Is Research on “Synthetic Cells” Moving to the Next Level?
Abstract
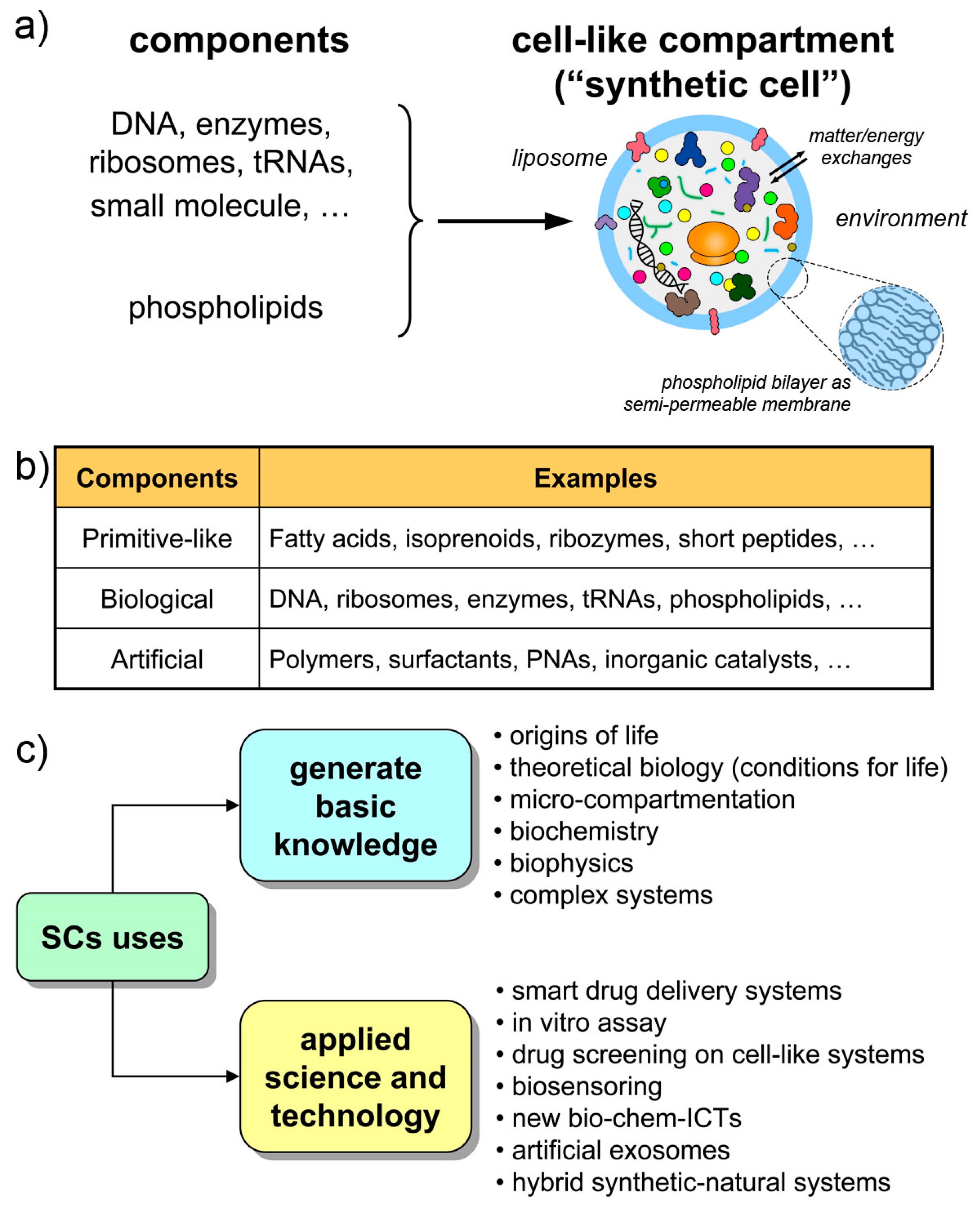
1. Introduction
2. Why and How SCs? Technologies and Theoretical Frameworks
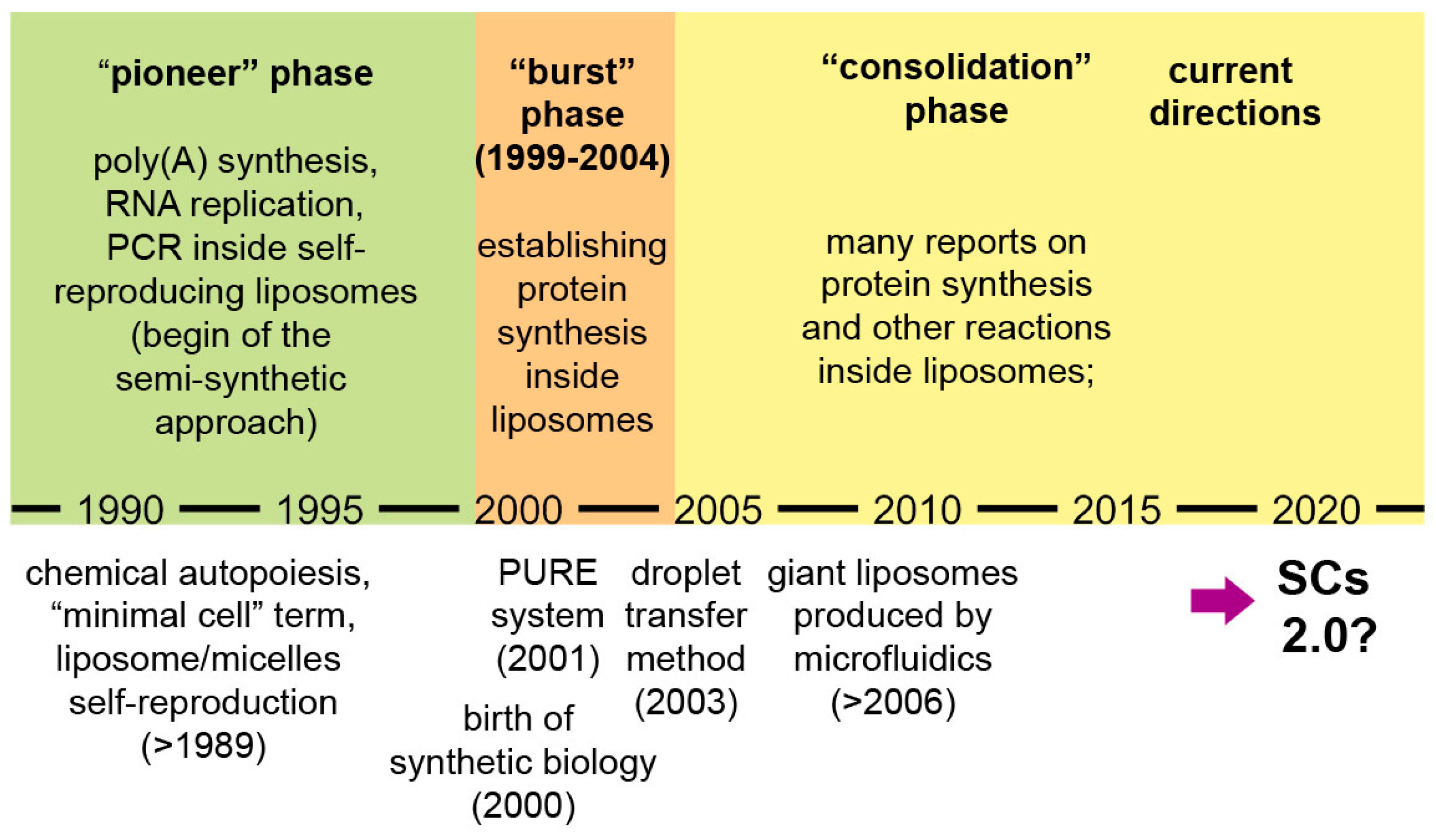
2.1. Progressing Phases of SCs Research
- a “consolidation” phase (after 2004): protein synthesis and other enzymatic reactions inside liposomes and other compartments have been studied in great detail.
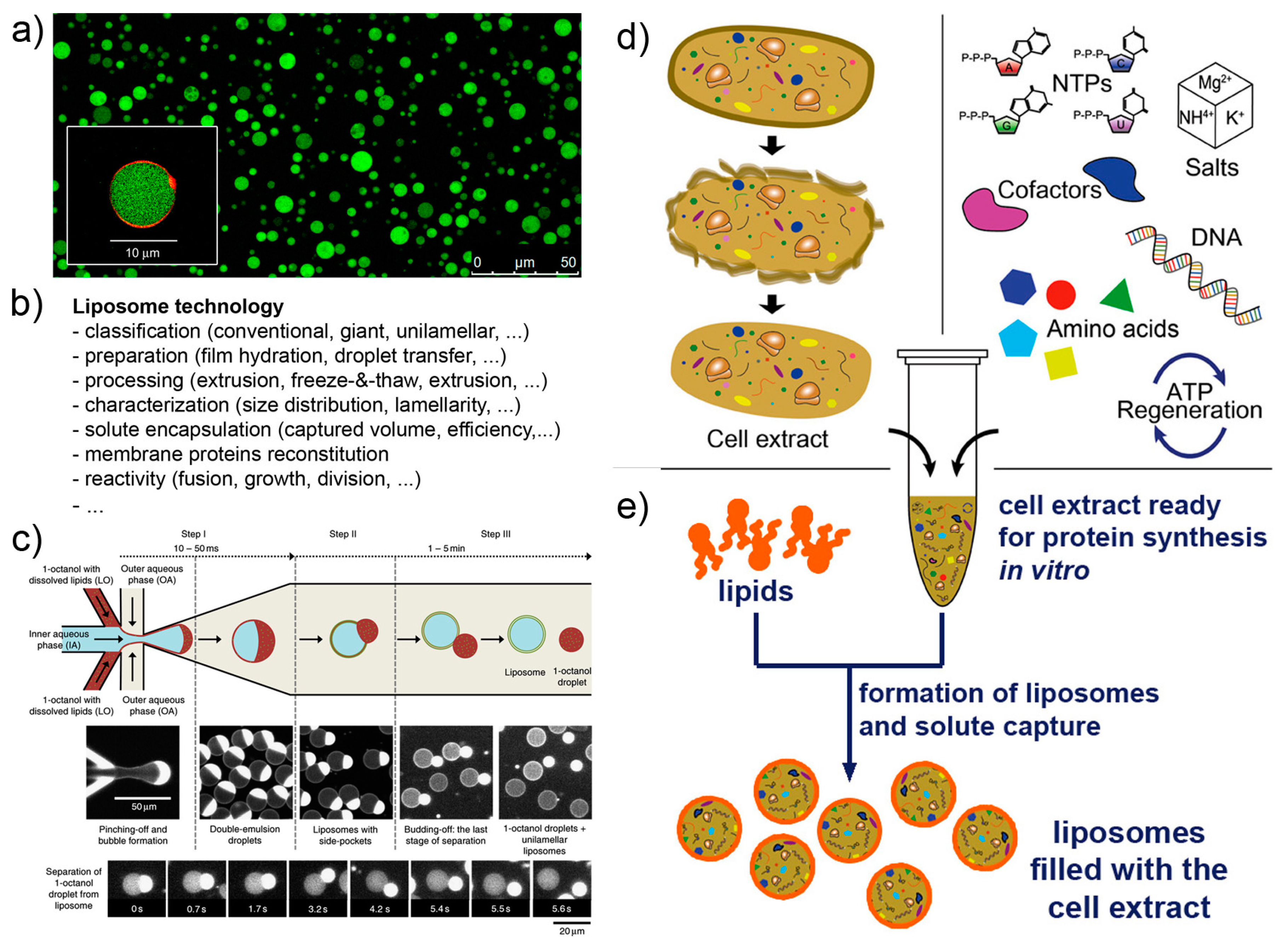
2.2. Enabling Technologies
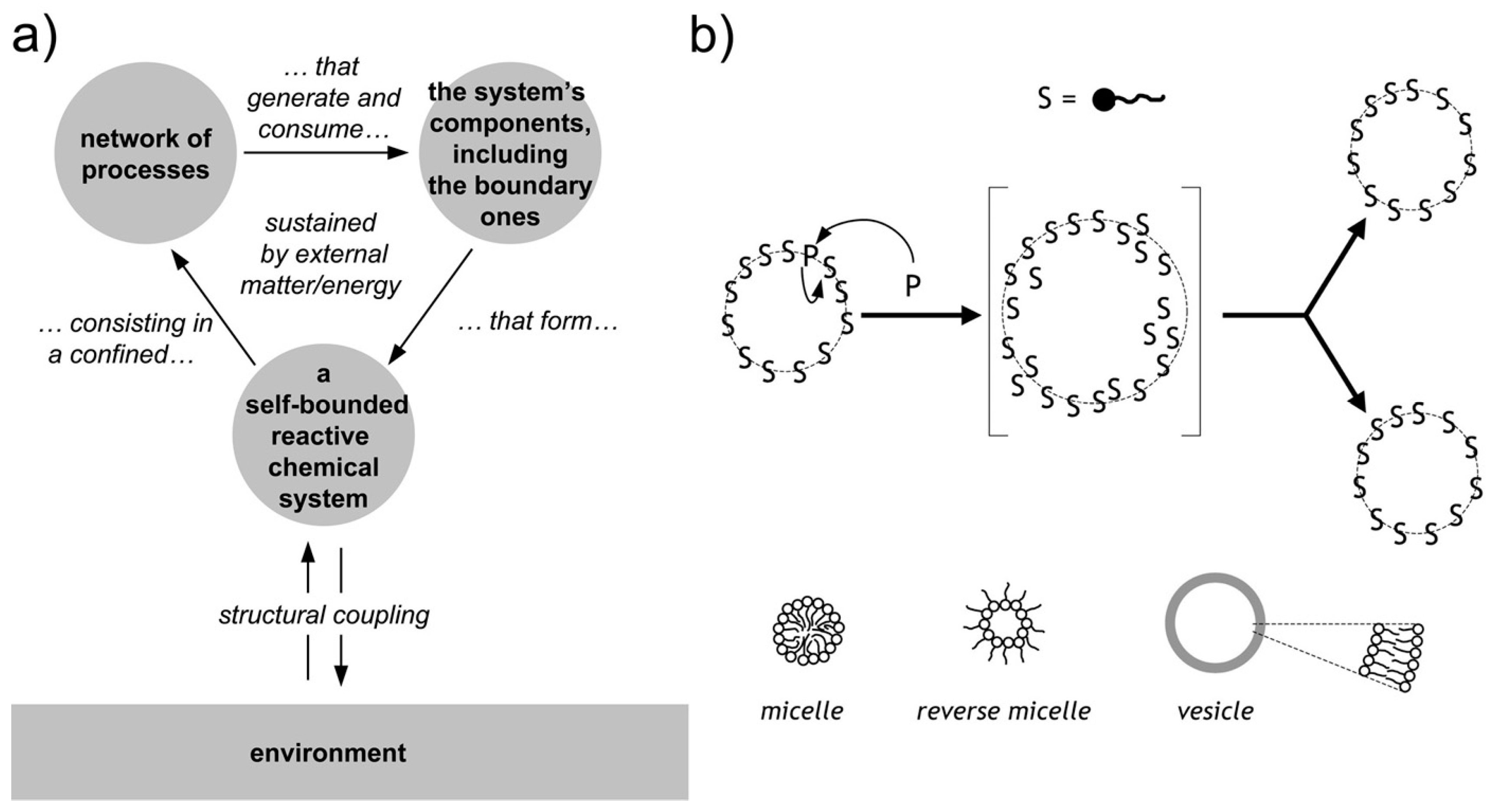
2.3. Theoretical Frameworks
3. Current Directions in SCs Research
- the functionalization of SC membrane
- the vesosome architecture
- the community perspective
- the exchange of chemical information
3.1. Functionalization of SC Membrane
3.2. The Vesosome Architecture
3.3. The Community Perspective
3.4. Exchange of Chemical Information
4. A Qualitative Jump toward SCs 2.0?
4.1. Structure
- SC ensemble/community
- individual SCs
- intra-SC compartments
4.2. Organization
5. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
Appendix A. Autopoietic SCs Based on Gene Expression?
- 36 proteins (20 amino acyl-tRNA synthetase, 10 translation factors, 4 energy-related enzymes, T7 RNA polymerase, methionine trans-formilase)
- E. coli ribosomes (composed by 3 rRNAs and 55 ribosomal proteins)
- tRNA mix from E. coli (46 according to [196]);
- small molecular weight (MW) compounds (amino acids, nucleotides, etc).
Appendix B. SC Complexity: A Still Unexplored Topic
References
- Luisi, P.L. Toward the engineering of minimal living cells. Anat. Rec. 2002, 268, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Pohorille, A.; Deamer, D. Artificial cells: Prospects for biotechnology. Trends Biotechnol. 2002, 20, 123–128. [Google Scholar] [CrossRef]
- Nomura, S.; Tsumoto, K.; Hamada, T.; Akiyoshi, K.; Nakatani, Y.; Yoshikawa, K. Gene expression within cell-sized lipid vesicles. ChemBioChem 2003, 4, 1172–1175. [Google Scholar] [CrossRef] [PubMed]
- Noireaux, V.; Libchaber, A. A vesicle bioreactor as a step toward an artificial cell assembly. Proc. Natl. Acad. Sci. USA 2004, 101, 17669–17674. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.A.; Salehi-Ashtiani, K.; Szostak, J.W. RNA catalysis in model protocell vesicles. J. Am. Chem. Soc. 2005, 127, 13213–13219. [Google Scholar] [CrossRef] [PubMed]
- Luisi, P.L.; Ferri, F.; Stano, P. Approaches to semi-synthetic minimal cells: A review. Naturwissenschaften 2006, 93, 1–13. [Google Scholar] [CrossRef]
- Mansy, S.S.; Szostak, J.W. Reconstructing the emergence of cellular life through the synthesis of model protocells. Cold Spring Harb. Symp. Quant. Biol. 2009, 74, 47–54. [Google Scholar] [CrossRef]
- Ichihashi, N.; Matsuura, T.; Kita, H.; Sunami, T.; Suzuki, H.; Yomo, T. Constructing partial models of cells. Cold Spring Harb. Perspect. Biol. 2010, 2, a004945. [Google Scholar] [CrossRef]
- Stano, P.; Carrara, P.; Kuruma, Y.; de Souza, T.P.; Luisi, P.L. Compartmentalized reactions as a case of soft-matter biotechnology: Synthesis of proteins and nucleic acids inside lipid vesicles. J. Mater. Chem. 2011, 21, 18887–18902. [Google Scholar] [CrossRef]
- Dzieciol, A.J.; Mann, S. Designs for life: Protocell models in the laboratory. Chem. Soc. Rev. 2012, 41, 79–85. [Google Scholar] [CrossRef]
- Torino, D.; Martini, L.; Mansy, S.S. Piecing Together Cell-like Systems. Curr. Org. Chem. 2013, 17, 1751–1757. [Google Scholar] [CrossRef] [PubMed]
- Nourian, Z.; Scott, A.; Danelon, C. Toward the assembly of a minimal divisome. Syst. Synth. Biol. 2014, 8, 237–247. [Google Scholar] [CrossRef] [PubMed]
- Blain, J.C.; Szostak, J.W. Progress Toward Synthetic Cells. Ann. Rev. Biochem. 2014, 83, 615–640. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, K.; Okura, Y.; Matsuo, M.; Toyota, T.; Suzuki, K.; Sugawara, T. A recursive vesicle-based model protocell with a primitive model cell cycle. Nat. Commun. 2015, 6, 8352. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Yomo, T. Constructive Approaches for Understanding the Origin of Self-Replication and Evolution. Life 2016, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Salehi-Reyhani, A.; Ces, O.; Elani, Y. Artificial cell mimics as simplified models for the study of cell biology. Exp. Biol. Med. (Maywood) 2017, 242, 1309–1317. [Google Scholar] [CrossRef] [PubMed]
- Adamala, K.P.; Martin-Alarcon, D.A.; Guthrie-Honea, K.R.; Boyden, E.S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 2017, 9, 431–439. [Google Scholar] [CrossRef] [PubMed]
- Schwille, P.; Spatz, J.; Landfester, K.; Bodenschatz, E.; Herminghaus, S.; Sourjik, V.; Erb, T.J.; Bastiaens, P.; Lipowsky, R.; Hyman, A.; et al. MaxSynBio: Avenues Towards Creating Cells from the Bottom Up. Angew. Chem. Int. Ed. Engl. 2018, 57, 13382–13392. [Google Scholar] [CrossRef]
- Göpfrich, K.; Platzman, I.; Spatz, J.P. Mastering Complexity: Towards Bottom-up Construction of Multifunctional Eukaryotic Synthetic Cells. Trends Biotechnol. 2018, 36, 938–951. [Google Scholar] [CrossRef]
- Spoelstra, W.K.; Deshpande, S.; Dekker, C. Tailoring the appearance: What will synthetic cells look like? Curr. Opin. Biotechnol. 2018, 51, 47–56. [Google Scholar] [CrossRef]
- Chandrawati, R.; Caruso, F. Biomimetic liposome- and polymersome-based multicompartmentalized assemblies. Langmuir 2012, 28, 13798–13807. [Google Scholar] [CrossRef] [PubMed]
- Brea, R.J.; Hardy, M.D.; Devaraj, N.K. Towards self-assembled hybrid artificial cells: Novel bottom-up approaches to functional synthetic membranes. Chemistry 2015, 21, 12564–12570. [Google Scholar] [CrossRef]
- Rideau, E.; Dimova, R.; Schwille, P.; Wurm, F.R.; Landfester, K. Liposomes and polymersomes: A comparative review towards cell mimicking. Chem. Soc. Rev. 2018. [Google Scholar] [CrossRef]
- Szostak, J.W.; Bartel, D.P.; Luisi, P.L. Synthesizing life. Nature 2001, 409, 387–390. [Google Scholar] [CrossRef] [PubMed]
- Deplazes, A.; Huppenbauer, M. Synthetic organisms and living machines: Positioning the products of synthetic biology at the borderline between living and non-living matter. Syst. Synth. Biol. 2009, 3, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Luisi, P.L.; Varela, F.J. Self-replicating micelles—A chemical version of a minimal autopoietic system. Orig. Life Evol. Biosph. 1989, 19, 633–643. [Google Scholar] [CrossRef]
- Bachmann, P.; Walde, P.; Luisi, P.; Lang, J. Self-replicating reverse micelles and chemical autopoiesis. J. Am. Chem. Soc. 1990, 112, 8200–8201. [Google Scholar] [CrossRef]
- Schmidli, P.K.; Schurtenberger, P.; Luisi, P.L. Liposome-mediated enzymatic synthesis of phosphatidylcholine as an approach to self-replicating liposomes. J. Am. Chem. Soc. 1991, 113, 8127–8130. [Google Scholar] [CrossRef]
- Walde, P.; Goto, A.; Monnard, P.; Wessicken, M.; Luisi, P. Oparins Reactions Revisited—Enzymatic-Synthesis of Poly(adenylic Acid). J. Am. Chem. Soc. 1994, 116, 7541–7547. [Google Scholar] [CrossRef]
- Oberholzer, T.; Wick, R.; Luisi, P.L.; Biebricher, C.K. Enzymatic RNA replication in self-reproducing vesicles: An approach to a minimal cell. Biochem. Biophys. Res. Commun. 1995, 207, 250–257. [Google Scholar] [CrossRef]
- Oberholzer, T.; Albrizio, M.; Luisi, P. Polymerase Chain-Reaction in Liposomes. Chem. Biol. 1995, 2, 677–682. [Google Scholar] [CrossRef]
- Oberholzer, T.; Nierhaus, K.H.; Luisi, P.L. Protein expression in liposomes. Biochem. Biophys. Res. Commun. 1999, 261, 238–241. [Google Scholar] [CrossRef] [PubMed]
- Oparin, A.I. The pathways of the primary development of metabolism and artificial modeling of this development in coacervate drops. In The Origins of Prebiological Systems and of Their Molecular Matrices; S. W. Fox: New York, NY, USA, 1965; pp. 331–345. [Google Scholar]
- Chakrabarti, A.C.; Breaker, R.R.; Joyce, G.F.; Deamer, D.W. Production of RNA by a polymerase protein encapsulated within phospholipid vesicles. J. Mol. Evol. 1994, 39, 555–559. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Huang, X.; Tang, T.-Y.D.; Mann, S. Synthetic cellularity based on non-lipid micro-compartments and protocell models. Curr. Opin. Chem. Biol. 2014, 22, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Dora Tang, T.-Y.; van Swaay, D.; deMello, A.; Ross Anderson, J.L.; Mann, S. In vitro gene expression within membrane-free coacervate protocells. Chem. Commun. (Camb.) 2015, 51, 11429–11432. [Google Scholar] [CrossRef] [PubMed]
- Frankel, E.A.; Bevilacqua, P.C.; Keating, C.D. Polyamine/Nucleotide Coacervates Provide Strong Compartmentalization of Mg2+, Nucleotides, and RNA. Langmuir 2016, 32, 2041–2049. [Google Scholar] [CrossRef] [PubMed]
- Budin, I.; Szostak, J.W. Physical effects underlying the transition from primitive to modern cell membranes. Proc. Natl. Acad. Sci. USA 2011, 108, 5249–5254. [Google Scholar] [CrossRef]
- Stano, P. Minimal cells: Relevance and interplay of physical and biochemical factors. Biotechol. J. 2011, 6, 850–859. [Google Scholar] [CrossRef]
- Engelhart, A.E.; Adamala, K.P.; Szostak, J.W. A simple physical mechanism enables homeostasis in primitive cells. Nat. Chem. 2016, 8, 448–453. [Google Scholar] [CrossRef]
- Luisi, P.L. The Emergence of Life: From Chemical Origins to Synthetic Biology, 1st ed.; Cambridge University Press: Cambridge, UK, 2006; ISBN 0-521-52801-1. [Google Scholar]
- Forster, A.C.; Church, G.M. Towards synthesis of a minimal cell. Mol. Syst. Biol. 2006, 2, 45. [Google Scholar] [CrossRef]
- Villarreal, F.; Tan, C. Cell-free systems in the new age of synthetic biology. Front. Chem. Sci. Eng. 2017, 11, 58–65. [Google Scholar] [CrossRef]
- Garenne, D.; Noireaux, V. Cell-free transcription-translation: Engineering biology from the nanometer to the millimeter scale. Curr. Opin. Biotechnol. 2018, 58, 19–27. [Google Scholar] [CrossRef]
- Forster, A.C.; Church, G.M. Synthetic biology projects in vitro. Genome Res. 2007, 17, 1–6. [Google Scholar] [CrossRef]
- Shi, T.; Han, P.; You, C.; Zhang, Y.-H.P.J. An in vitro synthetic biology platform for emerging industrial biomanufacturing: Bottom-up pathway design. Synth. Syst. Biotechnol. 2018, 3, 186–195. [Google Scholar] [CrossRef]
- Luisi, P.L. Chemical Aspects of Synthetic Biology. Chem. Biodiv. 2007, 4, 603–621. [Google Scholar] [CrossRef] [PubMed]
- Ashkenasy, G.; Hermans, T.M.; Otto, S.; Taylor, A.F. Systems chemistry. Chem. Soc. Rev. 2017, 46, 2543–2554. [Google Scholar] [CrossRef]
- Altamura, E.; Carrara, P.; D’Angelo, F.; Mavelli, F.; Stano, P. Extrinsic stochastic factors (solute partition) in gene expression inside lipid vesicles and lipid-stabilized water-in-oil droplets: A review. Synth. Biol. 2018, 3, ysy011. [Google Scholar] [CrossRef]
- Yu, W.; Sato, K.; Wakabayashi, M.; Nakaishi, T.; Ko-Mitamura, E.P.; Shima, Y.; Urabe, I.; Yomo, T. Synthesis of functional protein in liposome. J. Biosci. Bioeng. 2001, 92, 590–593. [Google Scholar] [CrossRef]
- Oberholzer, T.; Luisi, P.L. The use of liposomes for constructing cell models. J. Biol. Phys. 2002, 28, 733–744. [Google Scholar] [CrossRef]
- Ishikawa, K.; Sato, K.; Shima, Y.; Urabe, I.; Yomo, T. Expression of a cascading genetic network within liposomes. FEBS Lett. 2004, 576, 387–390. [Google Scholar] [CrossRef]
- Szoka, F.; Papahadjopoulos, D. Comparative properties and methods of preparation of lipid vesicles (liposomes). Annu. Rev. Biophys. Bioeng. 1980, 9, 467–508. [Google Scholar] [CrossRef] [PubMed]
- New, R.R.C. Liposomes: A Practical Approach, 1st ed.; IRL Press at Oxford University Press: Oxford, UK, 1990. [Google Scholar]
- Walde, P. Preparation of Vesicles (Liposomes). In Encyclopedia of Nanoscience and Nanotechnology; Nalwa, H.S., Ed.; American Scientific Publishers: Valencia, CA, USA, 2004; Volume 9, pp. 43–79. [Google Scholar]
- Walde, P.; Cosentino, K.; Engel, H.; Stano, P. Giant vesicles: Preparations and applications. ChemBioChem 2010, 11, 848–865. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Obinata, K.; Sugawara, T.; Urabe, I.; Yomo, T. Quantification of structural properties of cell-sized individual liposomes by flow cytometry. J. Biosci. Bioeng. 2006, 102, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Hosoi, T.; Sunami, T.; Toyota, T.; Fujinami, M.; Oguma, K.; Matsuura, T.; Suzuki, H.; Yomo, T. Population analysis of structural properties of giant liposomes by flow cytometry. Langmuir 2009, 25, 10439–10443. [Google Scholar] [CrossRef] [PubMed]
- Sakakura, T.; Nishimura, K.; Suzuki, H.; Yomo, T. Statistical analysis of discrete encapsulation of nanomaterials in colloidal capsules. Anal. Methods 2012, 4, 1648–1655. [Google Scholar] [CrossRef]
- Luisi, P.L.; Walde, P. (Eds.) Giant Vesicles; Wiley: Chichester, UK; New York, NY, USA, 2000; ISBN 0-471-97986-4. [Google Scholar]
- Fenz, S.F.; Sengupta, K. Giant vesicles as cell models. Integr. Biol. (Camb.) 2012, 4, 982–995. [Google Scholar] [CrossRef]
- Xiao, Z.; Huang, N.; Xu, M.; Lu, Z.; Wei, Y. Novel Preparation of Asymmetric Liposomes with Inner and Outer Layer of Different Materials. Chem. Lett. 1998, 27, 225–226. [Google Scholar] [CrossRef]
- Pautot, S.; Frisken, B.J.; Weitz, D.A. Production of unilamellar vesicles using an inverted emulsion. Langmuir 2003, 19, 2870–2879. [Google Scholar] [CrossRef]
- Pautot, S.; Frisken, B.J.; Weitz, D.A. Engineering asymmetric vesicles. Proc. Natl. Acad. Sci. USA 2003, 100, 10718–10721. [Google Scholar] [CrossRef]
- Dimova, R.; Marques, C. The Giant Vesicle Book, 1st ed.; CRC Press: Boca Raton, FL, USA, 2019; Chapter 1; ISBN 9781498752176. [Google Scholar]
- Fujii, S.; Matsuura, T.; Sunami, T.; Nishikawa, T.; Kazuta, Y.; Yomo, T. Liposome display for in vitro selection and evolution of membrane proteins. Nat. Protoc. 2014, 9, 1578–1591. [Google Scholar] [CrossRef]
- Rampioni, G.; D’Angelo, F.; Messina, M.; Zennaro, A.; Kuruma, Y.; Tofani, D.; Leoni, L.; Stano, P. Synthetic cells produce a quorum sensing chemical signal perceived by Pseudomonas aeruginosa. Chem. Commun. 2018, 54, 2090–2093. [Google Scholar] [CrossRef] [PubMed]
- Fayolle, D.; Altamura, E.; D’Onofrio, A.; Madanamothoo, W.; Fenet, B.; Mavelli, F.; Buchet, R.; Stano, P.; Fiore, M.; Strazewski, P. Crude phosphorylation mixtures containing racemic lipid amphiphiles self-assemble to give stable primitive compartments. Sci. Rep. 2017, 7, 18106. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Caspi, Y.; Meijering, A.E.C.; Dekker, C. Octanol-assisted liposome assembly on chip. Nat. Commun. 2016, 7, 10447. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.H.; Kwon, Y.-C.; Jewett, M.C. Non-standard amino acid incorporation into proteins using Escherichia coli cell-free protein synthesis. Front. Chem. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, Y.; Inoue, A.; Tomari, Y.; Suzuki, T.; Yokogawa, T.; Nishikawa, K.; Ueda, T. Cell-free translation reconstituted with purified components. Nat. Biotechnol. 2001, 19, 751–755. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, Y.; Kanamori, T.; Ueda, T. Protein synthesis by pure translation systems. Methods 2005, 36, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Damiano, L.; Stano, P. Synthetic Biology and Artificial Intelligence. Grounding a cross-disciplinary approach to the synthetic exploration of (embodied) cognition. Complex Syst. 2018, 27, 199–228. [Google Scholar] [CrossRef]
- Hillebrecht, J.R.; Chong, S. A comparative study of protein synthesis in in vitro systems: From the prokaryotic reconstituted to the eukaryotic extract-based. BMC Biotechnol. 2008, 8. [Google Scholar] [CrossRef]
- Koonin, E.V. How many genes can make a cell: The minimal-gene-set concept. Annu. Rev. Genom. Hum. Genet. 2000, 1, 99–116. [Google Scholar] [CrossRef]
- Gil, R.; Silva, F.J.; Peretó, J.; Moya, A. Determination of the core of a minimal bacterial gene set. Microbiol. Mol. Biol. Rev. 2004, 68, 518–537. [Google Scholar] [CrossRef]
- Caschera, F.; Noireaux, V. Synthesis of 2.3 mg/mL of protein with an all Escherichia coli cell-free transcription-translation system. Biochimie 2014, 99, 162–168. [Google Scholar] [CrossRef] [PubMed]
- Villarreal, F.; Contreras-Llano, L.E.; Chavez, M.; Ding, Y.; Fan, J.; Pan, T.; Tan, C. Synthetic microbial consortia enable rapid assembly of pure translation machinery. Nat. Chem. Biol. 2018, 14, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Van Swaay, D.; deMello, A. Microfluidic methods for forming liposomes. Lab Chip 2013, 13, 752–767. [Google Scholar] [CrossRef] [PubMed]
- Trantidou, T.; Friddin, M.S.; Salehi-Reyhani, A.; Ces, O.; Elani, Y. Droplet microfluidics for the construction of compartmentalised model membranes. Lab Chip 2018, 18, 2488–2509. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Dekker, C. On-chip microfluidic production of cell-sized liposomes. Nat. Protoc. 2018, 13, 856–874. [Google Scholar] [CrossRef] [PubMed]
- Booth, M.J.; Schild, V.R.; Graham, A.D.; Olof, S.N.; Bayley, H. Light-activated communication in synthetic tissues. Sci. Adv. 2016, 2, e1600056. [Google Scholar] [CrossRef] [PubMed]
- Martino, C.; Kim, S.-H.; Horsfall, L.; Abbaspourrad, A.; Rosser, S.J.; Cooper, J.; Weitz, D.A. Protein Expression, Aggregation, and Triggered Release from Polymersomes as Artificial Cell-like Structures. Angew. Chem. Int. Ed. 2012, 51, 6416–6420. [Google Scholar] [CrossRef] [PubMed]
- Petit, J.; Thomi, L.; Schultze, J.; Makowski, M.; Negwer, I.; Koynov, K.; Herminghaus, S.; Wurm, F.R.; Bäumchen, O.; Landfester, K. A modular approach for multifunctional polymersomes with controlled adhesive properties. Soft Matter 2018, 14, 894–900. [Google Scholar] [CrossRef]
- Jahn, A.; Vreeland, W.N.; Gaitan, M.; Locascio, L.E. Controlled vesicle self-assembly in microfluidic channels with hydrodynamic focusing. J. Am. Chem. Soc. 2004, 126, 2674–2675. [Google Scholar] [CrossRef]
- Siegal-Gaskins, D.; Tuza, Z.A.; Kim, J.; Noireaux, V.; Murray, R.M. Gene circuit performance characterization and resource usage in a cell-free “breadboard”. ACS Synth. Biol. 2014, 3, 416–425. [Google Scholar] [CrossRef]
- Karzbrun, E.; Shin, J.; Bar-Ziv, R.H.; Noireaux, V. Coarse-grained dynamics of protein synthesis in a cell-free system. Phys. Rev. Lett. 2011, 106, 048104. [Google Scholar] [CrossRef] [PubMed]
- Stögbauer, T.; Windhager, L.; Zimmer, R.; Rädler, J.O. Experiment and mathematical modeling of gene expression dynamics in a cell-free system. Integr. Biol. 2012, 4, 494–501. [Google Scholar] [CrossRef] [PubMed]
- Calviello, L.; Stano, P.; Mavelli, F.; Luisi, P.L.; Marangoni, R. Quasi-cellular systems: Stochastic simulation analysis at nanoscale range. BMC Bioinf. 2013, 14, S7. [Google Scholar] [CrossRef]
- Mavelli, F.; Marangoni, R.; Stano, P. A Simple Protein Synthesis Model for the PURE System Operation. Bull. Math. Biol. 2015, 77, 1185–1212. [Google Scholar] [CrossRef] [PubMed]
- Matsuura, T.; Tanimura, N.; Hosoda, K.; Yomo, T.; Shimizu, Y. Reaction dynamics analysis of a reconstituted Escherichia coli protein translation system by computational modeling. Proc. Natl. Acad. Sci. USA 2017, 114, E1336–E1344. [Google Scholar] [CrossRef] [PubMed]
- Matsuura, T.; Hosoda, K.; Shimizu, Y. Robustness of a Reconstituted Escherichia coli Protein Translation System Analyzed by Computational Modeling. ACS Synth. Biol. 2018, 7, 1964–1972. [Google Scholar] [CrossRef] [PubMed]
- Mavelli, F. Stochastic simulations of minimal cells: The Ribocell model. BMC Bioinf. 2012, 13 (Suppl. 4), S10. [Google Scholar] [CrossRef] [PubMed]
- Lazzerini-Ospri, L.; Stano, P.; Luisi, P.; Marangoni, R. Characterization of the emergent properties of a synthetic quasi-cellular system. BMC Bioinf. 2012, 13 (Suppl. 4), S9. [Google Scholar] [CrossRef]
- Kapsner, K.; Simmel, F.C. Partitioning Variability of a Compartmentalized In Vitro Transcriptional Thresholding Circuit. ACS Synth. Biol. 2015, 4, 1136–1143. [Google Scholar] [CrossRef]
- Fanti, A.; Gammuto, L.; Mavelli, F.; Stano, P.; Marangoni, R. Do protocells preferentially retain macromolecular solutes upon division/fragmentation? A study based on the extrusion of POPC giant vesicles. Integr. Biol. (Camb.) 2018, 10, 6–17. [Google Scholar] [CrossRef]
- Rampioni, G.; Mavelli, F.; Damiano, L.; D’Angelo, F.; Messina, M.; Leoni, L.; Stano, P. A synthetic biology approach to bio-chem-ICT: First moves towards chemical communication between synthetic and natural cells. Nat. Comput. 2014, 13, 1–17. [Google Scholar] [CrossRef]
- Bozic, B.; Svetina, S. A relationship between membrane properties forms the basis of a selectivity mechanism for vesicle self-reproduction. Eur. Biophys. J. 2004, 33, 565–571. [Google Scholar] [CrossRef] [PubMed]
- Mavelli, F.; Ruiz-Mirazo, K. ENVIRONMENT: A computational platform to stochastically simulate reacting and self-reproducing lipid compartments. Phys. Biol. 2010, 7, 036002. [Google Scholar] [CrossRef] [PubMed]
- Pereira de Souza, T.; Stano, P.; Luisi, P.L. The minimal size of liposome-based model cells brings about a remarkably enhanced entrapment and protein synthesis. ChemBioChem 2009, 10, 1056–1063. [Google Scholar] [CrossRef]
- Luisi, P.L.; Allegretti, M.; Pereira de Souza, T.; Steiniger, F.; Fahr, A.; Stano, P. Spontaneous protein crowding in liposomes: A new vista for the origin of cellular metabolism. ChemBioChem 2010, 11, 1989–1992. [Google Scholar] [CrossRef]
- Van Hoof, B.; Markvoort, A.J.; van Santen, R.A.; Hilbers, P.A.J. On protein crowding and bilayer bulging in spontaneous vesicle formation. J. Phys. Chem. B 2012, 116, 12677–12683. [Google Scholar] [CrossRef]
- Paradisi, P.; Allegrini, P.; Chiarugi, D. A renewal model for the emergence of anomalous solute crowding in liposomes. BMC Syst. Biol. 2015, 9, S7. [Google Scholar] [CrossRef]
- Liu, Y.; Tsao, C.-Y.; Kim, E.; Tschirhart, T.; Terrell, J.L.; Bentley, W.E.; Payne, G.F. Using a Redox Modality to Connect Synthetic Biology to Electronics: Hydrogel-Based Chemo-Electro Signal Transduction for Molecular Communication. Adv. Healthc. Mater. 2017, 6. [Google Scholar] [CrossRef]
- Selberg, J.; Gomez, M.; Rolandi, M. The Potential for Convergence between Synthetic Biology and Bioelectronics. Cell. Syst. 2018, 7, 231–244. [Google Scholar] [CrossRef]
- Bachmann, P.; Luisi, P.; Lang, J. Autocatalytic Self-Replicating Micelles as Models for Prebiotic Structures. Nature 1992, 357, 57–59. [Google Scholar] [CrossRef]
- Walde, P.; Wick, R.; Fresta, M.; Mangone, A.; Luisi, P. Autopoietic Self-Reproduction of Fatty-Acid Vesicles. J. Am. Chem. Soc. 1994, 116, 11649–11654. [Google Scholar] [CrossRef]
- Varela, F.G.; Maturana, H.R.; Uribe, R. Autopoiesis: The organization of living systems, its characterization and a model. Biosystems 1974, 5, 187–196. [Google Scholar] [CrossRef]
- Luisi, P.L. Autopoiesis: A review and a reappraisal. Naturwissenschaften 2003, 90, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Stano, P. Synthetic biology of minimal living cells: Primitive cell models and semi-synthetic cells. Syst. Synth. Biol. 2010, 4, 149–156. [Google Scholar] [CrossRef]
- Stano, P.; Luisi, P.L. Achievements and open questions in the self-reproduction of vesicles and synthetic minimal cells. Chem. Commun. (Camb.) 2010, 46, 3639–3653. [Google Scholar] [CrossRef] [PubMed]
- Letelier, J.C.; Marín, G.; Mpodozis, J. Autopoietic and (M,R) systems. J. Theor. Biol. 2003, 222, 261–272. [Google Scholar] [CrossRef]
- McMullin, B. Thirty years of computational autopoiesis: A review. Artif. Life 2004, 10, 277–295. [Google Scholar] [CrossRef]
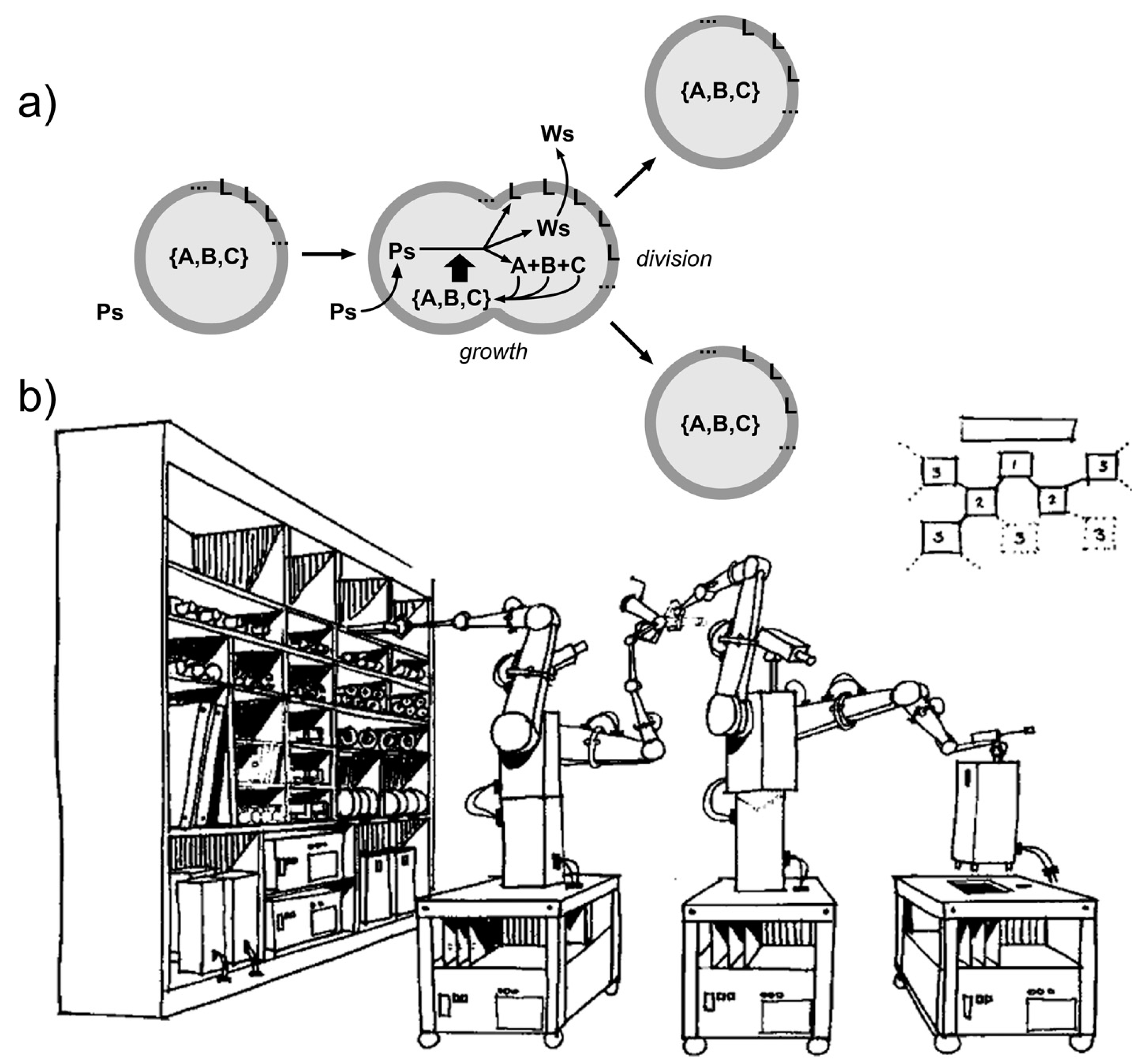
- Wikimedia Commons. “Advanced Automation for Space Missions Figure 5-29.gif”. Available online: https://commons.wikimedia.org/w/index.php?curid=1687447 (accessed on 26 November 2018).
- Gánti, T. Organization of chemical reactions into dividing and metabolizing units: The chemotons. Biosystems 1975, 7, 15–21. [Google Scholar] [CrossRef]
- Gànti, T. Chemoton Theory: Theory of Living Systems, 2004th ed.; Springer: Berlin, Germany, 2003; ISBN 0-306-47785-8. [Google Scholar]
- Studies in History and Philosophy of Science Part C: Studies in History and Philosophy of Biological and Biomedical Sciences; Elsevier: Amsterdam, The Netherlands, 2013; Volume 44, pp. 627–792.
- Damiano, L.; Stano, P. Understanding Embodied Cognition by Building Models of Minimal Life. In Artificial Life and Evolutionary Computation; Communications in Computer and Information Science; Springer: Cham, Switzerland, 2017; pp. 73–87. [Google Scholar]
- Bitbol, M.; Luisi, P.L. Autopoiesis with or without cognition: Defining life at its edge. J. R. Soc. Interface 2004, 1, 99–107. [Google Scholar] [CrossRef]
- Bourgine, P.; Stewart, J. Autopoiesis and cognition. Artif. Life 2004, 10, 327–345. [Google Scholar] [CrossRef]
- Ceruti, M.; Damiano, L. Plural Embodiment(s) of Mind. Genealogy and Guidelines for a Radically Embodied Approach to Mind and Consciousness. Front. Psychol. 2018, 9, 2204. [Google Scholar] [CrossRef] [PubMed]
- Stano, P.; Rampioni, G.; D’Angelo, F.; Altamura, E.; Mavelli, F.; Marangoni, R.; Rossi, F.; Damiano, L. Current Directions in Synthetic Cell Research. In Advances in Bionanomaterials; Lecture Notes in Bioengineering; Springer: Cham, Switzerland, 2018; pp. 141–154. ISBN 978-3-319-62026-8. [Google Scholar]
- Rigaud, J.-L.; Lévy, D. Reconstitution of membrane proteins into liposomes. Meth. Enzymol. 2003, 372, 65–86. [Google Scholar] [CrossRef]
- Dezi, M.; Cicco, A.D.; Bassereau, P.; Lévy, D. Detergent-mediated incorporation of transmembrane proteins in giant unilamellar vesicles with controlled physiological contents. Proc. Natl. Acad. Sci. USA 2013, 110, 7276–7281. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, I.L.; Kemmer, G.C.; Pomorski, T.G. Membrane protein reconstitution into giant unilamellar vesicles: A review on current techniques. Eur. Biophys. J. 2017, 46, 103–119. [Google Scholar] [CrossRef] [PubMed]
- Robertson, J.L. The lipid bilayer membrane and its protein constituents. J. Gen. Physiol. 2018, 150, 1472–1483. [Google Scholar] [CrossRef]
- Miller, C.; Racker, E. Reconstitution of Membrane Transport Functions. In The Receptors. General Principles and Procedures (A Comprehensive Treatise); O’Brien, R.D., Ed.; Springer: Boston, MA, USA, 1979; Volume 1, pp. 1–31. ISBN 978-1-4684-0981-9. [Google Scholar] [CrossRef]
- Kuruma, Y.; Stano, P.; Ueda, T.; Luisi, P.L. A synthetic biology approach to the construction of membrane proteins in semi-synthetic minimal cells. Biochim. Biophys. Acta 2009, 1788, 567–574. [Google Scholar] [CrossRef]
- Hamada, S.; Tabuchi, M.; Toyota, T.; Sakurai, T.; Hosoi, T.; Nomoto, T.; Nakatani, K.; Fujinami, M.; Kanzaki, R. Giant vesicles functionally expressing membrane receptors for an insect pheromone. Chem. Commun. (Camb.) 2014, 50, 2958–2961. [Google Scholar] [CrossRef]
- Soga, H.; Fujii, S.; Yomo, T.; Kato, Y.; Watanabe, H.; Matsuura, T. In vitro membrane protein synthesis inside cell-sized vesicles reveals the dependence of membrane protein integration on vesicle volume. ACS Synth. Biol. 2014, 3, 372–379. [Google Scholar] [CrossRef]
- Ohta, N.; Kato, Y.; Watanabe, H.; Mori, H.; Matsuura, T. In vitro membrane protein synthesis inside Sec translocon-reconstituted cell-sized liposomes. Sci. Rep. 2016, 6, 36466. [Google Scholar] [CrossRef]
- Uyeda, A.; Nakayama, S.; Kato, Y.; Watanabe, H.; Matsuura, T. Construction of an in Vitro Gene Screening System of the E. coli EmrE Transporter Using Liposome Display. Anal. Chem. 2016, 88, 12028–12035. [Google Scholar] [CrossRef]
- Furusato, T.; Horie, F.; Matsubayashi, H.T.; Amikura, K.; Kuruma, Y.; Ueda, T. De Novo Synthesis of Basal Bacterial Cell Division Proteins FtsZ, FtsA, and ZipA Inside Giant Vesicles. ACS Synth. Biol. 2018, 7, 953–961. [Google Scholar] [CrossRef]
- Yanagisawa, M.; Iwamoto, M.; Kato, A.; Yoshikawa, K.; Oiki, S. Oriented Reconstitution of a Membrane Protein in a Giant Unilamellar Vesicle: Experimental Verification with the Potassium Channel KcsA. J. Am. Chem. Soc. 2011, 133, 11774–11779. [Google Scholar] [CrossRef]
- Altamura, E.; Milano, F.; Tangorra, R.R.; Trotta, M.; Omar, O.H.; Stano, P.; Mavelli, F. Highly oriented photosynthetic reaction centers generate a proton gradient in synthetic protocells. Proc. Natl. Acad. Sci. USA 2017, 114, 3837–3842. [Google Scholar] [CrossRef] [PubMed]
- Sachse, R.; Dondapati, S.K.; Fenz, S.F.; Schmidt, T.; Kubick, S. Membrane protein synthesis in cell-free systems: From bio-mimetic systems to bio-membranes. FEBS Lett. 2014, 588, 2774–2781. [Google Scholar] [CrossRef] [PubMed]
- Kuruma, Y.; Suzuki, T.; Ono, S.; Yoshida, M.; Ueda, T. Functional analysis of membranous Fo-a subunit of F1Fo-ATP synthase by in vitro protein synthesis. Biochem. J. 2012, 442, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.-J.; Montemagno, C.D. Artificial Organelle: ATP Synthesis from Cellular Mimetic Polymersomes. Nano Lett. 2005, 5, 2538–2542. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Jia, Y.; Cai, P.; Fei, J.; Li, J. Coassembly of Photosystem II and ATPase as Artificial Chloroplast for Light-Driven ATP Synthesis. ACS Nano 2016, 10, 556–561. [Google Scholar] [CrossRef]
- Altamura, E.; Fiorentino, R.; Milano, F.; Trotta, M.; Palazzo, G.; Stano, P.; Mavelli, F. First moves towards photoautotrophic synthetic cells: In Vitro study of photosynthetic reaction centre and cytochrome bc1 complex interactions. Biophys. Chem. 2017, 229, 46–56. [Google Scholar] [CrossRef]
- Kumar, B.V.V.S.P.; Fothergill, J.; Bretherton, J.; Tian, L.; Patil, A.J.; Davis, S.A.; Mann, S. Chloroplast-containing coacervate micro-droplets as a step towards photosynthetically active membrane-free protocells. Chem. Commun. 2018, 54, 3594–3597. [Google Scholar] [CrossRef]
- Lee, K.Y.; Park, S.-J.; Lee, K.A.; Kim, S.-H.; Kim, H.; Meroz, Y.; Mahadevan, L.; Jung, K.-H.; Ahn, T.K.; Parker, K.K.; et al. Photosynthetic artificial organelles sustain and control ATP-dependent reactions in a protocellular system. Nat. Biotechnol. 2018, 36, 530–535. [Google Scholar] [CrossRef]
- Kim, T.; Murdande, S.; Gruber, A.; Kim, S. Sustained-release Morphine for Epidural Analgesia in Rats. Anesthesiology 1996, 85, 331–338. [Google Scholar] [CrossRef]
- Kisak, E.T.; Coldren, B.; Zasadzinski, J.A. Nanocompartments Enclosing Vesicles, Colloids, and Macromolecules via Interdigitated Lipid Bilayers. Langmuir 2002, 18, 284–288. [Google Scholar] [CrossRef]
- Kisak, E.T.; Coldren, B.; Evans, C.A.; Boyer, C.; Zasadzinski, J.A. The vesosome—A multicompartment drug delivery vehicle. Curr. Med. Chem. 2004, 11, 199–219. [Google Scholar] [CrossRef] [PubMed]
- Walker, S.A.; Kennedy, M.T.; Zasadzinski, J.A. Encapsulation of bilayer vesicles by self-assembly. Nature 1997, 387, 61–64. [Google Scholar] [CrossRef] [PubMed]
- Bolinger, P.-Y.; Stamou, D.; Vogel, H. An integrated self-assembled nanofluidic system for controlled biological chemistries. Angew. Chem. Int. Ed. Engl. 2008, 47, 5544–5549. [Google Scholar] [CrossRef] [PubMed]
- Paleos, C.M.; Tsiourvas, D.; Sideratou, Z. Interaction of vesicles: Adhesion, fusion and multicompartment systems formation. ChemBioChem 2011, 12, 510–521. [Google Scholar] [CrossRef] [PubMed]
- Hadorn, M.; Boenzli, E.; Eggenberger Hotz, P.; Hanczyc, M.M. Hierarchical Unilamellar Vesicles of Controlled Compositional Heterogeneity. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Deng, N.-N.; Yelleswarapu, M.; Zheng, L.; Huck, W.T.S. Microfluidic Assembly of Monodisperse Vesosomes as Artificial Cell Models. J. Am. Chem. Soc. 2017, 139, 587–590. [Google Scholar] [CrossRef]
- Haller, B.; Göpfrich, K.; Schröter, M.; Janiesch, J.-W.; Platzman, I.; Spatz, J.P. Charge-controlled microfluidic formation of lipid-based single- and multicompartment systems. Lab Chip 2018, 18, 2665–2674. [Google Scholar] [CrossRef] [PubMed]
- Pak, C.C.; Ali, S.; Janoff, A.S.; Meers, P. Triggerable liposomal fusion by enzyme cleavage of a novel peptide–lipid conjugate. Biochim. Biophys. Acta (BBA) Biomembr. 1998, 1372, 13–27. [Google Scholar] [CrossRef]
- Chaize, B.; Nguyen, M.; Ruysschaert, T.; le Berre, V.; Trévisiol, E.; Caminade, A.-M.; Majoral, J.P.; Pratviel, G.; Meunier, B.; Winterhalter, M.; et al. Microstructured liposome array. Bioconjug. Chem. 2006, 17, 245–247. [Google Scholar] [CrossRef]
- Liu, X.; Zhao, R.; Zhang, Y.; Jiang, X.; Yue, J.; Jiang, P.; Zhang, Z. Using giant unilamellar lipid vesicle micro-patterns as ultrasmall reaction containers to observe reversible ATP synthesis/hydrolysis of F0F1-ATPase directly. Biochim. Biophys. Acta 2007, 1770, 1620–1626. [Google Scholar] [CrossRef] [PubMed]
- Christensen, S.M.; Stamou, D.G. Sensing-applications of surface-based single vesicle arrays. Sensors 2010, 10, 11352–11368. [Google Scholar] [CrossRef] [PubMed]
- Osaki, T.; Kamiya, K.; Kawano, R.; Sasaki, H.; Takeuchi, S. Towards artificial cell array system: Encapsulation and hydration technologies integrated in liposome array. In Proceedings of the 2012 IEEE 25th International Conference on Micro Electro Mechanical Systems (MEMS), Paris, France, 29 January–2 February 2012; pp. 333–336. [Google Scholar]
- Kang, Y.J.; Wostein, H.S.; Majd, S. A simple and versatile method for the formation of arrays of giant vesicles with controlled size and composition. Adv. Mater. 2013, 25, 6834–6838. [Google Scholar] [CrossRef] [PubMed]
- Mantri, S.; Sapra, K.T. Evolving protocells to prototissues: Rational design of a missing link. Biochem. Soc. Trans. 2013, 41, 1159–1165. [Google Scholar] [CrossRef] [PubMed]
- Hamano, H.; Tonooka, T.; Osaki, T.; Takeuchi, S. Highly packed liposome assemblies toward synthetic tissue. In Proceedings of the 2014 IEEE 27th International Conference on Micro Electro Mechanical Systems (MEMS), San Francisco, CA, USA, 26–30 January 2014; pp. 17–19. [Google Scholar]
- Hamano, H.; Osaki, T.; Takeuchi, S. Liposome arrangement connected with avidin-biotin complex for constructing functional synthetic tissue. In Proceedings of the 2015 28th IEEE International Conference on Micro Electro Mechanical Systems (MEMS), Estoril, Portugal, 18–22 January 2015; pp. 336–339. [Google Scholar]
- Kazayama, Y.; Teshima, T.; Osaki, T.; Takeuchi, S.; Toyota, T. Integrated Microfluidic System for Size-Based Selection and Trapping of Giant Vesicles. Anal. Chem. 2016, 88, 1111–1116. [Google Scholar] [CrossRef]
- Hadorn, M.; Hotz, P.E. DNA-Mediated Self-Assembly of Artificial Vesicles. PLoS ONE 2010, 5. [Google Scholar] [CrossRef]
- Hadorn, M.; Boenzli, E.; Sørensen, K.T.; De Lucrezia, D.; Hanczyc, M.M.; Yomo, T. Defined DNA-Mediated Assemblies of Gene-Expressing Giant Unilamellar Vesicles. Langmuir 2013, 29, 15309–15319. [Google Scholar] [CrossRef]
- Carrara, P.; Stano, P.; Luisi, P.L. Giant Vesicles “Colonies”: A Model for Primitive Cell Communities. ChemBioChem 2012, 13, 1497–1502. [Google Scholar] [CrossRef]
- Nakano, T.; Eckford, A.W.; Haraguchi, T. Molecular Communications; Cambridge University Press: Cambridge, UK, 2013; ISBN 978-1-107-02308-6. [Google Scholar]
- Nakano, T. Molecular Communication: A 10 Year Retrospective. IEEE Trans. Mol. Biol. Multi-Scale Commun. 2017, 3, 71–78. [Google Scholar] [CrossRef]
- Amos, M.; Dittrich, P.; McCaskill, J.; Rasmussen, S. Biological and Chemical Information Technologies. Procedia Comput. Sci. 2011, 7, 56–60. [Google Scholar] [CrossRef]
- Stano, P.; Rampioni, G.; Carrara, P.; Damiano, L.; Leoni, L.; Luisi, P.L. Semi-synthetic minimal cells as a tool for biochemical ICT. BioSystems 2012, 109, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Cronin, L.; Krasnogor, N.; Davis, B.G.; Alexander, C.; Robertson, N.; Steinke, J.H.G.; Schroeder, S.L.M.; Khlobystov, A.N.; Cooper, G.; Gardner, P.M.; et al. The imitation game—A computational chemical approach to recognizing life. Nat. Biotechnol. 2006, 24, 1203–1206. [Google Scholar] [CrossRef] [PubMed]
- Leduc, P.R.; Wong, M.S.; Ferreira, P.M.; Groff, R.E.; Haslinger, K.; Koonce, M.P.; Lee, W.Y.; Love, J.C.; McCammon, J.A.; Monteiro-Riviere, N.A.; et al. Towards an in vivo biologically inspired nanofactory. Nat. Nanotechnol. 2007, 2, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Gardner, P.M.; Winzer, K.; Davis, B.G. Sugar synthesis in a protocellular model leads to a cell signalling response in bacteria. Nat. Chem. 2009, 1, 377–383. [Google Scholar] [CrossRef] [PubMed]
- Kaneda, M.; Nomura, S.M.; Ichinose, S.; Kondo, S.; Nakahama, K.; Akiyoshi, K.; Morita, I. Direct formation of proteo-liposomes by in vitro synthesis and cellular cytosolic delivery with connexin-expressing liposomes. Biomaterials 2009, 30, 3971–3977. [Google Scholar] [CrossRef] [PubMed]
- Rampioni, G.; D’Angelo, F.; Leoni, L.; Stano, P. Gene-expressing liposomes as synthetic cells for molecular communication studies. Front. Bioeng. Biotech. Synth. Biol. 2019, submitted. [Google Scholar]
- Lentini, R.; Santero, S.P.; Chizzolini, F.; Cecchi, D.; Fontana, J.; Marchioretto, M.; Del Bianco, C.; Terrell, J.L.; Spencer, A.C.; Martini, L.; et al. Integrating artificial with natural cells to translate chemical messages that direct E. coli behaviour. Nat. Commun. 2014, 5, 4012. [Google Scholar] [CrossRef]
- Lentini, R.; Martín, N.Y.; Forlin, M.; Belmonte, L.; Fontana, J.; Cornella, M.; Martini, L.; Tamburini, S.; Bentley, W.E.; Jousson, O.; et al. Two-Way Chemical Communication between Artificial and Natural Cells. ACS Cent. Sci. 2017, 3, 117–123. [Google Scholar] [CrossRef]
- Niederholtmeyer, H.; Chaggan, C.; Devaraj, N.K. Communication and quorum sensing in non-living mimics of eukaryotic cells. Nat. Commun. 2018, 9, 5027. [Google Scholar] [CrossRef]
- Tang, T.-Y.D.; Cecchi, D.; Fracasso, G.; Accardi, D.; Coutable-Pennarun, A.; Mansy, S.S.; Perriman, A.W.; Anderson, J.L.R.; Mann, S. Gene-Mediated Chemical Communication in Synthetic Protocell Communities. ACS Synth. Biol. 2018, 7, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Krinsky, N.; Kaduri, M.; Zinger, A.; Shainsky-Roitman, J.; Goldfeder, M.; Benhar, I.; Hershkovitz, D.; Schroeder, A. Synthetic Cells Synthesize Therapeutic Proteins inside Tumors. Adv. Healthc. Mater. 2018, 7, e1701163. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Contreras-Llano, L.E.; Morris, E.; Mao, M.; Tan, C. Minimizing Context Dependency of Gene Networks Using Artificial Cells. ACS Appl. Mater. Interfaces 2018, 10, 30137–30146. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Koo, H.; Na, J.H.; Lee, K.E.; Jeong, S.Y.; Choi, K.; Kim, S.H.; Kwon, I.C.; Kim, K. DNA amplification in neutral liposomes for safe and efficient gene delivery. ACS Nano 2014, 8, 4257–4267. [Google Scholar] [CrossRef] [PubMed]
- Tsugane, M.; Suzuki, H. Reverse Transcription Polymerase Chain Reaction in Giant Unilamellar Vesicles. Sci. Rep. 2018, 8, 9214. [Google Scholar] [CrossRef] [PubMed]
- Van Nies, P.; Westerlaken, I.; Blanken, D.; Salas, M.; Mencía, M.; Danelon, C. Self-replication of DNA by its encoded proteins in liposome-based synthetic cells. Nat. Commun. 2018, 9, 1583. [Google Scholar] [CrossRef] [PubMed]
- Scott, A.; Noga, M.J.; de Graaf, P.; Westerlaken, I.; Yildirim, E.; Danelon, C. Cell-Free Phospholipid Biosynthesis by Gene-Encoded Enzymes Reconstituted in Liposomes. PLoS ONE 2016, 11, e0163058. [Google Scholar] [CrossRef]
- Gell-Mann, M.; Lloyd, S. Information measures, effective complexity, and total information. Complexity 1996, 2, 44–52. [Google Scholar] [CrossRef]
- Emmeche, C. Aspects of Complexity in Life and Science. Philosophica 1997, 59, 41–68. [Google Scholar]
- Lloyd, S. Measures of complexity: A nonexhaustive list. IEEE Control Syst. Mag. 2001, 21, 7–8. [Google Scholar] [CrossRef]
- Cheng, Z.L.; Luisi, P.L. Coexistence and mutual competition of vesicles with different size distributions. J. Phys. Chem. B 2003, 107, 10940–10945. [Google Scholar] [CrossRef]
- Chen, I.A.; Roberts, R.W.; Szostak, J.W. The emergence of competition between model protocells. Science 2004, 305, 1474–1476. [Google Scholar] [CrossRef]
- Adamala, K.; Szostak, J.W. Competition between model protocells driven by an encapsulated catalyst. Nat. Chem. 2013, 5, 495–501. [Google Scholar] [CrossRef]
- Stano, P. Question 7: New aspects of interactions among vesicles. Orig. Life Evol. Biosph. 2007, 37, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Tsuru, S.; Suzuki, H.; Yomo, T. Stochasticity in Gene Expression in a Cell-Sized Compartment. ACS Synth. Biol. 2015, 4, 566–576. [Google Scholar] [CrossRef]
- Stano, P.; de Souza, T.P.; Carrara, P.; Altamura, E.; D’Aguanno, E.; Caputo, M.; Luisi, P.L.; Mavelli, F. Recent Biophysical Issues About the Preparation of Solute-Filled Lipid Vesicles. Mech. Adv. Mater. Struct. 2015, 22, 748–759. [Google Scholar] [CrossRef]
- Damiano, L.; Hiolle, A.; Cañamero, L. Grounding Synthetic Knowledge. In Advances in Artificial Life, ECAL 2011; Lenaerts, T., Giacobini, M., Bersini, H., Bourgine, P., Dorigo, M., Doursat, R., Eds.; MIT Press: Boston, MA, USA, 2011; pp. 200–207. [Google Scholar]
- Sunami, T.; Sato, K.; Matsuura, T.; Tsukada, K.; Urabe, I.; Yomo, T. Femtoliter compartment in liposomes for in vitro selection of proteins. Anal. Biochem. 2006, 357, 128–136. [Google Scholar] [CrossRef]
- Murtas, G.; Kuruma, Y.; Bianchini, P.; Diaspro, A.; Luisi, P.L. Protein synthesis in liposomes with a minimal set of enzymes. Biochem. Biophys. Res. Commun. 2007, 363, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Nilsson, L.; Kurland, C.G. Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J. Mol. Biol. 1996, 260, 649–663. [Google Scholar] [CrossRef]
- Gibson, D.G.; Glass, J.I.; Lartigue, C.; Noskov, V.N.; Chuang, R.-Y.; Algire, M.A.; Benders, G.A.; Montague, M.G.; Ma, L.; Moodie, M.M.; et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 2010, 329, 52–56. [Google Scholar] [CrossRef]
- Hutchison, C.A.; Chuang, R.-Y.; Noskov, V.N.; Assad-Garcia, N.; Deerinck, T.J.; Ellisman, M.H.; Gill, J.; Kannan, K.; Karas, B.J.; Ma, L.; et al. Design and synthesis of a minimal bacterial genome. Science 2016, 351, aad6253. [Google Scholar] [CrossRef] [PubMed]
- Jewett, M.C.; Fritz, B.R.; Timmerman, L.E.; Church, G.M. In vitro integration of ribosomal RNA synthesis, ribosome assembly, and translation. Mol. Syst. Biol. 2013, 9, 678. [Google Scholar] [CrossRef] [PubMed]
- Caschera, F.; Lee, J.W.; Ho, K.K.Y.; Liu, A.P.; Jewett, M.C. Cell-free compartmentalized protein synthesis inside double emulsion templated liposomes with in vitro synthesized and assembled ribosomes. Chem. Commun. (Camb.) 2016, 52, 5467–5469. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Haas, W.; Jackson, K.; Kuru, E.; Jewett, M.C.; Fan, Z.H.; Gygi, S.; Church, G.M. Cogenerating Synthetic Parts toward a Self-Replicating System. ACS Synth. Biol. 2017, 6, 1327–1336. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhang, Y.; Jia, X.; Hu, M.; Deng, Z.; Xu, Y.; Liu, T. In Vitro Reconstitution and Optimization of the Entire Pathway to Convert Glucose into Fatty Acid. ACS Synth. Biol. 2017, 6, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Exterkate, M.; Caforio, A.; Stuart, M.C.A.; Driessen, A.J.M. Growing Membranes In Vitro by Continuous Phospholipid Biosynthesis from Free Fatty Acids. ACS Synth. Biol. 2018, 7, 153–165. [Google Scholar] [CrossRef] [PubMed]
- Blöchliger, E.; Blocher, M.; Walde, P.; Luisi, P.L. Matrix effect in the size distribution of fatty acid vesicles. J. Phys. Chem. B 1998, 102, 10383–10390. [Google Scholar] [CrossRef]
- Lonchin, S.; Luisi, P.L.; Walde, P.; Robinson, B.H. A matrix effect in mixed phospholipid/fatty acid vesicle formation. J. Phys. Chem. B 1999, 103, 10910–10916. [Google Scholar] [CrossRef]
- Berclaz, N.; Muller, M.; Walde, P.; Luisi, P.L. Growth and transformation of vesicles studied by ferritin labeling and cryotransmission electron microscopy. J. Phys. Chem. B 2001, 105, 1056–1064. [Google Scholar] [CrossRef]
- Rasi, S.; Mavelli, F.; Luisi, P.L. Cooperative micelle binding and matrix effect in oleate vesicle formation. J. Phys. Chem. B 2003, 107, 14068–14076. [Google Scholar] [CrossRef]
- Stano, P.; Wehrli, E.; Luisi, P.L. Insights into the self-reproduction of oleate vesicles. J. Phys. Condens. Matter 2006, 18, S2231. [Google Scholar] [CrossRef]
- Zhu, T.F.; Szostak, J.W. Coupled Growth and Division of Model Protocell Membranes. J. Am. Chem. Soc. 2009, 131, 5705–5713. [Google Scholar] [CrossRef] [PubMed]
- Carello, C.; Turvey, M.T.; Kugler, P.N.; Shaw, R.E. Inadequacies of the Computer Metaphor. In Handbook of Cognitive Neuroscience; Gazzaniga, M.S., Ed.; Springer: Boston, MA, USA, 1984; pp. 229–248. ISBN 978-1-4899-2177-2. [Google Scholar]
- Danchin, A. Bacteria as computers making computers. FEMS Microbiol. Rev. 2009, 33, 3–26. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, E. A mechanical Turing machine: Blueprint for a biomolecular computer. Interface Focus 2012, 2, 497–503. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, D.J. Organisms≠Machines. Stud. Hist. Philos. Biol. Biomed. Sci. 2013, 44, 669–678. [Google Scholar] [CrossRef]
- Boldt, J. Machine metaphors and ethics in synthetic biology. Life Sci. Soc. Policy 2018, 14, 12. [Google Scholar] [CrossRef]
- Jiang, Y.; Xu, C. The calculation of information and organismal complexity. Biol. Direct 2010, 5, 59. [Google Scholar] [CrossRef]
- Kolmogorov, A.N. Three approaches to the quantitative definition of information. Int. J. Comp. Math. 1968, 2, 157–168. [Google Scholar] [CrossRef]
- McCabe, T.J. A complexity measure. IEEE Trans. Soft. Eng. 1976, SE-2, 308–320. [Google Scholar] [CrossRef]
- Shin, J.; Noireaux, V. An E. coli Cell-Free Expression Toolbox: Application to Synthetic Gene Circuits and Artificial Cells. ACS Synth. Biol. 2012, 1, 29–41. [Google Scholar] [CrossRef]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stano, P. Is Research on “Synthetic Cells” Moving to the Next Level? Life 2019, 9, 3. https://doi.org/10.3390/life9010003
Stano P. Is Research on “Synthetic Cells” Moving to the Next Level? Life. 2019; 9(1):3. https://doi.org/10.3390/life9010003
Chicago/Turabian StyleStano, Pasquale. 2019. "Is Research on “Synthetic Cells” Moving to the Next Level?" Life 9, no. 1: 3. https://doi.org/10.3390/life9010003
APA StyleStano, P. (2019). Is Research on “Synthetic Cells” Moving to the Next Level? Life, 9(1), 3. https://doi.org/10.3390/life9010003




