Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis
Abstract
1. Introduction
2. Materials and Methods
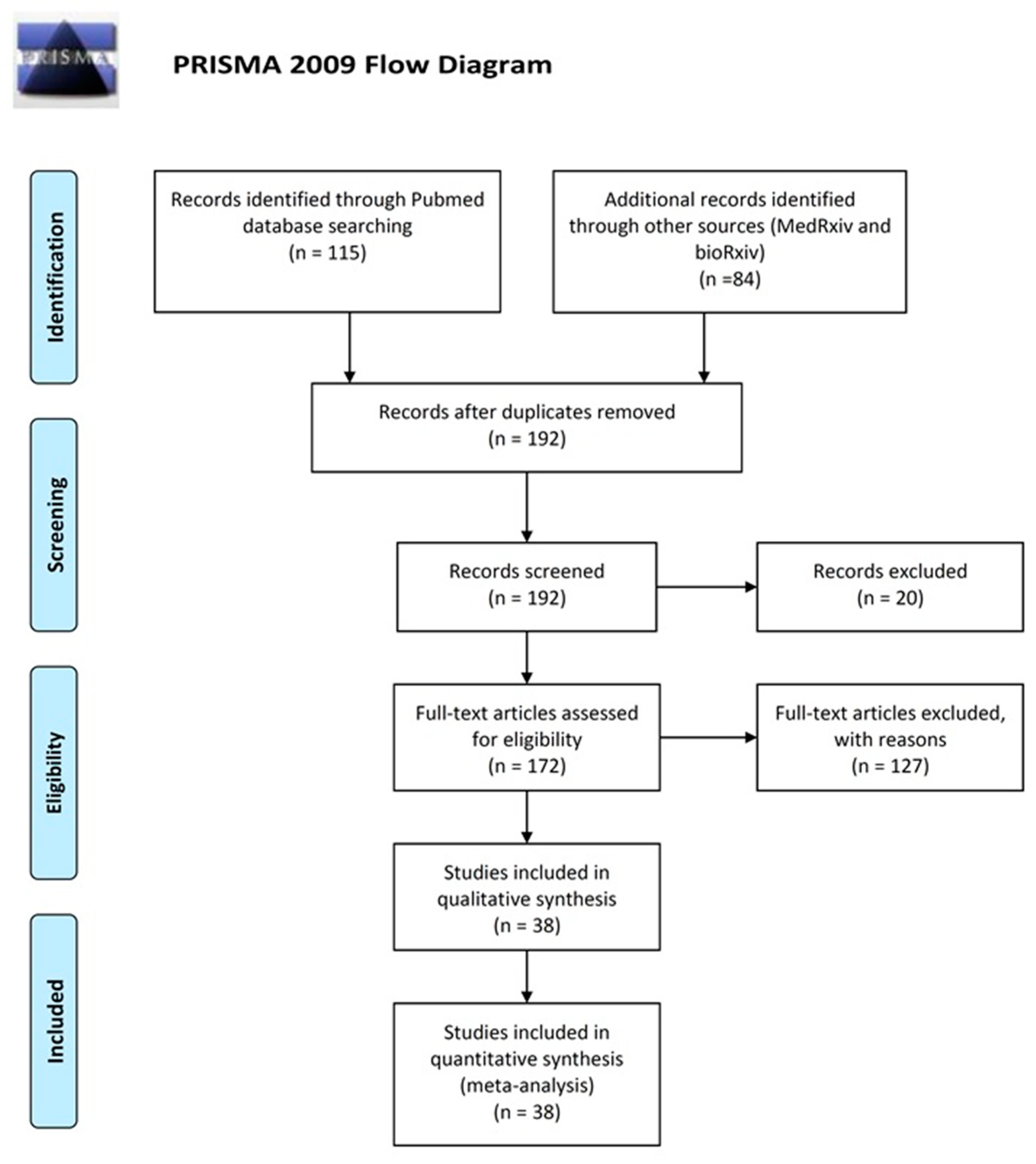
2.1. Search Strategy and Selection Criteria
2.2. Data Analysis
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Disclaimer
References
- Chan, J.F.-W.; Yuan, S.; Kok, K.-H.; To, K.K.-W.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.-Y.; Poon, R.W.-S.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.M.; Wang, W.; Song, Z.G.; Hu, Y.; Tao, Z.W.; Tian, J.H.; Pei, Y.Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef]
- Sohrabi, C.; Alsafi, Z.; O’Neill, N.; Khan, M.; Kerwan, A.; Al-Jabir, A.; Iosifidis, C.; Agha, R. World Health Organization declares global emergency: A review of the 2019 novel coronavirus (COVID-19). Int. J. Surg. 2020, 76, 71–76. [Google Scholar] [CrossRef]
- Liu, J.; Zheng, X.; Tong, Q.; Li, W.; Wang, B.; Sutter, K.; Trilling, M.; Lu, M.; Dittmer, U.; Yang, D. Overlapping and discrete aspects of the pathology and pathogenesis of the emerging human pathogenic coronaviruses SARS-CoV, MERS-CoV, and 2019-nCoV. J. Med. Virol. 2020, 92, 491–494. [Google Scholar] [CrossRef]
- Promislow, D.E.L. A geroscience perspective on COVID-19 mortality. J. Gerontol. Ser. A Biol. Sci. Med. Sci. 2020. [Google Scholar] [CrossRef]
- Wang, B.; Li, R.; Lu, Z.; Huang, Y. Does comorbidity increase the risk of patients with COVID-19: Evidence from meta-analysis. Aging 2020, 12, 6049–6057. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, X.; Jia, X.; Li, J.; Hu, K.; Chen, G.; Wei, J.; Gong, Z.; Zhou, C.; Yu, H.; et al. Risk factors for disease severity, unimprovement, and mortality of COVID-19 patients in Wuhan, China. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Wong, B.H.; Tsoi, H.W.; Fung, A.M.; Kao, R.Y.; Chan, K.H.; Peiris, J.S.; Yuen, K.Y. Differential sensitivities of severe acute respiratory syndrome (SARS) coronavirus spike polypeptide enzyme-linked immunosorbent assay (ELISA) and SARS coronavirus nucleocapsid protein ELISA for serodiagnosis of SARS coronavirus pneumonia. J. Clin. Microbiol. 2005, 43, 3054–3058. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.K.; Sue, S.C.; Yu, T.H.; Hsieh, C.M.; Tsai, C.K.; Chiang, Y.C.; Lee, S.J.; Hsiao, H.H.; Wu, W.J.; Chang, W.L.; et al. Modular organization of SARS coronavirus nucleocapsid protein. J. Biomed. Sci. 2006, 13, 59–72. [Google Scholar] [CrossRef] [PubMed]
- Hurst, K.R.; Koetzner, C.A.; Masters, P.S. Identification of in vivo-interacting domains of the murine coronavirus nucleocapsid protein. J. Virol. 2009, 83, 7221–7234. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. The Laboratory Diagnosis of COVID-19 Infection: Current Issues and Challenges. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e286. [Google Scholar] [CrossRef]
- Zheng, M.; Song, L. Novel antibody epitopes dominate the antigenicity of spike glycoprotein in SARS-CoV-2 compared to SARS-CoV. Cell Mol. Immunol. 2020. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brunink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eur. Surveill. Bull. Eur. Sur Les Mal. Transm. Eur. Commun. Dis. Bull. 2020, 25. [Google Scholar] [CrossRef]
- ECDC. Novel Coronavirus Disease 2019 (COVID-19) Pandemic: Increased Transmission in the EU/EEA and the UK—Sixth Update—12 March 2020; European Centre for Disease Prevention and Control: Stockholm, Sweden, 2020. [Google Scholar]
- Service, R.F. The standard coronavirus test, if available, works well—But can new diagnostics help in this pandemic? ScienceMag 2020. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. New Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Pang, J.; Wang, M.X.; Ang, I.Y.H.; Tan, S.H.X.; Lewis, R.F.; Chen, J.I.; Gutierrez, R.A.; Gwee, S.X.W.; Chua, P.E.Y.; Yang, Q.; et al. Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review. J. Clin. Med. 2020, 9, 623. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef]
- Forero, D.A.; Lopez-Leon, S.; González-Giraldo, Y.; Bagos, P.G. Ten simple rules for carrying out and writing meta-analyses. PLoS Comput. Biol. 2019, 15, e1006922. [Google Scholar] [CrossRef] [PubMed]
- Du, Z.; Zhu, F.; Guo, F.; Yang, B.; Wang, T. Detection of antibodies against SARS-CoV-2 in patients with COVID-19. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Yi, Y.; Luo, X.; Xiong, N.; Liu, Y.; Li, S.; Sun, R.; Wang, Y.; Hu, B.; Chen, W.; et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. 2020. [Google Scholar] [CrossRef]
- Liu, R.; Liu, X.; Han, H.; Shereen, M.A.; Niu, Z.; Li, D.; Liu, F.; Wu, K.; Luo, Z.; Zhu, C. The comparative superiority of IgM-IgG antibody test to real-time reverse transcriptase PCR detection for SARS-CoV-2 infection diagnosis. medRxiv 2020. [Google Scholar] [CrossRef]
- Xiang, J.; Yan, M.; Li, H.; Liu, T.; Lin, C.; Huang, S.; Shen, C. Evaluation of Enzyme-Linked Immunoassay and Colloidal Gold—Immunochromatographic Assay Kit for Detection of Novel Coronavirus (SARS-Cov-2) Causing an Outbreak of Pneumonia (COVID-19). medRxiv 2020. [Google Scholar] [CrossRef]
- Xiao, D.A.T.; Gao, D.C.; Zhang, D.S. Profile of Specific Antibodies to SARS-CoV-2: The First Report. J. Infect. 2020. [Google Scholar] [CrossRef]
- Lassaunière, R.; Frische, A.; Harboe, Z.B.; Nielsen, A.C.Y.; Fomsgaard, A.; Krogfelt, K.A.; Jørgensen, C.S. Evaluation of nine commercial SARS-CoV-2 immunoassays. medRxiv 2020. [Google Scholar] [CrossRef]
- Lou, B.; Li, T.; Zheng, S.; Su, Y.; Li, Z.; Liu, W.; Yu, F.; Ge, S.; Zou, Q.; Yuan, Q.; et al. Serology characteristics of SARS-CoV-2 infection since the exposure and post symptoms onset. medRxiv 2020. [Google Scholar] [CrossRef]
- Zhang, B.; Zhou, X.; Zhu, C.; Feng, F.; Qiu, Y.; Feng, J.; Jia, Q.; Song, Q.; Zhu, B.; Wang, J. Immune phenotyping based on neutrophil-to-lymphocyte ratio and IgG predicts disease severity and outcome for patients with COVID-19. medRxiv 2020. [Google Scholar] [CrossRef]
- Xu, Y. Dynamic profile of severe or critical COVID-19 cases. medRxiv 2020. [Google Scholar] [CrossRef]
- Wang, Z.; Li, H.; Li, J.; Yang, C.; Guo, X.; Hu, Z.; Chen, Z.; Wang, S.; Liu, J. Elevated serum IgM levels indicate poor outcome in patients with coronavirus disease 2019 pneumonia: A retrospective case-control study. medRxiv 2020. [Google Scholar] [CrossRef]
- Lin, D.; Liu, L.; Zhang, M.; Hu, Y.; Yang, Q.; Guo, J.; Dai, Y.; Xu, Y.; Cai, Y.; Chen, X.; et al. Evaluations of serological test in the diagnosis of 2019 novel coronavirus (SARS-CoV-2) infections during the COVID-19 outbreak. medRxiv 2020. [Google Scholar] [CrossRef]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020. [Google Scholar] [CrossRef]
- Hu, X.; An, T.; Situ, B.; Hu, Y.; Ou, Z.; Li, Q.; He, X.; Zhang, Y.; Tian, P.; Sun, D.; et al. Heat inactivation of serum interferes with the immunoanalysis of antibodies to SARS-CoV-2. medRxiv 2020. [Google Scholar] [CrossRef]
- Jia, X.; Zhang, P.; Tian, Y.; Wang, J.; Zeng, H.; Wang, J.; Jiao, L.; Chen, Z.; Zhang, L.; He, H.; et al. Clinical significance of IgM and IgG test for diagnosis of highly suspected COVID-19 infection. medRxiv 2020. [Google Scholar] [CrossRef]
- Van Houwelingen, H.C.; Zwinderman, K.H.; Stijnen, T. A bivariate approach to meta-analysis. Stat. Med. 1993, 12, 2273–2284. [Google Scholar] [CrossRef]
- Arends, L.R.; Hamza, T.H.; van Houwelingen, J.C.; Heijenbrok-Kal, M.H.; Hunink, M.G.; Stijnen, T. Bivariate random effects meta-analysis of ROC curves. Med. Decis. Mak. Int. J. Soc. Med. Decis. Mak. 2008, 28, 621–638. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, Y.; Chu, H. A unification of models for meta-analysis of diagnostic accuracy studies without a gold standard. Biometrics 2015, 71, 538–547. [Google Scholar] [CrossRef]
- Begg, C.B.; Mazumdar, M. Operating characteristics of a rank correlation test for publication bias. Biometrics 1994, 50, 1088–1101. [Google Scholar] [CrossRef] [PubMed]
- Egger, M.; Smith, G.D.; Schneider, M.; Minder, C. Bias in meta-analysis detected by a simple, graphical test. BMJ 1997, 315, 629. [Google Scholar] [CrossRef]
- White, I.R. Multivariate random-effects meta-regression: Updates to mvmeta. Stata J. 2011, 11, 255–270. [Google Scholar] [CrossRef]
- Bai, S.L.; Wang, J.Y.; Zhou, Y.Q.; Yu, D.S.; Gao, X.M.; Li, L.L.; Yang, F. Analysis of the first cluster of cases in a family of novel coronavirus pneumonia in Gansu Province. Zhonghua Yu Fang Yi Xue Za Zhi Chin. J. Prev. Med. 2020, 54, E005. [Google Scholar] [CrossRef]
- Bendavid, E.; Mulaney, B.; Sood, N.; Shah, S.; Ling, E.; Bromley-Dulfano, R.; Lai, C.; Weissberg, Z.; Saavedra, R.; Tedrow, J.; et al. COVID-19 Antibody Seroprevalence in Santa Clara County, California. medRxiv 2020. [Google Scholar] [CrossRef]
- Cassaniti, I.; Novazzi, F.; Giardina, F.; Salinaro, F.; Sachs, M.; Perlini, S.; Bruno, R.; Mojoli, F.; Baldanti, F. Performance of VivaDiag COVID-19 IgM/IgG Rapid Test is inadequate for diagnosis of COVID-19 in acute patients referring to emergency room department. J. Med. Virol. 2020. [Google Scholar] [CrossRef]
- Gao, Y.; Yuan, Y.; Li, T.T.; Wang, W.X.; Li, Y.X.; Li, A.; Han, F.M. Evaluation the auxiliary diagnosis value of antibodies assays for detection of novel coronavirus (SARS-Cov-2) causing an outbreak of pneumonia (COVID-19). medRxiv 2020. [Google Scholar] [CrossRef]
- Garcia, F.P.; Perez Tanoira, R.; Romanyk Cabrera, J.P.; Arroyo Serrano, T.; Gomez Herruz, P.; Cuadros Gonzalez, J. Rapid diagnosis of SARS-CoV-2 infection by detecting IgG and IgM antibodies with an immunochromatographic device: A prospective single-center study. medRxiv 2020. [Google Scholar] [CrossRef]
- Guo, L.; Ren, L.; Yang, S.; Xiao, M.; Chang, D.; Yang, F.; Dela Cruz, C.S.; Wang, Y.; Wu, C.; Xiao, Y.; et al. Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19). Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2020. [Google Scholar] [CrossRef]
- Jin, Y.; Wang, M.; Zuo, Z.; Fan, C.; Ye, F.; Cai, Z.; Wang, Y.; Cui, H.; Pan, K.; Xu, A. Diagnostic value and dynamic variance of serum antibody in coronavirus disease 2019. Int. J. Infect. Dis. IJID Off. Publ. Int. Soc. Infect. Dis. 2020, 94, 49–52. [Google Scholar] [CrossRef]
- Liu, L.; Liu, W.; Wang, S.; Zheng, S. A preliminary study on serological assay for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in 238 admitted hospital patients. medRxiv 2020. [Google Scholar] [CrossRef]
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; et al. Evaluation of Nucleocapsid and Spike Protein-based ELISAs for detecting antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, Y.; Diao, B.; Ren, F.; Wang, Y.; Ding, J.; Huang, Q. Diagnostic Indexes of a Rapid IgG/IgM Combined Antibody Test for SARS-CoV-2. medRxiv 2020. [Google Scholar] [CrossRef]
- Long, Q.-X.; Deng, H.-J.; Chen, J.; Hu, J.; Liu, B.-Z.; Liao, P.; Lin, Y.; Yu, L.-H.; Mo, Z.; Xu, Y.-Y.; et al. Antibody responses to SARS-CoV-2 in COVID-19 patients: The perspective application of serological tests in clinical practice. medRxiv 2020. [Google Scholar] [CrossRef]
- Okba, N.M.A.; Muller, M.A.; Li, W.; Wang, C.; GeurtsvanKessel, C.H.; Corman, V.M.; Lamers, M.M.; Sikkema, R.S.; de Bruin, E.; Chandler, F.D.; et al. Severe Acute Respiratory Syndrome Coronavirus 2-Specific Antibody Responses in Coronavirus Disease 2019 Patients. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Pan, Y.; Li, X.; Yang, G.; Fan, J.; Tang, Y.; Zhao, J.; Long, X.; Guo, S.; Zhao, Z.; Liu, Y.; et al. Serological immunochromatographic approach in diagnosis with SARS-CoV-2 infected COVID-19 patients. J. Infect. 2020. [Google Scholar] [CrossRef]
- Paradiso, A.V.; De Summa, S.; Loconsole, D.; Procacci, V.; Sallustio, A.; Centrone, F.; Silvestris, N.; Cafagna, V.; De Palma, G.; Tufaro, A.; et al. Clinical meanings of rapid serological assay in patients tested for SARS-Co2 RT-PCR. medRxiv 2020. [Google Scholar] [CrossRef]
- Tan, W.; Lu, Y.; Zhang, J.; Wang, J.; Dan, Y.; Tan, Z.; He, X.; Qian, C.; Sun, Q.; Hu, Q.; et al. Viral Kinetics and Antibody Responses in Patients with COVID-19. medRxiv 2020. [Google Scholar] [CrossRef]
- To, K.K.; Tsang, O.T.; Leung, W.S.; Tam, A.R.; Wu, T.C.; Lung, D.C.; Yip, C.C.; Cai, J.P.; Chan, J.M.; Chik, T.S.; et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Zeng, H.; Xu, C.; Fan, J.; Tang, Y.; Deng, Q.; Zhang, W.; Long, X. Antibodies in Infants Born to Mothers With COVID-19 Pneumonia. JAMA 2020. [Google Scholar] [CrossRef]
- Zeng, Z.; Chen, L.; Pan, Y.; Deng, Q.; Ye, G.; Li, Y.; Wang, X. Re: Profile of specific antibodies to SARS-CoV-2: The first report. J. Infect. 2020. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, J.; Li, N.; Liu, Y.; Ye, R.; Qin, X.; Zheng, R. Serological detection of 2019-nCoV respond to the epidemic: A useful complement to nucleic acid testing. medRxiv 2020. [Google Scholar] [CrossRef]
- Zhang, P.; Gao, Q.; Wang, T.; Ke, Y.; Mo, F.; Jia, R.; Liu, W.; Liu, L.; Zheng, S.; Liu, Y.; et al. Evaluation of recombinant nucleocapsid and spike proteins for serological diagnosis of novel coronavirus disease 2019 (COVID-19). medRxiv 2020. [Google Scholar] [CrossRef]
- Zhao, J.; Yuan, Q.; Wang, H.; Liu, W.; Liao, X.; Su, Y.; Wang, X.; Yuan, J.; Li, T.; Li, J.; et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2020. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Li, M.; Song, H.; Chen, J.; Ren, W.; Feng, Y.; Song, J.-W.; Peng, Y.; Su, B.; Guo, X.; et al. Serological diagnostic kit of SARS-CoV-2 antibodies using CHO-expressed full-length SARS-CoV-2 S1 proteins. medRxiv 2020. [Google Scholar] [CrossRef]
- Zheng, C.; Wang, J.; Guo, H.; Lu, Z.; Ma, Y.; Zhu, Y.; Xia, D.; Wang, Y.; He, H.; Zhou, J.; et al. Risk-adapted Treatment Strategy For COVID-19 Patients. Int. J. Infect. Dis. IJID Off. Publ. Int. Soc. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Zhong, L.; Chuan, J.; Gong, B.O.; Shuai, P.; Zhou, Y.; Zhang, Y.; Jiang, Z.; Zhang, D.; Liu, X.; Ma, S.; et al. Detection of serum IgM and IgG for COVID-19 diagnosis. Sci. China Life Sci. 2020. [Google Scholar] [CrossRef]
- Pan, A.; Liu, L.; Wang, C.; Guo, H.; Hao, X.; Wang, Q.; Huang, J.; He, N.; Yu, H.; Lin, X.; et al. Association of Public Health Interventions With the Epidemiology of the COVID-19 Outbreak in Wuhan, China. JAMA 2020. [Google Scholar] [CrossRef]
- Sethuraman, N.; Jeremiah, S.S.; Ryo, A. Interpreting Diagnostic Tests for SARS-CoV-2. JAMA 2020. [Google Scholar] [CrossRef]
- Wu, F.; Wang, A.; Liu, M.; Wang, Q.; Chen, J.; Xia, S.; Ling, Y.; Zhang, Y.; Xun, J.; Lu, L.; et al. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. medRxiv 2020. [Google Scholar] [CrossRef]
- Bao, L.; Deng, W.; Gao, H.; Xiao, C.; Liu, J.; Xue, J.; Lv, Q.; Liu, J.; Yu, P.; Xu, Y.; et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. BioRxiv 2020. [Google Scholar] [CrossRef]
| Author [Ref] | Covid19/Healthy | Covid19 Ascertainment | Severe Covid19 (%) | Male Cases (%) | Cases Age | Days from Onset | Antibodies | Method | Company | Limit of Detection IgM/IgG | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Liu [27] | 238/120 | RT-PCR/clinical features | NR | 58 | 55 | 14 | IgM (N)/IgG (N) | ELISA | ZhuHai LivZon, Diagnostics | A450: 0.100/0.130 | 0.11–0.81 | 0.96–0.99 |
| Long [55] | 262/148 | RT-PCR | 13.6 | 55.4 | 47 | 13 | IgM (N,S)/IgG (N,S) | CLIA | Bioscience (Chongqing) Co., Ltd. | NR | 0.67–0.80 | 0.95 |
| Jia [38] | 33/242 | NR-NAT/clinical features | NR | NR | NR | 15 | IgM (N,S)/IgG (N,S) | FIA | Beijing Diagreat Biotechnologies Co., Ltd. | Fluorescence Intensity: 0.88/1.02 (Flu units) | 0.45–0.72 | 0.95 |
| Liu [54] | 95/84 | RT-PCR | 49 | 70 | 76 | 18 | IgM (N)/IgG (N) | LFIA | Not Reported (a Chinese Company) | NA | 0.37–0.86 | 0.93–0.94 |
| Xu [33] | 10/0 | NAT/sequencing | 100 | 60 | NR | 22 | IgM (S)/IgG (S) | LFIA | In-house test | NA | 0.3–0.9 | NA |
| Wang [34] | 116/0 | RT-PCR/clinical features | 12.9 | 56 | 68.8 | 31 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 1 | NA |
| Xiang [28] | 63/35 ELISA, 91/35 LFIA | RT-PCR/clinical features | 6.3 | 55.5 | 57.82 | NR | IgM (N,S)/IgG (N,S) | ELISA/LFIA | ZhuHai LivZon, Diagnostics Inc.BioEasy/Shenzhen BioEasy Biotechnology Co. | NR/NA | 0.44–0.87 | 1 |
| Zhang [64] | 154/660 | RT-PCR/clinical features | NR | NR | NR | NR | IgM (S)/IgG (S) | LFIA | In-house test | NA | 0.82 | 0.99 |
| Lin [35] | 79/80 | RT-PCR/clinical features | NR | 35 | 43.6 | 14 | IgM (N)/IgG (N) | ELISA/CLIA | Darui Biotech, China/Tianshen Tech, Shenzhen, China | NR/NR | 0.23–0.91 | 0.78–1 |
| Hu [37] | 34/9 | RT-PCR | NR | NR | NR | NR | IgM (N,S)/IgG (N,S) | FIA | KingFocus Biomedical engineering Co., Ltd. | Cutoff values were based on of seronegative samples | 0.97-1 | 1 |
| Zhang [32] | 222/0 | RT-PCR | 39.2 | 48.2 | 64 | 20 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) and the high-speed CLIA system iFlash 3000, BATCH ANALYZER | Cutoff values were based on of seronegative samples | 0.83–0.99 | NA |
| Okba [56] | 12/0 | RT-PCR | NR | NR | NR | 11 | IgG (S) | ELISA | EUROIMMUN Medizinische Labordiagnostika AG | Cutoff values set by mean of seronegative samples plus 6SD | 0.92 | 1 |
| Zhang [63] | 3/733 | RT-PCR/clinical features | 66.6 | 66.6 | 50.67 | 10 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 1 | 0.98 |
| Zhao [66] | 69/412 | NR-NAT/clinical features | NR | NR | NR | NR | IgM (S)/IgG (S) | ELISA | In-house test | Cutoff values were based on seronegative samples | 0.97 | 0.97 |
| Pan [57] | 86/0 | RT-PCR/clinical features | NR | 45.7 | 58 | 12 | IgM (N,S)/IgG (N,S) | LFIA | ZhuHai LivZon, Diagnostics | NA | 0.55–0.69 | NA |
| Lou [31] | 80/300 | RT-PCR/clinical features | 33 | 61.3 | 55 | 15 | IgM (N,S)/IgG (N,S) | ELISA/CLIA/LFIA | Beijing Wantai Biological Pharmacy Enterprise Co., Ltd., China (Beijing, China)/Xiamen InnoDx Biotech Co., Ltd. | NR/NR/NA | 0.86–0.97 | 0.95–1 |
| Liu [27] | 133/0 | RT-PCR/clinical features | 66.9 | 52.6 | 68.5 | NR | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 0.79–0.97 | NA |
| Tan [59] | 65/0 | RT-PCR/clinical features | 43.3 | 52.2 | 49 | 15 | IgM (N)/IgG (N) | ELISA | ZhuHai LivZon, Diagnostics | Titer cutoff value set according to non-responders | 0.43–0.78 | NA |
| To [60] | 16/0 | RT-PCR/sequencing/clinical features | 43.5 | 56.5 | 62 | 20 | IgM (N,S)/IgG (N,S) | ELISA | In-house test | Cutoff set by mean of seronegative samples plus 3SD | 0.87–1 | NA |
| Xiao [29] | 34/0 | RT-PCR/clinical features | NR | 64.7 | 55 | 25 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 0.82–0.94 | NA |
| Cassaniti [47] | 30/38 | RT-PCR | NR | 83.3 | 73.5/61.5 | 7 | IgM (N,S)/IgG (N,S) | LFIA | VivaChekTM | NA | 0.13–0.83 | 1 |
| Liu [53] | 214/100 | RT-PCR | NR | NR | NR | 15 | IgM (N,S)/IgG (N,S) | ELISA | ZhuHai LivZon, Diagnostics | A450: 0.100/0.130 | 0.68–0.77 | 1 |
| Li [26] | 397/128 | RT-PCR | NR | NR | NR | 20 | IgM (S)/IgG (S) | LFIA | Jiangsu Medomics Medical Technologies | NA | 0.7–0.82 | 0.91 |
| Zhao [65] | 173/0 | RT-PCR/clinical features | 18.5 | 48.5 | 48 | 7 | IgM (S)/IgG (S) | ELISA | Beijing Wantai Biological Pharmacy Enterprise Co., Ltd. | Cutoff value set by seronegative samples | 0.65–0.93 | NA |
| Bai [45] | 6/0 | RT-PCR/clinical features | 16.7 | 50 | 49 | 2 | IgM (N,S) | LFIA | Institute of Microbiology and Epidemiology of the Military Medical Research Institute and Beijingh Rejing Biotecnology Co., Ltd. | NA | 0.83 | NA |
| Zheng [67] | 55/0 | RT-PCR/clinical features | 40 | 43.6 | 60 | 11 | IgM (N,S)/IgG (N,S) | CLIA | Not Reported | NR | 0.82–0.98 | NA |
| Zeng [61] | 6/0 | RT-PCR/clinical features | 0 | 0 | NR | NR | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 0.83 | 1 |
| Guo [50] | 140/285 | RT-PCR/sequencing/clinical features | 23.6 | NR | NR | 13 | IgM (N) | ELISA | In-house test | A450: 0.130/0.300 | 0.83 | 1 |
| Jin [51] | 27/33 | RT-PCR | 0 | 39.5 | 47 | 16 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 0.48–0.89 | 0.9-1 |
| Du [25] | 60/0 | NR-NAT/clinical features | NR | NR | NR | 43 | IgM (N,S)/IgG (N,S) | CLIA | YHLO Biotechnology (Shenzhen, China) | 10 AU/mL | 0.78–1 | NA |
| Wölfel [36] | 9/0 | RT-PCR/clinical features | 0 | NR | NR | 18 | IgM (S)/IgG (S) | FIA | In-house with reagents from Euroimmun AG, Lübeck, Germany | NR | 0.66–1 | NA |
| Zhong [68] | 47/300 | NR-NAT | 23.4 | 34 | 48.21 | 15 | IgM (N,S)/IgG (N,S) | ELISA / CLIA | In-house test | A450: IgM(N) 0.059, IgM(S) 0.167/IgG(N) 0.036, IgG(S) 0.079/NR | 0.89–0.98 | 0.85–0.97 |
| Lassaunière [30] | 30/82 | RT-PCR | 100 | NR | NR | NR | IgM (N,S)/IgG (N,S) | ELISA / LFIA | Εuroimmun Medizinische Labordiagnostika, Lübeck, Germany/Beijing Wantai Biological Pharmacy Enterprise, Beijing, China/Dynamiker Biotechnology, Tianjin, China/CTK Biotech, Poway, CA, USA/AutoBio Diagnostics, Zhengzhou, China/Artron, Laboratories, Burnaby, Canada | NR/NA | 0.66–0.93 | 0.95–1 |
| Gao [48] | 38/0 | RT-PCR/clinical features | 7.9 | 55.3 | 40.5 | 16 | IgM (N,S)/IgG (N,S) | LFIA | Innovita Biological Technology Co., Ltd. | NA | 0.51–0.92 | NA |
| Zeng [62] | 27/36 | RT-PCR/clinical features | 63 | 51.8 | 62 | 18 | IgM (N)/IgG (N) | ELISA | ZhuHai LivZon, Diagnostics | A450: 0.105/0.105 | 1 | 1 |
| Garcia [49] | 118/45 | RT-PCR/clinical features | NR | 67.8 | 65.14 | 14 | IgM (N,S)/IgG (N,S) | LFIA | Biotech AllTest, Hangzhou, China | NA | 0.31–0.69 | 1 |
| Paradiso [58] | 191/0 | RT-PCR/clinical features | NR | 60.62 | 58.5 | 4 | IgM (N,S)/IgG (N,S) | LFIA | VivaChekTM | NA | 0.14–0.16 | NA |
| Bendavid [46] | 122/456 | RT-PCR | NR | NR | NR | NR | IgM (N,S)/IgG (N,S) | LFIA | Premier Biotech | NA | 0.67–0.92 | 0.99–1 |
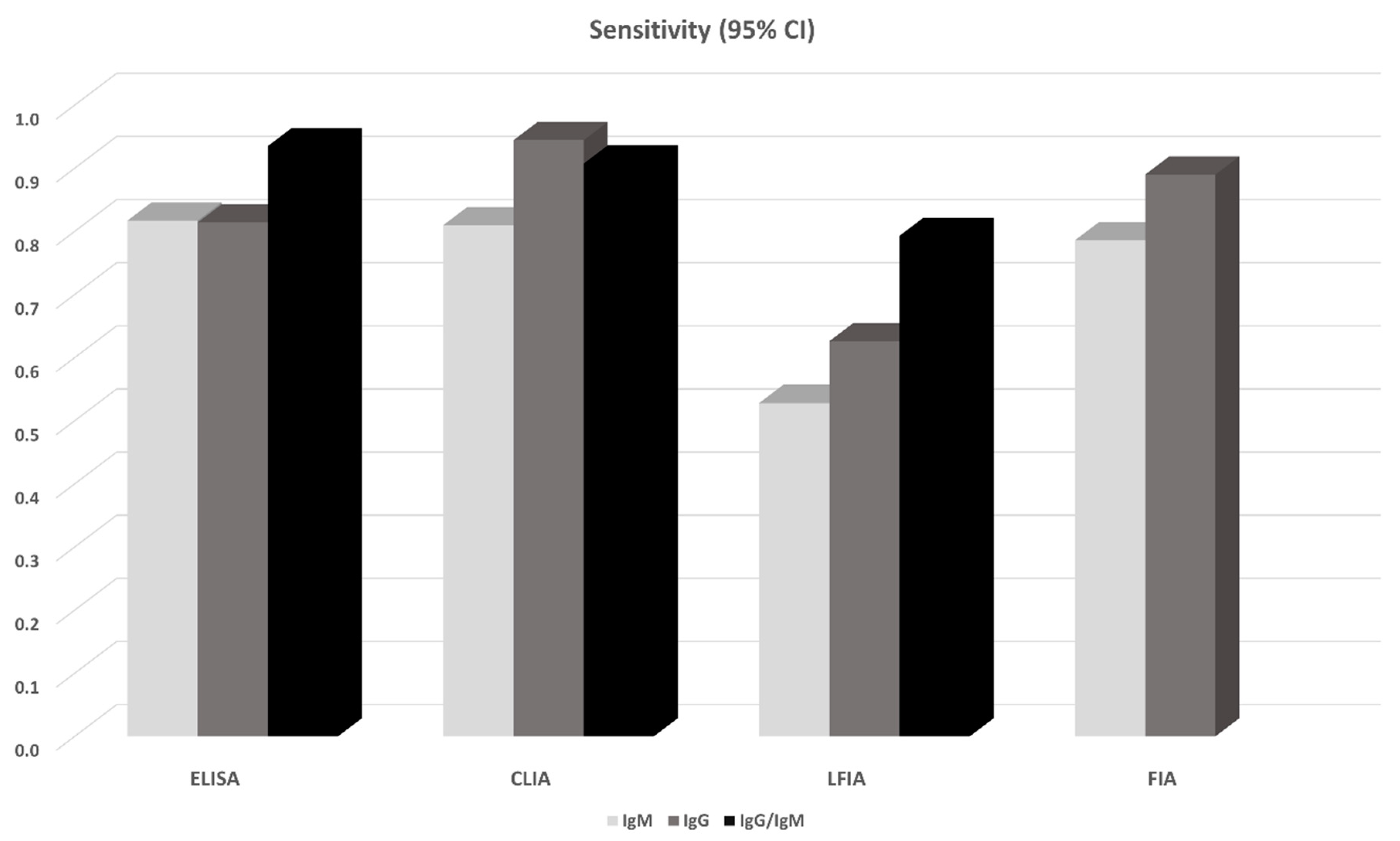
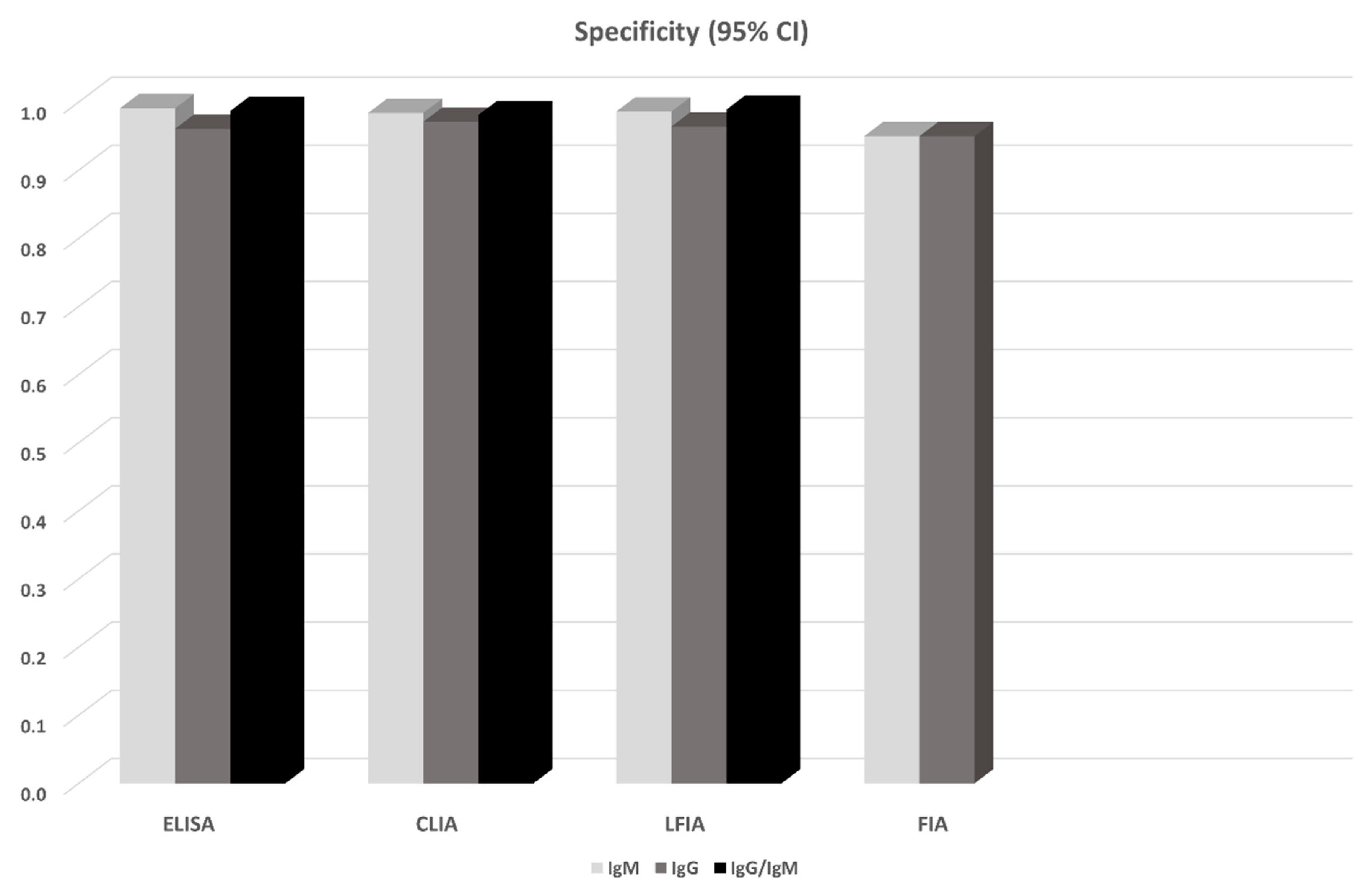
| Method | Ab | Ag | Studies/Patients | Sensitivity (95% CI) | Specificity (95% CI) | Covariates | Begg’s/Egger’s |
|---|---|---|---|---|---|---|---|
| ELISA | IgG | N | 8/1472 | 0.747 (0.509, 0.984) | 0.994 (0.988, 0.999) | mdfo, severe | -/- |
| ELISA | IgG | S | 7/1072 | 0.814 (0.688, 0.940) | 0.961 (0.910, 1.000) | - | -/- |
| ELISA | IgM | N | 8/1717 | 0.722 (0.449, 0.996) | 0.995 (0.989, 1.000) | - | -/- |
| ELISA | IgM | S | 6/1328 | 0.817 (0.704, 0.931) | 0.991 (0.976, 1.000) | - | -/- |
| ELISA | IgG/IgM | N | 2/423 | 0.808 (0.764, 0.853) | 0.967 (0.915, 0.987) | NA | NA |
| ELISA | IgG/IgM | S | 5/1244 | 0.935 (0.900, 0.971) | 0.987 (0.973, 1.000) | - | -/- |
| LFIA | IgG | S | 2/535 | 0.537 (0.123, 0.951) | 0.914 (0.853, 0.951) | NA | NA |
| LFIA | IgG | NS | 8/944 | 0.650 (0.404, 0.895) | 0.988 (0.973, 1.000) | mdfo | -/- |
| LFIA | IgG | S/NS | 10/1479 | 0.626 (0.439, 0.814) | 0.964 (0.922, 1.000) | - | -/- |
| LFIA | IgM | S | 2/535 | 0.663 (0.236, 1.000) | 0.914 (0.852, 0.951) | NA | NA |
| LFIA | IgM | NS | 9/1059 | 0.528 (0.329, 0.726) | 0.986 (0 974, 0.998) | - | -/- |
| LFIA | IgM | S/NS | 11/1594 | 0.555 (0.352, 0.758) | 0.979 (0.958, 0.999) | - | -/- |
| LFIA | IgG/IgM | S | 2/824 | 0.828 (0.770, 0.886) | 0.994 (0.984, 0.998) | NA | NA |
| LFIA | IgG/IgM | NS | 8/1373 | 0.777 (0.592. 0.962) | 0.986 (0.973, 1.000) | mdfo | -/- |
| LFIA | IgG/IgM | S/NS | 10/2197 | 0.793 (0.643, 0.942) | 0.989 (0.978, 0.999) | mdfo | -/- |
| LFIA | IgG/IgM | S/N/NS | 11/2376 | 0.800 (0.663, 0.935) | 0.984 (0.969, 0.999) | mdfo | -/- |
| CLIA | IgG | NS | 12/2320 | 0.944 (0.906, 0.983) | 0 971 (0.931, 1.000) | mdfo | -/+ |
| CLIA | IgG | N/NS | 13/2479 | 0.935 (0.896, 0.975) | 0.974 (0.953, 0.994) | mdfo | -/+ |
| CLIA | IgM | NS | 12/2411 | 0.810 (0.722, 0.897) | 0.984 (0.970, 0.999) | - | -/- |
| CLIA | IgM | N/NS | 13/2570 | 0.799 (0.737, 0.860) | 0.967 (0.927, 1.000) | - | -/- |
| CLIA | IgG/IgM | NS | 2/790 | 0.907 (0.753, 1.000) | 0.981 (0.944, 1.000) | NA | NA |
| CLIA | IgG/IgM | N/NS | 3/949 | 0.902 (0.811, 0.993) | 0.954 (0.875, 1.000) | NA | NA |
| FIA | IgG | NS | 2/318 | 0.859 (0.339, 1.000) | 0.950 (0.923, 0.977) | NA | NA |
| FIA | IgG | S/NS | 3/327 | 0.890 (0.591, 1.000) | 0.950 (0.923, 0.977) | NA | NA |
| FIA | IgM | NS | 2/318 | 0.860 (0.500, 1.000) | 0.950 (0.923, 0.977) | NA | NA |
| FIA | IgM | S/NS | 3/327 | 0.786 (0.531, 1.000) | 0.950 (0.923, 0.977) | NA | NA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kontou, P.I.; Braliou, G.G.; Dimou, N.L.; Nikolopoulos, G.; Bagos, P.G. Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis. Diagnostics 2020, 10, 319. https://doi.org/10.3390/diagnostics10050319
Kontou PI, Braliou GG, Dimou NL, Nikolopoulos G, Bagos PG. Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis. Diagnostics. 2020; 10(5):319. https://doi.org/10.3390/diagnostics10050319
Chicago/Turabian StyleKontou, Panagiota I., Georgia G. Braliou, Niki L. Dimou, Georgios Nikolopoulos, and Pantelis G. Bagos. 2020. "Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis" Diagnostics 10, no. 5: 319. https://doi.org/10.3390/diagnostics10050319
APA StyleKontou, P. I., Braliou, G. G., Dimou, N. L., Nikolopoulos, G., & Bagos, P. G. (2020). Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis. Diagnostics, 10(5), 319. https://doi.org/10.3390/diagnostics10050319







