Biota and Biomolecules in Extreme Environments on Earth: Implications for Life Detection on Mars
Abstract
:1. Introduction
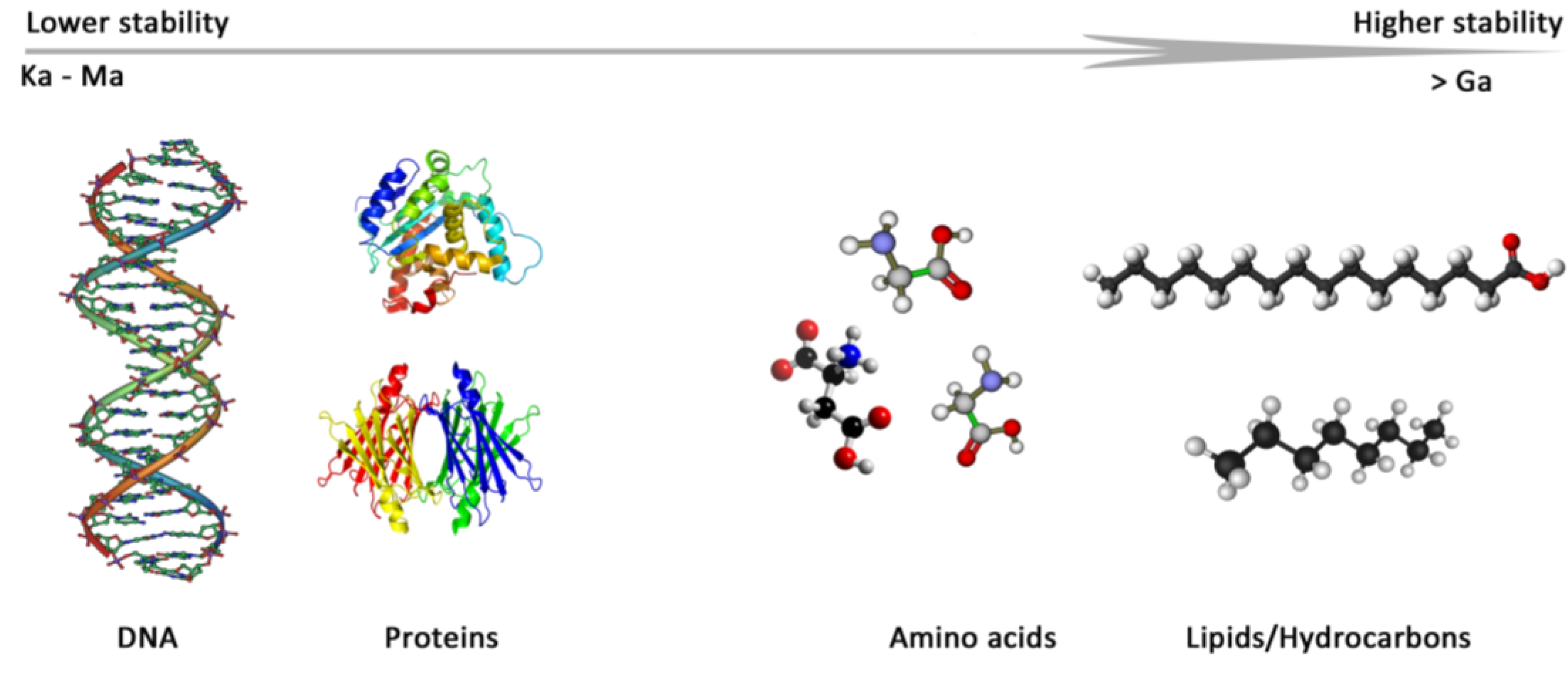
2. Biomarkers: What to Look for?
2.1. Deoxyribonucleic Acid (DNA)
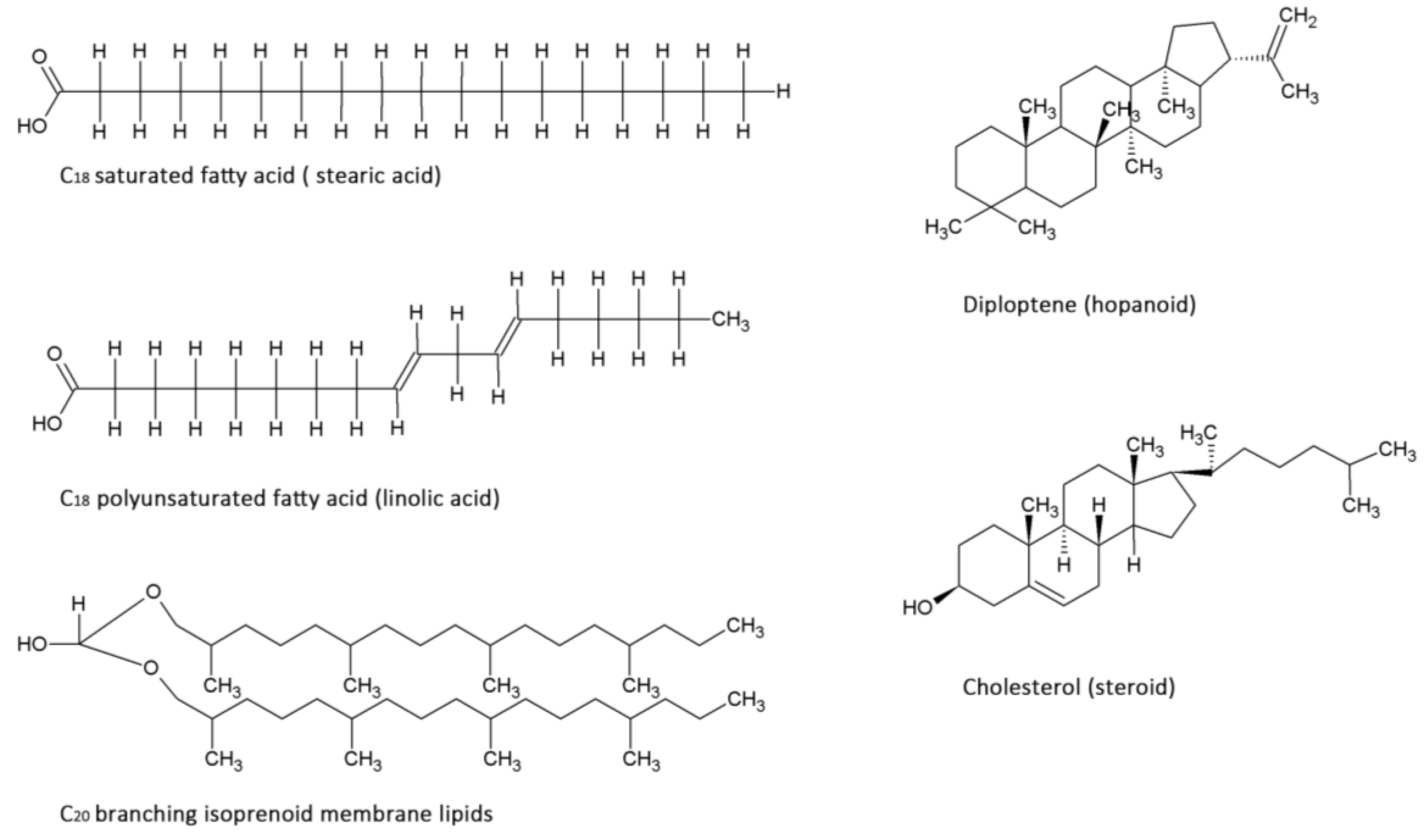
2.2. Lipids
| Domain | Dominant Membrane molecule | Molecule build-up/adaptations | Typical chain length |
|---|---|---|---|
| Archaea | Isoprenoids | 5-carbon isoprene unit incorporation, unsaturated branched side chains | 20 carbon atoms |
| Bacteria | Fatty acids, Hopanoids | 2-carbon acetyl incorporation, unsaturated cis-double bonds, addition of methyl groups | 14–18 carbon atoms |
| Eukarya | Fatty acids, Steroids | 2-carbon acetyl incorporation, unsaturated cis-double bonds | 14–18 carbon atoms |
2.3. Amino Acids
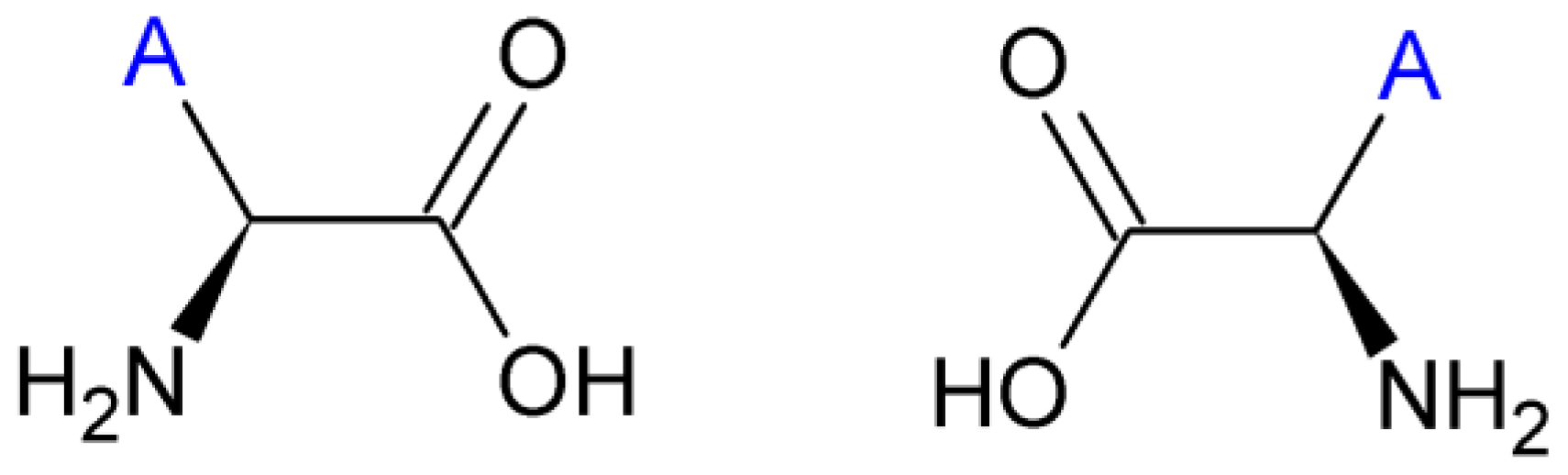
| Amino Acid | Side chain properties | Chemical structure | Amino Acid | Side chain properties | Chemical structure |
|---|---|---|---|---|---|
| Alanine | Hydrophobic side chain | 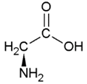 | Serine | Polar, uncharged side chain | 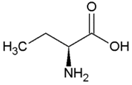 |
| Valine | Hydrophobic side chain | 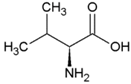 | Threonine | Polar, uncharged side chain | 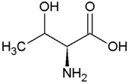 |
| Leucine | Hydrophobic side chain | 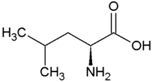 | Asparagine | Polar, uncharged side chain | 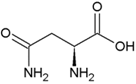 |
| Isoleucine | Hydrophobic side chain | 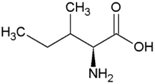 | Methionine | Polar, uncharged side chain | 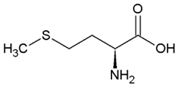 |
| Phenyl-alanine | Hydrophobic side chain | 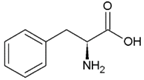 | Lysine | Positively charged side chain |  |
| Tyrosine | Hydrophobic side chain | 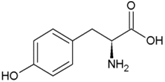 | Arginine | Positively charged side chain | 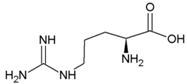 |
| Tryptophan | Hydrophobic side chain | 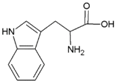 | Histidine | Positively charged side chain | 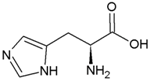 |
| Proline | Hydrophobic side chain | 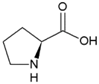 | Aspartic acid | Negatively charged side chain | 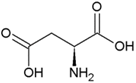 |
| Glycine | Polar, uncharged side chain | 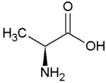 | Glutamic acid | Negatively charged side chain | 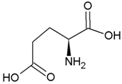 |
| Glutamine | Polar, uncharged side chain | 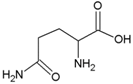 | Pyrrolysine | Positively charged side chain | 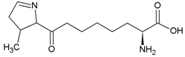 |
| Cysteine | Polar, uncharged side chain | 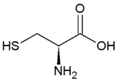 | Seleno-Cysteine | Hydrophobic side chain | 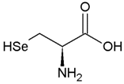 |
3. Mars and Terrestrial Analogues: Where to Look?
3.1. Mars: Past and Present
3.2. Terrestrial Extreme Environments
3.2.1. Hot Deserts
3.2.2. Subsurface Environments
3.2.3. Polar Region: Antarctica
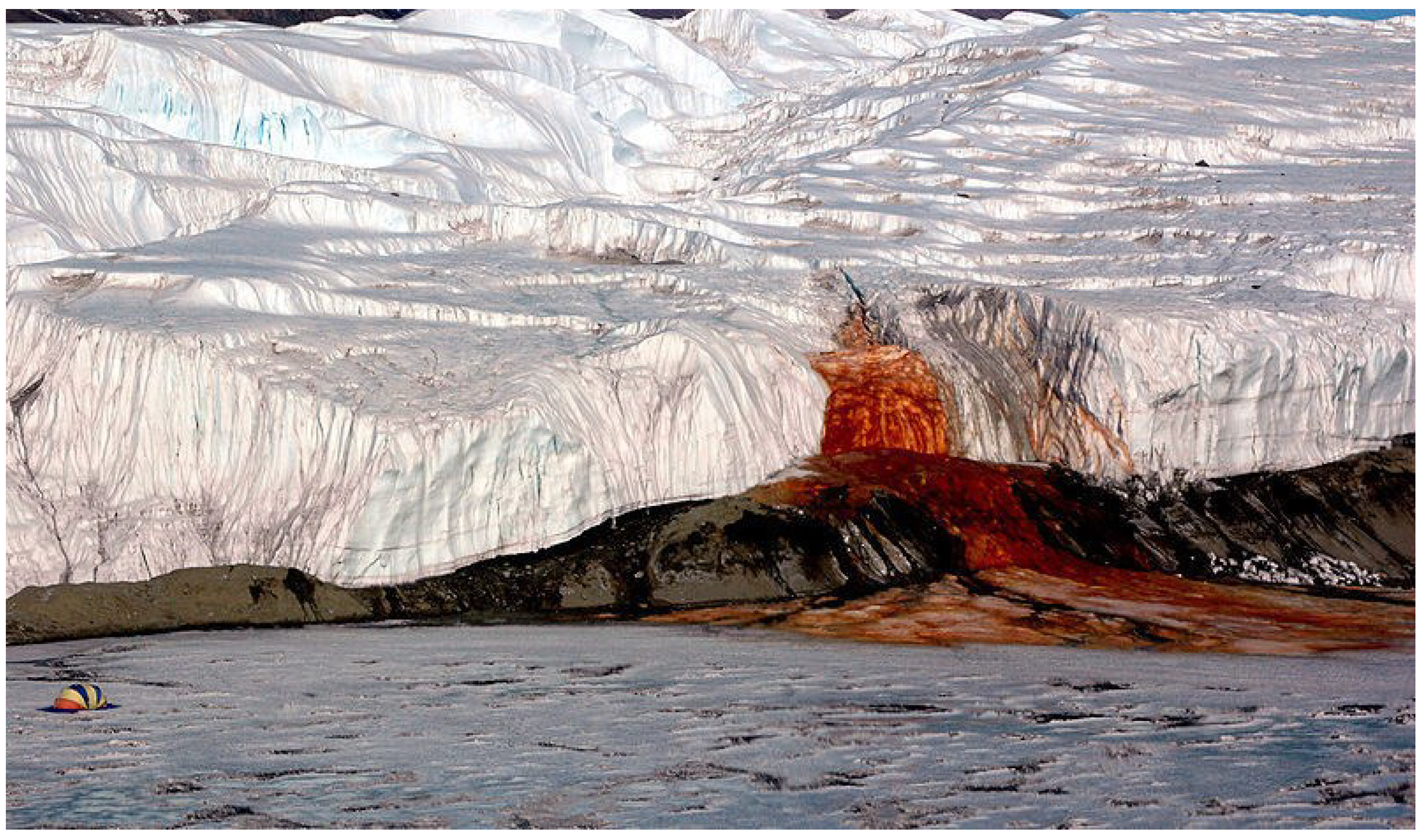

3.3. C-Type Meteorites
4. Techniques Currently in Use for Biomarker Detection: How to Look Here?
5. Current Instrumentation on Mars Life Detection Missions: How to Look There?
6. Conclusions and Looking forward
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ehrenfreund, P.; Rasmussen, S.; Cleaves, J.; Chen, L. Experimentally tracing the key steps in the origin of life: The aromatic world. Astrobiology 2006, 6, 490–520. [Google Scholar] [PubMed]
- Bada, J.L.; Lazcano, A. The Origin of Life; Courier Dover Publications: Mineola, NY, USA, 2001. [Google Scholar]
- Miller, S.L. A Production of Amino Acids Under Possible Primitive Earth Conditions. Science 1953, 117, 528–529. [Google Scholar] [CrossRef] [PubMed]
- Parker, E.T.; Zhou, M.; Burton, A.S.; Glavin, D.P.; Dworkin, J.P.; Krishnamurthy, R.; Fernández, F.M.; Bada, J.L. A Plausible Simultaneous Synthesis of Amino Acids and Simple Peptides on the Primordial Earth. Angewandte Chemie 2014, 126, 8270–8274. [Google Scholar] [CrossRef]
- Levy, M.; Miller, S.L.; Brinton, K.; Bada, J.L. Prebiotic Synthesis of Adenine and Amino Acids Under Europa-like Conditions. Icarus 2000, 145, 609–613. [Google Scholar] [CrossRef] [PubMed]
- Martins, Z.; Price, M.C.; Goldman, N.; Sephton, M.A.; Burchell, M.J. Shock synthesis of amino acids from impacting cometary and icy planet surface analogues. Nat. Geosci. 2013, 6, 1045–1049. [Google Scholar] [CrossRef]
- Simoneit, B.R.T. Biomarkers (molecular fossils) as geochemical indicators of life. Adv. Space Res. 2004, 33, 1255–1261. [Google Scholar] [CrossRef]
- Rushdi, A.I.; Simoneit, B.R. Lipid formation by aqueous Fischer-Tropsch-type synthesis over a temperature range of 100 to 400 degrees C. Orig. Life Evol. Biosph. 2001, 31, 103–118. [Google Scholar] [CrossRef] [PubMed]
- McCollom, T.; Ritter, G.; Simoneit, B.T. Lipid Synthesis Under Hydrothermal Conditions by Fischer-Tropsch-Type Reactions. Orig. Life Evol. Biosph. 1999, 29, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Saladino, R.; Ciambecchini, U.; Crestini, C.; Costanzo, G.; Negri, R.; Di Mauro, E. One-Pot TiO2-Catalyzed Synthesis of Nucleic Bases and Acyclonucleosides from Formamide: Implications for the Origin of Life. ChemBioChem 2003, 4, 514–521. [Google Scholar] [CrossRef] [PubMed]
- Barks, H.L.; Buckley, R.; Grieves, G.A.; Di Mauro, E.; Hud, N.V.; Orlando, T.M. Guanine, Adenine, and Hypoxanthine Production in UV-Irradiated Formamide Solutions: Relaxation of the Requirements for Prebiotic Purine Nucleobase Formation. ChemBioChem 2010, 11, 1240–1243. [Google Scholar] [CrossRef] [PubMed]
- Saladino, R.; Crestini, C.; Ciciriello, F.; Costanzo, G.; Di Mauro, E. Formamide Chemistry and the Origin of Informational Polymers. Chem. Biodivers. 2007, 4, 694–720. [Google Scholar] [CrossRef] [PubMed]
- Sephton, M.A. Organic matter in carbonaceous meteorites: Past, present and future research. Phil. Trans. R. Soc. 2005, 363, 2729–2742. [Google Scholar]
- Parnell, J.; Cullen, D.; Sims, M.R.; Bowden, S.; Cockell, C.S.; Court, R.; Ehrenfreund, P.; Gaubert, F.; Grant, W.; Parro, V.; et al. Searching for life on Mars: Selection of molecular targets for ESA’s aurora ExoMars mission. Astrobiology 2007, 7, 578–604. [Google Scholar] [CrossRef] [PubMed]
- Kanavarioti, A.; Mancinelli, R.L. Could organic matter have been preserved on Mars for 3.5 billion years? Icarus 1990, 84, 196–202. [Google Scholar] [CrossRef]
- Pavlov, A.K.; Blinov, A.V.; Konstantinov, A.N. Sterilization of Martian surface by cosmic radiation. Planet. Space Sci. 2002, 50, 669–673. [Google Scholar] [CrossRef]
- Jones, B.W. Mars before the Space Age. Int. J. Astrobiol. 2008, 7, 143–155. [Google Scholar]
- Rothschild, L.J.; Mancinelli, R.L. Life in extreme environments. Nature 2001, 409, 1092–1101. [Google Scholar] [CrossRef] [PubMed]
- Satyanarayana, T.; Raghukumar, C.; Shivaji, S. Extremophilic microbes: Diversity and perspectives. Curr. Sci. 2005, 89, 78–90. [Google Scholar]
- Lin, L.-H.; Wang, P.-L.; Rumble, D.; Lippmann-Pipke, J.; Boice, E.; Pratt, L.M.; Lollar, B.S.; Brodie, E.L.; Hazen, T.C.; Andersen, G.L.; et al. Long-Term Sustainability of a High-Energy, Low-Diversity Crustal Biome. Science 2006, 314, 479–482. [Google Scholar] [CrossRef] [PubMed]
- Pointing, S.B.; Chan, Y.; Lacap, D.C.; Lau, M.C.Y.; Jurgens, J.A.; Farrell, R.L. Highly specialized microbial diversity in hyper-arid polar desert. Proc. Natl. Acad. Sci. USA 2009, 106, 19964–19969. [Google Scholar] [CrossRef] [PubMed]
- Shtarkman, Y.M.; Kocer, Z.A.; Edgar, R.; Veerapaneni, R.S.; D’Elia, T.; Morris, P.F.; Rogers, S.O. Subglacial Lake Vostok (Antarctica) accretion ice contains a diverse set of sequences from aquatic, marine and sediment-inhabiting bacteria and eukarya. PLoS One 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Rohwerder, T.; Sand, W.; Lascu, C. Preliminary Evidence for a Sulphur Cycle in Movile Cave, Romania. Acta Biotechnol. 2003, 23, 101–107. [Google Scholar]
- Brocks, J.J.; Logan, G.A.; Buick, R.; Summons, R.E. Archean Molecular Fossils and the Early Rise of Eukaryotes. Science 1999, 285, 1033–1036. [Google Scholar] [CrossRef] [PubMed]
- Aubrey, A.; Cleaves, H.J.; Chalmers, J.H.; Skelley, A.M.; Mathies, R.A.; Grunthaner, F.J.; Ehrenfreund, P.; Bada, J.L. Sulfate minerals and organic compounds on Mars. Geology 2006, 34, 357–360. [Google Scholar] [CrossRef]
- Kminek, G.; Bada, J. The effect of ionizing radiation on the preservation of amino acids on Mars. Earth Planet. Sci. Lett. 2006, 245, 1–5. [Google Scholar] [CrossRef]
- Paabo, S.; Poinar, H.; Serre, D.; Jaenicke-Despres, V.; Hebler, J.; Rohland, N.; Kuch, M.; Krause, J.; Vigilant, L.; Hofreiter, M. Genetic analyses from ancient DNA. Ann. Rev. Genet. 2004, 38, 645–679. [Google Scholar] [CrossRef] [PubMed]
- Orlando, L.; Ginolhac, A.; Zhang, G.; Froese, D.; Albrechtsen, A.; Stiller, M.; Schubert, M.; Cappellini, E.; Petersen, B.; Moltke, I.; et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 2013, 499, 74–78. [Google Scholar] [CrossRef] [PubMed]
- Ehrenfreund, P.; Röling, W.F.M.; Thiel, C.S.; Quinn, R.; Sephton, M.A.; Stoker, C.; Kotler, J.M.; Direito, S.O.L.; Martins, Z.; Orzechowska, G.E.; et al. Astrobiology and habitability studies in preparation for future Mars missions: Trends from investigating minerals, organics and biota. Int. J. Astrobiol. 2011, 10, 239–253. [Google Scholar] [CrossRef]
- Direito, S.O.L.; Marees, A.; Röling, W.F.M. Sensitive life detection strategies for low-biomass environments: Optimizing extraction of nucleic acids adsorbing to terrestrial and Mars analogue minerals. FEMS Microbiol. Ecol. 2012, 81, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Lambert, J.-F.; Stievano, L.; Lopes, I.; Gharsallah, M.; Piao, L. The fate of amino acids adsorbed on mineral matter. Planet. Space Sci. 2009, 57, 460–467. [Google Scholar] [CrossRef]
- Lambert, J.F. Adsorption and polymerization of amino acids on mineral surfaces: A review. Orig. Life Evol. Biosph. 2008, 38, 211–242. [Google Scholar] [CrossRef]
- Hazen, R.M.; Sverjensky, D.A. Mineral Surfaces, Geochemical Complexities, and the Origins of Life. Cold Spring Harb. Perspect. Biol. 2010, 2. [Google Scholar] [CrossRef]
- Carson, J.K.; Campbell, L.; Rooney, D.; Clipson, N.; Gleeson, D.B. Minerals in soil select distinct bacterial communities in their microhabitats. FEMS Microbiol. Ecol. 2009, 67, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.A.; Bennett, P.C. Mineral Microniches Control the Diversity of Subsurface Microbial Populations. Geomicrobiol. J. 2014, 31, 246–261. [Google Scholar] [CrossRef]
- Hassink, J. The capacity of soils to preserve organic C and N by their association with clay and silt particles. Plant Soil 1997, 191, 77–87. [Google Scholar] [CrossRef]
- Martins, Z. In situ biomarkers and the Life Marker Chip. Astron. Geophys. 2011, 52, 34–35. [Google Scholar] [CrossRef]
- Gallori, E. Astrochemistry and the origin of genetic material. Rendiconti Lincei 2011, 22, 113–118. [Google Scholar] [CrossRef]
- Röling, W.F.M.; Head, I.M. Prokaryotic Systematics: PCR and Sequence Analysis of Amplified 16S rRNA Genes. In Molecular Microbiology Ecology; Garland Science: New York, NY, USA, 2005. [Google Scholar]
- Saeki, K.; Sakai, M. The Influence of Soil Organic Matter on DNA Adsorptions on Andosols. Microbes Environ. 2009, 24, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Aardema, B.W.; Lorenz, M.G.; Krumbein, W.E. Protection of Sediment-Adsorbed Transforming DNA Against Enzymatic Inactivation. Appl. Environ. Microbiol. 1983, 46, 417–420. [Google Scholar] [PubMed]
- Scappini, F.; Casadei, F.; Zamboni, R.; Franchi, M.; Gallori, E.; Monti, S. Protective effect of clay minerals on adsorbed nucleic acid against UV radiation: possible role in the origin of life. Int. J. Astrobiol. 2004, 3, 17–19. [Google Scholar] [CrossRef]
- Ciaravella, A.; Scappini, F.; Franchi, M.; Cecchi-Pestellini, C.; Barbera, M.; Candia, R.; Gallori, E.; Micela, G. Role of clays in protecting adsorbed DNA against X-ray radiation. Int. J. Astrobiol. 2004, 3, 31–35. [Google Scholar] [CrossRef]
- Lindahl, T. Instability and decay of the primary structure of DNA. Nature 1993, 362, 709–715. [Google Scholar] [CrossRef] [PubMed]
- Wayne, R.K.; Leonard, J.A.; Cooper, A. Full of Sound and Fury: History of Ancient DNA. Ann. Rev. Ecol. Syst. 1999, 30, 457–477. [Google Scholar] [CrossRef]
- Vreeland, R.H.; Rosenzweig, W.D.; Powers, D.W. Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal. Nature 2000, 407, 897–900. [Google Scholar] [CrossRef] [PubMed]
- Panieri, G.; Lugli, S.; Manzi, V.; Roveri, M.; Schreiber, B.C.; Palinska, K.A. Ribosomal RNA gene fragments from fossilized cyanobacteria identified in primary gypsum from the late Miocene, Italy. Geobiology 2010, 8, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Sephton, M.A. Organic geochemistry and the exploration of Mars. J. Cosmol. 2010, 5, 1141–1149. [Google Scholar]
- Mullis, K.; Faloona, F.; Scharf, S.; Saiki, R.; Horn, G.; Erlich, H. Specific enzymatic amplification of DNA in vitro: The polymerase chain reaction. Cold Spring Harb. Symp. Quant. Biol. 1986, 51, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Callahan, M.P.; Smith, K.E.; Cleaves, H.J.; Ruzicka, J.; Stern, J.C.; Glavin, D.P.; House, C.H.; Dworkin, J.P. Carbonaceous meteorites contain a wide range of extraterrestrial nucleobases. Proc. Natl. Acad. Sci. USA 2011, 108, 13995–13998. [Google Scholar] [CrossRef] [PubMed]
- Martins, Z.; Botta, O.; Fogel, M.L.; Sephton, M.A.; Glavin, D.P.; Watson, J.S.; Dworkin, J.P.; Schwartz, A.W.; Ehrenfreund, P. Extraterrestrial nucleobases in the Murchison meteorite. Earth Planet. Sci. Lett. 2008, 270, 130–136. [Google Scholar] [CrossRef]
- Yu, H.; Zhang, S.; Chaput, J.C. Darwinian evolution of an alternative genetic system provides support for TNA as an RNA progenitor. Nat. Chem. 2012, 4, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Ourisson, G.; Albrecht, P.; Rohmer, M. Predictive microbial biochemistry—from molecular fossils to procaryotic membranes. Trends Biochem. Sci. 1982, 7, 236–239. [Google Scholar]
- Eigenbrode, J. Fossil Lipids for Life-Detection: A Case Study from the Early Earth Record. Space Sci. Rev. 2008, 135, 161–185. [Google Scholar] [CrossRef]
- Georgiou, C.D.; Deamer, D.W. Lipids as universal biomarkers of extraterrestrial life. Astrobiology 2014, 14, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.S.; Miller, M.R.; Davies, N.W.; Goodchild, A.; Raftery, M.; Cavicchioli, R. Cold adaptation in the Antarctic Archaeon Methanococcoides burtonii involves membrane lipid unsaturation. J. Bacteriol. 2004, 186, 8508–8515. [Google Scholar] [CrossRef] [PubMed]
- Cook, H.W.; McMaster, C.R. Fatty Acid Desaturation and Chain Elongation in Eukaryotes. In Biochemistry of Lipids, Lipoproteins and Membranes; Vance, J.E., Vance, D.E., Eds.; Elsevier: New York, NY, USA, 2002. [Google Scholar]
- Kaneda, T. Iso- and anteiso-fatty acids in bacteria: Biosynthesis, function, and taxonomic significance. Microbiol. Rev. 1991, 55, 288–302. [Google Scholar] [PubMed]
- Simoneit, B.R.T. Molecular indicators (biomarkers) of past life. Anat. Rec. 2002, 268, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Hebting, Y.; Schaeffer, P.; Behrens, A.; Adam, P.; Schmitt, G.; Schneckenburger, P.; Bernasconi, S.M.; Albrecht, P. Biomarker Evidence for a Major Preservation Pathway of Sedimentary Organic Carbon. Science 2006, 312, 1627–1631. [Google Scholar] [CrossRef] [PubMed]
- Tegelaar, E.W.; de Leeuw, J.W.; Derenne, S.; Largeau, C. A reappraisal of kerogen formation. Geochimica et Cosmochimica Acta 1989, 53, 3103–3106. [Google Scholar] [CrossRef]
- Das, S.K.; Harris, R.S. Lipids and Fatty Acids in Fossil Teeth. J. Dent. Res. 1970, 49, 126–130. [Google Scholar] [CrossRef] [PubMed]
- Guido, A.; Jacob, J.; Gautret, P.; Laggoun-Défarge, F.; Mastandrea, A.; Russo, F. Molecular fossils and other organic markers as palaeoenvironmental indicators of the Messinian Calcare di Base Formation: Normal versus stressed marine deposition (Rossano Basin, northern Calabria, Italy). Palaeogeography Palaeoclimatology Palaeoecology 2007, 255, 265–283. [Google Scholar] [CrossRef] [Green Version]
- Woese, C.R.; Kandler, O.; Wheelis, M.L. Towards a natural system of organisms: Proposal for the domains Archaea, Bacteria, and Eucarya. Proc. Natl. Acad. Sci. USA 1990, 87, 4576–4579. [Google Scholar] [CrossRef] [PubMed]
- Botta, O.; Bada, J. Extraterrestrial Organic Compounds in Meteorites. Surv. Geophys. 2002, 23, 411–467. [Google Scholar] [CrossRef]
- Tielens, A.G.G.M. Interstellar Polycyclic Aromatic Hydrocarbon Molecules. Annu. Rev. Astron. Astrophys. 2008, 46, 289–337. [Google Scholar] [CrossRef]
- Ehrenfreund, P.; Sephton, M.A. Carbon molecules in space: from astrochemistry to astrobiology. Faraday Discuss. 2006, 133, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Sephton, M.A.; Botta, O. Recognizing life in the Solar System: guidance from meteoritic organic matter. Int. J. Astrobiol. 2005, 4, 269–276. [Google Scholar] [CrossRef]
- Van Hoeven, W.; Maxwell, J.R.; Calvin, M. Fatty acids and hydrocarbons as evidence of life processes in ancient sediments and crude oils. Geochimica et Cosmochimica Acta 1969, 33, 877–881. [Google Scholar] [CrossRef]
- Wagner, I.; Musso, H. New Naturally Occurring Amino Acids. Angewandte Chemie 1983, 22, 816–828. [Google Scholar] [CrossRef]
- Creighton, T.E. Proteins: Structures and Molecular Properties; W.H. Freeman: San Francisco, CA, USA, 1993. [Google Scholar]
- Churchill, H.; Teng, H.; Hazen, R.M. Correlation of pH-dependent surface interaction forces to amino acid adsorption: Implications for the origin of life. Am. Mineral. 2004, 89, 1048–1055. [Google Scholar]
- Ben-Taleb, A.; Vera, P.; Delgado, A.V.; Gallardo, V. Electrokinetic studies of monodisperse hematite particles: Effects of inorganic electrolytes and amino acids. Mater. Chem. Phys. 1994, 37, 68–75. [Google Scholar] [CrossRef]
- Meierhenrich, U. Amino Acids and the Asymetry of Life; Springer: Berlin/Heidelberg, Germany, 2008. [Google Scholar]
- Hazen, R.M.; Filley, T.R.; Goodfriend, G.A. Selective adsorption of l- and d-amino acids on calcite: Implications for biochemical homochirality. Proc. Natl. Acad. Sci. USA 2001, 98, 5487–5490. [Google Scholar] [CrossRef] [PubMed]
- Cerf, C.; Jorissen, A. Is amino-acid homochirality due to asymmetric photolysis in space? Space Sci. Rev. 2000, 92, 603–612. [Google Scholar]
- Bonner, W.A. Homochirality and life. EXS-BASEL 1998, 85, 159–188. [Google Scholar]
- Glavin, D.P.; Dworkin, J.P. Enrichment of the amino acid l-isovaline by aqueous alteration on CI and CM meteorite parent bodies. Proc. Natl. Acad. Sci. USA 2009, 106, 5487–5492. [Google Scholar] [CrossRef] [PubMed]
- Steendam, R.R.E.; Harmsen, B.; Meekes, H.; van Enckevort, W.J.P.; Kaptein, B.; Kellogg, R.M.; Raap, J.; Rutjes, F.P.J.T.; Vlieg, E. Controlling the Effect of Chiral Impurities on Viedma Ripening. Cryst. Growth Des. 2013, 13, 4776–4780. [Google Scholar] [CrossRef]
- Botta, O.; Martins, Z.; Ehrenfreund, P. Amino acids in Antarctic CM1 meteorites and their relationship to other carbonaceous chondrites. Meteorit. Planet. Sci. 2007, 42, 81–92. [Google Scholar] [CrossRef]
- Ehrenfreund, P.; Glavin, D.P.; Botta, O.; Cooper, G.; Bada, J.L. Extraterrestrial amino acids in Orgueil and Ivuna: Tracing the parent body of CI type carbonaceous chondrites. Proc. Natl. Acad. Sci. USA 2001, 98, 2138–2141. [Google Scholar] [CrossRef] [PubMed]
- Martins, Z.; Sephton, M.A.; Foing, B.H.; Ehrenfreund, P. Extraction of amino acids from soils close to the Mars Desert Research Station (MDRS), Utah. Int. J. Astrobiol. 2011, 10, 231–238. [Google Scholar] [CrossRef]
- Ten Kate, I.L.; Garry, J.R.C.; Peeters, Z.; Quinn, R.; Foing, B.; Ehrenfreund, P. Amino acid photostability on the Martian surface. Meteorit. Planet. Sci. 2005, 40, 1185–1193. [Google Scholar]
- Ten Kate, I.L.; Garry, J.R.C.; Peeters, Z.; Foing, B.; Ehrenfreund, P. The effects of Martian near surface conditions on the photochemistry of amino acids. Planet. Space Sci. 2006, 54, 296–302. [Google Scholar]
- Pollack, J.B.; Kasting, J.F.; Richardson, S.M.; Poliakoff, K. The case for a wet, warm climate on early Mars. Icarus 1987, 71, 203–224. [Google Scholar] [CrossRef] [PubMed]
- Carr, M.H. Water erosion on Mars and its biologic implications. Endeavour 1996, 20, 56–60. [Google Scholar] [CrossRef] [PubMed]
- Carr, M.H. Water on Early Mars. In Ciba Foundation Symposium 202—Evolution of Hydrothermal Ecosystems on Earth (And Mars?); Wiley: Hoboken, NJ, USA, 2007; pp. 249–272. [Google Scholar]
- Baker, V.R. Water and the martian landscape. Nature 2001, 412, 228–236. [Google Scholar] [CrossRef] [PubMed]
- Bibring, J.-P.; Langevin, Y.; Gendrin, A.; Gondet, B.; Poulet, F.; Berthé, M.; Soufflot, A.; Arvidson, R.; Mangold, N.; Mustard, J.; et al. Mars Surface Diversity as Revealed by the OMEGA/Mars Express Observations. Science 2005, 307, 1576–1581. [Google Scholar] [CrossRef] [PubMed]
- Squyres, S.W.; Grotzinger, J.P.; Arvidson, R.E.; Bell, J.F.; Calvin, W.; Christensen, P.R.; Clark, B.C.; Crisp, J.A.; Farrand, W.H.; Herkenhoff, K.E.; et al. In Situ Evidence for an Ancient Aqueous Environment at Meridiani Planum, Mars. Science 2004, 306, 1709–1714. [Google Scholar] [CrossRef] [PubMed]
- Tuff, J.; Wade, J.; Wood, B.J. Volcanism on Mars controlled by early oxidation of the upper mantle. Nature 2013, 498, 342–345. [Google Scholar] [CrossRef] [PubMed]
- Poulet, F.; Bibring, J.P.; Mustard, J.F.; Gendrin, A.; Mangold, N.; Langevin, Y.; Arvidson, R.E.; Gondet, B.; Gomez, C. Phyllosilicates on Mars and implications for early martian climate. Nature 2005, 438, 623–627. [Google Scholar] [PubMed]
- Ehlmann, B.L.; Mustard, J.F.; Murchie, S.L.; Bibring, J.-P.; Meunier, A.; Fraeman, A.A.; Langevin, Y. Subsurface water and clay mineral formation during the early history of Mars. Nature 2011, 479, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Malin, M.C.; Edgett, K.S. Evidence for Recent Groundwater Seepage and Surface Runoff on Mars. Science 2000, 288, 2330–2335. [Google Scholar] [CrossRef] [PubMed]
- Ojha, L.; McEwen, A.; Dundas, C.; Byrne, S.; Mattson, S.; Wray, J.; Masse, M.; Schaefer, E. HiRISE observations of Recurring Slope Lineae (RSL) during southern summer on Mars. Icarus 2014, 231, 365–376. [Google Scholar] [CrossRef]
- Benner, S.A.; Devine, K.G.; Matveeva, L.N.; Powell, D.H. The missing organic molecules on Mars. Proc. Natl. Acad. Sci. USA 2000, 97, 2425–2430. [Google Scholar] [CrossRef] [PubMed]
- Yen, A.S.; Kim, S.S.; Hecht, M.H.; Frant, M.S.; Murray, B. Evidence That the Reactivity of the Martian Soil Is Due to Superoxide Ions. Science 2000, 289, 1909–1912. [Google Scholar] [CrossRef] [PubMed]
- Hecht, M.H.; Kounaves, S.P.; Quinn, R.C.; West, S.J.; Young, S.M.M.; Ming, D.W.; Catling, D.C.; Clark, B.C.; Boynton, W.V.; Hoffman, J.; et al. Detection of Perchlorate and the Soluble Chemistry of Martian Soil at the Phoenix Lander Site. Science 2009, 325, 64–67. [Google Scholar]
- Hassler, D.M.; Zeitlin, C.; Wimmer-Schweingruber, R.F.; Ehresmann, B.; Rafkin, S.; Eigenbrode, J.L.; Brinza, D.E.; Weigle, G.; Böttcher, S.; Böhm, E.; et al. Mars’ Surface Radiation Environment Measured with the Mars Science Laboratory’s Curiosity Rover. Science 2014, 343. [Google Scholar] [CrossRef] [PubMed]
- Committee on the Planetary Science Decadal SurveyNational Research CouncilMars: Evolution of an Earth-Like World. In Vision and Voyages for Planetary Science in the Decade 2013–2022; National Academies Press: Washington, DC, USA, 2011.
- Cushing, G.E.; Titus, T.N.; Wynne, J.J.; Christensen, P.R. THEMIS observes possible cave skylights on Mars. Geophys. Res. Lett. 2007, 34, L17201. [Google Scholar] [CrossRef]
- Navarro-González, R.; Vargas, E.; de la Rosa, J.; Raga, A.C.; McKay, C.P. Reanalysis of the Viking results suggests perchlorate and organics at midlatitudes on Mars. J. Geophys. Planet. 2010, 115, E12010. [Google Scholar] [CrossRef]
- Biemann, K.; Oro, J.; Toulmin, P.; Orgel, L.E.; Nier, A.O.; Anderson, D.M.; Simmonds, P.G.; Flory, D.; Diaz, A.V.; Rushneck, D.R.; et al. The search for organic substances and inorganic volatile compounds in the surface of Mars. J. Geophys. Res. 1977, 82, 4641–4658. [Google Scholar] [CrossRef]
- Keller, J.M.; Boynton, W.V.; Karunatillake, S.; Baker, V.R.; Dohm, J.M.; Evans, L.G.; Finch, M.J.; Hahn, B.C.; Hamara, D.K.; Janes, D.M.; et al. Equatorial and midlatitude distribution of chlorine measured by Mars Odyssey GRS. J. Geophys. Res. Planet. 2006, 111, E03S08. [Google Scholar] [CrossRef]
- Glavin, D.P.; Freissinet, C.; Miller, K.E.; Eigenbrode, J.L.; Brunner, A.E.; Buch, A.; Sutter, B.; Archer, P.D.; Atreya, S.K.; Brinckerhoff, W.B.; et al. Evidence for perchlorates and the origin of chlorinated hydrocarbons detected by SAM at the Rocknest aeolian deposit in Gale Crater. J. Geophys. Res. Planet. 2013, 118, 1955–1973. [Google Scholar] [CrossRef]
- De Vera, J.-P.; Dulai, S.; Kereszturi, A.; Koncz, L.; Lorek, A.; Mohlmann, D.; Marschall, M.; Pocs, T. Results on the survival of cryptobiotic cyanobacteria samples after exposure to Mars-like environmental conditions. Int. J. Astrobiol. 2014, 13, 35–44. [Google Scholar]
- De Vera, J.P.; Mohlmann, D.; Butina, F.; Lorek, A.; Wernecke, R.; Ott, S. Survival potential and photosynthetic activity of lichens under Mars-like conditions: A laboratory study. Astrobiology 2010, 10, 215–227. [Google Scholar] [CrossRef] [PubMed]
- Schirmack, J.; Böhm, M.; Brauer, C.; Löhmannsröben, H.-G.; de Vera, J.-P.; Möhlmann, D.; Wagner, D. Laser spectroscopic real time measurements of methanogenic activity under simulated Martian subsurface analog conditions. Planet. Space Sci. 2014, 98, 198–204. [Google Scholar] [CrossRef]
- Friedmann, E.I. Endolithic Microorganisms in the Antarctic Cold Desert. Science 1982, 215, 1045–1053. [Google Scholar] [CrossRef] [PubMed]
- Boison, G.; Mergel, A.; Jolkver, H.; Bothe, H. Bacterial Life and Dinitrogen Fixation at a Gypsum Rock. Appl. Environ. Microbiol. 2004, 70, 7070–7077. [Google Scholar] [CrossRef] [PubMed]
- Lester, E.D.; Satomi, M.; Ponce, A. Microflora of extreme arid Atacama Desert soils. Soil Biol. Biochem. 2007, 39, 704–708. [Google Scholar] [CrossRef]
- Navarro-Gonzalez, R.; Iniguez, E.; de la Rosa, J.; McKay, C.P. Characterization of organics, microorganisms, desert soils, and Mars-like soils by thermal volatilization coupled to mass spectrometry and their implications for the search for organics on Mars by Phoenix and future space missions. Astrobiology 2009, 9, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Navarro-González, R.; Rainey, F.A.; Molina, P.; Bagaley, D.R.; Hollen, B.J.; de la Rosa, J.; Small, A.M.; Quinn, R.; Grunthaner, F.J.; Cáceres, L.; et al. Mars-Like Soils in the Atacama Desert, Chile, and the Dry Limit of Microbial Life. Science 2003, 302, 1018–1021. [Google Scholar] [PubMed]
- Direito, S.O.L.; Ehrenfreund, P.; Marees, A.; Staats, M.; Foing, B.; Röling, W.F.M. A wide variety of putative extremophiles and large beta-diversity at the Mars Desert Research Station (Utah). Int. J. Astrobiol. 2011, 10, 191–207. [Google Scholar] [CrossRef]
- Carson, J.K.; Gonzalez-Quiñones, V.; Murphy, D.V.; Hinz, C.; Shaw, J.A.; Gleeson, D.B. Low Pore Connectivity Increases Bacterial Diversity in Soil. Appl. Environ. Microbiol. 2010, 76, 3936–3942. [Google Scholar] [CrossRef] [PubMed]
- Parro, V.; de Diego-Castilla, G.; Moreno-Paz, M.; Blanco, Y.; Cruz-Gil, P.; Rodriguez-Manfredi, J.A.; Fernandez-Remolar, D.; Gomez, F.; Gomez, M.J.; Rivas, L.A.; et al. A microbial oasis in the hypersaline Atacama subsurface discovered by a life detector chip: Implications for the search for life on Mars. Astrobiology 2011, 11, 969–996. [Google Scholar] [CrossRef] [PubMed]
- Stivaletta, N.; López-García, P.; Boihem, L.; Millie, D.F.; Barbieri, R. Biomarkers of Endolithic Communities within Gypsum Crusts (Southern Tunisia). Geomicrobiol. J. 2010, 27, 101–110. [Google Scholar] [CrossRef]
- Pedersen, K. Exploration of deep intraterrestrial microbial life: Current perspectives. FEMS Microbiol. Lett. 2000, 185, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Roberts, J.A. Inhibition and enhancement of microbial surface colonization: The role of silicate composition. Chem. Geol. 2004, 212, 313–327. [Google Scholar] [CrossRef]
- Lengeler, J.W.; Drews, G.; Schlegel, H.G. Front Matter. In Biology of the Prokaryotes; Wiley: River Street Hoboken, NJ, USA, 2009; pp. 234–257. [Google Scholar]
- Sarbu, S.M.; Kane, T.C.; Kinkle, B.K. A Chemoautotrophically Based Cave Ecosystem. Science 1996, 272, 1953–1955. [Google Scholar] [CrossRef] [PubMed]
- Sarbu, S.M.; Kinkle, B.K.; Vlasceanu, L.; Kane, T.C.; Popa, R. Microbiological characterization of a sulfide-rich groundwater ecosystem. Geomicrobiol. J. 1994, 12, 175–182. [Google Scholar] [CrossRef]
- Hutchens, E.; Radajewski, S.; Dumont, M.G.; McDonald, I.R.; Murrell, J.C. Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environ. Microbiol. 2004, 6, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wu, L.; Boden, R.; Hillebrand, A.; Kumaresan, D.; Moussard, H.; Baciu, M.; Lu, Y.; Colin Murrell, J. Life without light: microbial diversity and evidence of sulfur- and ammonium-based chemolithotrophy in Movile Cave. ISME J. 2009, 3, 1093–1104. [Google Scholar] [CrossRef] [PubMed]
- McGenity, T.J.; Gemmell, R.T.; Grant, W.D. Proposal of a new halobacterial genus Natrinema gen. nov., with two species Natrinema pellirubrum nom. nov. and Natrinema pallidum nom. nov. Int. J. Syst. Bacteriol. 1998, 48, 1187–1196. [Google Scholar] [CrossRef] [PubMed]
- McGenity, T.J.; Gemmell, R.T.; Grant, W.D.; Stan-Lotter, H. Origins of halophilic microorganisms in ancient salt deposits. Environ. Microbiol. 2000, 2, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Lide, D.R. CRC Handbook of Chemistry and Physics; CRC press: Boca Raton, FL, USA, 2004. [Google Scholar]
- Tanaka, T.; Burgess, J.; Wright, P. High-pressure adaptation by salt stress in a moderately halophilic bacterium obtained from open seawater. Appl. Microbiol. Biotechnol. 2001, 57, 200–204. [Google Scholar] [CrossRef] [PubMed]
- Tedrow, J.C.F. Antarctic Soils and Soil Forming Processes; Antarctic Research Series; the American Geophysical Union: Washington, DC, USA, 2013; pp. 125–159. [Google Scholar]
- Heldmann, J.L.; Pollard, W.; McKay, C.P.; Marinova, M.M.; Davila, A.; Williams, K.E.; Lacelle, D.; Andersen, D.T. The high elevation Dry Valleys in Antarctica as analog sites for subsurface ice on Mars. Planet. Space Sci. 2013, 85, 53–58. [Google Scholar] [CrossRef]
- Brass, G.W. Stability of brines on Mars. Icarus 1980, 42, 20–28. [Google Scholar]
- Murray, A.E.; Kenig, F.; Fritsen, C.H.; McKay, C.P.; Cawley, K.M.; Edwards, R.; Kuhn, E.; McKnight, D.M.; Ostrom, N.E.; Peng, V.; et al. Microbial life at −13 °C in the brine of an ice-sealed Antarctic lake. Proc. Natl. Acad. Sci. USA 2012, 109, 20626–20631. [Google Scholar] [CrossRef] [PubMed]
- Kivelson, M.G.; Khurana, K.K.; Russell, C.T.; Volwerk, M.; Walker, R.J.; Zimmer, C. Galileo Magnetometer Measurements: A Stronger Case for a Subsurface Ocean at Europa. Science 2000, 289, 1340–1343. [Google Scholar] [CrossRef] [PubMed]
- Kuskov, O.L.; Kronrod, V.A. Internal structure of Europa and Callisto. Icarus 2005, 177, 550–569. [Google Scholar] [CrossRef]
- Showman, A.P.; Malhotra, R. The Galilean Satellites. Science 1999, 286, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Khurana, K.K.; Kivelson, M.G.; Stevenson, D.J.; Schubert, G.; Russell, C.T.; Walker, R.J.; Polanskey, C. Induced magnetic fields as evidence for subsurface oceans in Europa and Callisto. Nature 1998, 395, 777–780. [Google Scholar] [CrossRef] [PubMed]
- Mikucki, J.A.; Pearson, A.; Johnston, D.T.; Turchyn, A.V.; Farquhar, J.; Schrag, D.P.; Anbar, A.D.; Priscu, J.C.; Lee, P.A. A Contemporary Microbially Maintained Subglacial Ferrous “Ocean”. Science 2009, 324, 397–400. [Google Scholar] [CrossRef] [PubMed]
- Doran, P.T.; Fritsen, C.H.; McKay, C.P.; Priscu, J.C.; Adams, E.E. Formation and character of an ancient 19-m ice cover and underlying trapped brine in an “ice-sealed” east Antarctic lake. Proc. Natl. Acad. Sci. USA 2003, 100, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Bell, R.E.; Studinger, M.; Tikku, A.A.; Clarke, G.K.C.; Gutner, M.M.; Meertens, C. Origin and fate of Lake Vostok water frozen to the base of the East Antarctic ice sheet. Nature 2002, 416, 307–310. [Google Scholar] [CrossRef]
- Bridges, J.C.; Grady, M.M. Evaporite mineral assemblages in the nakhlite (martian) meteorites. Earth Planet. Sci. Lett. 2000, 176, 267–279. [Google Scholar] [CrossRef]
- Weiss, B.P.; Kirschvink, J.L.; Baudenbacher, F.J.; Vali, H.; Peters, N.T.; Macdonald, F.A.; Wikswo, J.P. A Low Temperature Transfer of ALH84001 from Mars to Earth. Science 2000, 290, 791–795. [Google Scholar] [CrossRef] [PubMed]
- Folinsbee, R.E.; Douglas, J.A.V.; Maxwell, J.A. Revelstoke, a new Type I carbonaceous chondrite. Geochimica et Cosmochimica Acta 1967, 31, 1625–1635. [Google Scholar] [CrossRef]
- Hoover, R.B. Microfossils of cyanobacteria in carbonaceous meteorites. Proc. SPIE 2007, 6694. [Google Scholar] [CrossRef]
- Tipler, F.J. Discovery of Cyanobacteria in Meteorites: Implications for Astrobiology and Cosmology. Available online: http://journalofcosmology.com/Life101.html#17 (accessed on 17 September 2014).
- Wickramasinghe, N.C.; Wallis, J.; Wallis, D.H.; Samaranayake, A. Fossil diatoms in a new carbonaceous meteorite. Available online: http://arxiv.org/abs/1303.2398 (accessed on 17 September 2014).
- Sephton, M.A. Organic compounds in carbonaceous meteorites. Nat. Prod. Rep. 2002, 19, 292–311. [Google Scholar] [CrossRef] [PubMed]
- Cronin, J.R.; Pizzarello, S.; Epstein, S.; Krishnamurthy, R.V. Molecular and isotopic analyses of the hydroxy acids, dicarboxylic acids, and hydroxydicarboxylic acids of the Murchison meteorite. Geochimica et Cosmochimica Acta 1993, 57, 4745–4752. [Google Scholar] [CrossRef] [PubMed]
- Botta, O.; Martins, Z.; Emmenegger, C.; Dworkin, J.P.; Glavin, D.P.; Harvey, R.P.; Zenobi, R.; Bada, J.L.; Ehrenfreund, P. Polycyclic aromatic hydrocarbons and amino acids in meteorites and ice samples from LaPaz Icefield, Antarctica. Meteorit. Planet. Sci. 2008, 43, 1465–1480. [Google Scholar] [CrossRef]
- Martins, Z.; Alexander, C.M.O.D.; Orzechowska, G.E.; Fogel, M.L.; Ehrenfreund, P. Indigenous amino acids in primitive CR meteorites. Meteorit. Planet. Sci. 2007, 42, 2125–2136. [Google Scholar] [CrossRef]
- Cronin, J.; Chang, S. Organic Matter in Meteorites: Molecular and Isotopic Analyses of the Murchison Meteorite. In The Chemistry of Life’s Origins; Greenberg, J.M., Mendoza-Gómez, C.X., Pirronello, V., Eds.; Springer: Berlin/Heidelberg, Germany, 1993; Volume 416, pp. 209–258. [Google Scholar]
- Chyba, C.; Sagan, C. Endogenous production, exogenous delivery and impact-shock synthesis of organic molecules: An inventory for the origins of life. Nature 1992, 355, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Elsila, J.E.; Glavin, D.P.; Dworkin, J.P. Cometary glycine detected in samples returned by Stardust. Meteorit. Planet. Sci. 2009, 44, 1323–1330. [Google Scholar] [CrossRef]
- Sandford, S.A.; Aléon, J.; Alexander, C.M.O.D.; Araki, T.; Bajt, S.; Baratta, G.A.; Borg, J.; Bradley, J.P.; Brownlee, D.E.; Brucato, J.R.; et al. Organics Captured from Comet 81P/Wild 2 by the Stardust Spacecraft. Science 2006, 314, 1720–1724. [Google Scholar] [CrossRef] [PubMed]
- Department of Earth, Environmental and Planetary Sciences at Case Western Reserve University, ANSMET. Available online: http://artscilabs.case.edu/ansmet/ (accessed on 25 June 2014).
- Hedges, J.I.; Hare, P.E. Amino acid adsorption by clay minerals in distilled water. Geochimica et Cosmochimica Acta 1987, 51, 255–259. [Google Scholar] [CrossRef]
- Friebele, E.; Shimoyama, A.; Hare, P.E.; Ponnamperuma, C. Adsorption of amino acid entantiomers by Na-montmorillonite. Orig. Life Evol. Biosph. 1981, 11, 173–184. [Google Scholar] [CrossRef]
- Kennedy, M.J.; Pevear, D.R.; Hill, R.J. Mineral Surface Control of Organic Carbon in Black Shale. Science 2002, 295, 657–660. [Google Scholar] [CrossRef] [PubMed]
- Ehlmann, B.L.; Mustard, J.F.; Fassett, C.I.; Schon, S.C.; Head Iii, J.W.; Des Marais, D.J.; Grant, J.A.; Murchie, S.L. Clay minerals in delta deposits and organic preservation potential on Mars. Nat. Geosci. 2008, 1, 355–358. [Google Scholar] [CrossRef]
- Wattel-Koekkoek, E.J.W.; Buurman, P.; van der Plicht, J.; Wattel, E.; van Breemen, N. Mean residence time of soil organic matter associated with kaolinite and smectite. Eur. J. Soil Sci. 2003, 54, 269–278. [Google Scholar] [CrossRef]
- Beaty, D.W.; Miller, S.; Zimmerman, W.; Bada, J.; Conrad, P.; Dupuis, E.; Huntsberger, T.; Ivlev, R.; Kim, S.S.; Lee, B.G.; et al. Planning for a Mars in situ sample preparation and distribution (SPAD) system. Planet. Space Sci. 2004, 52, 55–66. [Google Scholar] [CrossRef]
- Bowden, S.A.; Wilson, R.; Taylor, C.; Cooper, J.M.; Parnell, J. The extraction of intracrystalline biomarkers and other organic compounds from sulphate minerals using a microfluidic format—a feasibility study for remote fossil-life detection using a microfluidic H-cell. Int. J. Astrobiol. 2007, 6, 27–36. [Google Scholar] [CrossRef]
- Shen, Y.; Buick, R.; Canfield, D.E. Isotopic evidence for microbial sulphate reduction in the early Archaean era. Nature 2001, 410, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Schulze, W.X.; Gleixner, G.; Kaiser, K.; Guggenberger, G.; Mann, M.; Schulze, E.D. A proteomic fingerprint of dissolved organic carbon and of soil particles. Oecologia 2005, 142, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Court, R.W.; Baki, A.O.; Sims, M.R.; Cullen, D.; Sephton, M.A. Novel solvent systems for in situ extraterrestrial sample analysis. Planet. Space Sci. 2010, 58, 1470–1474. [Google Scholar] [CrossRef] [Green Version]
- Court, R.W.; Baki, A.O.; Sims, M.R.; Cullen, D.; Sephton, M.A. The Life Marker Chip-Extracting Polar and Nonpolar Biomarkers from the Martian Soil Using a Surfactant-Based Solvent. In Proceedings of 74th Annual Meeting of the Meteoritical Society, London, UK, 8–12 August 2011; Wiley: Hoboken, NJ, USA, 2001; Volume 46, p. A50. [Google Scholar]
- Sims, M.R.; Cullen, D.C.; Rix, C.S.; Buckley, A.; Derveni, M.; Evans, D.; Miguel García-Con, L.; Rhodes, A.; Rato, C.C.; Stefinovic, M.; et al. Development status of the life marker chip instrument for ExoMars. Planet. Space Sci. 2012, 72, 129–137. [Google Scholar] [CrossRef]
- Hosseinzadeh, R.; Khorsandi, K.; Hemmaty, S. Study of the Effect of Surfactants on Extraction and Determination of Polyphenolic Compounds and Antioxidant Capacity of Fruits Extracts. PLoS One 2013, 8, e57353. [Google Scholar] [CrossRef] [PubMed]
- Uematsu, M.; Frank, E.U. Static Dielectric Constant of Water and Steam. J. Phys. Chem. Ref. Data 1980, 9, 1291–1306. [Google Scholar] [CrossRef]
- Amashukeli, X.; Pelletier, C.C.; Kirby, J.P.; Grunthaner, F.J. Subcritical water extraction of amino acids from Atacama Desert soils. J. Geophys. Res. Biogeosci. 2007, 112. [Google Scholar] [CrossRef]
- Hawthorne, S.B.; Miller, D.J. Direct Comparison of Soxhlet and Low- and High-Temperature Supercritical CO2 Extraction Efficiencies of Organics from Environmental Solids. Anal. Chem. 1994, 66, 4005–4012. [Google Scholar] [CrossRef]
- Hawthorne, S.B.; Grabanski, C.B.; Martin, E.; Miller, D.J. Comparisons of Soxhlet extraction, pressurized liquid extraction, supercritical fluid extraction and subcritical water extraction for environmental solids: Recovery, selectivity and effects on sample matrix. J. Chromatogr. A 2000, 892, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Parro, V.; de Diego-Castilla, G.; Rodriguez-Manfredi, J.A.; Rivas, L.A.; Blanco-Lopez, Y.; Sebastian, E.; Romeral, J.; Compostizo, C.; Herrero, P.L.; Garcia-Marin, A.; et al. SOLID3: A multiplex antibody microarray-based optical sensor instrument for in situ life detection in planetary exploration. Astrobiology 2011, 11, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Parro, V.; Fernandez-Calvo, P.; Rodriguez-Manfredi, J.A.; Moreno-Paz, M.; Rivas, L.A.; Garcia-Villadangos, M.; Bonaccorsi, R.; Gonzalez-Pastor, J.E.; Prieto-Ballesteros, O.; Schuerger, A.C.; et al. SOLID2: An antibody array-based life-detector instrument in a Mars Drilling Simulation Experiment (MARTE). Astrobiology 2008, 8, 987–999. [Google Scholar] [CrossRef] [PubMed]
- Snyder, L.R.; Kirkland, J.J.; Dolan, J.W. Basic Concepts and the Control of Separation. In Introduction to Modern Liquid Chromatography; Wiley: Hoboken, NJ, USA, 2010; pp. 19–86. [Google Scholar]
- Snyder, L.R.; Kirkland, J.J.; Dolan, J.W. Detection. In Introduction to Modern Liquid Chromatography; Wiley: Hoboken, NJ, USA, 2010; pp. 147–197. [Google Scholar]
- De Hoffmann, E. Mass Spectrometry. In Kirk-Othmer Encyclopedia of Chemical Technology; Wiley: Hoboken, NJ, USA, 2000. [Google Scholar]
- Bidlingmeyer, B.A.; Cohen, S.A.; Tarvin, T.L. Rapid analysis of amino acids using pre-column derivatization. J. Chromatogr. B Biomed. Sci. Appl. 1984, 336, 93–104. [Google Scholar] [CrossRef]
- Lindroth, P.; Mopper, K. High performance liquid chromatographic determination of subpicomole amounts of amino acids by precolumn fluorescence derivatization with o-phthaldialdehyde. Anal. Chem. 1979, 51, 1667–1674. [Google Scholar] [CrossRef]
- Einarsson, S.; Josefsson, B.; Moeller, P.; Sanchez, D. Separation of amino acid enantiomers and chiral amines using precolumn derivatization with (+)-1-(9-fluorenyl)ethyl chloroformate and reversed-phase liquid chromatography. Anal. Chem. 1987, 59, 1191–1195. [Google Scholar] [CrossRef] [PubMed]
- Pirkle, W.H.; Pochapsky, T.C.; Mahler, G.S.; Corey, D.E.; Reno, D.S.; Alessi, D.M. Useful and easily prepared chiral stationary phases for the direct chromatographic separation of the enantiomers of a variety of derivatized amines, amino acids, alcohols, and related compounds. J. Org. Chem. 1986, 51, 4991–5000. [Google Scholar] [CrossRef]
- McNair, H.M.; Miller, J.M. Basic Gas Chromatography, 2nd ed.; Wiley: Hoboken, NJ, USA, 2011. [Google Scholar]
- Ward, D.M.; Weller, R.; Bateson, M.M. 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature 1990, 345, 63–65. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, H.; Poly, F.; Van, V.T.; Lombard, N.; Nalin, R.; Vogel, T.M.; Simonet, P. High molecular weight DNA recovery from soils prerequisite for biotechnological metagenomic library construction. J. Microbiol. Methods 2005, 62, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rappe, M.S.; Giovannoni, S.J. The uncultured microbial majority. Annu. Rev. Microbiol. 2003, 57, 369–394. [Google Scholar] [CrossRef] [PubMed]
- Takada-Hoshino, Y.; Matsumoto, N. An Improved DNA Extraction Method Using Skim Milk from Soils That Strongly Adsorb DNA. Microbes Environ. 2004, 19, 13–19. [Google Scholar] [CrossRef]
- Huber, H.; Hohn, M.J.; Rachel, R.; Fuchs, T.; Wimmer, V.C.; Stetter, K.O. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 2002, 417, 63–67. [Google Scholar] [CrossRef] [PubMed]
- Binga, E.K.; Lasken, R.S.; Neufeld, J.D. Something from (almost) nothing: The impact of multiple displacement amplification on microbial ecology. ISME J. 2008, 2, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Schuster, S.C. Next-generation sequencing transforms today’s biology. Nat. Method 2008, 5, 16–18. [Google Scholar] [CrossRef]
- Mahaffy, P.; Webster, C.; Cabane, M.; Conrad, P.; Coll, P.; Atreya, S.; Arvey, R.; Barciniak, M.; Benna, M.; Bleacher, L.; et al. The Sample Analysis at Mars Investigation and Instrument Suite. Space Sci. Rev. 2012, 170, 401–478. [Google Scholar] [CrossRef]
- Grotzinger, J.P.; Sumner, D.Y.; Kah, L.C.; Stack, K.; Gupta, S.; Edgar, L.; Rubin, D.; Lewis, K.; Schieber, J.; Mangold, N.; et al. A Habitable Fluvio-Lacustrine Environment at Yellowknife Bay, Gale Crater, Mars. Science 2014, 343. [Google Scholar] [CrossRef] [PubMed]
- Leshin, L.A.; Mahaffy, P.R.; Webster, C.R.; Cabane, M.; Coll, P.; Conrad, P.G.; Archer, P.D., Jr.; Atreya, S.K.; Brunner, A.E.; Buch, A.; et al. Volatile, isotope, and organic analysis of martian fines with the Mars Curiosity rover. Science 2013, 341. [Google Scholar] [CrossRef]
- Evans-Nguyen, T.; Becker, L.; Doroshenko, V.; Cotter, R.J. Development of a low power, high mass range mass spectrometer for Mars surface analysis. Int. J. Mass Spectrom. 2008, 278, 170–177. [Google Scholar] [CrossRef]
- Jorge Villar, S.; Edwards, H.M. Raman spectroscopy in astrobiology. Anal. Bioanal. Chem. 2006, 384, 100–113. [Google Scholar] [CrossRef] [PubMed]
- Aubrey, A.D.; Chalmers, J.H.; Bada, J.L.; Grunthaner, F.J.; Amashukeli, X.; Willis, P.; Skelley, A.M.; Mathies, R.A.; Quinn, R.C.; Zent, A.P.; et al. The Urey instrument: an advanced in situ organic and oxidant detector for Mars exploration. Astrobiology 2008, 8, 583–595. [Google Scholar] [CrossRef] [PubMed]
- Direito, S.O.L.; Zaura, E.; Little, M.; Ehrenfreund, P.; Röling, W.F.M. Systematic evaluation of bias in microbial community profiles induced by whole genome amplification. Environ. Microbiol. 2014, 16, 643–657. [Google Scholar] [CrossRef] [PubMed]
- Skládal, P. Effect of methanol on the interaction of monoclonal antibody with free and immobilized atrazine studied using the resonant mirror-based biosensor. Biosens. Bioelectron. 1999, 14, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Ruigrok, V.J.B.; Levisson, M.; Eppink, M.H.M.; Smidt, H.; van der Oost, J. Alternative affinity tools: More attractive than antibodies? Biochem. J. 2011, 436, 1–13. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aerts, J.W.; Röling, W.F.M.; Elsaesser, A.; Ehrenfreund, P. Biota and Biomolecules in Extreme Environments on Earth: Implications for Life Detection on Mars. Life 2014, 4, 535-565. https://doi.org/10.3390/life4040535
Aerts JW, Röling WFM, Elsaesser A, Ehrenfreund P. Biota and Biomolecules in Extreme Environments on Earth: Implications for Life Detection on Mars. Life. 2014; 4(4):535-565. https://doi.org/10.3390/life4040535
Chicago/Turabian StyleAerts, Joost W., Wilfred F.M. Röling, Andreas Elsaesser, and Pascale Ehrenfreund. 2014. "Biota and Biomolecules in Extreme Environments on Earth: Implications for Life Detection on Mars" Life 4, no. 4: 535-565. https://doi.org/10.3390/life4040535
APA StyleAerts, J. W., Röling, W. F. M., Elsaesser, A., & Ehrenfreund, P. (2014). Biota and Biomolecules in Extreme Environments on Earth: Implications for Life Detection on Mars. Life, 4(4), 535-565. https://doi.org/10.3390/life4040535




