Identification of Potential Genes Encoding Protein Transporters in Arabidopsis thaliana Glucosinolate (GSL) Metabolism
Abstract
:1. Introduction
2. Results
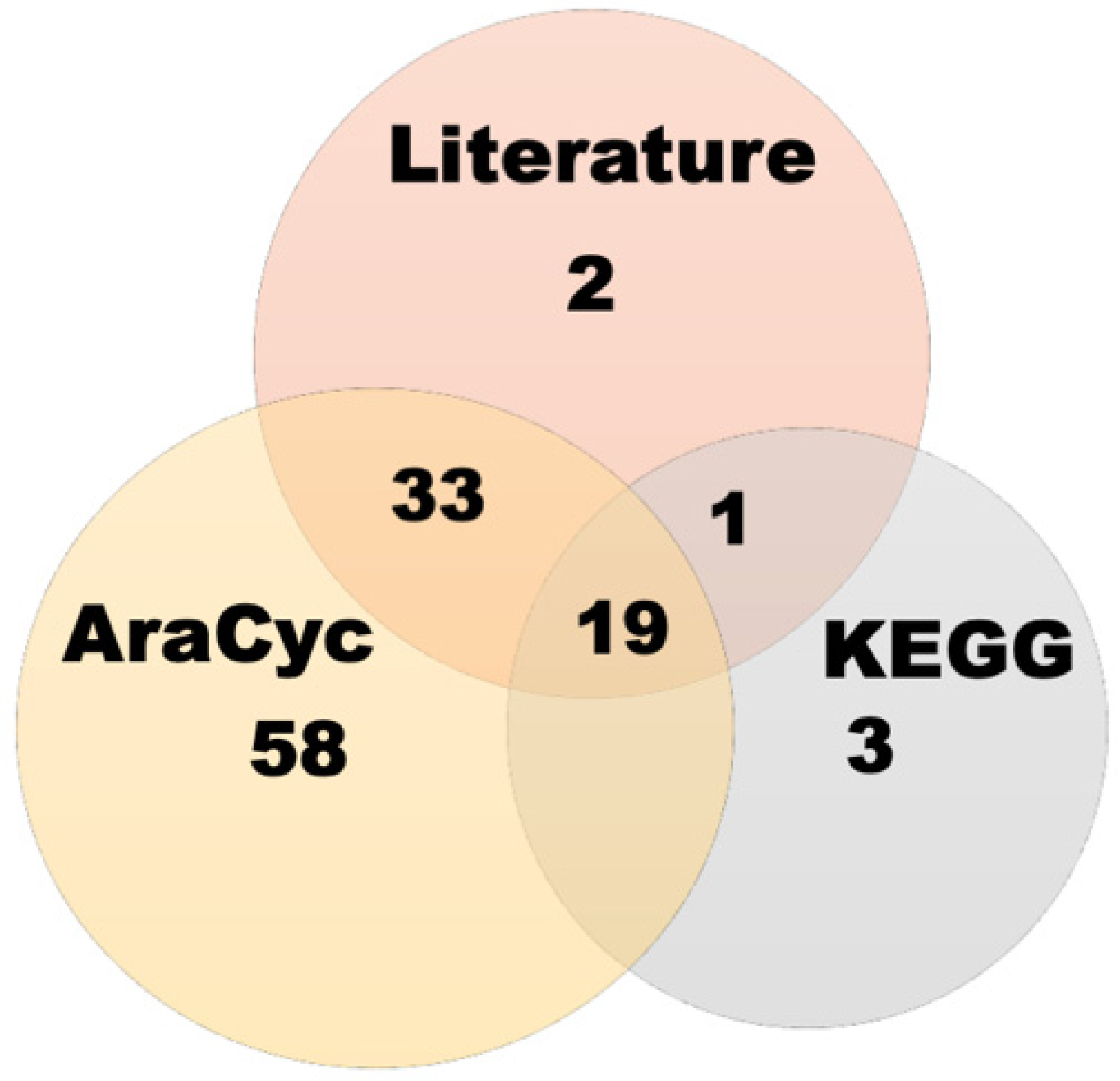
2.1. Data Collection and Establishment
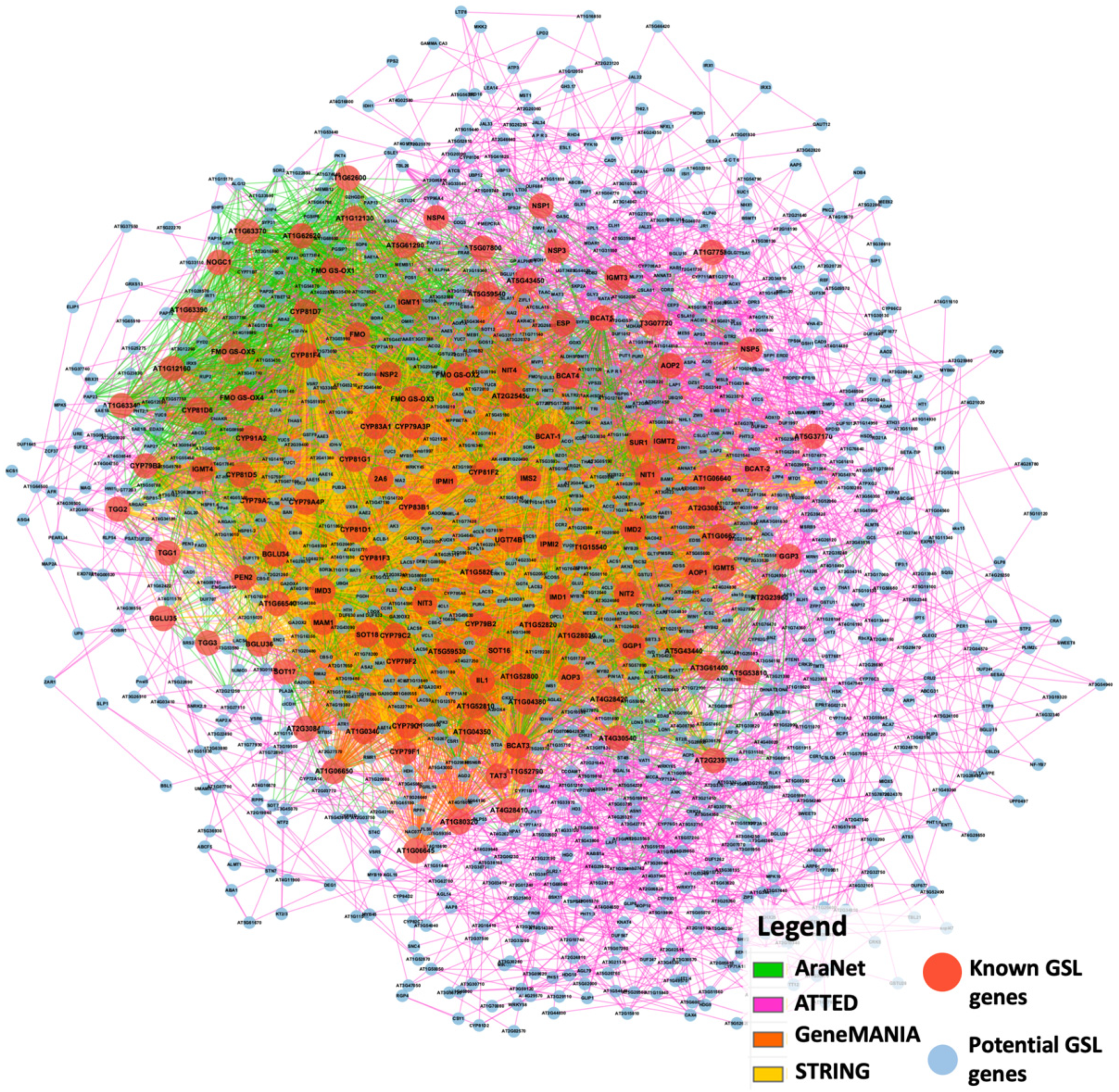
2.2. Gene Co-Expression Network Analysis
2.3. GO Enrichment
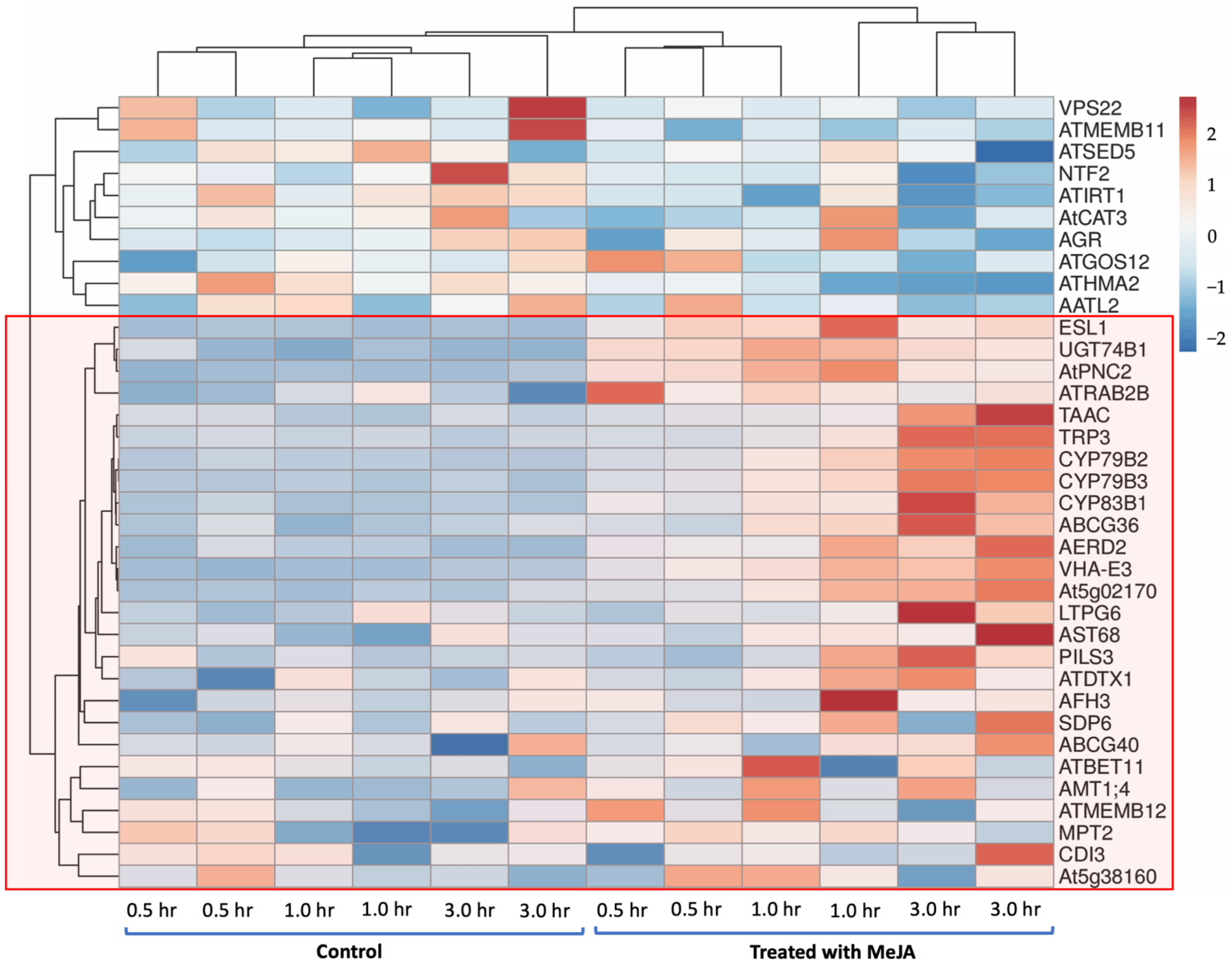
2.4. Gene-Expression Pattern Analysis and Visualisation
2.5. Sequence Analysis of Potential GSL TPs
2.6. Evolutionary Relationship Analysis of the Potential GSL TPs
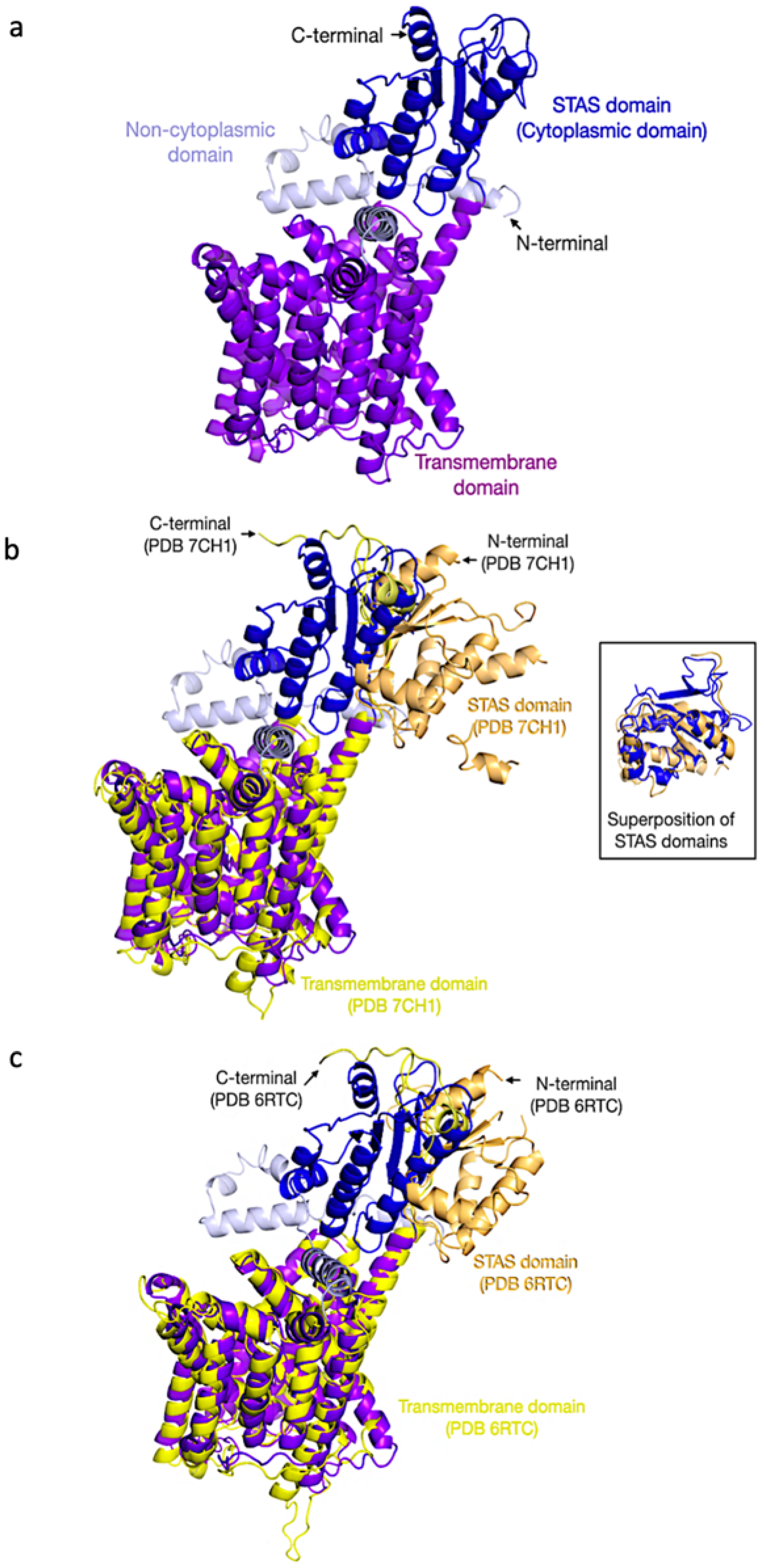
2.7. Protein Structure Prediction and Model Evaluation
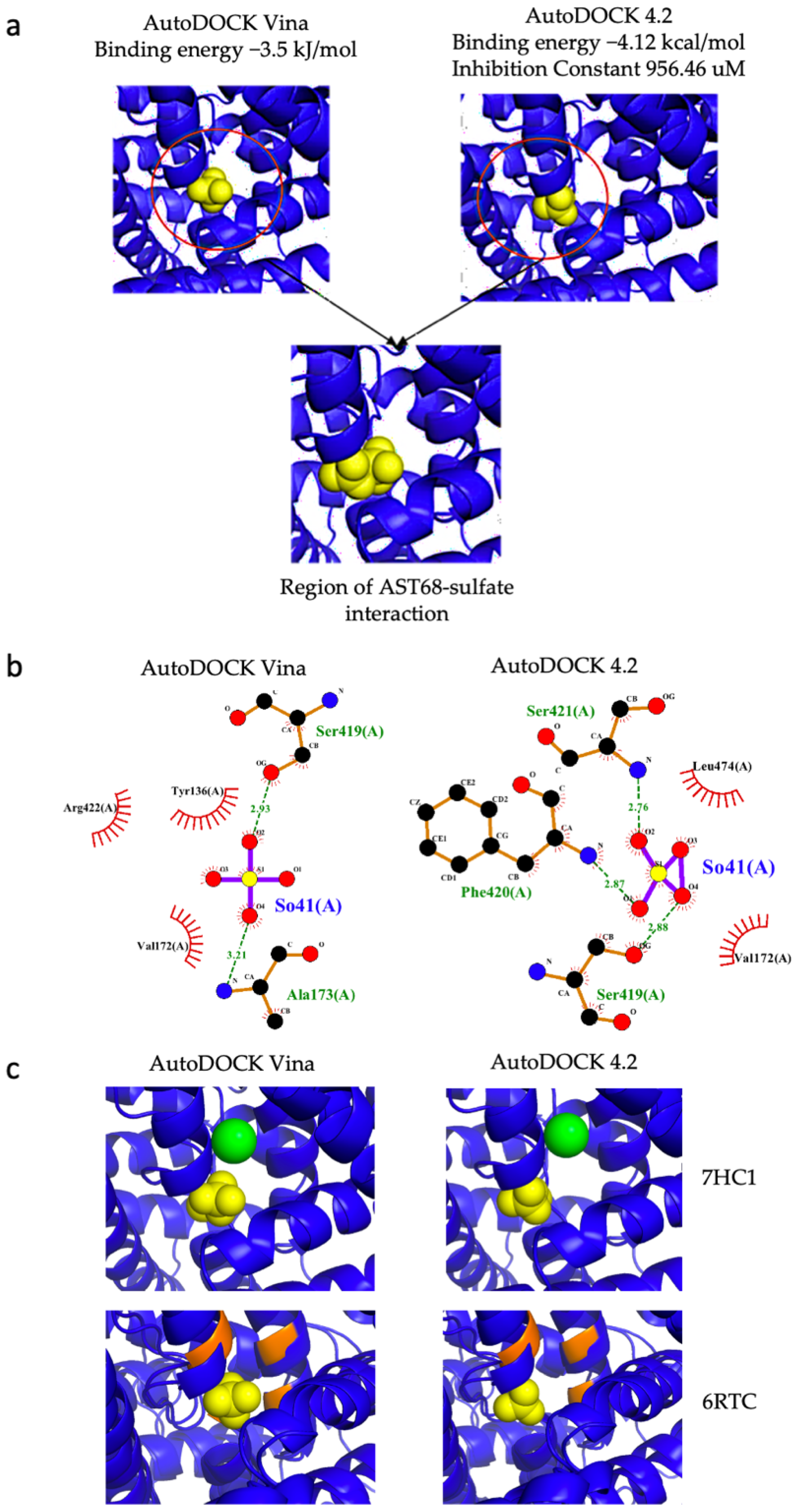
2.8. Molecular Docking of AST68 with Sulphate Ion
3. Discussion
4. Materials and Methods
4.1. Data Collection and Construction of the Gene-Co-Expression Network
4.2. GO Enrichment Analysis
4.3. Expression-Pattern Analysis
4.4. Characterisation of Potential GSL TPs
4.5. Sequence Analysis of GSL TPs
4.6. Construction of Phylogenetic Tree
4.7. Protein-Structure Prediction and Model Evaluations
4.8. Molecular Docking of Potential GSL Transporters
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Manghwar, H.; Hussain, A.; Ullah, A.; Gul, S.; Shaban, M.; Khan, A.H.; Ali, M.; Sani, S.G.A.S.; Chaudhary, H.J.; Munis, M.F.H. Expression analysis of defense related genes in wheat and maize against Bipolaris sorokiniana. Physiol. Mol. Plant Pathol. 2018, 103, 36–46. [Google Scholar] [CrossRef]
- Manghwar, H.; Hussain, A. Mechanism of tobacco osmotin gene in plant responses to biotic and abiotic stress tolerance: A brief history. Biocell 2022, 46, 623–632. [Google Scholar] [CrossRef]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52, 39. [Google Scholar] [CrossRef] [Green Version]
- Chhajed, S.; Misra, B.B.; Tello, N.; Chen, S. Chemodiversity of the Glucosinolate-Myrosinase System at the Single Cell Type Resolution. Front. Plant Sci. 2019, 10, 618. [Google Scholar] [CrossRef] [PubMed]
- Clay, N.K.; Adio, A.M.; Denoux, C.; Jander, G.; Ausubel, F.M. Glucosinolate Metabolites Required for an Arabidopsis Innate Immune Response. Science 2009, 323, 95–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fahey, J.W.; Zalcmann, A.T.; Talalay, P. The chemical diversity and distribution of glucosinolates and isothiocyanates among plants. Phytochemistry 2001, 56, 5–51. [Google Scholar] [CrossRef]
- Reichelt, M.; Brown, P.D.; Schneider, B.; Oldham, N.; Stauber, E.; Tokuhisa, J.; Kliebenstein, D.; Mitchell-Olds, T.; Gershenzon, J. Benzoic acid glucosinolate esters and other glucosinolates from Arabidopsis thaliana. Phytochemistry 2002, 59, 663–671. [Google Scholar] [CrossRef]
- Ishida, M.; Hara, M.; Fukino, N.; Kakizaki, T.; Morimitsu, Y. Glucosinolate metabolism, functionality and breeding for the improvement of Brassicaceae vegetables. Breed. Sci. 2014, 64, 48–59. [Google Scholar] [CrossRef] [Green Version]
- Falk, K.L.; Tokuhisa, J.; Gershenzon, J. The Effect of Sulfur Nutrition on Plant Glucosinolate Content: Physiology and Molecular Mechanisms. Plant Biol. 2007, 9, 573–581. [Google Scholar] [CrossRef]
- Barba, F.J.; Nikmaram, N.; Roohinejad, S.; Khelfa, A.; Zhu, Z.; Koubaa, M. Bioavailability of Glucosinolates and Their Breakdown Products: Impact of Processing. Front. Nutr. 2016, 3, 24. [Google Scholar] [CrossRef] [Green Version]
- Seo, M.-S.; Kim, J.S. Understanding of MYB Transcription Factors Involved in Glucosinolate Biosynthesis in Brassicaceae. Molecules 2017, 22, 1549. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harun, S.; Abdullah-Zawawi, M.-R.; Goh, H.-H.; Mohamed-Hussein, Z.-A. A Comprehensive Gene Inventory for Glucosinolate Biosynthetic Pathway in Arabidopsis thaliana. J. Agric. Food Chem. 2020, 68, 7281–7297. [Google Scholar] [CrossRef] [PubMed]
- Blažević, I.; Montaut, S.; Burčul, F.; Olsen, C.E.; Burow, M.; Rollin, P.; Agerbirk, N. Glucosinolate structural diversity, identification, chemical synthesis and metabolism in plants. Phytochemistry 2019, 169, 112100. [Google Scholar] [CrossRef]
- Halkier, B.A.; Gershenzon, J. Biology and Biochemistry of Glucosinolates. Annu. Rev. Plant Biol. 2006, 57, 303–333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Y.; Rossi, M.; Liang, X.; Zhang, H.; Zou, L.; Ong, C.N. An Integrated Metabolomics Study of Glucosinolate Metabolism in Different Brassicaceae Genera. Metabolites 2020, 10, 313. [Google Scholar] [CrossRef]
- Wittstock, U.; Meier, K.; Dörr, F.; Ravindran, B.M. NSP-Dependent Simple Nitrile Formation Dominates upon Breakdown of Major Aliphatic Glucosinolates in Roots, Seeds, and Seedlings of Arabidopsis thaliana Columbia-0. Front. Plant Sci. 2016, 7, 1821. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chhajed, S.; Mostafa, I.; He, Y.; Abou-Hashem, M.; El-Domiaty, M.; Chen, S. Glucosinolate Biosynthesis and the Glucosinolate–Myrosinase System in Plant Defense. Agronomy 2020, 10, 1786. [Google Scholar] [CrossRef]
- Winde, I.; Wittstock, U. Insect herbivore counteradaptations to the plant glucosinolate–myrosinase system. Phytochemistry 2011, 72, 1566–1575. [Google Scholar] [CrossRef]
- Koroleva, O.A.; Davies, A.; Deeken, R.; Thorpe, M.R.; Tomos, A.D.; Hedrich, R. Identification of a New Glucosinolate-Rich Cell Type in Arabidopsis Flower Stalk. Plant Physiol. 2000, 124, 599–608. [Google Scholar] [CrossRef] [Green Version]
- Koroleva, O.A.; Cramer, R. Single-cell proteomic analysis of glucosinolate-rich S-cells in Arabidopsis thaliana. Methods 2011, 54, 413–423. [Google Scholar] [CrossRef]
- Kissen, R.; Bones, A.M. Nitrile-specifier Proteins Involved in Glucosinolate Hydrolysis in Arabidopsis thaliana. J. Biol. Chem. 2009, 284, 12057–12070. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petersen, B.L.; Chen, S.; Hansen, C.H.; Olsen, C.E.; Halkier, B. Composition and content of glucosinolates in developing Arabidopsis thaliana. Planta 2002, 214, 562–571. [Google Scholar] [CrossRef] [PubMed]
- Magrath, R.; Mithen, R. Maternal Effects on the Expression of Individual Aliphatic Glucosinolates in Seeds and Seedlings of Brassica napus. Plant Breed. 1993, 111, 249–252. [Google Scholar] [CrossRef]
- van Dam, N.M.; Tytgat, T.O.G.; Kirkegaard, J.A. Root and shoot glucosinolates: A comparison of their diversity, function and interactions in natural and managed ecosystems. Phytochem. Rev. 2008, 8, 171–186. [Google Scholar] [CrossRef] [Green Version]
- Tsunoda, T.; Grosser, K.; Van Dam, N.M. Locally and systemically induced glucosinolates follow optimal defence allocation theory upon root herbivory. Funct. Ecol. 2018, 32, 2127–2137. [Google Scholar] [CrossRef]
- Meldau, S.; Erb, M.; Baldwin, I.T. Defence on demand: Mechanisms behind optimal defence patterns. Ann. Bot. 2012, 110, 1503–1514. [Google Scholar] [CrossRef] [Green Version]
- Touw, A.J.; Mogena, A.V.; Maedicke, A.; Sontowski, R.; van Dam, N.M.; Tsunoda, T. Both Biosynthesis and Transport Are Involved in Glucosinolate Accumulation During Root-Herbivory in Brassica rapa. Front. Plant Sci. 2020, 10, 1653. [Google Scholar] [CrossRef] [Green Version]
- Tsunoda, T.; Krosse, S.; Van Dam, N.M. Root and shoot glucosinolate allocation patterns follow optimal defence allocation theory. J. Ecol. 2017, 105, 1256–1266. [Google Scholar] [CrossRef]
- Hunziker, P.; Halkier, B.A.; Schulz, A. Arabidopsis glucosinolate storage cells transform into phloem fibres at late stages of development. J. Exp. Bot. 2019, 70, 4305–4317. [Google Scholar] [CrossRef]
- Koroleva, O.A.; Gibson, T.M.; Cramer, R.; Stain, C. Glucosinolate-accumulating S-cells in Arabidopsis leaves and flower stalks undergo programmed cell death at early stages of differentiation. Plant J. 2010, 64, 456–469. [Google Scholar] [CrossRef]
- Borpatragohain, P.; Rose, T.; King, G.J. Fire and Brimstone: Molecular Interactions between Sulfur and Glucosinolate Biosynthesis in Model and Crop Brassicaceae. Front. Plant Sci. 2016, 7, 1735. [Google Scholar] [CrossRef] [Green Version]
- David, R.; Byrt, C.S.; Tyerman, S.D.; Gilliham, M.; Wege, S. Roles of membrane transporters: Connecting the dots from sequence to phenotype. Ann. Bot. 2019, 124, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Nour-Eldin, H.H.; Andersen, T.; Burow, M.; Madsen, S.R.; Jørgensen, M.E.; Olsen, C.E.; Dreyer, I.; Hedrich, R.; Geiger, D.; Halkier, B. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 2012, 488, 531–534. [Google Scholar] [CrossRef]
- Andersen, T.G.; Nour-Eldin, H.H.; Fuller, V.L.; Olsen, C.E.; Burow, M.; Halkier, B.A. Integration of Biosynthesis and Long-Distance Transport Establish Organ-Specific Glucosinolate Profiles in Vegetative Arabidopsis. Plant Cell 2013, 25, 3133–3145. [Google Scholar] [CrossRef] [Green Version]
- Madsen, S.R.; Olsen, C.E.; Nour-Eldin, H.H.; Halkier, B.A. Elucidating the Role of Transport Processes in Leaf Glucosinolate Distribution. Plant Physiol. 2014, 166, 1450–1462. [Google Scholar] [CrossRef] [PubMed]
- Moussaieff, A.; Rogachev, I.; Brodsky, L.; Malitsky, S.; Toal, T.W.; Belcher, H.; Yativ, M.; Brady, S.M.; Benfey, P.N.; Aharoni, A. High-resolution metabolic mapping of cell types in plant roots. Proc. Natl. Acad. Sci. USA 2013, 110, E1232–E1241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andersen, T.G.; Halkier, B.A. Upon bolting the GTR1 and GTR2 transporters mediate transport of glucosinolates to the inflorescence rather than roots. Plant Signal. Behav. 2014, 9, e27740. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madsen, S.R.; Kunert, G.; Reichelt, M.; Gershenzon, J.; Halkier, B.A. Feeding on Leaves of the Glucosinolate Transporter Mutant gtr1gtr2 Reduces Fitness of Myzus persicae. J. Chem. Ecol. 2015, 41, 975–984. [Google Scholar] [CrossRef] [Green Version]
- Koprivova, A.; Kopriva, S. Molecular mechanisms of regulation of sulfate assimilation: First steps on a long road. Front. Plant Sci. 2014, 5, 589. [Google Scholar] [CrossRef] [Green Version]
- Morikawa-Ichinose, T.; Kim, S.-J.; Allahham, A.; Kawaguchi, R.; Maruyama-Nakashita, A. Glucosinolate Distribution in the Aerial Parts of sel1-10, a Disruption Mutant of the Sulfate Transporter SULTR1;2, in Mature Arabidopsis thaliana Plants. Plants 2019, 8, 95. [Google Scholar] [CrossRef] [Green Version]
- Hirai, M.Y.; Fujiwara, T.; Awazuhara, M.; Kimura, T.; Noji, M.; Saito, K. Global expression profiling of sulfur-starved Arabidopsis by DNA macroarray reveals the role of O -acetyl-l -serine as a general regulator of gene expression in response to sulfur nutrition. Plant J. 2003, 33, 651–663. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nikiforova, V.; Freitag, J.; Kempa, S.; Adamik, M.; Hesse, H.; Hoefgen, R. Transcriptome analysis of sulfur depletion inArabidopsis thaliana: Interlacing of biosynthetic pathways provides response specificity. Plant J. 2003, 33, 633–650. [Google Scholar] [CrossRef]
- Aarabi, F.; Kusajima, M.; Tohge, T.; Konishi, T.; Gigolashvili, T.; Takamune, M.; Sasazaki, Y.; Watanabe, M.; Nakashita, H.; Fernie, A.R.; et al. Sulfur deficiency–induced repressor proteins optimize glucosinolate biosynthesis in plants. Sci. Adv. 2016, 2, e1601087. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maruyama-Nakashita, A.; Nakamura, Y.; Tohge, T.; Saito, K.; Takahashi, H. Arabidopsis SLIM1 Is a Central Transcriptional Regulator of Plant Sulfur Response and Metabolism. Plant Cell 2006, 18, 3235–3251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gigolashvili, T.; Yatusevich, R.; Rollwitz, I.; Humphry, M.; Gershenzon, J.; Flügge, U.-I. The Plastidic Bile Acid Transporter 5 Is Required for the Biosynthesis of Methionine-Derived Glucosinolates inArabidopsis thaliana. Plant Cell 2009, 21, 1813–1829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sawada, Y.; Toyooka, K.; Kuwahara, A.; Sakata, A.; Nagano, M.; Saito, K.; Hirai, M.Y. Arabidopsis Bile Acid:Sodium Symporter Family Protein 5 is Involved in Methionine-Derived Glucosinolate Biosynthesis. Plant Cell Physiol. 2009, 50, 1579–1586. [Google Scholar] [CrossRef] [Green Version]
- Pushparaj, P.N. Introduction to Functional Bioinformatics; Springer: Berlin/Heidelberg, Germany, 2019; pp. 235–254. [Google Scholar] [CrossRef]
- Ayaz, A.; Saqib, S.; Huang, H.; Zaman, W.; Lü, S.; Zhao, H. Genome-wide comparative analysis of long-chain acyl-CoA synthetases (LACSs) gene family: A focus on identification, evolution and expression profiling related to lipid synthesis. Plant Physiol. Biochem. 2021, 161, 1–11. [Google Scholar] [CrossRef]
- Ayaz, A.; Huang, H.; Zheng, M.; Zaman, W.; Li, D.; Saqib, S.; Zhao, H.; Lü, S. Molecular Cloning and Functional Analysis of GmLACS2-3 Reveals Its Involvement in Cutin and Suberin Biosynthesis along with Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 9175. [Google Scholar] [CrossRef]
- Ashari, K.-S.; Abdullah-Zawawi, M.-R.; Harun, S.; Mohamed-Hussein, Z.-A. Reconstruction of the Transcriptional Regulatory Network in Arabidopsis thaliana Aliphatic Glucosinolate Biosynthetic Pathway. Sains Malays. 2018, 47, 2993–3002. [Google Scholar] [CrossRef]
- Hirai, M.; Sugiyama, K.; Sawada, Y.; Tohge, T.; Obayashi, T.; Suzuki, A.; Araki, R.; Sakurai, N.; Suzuki, H.; Aoki, K.; et al. Omics-based identification of Arabidopsis Myb transcription factors regulating aliphatic glucosinolate biosynthesis. Proc. Natl. Acad. Sci. USA 2007, 104, 6478–6483. [Google Scholar] [CrossRef] [Green Version]
- Gigolashvili, T.; Berger, B.; Mock, H.-P.; Müller, C.; Weisshaar, B.; Flügge, U.-I. The transcription factor HIG1/MYB51 regulates indolic glucosinolate biosynthesis in Arabidopsis thaliana. Plant J. 2007, 50, 886–901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gigolashvili, T.; Yatusevich, R.; Berger, B.; Müller, C.; Flügge, U.-I. The R2R3-MYB transcription factor HAG1/MYB28 is a regulator of methionine-derived glucosinolate biosynthesis inArabidopsis thaliana. Plant J. 2007, 51, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Sønderby, I.E.; Hansen, B.G.; Bjarnholt, N.; Ticconi, C.; Halkier, B.A.; Kliebenstein, D.J. A Systems Biology Approach Identifies a R2R3 MYB Gene Subfamily with Distinct and Overlapping Functions in Regulation of Aliphatic Glucosinolates. PLoS ONE 2007, 2, e1322. [Google Scholar] [CrossRef]
- Harun, S.; Rohani, E.R.; Ohme-Takagi, M.; Goh, H.-H.; Mohamed-Hussein, Z.-A. ADAP is a possible negative regulator of glucosinolate biosynthesis in Arabidopsis thaliana based on clustering and gene expression analyses. J. Plant Res. 2021, 134, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Harun, S.; Afiqah-Aleng, N.; Karim, M.B.; Amin, A.U.; Kanaya, S.; Mohamed-Hussein, Z.-A. Potential Arabidopsis thaliana glucosinolate genes identified from the co-expression modules using graph clustering approach. PeerJ 2021, 9, e11876. [Google Scholar] [CrossRef]
- Knill, T.; Schuster, J.; Reichelt, M.; Gershenzon, J.; Binder, S. Arabidopsis Branched-Chain Aminotransferase 3 Functions in Both Amino Acid and Glucosinolate Biosynthesis. Plant Physiol. 2007, 146, 1028–1039. [Google Scholar] [CrossRef] [Green Version]
- Sawada, Y.; Kuwahara, A.; Nagano, M.; Narisawa, T.; Sakata, A.; Saito, K.; Hirai, M.Y. Omics-Based Approaches to Methionine Side Chain Elongation in Arabidopsis: Characterization of the Genes Encoding Methylthioalkylmalate Isomerase and Methylthioalkylmalate Dehydrogenase. Plant Cell Physiol. 2009, 50, 1181–1190. [Google Scholar] [CrossRef] [Green Version]
- Geu-Flores, F.; Nielsen, M.T.; Nafisi, M.; Møldrup, M.E.; Olsen, C.E.; Motawia, M.S.; Halkier, B. Glucosinolate engineering identifies a γ-glutamyl peptidase. Nat. Chem. Biol. 2009, 5, 575–577. [Google Scholar] [CrossRef]
- Harun, S.; Abdullah-Zawawi, M.-R.; A-Rahman, M.R.A.; Muhammad, N.A.N.; Mohamed-Hussein, Z.-A. SuCComBase: A manually curated repository of plant sulfur-containing compounds. Database 2019, 2019, baz021. [Google Scholar] [CrossRef]
- Aoki, Y.; Okamura, Y.; Tadaka, S.; Kinoshita, K.; Obayashi, T. ATTED-II in 2016: A Plant Coexpression Database Towards Lineage-Specific Coexpression. Plant Cell Physiol. 2015, 57, e5. [Google Scholar] [CrossRef]
- Lee, T.; Yang, S.; Kim, E.; Ko, Y.; Hwang, S.; Shin, J.; Shim, J.E.; Shim, H.; Kim, H.; Kim, C.; et al. AraNet v2: An improved database of co-functional gene networks for the study of Arabidopsis thaliana and 27 other nonmodel plant species. Nucleic Acids Res. 2014, 43, D996–D1002. [Google Scholar] [CrossRef] [PubMed]
- Montojo, J.; Zuberi, K.; Rodriguez, H.; Bader, G.D.; Morris, Q. GeneMANIA: Fast gene network construction and function prediction for Cytoscape. F1000Research 2014, 3, 153. [Google Scholar] [CrossRef] [PubMed]
- Warde-Farley, D.; Donaldson, S.L.; Comes, O.; Zuberi, K.; Badrawi, R.; Chao, P.; Franz, M.; Grouios, C.; Kazi, F.; Lopes, C.T.; et al. The GeneMANIA prediction server: Biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010, 38, W214–W220. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P.; et al. STRING v10: Protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015, 43, D447–D452. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Metsalu, T.; Vilo, J. ClustVis: A web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef]
- Larsen, B.; Xu, D.; Halkier, B.A.; Nour-Eldin, H.H. Advances in methods for identification and characterization of plant transporter function. J. Exp. Bot. 2017, 68, 4045–4056. [Google Scholar] [CrossRef]
- Takano, J.; Wada, M.; Ludewig, U.; Schaaf, G.; von Wirén, N.; Fujiwara, T. The Arabidopsis Major Intrinsic Protein NIP5;1 Is Essential for Efficient Boron Uptake and Plant Development under Boron Limitation. Plant Cell 2006, 18, 1498–1509. [Google Scholar] [CrossRef] [Green Version]
- Payne, R.; Xu, D.; Foureau, E.; Carqueijeiro, I.; Oudin, A.; De Bernonville, T.D.; Novak, V.; Burow, M.; Olsen, C.-E.; Jones, M.; et al. An NPF transporter exports a central monoterpene indole alkaloid intermediate from the vacuole. Nat. Plants 2017, 3, 16208. [Google Scholar] [CrossRef]
- De Geyter, N.; Gholami, A.; Goormachtig, S.; Goossens, A. Transcriptional machineries in jasmonate-elicited plant secondary metabolism. Trends Plant Sci. 2012, 17, 349–359. [Google Scholar] [CrossRef]
- Kang, J.; Park, J.; Choi, H.; Burla, B.; Kretzschmar, T.; Lee, Y.; Martinoia, E. Plant ABC Transporters. Arab. Book 2011, 9, e0153. [Google Scholar] [CrossRef] [Green Version]
- Biemans-Oldehinkel, E.; Doeven, M.K.; Poolman, B. ABC transporter architecture and regulatory roles of accessory domains. FEBS Lett. 2005, 580, 1023–1035. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gräfe, K.; Schmitt, L. The ABC transporter G subfamily in Arabidopsis thaliana. J. Exp. Bot. 2020, 72, 92–106. [Google Scholar] [CrossRef]
- Lu, X.; Dittgen, J.; Piślewska-Bednarek, M.; Molina, A.; Schneider, B.; Svatoš, A.; Doubský, J.; Schneeberger, K.; Weigel, D.; Bednarek, P.; et al. Mutant Allele-Specific Uncoupling of PENETRATION3 Functions Reveals Engagement of the ATP-Binding Cassette Transporter in Distinct Tryptophan Metabolic Pathways. Plant Physiol. 2015, 168, 814–827. [Google Scholar] [CrossRef]
- Bednarek, P.; Piślewska-Bednarek, M.; Svatoš, A.; Schneider, B.; Doubský, J.; Mansurova, M.; Humphry, M.; Consonni, C.; Panstruga, R.; Sanchez-Vallet, A.; et al. A Glucosinolate Metabolism Pathway in Living Plant Cells Mediates Broad-Spectrum Antifungal Defense. Science 2009, 323, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Chi, X.; Jin, X.; Chen, Y.; Lu, X.; Tu, X.; Li, X.; Zhang, Y.; Lei, J.; Huang, J.; Huang, Z.; et al. Structural insights into the gating mechanism of human SLC26A9 mediated by its C-terminal sequence. Cell Discov. 2020, 6, 55. [Google Scholar] [CrossRef]
- Walter, J.D.; Sawicka, M.; Dutzler, R. Cryo-EM structures and functional characterization of murine Slc26a9 reveal mechanism of uncoupled chloride transport. eLife 2019, 8, e46986. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Sato, Y.; Furumichi, M.; Morishima, K.; Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 2019, 47, D590–D595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogata, H.; Goto, S.; Sato, K.; Fujibuchi, W.; Bono, H.; Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 1999, 27, 29–34. [Google Scholar] [CrossRef] [Green Version]
- Mueller, L.A.; Zhang, P.; Rhee, S.Y. AraCyc: A Biochemical Pathway Database for Arabidopsis. Plant Physiol. 2003, 132, 453–460. [Google Scholar] [CrossRef] [Green Version]
- Maere, S.; Heymans, K.; Kuiper, M. BiNGO: A Cytoscape plugin to assess overrepresentation of Gene Ontology categories in Biological Networks. Bioinformatics 2005, 21, 3448–3449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benjamini, Y.; Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 2001, 29, 1165–1188. [Google Scholar] [CrossRef]
- Toufighi, K.; Brady, S.M.; Austin, R.; Ly, E.; Provart, N.J. The Botany Array Resource: E-Northerns, Expression Angling, and promoter analyses. Plant J. 2005, 43, 153–163. [Google Scholar] [CrossRef]
- Bateman, A.; Martin, M.J.; Orchard, S.; Magrane, M.; Agivetova, R.; Ahmad, S.; Alpi, E.; Bowler-Barnett, E.H.; Britto, R.; Bursteinas, B.; et al. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D480–D489. [Google Scholar] [CrossRef]
- Hooper, C.M.; Castleden, I.R.; Tanz, S.; Aryamanesh, N.; Millar, A.H. SUBA4: The interactive data analysis centre for Arabidopsis subcellular protein locations. Nucleic Acids Res. 2016, 45, D1064–D1074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Stecher, G.; Tamura, K.; Kumar, S. Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Mol. Biol. Evol. 2020, 37, 1237–1239. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.; Anishchenko, I.; Park, H.; Peng, Z.; Ovchinnikov, S.; Baker, D. Improved protein structure prediction using predicted interresidue orientations. Proc. Natl. Acad. Sci. USA 2020, 117, 1496–1503. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; McPartlon, M.; Li, J. Improved protein structure prediction by deep learning irrespective of co-evolution information. Nat. Mach. Intell. 2021, 3, 601–609. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.J.; Headd, J.J.; Moriarty, N.W.; Prisant, M.G.; Videau, L.L.; Deis, L.N.; Verma, V.; Keedy, D.A.; Hintze, B.J.; Chen, V.B.; et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 2018, 27, 293–315. [Google Scholar] [CrossRef]
- McGuffin, L.J.; Aldowsari, F.M.F.; Alharbi, S.M.A.; Adiyaman, R. ModFOLD8: Accurate global and local quality estimates for 3D protein models. Nucleic Acids Res. 2021, 49, W425–W430. [Google Scholar] [CrossRef]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem in 2021: New data content and improved web interfaces. Nucleic Acids Res. 2021, 49, D1388–D1395. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [Green Version]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [Green Version]


| Potential GSL TP | Server | MolProbity Score | Percentage of Residues That Fall Inside Ramachandran-Favoured Regions (%) | Clashscore, All Atoms |
|---|---|---|---|---|
| AST68 | I-TASSER | 3.29 | 80.15 | 15.6 |
| Robetta server (trRosetta) | 1.38 | 96.89 | 4.13 | |
| Raptor-X | 5.00 | 94.67 | 488.12 | |
| ABCG40 (506–1415) | I-TASSER | 3.60 | 65.94 | 18.74 |
| Robetta server (trRosetta) | 1.37 | 97.49 | 5.04 | |
| Raptor-X | 4.93 | 95.31 | 448.78 |
| Known GSL TP (TAIR ID/UniProt ID) | Potential GSL TP (TAIR ID/UniProt ID) | Localisation | Possible Substrate |
|---|---|---|---|
| SULTR1;1 (At4g08620/Q9SAY1) and SULTR1;2 (At1g78000/Q9MAX3) | AST68 (At5g10180/O04722) | Plasma membrane; sulphate transporter 2;1 | Sulphate |
| ABCG36/PEN3 (At1g59870/Q9XIE2) | ABCG40 (At1g15520/Q9M9E1) | Plasma membrane; ABC transporter G family member 40 | 4OH-X |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harun, S.; Afiqah-Aleng, N.; Abdul Hadi, F.I.; Lam, S.D.; Mohamed-Hussein, Z.-A. Identification of Potential Genes Encoding Protein Transporters in Arabidopsis thaliana Glucosinolate (GSL) Metabolism. Life 2022, 12, 326. https://doi.org/10.3390/life12030326
Harun S, Afiqah-Aleng N, Abdul Hadi FI, Lam SD, Mohamed-Hussein Z-A. Identification of Potential Genes Encoding Protein Transporters in Arabidopsis thaliana Glucosinolate (GSL) Metabolism. Life. 2022; 12(3):326. https://doi.org/10.3390/life12030326
Chicago/Turabian StyleHarun, Sarahani, Nor Afiqah-Aleng, Fatin Izzati Abdul Hadi, Su Datt Lam, and Zeti-Azura Mohamed-Hussein. 2022. "Identification of Potential Genes Encoding Protein Transporters in Arabidopsis thaliana Glucosinolate (GSL) Metabolism" Life 12, no. 3: 326. https://doi.org/10.3390/life12030326
APA StyleHarun, S., Afiqah-Aleng, N., Abdul Hadi, F. I., Lam, S. D., & Mohamed-Hussein, Z.-A. (2022). Identification of Potential Genes Encoding Protein Transporters in Arabidopsis thaliana Glucosinolate (GSL) Metabolism. Life, 12(3), 326. https://doi.org/10.3390/life12030326







