Status and Challenges for Vaccination against Avian H9N2 Influenza Virus in China
Abstract
1. Introduction
2. Prevalence and Antigenic Drift of H9N2 AIVs under Vaccination in China
2.1. H9N2 AIV in Avian and Human
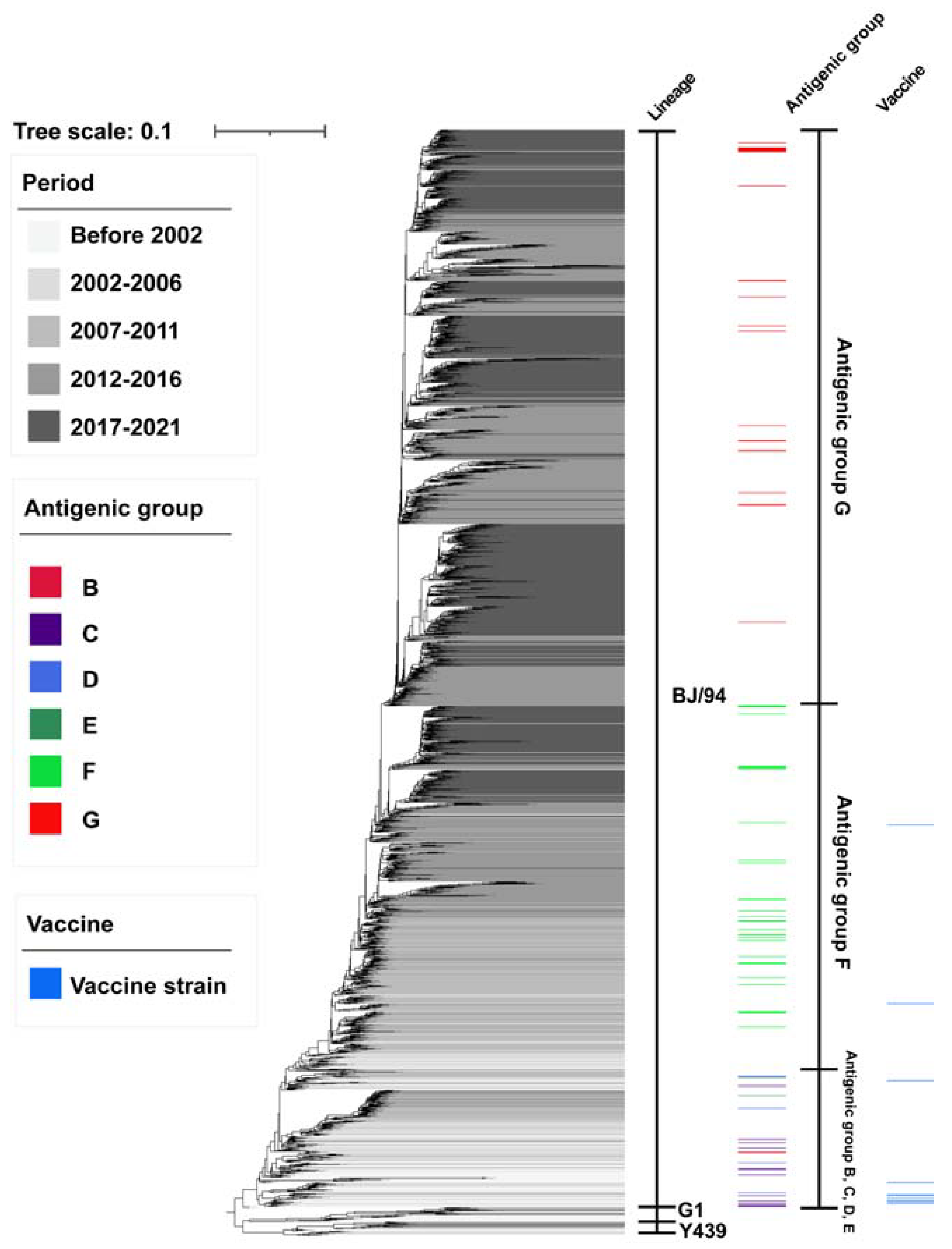
2.2. Antigenic Drift of H9N2 AIV under Vaccination in China
3. Vaccination Strategy for H9N2 AIV in China
4. Vaccine Development against H9N2-Subtype Avian Influenza
4.1. Inactivated Whole-Virus Vaccines
4.2. Vector Vaccine
4.3. Live-Attenuated Vaccine
4.4. DNA and mRNA Vaccines
4.5. Virus-like Particle Vaccine
4.6. Recombinant Protein Vaccine
5. Conclusions and Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Homme, P.J.; Easterday, B.C. Avian influenza virus infections. I. Characteristics of influenza A-turkey-Wisconsin-1966 virus. Avian Dis. 1970, 14, 66–74. [Google Scholar] [CrossRef]
- Lee, C.W.; Song, C.S.; Lee, Y.J.; Mo, I.P.; Garcia, M.; Suarez, D.L.; Kim, S.J. Sequence analysis of the hemagglutinin gene of H9N2 Korean avian influenza viruses and assessment of the pathogenic potential of isolate MS96. Avian Dis. 2000, 44, 527–535. [Google Scholar] [CrossRef]
- Naeem, K.; Ullah, A.; Manvell, R.J.; Alexander, D.J. Avian influenza A subtype H9N2 in poultry in Pakistan. Vet. Rec. 1999, 145, 560. [Google Scholar] [CrossRef]
- Nili, H.; Asasi, K. Natural cases and an experimental study of H9N2 avian influenza in commercial broiler chickens of Iran. Avian Pathol. 2002, 31, 247–252. [Google Scholar] [CrossRef]
- Chen, B.L.; Zhang, Z.J.; Chen, W.B. Study on avian influenza I. Isolation and preliminary serological identification of avian influenza A virus in chickens. China J. Vet. Med. 1994, 10, 3–5. [Google Scholar]
- Nagy, A.; Mettenleiter, T.C.; Abdelwhab, E.M. A brief summary of the epidemiology and genetic relatedness of avian influenza H9N2 virus in birds and mammals in the Middle East and North Africa. Epidemiol Infect. 2017, 145, 3320–3333. [Google Scholar] [CrossRef]
- Arafat, N.; Eladl, A.H.; Marghani, B.H.; Saif, M.A.; El-Shafei, R.A. Enhanced infection of avian influenza virus H9N2 with infectious laryngeotracheitis vaccination in chickens. Vet. Microbiol. 2018, 219, 8–16. [Google Scholar] [CrossRef]
- Sun, Y.; Liu, J. H9N2 influenza virus in China: A cause of concern. Protein Cell. 2015, 6, 18–25. [Google Scholar] [CrossRef]
- Jiang, W.; Liu, S.; Hou, G.; Li, J.; Zhuang, Q.; Wang, S.; Zhang, P.; Chen, J. Chinese and global distribution of H9 subtype avian influenza viruses. PLoS ONE 2012, 7, e52671. [Google Scholar] [CrossRef]
- Cong, Y.L.; Pu, J.; Liu, Q.F.; Wang, S.; Zhang, G.Z.; Zhang, X.L.; Fan, W.X.; Brown, E.G.; Liu, J.H. Antigenic and genetic characterization of H9N2 swine influenza viruses in China. J. Gen. Virol. 2007, 88, 2035–2041. [Google Scholar] [CrossRef]
- Sun, X.; Xu, X.; Liu, Q.; Liang, D.; Li, C.; He, Q.; Jiang, J.; Cui, Y.; Li, J.; Zheng, L. Evidence of avian-like H9N2 influenza A virus among dogs in Guangxi, China. Infect. Genet. Evol. 2013, 20, 471–475. [Google Scholar] [CrossRef]
- Song, W.; Qin, K. Human-infecting influenza A (H9N2) virus: A forgotten potential pandemic strain? Zoonoses Public Health 2020, 67, 203–212. [Google Scholar] [CrossRef]
- Lam, T.T.-Y.; Wang, J.; Shen, Y.; Zhou, B.; Duan, L.; Cheung, C.-L.; Ma, C.; Lycett, S.J.; Leung, C.Y.-H.; Chen, X. The genesis and source of the H7N9 influenza viruses causing human infections in China. Nature 2013, 502, 241–244. [Google Scholar] [CrossRef]
- Kageyama, T.; Fujisaki, S.; Takashita, E.; Xu, H.; Yamada, S.; Uchida, Y.; Neumann, G.; Saito, T.; Kawaoka, Y.; Tashiro, M. Genetic analysis of novel avian A (H7N9) influenza viruses isolated from patients in China, February to April 2013. Eurosurveillance. 2013, 18, 20453. [Google Scholar] [CrossRef]
- Chen, H.; Yuan, H.; Gao, R.; Zhang, J.; Wang, D.; Xiong, Y.; Fan, G.; Yang, F.; Li, X.; Zhou, J.; et al. Clinical and epidemiological characteristics of a fatal case of avian influenza A H10N8 virus infection: A descriptive study. Lancet 2014, 383, 714–721. [Google Scholar] [CrossRef]
- Shen, Y.-Y.; Ke, C.-W.; Li, Q.; Yuan, R.-Y.; Xiang, D.; Jia, W.-X.; Yu, Y.-D.; Liu, L.; Huang, C.; Qi, W.-B. Novel reassortant avian influenza A (H5N6) viruses in humans, Guangdong, China, 2015. Emerg. Infect. Dis. 2016, 22, 1507. [Google Scholar] [CrossRef]
- Burlington, D.; Clements, M.; Meiklejohn, G.; Phelan, M.; Murphy, B. Hemagglutinin-specific antibody responses in immunoglobulin G, A, and M isotypes as measured by enzyme-linked immunosorbent assay after primary or secondary infection of humans with influenza A virus. Infect. Immun. 1983, 41, 540–545. [Google Scholar] [CrossRef]
- Eichelberger, M.C.; Morens, D.M.; Taubenberger, J.K. Neuraminidase as an influenza vaccine antigen: A low hanging fruit, ready for picking to improve vaccine effectiveness. Curr. Opin. Immunol. 2018, 53, 38–44. [Google Scholar] [CrossRef]
- Memoli, M.J.; Shaw, P.A.; Han, A.; Czajkowski, L.; Reed, S.; Athota, R.; Bristol, T.; Fargis, S.; Risos, K.; Powers, J.H. Evaluation of antihemagglutinin and antineuraminidase antibodies as correlates of protection in an influenza A/H1N1 virus healthy human challenge model. Mbio 2016, 7, e00417-16. [Google Scholar] [CrossRef]
- Moskophidis, D.; Kioussis, D. Contribution of virus-specific CD8+ cytotoxic T cells to virus clearance or pathologic manifestations of influenza virus infection in a T cell receptor transgenic mouse model. J. Exp. Med. 1998, 188, 223–232. [Google Scholar] [CrossRef]
- Jansen, J.M.; Gerlach, T.; Elbahesh, H.; Rimmelzwaan, G.F.; Saletti, G. Influenza virus-specific CD4+ and CD8+ T cell-mediated immunity induced by infection and vaccination. J. Clin. Virol. 2019, 119, 44–52. [Google Scholar] [CrossRef]
- Sridhar, S. Heterosubtypic T-cell immunity to influenza in humans: Challenges for universal T-cell influenza vaccines. Front. Immunol. 2016, 7, 195. [Google Scholar] [CrossRef]
- Mettelman, R.C.; Allen, E.K.; Thomas, P.G. Mucosal immune responses to infection and vaccination in the respiratory tract. Immunity 2022, 55, 749–780. [Google Scholar] [CrossRef]
- Zens, K.D.; Chen, J.K.; Farber, D.L. Vaccine-generated lung tissue–resident memory T cells provide heterosubtypic protection to influenza infection. JCI Insight 2016, 1, e85832. [Google Scholar] [CrossRef]
- Pizzolla, A.; Nguyen, T.H.; Smith, J.M.; Brooks, A.G.; Kedzierska, K.; Heath, W.R.; Reading, P.C.; Wakim, L.M. Resident memory CD8+ T cells in the upper respiratory tract prevent pulmonary influenza virus infection. Sci. Immunol. 2017, 2, eaam6970. [Google Scholar] [CrossRef]
- Guan, Y.; Shortridge, K.F.; Krauss, S.; Chin, P.S.; Dyrting, K.C.; Ellis, T.M.; Webster, R.G.; Peiris, M. H9N2 influenza viruses possessing H5N1-like internal genomes continue to circulate in poultry in southeastern China. J. Virol. 2000, 74, 9372–9380. [Google Scholar] [CrossRef]
- Guo, Y.J.; Krauss, S.; Senne, D.A.; Mo, I.P.; Lo, K.S.; Xiong, X.P.; Norwood, M.; Shortridge, K.F.; Webster, R.G.; Guan, Y. Characterization of the pathogenicity of members of the newly established H9N2 influenza virus lineages in Asia. Virology 2000, 267, 279–288. [Google Scholar] [CrossRef]
- Liu, H.; Liu, X.; Cheng, J.; Peng, D.; Jia, L.; Huang, Y. Phylogenetic analysis of the hemagglutinin genes of twenty-six avian influenza viruses of subtype H9N2 isolated from chickens in China during 1996-2001. Avian Dis. 2003, 47, 116–127. [Google Scholar] [CrossRef]
- Liu, J.H.; Okazaki, K.; Shi, W.M.; Wu, Q.M.; Mweene, A.S.; Kida, H. Phylogenetic analysis of neuraminidase gene of H9N2 influenza viruses prevalent in chickens in China during 1995-2002. Virus Genes 2003, 27, 197–202. [Google Scholar] [CrossRef]
- Sun, Y.; Pu, J.; Jiang, Z.; Guan, T.; Xia, Y.; Xu, Q.; Liu, L.; Ma, B.; Tian, F.; Brown, E.G.; et al. Genotypic evolution and antigenic drift of H9N2 influenza viruses in China from 1994 to 2008. Vet. Microbiol. 2010, 146, 215–225. [Google Scholar] [CrossRef]
- Ge, F.F.; Zhou, J.P.; Liu, J.; Wang, J.; Zhang, W.Y.; Sheng, L.P.; Xu, F.; Ju, H.B.; Sun, Q.Y.; Liu, P.H. Genetic evolution of H9 subtype influenza viruses from live poultry markets in Shanghai, China. J. Clin. Microbiol. 2009, 47, 3294–3300. [Google Scholar] [CrossRef]
- Xu, K.M.; Smith, G.J.; Bahl, J.; Duan, L.; Tai, H.; Vijaykrishna, D.; Wang, J.; Zhang, J.X.; Li, K.S.; Fan, X.H.; et al. The genesis and evolution of H9N2 influenza viruses in poultry from southern China, 2000 to 2005. J. Virol. 2007, 81, 10389–10401. [Google Scholar] [CrossRef]
- Li, C.; Yu, K.; Tian, G.; Yu, D.; Liu, L.; Jing, B.; Ping, J.; Chen, H. Evolution of H9N2 influenza viruses from domestic poultry in Mainland China. Virology 2005, 340, 70–83. [Google Scholar] [CrossRef]
- Zhang, Y.; Yin, Y.; Bi, Y.; Wang, S.; Xu, S.; Wang, J.; Zhou, S.; Sun, T.; Yoon, K.J. Molecular and antigenic characterization of H9N2 avian influenza virus isolates from chicken flocks between 1998 and 2007 in China. Vet. Microbiol. 2012, 156, 285–293. [Google Scholar] [CrossRef]
- Pu, J.; Wang, S.; Yin, Y.; Zhang, G.; Carter, R.A.; Wang, J.; Xu, G.; Sun, H.; Wang, M.; Wen, C.; et al. Evolution of the H9N2 influenza genotype that facilitated the genesis of the novel H7N9 virus. Proc. Natl. Acad. Sci. USA 2015, 112, 548–553. [Google Scholar] [CrossRef]
- Pu, J.; Sun, H.; Qu, Y.; Wang, C.; Gao, W.; Zhu, J.; Sun, Y.; Bi, Y.; Huang, Y.; Chang, K.C.; et al. M Gene Reassortment in H9N2 Influenza Virus Promotes Early Infection and Replication: Contribution to Rising Virus Prevalence in Chickens in China. J. Virol. 2017, 91, e02055-16. [Google Scholar] [CrossRef]
- Pu, J.; Yin, Y.; Liu, J.; Wang, X.; Zhou, Y.; Wang, Z.; Sun, Y.; Sun, H.; Li, F.; Song, J.; et al. Reassortment with dominant chicken H9N2 influenza virus contributed to the fifth H7N9 virus human epidemic. J. Virol. 2021, 95, e01578-20. [Google Scholar] [CrossRef]
- Bi, Y.; Chen, Q.; Wang, Q.; Chen, J.; Jin, T.; Wong, G.; Quan, C.; Liu, J.; Wu, J.; Yin, R.; et al. Genesis, Evolution and Prevalence of H5N6 Avian Influenza Viruses in China. Cell Host Microbe. 2016, 20, 810–821. [Google Scholar] [CrossRef]
- Bi, Y.; Li, J.; Li, S.; Fu, G.; Jin, T.; Zhang, C.; Yang, Y.; Ma, Z.; Tian, W.; Li, J.; et al. Dominant subtype switch in avian influenza viruses during 2016-2019 in China. Nat. Commun. 2020, 11, 5909. [Google Scholar] [CrossRef]
- Guo, Y.J.; Li, J.G.; Cheng, X.W.; Wang, M.; Zou, Y.; Li, Z.H.; Cai, F.C.; Liao, H.L.; Zhang, Y.; Guo, J.F.; et al. Discovery that avian H9N2 subtype influenza virus can infect humans. Zhonghua Shi Yan He Lin Chuang Bing Du Xue Za Zhi. 1999, 5–8. [Google Scholar]
- Adlhoch, C.; Fusaro, A.; Gonzales, J.L.; Kuiken, T.; Marangon, S.; Niqueux, É.; Staubach, C.; Terregino, C.; Aznar, I.; Muñoz Guajardo, I.; et al. Avian influenza overview September—December 2021. EFSA J. Eur. Food Saf. Auth. 2021, 19, e07108. [Google Scholar] [CrossRef]
- Guan, Y.; Shortridge, K.F.; Krauss, S.; Webster, R.G. Molecular characterization of H9N2 influenza viruses: Were they the donors of the "internal" genes of H5N1 viruses in Hong Kong? Proc. Natl. Acad. Sci. USA 1999, 96, 9363–9367. [Google Scholar] [CrossRef]
- Qi, W.; Zhou, X.; Shi, W.; Huang, L.; Xia, W.; Liu, D.; Li, H.; Chen, S.; Lei, F.; Cao, L.; et al. Genesis of the novel human-infecting influenza A(H10N8) virus and potential genetic diversity of the virus in poultry, China. Euro Surveill. Bull. Eur. Sur Les Mal. Transm. = Eur. Commun. Dis. Bull. 2014, 19, 20841. [Google Scholar] [CrossRef]
- Li, F.; Liu, J.; Yang, J.; Sun, H.; Jiang, Z.; Wang, C.; Zhang, X.; Yu, Y.; Zhao, C.; Pu, J.; et al. H9N2 virus-derived M1 protein promotes H5N6 virus release in mammalian cells: Mechanism of avian influenza virus inter-species infection in humans. PLoS Pathog. 2021, 17, e1010098. [Google Scholar] [CrossRef]
- Liu, D.; Shi, W.; Gao, G.F. Poultry carrying H9N2 act as incubators for novel human avian influenza viruses. Lancet 2014, 383, 869. [Google Scholar] [CrossRef]
- Wiley, D.C.; Wilson, I.A.; Skehel, J.J. Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 1981, 289, 373–378. [Google Scholar] [CrossRef]
- Caton, A.J.; Brownlee, G.G.; Yewdell, J.W.; Gerhard, W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 1982, 31 Pt 1, 417–427. [Google Scholar] [CrossRef]
- Okamatsu, M.; Sakoda, Y.; Kishida, N.; Isoda, N.; Kida, H. Antigenic structure of the hemagglutinin of H9N2 influenza viruses. Arch. Virol. 2008, 153, 2189–2195. [Google Scholar] [CrossRef]
- Peacock, T.P.; Harvey, W.T.; Sadeyen, J.R.; Reeve, R.; Iqbal, M. The molecular basis of antigenic variation among A(H9N2) avian influenza viruses. Emerg. Microbes Infect. 2018, 7, 176. [Google Scholar] [CrossRef]
- Wan, Z.; Ye, J.; Xu, L.; Shao, H.; Jin, W.; Qian, K.; Wan, H.; Qin, A. Antigenic mapping of the hemagglutinin of an H9N2 avian influenza virus reveals novel critical amino acid positions in antigenic sites. J. Virol. 2014, 88, 3898–3901. [Google Scholar] [CrossRef]
- Peacock, T.; Reddy, K.; James, J.; Adamiak, B.; Barclay, W.; Shelton, H.; Iqbal, M. Antigenic mapping of an H9N2 avian influenza virus reveals two discrete antigenic sites and a novel mechanism of immune escape. Sci. Rep. 2016, 6, 18745. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Yang, D.; Ren, Q.; Yang, Y.; Liu, X.; Xu, X.; Liu, W.; Chen, S.; Peng, D.; Liu, X. Identification and characterization of a novel antigenic epitope in the hemagglutinin of the escape mutants of H9N2 avian influenza viruses. Vet. Microbiol. 2015, 178, 144–149. [Google Scholar] [CrossRef]
- Yan, W.; Cui, H.; Engelsma, M.; Beerens, N.; van Oers, M.M.; de Jong, M.C.M.; Li, X.; Liu, Q.; Yang, J.; Teng, Q.; et al. Molecular and Antigenic Characterization of Avian H9N2 Viruses in Southern China. Microbiol. Spectr. 2022, 10, e0082221. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Guo, Y.; Li, Y.; Liang, B.; Sun, X.; Li, S.; Xia, H.; Ping, J. The molecular determinants of antigenic drift in a novel avian influenza A (H9N2) variant virus. Virol. J. 2022, 19, 26. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Zhao, L.; Guo, Y.; Zhao, Y.; Li, Y.; Chen, N.; Lu, Y.; Yu, M.; Deng, L.; Ping, J. Antigenic Evolution Characteristics and Immunological Evaluation of H9N2 Avian Influenza Viruses from 1994-2019 in China. Viruses 2022, 14. [Google Scholar] [CrossRef]
- Wei, Y.; Xu, G.; Zhang, G.; Wen, C.; Anwar, F.; Wang, S.; Lemmon, G.; Wang, J.; Carter, R.; Wang, M.; et al. Antigenic evolution of H9N2 chicken influenza viruses isolated in China during 2009-2013 and selection of a candidate vaccine strain with broad cross-reactivity. Vet. Microbiol. 2016, 182, 1–7. [Google Scholar] [CrossRef]
- Liu, S.; Zhuang, Q.; Wang, S.; Jiang, W.; Jin, J.; Peng, C.; Hou, G.; Li, J.; Yu, J.; Yu, X.; et al. Control of avian influenza in China: Strategies and lessons. Transbound. Emerg. Dis. 2020, 67, 1463–1471. [Google Scholar] [CrossRef]
- Liu, L.; Zeng, X.; Chen, P.; Deng, G.; Li, Y.; Shi, J.; Gu, C.; Kong, H.; Suzuki, Y.; Jiang, Y. Characterization of clade 7.2 H5 avian influenza viruses that continue to circulate in chickens in China. J. Virol. 2016, 90, 9797–9805. [Google Scholar] [CrossRef]
- Sun, Y.; Pu, J.; Fan, L.; Sun, H.; Wang, J.; Zhang, Y.; Liu, L.; Liu, J. Evaluation of the protective efficacy of a commercial vaccine against different antigenic groups of H9N2 influenza viruses in chickens. Vet. Microbiol. 2012, 156, 193–199. [Google Scholar] [CrossRef]
- Zhong, L.; Wang, X.; Li, Q.; Liu, D.; Chen, H.; Zhao, M.; Gu, X.; He, L.; Liu, X.; Gu, M. Molecular mechanism of the airborne transmissibility of H9N2 avian influenza A viruses in chickens. J. Virol. 2014, 88, 9568–9578. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Z.; Su, X.; Qiao, Y.; Fan, W.; Li, H.; Shi, B.; Qin, T.; Chen, S.; Peng, D.; et al. Enhanced cross-lineage protection induced by recombinant H9N2 avian influenza virus inactivated vaccine. Vaccine 2019, 37, 1736–1742. [Google Scholar] [CrossRef]
- Du, X.; Dong, L.; Lan, Y.; Peng, Y.; Wu, A.; Zhang, Y.; Huang, W.; Wang, D.; Wang, M.; Guo, Y.; et al. Mapping of H3N2 influenza antigenic evolution in China reveals a strategy for vaccine strain recommendation. Nat. Commun. 2012, 3, 709. [Google Scholar] [CrossRef] [PubMed]
- Radvak, P.; Kosikova, M.; Kuo, Y.C.; Li, X.; Garner, R.; Schmeisser, F.; Kosik, I.; Ye, Z.; Weir, J.P.; Yewdell, J.W.; et al. Highly pathogenic avian influenza A/Guangdong/17SF003/2016 is immunogenic and induces cross-protection against antigenically divergent H7N9 viruses. NPJ Vaccines. 2021, 6, 30. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Tang, G.Y.; Feng, H.L.; Xue, Y.H.; Ren, Z.; Wang, G.K.; Jia, M.M.; Shang, Y.; Luo, Q.P.; Shao, H.B.; et al. Evaluation of Immune Efficacy of H9 Subtype Avian Influenza Virus Inactivated Vaccine Based on Mosaic HA Sequence. Acta Vet. Et Zootech. Sin. 2021, 52, 3569–3577. [Google Scholar]
- Bullard, B.L.; Corder, B.N.; DeBeauchamp, J.; Rubrum, A.; Korber, B.; Webby, R.J.; Weaver, E.A. Epigraph hemagglutinin vaccine induces broad cross-reactive immunity against swine H3 influenza virus. Nat. Commun. 2021, 12, 1203. [Google Scholar] [CrossRef]
- Bullard, B.L.; DeBeauchamp, J.; Pekarek, M.J.; Petro-Turnquist, E.; Vogel, P.; Webby, R.J.; Weaver, E.A. An epitope-optimized human H3N2 influenza vaccine induces broadly protective immunity in mice and ferrets. NPJ Vaccines 2022, 7, 65. [Google Scholar] [CrossRef]
- Kim, S.M.; Kim, Y.I.; Park, S.J.; Kim, E.H.; Kwon, H.I.; Si, Y.J.; Lee, I.W.; Song, M.S.; Choi, Y.K. Vaccine Efficacy of Inactivated, Chimeric Hemagglutinin H9/H5N2 Avian Influenza Virus and Its Suitability for the Marker Vaccine Strategy. J. Virol. 2017, 91, e01693-16. [Google Scholar] [CrossRef]
- Ping, J.; Lopes, T.J.S.; Nidom, C.A.; Ghedin, E.; Macken, C.A.; Fitch, A.; Imai, M.; Maher, E.A.; Neumann, G.; Kawaoka, Y. Development of high-yield influenza A virus vaccine viruses. Nat. Commun. 2015, 6, 8148. [Google Scholar] [CrossRef]
- An, S.H.; Lee, C.Y.; Choi, J.G.; Lee, Y.J.; Kim, J.H.; Kwon, H.J. Generation of highly productive and mammalian nonpathogenic recombinant H9N2 avian influenza viruses by optimization of 3’end promoter and NS genome. Vet. Microbiol. 2019, 228, 213–218. [Google Scholar] [CrossRef]
- Song, C.L.; Liao, Z.H.; Shen, Y.; Wang, H.; Lin, W.C.; Li, H.; Chen, W.G.; Xie, Q.M. Assessing the efficacy of a recombinant H9N2 avian influenza virus-inactivated vaccine. Poult Sci. 2020, 99, 4334–4342. [Google Scholar] [CrossRef]
- Astill, J.; Alkie, T.; Yitbarek, A.; Taha-Abdelaziz, K.; Bavananthasivam, J.; Nagy, E.; Petrik, J.J.; Sharif, S. Examination of the effects of virus inactivation methods on the induction of antibody- and cell-mediated immune responses against whole inactivated H9N2 avian influenza virus vaccines in chickens. Vaccine 2018, 36, 3908–3916. [Google Scholar] [CrossRef]
- Qin, T.; Yin, Y.; Yu, Q.; Huang, L.; Wang, X.; Lin, J.; Yang, Q. CpG Oligodeoxynucleotides Facilitate Delivery of Whole Inactivated H9N2 Influenza Virus via Transepithelial Dendrites of Dendritic Cells in Nasal Mucosa. J. Virol. 2015, 89, 5904–5918. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Qin, T.; Wang, X.; Lin, J.; Yu, Q.; Yang, Q. CpG DNA assists the whole inactivated H9N2 influenza virus in crossing the intestinal epithelial barriers via transepithelial uptake of dendritic cell dendrites. Mucosal Immunol. 2015, 8, 799–814. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Xue, M.; Wang, C.; Wang, J.; Chen, P. Bursopentine as a novel immunoadjuvant enhances both humoral and cell-mediated immune responses to inactivated H9N2 Avian Influenza virus in chickens. Clin. Vaccine Immunol. 2011, 18, 1497–1502. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, X.; Wu, T.; Li, D.; Niu, M.; Wang, Y.; Zhang, C.; Cheng, X.; Chen, P. Bursin-like peptide (BLP) enhances H9N2 influenza vaccine induced humoral and cell mediated immune responses. Cell Immunol. 2014, 292, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Lai, H.; Xu, J.; Huang, W.; Liu, Y.; Zhao, D.; Chen, R. Evaluation of the Protective Efficacy of Poly I:C as an Adjuvant for H9N2 Subtype Avian Influenza Inactivated Vaccine and Its Mechanism of Action in Ducks. PLoS ONE 2017, 12, e0170681. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, C.; Zhou, J.; Liu, Z.; Liu, Y.; Cai, K.; Shen, T.; Liao, C.; Wang, C. Comparison of immunoadjuvant activities of four bursal peptides combined with H9N2 avian influenza virus vaccine. J. Vet. Sci. 2018, 19, 817–826. [Google Scholar] [CrossRef]
- Khalili, I.; Ghadimipour, R.; Sadigh Eteghad, S.; Fathi Najafi, M.; Ebrahimi, M.M.; Godsian, N.; Sefidi Heris, Y.; Khalili, M.T. Evaluation of Immune Response Against Inactivated Avian Influenza (H9N2) Vaccine, by Using Chitosan Nanoparticles. Jundishapur. J. Microbiol. 2015, 8, e27035. [Google Scholar] [CrossRef]
- Lee, J.E.; Kye, Y.C.; Park, S.M.; Shim, B.S.; Yoo, S.; Hwang, E.; Kim, H.; Kim, S.J.; Han, S.H.; Park, T.S.; et al. Bacillus subtilis spores as adjuvants against avian influenza H9N2 induce antigen-specific antibody and T cell responses in White Leghorn chickens. Vet. Res. 2020, 51, 68. [Google Scholar] [CrossRef]
- Gu, P.; Wusiman, A.; Zhang, Y.; Cai, G.; Xu, S.; Zhu, S.; Liu, Z.; Hu, Y.; Liu, J.; Wang, D. Polyethylenimine-coated PLGA nanoparticles-encapsulated Angelica sinensis polysaccharide as an adjuvant for H9N2 vaccine to improve immune responses in chickens compared to Alum and oil-based adjuvants. Vet. Microbiol. 2020, 251, 108894. [Google Scholar] [CrossRef]
- Chen, H.Y.; Shang, Y.H.; Yao, H.X.; Cui, B.A.; Zhang, H.Y.; Wang, Z.X.; Wang, Y.D.; Chao, A.J.; Duan, T.Y. Immune responses of chickens inoculated with a recombinant fowlpox vaccine coexpressing HA of H9N2 avain influenza virus and chicken IL-18. Antivir. Res. 2011, 91, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Swayne, D.E.; Beck, J.R.; Kinney, N. Failure of a recombinant fowl poxvirus vaccine containing an avian influenza hemagglutinin gene to provide consistent protection against influenza in chickens preimmunized with a fowl pox vaccine. Avian Dis. 2000, 44, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Pan, Q.; Zhang, Y.; Liu, A.; Cui, H.; Gao, Y.; Qi, X.; Liu, C.; Zhang, Y.; Li, K.; Gao, L. Development of a Novel Avian Vaccine Vector Derived From the Emerging Fowl Adenovirus 4. Front. Microbiol. 2021, 12, 780978. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Zhang, Z.; Zhao, P.; Duan, L.; Zhang, Y.; Zhang, F.; Chen, W.; Cui, Z. Comparative transcriptional activity of five promoters in BAC-cloned MDV for the expression of the hemagglutinin gene of H9N2 avian influenza virus. J. Virol. Methods 2014, 206, 119–127. [Google Scholar] [CrossRef]
- Liu, L.; Wang, T.; Wang, M.; Tong, Q.; Sun, Y.; Pu, J.; Sun, H.; Liu, J. Recombinant turkey herpesvirus expressing H9 hemagglutinin providing protection against H9N2 avian influenza. Virology 2019, 529, 7–15. [Google Scholar] [CrossRef]
- Nagy, A.; Lee, J.; Mena, I.; Henningson, J.; Li, Y.; Ma, J.; Duff, M.; Li, Y.; Lang, Y.; Yang, J.; et al. Recombinant Newcastle disease virus expressing H9 HA protects chickens against heterologous avian influenza H9N2 virus challenge. Vaccine 2016, 34, 2537–2545. [Google Scholar] [CrossRef]
- Liu, J.; Xue, L.; Hu, S.; Cheng, H.; Deng, Y.; Hu, Z.; Wang, X.; Liu, X. Chimeric Newcastle disease virus-vectored vaccine protects chickens against H9N2 avian influenza virus in the presence of pre-existing NDV immunity. Arch. Virol. 2018, 163, 3365–3371. [Google Scholar] [CrossRef]
- Xu, X.; Xue, C.; Liu, X.; Li, J.; Fei, Y.; Liu, Z.; Mu, J.; Bi, Y.; Qian, J.; Yin, R.; et al. A novel recombinant attenuated Newcastle disease virus expressing H9 subtype hemagglutinin protected chickens from challenge by genotype VII virulent Newcastle disease virus and H9N2 avian influenza virus. Vet. Microbiol. 2019, 228, 173–180. [Google Scholar] [CrossRef]
- Zhang, X.; Bo, Z.; Meng, C.; Chen, Y.; Zhang, C.; Cao, Y.; Wu, Y. Generation and Evaluation of Recombinant Thermostable Newcastle Disease Virus Expressing the HA of H9N2 Avian Influenza Virus. Viruses 2021, 13, 1606. [Google Scholar] [CrossRef]
- Yang, W.; Dai, J.; Liu, J.; Guo, M.; Liu, X.; Hu, S.; Gu, M.; Hu, J.; Hu, Z.; Gao, R.; et al. Intranasal Immunization with a Recombinant Avian Paramyxovirus Serotypes 2 Vector-Based Vaccine Induces Protection against H9N2 Avian Influenza in Chicken. Viruses 2022, 14, 918. [Google Scholar] [CrossRef]
- Chen, H.; Matsuoka, Y.; Chen, Q.; Cox, N.; Murphy, B.; Subbarao, K. Generation and characterization of an H9N2 cold-adapted reassortant as a vaccine candidate. Avian Dis. 2003, 47, 1127–1130. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Kim, H.S.; Seo, S.H. Genetic characterization and protective immunity of cold-adapted attenuated avian H9N2 influenza vaccine. Vaccine 2008, 26, 6569–6576. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Pu, J.; Wei, Y.; Sun, Y.; Hu, J.; Liu, L.; Xu, G.; Gao, W.; Li, C.; Zhang, X.; et al. Highly Pathogenic Avian Influenza H5N6 Viruses Exhibit Enhanced Affinity for Human Type Sialic Acid Receptor and In-Contact Transmission in Model Ferrets. J. Virol. 2016, 90, 6235–6243. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhu, Y.; Yang, D.; Yang, Y.; Shi, S.; Qin, T.; Peng, D.; Liu, X. Efficacy of Live-Attenuated H9N2 Influenza Vaccine Candidates Containing NS1 Truncations against H9N2 Avian Influenza Viruses. Front. Microbiol. 2017, 8, 1086. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Quan, K.; Wang, H.; Li, S.; Xue, J.; Qin, T.; Chu, D.; Fan, G.; Du, Y.; Peng, D. A Live Attenuated H9N2 Avian Influenza Vaccine Prevents the Viral Reassortment by Exchanging the HA and NS1 Packaging Signals. Front. Microbiol. 2020, 11, 613437. [Google Scholar] [CrossRef] [PubMed]
- Si, L.; Xu, H.; Zhou, X.; Zhang, Z.; Tian, Z.; Wang, Y.; Wu, Y.; Zhang, B.; Niu, Z.; Zhang, C. Generation of influenza A viruses as live but replication-incompetent virus vaccines. Science 2016, 354, 1170–1173. [Google Scholar] [CrossRef] [PubMed]
- Dudek, T.; Knipe, D.M. Replication-defective viruses as vaccines and vaccine vectors. Virology 2006, 344, 230–239. [Google Scholar] [CrossRef]
- Victor, S.T.; Watanabe, S.; Katsura, H.; Ozawa, M.; Kawaoka, Y. A replication-incompetent PB2-knockout influenza A virus vaccine vector. J. Virol. 2012, 86, 4123–4128. [Google Scholar] [CrossRef]
- Zhao, K.; Rong, G.; Teng, Q.; Li, X.; Lan, H.; Yu, L.; Yu, S.; Jin, Z.; Chen, G.; Li, Z. Dendrigraft poly-L-lysines delivery of DNA vaccine effectively enhances the immunogenic responses against H9N2 avian influenza virus infection in chickens. Nanomed. Nanotechnol. Biol. Med. 2020, 27, 102209. [Google Scholar] [CrossRef]
- Feldman, R.A.; Fuhr, R.; Smolenov, I.; Ribeiro, A.M.; Panther, L.; Watson, M.; Senn, J.J.; Smith, M.; Almarsson, Ӧ.; Pujar, H.S. mRNA vaccines against H10N8 and H7N9 influenza viruses of pandemic potential are immunogenic and well tolerated in healthy adults in phase 1 randomized clinical trials. Vaccine 2019, 37, 3326–3334. [Google Scholar] [CrossRef]
- Li, X.; Ju, H.; Liu, J.; Yang, D.; Qi, X.; Yang, X.; Qiu, Y.; Zheng, J.; Ge, F.; Zhou, J. Influenza virus-like particles harboring H9N2 HA and NA proteins induce a protective immune response in chicken. Influenza Other Respi. Viruses 2017, 11, 518–524. [Google Scholar] [CrossRef] [PubMed]
- Tretyakova, I.; Pearce, M.B.; Florese, R.; Tumpey, T.M.; Pushko, P. Intranasal vaccination with H5, H7 and H9 hemagglutinins co-localized in a virus-like particle protects ferrets from multiple avian influenza viruses. Virology 2013, 442, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Pushko, P.; Tretyakova, I.; Hidajat, R.; Zsak, A.; Chrzastek, K.; Tumpey, T.M.; Kapczynski, D.R. Virus-like particles displaying H5, H7, H9 hemagglutinins and N1 neuraminidase elicit protective immunity to heterologous avian influenza viruses in chickens. Virology 2017, 501, 176–182. [Google Scholar] [CrossRef] [PubMed]
- Jáuregui-Zúñiga, D.; Pedraza-Escalona, M.; Espino-Solís, G.P.; Quintero-Hernández, V.; Olvera-Rodríguez, A.; Díaz-Salinas, M.A.; López, S.; Possani, L.D. Targeting antigens to Dec-205 on dendritic cells induces a higher immune response in chickens: Hemagglutinin of avian influenza virus example. Res. Vet. Sci. 2017, 111, 55–62. [Google Scholar] [CrossRef]
- Wang, T.; Wei, F.; Liu, L.; Sun, Y.; Song, J.; Wang, M.; Yang, J.; Li, C.; Liu, J. Recombinant HA1-DeltafliC enhances adherence to respiratory epithelial cells and promotes the superiorly protective immune responses against H9N2 influenza virus in chickens. Vet. Microbiol. 2021, 262, 109238. [Google Scholar] [CrossRef]
- Akauliya, M.; Gautam, A.; Maharjan, S.; Park, B.K.; Kim, J.; Kwon, H.-J. CD83 expression regulates antibody production in response to influenza A virus infection. Virol. J. 2020, 17, 1–11. [Google Scholar] [CrossRef]
- Aida, V.; Pliasas, V.C.; Neasham, P.J.; North, J.F.; McWhorter, K.L.; Glover, S.R.; Kyriakis, C.S. Novel Vaccine Technologies in Veterinary Medicine: A Herald to Human Medicine Vaccines. Front. Vet. Sci. 2021, 8, 654289. [Google Scholar] [CrossRef]
- Shi, S.H.; Yang, W.T.; Yang, G.L.; Cong, Y.L.; Huang, H.B.; Wang, Q.; Cai, R.P.; Ye, L.P.; Hu, J.T.; Zhou, J.Y.; et al. Immunoprotection against influenza virus H9N2 by the oral administration of recombinant Lactobacillus plantarumNC8 expressing hemagglutinin in BALB/c mice. Virology 2014, 464-465, 166–176. [Google Scholar] [CrossRef]
- Shi, S.H.; Yang, W.T.; Yang, G.L.; Zhang, X.K.; Liu, Y.Y.; Zhang, L.J.; Ye, L.P.; Hu, J.T.; Xing, X.; Qi, C.; et al. Lactobacillus plantarum vaccine vector expressing hemagglutinin provides protection against H9N2 challenge infection. Virus Res. 2016, 211, 46–57. [Google Scholar] [CrossRef]
- Bo, F.; Yang, W.T.; Shonyela, S.M.; Jin, Y.B.; Huang, K.Y.; Shao, L.N.; Wang, C.; Zhou, Y.; Li, Q.Y.; Jiang, Y.L.; et al. Immune responses of mice inoculated with recombinant Lactobacillus plantarum NC8 expressing the fusion gene HA2 and 3M2e of the influenza virus and protection against different subtypes of influenza virus. Virus Res. 2019, 263, 64–72. [Google Scholar] [CrossRef]
- Li, Q.Y.; Xu, M.M.; Dong, H.; Zhao, J.H.; Xing, J.H.; Wang, G.; Yao, J.Y.; Huang, H.B.; Shi, C.W.; Jiang, Y.L.; et al. Lactobacillus plantarum surface-displayed influenza antigens (NP-M2) with FliC flagellin stimulate generally protective immune responses against H9N2 influenza subtypes in chickens. Vet. Microbiol. 2020, 249, 108834. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Tang, X.; Wang, S.; Shi, F.; Duan, C.; Bi, F.; Suo, J.; Hu, D.; Liu, J.; Wang, C.; et al. Establishment of Recombinant Eimeria acervulina Expressing Multi-Copies M2e Derived from Avian Influenza Virus H9N2. Vaccines 2021, 9, 791. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, Y.; Huang, Y.; Liu, X. Protection conferred by a recombinant Marek’s disease virus that expresses the spike protein from infectious bronchitis virus in specific pathogen-free chicken. Virol. J. 2012, 9, 1–10. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, D.; Xiong, J.; Zhang, P.; Li, Y.; She, R. Protection of chickens, with or without maternal antibodies, against IBDV infection by a recombinant IBDV-VP2 protein. Vaccine 2010, 28, 3990–3996. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Liu, Y.; Liu, C.; Gao, L.; Zhang, Y.; Cui, H.; Gao, Y.; Qi, X.; Zhong, L.; Wang, X. Recombinant Marek’s disease virus type 1 provides full protection against very virulent Marek’s and infectious bursal disease viruses in chickens. Sci. Rep. 2016, 6, 1–10. [Google Scholar] [CrossRef]
- Liu, Y.; Li, K.; Gao, Y.; Gao, L.; Zhong, L.; Zhang, Y.; Liu, C.; Zhang, Y.; Wang, X. Recombinant Marek’s disease virus as a vector-based vaccine against avian leukosis virus subgroup J in chicken. Viruses 2016, 8, 301. [Google Scholar] [CrossRef]
- Cui, H.; Gao, H.; Cui, X.; Zhao, Y.; Shi, X.; Li, Q.; Yan, S.; Gao, M.; Wang, M.; Liu, C. Avirulent Marek’s disease virus type 1 strain 814 vectored vaccine expressing avian influenza (AI) virus H5 haemagglutinin induced better protection than turkey herpesvirus vectored AI vaccine. PloS ONE 2013, 8, e53340. [Google Scholar]
- Sun, Y.; Yang, C.; Li, J.; Li, L.; Cao, M.; Li, Q.; Li, H. Construction of a recombinant duck enteritis virus vaccine expressing hemagglutinin of H9N2 avian influenza virus and evaluation of its efficacy in ducks. Arch. Virol. 2017, 162, 171–179. [Google Scholar] [CrossRef]
- Hu, Z.; Ni, J.; Cao, Y.; Liu, X. Newcastle Disease Virus as a Vaccine Vector for 20 Years: A Focus on Maternally Derived Antibody Interference. Vaccines 2020, 8, 222. [Google Scholar] [CrossRef]
- Park, M.-S.; Steel, J.; García-Sastre, A.; Swayne, D.; Palese, P. Engineered viral vaccine constructs with dual specificity: Avian influenza and Newcastle disease. Proc. Natl. Acad. Sci. USA 2006, 103, 8203–8208. [Google Scholar] [CrossRef]
- Falkenhorst, G.; Harder, T.; Remschmidt, C.; Terhardt, M.; Zepp, F.; Ledig, T.; Wicker, S.; Keller-Stanislawski, B.; Mertens, T. Background paper to the recommendation for the preferential use of live-attenuated influenza vaccine in children aged 2–6 years in Germany. Bundesgesundheitsblatt-Gesundh.-Gesundheitsschutz. 2013, 56, 1557–1564. [Google Scholar]
- Control, C.F.D. Prevention, Prevention and control of seasonal influenza with vaccines. Recommendations of the Advisory Committee on Immunization Practices—United States, 2013–2014. MMWR. Recomm. Rep. Morb. Mortal. Wkly. Rep. Recomm. Rep. 2013, 62, 1–43. [Google Scholar]
- Tisoncik, J.R.; Billharz, R.; Burmakina, S.; Belisle, S.E.; Proll, S.C.; Korth, M.J.; García-Sastre, A.; Katze, M.G. The NS1 protein of influenza A virus suppresses interferon-regulated activation of antigen-presentation and immune-proteasome pathways. J. Gen. Virol. 2011, 92, 2093. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Lobo, P.; Nogales, A.; Rodriguez, L.; Martinez-Sobrido, L. Novel Approaches for The Development of Live Attenuated Influenza Vaccines. Viruses 2019, 11, 190. [Google Scholar] [CrossRef]
- Nogales, A.; DeDiego, M.L.; Topham, D.J.; Martínez-Sobrido, L. Rearrangement of influenza virus spliced segments for the development of live-attenuated vaccines. J. Virol. 2016, 90, 6291–6302. [Google Scholar] [PubMed]
- Coleman, J.R.; Papamichail, D.; Skiena, S.; Futcher, B.; Wimmer, E.; Mueller, S. Virus attenuation by genome-scale changes in codon pair bias. Science 2008, 320, 1784–1787. [Google Scholar] [PubMed]
- Mueller, S.; Coleman, J.R.; Papamichail, D.; Ward, C.B.; Nimnual, A.; Futcher, B.; Skiena, S.; Wimmer, E. Live attenuated influenza virus vaccines by computer-aided rational design. Nat. Biotechnol. 2010, 28, 723–726. [Google Scholar]
- Yang, C.; Skiena, S.; Futcher, B.; Mueller, S.; Wimmer, E. Deliberate reduction of hemagglutinin and neuraminidase expression of influenza virus leads to an ultraprotective live vaccine in mice. Proc. Natl. Acad. Sci. USA 2013, 110, 9481–9486. [Google Scholar]
- Broadbent, A.J.; Santos, C.P.; Anafu, A.; Wimmer, E.; Mueller, S.; Subbarao, K. Evaluation of the attenuation, immunogenicity, and efficacy of a live virus vaccine generated by codon-pair bias de-optimization of the 2009 pandemic H1N1 influenza virus, in ferrets. Vaccine 2016, 34, 563–570. [Google Scholar]
- Eickhoff, C.S.; Terry, F.E.; Peng, L.; Meza, K.A.; Sakala, I.G.; Van Aartsen, D.; Moise, L.; Martin, W.D.; Schriewer, J.; Buller, R.M.; et al. Highly conserved influenza T cell epitopes induce broadly protective immunity. Vaccine 2019, 37, 5371–5381. [Google Scholar] [CrossRef]
- Li, L.; Petrovsky, N. Molecular mechanisms for enhanced DNA vaccine immunogenicity. Expert Rev. Vaccines 2016, 15, 313–329. [Google Scholar] [PubMed]
- Karlsson, I.; Borggren, M.; Nielsen, J.; Christensen, D.; Williams, J.; Fomsgaard, A. Increased humoral immunity by DNA vaccination using an α-tocopherol-based adjuvant. Hum. Vaccin. Immunother. 2017, 13, 1823–1830. [Google Scholar] [PubMed]
- Zhou, F.; Wang, G.; Buchy, P.; Cai, Z.; Chen, H.; Chen, Z.; Cheng, G.; Wan, X.-F.; Deubel, V.; Zhou, P. A triclade DNA vaccine designed on the basis of a comprehensive serologic study elicits neutralizing antibody responses against all clades and subclades of highly pathogenic avian influenza H5N1 viruses. J. Virol. 2012, 86, 6970–6978. [Google Scholar] [PubMed]
- Espeseth, A.S.; Cejas, P.J.; Citron, M.P.; Wang, D.; DiStefano, D.J.; Callahan, C.; Donnell, G.O.; Galli, J.D.; Swoyer, R.; Touch, S. Modified mRNA/lipid nanoparticle-based vaccines expressing respiratory syncytial virus F protein variants are immunogenic and protective in rodent models of RSV infection. Npj Vaccines 2020, 5, 1–14. [Google Scholar]
- Richner, J.M.; Himansu, S.; Dowd, K.A.; Butler, S.L.; Salazar, V.; Fox, J.M.; Julander, J.G.; Tang, W.W.; Shresta, S.; Pierson, T.C. Modified mRNA vaccines protect against Zika virus infection. Cell 2017, 168, 1114–1125.e10. [Google Scholar]
- Corbett, K.S.; Edwards, D.K.; Leist, S.R.; Abiona, O.M.; Boyoglu-Barnum, S.; Gillespie, R.A.; Himansu, S.; Schäfer, A.; Ziwawo, C.T.; DiPiazza, A.T. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature 2020, 586, 567–571. [Google Scholar]
- Meyer, M.; Huang, E.; Yuzhakov, O.; Ramanathan, P.; Ciaramella, G.; Bukreyev, A. Modified mRNA-based vaccines elicit robust immune responses and protect guinea pigs from Ebola virus disease. J. Infect. Dis. 2018, 217, 451–455. [Google Scholar]
- Mu, Z.; Haynes, B.F.; Cain, D.W. HIV mRNA vaccines—progress and future paths. Vaccines 2021, 9, 134. [Google Scholar]
- Qian, C.; Liu, X.; Xu, Q.; Wang, Z.; Chen, J.; Li, T.; Zheng, Q.; Yu, H.; Gu, Y.; Li, S.; et al. Recent Progress on the Versatility of Virus-Like Particles. Vaccines 2020, 8, 139. [Google Scholar] [CrossRef]
- Wu, P.; Lu, J.; Zhang, X.; Mei, M.; Feng, L.; Peng, D.; Hou, J.; Kang, S.-M.; Liu, X.; Tang, Y. Single dose of consensus hemagglutinin-based virus-like particles vaccine protects chickens against divergent H5 subtype influenza viruses. Front. Immunol. 2017, 8, 1649. [Google Scholar]
- Zhu, W.-Z.; Wen, Y.-C.; Lin, S.-Y.; Chen, T.-C.; Chen, H.-W. Anti-influenza protective efficacy of a H6 virus-like particle in chickens. Vaccines 2020, 8, 465. [Google Scholar]
- Hu, C.-M.J.; Chien, C.-Y.; Liu, M.-T.; Fang, Z.-S.; Chang, S.-Y.; Juang, R.-H.; Chang, S.-C.; Chen, H.-W. Multi-antigen avian influenza a (H7N9) virus-like particles: Particulate characterizations and immunogenicity evaluation in murine and avian models. BMC Biotechnol. 2017, 17, 1–12. [Google Scholar]
- Elaish, M.; Kang, K.I.; Xia, M.; Ali, A.; Shany, S.A.; Wang, L.; Jiang, X.; Lee, C.W. Immunogenicity and protective efficacy of the norovirus P particle-M2e chimeric vaccine in chickens. Vaccine 2015, 33, 4901–4909. [Google Scholar] [CrossRef]
- Graves, P.N.; Schulman, J.L.; Young, J.F.; Palese, P. Preparation of influenza virus subviral particles lacking the HA1 subunit of hemagglutinin: Unmasking of cross-reactive HA2 determinants. Virology 1983, 126, 106–116. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-H.; Abi-Ghanem, D.; Waghela, S.D.; Chou, W.-K.; Farnell, M.B.; Mwangi, W.; Berghman, L.R. Immunization of chickens with an agonistic monoclonal anti-chicken CD40 antibody–hapten complex: Rapid and robust IgG response induced by a single subcutaneous injection. J. Immunol. Methods 2012, 378, 116–120. [Google Scholar] [PubMed]
- Song, L.; Xiong, D.; Song, H.; Wu, L.; Zhang, M.; Kang, X.; Pan, Z.; Jiao, X. Mucosal and systemic immune responses to influenza H7N9 antigen HA1–2 co-delivered intranasally with flagellin or polyethyleneimine in mice and chickens. Front. Immunol. 2017, 8, 326. [Google Scholar] [PubMed]
- Park, H.Y.; Tan, P.S.; Kavishna, R.; Ker, A.; Lu, J.; Chan, C.E.Z.; Hanson, B.J.; MacAry, P.A.; Caminschi, I.; Shortman, K.; et al. Enhancing vaccine antibody responses by targeting Clec9A on dendritic cells. NPJ Vaccines 2017, 2, 31. [Google Scholar] [CrossRef]
- Kavishna, R.; Kang, T.Y.; Vacca, M.; Chua, B.Y.L.; Park, H.Y.; Tan, P.S.; Chow, V.T.; Lahoud, M.H.; Alonso, S. A single-shot vaccine approach for the universal influenza A vaccine candidate M2e. Proc. Natl. Acad. Sci. USA 2022, 119, e2025607119. [Google Scholar] [CrossRef]
- Lin, S.C.; Lin, Y.F.; Chong, P.; Wu, S.C. Broader neutralizing antibodies against H5N1 viruses using prime-boost immunization of hyperglycosylated hemagglutinin DNA and virus-like particles. PLoS ONE 2012, 7, e39075. [Google Scholar] [CrossRef]
- Lin, S.C.; Liu, W.C.; Jan, J.T.; Wu, S.C. Glycan masking of hemagglutinin for adenovirus vector and recombinant protein immunizations elicits broadly neutralizing antibodies against H5N1 avian influenza viruses. PLoS ONE 2014, 9, e92822. [Google Scholar] [CrossRef]
- Krammer, F.; Pica, N.; Hai, R.; Margine, I.; Palese, P. Chimeric hemagglutinin influenza virus vaccine constructs elicit broadly protective stalk-specific antibodies. J. Virol. 2013, 87, 6542–6550. [Google Scholar] [CrossRef]
- Li, M.; Wang, Y.; Sun, Y.; Cui, H.; Zhu, S.J.; Qiu, H.J. Mucosal vaccines: Strategies and challenges. Immunol. Lett. 2020, 217, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Mohan, T.; Berman, Z.; Luo, Y.; Wang, C.; Wang, S.; Compans, R.W.; Wang, B.-Z. Chimeric virus-like particles containing influenza HA antigen and GPI-CCL28 induce long-lasting mucosal immunity against H3N2 viruses. Sci. Rep. 2017, 7, 1–11. [Google Scholar]
- Lapuente, D.; Storcksdieck Genannt Bonsmann, M.; Maaske, A.; Stab, V.; Heinecke, V.; Watzstedt, K.; Heß, R.; Westendorf, A.; Bayer, W.; Ehrhardt, C. IL-1β as mucosal vaccine adjuvant: The specific induction of tissue-resident memory T cells improves the heterosubtypic immunity against influenza A viruses. Mucosal. Immunol. 2018, 11, 1265–1278. [Google Scholar] [CrossRef] [PubMed]
| Vaccine Strain Abbreviation | Commercial Vaccine | Announcement No. | Announcement Date | Notes |
|---|---|---|---|---|
| F strain | AI (H9 subtype) IV (F/98 strain) | N/A | N/A | A/Chicken/Shanghai/F/98 |
| ND and AI (H9 subtype) combined IV (La Sota strain + F strain) | N/A | N/A | ||
| HN03 strain | AI (H9 subtype) IV (HN03 strain) | 2534 | 23 May 2017 | N/A |
| HN106 strain | AI (H9 subtype) IV (HN106 strain) | 2306 | 8 October 2015 | A/chicken/Henan/01/2006 |
| ND and AI (H9 subtype) combined IV (La Sota strain + HN106 strain) | 2390 | 15 April 2016 | ||
| ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + HN106 strain) | 1489 | 26 November 2010 | ||
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + HE02 strain + HN106 strain) | 1883 | 7 January 2013 | ||
| LG1 strain | AI (H9 subtype) IV (LG1 strain) | N/A | N/A | A/Chicken/Shandong/LG1/2000 |
| ND and AI (H9 subtype) combined IV (La Sota strain + LG1 strain) | 1322 | 11 January 2010 | ||
| ND, IB, and AI (H9 subtype) triple IV (La Sota strain + M41 strain + LG1 strain) | 1507 | 30 November 2010 | ||
| NJ01 strain | AI (H9 subtype) IV (NJ01 strain) | 1938 | 6 May 2013 | A/chicken/Nanjing/01/99 |
| SD696 strain | AI (H9 subtype) IV (SD696 strain) | N/A | N/A | A/Chicken/Shandong/6/96 |
| SS strain | AI (H9 subtype) IV (SS/94 strain) | N/A | N/A | A/Chicken/Guangdong/SS/94 |
| ND and AI (H9 subtype) combined IV ((La Sota strain + SS/94 strain) | N/A | N/A | ||
| ND, IB and AI (H9 subtype) triple IV (LaSota strain + M41 strain + SS/94 strain) | 1530 | 19 January 2011 | ||
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + K-11 strain + SS/94 strain) | 2083 | 26 March 2014 | ||
| SY strain | AI IV (H9 subtype, SY strain) | N/A | N/A | A/chicken/Shaanxi/SY/97 |
| ND and AI (H9 subtype) combined IV (La Sota strain + SY strain) | 1821 | 22 August 2012 | ||
| ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + SY strain) | 1779 | 4 June 2012 | ||
| SZ strain | AI (H9 subtype) IV (SZ strain) | 2270 | 24 June 2015 | A/chicken/Shandong/SZ/2008 |
| ND and AI (H9 subtype) combined IV (La Sota strain + SZ strain) | 2506 | 14 March 2017 | ||
| ND, AI (H9 subtype) and IBD triple IV (La Sota strain + SZ strain + rVP2 protein) | 2525 | 3 May 2017 | ||
| ND, IB, AI (H9 subtype) and IBD quadruple IV (La Sota strain + M41 strain + SZ strain + rVP2 protein) | 2400 | 5 May 2016 | ||
| HL strain | ND and AI (H9 subtype) double IV (La Sota strain + HL strain) | N/A | N/A | HL strain isolated from clinical cases in Luoyang, Henan, China in 2001 |
| ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + HL strain) | N/A | N/A | ||
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + AV127 strain + HL strain) | N/A | N/A | ||
| HP strain | ND and AI (H9 subtype) combined IV (La Sota strain + HP strain) | N/A | N/A | HP strain was isolated from diseased chicken flocks in Puyang, Henan, China in 1998 |
| ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + HP strain) | 1335 | 1 February 2010 | ||
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + Z16 strain + HP strain) | N/A | N/A | ||
| JD strain | ND and AI (H9 subtype) combined IV (La Sota strain + JD strain) | 2577 | 31 August 2017 | N/A |
| WD strain | ND and AI (H9 subtype) combined IV (La Sota strain + WD strain) | 136 | 1 February 2019 | WD strain was isolated from Wangdu, Hebei, China in 1998 |
| ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + WD strain) | 1489 | 26 November 2010 | ||
| ND, IBD and AI (H9 subtype) triple IV (La Sota strain + BJQ902 strain + WD strain) | 2557 | 2 August 2017 | ||
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + HSH23 strain + WD strain) | 2268 | 17 June 2015 | ||
| G strain | ND and AI (H9 subtype) combined IV (aSG10 strain + G strain) | 164 | 16 April 2019 | A/chicken/Hebei/G/2012 |
| WJ57 strain | ND and AI (H9 subtype) combined IV (A-VII strain + WJ57 strain) | registered | 15 October 2018 | A/chicken/Jiangsu/WJ57/2012 |
| D1 strain | DP and AI (H9 subtype) double IV (AV1221 strain + D1 strain) | 2557 | 2 August 2017 | N/A |
| HZ strain | ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + HZ strain) | 1556 | 13 April 2011 | N/A |
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + HS25 strain + HZ strain) | 2106 | 28 May 2014 | ||
| JY strain | ND and AI (H9 subtype) combined IV (La Sota strain + JY strain) | 1507 | 30 November 2010 | A/chicken/Jiangsu/JY/99 |
| L strain | ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + L strain) | N/A | N/A | N/A |
| NJ02 strain | ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + NJ02 strain) | 1448 | 27 August 2010 | A/chicken/Nanjing/02/2001 |
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + AV127 strain + NJ02 strain) | 1548 | 4 March 2011 | ||
| Re9 strain (HuN33 strain) | ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + Re9 strain) | 2324 | 18 November 2015 | A/chicken/Hunan/33/2008 |
| ND, AI (H9 subtype) and ED triple IV (La Sota strain + Re9 strain + Jing 911 strain) | 164 | 16 April 2019 | ||
| YBF003 strain | ND, IB and AI (H9 subtype) triple IV (La Sota strain + M41 strain + YBF003 strain) | N/A | N/A | N/A |
| ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + NE4 strain + YBF003 strain) | 1908 | 1 March 2013 | ||
| ND, AI (H9 subtype) and IBD triple IV (La Sota strain + YBF003 strain + VP2 protein) | 1865 | 3 December 2012 | ||
| ND, IB, AI (H9 subtype) and IBD quadruple IV (La Sota strain + M41 strain + YBF003 strain + SVP-2 protein) | 1532 | 21 January 2011 | ||
| S2 strain | ND, IB, ED and AI (H9 subtype) quadruple IV (La Sota strain + M41 strain + AV-127 strain + S2 strain) | 1532 | 21 January 2011 | A/chicken/Shandong/S2/2005 |
| YT strain | ND, AI and IC triple IV (La Sota strain + YT strain + DY2 strain) | 582 | 24 July 2022 | N/A |
| ND, AI and IC triple IV (La Sota strain + YT strain + QD strain) | 463 | 29 August 2021 | ||
| YBF13 | ND, AI and IC triple IV (La Sota strain + YBF13 strain + YBAV-4 strain) | 441 | 29 June 2021 | N/A |
| Type of Vaccine | Development and Application | Notes | References |
|---|---|---|---|
| Inactivated whole-virus vaccines | Change vaccine strains | Enhanced vaccine compatibility | [61] |
| Universal vaccine | Provides cross-protection | [63,64,65,66,67] | |
| Recombinant with PR8 virus | Improved production efficiency | [68] | |
| Gene modification | Modified HA sequence | [69,70] | |
| Inactivation method | Antibody-mediated immune responses were increased in chickens that received the BPL and gamma IWVs compared to the formaldehyde IWV | [71] | |
| Development of new adjuvants | CpG oligodeoxynucleotides, bursopentine, bursin-like epitope peptide, poly I:C, bursal peptides, chitosan, Bacillus subtilis spores, polyethylenimine-coated PLGA nanoparticles | [72,73,74,75,76,77,78,79,80] | |
| Vector vaccine | Fowlpox virus | Affected by preexisting immunity | [81,82] |
| Fowl adenoviruses | Application in IBD, not yet developed in influenza | [83] | |
| Marek’s disease virus | Could not induce robust local mucosal immunity in the respiratory tract | [84,85] | |
| Newcastle-disease virus | Interference from maternal antibodies greatly hinders clinical application | [86,87,88,89] | |
| APMV-2 | Conferred complete immune protection | [90] | |
| Live-attenuated vaccine | Attenuated cold-adapted live H9N2-subtype AIV vaccine | Provides better protection, but carries a biological risk | [91,92,93] |
| Recombinant influenza virus with modified, truncated or absent NS1 | Effectively reduced viral replication | [94,95] | |
| Codon-pair bias | Effectively reduced viral replication | [96,97,98] | |
| DNA and mRNA vaccine | DNA vaccines | Conferred complete immune protection | [99] |
| mRNA vaccine | Inability to induce strong immunity and unsuitable for mass vaccination | [100] | |
| Virus-like particle vaccine | H9N2 VLP | Reduced biosecurity threats and costs | [101] |
| Universal vaccination | Provides cross protection | [102,103] | |
| Recombinant protein vaccine | conjugated to anti-chicken Dec205 monoclonal antibody | Strong immune protection 14 days after initial immunization | [104] |
| H9 HA1-fliC | Promotes superior protective immune responses | [105] | |
| Chickens vaccinated with CD83 scFv targeted H9 HA | Effectively reduced viral replication | [106] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, J.; Zhou, Y.; Pu, J.; Liu, L. Status and Challenges for Vaccination against Avian H9N2 Influenza Virus in China. Life 2022, 12, 1326. https://doi.org/10.3390/life12091326
Dong J, Zhou Y, Pu J, Liu L. Status and Challenges for Vaccination against Avian H9N2 Influenza Virus in China. Life. 2022; 12(9):1326. https://doi.org/10.3390/life12091326
Chicago/Turabian StyleDong, Jinze, Yong Zhou, Juan Pu, and Litao Liu. 2022. "Status and Challenges for Vaccination against Avian H9N2 Influenza Virus in China" Life 12, no. 9: 1326. https://doi.org/10.3390/life12091326
APA StyleDong, J., Zhou, Y., Pu, J., & Liu, L. (2022). Status and Challenges for Vaccination against Avian H9N2 Influenza Virus in China. Life, 12(9), 1326. https://doi.org/10.3390/life12091326






