Metatranscriptomic Analysis of the Bacterial Symbiont Dactylopiibacterium carminicum from the Carmine Cochineal Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae)
Abstract
1. Introduction
2. Materials and Methods
2.1. Tissue Dissection, RNA Extraction, and Sequencing
2.2. Bioinformatic Analyses
2.2.1. Mapping to Reference Genome, Core Transcriptome, and Differential Expression Analysis
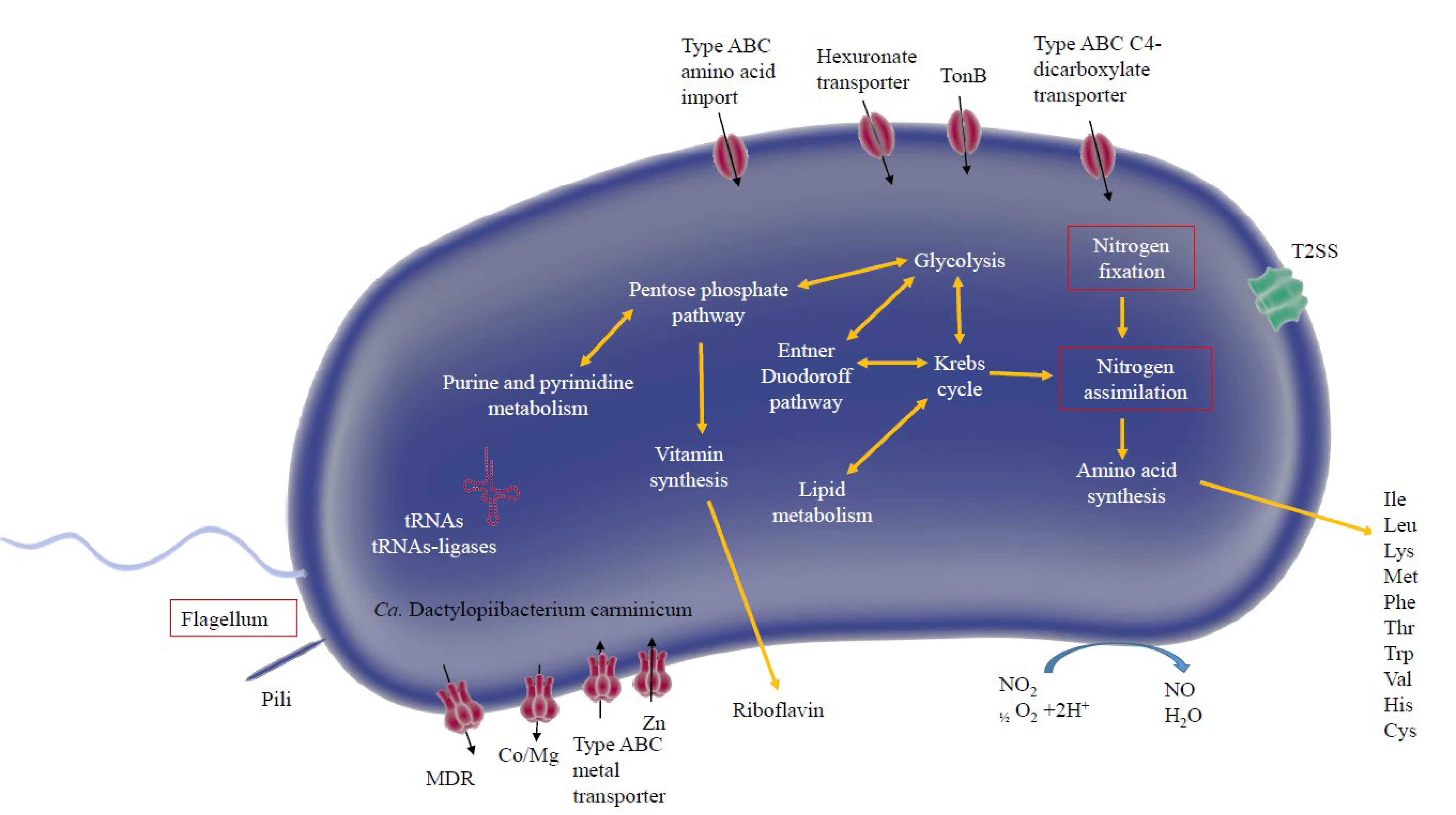
2.2.2. Metabolic Analysis and Construction of a Metabolic Model
2.2.3. Quantitative Analysis
3. Results
3.1. Dactylopiibacterium Expression in the Carmine Cochineal with Core Transcriptome Analysis
3.2. Differential Expression Analysis with NOISeq
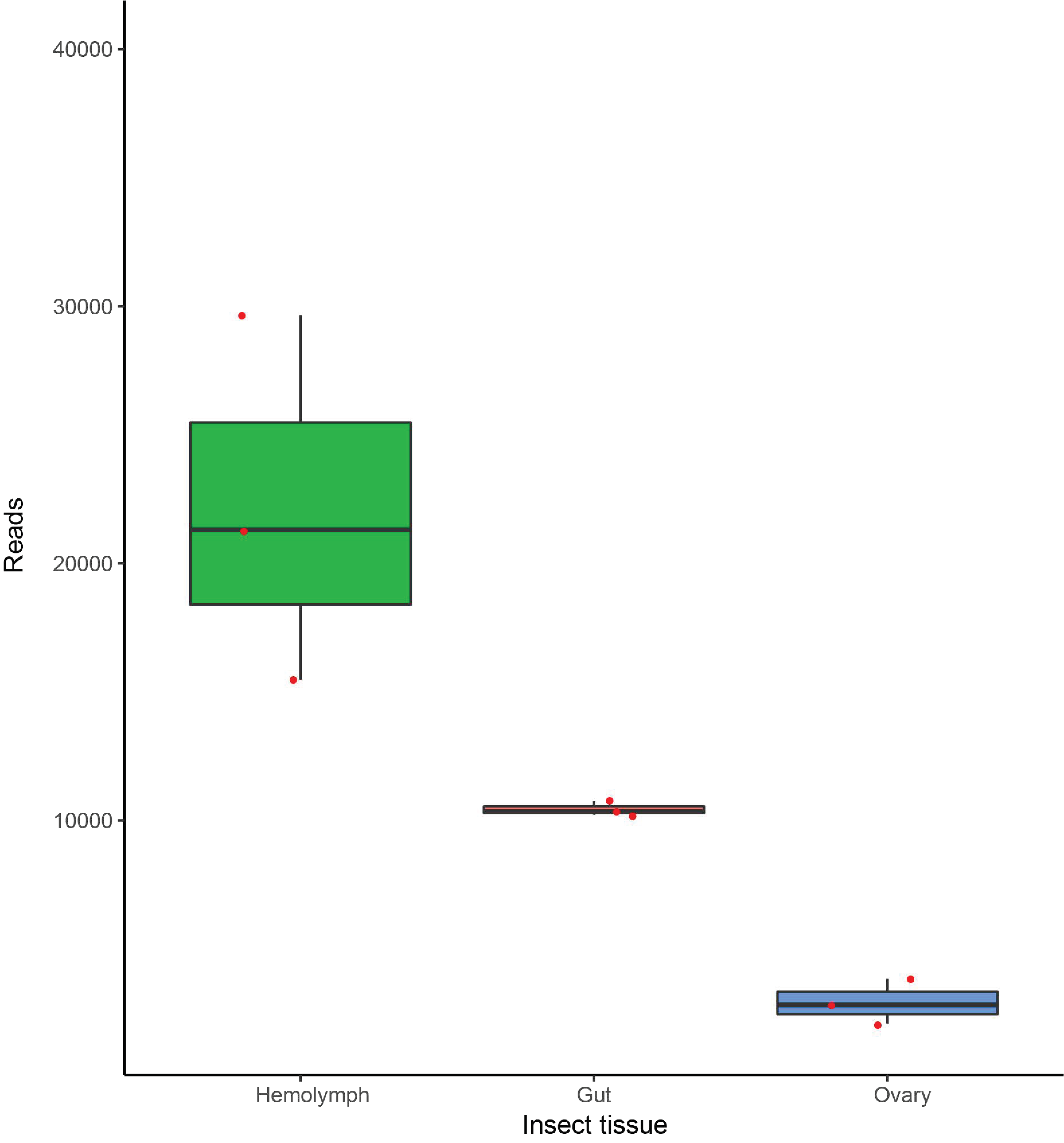
3.3. Dactylopiibacterium Quantitative Transcript Analysis
4. Discussion
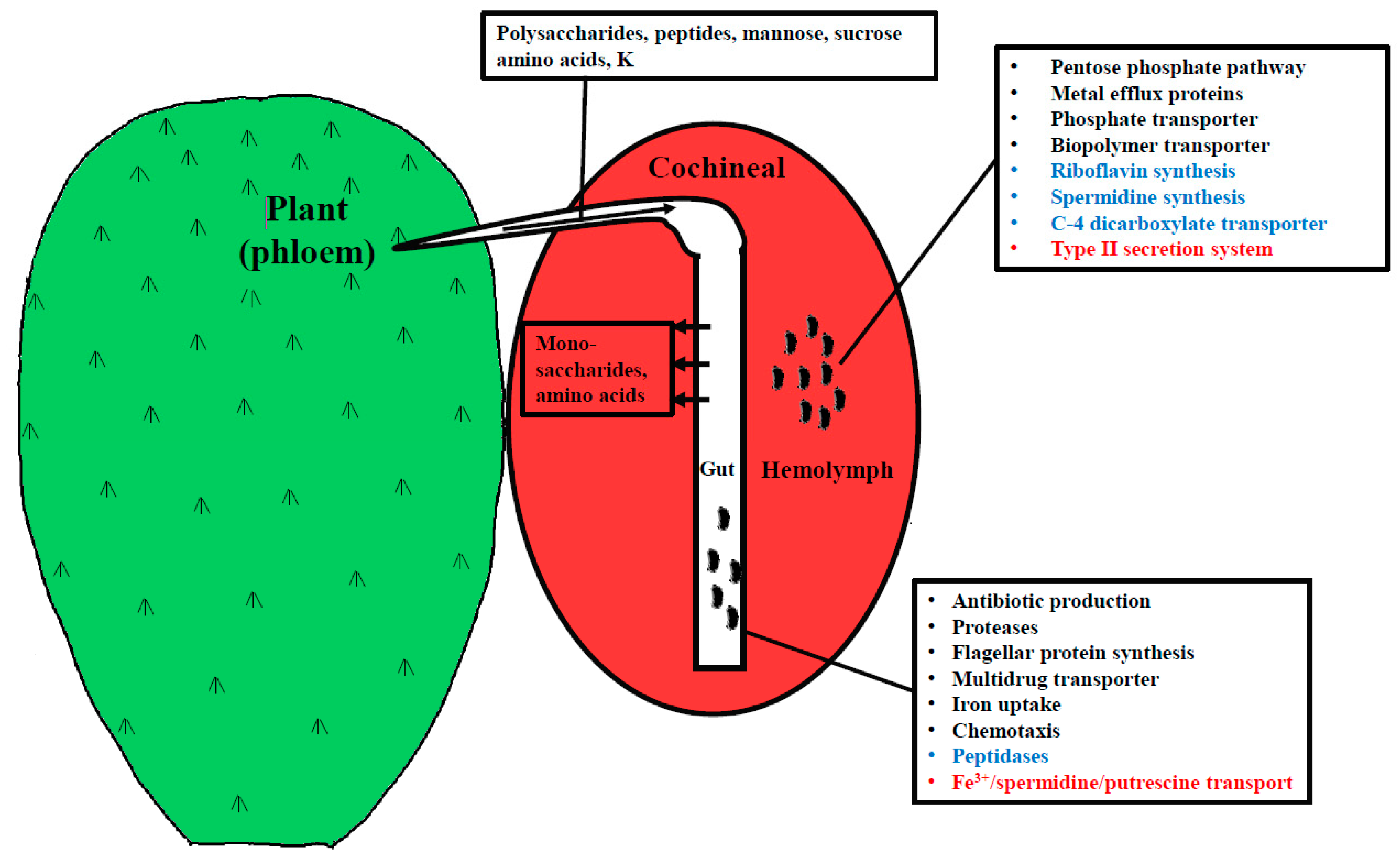
4.1. Genes Expressed in the Hemolymph and Gut
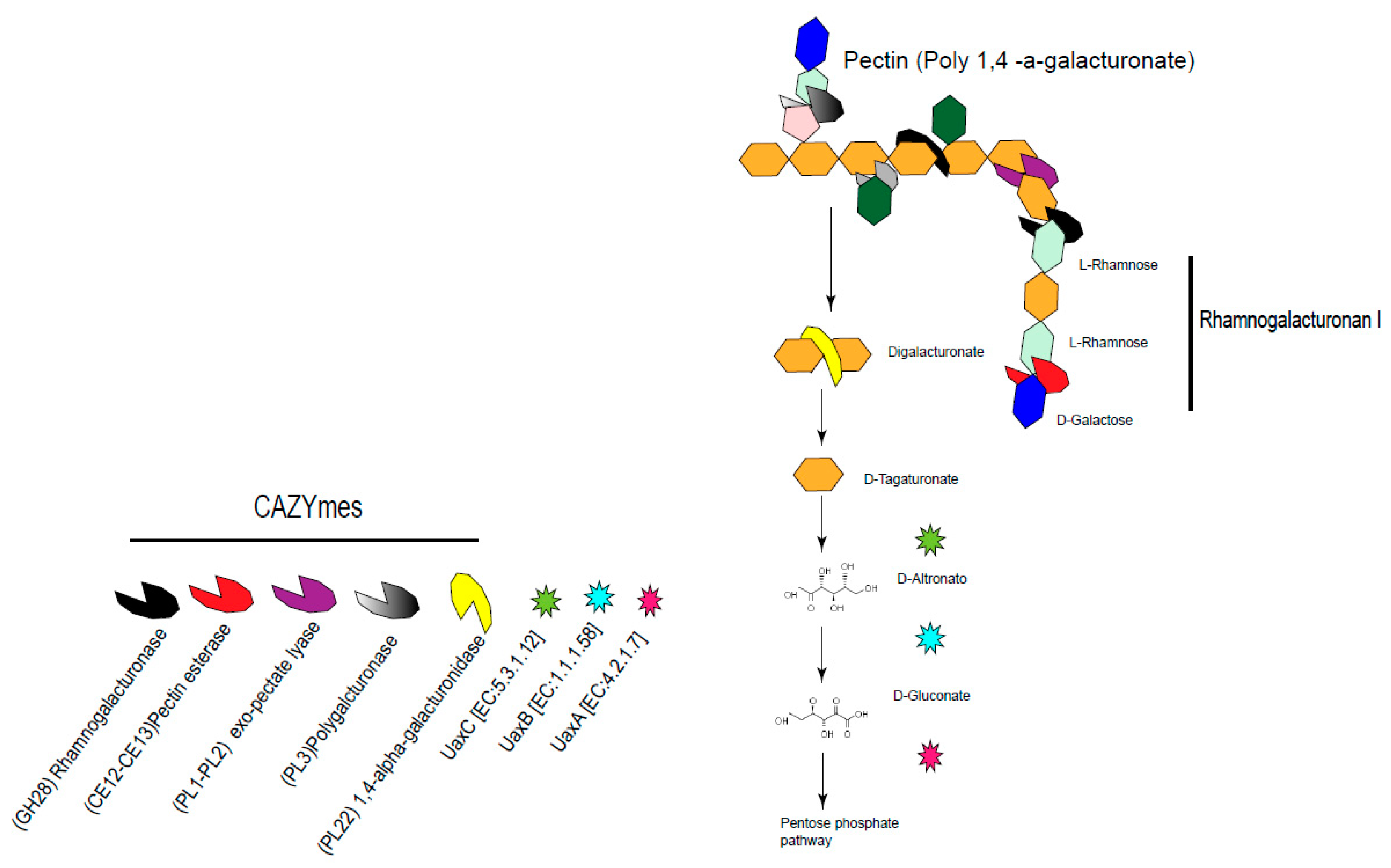
4.2. Up-Regulated Genes in the Gut
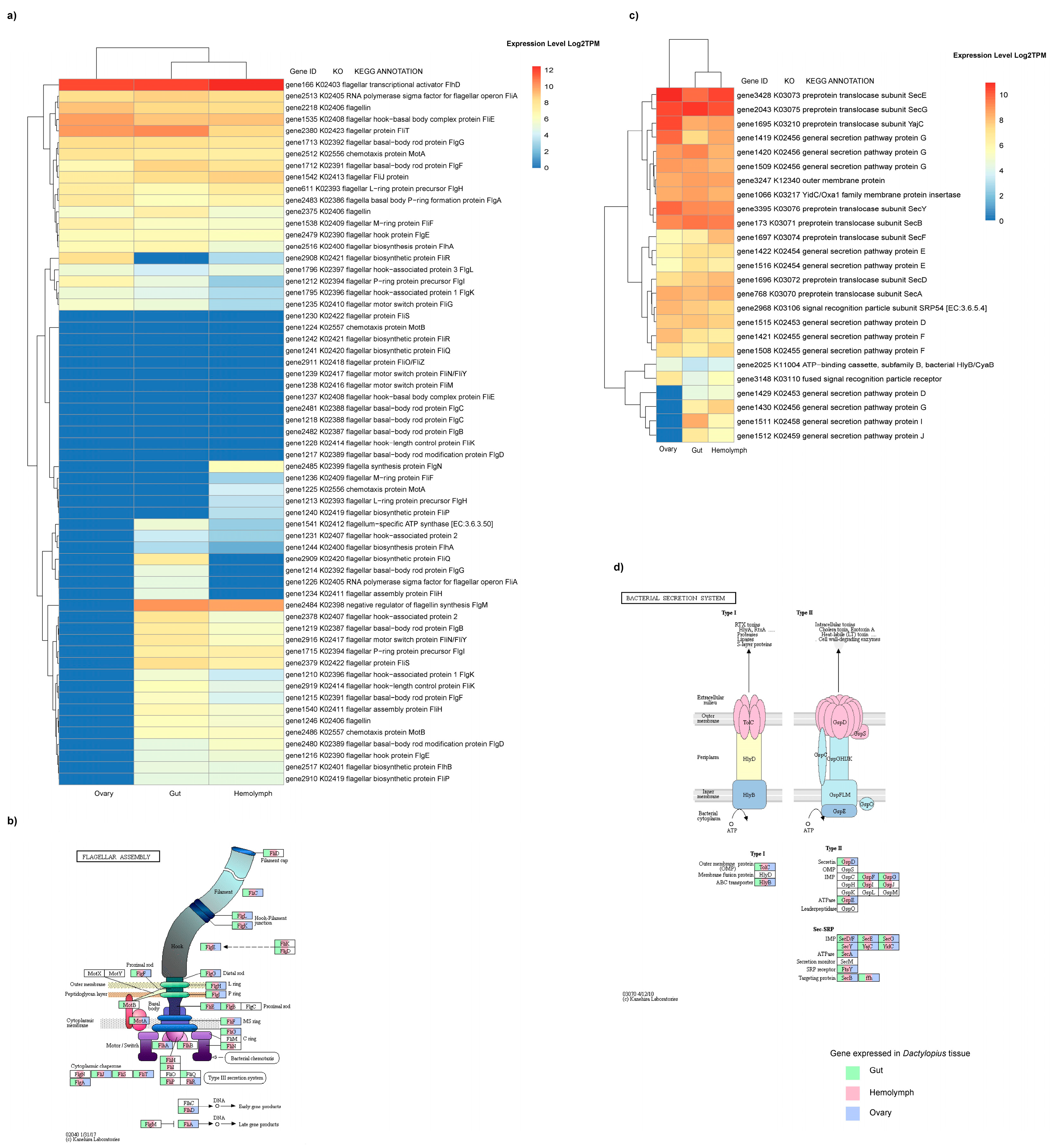
4.3. Differentially Expressed Genes in Hemolymph
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gullan, P.J.; Kosztarab, M. Adaptations in scale insects. Annu. Rev. Entomol. 1997, 42, 23–50. [Google Scholar] [CrossRef]
- Rosenblueth, M.; Martínez-Romero, J.; Ramírez-Puebla, S.T.; Vera-Ponce de León, A.; Rosas-Pérez, T.; Bustamante-Brito, R.; Rincón-Rosales, R.; Martínez-Romero, E. Endosymbiotic microorganisms of scale insects. TIP Revista Especializada en Ciencias Químico-Biológicas 2018, 21, 53–69. [Google Scholar] [CrossRef]
- Campana, M.G.; Robles García, N.M.; Tuross, N. America’s red gold: Multiple lineages of cultivated cochineal in Mexico. Ecol. Evol. 2015, 5, 607–617. [Google Scholar] [CrossRef]
- Eisner, T.; Nowicki, S.; Goetz, M.; Meinwald, J. Red cochineal dye (carminic acid): Its role in nature. Science 1980, 208, 1039–1042. [Google Scholar] [CrossRef] [PubMed]
- Morrison, J.F. Protection from Beetle-Predation in Cochineal Insects (Dactylopiidae: Homoptera). Master’s Thesis, Master of Science, Rhodes University, Grahamstown, South Africa, January 1984. [Google Scholar]
- Chávez-Moreno, C.K.; Tecante, A.; Casas, A. The Opuntia (Cactaceae) and Dactylopius (Hemiptera: Dactylopiidae) in Mexico: A historical perspective of use, interaction and distribution. Biodivers. Conserv. 2009, 18, 3337–3355. [Google Scholar] [CrossRef]
- Portillo, L. Origen de Dactylopius coccus Costa (Hemiptera: Dactylopiidae): Norte o Sudamérica? Dugesiana 2005, 12, 1–8. [Google Scholar]
- Van Dam, A.R.; Portillo Martinez, L.; Chavez, A.J.; May, B.P. Range Wide Phylogeography of Dactylopius coccus (Hemiptera: Dactylopiidae). Ann. Entomol. Soc. Am. 2015, 108, 299–310. [Google Scholar] [CrossRef]
- Chávez-Moreno, C.K.; Tecante, A.; Fragoso-Serrano, M.; Pereda-Miranda, R. Metabolic profiling of Dactylopius (Hemiptera: Dactylopiidae) species pigments by geographical origin and hosts using multivariate data analysis. Biochem. Syst. Ecol. 2010, 38, 671–679. [Google Scholar] [CrossRef]
- Wang, N.; Nobel, P.S. Phloem exudate collected via scale insect stylets for the CAM species Opuntia ficus-indica under current and doubled CO2 concentrations. Ann. Bot. 1995, 75, 525–532. [Google Scholar] [CrossRef]
- Lough, T.J.; Lucas, W.J. Integrative plant biology: Role of phloem long-distance macromolecular trafficking. Annu. Rev. Plant Biol. 2006, 57, 203–232. [Google Scholar] [CrossRef]
- Aki, T.; Shigyo, M.; Nakano, R.; Yoneyama, T.; Yanagisawa, S. Nano scale proteomics revealed the presence of regulatory proteins including three FT-Like proteins in phloem and xylem saps from rice. Plant Cell Physiol. 2008, 49, 767–790. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Sun, L.; Kragler, F. The phloem-delivered RNA pool contains small noncoding RNAs and interferes with translation. Plant Physiol. 2009, 150, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Medina, C.; Atkins, C.A.; Mann, A.J.; Jordan, M.E.; Smith, P.M. Macromolecular composition of phloem exudate from white lupin (Lupinus albus L.). BMC Plant Biol. 2011, 11, 36. [Google Scholar] [CrossRef]
- Rodriguez, L.C.; Niemeyer, H.M. Cochineal production: A reviving Precolumbian industry. Athena Rev. 2001, 2, 76–78. [Google Scholar]
- Ramírez-Puebla, S.T.; Ormeño-Orrillo, E.; Vera-Ponce de León, A.; Lozano, L.; Sánchez, A.; Rosenblueth, M.; Martínez-Romero, E. Genomes of Candidatus Wolbachia bourtzisii wDacA and Candidatus Wolbachia pipientis wDacB from the Cochineal insect Dactylopius coccus (Hemiptera: Dactylopiidae). G3 2016, 6, 3343–3349. [Google Scholar] [CrossRef] [PubMed]
- Vera-Ponce de León, A.; Ormeño-Orrillo, E.; Ramírez-Puebla, S.T.; González-Román, P.; Rosenblueth, M.; Degli Esposti, M.; Martinez, J.; Martínez-Romero, E. Candidatus Dactylopiibacterium carminicum, a nitrogen-fixing symbiont of the cochineal insect Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae). Genome Biol. Evol. 2017, 9, 2237–2250. [Google Scholar] [CrossRef]
- Vera-Ponce de León, A.; Sanchez-Flores, A.; Rosenblueth, M.; Martínez-Romero, E. Fungal community associated with Dactylopius (Hemiptera: Coccoidea: Dactylopiidae) and its role in uric acid metabolism. Front. Microbiol. 2016, 7, 1–15. [Google Scholar] [CrossRef]
- Ramírez-Puebla, S.T.; Rosenblueth, M.; Chávez-Moreno, C.K.; de Lyra, M.C.; Tecante, A.; Martínez-Romero, E. Molecular phylogeny of the genus Dactylopius (Hemiptera: Dactylopiidae) and identification of the symbiotic bacteria. Environ. Entomol. 2010, 39, 1178–1183. [Google Scholar] [CrossRef]
- Hurek, T.; Reinhold-Hurek, B. Azoarcus sp. strain BH72 as a model for nitrogen-fixing grass endophytes. J. Biotechnol. 2003, 106, 169–178. [Google Scholar] [CrossRef] [PubMed]
- van Borm, S.; Buschinger, A.; Boomsma, J.J.; Billen, J. Tetraponera ants have gut symbionts related to nitrogen-fixing root-nodule bacteria. Proc. Biol. Sci. 2002, 269, 2023–2027. [Google Scholar] [CrossRef]
- Behar, A.; Yuval, B.; Jurkevitch, E. Enterobacteria-mediated nitrogen fixation in natural populations of the fruit fly Ceratitis capitata. Mol. Ecol. 2005, 14, 2637–2643. [Google Scholar] [CrossRef]
- Pinto-Tomás, A.A.; Anderson, M.A.; Suen, G.; Stevenson, D.M.; Chu, F.S.; Cleland, W.W.; Weimer, P.J.; Currie, C.R. Symbiotic nitrogen fixation in the fungus gardens of leaf-cutter ants. Science 2009, 326, 1120–1123. [Google Scholar] [CrossRef] [PubMed]
- Morales-Jiménez, J.; Zúñiga, G.; Villa-Tanaca, L.; Hernández-Rodríguez, C. Bacterial community and nitrogen fixation in the red turpentine beetle, Dendroctonus valens LeConte (Coleoptera: Curculionidae: Scolytinae). Microb. Ecol. 2009, 58, 879–891. [Google Scholar] [CrossRef] [PubMed]
- Morales-Jiménez, J.; Vera-Ponce de León, A.; García-Domínguez, A.; Martínez-Romero, E.; Zúñiga, G.; Hernández-Rodríguez, C. Nitrogen-fixing and uricolytic bacteria associated with the gut of Dendroctonus rhizophagus and Dendroctonus valens (Curculionidae: Scolytinae). Microb. Ecol. 2013, 66, 200–210. [Google Scholar] [CrossRef] [PubMed]
- Desai, M.S.; Brune, A. Bacteroidales ectosymbionts of gut flagellates shape the nitrogen-fixing community in dry-wood termites. ISME J. 2012, 6, 1302–1313. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Castro, J.; Lozano, L.; Sohlenkamp, C. Dissectin the Acid Stress Response of Rhizobium tropici CIAT 899. Front. Microbiol. 2018, 9, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast grapped-read alignment with Bowtie2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.; Harris, S.R.; Berriman, M.; Parkhill, J.; McQuillan, J.A. Artemis: An integrated plataform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 2012, 28, 464–469. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Tarazona, S.; Furió-Tarí, P.; Turrá, D.; Di Pietro, A.; Nueda, M.J.; Ferrer, A.; Conesa, A. Data quality aware analysis of differential expression in RNA-seq with NOISeq R/Bioc package. Nucleic Acids Res. 2015, 43, e140. [Google Scholar] [CrossRef]
- Yamada, T.; Letunic, I.; Okuda, S.; Kanehisa, M.; Bork, P. iPath2.0: Interactive pathway explorer. Nucleic Acids Res. 2011, 39, W412–W415. [Google Scholar] [CrossRef] [PubMed]
- Lahortiga, I.; Cox, L. somersault18:24. Grote Markt, Belgium. 2018. Available online: http://www.somersault1824.com/resources/ (accessed on 8 August 2018).
- Kanehisa, M.; Sato, Y.; Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 2016, 428, 726–731. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Mao, X.; Yang, J.; Chen, X.; Mao, F.; Xu, Y. DbCAN: A web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2012, 40, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Dressaire, C.; Moreira, R.N.; Barahona, S.; Alves de Matos, A.P.; Arraiano, C.M. BolA is a transcriptional switch that turns off motility and turns on biofilm development. MBio 2015, 6, e02352-14. [Google Scholar] [CrossRef] [PubMed]
- Soneson, C.; Delorenzi, M. A comparison of methods for differential expression analysis of RNA-seq data. BMC Bioinform. 2013, 14, 91. [Google Scholar] [CrossRef] [PubMed]
- Costa-Silva, J.; Domingues, D.; Martins Lopes, F. RNA-Seq differential expression analysis: An extendes review and a software tool. PLoS ONE 2017, 12, e0190152. [Google Scholar] [CrossRef] [PubMed]
- Goethals, K.; Van Montagu, M.; Holsters, M. Conserved motifs in a divergent nod box of Azorhizobium caulinodans ORS571 reveal a common structure in promoters regulated by LysR-type proteins. Proc. Natl. Acad. Sci. USA 1992, 89, 1646–1650. [Google Scholar] [CrossRef]
- Schlaman, H.R.; Okker, R.J.; Lugtenberg, B.J. Regulation of nodulation gene expression by NodD in rhizobia. J. Bacteriol. 1992, 174, 5177–5182. [Google Scholar] [CrossRef] [PubMed]
- Bernier, S.P.; Nguyen, D.T.; Sokol, P.A. A LysR-type transcriptional regulator in Burkholderia cenocepacia influences colony morphology and virulence. Infect. Immun. 2008, 76, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Maddocks, S.E.; Oyston, P.C. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 2008, 154, 3609–3623. [Google Scholar] [CrossRef] [PubMed]
- Balakrishnan, L.; Venter, H.; Shilling, R.A.; Veen, H.W. Reversible transport by the ATP-binding cassette multidrug export pump LmrA: ATP synthesis at the expense of downhill ethidium uptake. J. Biol. Chem. 2004, 279, 11273–11280. [Google Scholar] [CrossRef] [PubMed]
- Charles, H.; Heddi, A.; Guillaud, J.; Nardon, C.; Nardon, P. A molecular aspect of symbiotic interactions between the weevil Sitophilus oryzae and its endosymbiotic bacteria: Over-expression of a chaperonin. Biochem. Biophys. Res. Commun. 1997, 239, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Fares, M.A.; Moya, A.; Barrio, E. GroEL and the maintenance of bacterial endosymbiosis. TRENDS Genet. 2004, 20, 413–416. [Google Scholar] [CrossRef]
- Poliakov, A.; Russell, C.W.; Ponnala, L.; Hoops, H.J.; Sun, Q.; Douglas, A.E.; van Wijk, K.J. Large-scale label-free quantitative proteomics of the pea aphid-Buchnera symbiosis. Mol. Cell Proteomics 2011, 10, M110.007039. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Thompson, J.W.; Dubois, L.G.; Moseley, M.A.; Wernegreen, J.J. Proteomic analysis of an unculturable bacterial endosymbiont (Blochmannia) reveals high abundance of chaperonins and biosynthetic enzymes. J. Proteome Res. 2013, 12, 704–718. [Google Scholar] [CrossRef] [PubMed]
- Kühner, S.; van Noort, V.; Betts, M.J.; Leo-Macias, A.; Batisse, C.; Rode, M.; Yamada, T.; Maier, T.; Bader, S.; Beltran-Alvarez, P.; et al. Proteome organization in a genome-reduced bacterium. Science 2009, 326, 1235–1240. [Google Scholar] [CrossRef] [PubMed]
- Kupper, M.; Gupta, S.K.; Feldhaar, H.; Gross, R. Versatile roles of the chaperonin GroEL in microorganism-insect interactions. FEMS Microbiol. Lett. 2014, 353, 1–10. [Google Scholar] [CrossRef]
- López-Guerrero, M.G.; Ormeño-Orrillo, E.; Rosenblueth, M.; Martínez-Romero, J.; Martínez-Romero, E. Buffet hypothesis for microbial nutrition at the rhizosphere. Front. Plant Sci. 2013, 4, 188. [Google Scholar] [CrossRef] [PubMed]
- Degli Esposti, M.; Martínez-Romero, E. The functional microbiome of arthropods. PLoS ONE 2017, 12, e0176573. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.E.; Reinhold-Hurek, B. Distinct roles of P(II)-like signal transmitter proteins and amtB in regulation of nif gene expression, nitrogenase activity, and posttranslational modification of NifH in Azoarcus sp. strain BH72. J. Bacteriol. 2002, 184, 2251–2259. [Google Scholar] [CrossRef] [PubMed]
- Sickerman, N.S.; Rettberg, L.A.; Lee, C.C.; Hu, Y.; Ribbe, M.W. Cluster assembly in nitrogenase. Essays Biochem. 2017, 61, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Kaminski, P.A.; Kitts, CL.; Zimmerman, Z.; Ludwig, R.A. Azorhizobium caulinodans uses both cytochrome bd (quinol) and cytochrome cbb3 (cytochrome c) terminal oxidases for symbiotic N2 fixation. J. Bacteriol. 1996, 178, 5989–5994. [Google Scholar] [CrossRef] [PubMed]
- Dinant, S.; Bonnemain, J.L.; Girousse, C.; Kehr, J. Phloem sap intricacy and interplay with aphid feeding. C. R. Biol. 2010, 333, 504–515. [Google Scholar] [CrossRef] [PubMed]
- Gaille, C.; Reimmann, C.; Haas, D. Isochorismate synthase (PchA), the first and rate-limiting enzyme in salicylate biosynthesis of Pseudomonas aeruginosa. J. Biol. Chem. 2003, 278, 16893–16898. [Google Scholar] [CrossRef] [PubMed]
- Skaar, E.P. The Battle for Iron between Bacterial Pathogens and Their Vertebrate Hosts. PLoS Pathog. 2010, 6, e1000949. [Google Scholar] [CrossRef] [PubMed]
- Barka, N.; Abdennouri, M.; El Makhfouk, M.; Qourzal, S. Biosorption characteristics of cadmium and lead onto eco-friendly dried cactus (Opuntia ficus indica) cladodes. J. Environ. Chem. Eng. 2013, 1, 144–149. [Google Scholar] [CrossRef]
- Nharingo, T.; Moyo, M. Application of Opuntia ficus-indica in bioremediation of wastewaters. A critical review. J. Environ. Manag. 2016, 166, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Domozych, D.S.; Sørensen, I.; Popper, Z.A.; Ochs, J.; Andreas, A.; Fangel, J.U.; Pielach, A.; Sacks, C.; Brechka, H.; Ruisi-Besares, P.; et al. Pectin Metabolism and Assembly in the Cell Wall of the Charophyte Green Alga Penium margaritaceum. Plant Physiol. 2014, 165, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Yapo, B.M. Rhamnogalacturonan-I: A structurally puzzling and functionally versatile polysaccharide from plant cell walls and mucilages. Polym. Rev. 2011, 51, 391–413. [Google Scholar] [CrossRef]
- Azadi, P.; O’Neill, M.A.; Bergmann, C.; Darvill, A.G.; Albersheim, P. The backbone of the pectic polysaccharide rhamnogalacturonan I is cleaved by an endohydrolase and an endolyase. Glycobiology 1995, 5, 783–789. [Google Scholar] [CrossRef] [PubMed]
- Lefsih, K.; Giacomazza, D.; Passantino, R.; Costa, M.A.; Bulone, D.; Mangione, M.R.; Guarrasi, V.; Mingoia, F.; San Biagio, P.L.; Madani, K. Biochemical and biophysical characterization of water-soluble pectin from Opuntia ficus-indica and its potential cytotoxic activity. Phytochemistry 2018, 154, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Ndeh, D.; Rogowski, A.; Cartmell, A.; Luis, A.S.; Baslé, A.; Gray, J.; Venditto, I.; Briggs, J.; Zhang, X.; Labourel, A.; et al. Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature 2017, 544, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.; Martinson, V.G.; Moran, N.A. Functional diversity within the simple gut microbiota of the honey bee. Proc. Natl. Acad. Sci. USA 2012, 109, 11002–11007. [Google Scholar] [CrossRef] [PubMed]
- James, V.; Hugouvieux-Cotte-Pattat, N. Regulatory systems modulating the transcription of the pectinase genes of Erwinia chrysanthemi are conserved in Escherichia coli. Microbiology 1996, 142, 2613–2619. [Google Scholar] [CrossRef]
- Reinhold-Hurek, B.; Hurek, T.; Claeyssens, M.; van Montagu, M. Cloning, expression in Escherichia coli, and characterization of cellulolytic enzymes of Azoarcus sp., a root-invading diazotroph. J. Bacteriol. 1993, 175, 7056–7065. [Google Scholar] [CrossRef] [PubMed]
- Garg, N.; Singla, R. Nitrogen fixation and carbon metabolism in legume nodules. Indian J. Exp. Biol. 2004, 42, 138–142. [Google Scholar] [PubMed]
- Iyer, B.; Rajput, M.S.; Jog, R.; Joshi, E.; Bharwad, K.; Rajkumar, S. Organic acid mediated repression of sugar utilization in rhizobia. Microbiol. Res. 2016, 192, 211–220. [Google Scholar] [CrossRef]
- Bursell, E. The Role of Proline in Energy Metabolism. In Energy Metabolism in Insects; Downer, R.G.H., Ed.; Plenum Press: New York, NY, USA, 1980; pp. 135–154. [Google Scholar]
- Phillips, D.A.; Joseph, C.M.; Yang, G.P.; Martinez-Romero, E.; Sanborn, J.R.; Volpin, H. Identification of lumichrome as a sinorhizobium enhancer of alfalfa root respiration and shoot growth. Proc. Natl. Acad. Sci. USA 1999, 96, 12275–12280. [Google Scholar] [CrossRef]
- Phillips, D.A.; Martinez-Romero, E.; Yang, G.P.; Joseph, C.M. Release of nitrogen: A key trait in selecting bacterial endophytes for agronomically useful nitrogen fixation. In The Quest for Nitrogen Fixation in Rice; Ladha, J.K., Reddy, P.M., Eds.; IRRI: Laguna, Philippines, 2000; pp. 205–217. [Google Scholar]
- Ramírez-Puebla, S.T.; Servín-Garcidueñas, L.E.; Jiménez-Marín, B.; Bolaños, L.M.; Rosenblueth, M.; Martínez, J.; Rogel, M.A.; Ormeño-Orrillo, E.; Martínez-Romero, E. Gut and root microbiota commonalities. Appl. Environ. Microbiol. 2013, 79, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Hümbelin, M.; Griesser, V.; Keller, T.; Schurter, W.; Haiker, M.; Hohmann, H.P.; Ritz, H.; Richter Bacher, G.A.; van Loon, A.P.G.M. GTP cyclohydrolase II and 3,4-dihydroxy-2-butanone 4-phosphate synthase are rate-limiting enzymes in riboflavin synthesis of an industrial Bacillus subtilis strain used for riboflavin production. J. Ind. Microbiol. Biotechnol. 1999, 22, 1. [Google Scholar] [CrossRef]
- Draghi, W.O.; Del Papa, M.F.; Hellweg, C.; Watt, S.A.; Watt, T.F.; Barsch, A.; Lozano, M.J.; Lagares, A., Jr.; Salas, M.E.; López, J.L.; et al. A consolidated analysis of the physiologic and molecular responses induced under acid stress in the legume-symbiont model-soil bacterium Sinorhizobium meliloti. Sci. Rep. 2016, 6, 29278. [Google Scholar] [CrossRef] [PubMed]
- Rosas-Pérez, T.; Vera-Ponce de León, A.; Rosenblueth, M.; Ramírez-Puebla, S.T.; Rincón-Rosales, R.; Martínez-Romero, J.; Dunn, M.F.; Kondorosi, E.; Martínez-Romero, E. Chapter 5. The Symbiome of Llaveia cochineals (Hemiptera: Coccoidea: Monophlebidae) includes a Gammaproteobacterial cosymbiont Sodalis TME1 and the known Candidatus Walczuchella monophlebidarum. In Agricultural and Biological Sciences "Insect Physiology and Ecology"; Shields, V.D.C., Ed.; InTech: Pleasanton, CA, USA, 2017; pp. 115–134. [Google Scholar]
- Jelsbak, L.; Thomsen, L.E.; Wallrodt, I.; Jensen, P.R.; Olsen, J.E. Polyamines are required for virulence in Salmonella enterica serovar Typhimurium. PLoS ONE 2012, 7, e36149. [Google Scholar] [CrossRef] [PubMed]
- Böhm, M.; Hurek, T.; Reinhold-Hurek, B. Twitching motility is essential for endophytic rice colonization by the N2-fixing endophyte Azoarcus sp. strain BH72. Mol. Plant Microbe Interact. 2007, 20, 526–533. [Google Scholar] [CrossRef] [PubMed]
- Pelicic, V. Type IV pili: E pluribus unum? Mol. Microbiol. 2008, 68, 827–837. [Google Scholar] [CrossRef] [PubMed]
- Powell, J.E.; Leonard, S.P.; Kwong, W.K.; Engel, P.; Moran, N.A. Genome-wide screen identifies host colonization determinants in a bacterial gut symbiont. Proc. Natl. Acad. Sci. USA 2016, 113, 13887–13892. [Google Scholar] [CrossRef]
- Cianciotto, N.P.; White, R.C. Expanding role of type II secretion in bacterial pathogenesis and beyond. Infect Immun. 2017, 85, e00014-17. [Google Scholar] [CrossRef]
- Wong, E.; Vaaje-Kolstad, G.; Ghosh, A.; Hurtado-Guerrero, R.; Konarev, P.V.; Ibrahim, A.F.; Svergun, D.I.; Eijsink, V.G.; Chatterjee, N.S.; van Aalten, D.M. The Vibrio cholerae colonization factor GbpA possesses a modular structure that governs binding to different host surfaces. PLoS Pathog. 2012, 8, e1002373. [Google Scholar] [CrossRef]
- Baldi, D.; Higginson, E.; Hocking, D.; Praszkier, J.; Cavaliere, R.; James, C.; Bennett-Wood, V.; Azzopardi, K.; Turnbull, L.; Lithgow, T.; et al. The type II secretion system and its ubiquitous lipoprotein substrate, SslE, are required for biofilm formation and virulence of enteropathogenic Escherichia coli. Infect. Immun. 2012, 80, 2042–2052. [Google Scholar] [CrossRef]
- Lee, D.J.; Wong, B.T.; Adav, S.S. Azoarcus taiwanensis sp. nov., a denitrifying species isolated from a hot spring. Appl. Microbiol. Biotechnol. 2014, 98, 1301–1307. [Google Scholar] [CrossRef]
- Macnab, R.M. How bacteria assemble flagella. Annu Rev Microbiol. 2003, 57, 77–100. [Google Scholar] [CrossRef]
- Sano, G.; Takada, Y.; Goto, S.; Maruyama, K.; Shindo, Y.; Oka, K.; Matsui, H.; Matsuo, K. Flagella facilitate escape of Salmonella from oncotic macrophages. J. Bacteriol. 2007, 189, 8224–8232. [Google Scholar] [CrossRef] [PubMed]
- Masson, F.; Zaidman-Rémy, A.; Heddi, A. Antimicrobial peptides and cell processes tracking endosymbiont dynamics. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2016, 371, 20150298. [Google Scholar] [CrossRef] [PubMed]
- Vigneron, A.; Masson, F.; Vallier, A.; Balmand, S.; Rey, M.; Vincent-Monégat, C.; Aksoy, E.; Aubailly-Giraud, E.; Zaidman-Rémy, A.; Heddi, A. Insects recycle endosymbionts when the benefit is over. Curr. Biol. 2014, 24, 2267–2273. [Google Scholar] [CrossRef] [PubMed]
- Barraquio, W.L.; Revilla, L.; Ladha, J.K. Isolation of endophytic diazotrophic bacteria from wetland rice. Plant Soil 1997, 194, 15–24. [Google Scholar] [CrossRef]
- Martínez, L.; Caballero, J.; Orozco, J.; Martínez-Romero, E. Diazotrophic bacteria associated with banana (Musa spp.). Plant Soil 2003, 257, 35–47. [Google Scholar] [CrossRef]

| Protein ID | Log2FC | COG | Function |
|---|---|---|---|
| Up-regulated in Gut | |||
| WP_095525391.1 | 4.10 | NA | Hypothetical protein |
| WP_095523710.1 | 3.51 | COG2081 | Aminoacetone oxidase family FAD-binding enzyme |
| WP_095525172.1 | 3.50 | COG0300 | Short-chain dehydrogenase |
| WP_095525875. | 3.17 | COG0583 | LysR family transcriptional regulator |
| WP_095523466.1 | 3.04 | COG2382 | Hypothetical protein |
| WP_095525673.1 | 3.01 | NA | Hypothetical protein |
| WP_095525264.1 | 2.98 | COG0643 | Chemotaxis protein CheA |
| WP_095524233.1 | 2.80 | NA | Hypothetical protein |
| WP_095525304.1 | 2.76 | COG2199 | Hypothetical protein |
| WP_095523625.1 | 2.67 | COG3419 | Pilus assembly protein |
| WP_095524852.1 | 2.65 | COG3895 | Hypothetical protein |
| WP_095526008.1 | 2.60 | COG3549 | Exonuclease ABC subunit A |
| WP_095523085.1 | 2.58 | COG0523 | Cobalamin biosynthesis protein CobW |
| WP_095523064.1 | 2.57 | COG0845 | Multidrug transporter subunit MdtA |
| WP_095524897.1 | 2.56 | COG2230 | SAM-dependent methylytransferase |
| Up-regulated in Hemolymph | |||
| WP_095525565.1 | 3.18 | NA | Hypothetical protein |
| WP_095525317.1 | 2.52 | COG0120 | Ribose 5-phosphate isomerase A |
| WP_095523550.1 | 2.47 | COG1702 | Phosphate starvation-inducible protein PhoH |
| WP_095525561.1 | 2.41 | NA | Hypothetical protein |
| WP_095523253.1 | 2.39 | COG0837 | Glucokinase |
| WP_095525928.1 | 2.36 | COG0171 | NAD+ synthase |
| WP_095523935.1 | 2.31 | COG1354 | Segregation/condensation protein A |
| WP_095522941. | 2.13 | COG0424 | Septum formation inhibitor Maf |
| WP_095523049.1 | 2.11 | COG2200 | Hypothetical protein |
| WP_095525579.1 | 2.11 | COG0848 | Biopolymer transporter ExbD |
| WP_095525717. | 2.09 | NA | Thioredoxin |
| WP_095526004.1 | 2.08 | NA | Hypothetical protein |
| WP_095524367.1 | 2.08 | COG1391 | Bifunctional [glutamate-amonia ligase]-Adenylyl-l-tyrosine phosphorylase/[glutamate-ammonia-ligase] adenylyltransferase |
| WP_095523236.1 | 2.06 | COG2010 | Cytochrome C |
| WP_095524731.1 | 2.05 | NA | GNAT family N-acetyltransferase |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bustamante-Brito, R.; Vera-Ponce de León, A.; Rosenblueth, M.; Martínez-Romero, J.C.; Martínez-Romero, E. Metatranscriptomic Analysis of the Bacterial Symbiont Dactylopiibacterium carminicum from the Carmine Cochineal Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae). Life 2019, 9, 4. https://doi.org/10.3390/life9010004
Bustamante-Brito R, Vera-Ponce de León A, Rosenblueth M, Martínez-Romero JC, Martínez-Romero E. Metatranscriptomic Analysis of the Bacterial Symbiont Dactylopiibacterium carminicum from the Carmine Cochineal Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae). Life. 2019; 9(1):4. https://doi.org/10.3390/life9010004
Chicago/Turabian StyleBustamante-Brito, Rafael, Arturo Vera-Ponce de León, Mónica Rosenblueth, Julio César Martínez-Romero, and Esperanza Martínez-Romero. 2019. "Metatranscriptomic Analysis of the Bacterial Symbiont Dactylopiibacterium carminicum from the Carmine Cochineal Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae)" Life 9, no. 1: 4. https://doi.org/10.3390/life9010004
APA StyleBustamante-Brito, R., Vera-Ponce de León, A., Rosenblueth, M., Martínez-Romero, J. C., & Martínez-Romero, E. (2019). Metatranscriptomic Analysis of the Bacterial Symbiont Dactylopiibacterium carminicum from the Carmine Cochineal Dactylopius coccus (Hemiptera: Coccoidea: Dactylopiidae). Life, 9(1), 4. https://doi.org/10.3390/life9010004






