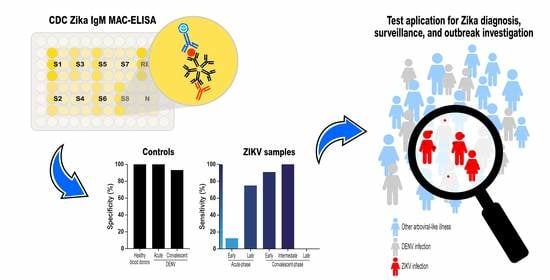
Accuracy of the Zika IgM Antibody Capture Enzyme-Linked Immunosorbent Assay from the Centers for Disease Control and Prevention (CDC Zika MAC-ELISA) for Diagnosis of Zika Virus Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Sample Selection
2.2. Laboratorial Tests
2.3. Data Analysis
3. Results
3.1. Participant Characteristics
3.2. CDC Zika MAC-ELISA Performance
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Test Result | % (95% CI) | McNemar p Value | ||
|---|---|---|---|---|
| Pos. | Neg. | |||
| Sensitivity | ||||
| Acute-phase samples | ||||
| CDC ZIKV IgM MAC-ELISA | 1 | 13 | 7.1 (0.2–33.9) | 0.48 |
| Euroimmun anti-ZikaVirusIgM ELISA | 0 | 14 | 0.0 (0.0–23.2) | |
| Convalescent-phase samples | ||||
| CDC ZIKV IgM MAC-ELISA | 8 | 0 | 100 (63.1–100) | 0.01 |
| Euroimmun anti-ZikaVirusIgM ELISA | 1 | 7 | 12.5 (0.3–52.6) | |
| Specificity against RT-PCR-confirmed dengue cases | ||||
| Acute-phase samples | ||||
| CDC ZIKV IgM MAC-ELISA a | 0 | 59 | 100 (93.9–100) | 1.00 c |
| Euroimmun anti-ZikaVirusIgMELISA b | 0 | 58 | 100 (93.8–100) | |
| Convalescent-phase samples | ||||
| CDC ZIKV IgM MAC-ELISA d | 4 | 55 | 93.2 (83.5–98.1) | 0.18 f |
| Euroimmun anti-ZikaVirusIgMELISA e | 1 | 58 | 98.3 (90.9–100) | |
| Specificity against blood donors’ samples | ||||
| CDC ZIKV IgM MAC-ELISA | 0 | 23 | 100 (85.2–100) | 1.00 |
| Euroimmun anti-ZikaVirusIgM ELISA | 0 | 23 | 100 (85.2–100) | |
References
- Cao-Lormeau, V.-M.; Blake, A.; Mons, S.; Lastère, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-Barré Syndrome outbreak caused by ZIKA virus infection in French Polynesia: A case-control study. Lancet 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- De Araújo, T.V.B.; Rodrigues, L.C.; Ximenes, R.A.D.A.; Miranda-Filho, D.D.B.; Montarroyos, U.R.; De Melo, A.P.L.; Valongueiro, S.; Albuquerque, M.F.P.M.; De Souza, W.V.; Braga, M.C.; et al. Association between Zika virus infection and microcephaly in Brazil, January to May, 2016: Preliminary report of a case-control study. Lancet Infect. Dis. 2016, 16, 1356–1363. [Google Scholar] [CrossRef]
- Paploski, I.A.D.; Prates, A.P.P.; Cardoso, C.W.; Kikuti, M.; Silva, M.M.O.; Waller, L.A.; Reis, M.G.; Kitron, U.; Ribeiro, G.S. Time Lags between Exanthematous Illness Attributed to Zika Virus, Guillain-Barré Syndrome, and Microcephaly, Salvador, Brazil. Emerg. Infect. Dis. 2016, 22, 1438–1444. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Zika MAC-ELISA—Instructions for Use. 2018. Available online: https://www.cdc.gov/zika/pdfs/zika-mac-elisa-instructions-for-use.pdf. (accessed on 8 June 2020).
- Landry, M.L.; George, K.S. Laboratory Diagnosis of Zika Virus Infection. Arch. Pathol. Lab. Med. 2017, 141, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Stettler, K.; Beltramello, M.; Espinosa, D.A.; Graham, V.; Cassotta, A.; Bianchi, S.; Vanzetta, F.; Minola, A.; Jaconi, S.; Mele, F.; et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 2016, 353, 823–827. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.M.O.; Tauro, L.B.; Kikuti, M.; O Anjos, R.; Santos, V.C.; Gonçalves, T.S.F.; Paploski, I.A.D.; Moreira, P.S.S.; Nascimento, L.C.J.; Campos, G.S.; et al. Concomitant Transmission of Dengue, Chikungunya, and Zika Viruses in Brazil: Clinical and Epidemiological Findings From Surveillance for Acute Febrile Illness. Clin. Infect. Dis. 2019, 69, 1353–1359. [Google Scholar] [CrossRef] [PubMed]
- Pessôa, R.; Patriota, J.V.; Souza, M.D.L.D.; Felix, A.C.; Mamede, N.; Sanabani, S.S. Investigation Into an Outbreak of Dengue-like Illness in Pernambuco, Brazil, revealed a co-circulation of Zika, chikungunya, and Dengue virus type 1. Medicine 2016, 95, 1–9. [Google Scholar] [CrossRef]
- Kikuti, M.; Cunha, G.M.; Paploski, I.A.D.; Kasper, A.M.; Silva, M.M.O.; Tavares, A.S.; Cruz, J.S.; Queiroz, T.L.; Rodrigues, M.S.; Santana, P.M.; et al. Spatial Distribution of Dengue in a Brazilian Urban Slum Setting: Role of Socioeconomic Gradient in Disease Risk. PLoS Negl. Trop. Dis. 2015, 9, 1–18. [Google Scholar] [CrossRef]
- Ribeiro, G.S.; Kikuti, M.; Tauro, L.B.; Nascimento, L.C.J.; Cardoso, C.W.; Campos, G.S.; Ko, A.I.; Weaver, S.C.; Dos Reis, M.G.; Kitron, U.D.; et al. Does immunity after Zika virus infection cross-protect against Dengue? Lancet Glob. Health 2017, 6, e140–e141. [Google Scholar] [CrossRef]
- Cardoso, C.W.; Paploski, I.A.D.; Kikuti, M.; Rodrigues, M.M.D.S.; Silva, M.M.; Campos, G.S.; Sardi, S.I.; Kitron, U.; Reis, M.G.; Ribeiro, G.S. Outbreak of Exanthematous Illness Associated with Zika, Chikungunya, and Dengue Viruses, Salvador, Brazil. Emerg. Infect. Dis. 2015, 21, 2274–2276. [Google Scholar] [CrossRef]
- Faria, N.R.; Azevedo, R.D.S.D.S.; Kraemer, M.U.G.; Souza, R.; Cunha, M.S.; Hill, S.; Theze, J.; Bonsall, M.B.; Bowden, T.A.; Rissanen, I.; et al. Zika virus in the Americas: Early epidemiological and genetic findings. Science 2016, 352, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Ayllón, T.; Campos, R.D.M.; Brasil, P.; Morone, F.C.; Câmara, D.C.P.; Meira, G.L.S.; Tannich, E.; Yamamoto, K.A.; Carvalho, M.S.; Pedro, R.S.; et al. Early evidence for zika virus circulation among aedes aegypti mosquitoes, Rio de Janeiro, Brazil. Emerg. Infect. Dis. 2017, 23, 1411–1412. [Google Scholar] [CrossRef] [PubMed]
- Passos, S.R.L.; Dos Santos, M.A.B.; Neto, J.C.; Buonora, S.; Souza, T.M.L.; De Oliveira, R.V.; Vizzoni, A.; Barbosa-Lima, G.; Vieira, Y.R.; De Lima, M.S.; et al. Detection of Zika virus in April 2013 patient samples, Rio de Janeiro, Brazil. Emerg. Infect. Dis. 2017, 23, 2120–2121. [Google Scholar] [CrossRef] [PubMed]
- Balm, M.N.; Lee, C.K.; Lee, H.K.; Chiu, L.; Koay, E.S.-C.; Tang, J.W. A Diagnostic Polymerase Chain Reaction Assay for Zika Virus. J. Med Virol. 2012, 84, 1501–1505. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.S.; Bandeira, A.C.; Sardi, S.I. Zika Virus Outbreak, Bahia, Brazil. Emerg. Infect. Dis. 2015, 21, 1885–1886. [Google Scholar] [CrossRef] [PubMed]
- Lanciotti, R.S.; Calisher, C.H.; Gubler, D.J.; Chang, G.J.; Vorndam, A.V. Rapid Detection and Typing of Dengue Viruses from Clinical Samples by Using Reverse Transcriptase-Polymerase Chain Reaction. J. Clin. Microbiol. 1992, 30, 545–551. [Google Scholar] [CrossRef]
- Santiago, G.A.; Vazquez, J.; Courtney, S.; Matías, K.Y.; Andersen, L.E.; Colón, C.; Butler, A.E.; Roulo, R.; Bowzard, J.; Villanueva, J.M.; et al. Performance of the Trioplex Real-Time RT-PCR assay for detection of Zika, dengue and chikungunya viruses. Nat. Commun. 2018, 9, 1391. [Google Scholar] [CrossRef]
- Lanciotti, R.S.; Kosoy, O.L.; Laven, J.J.; Velez, J.O.; Lambert, A.J.; Johnson, A.J.; Stanfield, S.M.; Duffy, M.R. Genetic and Serologic Properties of Zika Virus Associated with an Epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 2008, 14, 1232–1239. [Google Scholar] [CrossRef]
- Griffin, I.; Martin, S.W.; Fischer, M.; Chambers, T.V.; Kosoy, O.; Falise, A.; Ponomareva, O.; Gillis, L.D.; Blackmore, C.; Jean, R. Zika Virus IgM Detection and Neutralizing Antibody Profiles 12–19 Months after Illness Onset. Emerg. Infect. Dis. 2019, 25, 299–303. [Google Scholar] [CrossRef]
- Lee, W.T.; Wong, S.J.; Kulas, K.E.; Dupuis, A.P.; Payne, A.F.; Kramer, L.D.; Dean, A.B.; George, K.S.; White, J.L.; Sommer, J.N.; et al. Development of Zika Virus Serological Testing Strategies in New York State. J. Clin. Microbiol. 2017, 56, e01591-17. [Google Scholar] [CrossRef]
- L’huillier, A.G.; Hamid-Allie, A.; Kristjanson, E.; Papageorgiou, L.; Hung, S.; Wong, C.F.; Stein, D.R.; Olsha, R.; Goneau, L.W.; Dimitrova, K.; et al. Evaluation of Euroimmun anti-Zika virus IgM and IgG Enzyme-linked immunosorbent assays for Zika Virus serologic testing. J. Clin. Microbiol. 2017, 55, 2462–2471. [Google Scholar] [CrossRef]
- Granger, D.; Hilgart, H.; Misner, L.; Christensen, J.; Bistodeau, S.; Palm, J.; Strain, A.K.; Konstantinovski, M.; Liu, D.; Tran, A.; et al. Serologic testing for zika virus: Comparison of three zika virus IgM screening enzyme-linked immunosorbent assays and initial laboratory experiences. J. Clin. Microbiol. 2017, 55, 2127–2136. [Google Scholar] [CrossRef] [PubMed]
- Safronetz, D.; Sloan, A.; Stein, D.R.; Mendoza, E.; Barairo, N.; Ranadheera, C.; Scharikow, L.; Holloway, K.; Robinson, A.; Traykova-Andonova, M.; et al. Evaluation of 5 commercially available Zika Virus Immunoassays. Emerg. Infect. Dis. 2017, 23, 1577–1580. [Google Scholar] [CrossRef] [PubMed]
- Balmaseda, A.; Zambrana, J.V.; Collado, D.; Garcia, D.L.C.; Saborío, S.; Elizondo, D.; Mercado, J.C.; González, K.; Cerpas, C.; Nuñez, A.; et al. Comparison of Four Serological Methods and Two Reverse Transcription-PCR Assays for Diagnosis and Surveillance of Zika Virus Infection. J. Clin. Microbiol. 2018, 56, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kikuti, M.; Tauro, L.B.; Moreira, P.S.S.; Campos, G.S.; Paploski, I.A.D.; Weaver, S.C.; Reis, M.G.; Kitron, U.; Ribeiro, G.S. Diagnostic performance of commercial IgM and IgG enzyme-linked immunoassays (ELISAs) for diagnosis of Zika virus infection. Virol. J. 2018, 15, 1–7. [Google Scholar] [CrossRef]
| Characteristics | Zika Cases | Dengue Cases from Salvador | |||||
|---|---|---|---|---|---|---|---|
| from Salvador (n = 14) | from Campo Formoso (n = 7) | Total (n = 21) | DENV-1 (n = 20) | DENV-2 (n = 20) | DENV-4 (n = 20) | Total (n = 60) | |
| Number (%) or median (interquartile range) | |||||||
| Demographic | |||||||
| Female | 8 (57.1) | 5 (71.4) | 13 (61.9) | 9 (45.0) | 10 (50.0) | 10 (50.0) | 29 (48.3) |
| Age | 22.5 (15–41) | 24 (10–34) | 24 (14–38) | 20 (10.5–37) | 13.5 (9.5–22) | 22.5 (10.5–34.5) | 18 (10–34) |
| Clinical manifestation | |||||||
| Fever | 14 (100) | 7 (100) | 21 (100) | 20 (100) | 20 (100) | 20 (100) | 60 (100) |
| Headache | 13 (92.9) | 7 (100) | 20 (95.2) | 19 (95.0) | 19 (95.0) | 20 (100) | 58 (96.7) |
| Myalgia | 12 (85.7) | 5 (71.4) | 17 (80.9) | 18 (90.0) | 14 (70.0) | 14 (70.0) | 46 (76.7) |
| Rash | 10 (71.4) | 6 (85.7) | 16 (76.2) | 1 (5.0) | 5 (25.0) | 0 | 6 (10.0) |
| Pruritus | 10 (71.4) | 6 (85.7) | 16 (76.2) | NA | NA | NA | NA |
| Retro-orbital pain | 10 (71.4) | 6 (85.7) | 16 (76.2) | 14 (70.0) | 9 (45.0) | 11 (55.0) | 34 (56.7) |
| Arthralgia | 8 (57.1) | 6 (85.7) | 14 (66.7) | 10 (50.0) | 8 (40.0) | 10 (50.0) | 28 (46.7) |
| Vomiting | 2 (14.3) | 5 (71.4) | 7 (33.3) | 6 (30.0) | 2 (10.0) | 4 (20.0) | 12 (20.0) |
| Blood sample collection | |||||||
| Acute-phase sample | |||||||
| Early (0–4 DPSO) | 14 (100) | 2 (28.6) | 16 (76.2) | 19 (95.0) | 20 (100) | 20 (100) | 59 (98.3) |
| Late (5–9 DPSO) | 0 | 5 (71.4) | 5 (23.8) | 1 (5.0) | 0 | 0 | 1 (1.7) |
| Convalescent-phase sample a | |||||||
| Early (12–102 DPSO) | 8 (57.1) | 3 (42.9) | 11 (52.4) | 20 (100) | 20 (100) | 20 (100) | 60 (100) |
| Intermediate (258–260 DPSO) | 0 | 4 (57.1) | 4 (57.1) | 0 | 0 | 0 | 0 |
| Late (722–727 DPSO) | 0 | 4 (57.1) | 4 (57.1) | 0 | 0 | 0 | 0 |
| ZIKV diagnosis | |||||||
| RT-PCR positivity | 14 (100) | NA | 14 (100) | 0 | 0 | 0 | 0 b |
| qRT-PCR positivity | 12 (85.7) | 7 (100) | 19 (90.5) | NA | NA | NA | NA |
| Ct value on qRT-PCR | 32.3 (30.0–36.7) | 34.3 (26.3–36.8) | 33.6 (30.0–36.8) | NA | NA | NA | NA |
| DENV diagnosis | |||||||
| RT-PCR positivity c | 1 (7.1) | 1 (14.3) | 2 (9.5) | 20 (100) | 20 (100) | 20 (100) | 60 (100) |
| NS1-ELISA positivity | 0 | 0 | 0 | 15 (75.0) | 6 (30.0) | 15 (75.0) | 36 (60.0) |
| IgM-ELISA positivity d | 1 (7.1) | 0 | 1 (4.8) | 4 (20.0) | 0 | 2 (10.0) | 6 (10.0) |
| Prior DENV infection e | 11 (84.6) | 4 (57.1) | 15 (75.0) | 13 (65.0) | 14 (70.0) | 20 (100) | 47 (78.3) |
| Performance according to the Type of Serum Samples | Tested Samples | ZIKV IgM-Mac-ELISA Result | % (95% CI) | |
|---|---|---|---|---|
| Pos. | Neg. | |||
| Sensitivity against RT-PCR-confirmed Zika cases | ||||
| Acute-phase samples | ||||
| Overall a | 20 | 5 | 15 | 25.0 (8.7–49.1) |
| Early (0–4 DPSO) | 16 | 2 | 14 | 12.5 (1.5–38.3) |
| Late (5–9 DPSO) a | 4 | 3 | 1 | 75.0 (19.4–99.4) |
| CT value on qRT-PCR <34 b | 11 | 1 | 10 | 9.1 (0.2–41.3) |
| CT value on qRT-PCR ≥34 a,b | 7 | 3 | 4 | 42.9 (9.9–81.6) |
| Convalescent-phase samples | ||||
| Early (12–102 DPSO) | 11 | 10 | 1 | 90.9 (58.7–99.8) |
| Intermediate (258–260 DPSO) c | 2 | 2 | 0 | 100 (15.8–100) |
| Late convalescent-phase samples (722–727 DPSO) d | 2 | 0 | 2 | 0.0 (0.0–84.2) |
| CT value on qRT-PCR <34 b,e | 5 | 5 | 0 | 100 (47.9–100) |
| CT value on qRT-PCR ≥34 b,e | 4 | 3 | 1 | 75.0 (19.4–99.4) |
| Specificity against RT-PCR-confirmed dengue cases | ||||
| Acute-phase samples | ||||
| Overall f | 59 | 0 | 59 | 100 (93.9–100) |
| DENV-1 cases f | 19 | 0 | 19 | 100 (82.3–100) |
| DENV-2 cases | 20 | 0 | 20 | 100 (83.2–100) |
| DENV-4 cases | 20 | 0 | 20 | 100 (83.2–100) |
| Primary DENV infection | 13 | 0 | 13 | 100 (75.3–100) |
| Secondary DENV infection f | 46 | 0 | 46 | 100 (92.3–100) |
| Convalescent-phase samples | ||||
| Overall g | 59 | 4 | 55 | 93.2 (83.5–98.1) |
| DENV-1 cases | 20 | 2 | 18 | 90.0 (68.3–98.8) |
| DENV-2 cases | 20 | 1 | 19 | 95.0 (75.1–99.9) |
| DENV-4 cases g | 19 | 1 | 18 | 94.7 (74.0–99.9) |
| Primary DENV infection | 13 | 1 | 12 | 92.3 (64.0–99.8) |
| Secondary DENV infection g | 46 | 3 | 43 | 93.5 (82.1–98.6) |
| Specificity against blood donors’ samples | 23 | 0 | 23 | 100 (85.2–100) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Machado Portilho, M.; de Moraes, L.; Kikuti, M.; Jacob Nascimento, L.C.; Galvão Reis, M.; Sampaio Boaventura, V.; Khouri, R.; Sousa Ribeiro, G. Accuracy of the Zika IgM Antibody Capture Enzyme-Linked Immunosorbent Assay from the Centers for Disease Control and Prevention (CDC Zika MAC-ELISA) for Diagnosis of Zika Virus Infection. Diagnostics 2020, 10, 835. https://doi.org/10.3390/diagnostics10100835
Machado Portilho M, de Moraes L, Kikuti M, Jacob Nascimento LC, Galvão Reis M, Sampaio Boaventura V, Khouri R, Sousa Ribeiro G. Accuracy of the Zika IgM Antibody Capture Enzyme-Linked Immunosorbent Assay from the Centers for Disease Control and Prevention (CDC Zika MAC-ELISA) for Diagnosis of Zika Virus Infection. Diagnostics. 2020; 10(10):835. https://doi.org/10.3390/diagnostics10100835
Chicago/Turabian StyleMachado Portilho, Moyra, Laise de Moraes, Mariana Kikuti, Leile Camila Jacob Nascimento, Mitermayer Galvão Reis, Viviane Sampaio Boaventura, Ricardo Khouri, and Guilherme Sousa Ribeiro. 2020. "Accuracy of the Zika IgM Antibody Capture Enzyme-Linked Immunosorbent Assay from the Centers for Disease Control and Prevention (CDC Zika MAC-ELISA) for Diagnosis of Zika Virus Infection" Diagnostics 10, no. 10: 835. https://doi.org/10.3390/diagnostics10100835
APA StyleMachado Portilho, M., de Moraes, L., Kikuti, M., Jacob Nascimento, L. C., Galvão Reis, M., Sampaio Boaventura, V., Khouri, R., & Sousa Ribeiro, G. (2020). Accuracy of the Zika IgM Antibody Capture Enzyme-Linked Immunosorbent Assay from the Centers for Disease Control and Prevention (CDC Zika MAC-ELISA) for Diagnosis of Zika Virus Infection. Diagnostics, 10(10), 835. https://doi.org/10.3390/diagnostics10100835