Identification of Colon Immune Cell Marker Genes Using Machine Learning Methods
Abstract
:1. Introduction
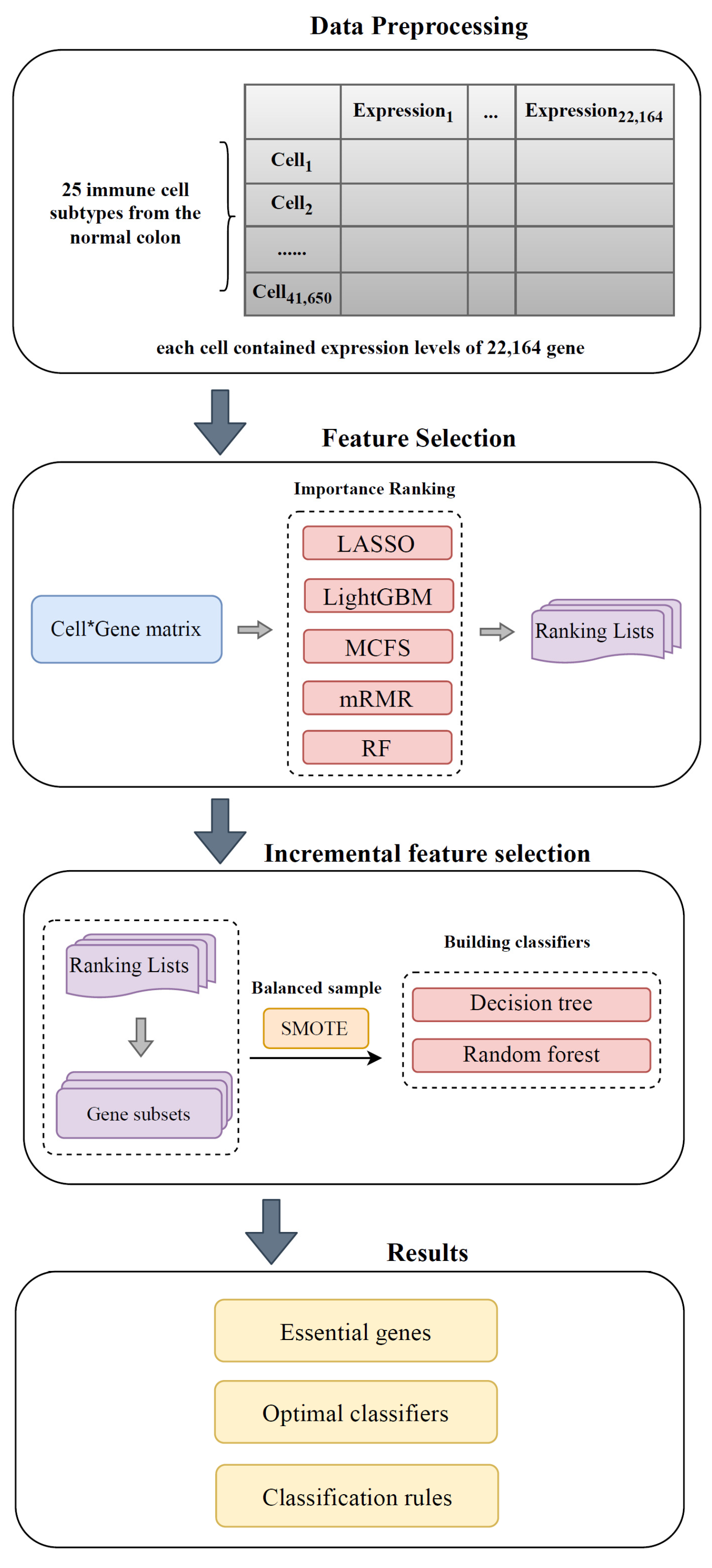
2. Materials and Methods
2.1. Data
2.2. Feature Ranking Algorithms
2.2.1. Last Absolute Shrinkage and Selection Operator
2.2.2. Light Gradient Boosting Machine
2.2.3. Monte Carlo Feature Selection
2.2.4. Minimum Redundancy Maximum Relevance
2.2.5. Random Forest
2.3. Incremental Feature Selection
2.4. Synthetic Minority Oversampling Technique
2.5. Classification Algorithm
2.5.1. Decision Tree
2.5.2. Random Forest
2.6. Performance Evaluation
3. Results
3.1. Dynamics of Classifier Performance
3.2. Relationships between the Most Essential Genes Extracted from Five Lists
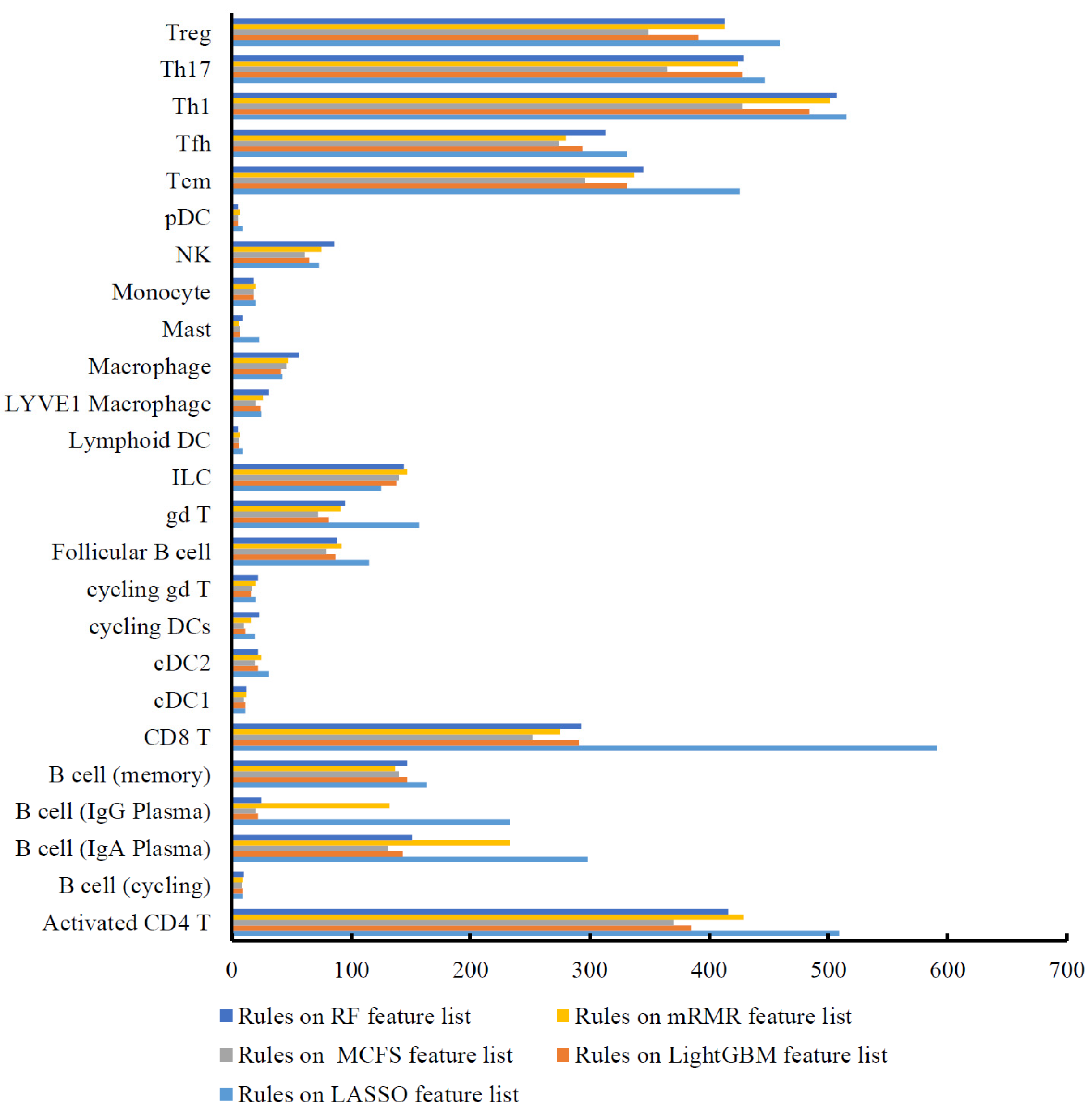
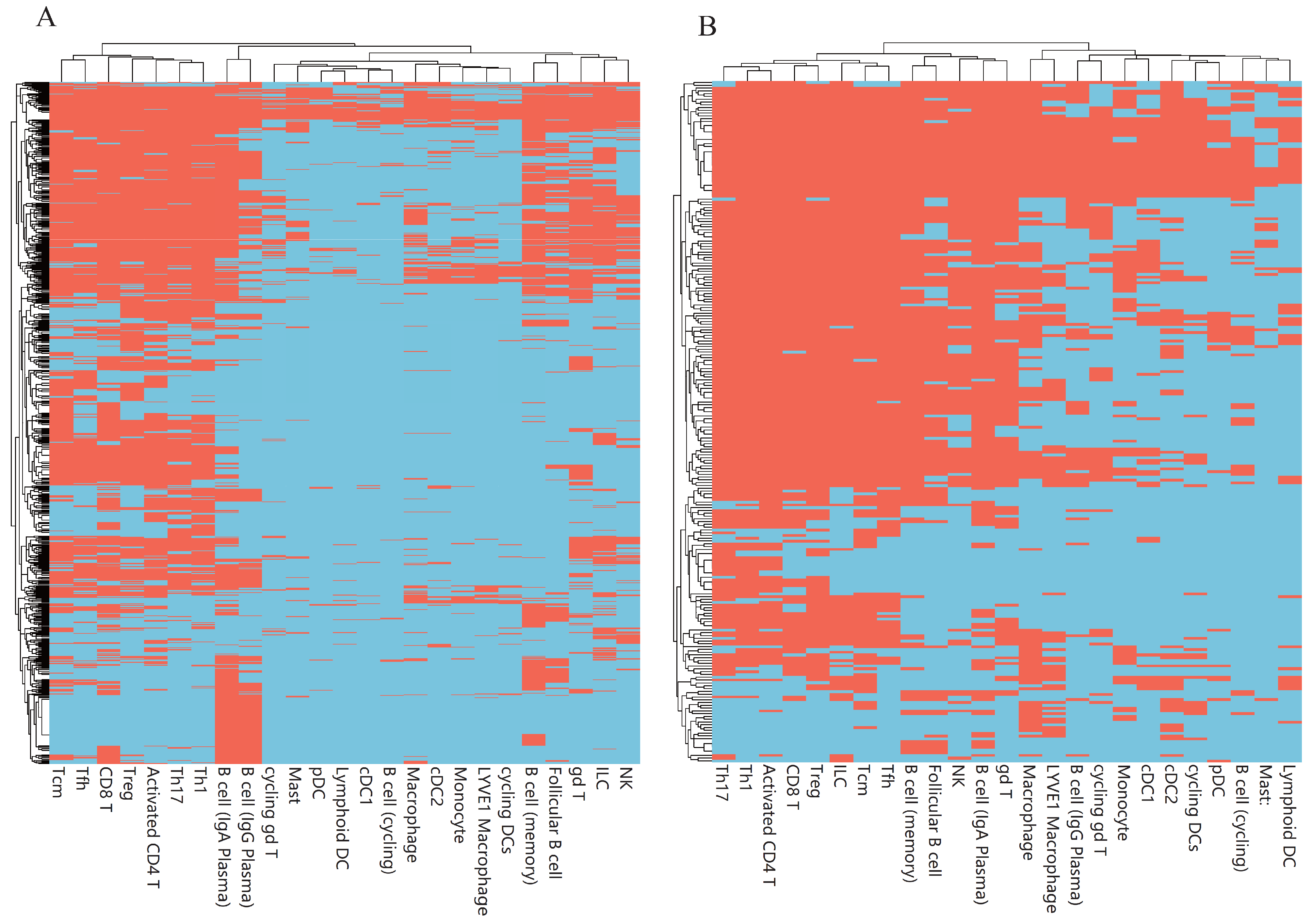
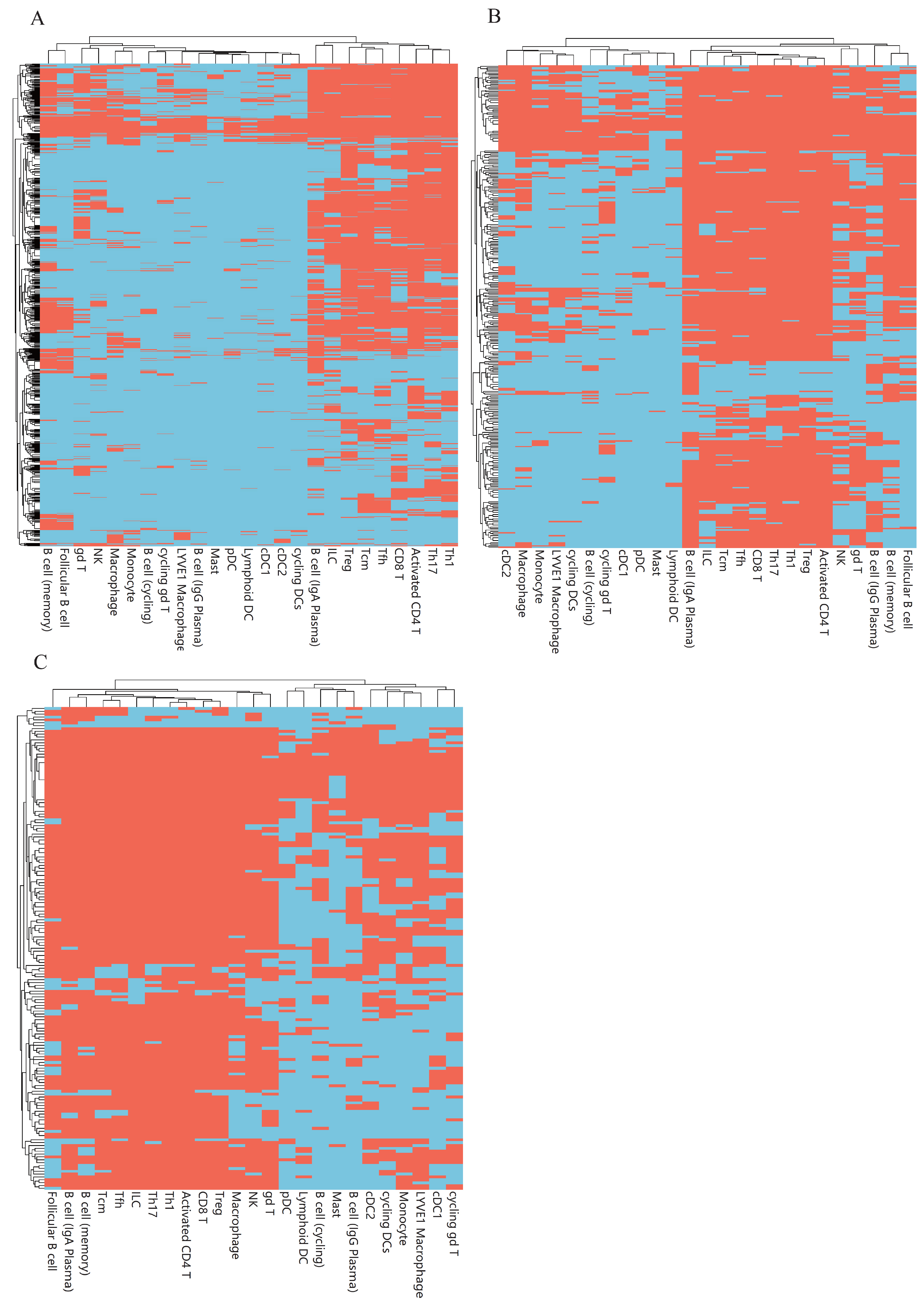
3.3. Classification Rules
4. Discussion
4.1. T Cell Family
4.2. B Cell Family
4.3. Other Cells
4.4. Limitation of This Study
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Azzouz, L.L.; Sharma, S. Physiology, Large Intestine; StatPearls Publishing: St. Petersburg, FL, USA, 2018. [Google Scholar]
- Kahai, P.; Mandiga, P.; Wehrle, C.J.; Lobo, S. Anatomy, abdomen and pelvis, large intestine. In Statpearls; StatPearls Publishing: St. Petersburg, FL, USA, 2021. [Google Scholar]
- Nigam, Y.; Knight, J.; Williams, N. Gastrointestinal tract 5: The anatomy and functions of the large intestine. Nurs. Times 2019, 115, 50–53. [Google Scholar]
- Louis, P.; Scott, K.P.; Duncan, S.H.; Flint, H.J. Understanding the effects of diet on bacterial metabolism in the large intestine. J. Appl. Microbiol. 2007, 102, 1197–1208. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.; Blaser, M.J. The human microbiome: At the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Börnigen, D.; Morgan, X.C.; Franzosa, E.A.; Ren, B.; Xavier, R.J.; Garrett, W.S.; Huttenhower, C. Functional profiling of the gut microbiome in disease-associated inflammation. Genome Med. 2013, 5, 65. [Google Scholar] [CrossRef]
- Germain, R.N. T-cell development and the CD4–CD8 lineage decision. Nat. Rev. Immunol. 2002, 2, 309–322. [Google Scholar] [CrossRef]
- Amadori, A.; Zamarchi, R.; De Silvestro, G.; Forza, G.; Cavatton, G.; Danieli, G.A.; Clementi, M.; Chieco-Bianchi, L. Genetic control of the CD4/CD8 T-cell ratio in humans. Nat. Med. 1995, 1, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- James, K.R.; Gomes, T.; Elmentaite, R.; Kumar, N.; Gulliver, E.L.; King, H.W.; Stares, M.D.; Bareham, B.R.; Ferdinand, J.R.; Petrova, V.N.; et al. Distinct microbial and immune niches of the human colon. Nat. Immunol. 2020, 21, 343–353. [Google Scholar] [CrossRef]
- Elmentaite, R.; Kumasaka, N.; Roberts, K.; Fleming, A.; Dann, E.; King, H.W.; Kleshchevnikov, V.; Dabrowska, M.; Pritchard, S.; Bolt, L.; et al. Cells of the human intestinal tract mapped across space and time. Nature 2021, 597, 250–255. [Google Scholar] [CrossRef] [PubMed]
- Ranstam, J.; Cook, J. Lasso regression. J. Br. Surg. 2018, 105, 1348. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.-Y. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Draminski, M.; Rada-Iglesias, A.; Enroth, S.; Wadelius, C.; Koronacki, J.; Komorowski, J. Monte carlo feature selection for supervised classification. Bioinformatics 2008, 24, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Long, F.; Ding, C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans. Pattern Anal. Mach. Intell. 2005, 27, 1226–1238. [Google Scholar] [CrossRef] [PubMed]
- Fisher, A.; Rudin, C.; Dominici, F. All models are wrong, but many are useful: Learning a variable’s importance by studying an entire class of prediction models simultaneously. J. Mach. Learn. Res. 2019, 20, 1–81. [Google Scholar]
- Liu, H.; Setiono, R. Incremental feature selection. Appl. Intell. 1998, 9, 217–230. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. Smote: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Safavian, S.R.; Landgrebe, D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man Cybern. 1991, 21, 660–674. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Shah, H.A.; Liu, J.; Yang, Z.; Feng, J. Review of machine learning methods for the prediction and reconstruction of metabolic pathways. Front. Mol. Biosci. 2021, 8, 634141. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Morishima, K. Blastkoala and ghostkoala: Kegg tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 2016, 428, 726–731. [Google Scholar] [CrossRef]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. Kaas: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M. The rast server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zhang, S.; Chen, L.; Pan, X.; Li, Z.; Huang, T.; Cai, Y.D. Identifying functions of proteins in mice with functional embedding features. Front. Genet. 2022, 13, 909040. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Huang, F.; Liao, H.; Li, Z.; Feng, K.; Huang, T.; Cai, Y.D. Identification of covid-19-specific immune markers using a machine learning method. Front. Mol. Biosci. 2022, 9, 952626. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Guo, W.; Ding, S.; Chen, L.; Feng, K.; Huang, T.; Cai, Y.D. Identifying key microrna signatures for neurodegenerative diseases with machine learning methods. Front. Genet. 2022, 13, 880997. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Meng, M.; Zhou, X.; Ding, S.; Feng, K.; Zeng, Z.; Huang, T.; Cai, Y.D. Identification of Covid-19 severity biomarkers based on feature selection on single-cell rna-seq data of CD8+ T cells. Front. Genet. 2022, 13, 1053772. [Google Scholar] [CrossRef]
- Huang, F.; Fu, M.; Li, J.; Chen, L.; Feng, K.; Huang, T.; Cai, Y.-D. Analysis and prediction of protein stability based on interaction network, gene ontology, and kegg pathway enrichment scores. BBA Proteins Proteom. 2023, 1871, 140889. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Ma, Q.; Ren, J.; Li, J.; Wang, F.; Huang, T.; Cai, Y.-D. Identification of smoking associated transcriptome aberration in blood with machine learning methods. BioMed Res. Int. 2023, 2023, 5333361. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Zhang, Y.; Guo, W.; Feng, K.; Yuan, Y.; Huang, T.; Cai, Y.-D. Identification of genes associated with the impairment of olfactory and gustatory functions in covid-19 via machine-learning methods. Life 2023, 13, 798. [Google Scholar] [CrossRef]
- Zhao, X.; Chen, L.; Lu, J. A similarity-based method for prediction of drug side effects with heterogeneous information. Math. Biosci. 2018, 306, 136–144. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Kohavi, R. A study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection. In Proceedings of the 14th International joint Conference on artificial intelligence, Montreal, QC, Canada, 20–25 August 1995; Lawrence Erlbaum Associates Ltd.: Mahwah, NJ, USA, 1995; pp. 1137–1145. [Google Scholar]
- Powers, D. Evaluation: From precision, recall and f-measure to roc., informedness, markedness & correlation. J. Mach. Learn. Technol. 2011, 2, 37–63. [Google Scholar]
- Tang, S.; Chen, L. iATC-NFMLP: Identifying classes of anatomical therapeutic chemicals based on drug networks, fingerprints and multilayer perceptron. Curr. Bioinform. 2022, 17, 814–824. [Google Scholar] [CrossRef]
- Wu, C.; Chen, L. A model with deep analysis on a large drug network for drug classification. Math. Biosci. Eng. 2023, 20, 383–401. [Google Scholar] [CrossRef]
- Wu, Z.; Chen, L. Similarity-based method with multiple-feature sampling for predicting drug side effects. Comput. Math. Methods Med. 2022, 2022, 9547317. [Google Scholar] [CrossRef] [PubMed]
- Liang, H.; Chen, L.; Zhao, X.; Zhang, X. Prediction of drug side effects with a refined negative sample selection strategy. Comput. Math. Methods Med. 2020, 2020, 1573543. [Google Scholar] [CrossRef]
- Zhao, X.; Chen, L.; Guo, Z.-H.; Liu, T. Predicting drug side effects with compact integration of heterogeneous networks. Curr. Bioinform. 2019, 14, 709–720. [Google Scholar] [CrossRef]
- Chen, L.; Chen, K.; Zhou, B. Inferring drug-disease associations by a deep analysis on drug and disease networks. Math. Biosci. Eng. 2023, 20, 14136–14157. [Google Scholar] [CrossRef]
- Gorodkin, J. Comparing two k-category assignments by a k-category correlation coefficient. Comput. Biol. Chem. 2004, 28, 367–374. [Google Scholar] [CrossRef]
- Pelletier, J.; Thomas, G.; Volarević, S. Ribosome biogenesis in cancer: New players and therapeutic avenues. Nat. Rev. Cancer 2018, 18, 51–63. [Google Scholar] [CrossRef]
- Pecoraro, A.; Pagano, M.; Russo, G.; Russo, A. Ribosome biogenesis and cancer: Overview on ribosomal proteins. Int. J. Mol. Sci. 2021, 22, 5496. [Google Scholar] [CrossRef]
- Lee, M.Y.; Jeon, J.W.; Sievers, C.; Allen, C.T. Antigen processing and presentation in cancer immunotherapy. J. Immunother. Cancer 2020, 8, e001111. [Google Scholar] [CrossRef]
- Zhang, B.; Li, J.; Hua, Q.; Wang, H.; Xu, G.; Chen, J.; Zhu, Y.; Li, R.; Liang, Q.; Wang, L.; et al. Tumor cemip drives immune evasion of colorectal cancer via mhc-i internalization and degradation. J. Immunother. Cancer 2023, 11, e005592. [Google Scholar] [CrossRef]
- Kasajima, A.; Sers, C.; Sasano, H.; Jöhrens, K.; Stenzinger, A.; Noske, A.; Buckendahl, A.C.; Darb-Esfahani, S.; Müller, B.M.; Budczies, J.; et al. Down-regulation of the antigen processing machinery is linked to a loss of inflammatory response in colorectal cancer. Hum. Pathol. 2010, 41, 1758–1769. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.K.; Gupta, G. Klrb receptor family and human early activation antigen (CD69). In Animal Lectins: Form, Function and Clinical Applications; Springer: Berlin/Heidelberg, Germany, 2012; pp. 619–638. [Google Scholar]
- Boix, F.; Millan, O.; San Segundo, D.; Mancebo, E.; Rimola, A.; Fabrega, E.; Fortuna, V.; Mrowiec, A.; Castro-Panete, M.J.; de la Pena, J. High expression of CD38, CD69, CD95 AND CD154 biomarkers in cultured peripheral T lymphocytes correlates with an increased risk of acute rejection in liver allograft recipients. Immunobiology 2016, 221, 595–603. [Google Scholar] [CrossRef]
- Testi, R.; Phillips, J.; Lanier, L. T cell activation via leu-23 (CD69). J. Immunol. 1989, 143, 1123–1128. [Google Scholar] [CrossRef]
- Ziegler, S.F.; Ramsdell, F.; Alderson, M.R. The activation antigen CD69. Stem Cells 1994, 12, 456–465. [Google Scholar] [CrossRef]
- Song, F.; Zhang, Y.; Chen, Q.; Bi, D.; Yang, M.; Lu, L.; Li, M.; Zhu, H.; Liu, Y.; Wei, Q. Mast cells inhibit colorectal cancer development by inducing er stress through secreting cystatin c. Oncogene 2022, 42, 209–223. [Google Scholar] [CrossRef] [PubMed]
- Utans, U.; Arceci, R.J.; Yamashita, Y.; Russell, M.E. Cloning and characterization of allograft inflammatory factor-1: A novel macrophage factor identified in rat cardiac allografts with chronic rejection. J. Clin. Investig. 1995, 95, 2954–2962. [Google Scholar] [CrossRef]
- Vu, D.; Tellez-Corrales, E.; Shah, T.; Hutchinson, I.; Min, D.I. Influence of cyclooxygenase-2 (cox-2) gene promoter-1195 and allograft inflammatory factor-1 (aif-1) polymorphisms on allograft outcome in hispanic kidney transplant recipients. Hum. Immunol. 2013, 74, 1386–1391. [Google Scholar] [CrossRef] [PubMed]
- Sinigaglia, F.; Guttinger, M.; Kilgus, J.; Doran, D.; Matile, H.; Etlinger, H.; Trzeciak, A.; Gillessen, D.; Pink, J. A malaria T-cell epitope recognized in association with most mouse and human mhc class ii molecules. Nature 1988, 336, 778–780. [Google Scholar] [CrossRef]
- Kim, Y.-I.; Shin, H.-W.; Chun, Y.-S.; Park, J.-W. Cst3 and gdf15 ameliorate renal fibrosis by inhibiting fibroblast growth and activation. Biochem. Biophys. Res. Commun. 2018, 500, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Burnside, E.; Bradbury, E. Manipulating the extracellular matrix and its role in brain and spinal cord plasticity and repair. Neuropathol. Appl. Neurobiol. 2014, 40, 26–59. [Google Scholar] [CrossRef]
- Zhao, Y.-Y.; Yan, D.-J.; Chen, Z.-W. Role of aif-1 in the regulation of inflammatory activation and diverse disease processes. Cell Immunol. 2013, 284, 75–83. [Google Scholar] [CrossRef]
- Tau, G.; Rothman, P. Biologic functions of the ifn-γ receptors. Allergy 1999, 54, 1233. [Google Scholar] [CrossRef]
- Sun, H.; Hartigan, C.R.; Chen, C.w.; Sun, Y.; Tariq, M.; Robertson, J.M.; Krummey, S.M.; Mehta, A.K.; Ford, M.L. Tigit regulates apoptosis of risky memory T cell subsets implicated in belatacept-resistant rejection. Am. J. Transplant. 2021, 21, 3256–3267. [Google Scholar] [CrossRef] [PubMed]
- Fuhrman, C.A.; Yeh, W.-I.; Seay, H.R.; Lakshmi, P.S.; Chopra, G.; Zhang, L.; Perry, D.J.; McClymont, S.A.; Yadav, M.; Lopez, M.-C. Divergent phenotypes of human regulatory T cells expressing the receptors tigit and CD226. J. Immunol. 2015, 195, 145–155. [Google Scholar] [CrossRef]
- Milcent, B.; Josseaume, N.; Petitprez, F.; Riller, Q.; Amorim, S.; Loiseau, P.; Toubert, A.; Brice, P.; Thieblemont, C.; Teillaud, J.-L. Recovery of central memory and naive peripheral T cells in follicular lymphoma patients receiving rituximab-chemotherapy based regimen. Sci. Rep. 2019, 9, 13471. [Google Scholar] [CrossRef]
- Masuda, K.; Kornberg, A.; Miller, J.; Lin, S.; Suek, N.; Botella, T.; Secener, K.; Bacarella, A.M.; Cheng, L.; Ingham, M. Multiplexed single-cell analysis reveals prognostic and non-prognostic T cell types in human colorectal cancer. JCI Insight 2022, 7, e154646. [Google Scholar] [CrossRef]
- Morille, J.; Mandon, M.; Rodriguez, S.; Roulois, D.; Leonard, S.; Garcia, A.; Wiertlewski, S.; Le Page, E.; Berthelot, L.; Nicot, A. Multiple sclerosis csf is enriched with follicular T cells displaying a th1/eomes signature. Neurol. Neuroimmunol. Neuroinflammation 2022, 9, e200033. [Google Scholar] [CrossRef]
- Béguelin, W.; Teater, M.; Meydan, C.; Hoehn, K.B.; Phillip, J.M.; Soshnev, A.A.; Venturutti, L.; Rivas, M.A.; Calvo-Fernández, M.T.; Gutierrez, J. Mutant ezh2 induces a pre-malignant lymphoma niche by reprogramming the immune response. Cancer Cell 2020, 37, 655–673.e611. [Google Scholar] [CrossRef] [PubMed]
- Arjunaraja, S.; Nosé, B.D.; Sukumar, G.; Lott, N.M.; Dalgard, C.L.; Snow, A.L. Intrinsic plasma cell differentiation defects in b cell expansion with nf-κb and T cell anergy patient b cells. Front. Immunol. 2017, 8, 913. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Jones, M.; Jiang, J.; Zaprazna, K.; Yu, D.; Pear, W.; Maillard, I.; Atchison, M.L. Increased expression of PcG protein YY1 negatively regulates b cell development while allowing accumulation of myeloid cells and LT-HSC cells. PLoS ONE 2012, 7, e30656. [Google Scholar] [CrossRef] [PubMed]
- Snir, O.; Kanduri, C.; Lundin, K.E.; Sandve, G.K.; Sollid, L.M. Transcriptional profiling of human intestinal plasma cells reveals effector functions beyond antibody production. United Eur. Gastroenterol. J. 2019, 7, 1399–1407. [Google Scholar] [CrossRef]
- Johansen, F.; Braathen, R.; Brandtzaeg, P. Role of j chain in secretory immunoglobulin formation. Scand. J. Immunol. 2000, 52, 240–248. [Google Scholar] [CrossRef]
- Brandtzaeg, P.; Korsrud, F. Significance of different J chain profiles in human tissues: Generation of IgA and IgM with binding site for secretory component is related to the J chain expressing capacity of the total local immunocyte population, including IgG and IgD producing cells, and depends on the clinical state of the tissue. Clin. Exp. Immunol. 1984, 58, 709. [Google Scholar]
- Bjerke, K.; Brandtzaeg, P. Terminally differentiated human intestinal B cells. J chain expression of IgA and IgG subclass-producing immunocytes in the distal ileum compared with mesenteric and peripheral lymph nodes. Clin. Exp. Immunol. 1990, 82, 411–415. [Google Scholar] [CrossRef]
- Stockis, J.; Fink, W.; François, V.; Connerotte, T.; De Smet, C.; Knoops, L.; van der Bruggen, P.; Boon, T.; Coulie, P.G.; Lucas, S. Comparison of stable human treg and th clones by transcriptional profiling. Eur. J. Immunol. 2009, 39, 869–882. [Google Scholar] [CrossRef] [PubMed]
- Höllbacher, B.; Duhen, T.; Motley, S.; Klicznik, M.M.; Gratz, I.K.; Campbell, D.J. Transcriptomic profiling of human effector and regulatory T cell subsets identifies predictive population signatures. Immunohorizons 2020, 4, 585–596. [Google Scholar] [CrossRef]
- Lin, C.; Yang, H.; Zhao, W.; Wang, W. Ctsb+ macrophage repress memory immune hub in the liver metastasis site of colorectal cancer patient revealed by multi-omics analysis. Biochem. Biophys. Res. Commun. 2022, 626, 8–14. [Google Scholar] [CrossRef]
- Duan, R.; Liu, Y.; Tang, D.; Xiao, S.; Lin, R.; Zhao, M. Single-cell rna-seq reveals collagen vi antibody-induced expressing lyve1 m2-like macrophages reduce atherosclerotic plaque area on apoe-/-mice. Int. Immunopharmacol. 2023, 116, 109794. [Google Scholar] [CrossRef]
- Wan, J.; Lv, J.; Wang, C.; Zhang, L. Rps27 selectively regulates the expression and alternative splicing of inflammatory and immune response genes in thyroid cancer cells. Adv. Clin. Exp. Med. 2022, 31, 889–901. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Liu, X.; Han, C.; Xu, S.; Xie, B.; Zhang, Q.; Gu, Y.; Hou, J.; Qian, L.; Qian, C. Identification of ifn-γ-producing innate b cells. Cell Res. 2014, 24, 161–176. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.Q.; Correia, D.V.; Grosso, A.R.; Lança, T.; Ferreira, C.; Lacerda, J.F.; Barata, J.T.; da Silva, M.G.; Silva-Santos, B. Identification of a panel of ten cell surface protein antigens associated with immunotargeting of leukemias and lymphomas by peripheral blood γδ T cells. Haematologica 2010, 95, 1397. [Google Scholar] [CrossRef] [PubMed]
- Gottenberg, J.-E.; Cagnard, N.; Lucchesi, C.; Letourneur, F.; Mistou, S.; Lazure, T.; Jacques, S.; Ba, N.; Ittah, M.; Lepajolec, C. Activation of ifn pathways and plasmacytoid dendritic cell recruitment in target organs of primary sjögren’s syndrome. Proc. Natl. Acad. Sci. USA 2006, 103, 2770–2775. [Google Scholar] [CrossRef] [PubMed]
- Ascough, S.; Paterson, S.; Chiu, C. Induction and subversion of human protective immunity: Contrasting influenza and respiratory syncytial virus. Front. Immunol. 2018, 9, 323. [Google Scholar] [CrossRef]
- Smillie, C.S.; Biton, M.; Ordovas-Montanes, J.; Sullivan, K.M.; Burgin, G.; Graham, D.B.; Herbst, R.H.; Rogel, N.; Slyper, M.; Waldman, J. Intra-and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 2019, 178, 714–730.e722. [Google Scholar] [CrossRef]
- Knobeloch, K.-P.; Wright, M.D.; Ochsenbein, A.F.; Liesenfeld, O.; Löhler, J.r.; Zinkernagel, R.M.; Horak, I.; Orinska, Z. Targeted inactivation of the tetraspanin CD37 impairs T-cell-dependent b-cell response under suboptimal costimulatory conditions. Mol. Cell. Biol. 2000, 20, 5363–5369. [Google Scholar] [CrossRef]
- Komili, S.; Farny, N.G.; Roth, F.P.; Silver, P.A. Functional specificity among ribosomal proteins regulates gene expression. Cell 2007, 131, 557–571. [Google Scholar] [CrossRef] [PubMed]
- Benvenuto, M.; Sileri, P.; Rossi, P.; Masuelli, L.; Fantini, M.; Nanni, M.; Franceschilli, L.; Sconocchia, G.; Lanzilli, G.; Arriga, R. Natural humoral immune response to ribosomal p0 protein in colorectal cancer patients. J. Transl. Med. 2015, 13, 101. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, G.; Liu, Y.; Chen, R.; Zhao, D.; McAlister, V.; Mele, T.; Liu, K.; Zheng, X. Gdf15 regulates malat-1 circular rna and inactivates nfκb signaling leading to immune tolerogenic dcs for preventing alloimmune rejection in heart transplantation. Front. Immunol. 2018, 9, 2407. [Google Scholar] [CrossRef]
- Kolseth, I.B.; Førland, D.T.; Risøe, P.K.; Flood-Kjeldsen, S.; Ågren, J.; Reseland, J.E.; Lyngstadaas, S.P.; Johnson, E.; Dahle, M.K. Human monocyte responses to lipopolysaccharide and 9-cis retinoic acid after laparoscopic surgery for colon cancer. Scand. J. Clin. Lab. Investig. 2012, 72, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, Y.; Yuan, J.; Li, N.; Pei, S.; Xu, J.; Luo, X.; Mao, C.; Liu, J.; Yu, T. Macrophage/microglial ezh2 facilitates autoimmune inflammation through inhibition of socs3. J. Exp. Med. 2018, 215, 1365–1382. [Google Scholar] [CrossRef] [PubMed]
- Lechner, M.G.; Cheng, M.I.; Patel, A.Y.; Hoang, A.T.; Yakobian, N.; Astourian, M.; Pioso, M.S.; Rodriguez, E.D.; McCarthy, E.C.; Hugo, W. Inhibition of il-17a protects against thyroid immune-related adverse events while preserving checkpoint inhibitor antitumor efficacy. J. Immunol. 2022, 209, 696–709. [Google Scholar] [CrossRef] [PubMed]
- Xia, P.; Xu, X.-Y. Prognostic significance of CD44 in human colon cancer and gastric cancer: Evidence from bioinformatic analyses. Oncotarget 2016, 7, 45538. [Google Scholar] [CrossRef]
- Gold, D.V.; Stein, R.; Burton, J.; Goldenberg, D.M. Enhanced expression of CD74 in gastrointestinal cancers and benign tissues. Int. J. Clin. Exp. Pathol. 2011, 4, 1–12. [Google Scholar]
- Beswick, E.J.; Reyes, V.E. Cd74 in antigen presentation, inflammation, and cancers of the gastrointestinal tract. World J. Gastroenterol. WJG 2009, 15, 2855. [Google Scholar] [CrossRef] [PubMed]
- Edvardsson, K.; Ström, A.; Jonsson, P.; Gustafsson, J.-. Å.; Williams, C. Estrogen receptor β induces antiinflammatory and antitumorigenic networks in colon cancer cells. Mol. Endocrinol. 2011, 25, 969–979. [Google Scholar] [CrossRef]
- Labriet, A.; Lévesque, É.; Cecchin, E.; De Mattia, E.; Villeneuve, L.; Rouleau, M.; Jonker, D.; Couture, F.; Simonyan, D.; Allain, E.P. Germline variability and tumor expression level of ribosomal protein gene rpl28 are associated with survival of metastatic colorectal cancer patients. Sci. Rep. 2019, 9, 13008. [Google Scholar] [CrossRef] [PubMed]
- Labriet, A.; Lévesque, É.; Mattia, E.D.; Cecchin, E.; Jonker, D.; Couture, F.; Simonyan, D.; Buonadonna, A.; D’Andrea, M.; Villeneuve, L. Rpl28 promoter polymorphism rs4806668 is associated with reduced survival in folfiri-treated metastatic colorectal cancer patients. Cancer Res. 2018, 78, 3889. [Google Scholar] [CrossRef]
- Nirmal, A.J.; Regan, T.; Shih, B.B.; Hume, D.A.; Sims, A.H.; Freeman, T.C. Immune cell gene signatures for profiling the microenvironment of solid tumorsimmune cell gene signatures for profiling solid tumors. Cancer Immunol. Res. 2018, 6, 1388–1400. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.C.; Chang, C.; Boschetti, G.; Ungaro, R.; Giri, M.; Grout, J.A.; Gettler, K.; Chuang, L.-S.; Nayar, S.; Greenstein, A.J. Single-cell analysis of crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-tnf therapy. Cell 2019, 178, 1493–1508.e1420. [Google Scholar] [CrossRef] [PubMed]
- Ziembik, M.A.; Bender, T.P.; Larner, J.M.; Brautigan, D.L. Functions of protein phosphatase-6 in nf-κb signaling and in lymphocytes. Biochem. Soc. Trans. 2017, 45, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Björklund, Å.K.; Forkel, M.; Picelli, S.; Konya, V.; Theorell, J.; Friberg, D.; Sandberg, R.; Mjösberg, J. The heterogeneity of human CD127+ innate lymphoid cells revealed by single-cell rna sequencing. Nat. Immunol. 2016, 17, 451–460. [Google Scholar] [CrossRef] [PubMed]



| Index | Immune Cell Subtypes | Sample Size |
|---|---|---|
| 1 | Activated CD4 T | 1531 |
| 2 | B cell (cycling) | 15 |
| 3 | B cell (IgA Plasma) | 12,522 |
| 4 | B cell (IgG Plasma) | 342 |
| 5 | B cell (memory) | 4508 |
| 6 | CD8 T | 3145 |
| 7 | cDC1 | 38 |
| 8 | cDC2 | 107 |
| 9 | cycling DCs | 47 |
| 10 | cycling gd T | 25 |
| 11 | Follicular B cell | 2582 |
| 12 | gd T | 548 |
| 13 | ILC | 832 |
| 14 | Lymphoid DC | 10 |
| 15 | LYVE1 Macrophage | 91 |
| 16 | Macrophage | 268 |
| 17 | Mast | 1151 |
| 18 | Monocyte | 98 |
| 19 | NK | 452 |
| 20 | pDC | 13 |
| 21 | Tcm | 3042 |
| 22 | Tfh | 1786 |
| 23 | Th1 | 2833 |
| 24 | Th17 | 3432 |
| 25 | Treg | 2232 |
| Feature List | Classification Algorithm | Number of Features | ACC | MCC | Macro F1 | Weighted F1 |
|---|---|---|---|---|---|---|
| LASSO feature list | Decision tree | 1240 | 0.823 | 0.796 | 0.775 | 0.827 |
| Random forest | 1230 | 0.960 | 0.953 | 0.953 | 0.959 | |
| Random forest | 50 | 0.933 | 0.923 | 0.937 | 0.933 | |
| LightGBM feature list | Decision tree | 300 | 0.895 | 0.878 | 0.871 | 0.895 |
| Random forest | 1200 | 0.978 | 0.974 | 0.985 | 0.978 | |
| Random forest | 60 | 0.943 | 0.934 | 0.949 | 0.943 | |
| MCFS feature list | Decision tree | 1620 | 0.892 | 0.875 | 0.881 | 0.893 |
| Random forest | 1720 | 0.976 | 0.972 | 0.983 | 0.976 | |
| Random forest | 90 | 0.944 | 0.935 | 0.954 | 0.944 | |
| mRMR feature list | Decision tree | 350 | 0.873 | 0.853 | 0.855 | 0.875 |
| Random forest | 560 | 0.972 | 0.968 | 0.973 | 0.972 | |
| Random forest | 90 | 0.961 | 0.955 | 0.969 | 0.961 | |
| RF feature list | Decision tree | 170 | 0.891 | 0.874 | 0.863 | 0.892 |
| Random forest | 1510 | 0.977 | 0.974 | 0.984 | 0.978 | |
| Random forest | 70 | 0.962 | 0.956 | 0.970 | 0.962 |
| Category | Gene Symbol | Description |
|---|---|---|
| T cell family | KLRB1 | Killer Cell Lectin Like Receptor B1 |
| CST3 | Cystatin C | |
| AIF1 | Allograft Inflammatory Factor 1 | |
| JCHAIN | Joining Chain of Multimeric IgA and IgM | |
| HLA-DRA | Major Histocompatibility Complex, Class II, DR Alpha | |
| TIGIT | T Cell Immunoreceptor with Ig and ITIM Domains | |
| RPS12 | Ribosomal Protein S12 | |
| MS4A1 | Membrane Spanning 4-Domains A1 | |
| B cell family | RPL30 | Ribosomal Protein L30 |
| ANXA1 | Annexin A1 | |
| JCHAIN | Joining Chain of Multimeric IgA and IgM | |
| ICA1 | Islet Cell Autoantigen 1 | |
| TYROBP | Transmembrane Immune Signaling Adaptor TYROBP | |
| RPS27 | Ribosomal Protein S27 | |
| HLA-DRA | Major Histocompatibility Complex, Class II, DR Alpha | |
| IFITM1 | Interferon Induced Transmembrane Protein 1 | |
| IGHA1 | Immunoglobulin Heavy Constant Alpha 1 | |
| CD37 | CD37 Molecule | |
| Other cells | RPS13 | Ribosomal Protein S13 |
| RPL34 | Ribosomal Protein L34 | |
| MALAT1 | Metastasis Associated Lung Adenocarcinoma Transcript 1 | |
| TMSB4X | Thymosin Beta 4 X-Linked | |
| SOCS3 | Suppressor Of Cytokine Signaling 3 | |
| IL7R | Interleukin 7 Receptor | |
| CD-74 | CD74 Molecule | |
| RPL28 | Ribosomal Protein L28 | |
| ANKRD28 | Ankyrin Repeat Domain 28 | |
| TYROBP | Transmembrane Immune Signaling Adaptor TYROBP |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.; Zhang, Y.; Ren, J.; Feng, K.; Li, Z.; Huang, T.; Cai, Y. Identification of Colon Immune Cell Marker Genes Using Machine Learning Methods. Life 2023, 13, 1876. https://doi.org/10.3390/life13091876
Yang Y, Zhang Y, Ren J, Feng K, Li Z, Huang T, Cai Y. Identification of Colon Immune Cell Marker Genes Using Machine Learning Methods. Life. 2023; 13(9):1876. https://doi.org/10.3390/life13091876
Chicago/Turabian StyleYang, Yong, Yuhang Zhang, Jingxin Ren, Kaiyan Feng, Zhandong Li, Tao Huang, and Yudong Cai. 2023. "Identification of Colon Immune Cell Marker Genes Using Machine Learning Methods" Life 13, no. 9: 1876. https://doi.org/10.3390/life13091876
APA StyleYang, Y., Zhang, Y., Ren, J., Feng, K., Li, Z., Huang, T., & Cai, Y. (2023). Identification of Colon Immune Cell Marker Genes Using Machine Learning Methods. Life, 13(9), 1876. https://doi.org/10.3390/life13091876









