Computational Investigation Identifies mTOR as a Primary Binding Target of Medicarpin in Cholangiocarcinoma: Insights from Network Pharmacology and Molecular Docking
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemoinformatics, Drug Likeness, and ADME Prediction
2.2. Prediction of Target Proteins
2.3. Potential Targets Associated with Cholangiocarcinoma
2.4. Gene Ontology and Kyoto Encyclopaedia of Genes and Genomes Pathway Enrichment Analysis
2.5. Protein–Protein Interaction (PPI) Network Construction
2.6. Molecular Docking Studies Involving Medicarpin and Hub Genes
2.7. Molecular Docking (MD) Simulation
2.8. Survival Analysis of mTOR Expression in Cholangiocarcinoma Patients
3. Results
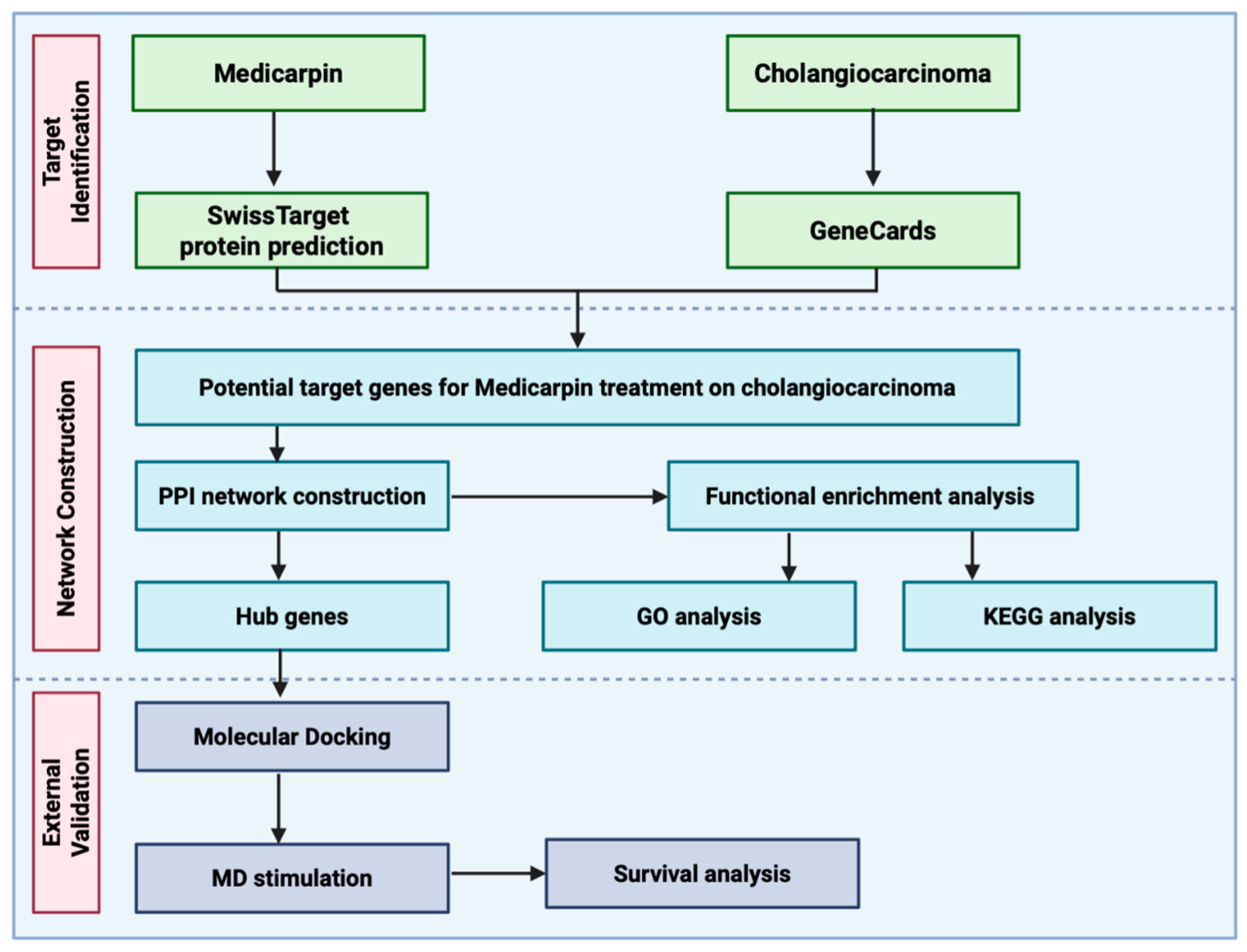
3.1. Procedure for Network Pharmacology Assessment of Medicarpin in Relation to Cholangiocarcinoma
3.2. Chemoinformatics, Drug-likeness, and ADME-Tox Profiling of Medicarpin
3.3. Identification of Targets and Analysis of Networks
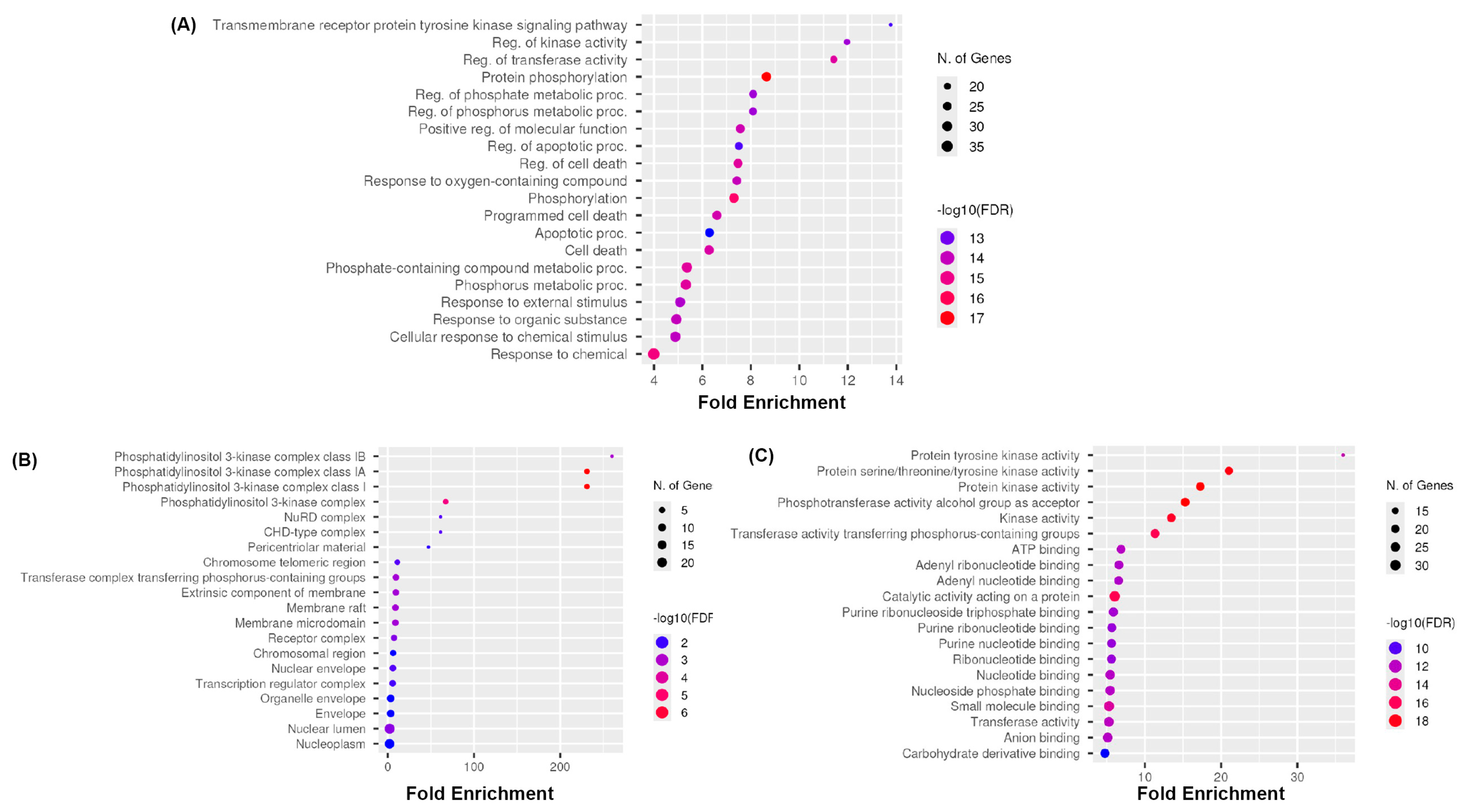
3.4. Gene Ontology Enrichment Analysis
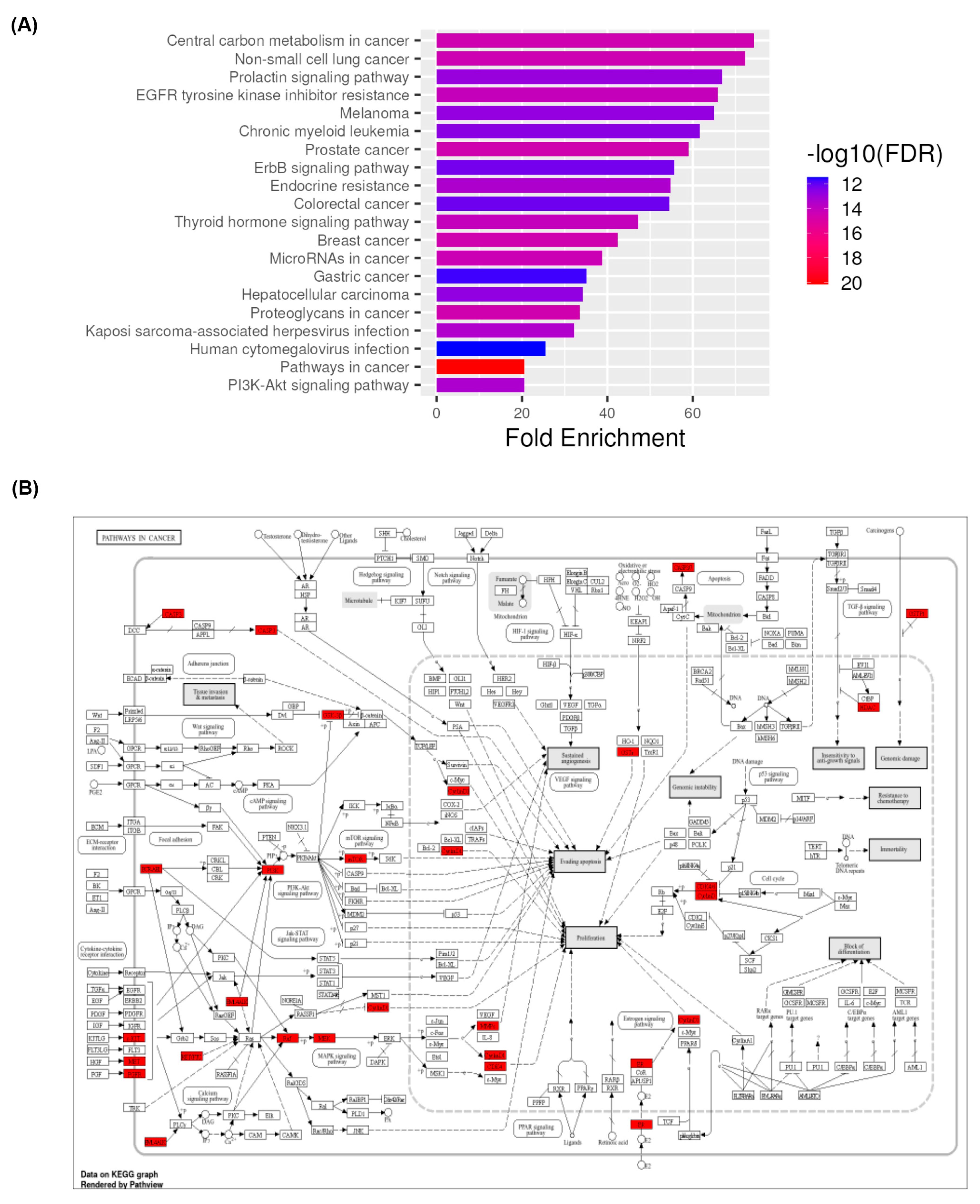
3.5. KEGG Pathway Enrichment Analysis
3.6. Mapping Pathways of Intersecting Targets
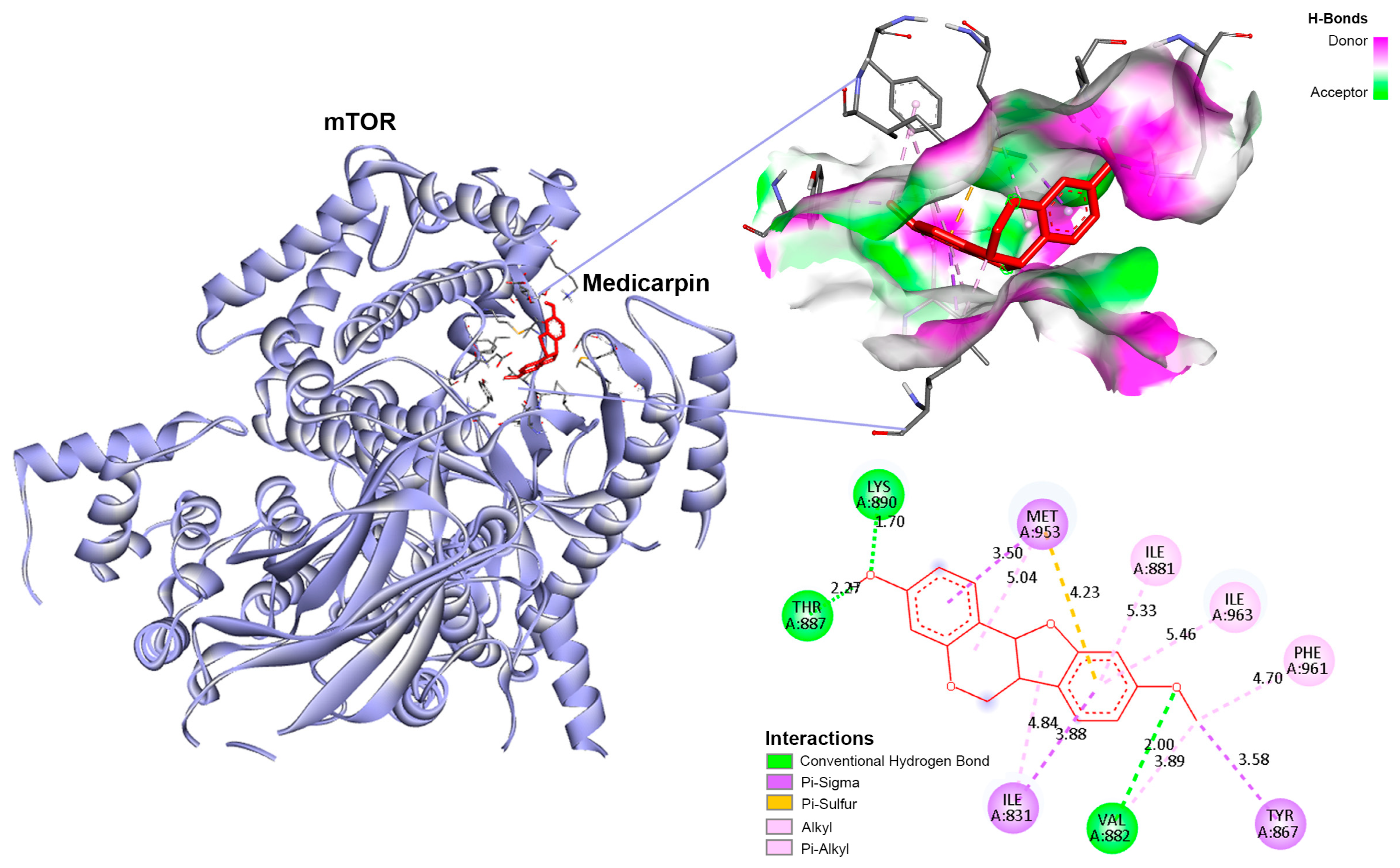
3.7. Validation of Hub Targets via Molecular Docking
3.8. Molecular Dynamics Simulation of the Medicarpin–mTOR Complex
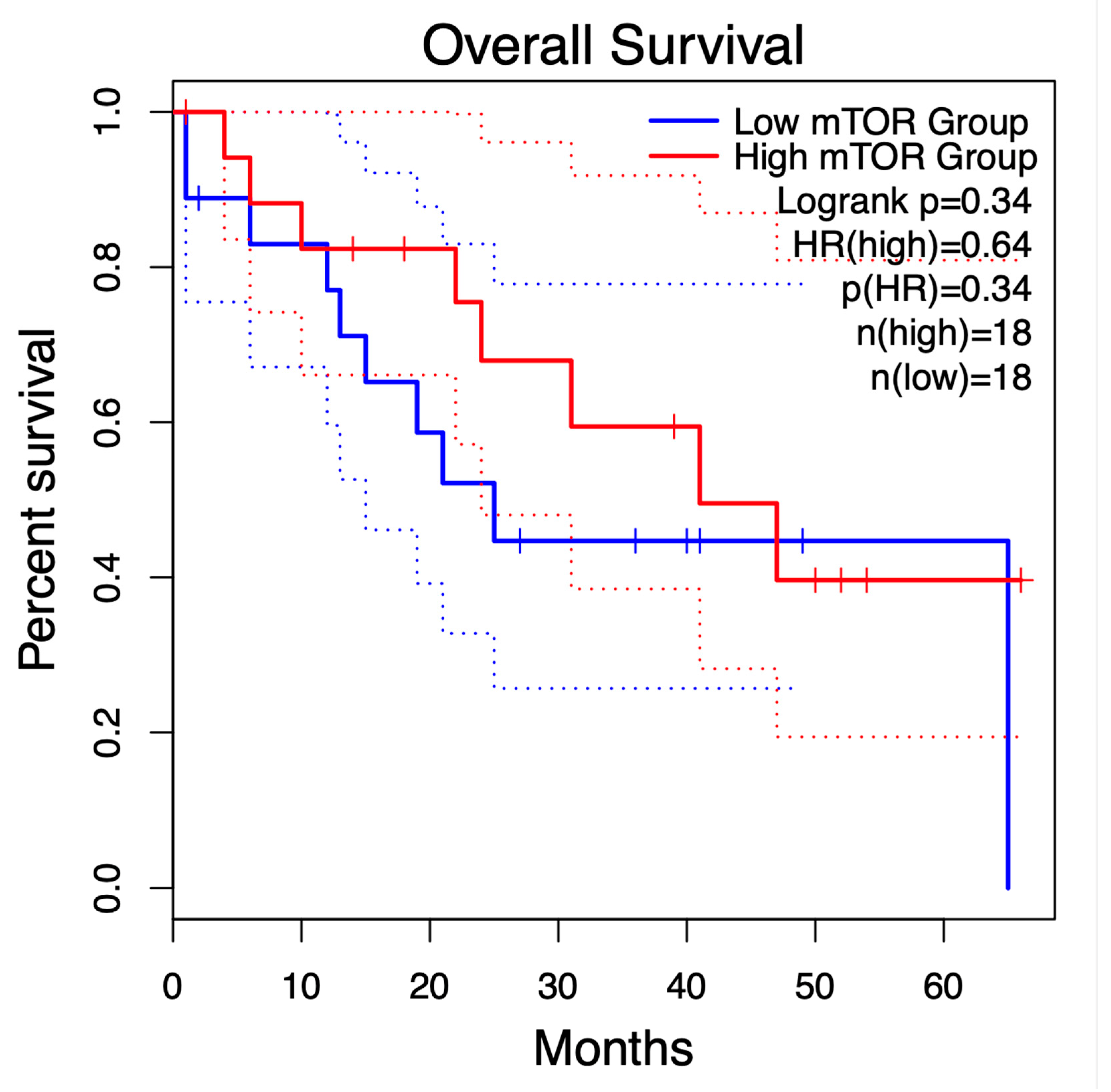
3.9. Prognostic Significance of mTOR Expression in Cholangiocarcinoma Patients
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
Abbreviations
| ADME | Absorption, Distribution, Metabolism, and Excretion |
| ADMET | Absorption, Distribution, Metabolism, Excretion, and Toxicity |
| Akt | Protein kinase B |
| AMP | Adenosine monophosphate |
| ATP | Adenosine triphosphate |
| BP | Biological Process |
| CCA | Cholangiocarcinoma |
| EGFR | Epidermal Growth Factor Receptor |
| EMT | Epithelial–Mesenchymal Transition |
| ER | Estrogen Receptor |
| ERα | Estrogen Receptor Alpha |
| FDR | False Discovery Rate |
| FGFR2 | Fibroblast Growth Factor Receptor 2 |
| GA | Genetic Algorithm |
| GO | Gene Ontology |
| HR | Hazard Ratio |
| IDH1 | Isocitrate Dehydrogenase 1 |
| KEGG | Kyoto Encyclopaedia of Genes and Genomes |
| LD50 | Median Lethal Dose |
| LogP | Logarithm of the Partition Coefficient (octanol/water) |
| LOAEL | Lowest Observed Adverse Effect Level |
| MAPK | Mitogen-Activated Protein Kinase |
| MD | Molecular Dynamics |
| MF | Molecular Function |
| mTOR | Mechanistic Target of Rapamycin |
| NADH | Nicotinamide Adenine Dinucleotide |
| NMDAR | N-Methyl-D-Aspartate Receptor |
| PI3K | Phosphatidylinositol 3-Kinase |
| PKA | Protein Kinase A |
| PPI | Protein–Protein Interaction |
| PTEN | Phosphatase and Tensin Homolog |
| RMSD | Root Mean Square Deviation |
| RMSF | Root Mean Square Fluctuation |
| TPSA | Topological Polar Surface Area |
| VEGFR | Vascular Endothelial Growth Factor Receptor |
References
- Kafle, A.; Suttiprapa, S. Current State of Knowledge on Blood and Tissue-Based Biomarkers for Opisthorchis viverrini-induced Cholangiocarcinoma: A Review of Prognostic, Predictive, and Diagnostic Markers. Asian Pac. J. Cancer Prev. 2024, 25, 25–41. [Google Scholar] [CrossRef]
- Tawarungruang, C.; Khuntikeo, N.; Chamadol, N.; Laopaiboon, V.; Thuanman, J.; Thinkhamrop, K.; Kelly, M.; Thinkhamrop, B. Survival after surgery among patients with cholangiocarcinoma in Northeast Thailand according to anatomical and morphological classification. BMC Cancer 2021, 21, 497. [Google Scholar] [CrossRef]
- Chen, Z.; Lin, T.; Liao, X.; Li, Z.; Lin, R.; Qi, X.; Chen, G.; Sun, L.; Lin, L. Network pharmacology based research into the effect and mechanism of Yinchenhao Decoction against Cholangiocarcinoma. Chin. Med. 2021, 16, 13. [Google Scholar] [CrossRef] [PubMed]
- Warren, E.A.K.; Maithel, S.K. Molecular pathology for cholangiocarcinoma: A review of actionable genetic targets and their relevance to adjuvant & neoadjuvant therapy, staging, follow-up, and determination of minimal residual disease. Hepatobiliary Surg. Nutr. 2024, 13, 29–38. [Google Scholar] [CrossRef]
- Song, F.; Lu, C.L.; Wang, C.G.; Hu, C.W.; Zhang, Y.; Wang, T.L.; Han, L.; Chen, Z. Uncovering the mechanism of Kang-ai injection for treating intrahepatic cholangiocarcinoma based on network pharmacology, molecular docking, and in vitro validation. Front. Pharmacol. 2023, 14, 1129709. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yu, J.; Zhang, Y.; Peng, C.; Song, Y.; Liu, S. Advances in targeted therapy of cholangiocarcinoma. Ann. Med. 2024, 56, 2310196. [Google Scholar] [CrossRef]
- Romero, D. Benefit from pemigatinib in cholangiocarcinoma. Nat. Rev. Clin. Oncol. 2020, 17, 337. [Google Scholar] [CrossRef] [PubMed]
- Abou-Alfa, G.K.; Macarulla, T.; Javle, M.M.; Kelley, R.K.; Lubner, S.J.; Adeva, J.; Cleary, J.M.; Catenacci, D.V.; Borad, M.J.; Bridgewater, J.; et al. Ivosidenib in IDH1-mutant, chemotherapy-refractory cholangiocarcinoma (ClarIDHy): A multicentre, randomised, double-blind, placebo-controlled, phase 3 study. Lancet Oncol. 2020, 21, 796–807. [Google Scholar] [CrossRef]
- Manogaran, P.; Beeraka, N.M.; Paulraj, R.S.; Sathiyachandran, P.; Thammaiappa, M. Impediment of Cancer by Dietary Plant-derived Alkaloids Through Oxidative Stress: Implications of PI3K/AKT Pathway in Apoptosis, Autophagy, and Ferroptosis. Curr. Top. Med. Chem. 2023, 23, 860–877. [Google Scholar] [CrossRef]
- Xie, S.; Zhan, F.; Zhu, J.; Xu, S.; Xu, J. The latest advances with natural products in drug discovery and opportunities for the future: A 2025 update. Expert Opin. Drug Discov. 2025, 20, 827–843. [Google Scholar] [CrossRef]
- Shang, X.; Dai, L.; Cao, X.; Ma, Y.; Gulnaz, I.; Miao, X.; Li, X.; Yang, X. Natural products in antiparasitic drug discovery: Advances, opportunities and challenges. Nat. Prod. Rep. 2025, 42, 1419–1458. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, F.; Sevindik, M.; Uysal, İ.; Şabik, A. Quercetin: Derivatives, biosynthesis, biological activity, pharmacological and therapeutic effects. Prospect. Pharm. Sci. 2023, 21, 49–56. [Google Scholar] [CrossRef]
- Mohammed, F.; Uysal, İ.; Yaz, H.; Sevindik, M. Papaver species: Usage areas, essential oil, nutrient and elements contents, biological activities. Prospect. Pharm. Sci. 2023, 21, 1–9. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, R.; Yan, F.; Jin, Y.; Liu, X.; Wang, T. Medicarpin Protects Cerebral Microvascular Endothelial Cells Against Oxygen-Glucose Deprivation/Reoxygenation-Induced Injury via the PI3K/Akt/FoxO Pathway: A Study of Network Pharmacology Analysis and Experimental Validation. Neurochem. Res. 2022, 47, 347–357. [Google Scholar] [CrossRef]
- Lu, C.; Du, R.; Fu, H.; Zhang, J.; Zhao, M.; Wei, Y.; Lin, W. Heterologous biosynthesis of medicarpin using engineered Saccharomyces cerevisiae. Synth. Syst. Biotechnol. 2023, 8, 749–756. [Google Scholar] [CrossRef]
- Trivedi, R.; Maurya, R.; Mishra, D.P. Medicarpin, a legume phytoalexin sensitizes myeloid leukemia cells to TRAIL-induced apoptosis through the induction of DR5 and activation of the ROS-JNK-CHOP pathway. Cell Death Dis. 2014, 5, e1465. [Google Scholar] [CrossRef] [PubMed]
- Banales, J.M.; Marin, J.J.G.; Lamarca, A.; Rodrigues, P.M.; Khan, S.A.; Roberts, L.R.; Cardinale, V.; Carpino, G.; Andersen, J.B.; Braconi, C.; et al. Cholangiocarcinoma 2020: The next horizon in mechanisms and management. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 557–588. [Google Scholar] [CrossRef]
- Zhang, Y.; Ma, Z.; Li, C.; Wang, C.; Jiang, W.; Chang, J.; Han, S.; Lu, Z.; Shao, Z.; Wang, Y.; et al. The genomic landscape of cholangiocarcinoma reveals the disruption of post-transcriptional modifiers. Nat. Commun. 2022, 13, 3061. [Google Scholar] [CrossRef]
- Porreca, V.; Barbagallo, C.; Corbella, E.; Peres, M.; Stella, M.; Mignogna, G.; Maras, B.; Ragusa, M.; Mancone, C. Unveil Intrahepatic Cholangiocarcinoma Heterogeneity through the Lens of Omics and Multi-Omics Approaches. Cancers 2024, 16, 2889. [Google Scholar] [CrossRef]
- Noor, F.; Tahir ul Qamar, M.; Ashfaq, U.A.; Albutti, A.; Alwashmi, A.S.S.; Aljasir, M.A. Network Pharmacology Approach for Medicinal Plants: Review and Assessment. Pharmaceuticals 2022, 15, 572. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, H.; Zhu, Z.; Yang, Y.; Xiong, Y.; Cui, X.; Liu, Y. Network Pharmacology-Driven Sustainability: AI and Multi-Omics Synergy for Drug Discovery in Traditional Chinese Medicine. Pharmaceuticals 2025, 18, 1074. [Google Scholar] [CrossRef]
- Sakle, N.S.; More, S.A.; Mokale, S.N. A network pharmacology-based approach to explore potential targets of Caesalpinia pulcherima: An updated prototype in drug discovery. Sci. Rep. 2020, 10, 17217. [Google Scholar] [CrossRef]
- Challapa-Mamani, M.R.; Tomás-Alvarado, E.; Espinoza-Baigorria, A.; León-Figueroa, D.A.; Sah, R.; Rodriguez-Morales, A.J.; Barboza, J.J. Molecular Docking and Molecular Dynamics Simulations in Related to Leishmania donovani: An Update and Literature Review. Trop. Med. Infect. Dis. 2023, 8, 457. [Google Scholar] [CrossRef] [PubMed]
- Jain, N.K.; Chandrasekaran, B.; Khazaleh, N.; Jain, H.K.; Lal, M.; Joshi, G.; Jha, V. Computational Network Pharmacology, Molecular Docking, and Molecular Dynamics to Decipher Natural Compounds of Alchornea laxiflora for Liver Cancer Chemotherapy. Pharmaceuticals 2025, 18, 508. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.; Yan, H.; Li, H.; Feng, Y.; Sun, W.; Ren, Y.; Ma, L.; Zeng, W.; Huang, F.; Jiang, Z.; et al. Use of network pharmacology and molecular docking to explore the mechanism of action of curcuma in the treatment of osteosarcoma. Sci. Rep. 2023, 13, 9569. [Google Scholar] [CrossRef]
- Safran, M.; Dalah, I.; Alexander, J.; Rosen, N.; Iny Stein, T.; Shmoish, M.; Nativ, N.; Bahir, I.; Doniger, T.; Krug, H.; et al. GeneCards Version 3: The human gene integrator. Database 2010, 2010, baq020. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Matsuura, Y.; Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 2025, 53, D672–D677. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef]
- Ni, B.; Wang, H.; Khalaf, H.K.S.; Blay, V.; Houston, D.R. AutoDock-SS: AutoDock for Multiconformational Ligand-Based Virtual Screening. J. Chem. Inf. Model. 2024, 64, 3779–3789. [Google Scholar] [CrossRef] [PubMed]
- Tedasen, A.; Chiabchalard, A.; Tencomnao, T.; Yamasaki, K.; Majima, H.J.; Phongphithakchai, A.; Chatatikun, M. Anti-Melanogenic Activity of Ethanolic Extract from Garcinia atroviridis Fruits Using In Vitro Experiments, Network Pharmacology, Molecular Docking, and Molecular Dynamics Simulation. Antioxidants 2024, 13, 713. [Google Scholar] [CrossRef]
- Tian, T.; Li, X.; Zhang, J. mTOR Signaling in Cancer and mTOR Inhibitors in Solid Tumor Targeting Therapy. Int. J. Mol. Sci. 2019, 20, 755. [Google Scholar] [CrossRef] [PubMed]
- Rascio, F.; Spadaccino, F.; Rocchetti, M.T.; Castellano, G.; Stallone, G.; Netti, G.S.; Ranieri, E. The Pathogenic Role of PI3K/AKT Pathway in Cancer Onset and Drug Resistance: An Updated Review. Cancers 2021, 13, 3949. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Pires, D.E.V.; Blundell, T.L.; Ascher, D.B. pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures. J. Med. Chem. 2015, 58, 4066–4072. [Google Scholar] [CrossRef]
- Banerjee, P.; Kemmler, E.; Dunkel, M.; Preissner, R. ProTox 3.0: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2024, 52, W513–W520. [Google Scholar] [CrossRef]
- Smolarz, B.; Durczyński, A.; Romanowicz, H.; Szyłło, K.; Hogendorf, P. miRNAs in Cancer (Review of Literature). Int. J. Mol. Sci. 2022, 23, 2805. [Google Scholar] [CrossRef]
- Jusakul, A.; Cutcutache, I.; Yong, C.H.; Lim, J.Q.; Huang, M.N.; Padmanabhan, N.; Nellore, V.; Kongpetch, S.; Ng, A.W.T.; Ng, L.M.; et al. Whole-Genome and Epigenomic Landscapes of Etiologically Distinct Subtypes of Cholangiocarcinoma. Cancer Discov. 2017, 7, 1116–1135. [Google Scholar] [CrossRef]
- Sitthirak, S.; Roytrakul, S.; Wangwiwatsin, A.; Namwat, N.; Klanrit, P.; Dokduang, H.; Sa-ngiamwibool, P.; Titapan, A.; Jareanrat, A.; Thanasukarn, V.; et al. Proteomic profiling reveals common and region-specific protein signatures underlying tumor heterogeneity in cholangiocarcinoma. Sci. Rep. 2025, 15, 17228. [Google Scholar] [CrossRef] [PubMed]
- Sitthirak, S.; Wangwiwatsin, A.; Jusakul, A.; Namwat, N.; Klanrit, P.; Dokduang, H.; Sa-ngiamwibool, P.; Titapun, A.; Jareanrat, A.; Thanasukarn, V.; et al. Whole exome sequencing of multi-regions reveals tumor heterogeneity in Opisthorchis viverrini-associated cholangiocarcinoma. Sci. Rep. 2025, 15, 10886. [Google Scholar] [CrossRef] [PubMed]
- Dokduang, H.; Juntana, S.; Techasen, A.; Namwat, N.; Yongvanit, P.; Khuntikeo, N.; Riggins, G.J.; Loilome, W. Survey of activated kinase proteins reveals potential targets for cholangiocarcinoma treatment. Tumor Biol. 2013, 34, 3519–3528. [Google Scholar] [CrossRef]
- Yothaisong, S.; Dokduang, H.; Techasen, A.; Namwat, N.; Yongvanit, P.; Bhudhisawasdi, V.; Puapairoj, A.; Riggins, G.J.; Loilome, W. Increased activation of PI3K/AKT signaling pathway is associated with cholangiocarcinoma metastasis and PI3K/mTOR inhibition presents a possible therapeutic strategy. Tumor Biol. 2013, 34, 3637–3648. [Google Scholar] [CrossRef]


| Properties | |
| Physicochemical Properties | |
| Formula | C16H14O4 |
| Molecular weight | 270.28 g/mol |
| Num. heavy atoms | 20 |
| Num. arom. heavy atoms | 12 |
| Fraction Csp3 | 0.25 |
| Num. rotatable bonds | 1 |
| Num. H-bond acceptors | 4 |
| Num. H-bond donors | 1 |
| Molar Refractivity | 73.17 |
| TPSA | 47.92 Å2 |
| Lipophilicity | |
| Log Po/w (iLOGP) | 2.57 |
| Log Po/w (XLOGP3) | 2.77 |
| Log Po/w (WLOGP) | 2.69 |
| Log Po/w (MLOGP) | 1.87 |
| Log Po/w (SILICOS-IT) | 2.75 |
| Consensus Log Po/w | 2.53 |
| Water Solubility | |
| Log S (ESOL) | −3.64 |
| Solubility | 6.21 × 10−2 mg/mL; 2.30 × 10−4 mol/L |
| Class | Soluble |
| Log S (Ali) | −3.43 |
| Solubility | 1.00 × 10−1 mg/mL; 3.70 × 10−4 mol/L |
| Class | Soluble |
| Log S (SILICOS-IT) | −4.31 |
| Solubility | 1.32 × 10−2 mg/mL; 4.90 × 10−5 mol/L |
| Class | Moderately soluble |
| Pharmacokinetics | |
| GI absorption | High |
| BBB permeant | Yes |
| P-gp substrate | Yes |
| CYP1A2 inhibitor | Yes |
| CYP2C19 inhibitor | Yes |
| CYP2C9 inhibitor | No |
| CYP2D6 inhibitor | Yes |
| CYP3A4 inhibitor | Yes |
| Log Kp (skin permeation) | −5.98 cm/s |
| Drug-likeness | |
| Lipinski | Yes; 0 violation |
| Ghose | Yes |
| Veber | Yes |
| Egan | Yes |
| Muegge | Yes |
| Bioavailability Score | 0.55 |
| Medicinal Chemistry | |
| PAINS | 0 alert |
| Brenk | 0 alert |
| Lead-likeness | Yes |
| Synthetic accessibility | 3.54 |
| Property | Model Name | Predicted Value | Unit |
|---|---|---|---|
| Absorption | Water solubility | −3.459 | Numeric (log mol/L) |
| Caco2 permeability | 1.246 | Numeric (log Papp in 10−6 cm/s) | |
| Intestinal absorption (human) | 95.188 | Numeric (% Absorbed) | |
| Skin Permeability | −2.819 | Numeric (log Kp) | |
| P-glycoprotein substrate | No | Categorical (Yes/No) | |
| P-glycoprotein I inhibitor | No | Categorical (Yes/No) | |
| P-glycoprotein II inhibitor | No | Categorical (Yes/No) | |
| Distribution | VDss (human) | 0.065 | Numeric (log L/kg) |
| Fraction unbound (human) | 0.04 | Numeric (Fu) | |
| BBB permeability | 0.324 | Numeric (log BB) | |
| CNS permeability | −1.838 | Numeric (log PS) | |
| Metabolism | CYP2D6 substrate | No | Categorical (Yes/No) |
| CYP3A4 substrate | Yes | Categorical (Yes/No) | |
| CYP1A2 inhibitor | Yes | Categorical (Yes/No) | |
| CYP2C19 inhibitor | Yes | Categorical (Yes/No) | |
| CYP2C9 inhibitor | Yes | Categorical (Yes/No) | |
| CYP2D6 inhibitor | No | Categorical (Yes/No) | |
| CYP3A4 inhibitor | Yes | Categorical (Yes/No) | |
| Excretion | Total Clearance | 0.273 | Numeric (log mL/min/kg) |
| Renal OCT2 substrate | No | Categorical (Yes/No) | |
| Toxicity | AMES toxicity | Yes | Categorical (Yes/No) |
| Max. tolerated dose (human) | −0.102 | Numeric (log mg/kg/day) | |
| hERG I inhibitor | No | Categorical (Yes/No) | |
| hERG II inhibitor | No | Categorical (Yes/No) | |
| Oral Rat Acute Toxicity (LD50) | 2.512 | Numeric (mol/kg) | |
| Oral Rat Chronic Toxicity (LOAEL) | 1.875 | Numeric (log mg/kg_bw/day) | |
| Hepatotoxicity | No | Categorical (Yes/No) | |
| Skin Sensitisation | No | Categorical (Yes/No) | |
| T. Pyriformis toxicity | 0.688 | Numeric (log ug/L) | |
| Minnow toxicity | 0.657 | Numeric (log mM) |
| Classification | Target | Prediction | Probability |
|---|---|---|---|
| Organ toxicity | Hepatotoxicity | Active | 0.69 |
| Organ toxicity | Neurotoxicity | Active | 0.87 |
| Organ toxicity | Nephrotoxicity | Inactive | 0.9 |
| Organ toxicity | Respiratory toxicity | Active | 0.98 |
| Organ toxicity | Cardiotoxicity | Inactive | 0.77 |
| Toxicity end points | Carcinogenicity | Inactive | 0.62 |
| Toxicity end points | Immunotoxicity | Active | 0.96 |
| Toxicity end points | Mutagenicity | Inactive | 0.97 |
| Toxicity end points | Cytotoxicity | Inactive | 0.93 |
| Toxicity end points | BBB-barrier | Inactive | 1 |
| Toxicity end points | Ecotoxicity | Active | 0.73 |
| Toxicity end points | Clinical toxicity | Inactive | 0.56 |
| Toxicity end points | Nutritional toxicity | Inactive | 0.74 |
| Tox21-Nuclear receptor signalling pathways | Aryl hydrocarbon Receptor (AhR) | Inactive | 0.97 |
| Tox21-Nuclear receptor signalling pathways | Androgen Receptor (AR) | Inactive | 0.99 |
| Tox21-Nuclear receptor signalling pathways | Androgen Receptor Ligand Binding Domain (AR-LBD) | Inactive | 0.99 |
| Tox21-Nuclear receptor signalling pathways | Aromatase | Active | 1 |
| Tox21-Nuclear receptor signalling pathways | Estrogen Receptor Alpha (ER) | Active | 0.99 |
| Tox21-Nuclear receptor signalling pathways | Estrogen Receptor Ligand Binding Domain (ER-LBD) | Active | 1 |
| Tox21-Nuclear receptor signalling pathways | Peroxisome Proliferator Activated Receptor Gamma (PPAR-Gamma) | Inactive | 0.99 |
| Tox21-Stress response pathways | Nuclear factor (erythroid-derived 2)-like 2/antioxidant responsive element (nrf2/ARE) | Inactive | 0.88 |
| Tox21-Stress response pathways | Heat shock factor response element (HSE) | Inactive | 0.88 |
| Tox21-Stress response pathways | Mitochondrial Membrane Potential (MMP) | Inactive | 0.7 |
| Tox21-Stress response pathways | Phosphoprotein (Tumor Supressor) p53 | Inactive | 0.96 |
| Tox21-Stress response pathways | ATPase family AAA domain-containing protein 5 (ATAD5) | Inactive | 0.99 |
| Molecular Initiating Events | Thyroid hormone receptor alpha (THRα) | Inactive | 0.9 |
| Molecular Initiating Events | Thyroid hormone receptor beta (THRβ) | Inactive | 0.78 |
| Molecular Initiating Events | Transtyretrin (TTR) | Inactive | 0.97 |
| Molecular Initiating Events | Ryanodine receptor (RYR) | Inactive | 0.98 |
| Molecular Initiating Events | GABA receptor (GABAR) | Inactive | 0.96 |
| Molecular Initiating Events | Glutamate N-methyl-D-aspartate receptor (NMDAR) | Inactive | 0.92 |
| Molecular Initiating Events | alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionate receptor (AMPAR) | Inactive | 0.97 |
| Molecular Initiating Events | Kainate receptor (KAR) | Inactive | 0.99 |
| Molecular Initiating Events | Achetylcholinesterase (AChE) | Active | 0.69 |
| Molecular Initiating Events | Constitutive androstane receptor (CAR) | Inactive | 0.98 |
| Molecular Initiating Events | Pregnane X receptor (PXR) | Inactive | 0.92 |
| Molecular Initiating Events | NADH-quinone oxidoreductase (NADHOX) | Inactive | 0.97 |
| Molecular Initiating Events | Voltage gated sodium channel (VGSC) | Inactive | 0.95 |
| Molecular Initiating Events | Na+/I- symporter (NIS) | Inactive | 0.98 |
| Metabolism | Cytochrome CYP1A2 | Inactive | 0.76 |
| Metabolism | Cytochrome CYP2C19 | Inactive | 0.87 |
| Metabolism | Cytochrome CYP2C9 | Active | 0.56 |
| Metabolism | Cytochrome CYP2D6 | Inactive | 0.63 |
| Metabolism | Cytochrome CYP3A4 | Active | 0.71 |
| Metabolism | Cytochrome CYP2E1 | Inactive | 0.98 |
| No. | Protein Name | PDB | Compound and Positive Control | Binding Energies (kcal/mol) | Inhibition Constant (nM) |
|---|---|---|---|---|---|
| 1 | CASP3 | 1RE1 | Medicarpin | −6.98 | 7.71 uM |
| NA3501 | −7.85 | 1.77 uM | |||
| 2 | ESR1 | 6VPF | Medicarpin | −7.65 | 2.47 uM |
| 53Q | −3.64 | 2.15 mM | |||
| 3 | SRC | 6WIW | Medicarpin | −7.08 | 6.51 uM |
| I14 | −6 | 40.11 uM | |||
| 4 | CCND1 | 9CSK | Medicarpin | −8.06 | 1.24 uM |
| Sancycline | −8.1 | 1.16 uM | |||
| 5 | MTOR | 5OQ4 | Medicarpin | −9.6 | 1.57 uM |
| A3W | −7.53 | 3.0 uM | |||
| 6 | PIK3CA | 7R9V | Medicarpin | −6.39 | 20.65 uM |
| 2Q7 | −11.31 | 5.11 nM | |||
| 7 | PARP1 | 7KK4 | Medicarpin | −7.32 | 4.33 uM |
| 09L | −11.5 | 3.73 nM | |||
| 8 | GSK3B | 4PTE | Medicarpin | −6.8 | 10.45 uM |
| 2WF | −7.2 | 5.26 uM | |||
| 9 | KDR | 3VHE | Medicarpin | −7.07 | 6.63 uM |
| 42Q | −11.34 | 4.91 nM | |||
| 10 | KIT | 4U0I | Medicarpin | −7.35 | 4.09 uM |
| 0LI | −13.29 | 180.87 pM |
| Medicarpin–mTOR complex | Hydrogen Bonds | ||||||
| Docking | MD Simulation Timeline | ||||||
| 50 ns | 100 ns | 150 ns | 200 ns | 250 ns | 300 ns | ||
| VAL882, THR887 and LYS890 | ALA805, ILE881, and THR887 | ALA805, SER806, VAL882 and THR887 | ALA805, SER806, VAL882 and THR887 | ALA805, SER806, VAL882, THR887, ASP950 and GLY970 | ALA805, SER806, VAL882, THR887 and ASP950 | ALA805, SER806, VAL882 THR887 and ASP950 | |
| Medicarpin–mTOR complex | Hydrophobic interactions | ||||||
| Docking | MD simulation timeline | ||||||
| 50 ns | 100 ns | 150 ns | 200 ns | 250 ns | 300 ns | ||
| ILE831, TYR867, ILE881, MET953, PHE961 and ILE963 | MET804, TRP812, ILE831, TYR867, ILE879, ILE881, MET953 and ILE963 | MET804, TRP802, ILE831, TYR867, ILE879, ILE881, MET953 and ILE963 | MET804, TRP812, ILE831, TYR867, ILE881, MET953 and ILE963 | MET804, TRP812, ILE831, TYR867, ILE879, ILE881, MET953, ILE963 and ILE968 | MET804, TRP812, ILE831, TYR867, ILE879, ILE881, MET953, ILE963 and ILE968 | MET804, TRP812, ILE831, TYR867, ILE879, ILE881, MET953, PHE961, ILE963 and ILE968 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sitthirak, S.; Tedasen, A.; Rattanapan, Y.; Duangchan, T.; Dokduang, H.; Pattaranggoon, N.C.; Saisuwan, K.; Loilome, W.; Namwat, N. Computational Investigation Identifies mTOR as a Primary Binding Target of Medicarpin in Cholangiocarcinoma: Insights from Network Pharmacology and Molecular Docking. Life 2025, 15, 1828. https://doi.org/10.3390/life15121828
Sitthirak S, Tedasen A, Rattanapan Y, Duangchan T, Dokduang H, Pattaranggoon NC, Saisuwan K, Loilome W, Namwat N. Computational Investigation Identifies mTOR as a Primary Binding Target of Medicarpin in Cholangiocarcinoma: Insights from Network Pharmacology and Molecular Docking. Life. 2025; 15(12):1828. https://doi.org/10.3390/life15121828
Chicago/Turabian StyleSitthirak, Sirinya, Aman Tedasen, Yanisa Rattanapan, Thitinat Duangchan, Hasaya Dokduang, Nawanwat C. Pattaranggoon, Krittamate Saisuwan, Watcharin Loilome, and Nisana Namwat. 2025. "Computational Investigation Identifies mTOR as a Primary Binding Target of Medicarpin in Cholangiocarcinoma: Insights from Network Pharmacology and Molecular Docking" Life 15, no. 12: 1828. https://doi.org/10.3390/life15121828
APA StyleSitthirak, S., Tedasen, A., Rattanapan, Y., Duangchan, T., Dokduang, H., Pattaranggoon, N. C., Saisuwan, K., Loilome, W., & Namwat, N. (2025). Computational Investigation Identifies mTOR as a Primary Binding Target of Medicarpin in Cholangiocarcinoma: Insights from Network Pharmacology and Molecular Docking. Life, 15(12), 1828. https://doi.org/10.3390/life15121828







