystem using culturing and pyrosequencing methods. Microb. Ecol. 2013, 65, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Malla, M.A.; Dubey, A.; Kumar, A.; Yadav, S.; Hashem, A.; Abd Allah, E.F. Exploring the human microbiome: The potential future role of next-generation sequencing in disease diagnosis and treatment. Front. Immunol. 2019, 9, 2868. [Google Scholar] [CrossRef] [PubMed]
- Kilian, M.; Chapple, I.L.; Hannig, M.; Marsh, P.D.; Meuric, V.; Pedersen, A.M.; Tonetti, M.S.; Wade, W.G.; Zaura, E. The oral microbiome—An update for oral healthcare professionals. Br. Dent. J. 2016, 221, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Marsh, P.D. Are dental diseases examples of ecological catastrophes? Microbiology 2003, 149, 279–294. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, F.E.; Chen, T.; Izard, J.; Paster, B.J.; Tanner, A.C.; Yu, W.H.; Lakshmanan, A.; Wade, W.G. The human oral microbiome. J. Bacteriol. 2010, 192, 5002–5017. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Xu, T.; Huang, G.; Jiang, S.; Gu, Y.; Chen, F. Oral microbiomes: More and more importance in oral cavity and whole body. Protein Cell 2018, 9, 488–500. [Google Scholar] [CrossRef]
- Bourgeois, D.; Inquimbert, C.; Ottolenghi, L.; Carrouel, F. Periodontal pathogens as risk factors of cardiovascular diseases, diabetes, rheumatoid arthritis, cancer, and chronic obstructive pulmonary disease—Is there cause for consideration? Microorganisms 2019, 7, 424. [Google Scholar] [CrossRef]
- Willis, J.R.; Gabaldón, T. The human oral microbiome in health and disease: From sequences to ecosystems. Microorganisms 2020, 8, 308. [Google Scholar] [CrossRef]
- Maukonen, J.; Mättö, J.; Suihko, M.L.; Saarela, M. Intra-individual diversity and similarity of salivary and faecal microbiota. J. Med. Microbiol. 2008, 57, 1560–1568. [Google Scholar] [CrossRef]
- Espinoza, J.L.; Harkins, D.M.; Torralba, M.; Gomez, A.; Highlander, S.K.; Jones, M.B.; Leong, P.; Saffery, R.; Bockmann, M.; Kuelbs, C.; et al. Supragingival plaque microbiome ecology and functional potential in the context of health and disease. mBio 2018, 9, e01631-18. [Google Scholar] [CrossRef] [PubMed]
- Camelo-Castillo, A.J.; Mira, A.; Pico, A.; Nibali, L.; Henderson, B.; Donos, N.; Tomás, I. Subgingival microbiota in health compared to periodontitis and the influence of smoking. Front. Microbiol. 2015, 6, 119. [Google Scholar] [CrossRef] [PubMed]
- Inquimbert, C.; Bourgeois, D.; Bravo, M.; Viennot, S.; Tramini, P.; Llodra, J.C.; Molinari, N.; Dussart, C.; Giraudeau, N.; Carrouel, F. The oral bacterial microbiome of interdental surfaces in adolescents according to carious risk. Microorganisms 2019, 7, 319. [Google Scholar] [CrossRef] [PubMed]
- Carrouel, F.; Viennot, S.; Santamaria, J.; Veber, P.; Bourgeois, D. Quantitative molecular detection of 19 major pathogens in the interdental biofilm of periodontally healthy young adults. Front. Microbiol. 2016, 7, 840. [Google Scholar] [CrossRef] [PubMed]
- Asakawa, M.; Takeshita, T.; Furuta, M.; Kageyama, S.; Takeuchi, K.; Hata, J.; Ninomiya, T.; Yamashita, Y. Tongue microbiota and oral health status in community-dwelling elderly adults. mSphere 2018, 3, e00332-18. [Google Scholar] [CrossRef] [PubMed]
- Faran Ali, S.M.; Tanwir, F. Oral microbial habitat a dynamic entity. J. Oral Biol. Craniofac. Res. 2012, 2, 181–187. [Google Scholar] [CrossRef]
- Deo, P.N.; Deshmukh, R. Oral microbiome: Unveiling the fundamentals. J. Oral Maxillofac. Pathol. 2019, 23, 122–128. [Google Scholar] [CrossRef]
- Lim, Y.; Totsika, M.; Morrison, M.; Punyadeera, C. Oral microbiome: A new biomarker reservoir for oral and oropharyngeal cancers. Theranostics 2017, 7, 4313–4321. [Google Scholar] [CrossRef]
- Baliga, S.; Muglikar, S.; Kale, R. Salivary pH: A diagnostic biomarker. J. Indian Soc. Periodontol. 2013, 17, 461–465. [Google Scholar] [CrossRef]
- Dickinson, B.C.; Moffatt, C.E.; Hagerty, D.; Whitmore, S.E.; Brown, T.A.; Graves, D.T.; Lamont, R.J. Interaction of oral bacteria with gingival epithelial cell multilayers. Mol. Oral Microbiol. 2011, 26, 210–220. [Google Scholar] [CrossRef]
- Peters, B.A.; Wu, J.; Hayes, R.B.; Ahn, J. The oral fungal mycobiome: Characteristics and relation to periodontitis in a pilot study. BMC Microbiol. 2017, 17, 157. [Google Scholar] [CrossRef] [PubMed]
- Bachrach, G.; Leizerovici-Zigmond, M.; Zlotkin, A.; Naor, R.; Steinberg, D. Bacteriophage isolation from human saliva. Lett. Appl. Microbiol. 2003, 36, 50–53. [Google Scholar] [CrossRef]
- Ly, M.; Abeles, S.R.; Boehm, T.K.; Robles-Sikisaka, R.; Naidu, M.; Santiago-Rodriguez, T.; Pride, D.T. Altered oral viral ecology in association with periodontal disease. mBio 2014, 5, e01133-14. [Google Scholar] [CrossRef]
- Pride, D.T.; Salzman, J.; Haynes, M.; Rohwer, F.; Davis-Long, C.; White, R.A., 3rd; Loomer, P.; Armitage, G.C.; Relman, D.A. Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J. 2012, 6, 915–926. [Google Scholar] [CrossRef]
- Naidu, M.; Robles-Sikisaka, R.; Abeles, S.R.; Boehm, T.K.; Pride, D.T. Characterization of bacteriophage communities and CRISPR profiles from dental plaque. BMC Microbiol. 2014, 14, 175. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Brocal, V.; Moya, A. The analysis of the oral DNA virome reveals which viruses are widespread and rare among healthy young adults in Valencia (Spain). PLoS ONE 2018, 13, e0191867. [Google Scholar] [CrossRef] [PubMed]
- Edlund, A.; Santiago-Rodriguez, T.M.; Boehm, T.K.; Pride, D.T. Bacteriophage and their potential roles in the human oral cavity. J. Oral Microbiol. 2015, 7, 27423. [Google Scholar] [CrossRef] [PubMed]
- To, K.K.; Tsang, O.T.; Yip, C.C.; Chan, K.H.; Wu, T.C.; Chan, J.M.; Leung, W.S.; Chik, T.S.; Choi, C.Y.; Kandamby, D.H.; et al. Consistent detection of 2019 novel coronavirus in Saliva. Clin. Infect. Dis. 2020, 71, 841–843. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.G.; Yoon, J.; Song, J.Y.; Yoon, S.Y.; Lim, C.S.; Seong, H.; Noh, J.Y.; Cheong, H.J.; Kim, W.J. Clinical significance of a high SARS-CoV-2 viral load in the Saliva. J. Korean Med. Sci. 2020, 35, e195. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Bhatia, S.; Sodhi, A.S.; Batra, N. Oral microbiome and health. AIMS Microbiol. 2018, 4, 42–66. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.C.; Shih, T.P.; Ko, W.C.; Tang, H.J.; Hsueh, P.R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int. J. Antimicrob. Agents 2020, 55, 105924. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Kang, H.; Liu, X.; Tong, Z. Combination of RT-qPCR testing and clinical features for diagnosis of COVID-19 facilitates management of SARS-CoV-2 outbreak. J. Med. Virol. 2020, 92, 538–539. [Google Scholar] [CrossRef]
- Sabino-Silva, R.; Jardim, A.C.G.; Siqueira, W.L. Coronavirus COVID-19 impacts to dentistry and potential salivary diagnosis. Clin. Oral Investig. 2020, 24, 1619–1621. [Google Scholar] [CrossRef]
- Medeiros da Silva, R.C.; Nogueira Marinho, L.C.; de Araújo Silva, D.N.; Costa de Lima, K.; Pirih, F.Q.; Luz de Aquino Martins, A.R. Saliva as a possible tool for the SARS-CoV-2 detection: A review. Travel Med. Infect. Dis. 2020, 38, 101920. [Google Scholar] [CrossRef] [PubMed]
- Zaura, E. Next-generation sequencing approaches to understanding the oral microbiome. Adv. Dent. Res. 2012, 24, 81–85. [Google Scholar] [CrossRef]
- Caselli, E.; Fabbri, C.; D’Accolti, M.; Soffritti, I.; Bassi, C.; Mazzacane, S.; Franchi, M. Defining the oral microbiome by whole-genome sequencing and resistome analysis: The complexity of the healthy picture. BMC Microbiol. 2020, 20, 120. [Google Scholar] [CrossRef] [PubMed]
- Clarridge, J.E., 3rd. Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev. 2004, 17, 840–862. [Google Scholar] [CrossRef] [PubMed]
- Lochman, J.; Zapletalova, M.; Poskerova, H.; Izakovicova Holla, L.; Borilova Linhartova, P. Rapid multiplex real-time PCR method for the detection and quantification of selected cariogenic and periodontal bacteria. Diagnostics 2019, 10, 8. [Google Scholar] [CrossRef] [PubMed]
- Melendez, J.H.; Frankel, Y.M.; An, A.T.; Williams, L.; Price, L.B.; Wang, N.Y.; Lazarus, G.S.; Zenilman, J.M. Real-time PCR assays compared to culture-based approaches for identification of aerobic bacteria in chronic wounds. Clin. Microbiol. Infect. 2010, 16, 1762–1769. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, M.; Umeda, M.; Benno, Y. Molecular analysis of human oral microbiota. J. Periodontal Res. 2005, 40, 277–285. [Google Scholar] [CrossRef]
- Huang, C.; Shi, G. Smoking and microbiome in oral, airway, gut and some systemic diseases. J. Transl. Med. 2019, 17, 225. [Google Scholar] [CrossRef]
- Martinon, P.; Fraticelli, L.; Giboreau, A.; Dussart, C.; Bourgeois, D.; Carrouel, F. Nutrition as a key modifiable factor for periodontitis and main chronic diseases. J. Clin. Med. 2021, 10, 197. [Google Scholar] [CrossRef] [PubMed]
- Lamont, R.J.; Koo, H.; Hajishengallis, G. The oral microbiota: Dynamic communities and host interactions. Nat. Rev. Microbiol. 2018, 16, 745–759. [Google Scholar] [CrossRef] [PubMed]
- Lönnroth, I.; Falsen, E.; Westin, J.; Lindholm, L. Two cases of “heavy chain disease”. Acta Pathol. Microbiol. Scand. Sect. B Microbiol. Immunol. 1971, 79, 443. [Google Scholar]
- Nociti, F.H., Jr.; Casati, M.Z.; Duarte, P.M. Current perspective of the impact of smoking on the progression and treatment of periodontitis. Periodontol. 2000 2015, 67, 187–210. [Google Scholar] [CrossRef]
- Wu, J.; Peters, B.A. Cigarette smoking and the oral microbiome in a large study of American adults. ISME J. 2016, 10, 2435–2446. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.S.; Matthews, C.R.; Joshi, V.; de Jager, M.; Aspiras, M. Tobacco smoking affects bacterial acquisition and colonization in oral biofilms. Infect. Immun. 2011, 79, 4730–4738. [Google Scholar] [CrossRef] [PubMed]
- Saadaoui, M.; Singh, P.; Al Khodor, S. Oral microbiome and pregnancy: A bidirectional relationship. J. Reprod. Immunol. 2021, 145, 103293. [Google Scholar] [CrossRef]
- Tettamanti, L.; Lauritano, D.; Nardone, M.; Gargari, M.; Silvestre-Rangil, J.; Gavoglio, P.; Tagliabue, A. Pregnancy and periodontal disease: Does exist a two-way relationship? ORAL Implantol. 2017, 10, 112–118. [Google Scholar] [CrossRef]
- Barak, S.; Oettinger-Barak, O.; Oettinger, M.; Machtei, E.E.; Peled, M.; Ohel, G. Common oral manifestations during pregnancy: A review. Obstet. Gynecol. Surv. 2003, 58, 624–628. [Google Scholar] [CrossRef]
- Madianos, P.N.; Bobetsis, Y.A.; Offenbacher, S. Adverse pregnancy outcomes (APOs) and periodontal disease: Pathogenic mechanisms. J. Periodontol. 2013, 84, S170–S180. [Google Scholar] [CrossRef] [PubMed]
- León, R.; Silva, N.; Ovalle, A.; Chaparro, A.; Ahumada, A.; Gajardo, M.; Martinez, M.; Gamonal, J. Detection of porphyromonas gingivalis in the amniotic fluid in pregnant women with a diagnosis of threatened premature labor. J. Periodontol. 2007, 78, 1249–1255. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, N.; Tsuruda, K.; Iwamoto, Y.; Kato, F.; Odaki, T.; Yamane, N.; Hori, Y.; Harashima, Y.; Sakoda, A.; Tagaya, A.; et al. Significant increase of oral bacteria in the early pregnancy period in Japanese women. J. Investig. Clin. Dent. 2017, 8, e12189. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Jiang, W.; Hu, X.; Gao, L.; Ai, D.; Pan, H.; Niu, C.; Yuan, K.; Zhou, X.; Xu, C.; et al. Ecological shifts of supragingival microbiota in association with pregnancy. Front. Cell. Infect. Microbiol. 2018, 8, 24. [Google Scholar] [CrossRef] [PubMed]
- Lazarevic, V.; Manzano, S.; Gaïa, N.; Girard, M.; Whiteson, K.; Hibbs, J.; François, P.; Gervaix, A.; Schrenzel, J. Effects of amoxicillin treatment on the salivary microbiota in children with acute otitis media. Clin. Microbiol. Infect. 2013, 19, e335–e342. [Google Scholar] [CrossRef] [PubMed]
- Raju, S.C.; Viljakainen, H.; Figueiredo, R.A.O.; Neuvonen, P.J.; Eriksson, J.G.; Weiderpass, E.; Rounge, T.B. Antimicrobial drug use in the first decade of life influences saliva microbiota diversity and composition. Microbiome 2020, 8, 121. [Google Scholar] [CrossRef]
- Gupta, P.; Gupta, N.; Pawar, A.P.; Birajdar, S.S.; Natt, A.S.; Singh, H.P. Role of sugar and sugar substitutes in dental caries: A review. ISRN Dent. 2013, 2013, 519421. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, N.; Nyvad, B. The role of bacteria in the caries process: Ecological perspectives. J. Dent. Res. 2011, 90, 294–303. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.L.; Leys, E.J.; Gasparovich, S.R.; Firestone, N.D.; Schwartzbaum, J.A.; Janies, D.A.; Asnani, K.; Griffen, A.L. Bacterial 16S sequence analysis of severe caries in young permanent teeth. J. Clin. Microbiol. 2010, 48, 4121–4128. [Google Scholar] [CrossRef]
- Han, S.J.; Bae, K.H.; Lee, H.J.; Kim, S.J.; Cho, H.J. Association between regular walking and periodontitis according to socioeconomic status: A cross-sectional study. Sci. Rep. 2019, 9, 12969. [Google Scholar] [CrossRef]
- Schwendicke, F.; Dörfer, C.E.; Schlattmann, P.; Foster Page, L.; Thomson, W.M.; Paris, S. Socioeconomic inequality and caries: A systematic review and meta-analysis. J. Dent. Res. 2015, 94, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Pihlstrom, B.L.; Michalowicz, B.S.; Johnson, N.W. Periodontal diseases. Lancet 2005, 366, 1809–1820. [Google Scholar] [CrossRef]
- FDI World Dental Federation. FDI policy statement on oral health and the social determinants of health. Adopted by the FDI General Assembly: 30 August 2013—Istanbul, Turkey. Int. Dent. J. 2013, 63, 287–288. [Google Scholar] [CrossRef]
- Kassebaum, N.J.; Smith, A.G.C.; Bernabe, E.; Fleming, T.D.; Reynolds, A.E.; Vos, T.; Murray, C.J.L.; Marcenes, W.; GBD 2015 Oral Health Collaborators. Global, regional, and national prevalence, incidence, and disability-adjusted life years for oral conditions for 195 countries, 1990–2015: A systematic analysis for the global burden of diseases, injuries, and risk factors. J. Dent. Res. 2017, 96, 380–387. [Google Scholar] [CrossRef] [PubMed]
- Tonetti, M.S.; Jepsen, S.; Jin, L.; Otomo-Corgel, J. Impact of the global burden of periodontal diseases on health, nutrition and wellbeing of mankind: A call for global action. J. Clin. Periodontol. 2017, 44, 456–462. [Google Scholar] [CrossRef] [PubMed]
- Loe, H.; Theilade, E.; Jensen, S.B. Experimental gingivitis in man. J. Periodontol. 1965, 36, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Darveau, R.P. Periodontitis: A polymicrobial disruption of host homeostasis. Nat. Rev. Microbiol. 2010, 8, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Liang, S.; Payne, M.A.; Hashim, A.; Jotwani, R.; Eskan, M.A.; McIntosh, M.L.; Alsam, A.; Kirkwood, K.L.; Lambris, J.D.; et al. Low-abundance biofilm species orchestrates inflammatory periodontal disease through the commensal microbiota and complement. Cell Host Microbe 2011, 10, 497–506. [Google Scholar] [CrossRef]
- Socransky, S.S.; Haffajee, A.D.; Cugini, M.A.; Smith, C.; Kent, R.L., Jr. Microbial complexes in subgingival plaque. J. Clin. Periodontol. 1998, 25, 134–144. [Google Scholar] [CrossRef]
- Samaranayake, L.; Matsubara, V.H. Normal oral flora and the oral ecosystem. Dent. Clin. N. Am. 2017, 61, 199–215. [Google Scholar] [CrossRef]
- Simón-Soro, A.; Tomás, I.; Cabrera-Rubio, R.; Catalan, M.D.; Nyvad, B.; Mira, A. Microbial geography of the oral cavity. J. Dent. Res. 2013, 92, 616–621. [Google Scholar] [CrossRef] [PubMed]
- Moore, W.E.; Moore, L.V. The bacteria of periodontal diseases. Periodontology 1994, 5, 66–77. [Google Scholar] [CrossRef] [PubMed]
- Abusleme, L.; Dupuy, A.K.; Dutzan, N.; Silva, N.; Burleson, J.A.; Strausbaugh, L.D.; Gamonal, J.; Diaz, P.I. The subgingival microbiome in health and periodontitis and its relationship with community biomass and inflammation. ISME J. 2013, 7, 1016–1025. [Google Scholar] [CrossRef] [PubMed]
- Curtis, M.A.; Diaz, P.I.; Van Dyke, T.E. The role of the microbiota in periodontal disease. Periodontology 2020, 83, 14–25. [Google Scholar] [CrossRef]
- Kistler, J.O.; Booth, V.; Bradshaw, D.J.; Wade, W.G. Bacterial community development in experimental gingivitis. PLoS ONE 2013, 8, e71227. [Google Scholar] [CrossRef] [PubMed]
- Schincaglia, G.P.; Hong, B.Y.; Rosania, A.; Barasz, J.; Thompson, A.; Sobue, T.; Panagakos, F.; Burleson, J.A.; Dongari-Bagtzoglou, A.; Diaz, P.I. Clinical, immune, and microbiome traits of gingivitis and peri-implant mucositis. J. Dent. Res. 2017, 96, 47–55. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Darveau, R.P.; Curtis, M.A. The keystone-pathogen hypothesis. Nat. Rev. Microbiol. 2012, 10, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Hasegawa, M.; Inohara, N. The role of oral pathobionts in dysbiosis during periodontitis development. J. Dent. Res. 2014, 93, 539–546. [Google Scholar] [CrossRef]
- Selwitz, R.H.; Ismail, A.I.; Pitts, N.B. Dental caries. Lancet 2007, 369, 51–59. [Google Scholar] [CrossRef]
- Featherstone, J.D. The continuum of dental caries—Evidence for a dynamic disease process. J. Dent. Res. 2004, 83 (Suppl. 1), C39–C42. [Google Scholar] [CrossRef]
- Takahashi, N.; Nyvad, B. Caries ecology revisited: Microbial dynamics and the caries process. Caries Res. 2008, 42, 409–418. [Google Scholar] [CrossRef]
- Kidd, E.A.; Fejerskov, O. What constitutes dental caries? Histopathology of carious enamel and dentin related to the action of cariogenic biofilms. J. Dent. Res. 2004, 83 (Suppl. 1), C35–C38. [Google Scholar] [CrossRef] [PubMed]
- Hopcraft, M.S.; Morgan, M.V. Pattern of dental caries experience on tooth surfaces in an adult population. Community Dent. Oral. Epidemiol. 2006, 34, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Almståhl, A.; Carlén, A.; Eliasson, L.; Lingström, P. Lactobacillus species in supragingival plaque in subjects with hyposalivation. Arch. Oral. Biol. 2010, 55, 255–259. [Google Scholar] [CrossRef]
- Simon-Soro, A.; Guillen-Navarro, M.; Mira, A. Metatranscriptomics reveals overall active bacterial composition in caries lesions. J. Oral. Microbiol. 2014, 6, 25443. [Google Scholar] [CrossRef] [PubMed]
- Simon-Soro, A.; Mira, A. Solving the etiology of dental caries. Trends Microbiol. 2015, 23, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Bowen, W.H.; Burne, R.A.; Wu, H.; Koo, H. Oral biofilms: Pathogens, matrix, and polymicrobial interactions in microenvironments. Trends Microbiol. 2018, 26, 229–242. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.L.; Beall, C.J.; Kutsch, S.R.; Firestone, N.D.; Leys, E.J.; Griffen, A.L. Beyond streptococcus mutans: Dental caries onset linked to multiple species by 16S rRNA community analysis. PLoS ONE 2012, 7, e47722. [Google Scholar] [CrossRef] [PubMed]
- Loesche, W.J. Role of streptococcus mutans in human dental decay. Microbiol. Rev. 1986, 50, 353–380. [Google Scholar] [CrossRef] [PubMed]
- Tanzer, J.M.; Livingston, J.; Thompson, A.M. The microbiology of primary dental caries in humans. J. Dent. Educ. 2001, 65, 1028–1037. [Google Scholar] [CrossRef]
- Mantzourani, M.; Fenlon, M.; Beighton, D. Association between bifidobacteriaceae and the clinical severity of root caries lesions. Oral Microbiol. Immunol. 2009, 24, 32–37. [Google Scholar] [CrossRef]
- Badet, C.; Thebaud, N.B. Ecology of lactobacilli in the oral cavity: A review of literature. Open Microbiol. J. 2008, 2, 38–48. [Google Scholar] [CrossRef]
- Tanner, A.C.; Mathney, J.M.; Kent, R.L.; Chalmers, N.I.; Hughes, C.V.; Loo, C.Y.; Pradhan, N.; Kanasi, E.; Hwang, J.; Dahlan, M.A.; et al. Cultivable anaerobic microbiota of severe early childhood caries. J. Clin. Microbiol. 2011, 49, 1464–1474. [Google Scholar] [CrossRef]
- Boorghani, M.; Gholizadeh, N.; Taghavi Zenouz, A.; Vatankhah, M.; Mehdipour, M. Oral lichen planus: Clinical features, etiology, treatment and management; a review of literature. J. Dent. Res. Dent. Clin. Dent. Prospect. 2010, 4, 3–9. [Google Scholar] [CrossRef]
- Bornstein, M.M.; Hakimi, B.; Persson, G.R.J. Microbiological findings in subjects with asymptomatic oral lichen planus: A cross-sectional comparative study. J. Periodontol. 2008, 79, 2347–2355. [Google Scholar] [CrossRef] [PubMed]
- Ertugrul, A.S.; Arslan, U.; Dursun, R.; Hakki, S.S. Periodontopathogen profile of healthy and oral lichen planus patients with gingivitis or periodontitis. Int. J. Oral Sci. 2013, 5, 92–97. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.S.; Kim, Y.; Yoon, H.-J.; Baek, K.J.; Alam, J.; Park, H.K.; Choi, Y. The presence of bacteria within tissue provides insights into the pathogenesis of oral lichen planus. Sci. Rep. 2016, 6, 1–13. [Google Scholar] [CrossRef]
- Metzemaekers, M.; Vanheule, V.; Janssens, R.; Struyf, S.; Proost, P. Overview of the mechanisms that may contribute to the non-redundant activities of interferon-inducible CXC chemokine receptor 3 ligands. Front. Immunol. 2018, 8, 1970. [Google Scholar] [CrossRef] [PubMed]
- Carrozzo, M.; Gandolfo, S.; Carbone, M.; Colombatto, P.; Broccoletti, R.; Garzino-Demo, P.; Ghisetti, V. Hepatitis C virus infection in Italian patients with oral lichen planus: A prospective case-control study. J. Oral Pathol. Med. 1996, 25, 527–533. [Google Scholar] [CrossRef]
- Mokni, M.; Rybojad, M.; Puppin, D., Jr.; Catala, S.; Venezia, F.; Djian, R.; Morel, P. Lichen planus and hepatitis C virus. J. Am. Acad. Dermatol. 1991, 24, 792. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, X.; Li, H.; Ni, C.; Du, Z.; Yan, F. Human oral microbiota and its modulation for oral health. Biomed. Pharmacother. 2018, 99, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Herrero, E.R.; Fernandes, S.; Verspecht, T.; Ugarte-Berzal, E.; Boon, N.; Proost, P.; Bernaerts, K.; Quirynen, M.; Teughels, W. Dysbiotic biofilms deregulate the periodontal inflammatory response. J. Dent. Res. 2018, 97, 547–555. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Roberts, J.S.; Atanasova, K.R.; Chowdhury, N.; Yilmaz, O. A novel kinase function of a nucleoside-diphosphate-kinase homologue in Porphyromonas gingivalis is critical in subversion of host cell apoptosis by targeting heat-shock protein 27. Cell Microbiol. 2018, 20, e12825. [Google Scholar] [CrossRef]
- Yilmaz, O.; Yao, L.; Maeda, K.; Rose, T.M.; Lewis, E.L.; Duman, M.; Lamont, R.J.; Ojcius, D.M. ATP scavenging by the intracellular pathogen Porphyromonas gingivalis inhibits P2X7-mediated host-cell apoptosis. Cell Microbiol. 2008, 10, 863–875. [Google Scholar] [CrossRef] [PubMed]
- Meurman, J.H.; Uittamo, J. Oral micro-organisms in the etiology of cancer. Acta Odontol. Scand. 2008, 66, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Marttila, E.; Bowyer, P.; Sanglard, D.; Uittamo, J.; Kaihovaara, P.; Salaspuro, M.; Richardson, M.; Rautemaa, R. Fermentative 2-carbon metabolism produces carcinogenic levels of acetaldehyde in Candida albicans. Mol. Oral Microbiol. 2013, 28, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Attene-Ramos, M.S.; Wagner, E.D.; Plewa, M.J.; Gaskins, H.R. Evidence that hydrogen sulfide is a genotoxic agent. Mol. Cancer Res. 2006, 4, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Yoneda, M.; Takeshita, T.; Hirofuji, T.; Hanioka, T. Induction and inhibition of oral malodor. Mol. Oral Microbiol. 2019, 34, 85–96. [Google Scholar] [CrossRef]
- Nakamura, S.; Shioya, K.; Hiraoka, B.Y.; Suzuki, N.; Hoshino, T.; Fujiwara, T.; Yoshinari, N.; Ansai, T.; Yoshida, A. Porphyromonas gingivalis hydrogen sulfide enhances methyl mercaptan-induced pathogenicity in mouse abscess formation. Microbiology 2018, 164, 529–539. [Google Scholar] [CrossRef]
- Naucler, C.S.; Geisler, J.; Vetvik, K. The emerging role of human cytomegalovirus infection in human carcinogenesis: A review of current evidence and potential therapeutic implications. Oncotarget 2019, 10, 4333–4347. [Google Scholar] [CrossRef]
- Foglio-Bonda, P.L.; Gabriele, M.; Graziani, F.; De Andrea, M.; Mondini, M.; Gariglio, M. High prevalence of human cytomegalovirus in a population of periodontally healthy subjects. Med. Oral Patol. Oral Cir. Bucal. 2010, 15, e292–e296. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Saboia-Dantas, C.J.; Coutrin de Toledo, L.F.; Sampaio-Filho, H.R.; Siqueira, J.F., Jr. Herpesviruses in asymptomatic apical periodontitis lesions: An immunohistochemical approach. Oral Microbiol. Immunol. 2007, 22, 320–325. [Google Scholar] [CrossRef] [PubMed]
- Sunde, P.T.; Olsen, I.; Enersen, M.; Beiske, K.; Grinde, B. Human cytomegalovirus and Epstein-Barr virus in apical and marginal periodontitis: A role in pathology? J. Med. Virol. 2008, 80, 1007–1011. [Google Scholar] [CrossRef] [PubMed]
- Nagy, K.N.; Sonkodi, I.; Szoke, I.; Nagy, E.; Newman, H.N. The microflora associated with human oral carcinomas. Oral Oncol. 1998, 34, 304–308. [Google Scholar] [CrossRef]
- Rosenbaum, J.T.; Asquith, M.J. The microbiome: A revolution in treatment for rheumatic diseases? Curr. Rheumatol. Rep. 2016, 18, 62. [Google Scholar] [CrossRef] [PubMed]
- Bawadekar, M.; De Andrea, M.; Gariglio, M.; Landolfo, S. Mislocalization of the interferon inducible protein IFI16 by environmental insults: Implications in autoimmunity. Cytokine Growth Factor Rev. 2015, 26, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, K.; Fei, G.; Lundmark, A.; Benchimol, D.; Lee, L.; Hu, Y.O.; Kats, A.; Saevarsdottir, S.; Catrina, A.I.; Klinge, B. Periodontal health and oral microbiota in patients with rheumatoid arthritis. J. Clin. Med. 2019, 8, 630. [Google Scholar] [CrossRef]
- Arvikar, S.L.; Collier, D.S.; Fisher, M.C.; Unizony, S.; Cohen, G.L.; McHugh, G.; Kawai, T.; Strle, K.; Steere, A.C. Clinical correlations with Porphyromonas gingivalis antibody responses in patients with early rheumatoid arthritis. Arthritis Res. Ther. 2013, 15, R109. [Google Scholar] [CrossRef]
- Hitchon, C.A.; Chandad, F.; Ferucci, E.D.; Willemze, A.; Ioan-Facsinay, A.; van der Woude, D.; Markland, J.; Robinson, D.; Elias, B.; Newkirk, M. Antibodies to porphyromonas gingivalis are associated with anticitrullinated protein antibodies in patients with rheumatoid arthritis and their relatives. J. Rheumatol. 2010, 37, 1105–1112. [Google Scholar] [CrossRef] [PubMed]
- Mikuls, T.R.; Payne, J.B.; Reinhardt, R.A.; Thiele, G.M.; Maziarz, E.; Cannella, A.C.; Holers, V.M.; Kuhn, K.A.; O’Dell, J.R. Antibody responses to porphyromonas gingivalis (P. gingivalis) in subjects with rheumatoid arthritis and periodontitis. Int. Immunopharmacol. 2009, 9, 38–42. [Google Scholar] [CrossRef]
- Cheng, Z.; Meade, J.; Mankia, K.; Emery, P.; Devine, D.A. Periodontal disease and periodontal bacteria as triggers for rheumatoid arthritis. Best Pract. Res. Clin. Rheumatol. 2017, 31, 19–30. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Tian, C.; Postlethwaite, A.; Jiao, Y.; Garcia-Godoy, F.; Pattanaik, D.; Wei, D.; Gu, W.; Li, J. Rheumatoid arthritis and periodontal disease: What are the similarities and differences? Int. J. Rheum. Dis. 2017, 20, 1887–1901. [Google Scholar] [CrossRef] [PubMed]
- Fuggle, N.R.; Smith, T.O.; Kaul, A.; Sofat, N. Hand to mouth: A systematic review and meta-analysis of the association between rheumatoid arthritis and periodontitis. Front. Immunol. 2016, 7, 80. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zou, Y.; Jiang, Q.; Jiang, L.; Yu, Q.; Ding, X.; Yu, Y. A preliminary study of the oral microbiota in Chinese patients with Sjögren’s syndrome. Arch. Oral Biol. 2016, 70, 143–148. [Google Scholar] [CrossRef] [PubMed]
- Rusthen, S.; Kristoffersen, A.K.; Young, A.; Galtung, H.; Petrovski, B.; Palm, Ø.; Enersen, M.; Jensen, J.L. Dysbiotic salivary microbiota in dry mouth and primary Sjögren’s syndrome patients. PLoS ONE 2019, 14, e0218319. [Google Scholar] [CrossRef]
- Sharma, D.; Sandhya, P.; Vellarikkal, S.K.; Surin, A.K.; Jayarajan, R.; Verma, A.; Kumar, A.; Ravi, R.; Danda, D.; Sivasubbu, S. Saliva microbiome in primary Sjögren’s syndrome reveals distinct set of disease-associated microbes. Oral Dis. 2020, 26, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, H.; Chen, T.; Aliko, A.; Mydel, P.M.; Jonsson, R.; Olsen, I. Microbiological and bioinformatics analysis of primary Sjögren’s syndrome patients with normal salivation§. J. Oral Microbiol. 2016, 8, 31119. [Google Scholar] [CrossRef]
- Van der Meulen, T.A.; Harmsen, H.J.; Bootsma, H.; Liefers, S.C.; Vich Vila, A.; Zhernakova, A.; Fu, J.; Wijmenga, C.; Spijkervet, F.K.; Kroese, F.G. Dysbiosis of the buccal mucosa microbiome in primary Sjögren’s syndrome patients. Rheumatology 2018, 57, 2225–2234. [Google Scholar] [CrossRef]
- Van der Meulen, T.A.; Harmsen, H.J.; Bootsma, H.; Liefers, S.C.; Vila, A.V.; Zhernakova, A.; Weersma, R.K.; Spijkervet, F.K.; Kroese, F.G.; Vissink, A. Reduced salivary secretion contributes more to changes in the oral microbiome of patients with primary Sjögren’s syndrome than underlying disease. Ann. Rheum. Dis. 2018, 77, 1542–1544. [Google Scholar] [CrossRef]
- De Paiva, C.S.; Jones, D.B.; Stern, M.E.; Bian, F.; Moore, Q.L.; Corbiere, S.; Streckfus, C.F.; Hutchinson, D.S.; Ajami, N.J.; Petrosino, J.F. Altered mucosal microbiome diversity and disease severity in Sjögren syndrome. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- Corrêa, J.D.; Calderaro, D.C.; Ferreira, G.A.; Mendonça, S.M.S.; Fernandes, G.R.; Xiao, E.; Teixeira, A.L.; Leys, E.J.; Graves, D.T.; Silva, T.A. Subgingival microbiota dysbiosis in systemic lupus erythematosus: Association with periodontal status. Microbiome 2017, 5, 34. [Google Scholar] [CrossRef]
- Fabbri, C.; Fuller, R.; Bonfá, E.; Guedes, L.K.; D’Alleva, P.S.R.; Borba, E.F. Periodontitis treatment improves systemic lupus erythematosus response to immunosuppressive therapy. Clin. Rheumatol. 2014, 33, 505–509. [Google Scholar] [CrossRef]
- Kobayashi, T.; Ito, S.; Yasuda, K.; Kuroda, T.; Yamamoto, K.; Sugita, N.; Tai, H.; Narita, I.; Gejyo, F.; Yoshie, H. The combined genotypes of stimulatory and inhibitory Fcγ receptors associated with systemic lupus erythematosus and periodontitis in Japanese adults. J. Periodontol. 2007, 78, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.; Sinha, R.; Pei, Z.; Dominianni, C.; Wu, J.; Shi, J.; Goedert, J.J.; Hayes, R.B.; Yang, L. Human gut microbiome and risk for colorectal cancer. J. Natl. Cancer Inst. 2013, 105, 1907–1911. [Google Scholar] [CrossRef] [PubMed]
- Castellarin, M.; Warren, R.L.; Freeman, J.D.; Dreolini, L.; Krzywinski, M.; Strauss, J.; Barnes, R.; Watson, P.; Allen-Vercoe, E.; Moore, R.A. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012, 22, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Kostic, A.D.; Gevers, D.; Pedamallu, C.S.; Michaud, M.; Duke, F.; Earl, A.M.; Ojesina, A.I.; Jung, J.; Bass, A.J.; Tabernero, J. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 2012, 22, 292–298. [Google Scholar] [CrossRef]
- Feng, Q.; Liang, S.; Jia, H.; Stadlmayr, A.; Tang, L.; Lan, Z.; Zhang, D.; Xia, H.; Xu, X.; Jie, Z. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 2015, 6, 6528. [Google Scholar] [CrossRef] [PubMed]
- Zackular, J.P.; Rogers, M.A.; Ruffin, M.T.; Schloss, P.D. The human gut microbiome as a screening tool for colorectal cancer. Cancer Prev. Res. 2014, 7, 1112–1121. [Google Scholar] [CrossRef] [PubMed]
- Kato, I.; Vasquez, A.A.; Moyerbrailean, G.; Land, S.; Sun, J.; Lin, H.-S.; Ram, J.L. Oral microbiome and history of smoking and colorectal cancer. J. Epidemiol. Res. 2016, 2, 92. [Google Scholar] [CrossRef]
- Farrell, J.J.; Zhang, L.; Zhou, H.; Chia, D.; Elashoff, D.; Akin, D.; Paster, B.J.; Joshipura, K.; Wong, D.T. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 2012, 61, 582–588. [Google Scholar] [CrossRef]
- Michaud, D.S.; Izard, J.; Wilhelm-Benartzi, C.S.; You, D.-H.; Grote, V.A.; Tjønneland, A.; Dahm, C.C.; Overvad, K.; Jenab, M.; Fedirko, V. Plasma antibodies to oral bacteria and risk of pancreatic cancer in a large European prospective cohort study. Gut 2013, 62, 1764–1770. [Google Scholar] [CrossRef]
- Zambirinis, C.P.; Levie, E.; Nguy, S.; Avanzi, A.; Barilla, R.; Xu, Y.; Seifert, L.; Daley, D.; Greco, S.H.; Deutsch, M. TLR9 ligation in pancreatic stellate cells promotes tumorigenesis. J. Exp. Med. 2015, 212, 2077–2094. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Alekseyenko, A.V.; Wu, J.; Peters, B.A.; Jacobs, E.J.; Gapstur, S.M.; Purdue, M.P.; Abnet, C.C.; Stolzenberg-Solomon, R.; Miller, G. Human oral microbiome and prospective risk for pancreatic cancer: A population-based nested case-control study. Gut 2018, 67, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Peters, B.A.; Wu, J.; Pei, Z.; Yang, L.; Purdue, M.P.; Freedman, N.D.; Jacobs, E.J.; Gapstur, S.M.; Hayes, R.B.; Ahn, J. Oral microbiome composition reflects prospective risk for esophageal cancers. Cancer Res. 2017, 77, 6777–6787. [Google Scholar] [CrossRef] [PubMed]
- Michaud, D.S.; Liu, Y.; Meyer, M.; Giovannucci, E.; Joshipura, K. Periodontal disease, tooth loss, and cancer risk in male health professionals: A prospective cohort study. Lancet Oncol. 2008, 9, 550–558. [Google Scholar] [CrossRef]
- Vogtmann, E.; Goedert, J.J. Epidemiologic studies of the human microbiome and cancer. Br. J. Cancer 2016, 114, 237–242. [Google Scholar] [CrossRef]
- Zeng, X.T.; Xia, L.Y.; Zhang, Y.G.; Li, S.; Leng, W.D.; Kwong, J.S. Periodontal disease and incident lung cancer risk: A meta-analysis of cohort studies. J. Periodontol. 2016, 87, 1158–1164. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Yang, M.; Liu, J.; Gao, R.; Hu, J.; Li, J.; Zhang, L.; Shi, Y.; Guo, H.; Cheng, J. Discovery and validation of potential bacterial biomarkers for lung cancer. Am. J. Cancer Res. 2015, 5, 3111. [Google Scholar]
- Ganu, R.S.; Ma, J.; Aagaard, K.M. The role of microbial communities in parturition: Is there evidence of association with preterm birth and perinatal morbidity and mortality? Am. J. Perinatol. 2013, 30, 613–624. [Google Scholar] [CrossRef] [PubMed]
- Mendz, G.L.; Kaakoush, N.O.; Quinlivan, J.A. Bacterial aetiological agents of intra-amniotic infections and preterm birth in pregnant women. Front. Cell. Infect. Microbiol. 2013, 3, 58. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.W.; Fardini, Y.; Chen, C.; Iacampo, K.G.; Peraino, V.A.; Shamonki, J.M.; Redline, R.W. Term stillbirth caused by oral Fusobacterium nucleatum. Obstet. Gynecol. 2010, 115, 442. [Google Scholar] [CrossRef]
- Katz, J.; Chegini, N.; Shiverick, K.; Lamont, R. Localization of P. gingivalis in preterm delivery placenta. J. Dent. Res. 2009, 88, 575–578. [Google Scholar] [CrossRef] [PubMed]
- Chapple, I.L.; Genco, R.; on behalf of working group 2 of the joint EFP/AAP workshop. Diabetes and periodontal diseases: Consensus report of the Joint EFP/AAP Workshop on Periodontitis and Systemic Diseases. J. Periodontol. 2013, 84, S106–S112. [Google Scholar] [CrossRef]
- Da Cruz, G.A.; de Toledo, S.; Sallum, E.A.; Sallum, A.W.; Ambrosano, G.M.; de Cassia Orlandi Sardi, J.; da Cruz, S.E.; Goncalves, R.B. Clinical and laboratory evaluations of non-surgical periodontal treatment in subjects with diabetes mellitus. J. Periodontol. 2008, 79, 1150–1157. [Google Scholar] [CrossRef]
- Ganesan, S.M.; Joshi, V.; Fellows, M.; Dabdoub, S.M.; Nagaraja, H.N.; O’Donnell, B.; Deshpande, N.R.; Kumar, P.S. A tale of two risks: Smoking, diabetes and the subgingival microbiome. ISME J. 2017, 11, 2075–2089. [Google Scholar] [CrossRef] [PubMed]
- Casarin, R.; Barbagallo, A.; Meulman, T.; Santos, V.; Sallum, E.; Nociti, F.; Duarte, P.; Casati, M.; Gonçalves, R. Subgingival biodiversity in subjects with uncontrolled type-2 diabetes and chronic periodontitis. J. Periodontal. Res. 2013, 48, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Spahr, A.; Klein, E.; Khuseyinova, N.; Boeckh, C.; Muche, R.; Kunze, M.; Rothenbacher, D.; Pezeshki, G.; Hoffmeister, A.; Koenig, W. Periodontal infections and coronary heart disease: Role of periodontal bacteria and importance of total pathogen burden in the Coronary Event and Periodontal Disease (CORODONT) study. Arch. Intern. Med. 2006, 166, 554–559. [Google Scholar] [CrossRef] [PubMed]
- Renvert, S.; Pettersson, T.; Ohlsson, O.; Persson, G.R. Bacterial profile and burden of periodontal infection in subjects with a diagnosis of acute coronary syndrome. J. Periodontol. 2006, 77, 1110–1119. [Google Scholar] [CrossRef]
- Andriankaja, O.; Trevisan, M.; Falkner, K.; Dorn, J.; Hovey, K.; Sarikonda, S.; Mendoza, T.; Genco, R. Association between periodontal pathogens and risk of nonfatal myocardial infarction. Community Dent. Oral Epidemiol. 2011, 39, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Fåk, F.; Tremaroli, V.; Bergström, G.; Bäckhed, F. Oral microbiota in patients with atherosclerosis. Atherosclerosis 2015, 243, 573–578. [Google Scholar] [CrossRef] [PubMed]
- Martini, A.M.; Moricz, B.S.; Ripperger, A.K.; Tran, P.M.; Sharp, M.E.; Forsythe, A.N.; Kulhankova, K.; Salgado-Pabón, W.; Jones, B.D. Association of novel streptococcus sanguinis virulence factors with pathogenesis in a native valve infective endocarditis model. Front. Microbiol. 2020, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Macleod, L.C.; Kitten, T.; Xu, P. Streptococcus sanguinis biofilm formation & interaction with oral pathogens. Future Microbiol. 2018, 13, 915–932. [Google Scholar] [CrossRef]
- Kallio, E. Lipopolysaccharide: A Link between Periodontitis and Cardiometabolic Disorders; University of Helsinki: Helsinki, Finland, 2014. [Google Scholar]
- Pussinen, P.J.; Tuomisto, K.; Jousilahti, P.; Havulinna, A.S.; Sundvall, J.; Salomaa, V. Endotoxemia, immune response to periodontal pathogens, and systemic inflammation associate with incident cardiovascular disease events. Arterioscler. Thromb. Vasc. Biol. 2007, 27, 1433–1439. [Google Scholar] [CrossRef]
- Haraguchi, A.; Miura, M.; Fujise, O.; Hamachi, T.; Nishimura, F. Porphyromonas gingivalis gingipain is involved in the detachment and aggregation of Aggregatibacter actinomycetemcomitans biofilm. Mol. Oral Microbiol. 2014, 29, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Jayaprakash, K.; Khalaf, H.; Bengtsson, T. Gingipains from porphyromonas gingivalis play a significant role in induction and regulation of CXCL8 in THP-1 cells. BMC Microbiol. 2014, 14, 193. [Google Scholar] [CrossRef]
- Dominy, S.S.; Lynch, C.; Ermini, F.; Benedyk, M.; Marczyk, A.; Konradi, A.; Nguyen, M.; Haditsch, U.; Raha, D.; Griffin, C. Porphyromonas gingivalis in Alzheimer’s disease brains: Evidence for disease causation and treatment with small-molecule inhibitors. Sci. Adv. 2019, 5, eaau3333. [Google Scholar] [CrossRef] [PubMed]
- Mihaila, D.; Donegan, J.; Barns, S.; LaRocca, D.; Du, Q.; Zheng, D.; Vidal, M.; Neville, C.; Uhlig, R.; Middleton, F.A. The oral microbiome of early stage Parkinson’s disease and its relationship with functional measures of motor and non-motor function. PLoS ONE 2019, 14, e0218252. [Google Scholar] [CrossRef] [PubMed]
- Miklossy, J. Bacterial amyloid and DNA are important constituents of senile plaques: Further evidence of the spirochetal and biofilm nature of senile plaques. J. Alzheimer Dis. 2016, 53, 1459–1473. [Google Scholar] [CrossRef] [PubMed]
| Factors | Samples | Major Finding |
|---|---|---|
| Smoking | Subgingival plaque | A high taxonomic diversity and richness in smokers |
| Higher abundances of anaerobes in smokers: Fusobacterium nucleatum, F. naviforme, Filifactor alocis, Dialister microaerophilus, Desulfobulbus sp. clone R004, Megasphaera sueciensis, M. geminatus, M. elsdenii, M. micronuciformis, Acinetobacter johnsonii, A. guillouiae, A. schindleri, A. baumannii, A. haemolyticus, Pseudomonas pseudoalcaligenes, and Pseudoramibacter alactolyticus | ||
| Saliva | Lower abundances of commensal microbes in smokers: Streptococcus sanguinis, S. parasanguinis, S. oralis, Granulicatella elegans, G. adiacens, Actinomyces viscosus, A. israelii, A. dentalis, Neisseria subflava and Hemophilus parainfluenzae | |
| Buccal mucosa | A lower taxonomic diversity in smokers | |
| Marginal and subgingival plaque and gingival crevicular fluid | High diversity in smokers Relatively unstable initial colonization of marginal and subgingival biofilms in smokers Periodontal pathogens belonging to the genera Fusobacterium, Cardiobacterium, Synergistes, and Selenomonas, as well as respiratory pathogens belonging to the genera Haemophilus and Pseudomonas, colonized the early biofilms of smokers and continued to persist over the observation period | |
| Oral wash samples | Lower abundance of the Proteobacteria phylum and genera belong to the Proteobacteria phylum in smokers Taxa not belonging to Proteobacteria: - increase: the Atopobium and Streptococcus genera in current smokers - decrease: the Capnocytophaga, Peptostreptococcus and Leptotrichia genera in current smokers | |
| Saliva | Increase: Streptococcus sobrinus and Eubacterium brachy in smokers | |
| Antimicrobial agent | Saliva | Reduced species richness and diversity At the phylum level: - increase: Proteobacteria at post-treatment - decrease: Actinobacteria at the end of treatment |
| Antibiotics | Saliva | A linear inverse association between the use of azithromycin and Shannon index Amoxicillin and amoxicillin-clavulanate use was associated with the largest decrease in abundance of Rikenellaceae Phenoxymethylpenicillin were associated with a decrease in Paludibacter |
| Pregnancy | Supragingival plaques | Significantly higher Shannon diversity in pregnant women More abundant: Neisseria, Porphyromonas, and Treponema in pregnant women Less abundant: Streptococcus and Veillonella in pregnant women |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, Y.-H.; Chung, S.W.; Auh, Q.-S.; Hong, S.-J.; Lee, Y.-A.; Jung, J.; Lee, G.-J.; Park, H.J.; Shin, S.-I.; Hong, J.-Y. Progress in Oral Microbiome Related to Oral and Systemic Diseases: An Update. Diagnostics 2021, 11, 1283. https://doi.org/10.3390/diagnostics11071283
Lee Y-H, Chung SW, Auh Q-S, Hong S-J, Lee Y-A, Jung J, Lee G-J, Park HJ, Shin S-I, Hong J-Y. Progress in Oral Microbiome Related to Oral and Systemic Diseases: An Update. Diagnostics. 2021; 11(7):1283. https://doi.org/10.3390/diagnostics11071283
Chicago/Turabian StyleLee, Yeon-Hee, Sang Wan Chung, Q-Schick Auh, Seung-Jae Hong, Yeon-Ah Lee, Junho Jung, Gi-Ja Lee, Hae Jeong Park, Seung-Il Shin, and Ji-Youn Hong. 2021. "Progress in Oral Microbiome Related to Oral and Systemic Diseases: An Update" Diagnostics 11, no. 7: 1283. https://doi.org/10.3390/diagnostics11071283
APA StyleLee, Y.-H., Chung, S. W., Auh, Q.-S., Hong, S.-J., Lee, Y.-A., Jung, J., Lee, G.-J., Park, H. J., Shin, S.-I., & Hong, J.-Y. (2021). Progress in Oral Microbiome Related to Oral and Systemic Diseases: An Update. Diagnostics, 11(7), 1283. https://doi.org/10.3390/diagnostics11071283