Sleep Apnea Detection Based on Multi-Scale Residual Network
Abstract
:1. Introduction
2. Materials and Methods
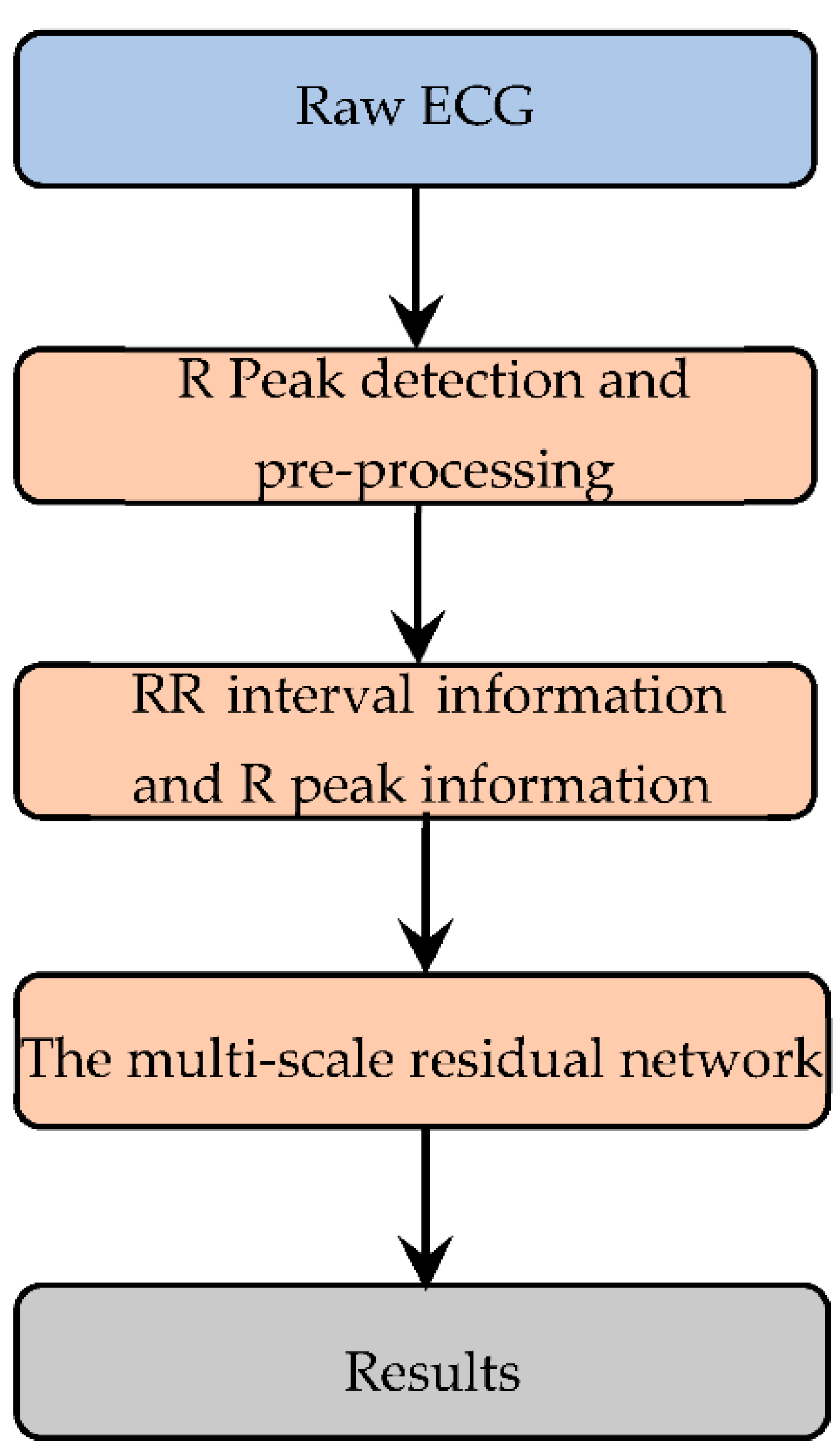
2.1. Flow Diagram of the Work
2.2. Experimental Data
2.3. Signal Denoising
- Baseline wandering—It is mainly caused by the low-frequency interference signals caused by poor contact of the measuring electrode or the patient’s breathing [21]. The frequency is between 0.05 Hz and 2 Hz, indicating that the ECG signals deviate from the normal baseline position.
- Power line interference—It is mainly 50 Hz/60 Hz noise generated by the power system, which will cause the entire waveform to be ambiguous and have a greater impact on the waveform.
- Electromyography noise—It is mainly caused by muscle fibrillation and contraction. The amplitude is small and the frequency is high [22]. The frequency is between 5 Hz and 2000 Hz, presenting an irregular and rapidly changing waveform.
2.4. R Peak Location and Signal Extraction
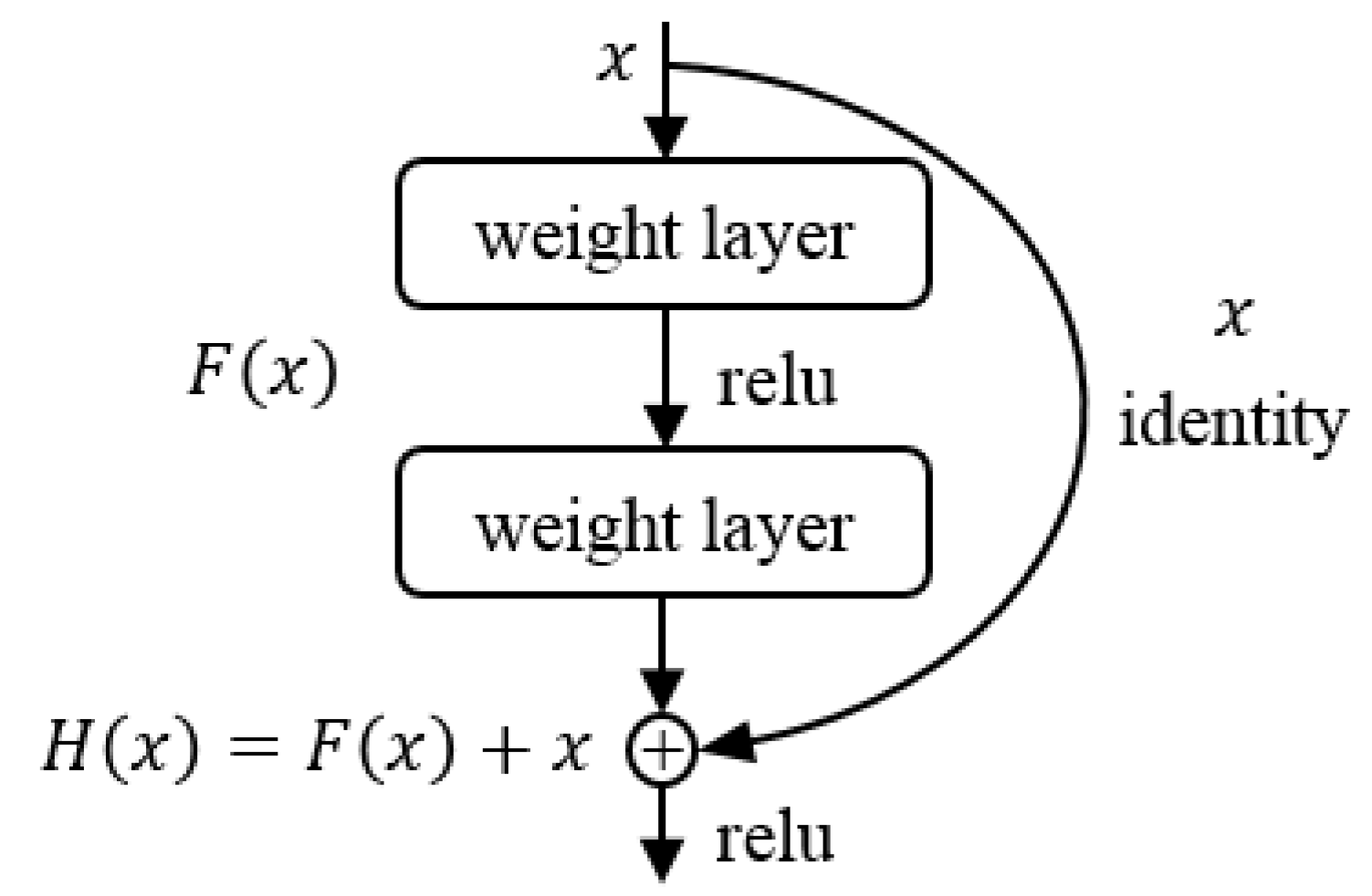
2.5. Residual Network
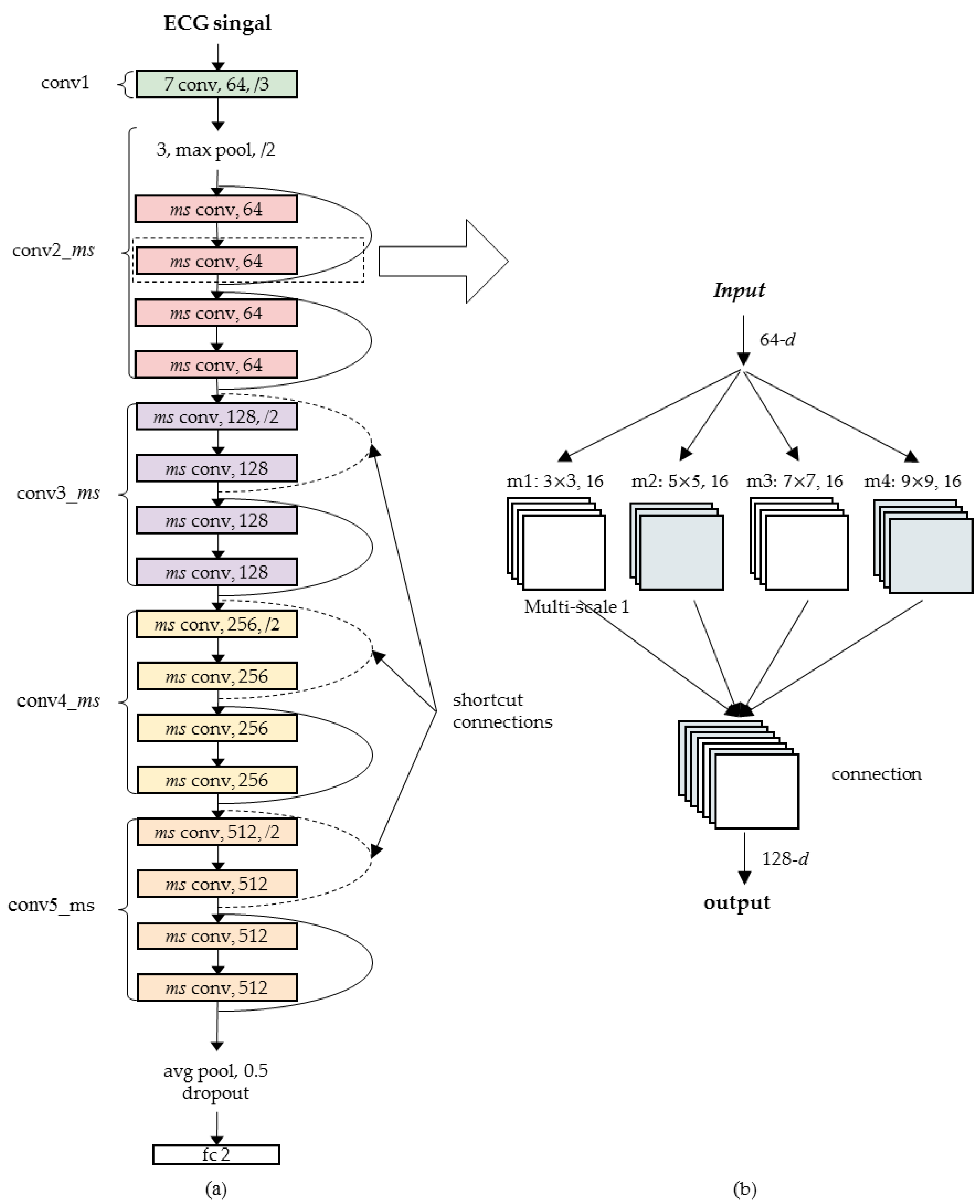
2.6. Construction of Multi-Scale Residual Network Model
2.7. Data Imbalance Processing
3. Experiment and Result Analysis
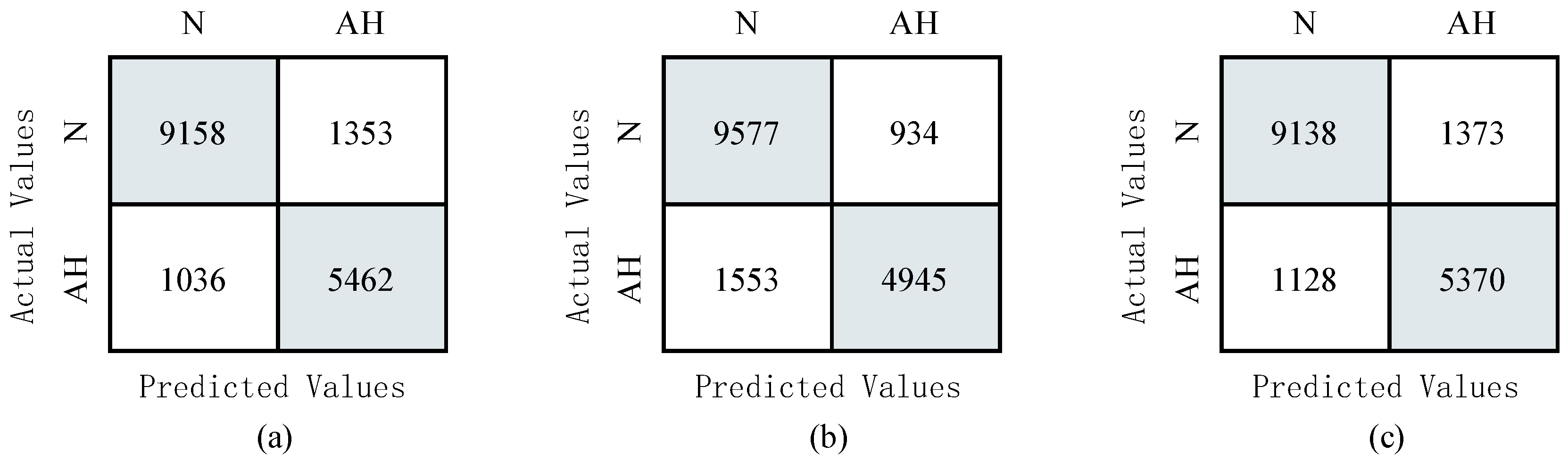
3.1. Sleep Apnea Detection Experiment
3.2. Per-Recording Classification
3.3. Test the Model on the UCD Database
3.4. Comparison of Similar Research Results
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sharma, M.; Patel, V.; Acharya, U.R. Automated identification of insomnia using optimal bi-orthogonal wavelet transform technique with single-channel EEG signals. Knowl.-Based Syst. 2021, 224, 107078–107083. [Google Scholar] [CrossRef]
- Sharma, M.; Patel, V.; Tiwari, J.; Acharya, U. Automated Characterization of Cyclic Alternating Pattern Using Wavelet-Based Features and Ensemble Learning Techniques with EEG Signals. Diagnostics 2021, 11, 1380. [Google Scholar] [CrossRef]
- Aurora, R.N.; Crainiceanu, C.; Gottlieb, D.J.; Kim, J.S.; Punjabi, N.M. Obstructive Sleep Apnea during REM Sleep and Cardiovascular Disease. Am. J. Respir. Crit. Care Med. 2018, 197, 653–660. [Google Scholar] [CrossRef]
- Butler, M.P.; Emch, J.T.; Rueschman, M.; Sands, S.A.; Shea, S.A.; Wellman, A.; Redline, S. Apnea–Hypopnea Event Duration Predicts Mortality in Men and Women in the Sleep Heart Health Study. Am. J. Respir. Crit. Care Med. 2018, 199, 903–912. [Google Scholar] [CrossRef]
- Benjafield, A.V.; Ayas, N.T.; Eastwood, P.R.; Heinzer, R.; Ip, M.S.M.; Morrell, M.J.; Nunez, C.M.; Patel, S.R.; Penzel, T.; Pépin, J.-L.; et al. Estimation of the global prevalence and burden of obstructive sleep apnoea: A literature-based analysis. Lancet Respir. Med. 2019, 7, 687–698. [Google Scholar] [CrossRef] [Green Version]
- Mostafa, S.S.; Morgado-Dias, F.; Ravelo-García, A.G. Comparison of SFS and mRMR for oximetry feature selection in obstructive sleep apnea detection. Neural Comput. Appl. 2018, 32, 15711–15731. [Google Scholar] [CrossRef]
- Romero-Ortuno, R.; O’Shea, D. Fitness and frailty: Opposite ends of a challenging continuum! Will the end of age discrimination make frailty assessments an imperative? Age Ageing 2013, 42, 279–280. [Google Scholar] [CrossRef] [Green Version]
- Jezzini, A.; Ayache, M.; Ibrahim, Z.; Elkhansa, L. ECG Classification for sleep apnea detection. In Proceedings of the Third International Conference on Advances in Biomedical Engineering (ICABME15), Beirut, Lebanon, 16–18 September 2015. [Google Scholar]
- Peppard, P.E.; Young, T.; Barnet, J.H.; Palta, M.; Hagen, E.W.; Hla, K.M. Increased Prevalence of Sleep-Disordered Breathing in Adults. Am. J. Epidemiol. 2013, 177, 1006–1014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, H.; Sharma, K.K. An algorithm for sleep apnea detection from single-lead ECG using Hermite Basis functions. Comput. Biol. Med. 2016, 77, 116–124. [Google Scholar] [CrossRef]
- Miao, K.; Ji, K.; Leng, X.; Xing, X.; Zou, H. Synthetic Aperture Radar Target Recognition with Feature Fusion Based on a Stacked Autoencoder. Sensors 2017, 17, 192. [Google Scholar]
- Ap, A.; Np, A.; Bmcsa, C.; Kb, B.; Ng, D. Towards an accurate sleep apnea detection based on ECG signal: The quintessential of a wise feature selection. Appl. Soft Comput. 2019, 83, 105568. [Google Scholar]
- Viswabhargav, C.; Tripathy, R.; Acharya, U.R. Automated detection of sleep apnea using sparse residual entropy features with various dictionaries extracted from heart rate and EDR signals. Comput. Biol. Med. 2019, 108, 20–30. [Google Scholar] [CrossRef]
- Feng, K.; Qin, H.; Wu, S.; Pan, W.; Liu, G. A Sleep Apnea Detection Method Based on Unsupervised Feature Learning and Single-Lead Electrocardiogram. IEEE Trans. Instrum. Meas. 2021, 70, 1–12. [Google Scholar] [CrossRef]
- Sharma, M.; Agarwal, S.; Acharya, U.R. Application of new class of antisymmetric wavelet filter banks for obstructive sleep apnea diagnosis using ECG signals. Comput. Biol. Med. 2018, 100, 100–113. [Google Scholar] [CrossRef]
- Li, K.; Pan, W.; Li, Y.; Jiang, Q.; Liu, G. A method to detect sleep apnea based on deep neural network and hidden Markov model using single-lead ECG signal. Neurocomputing 2018, 294, 94–101. [Google Scholar] [CrossRef]
- Urtnasan, E.; Park, J.-U.; Joo, E.Y.; Lee, K.-J. Automated Detection of Obstructive Sleep Apnea Events from a Single-Lead Electrocardiogram Using a Convolutional Neural Network. J. Med. Syst. 2018, 42, 104. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, X.; Song, C. An Automatic Screening Approach for Obstructive Sleep Apnea Diagnosis Based on Single-Lead Electrocardiogram. IEEE Trans. Autom. Sci. Eng. 2015, 12, 106–115. [Google Scholar] [CrossRef]
- Sharma, M.; Raval, M.; Acharya, U.R. A new approach to identify obstructive sleep apnea using an optimal orthogonal wavelet filter bank with ECG signals. Inform. Med. Unlocked 2019, 16, 100170. [Google Scholar] [CrossRef]
- Sharma, M.; Kumbhani, D.; Yadav, A.; Acharya, U.R. Automated Sleep apnea detection using optimal duration-frequency concentrated wavelet-based features of pulse oximetry signals. Appl. Intell. 2021, 1–13. [Google Scholar] [CrossRef]
- Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-Time Patient-Specific ECG Classification by 1-D Convolutional Neural Networks. IEEE Trans. Biomed. Eng. 2016, 63, 664–675. [Google Scholar] [CrossRef]
- Yea-Hoon, K.; Sae-Byuk, S.; Shin-Dug, K. Electroencephalography Based Fusion Two-Dimensional (2D)-Convolution Neural Networks (CNN) Model for Emotion Recognition System. Sensors 2018, 18, 1383. [Google Scholar]
- Pan, J.; Tompkins, W.J. A Real-Time QRS Detection Algorithm. IEEE Trans. Biomed. Eng. 2007, BME-32, 230–236. [Google Scholar] [CrossRef]
- Hamilton, P. Open Source ECG Analysis. Comput. IEEE Cardiol. 2002, 29, 101–104. [Google Scholar]
- Parhi, K.K.; Ayinala, M. Low-Complexity Welch Power Spectral Density Computation. IEEE Trans. Circuits Syst. I Regul. Pap. 2014, 61, 172–182. [Google Scholar] [CrossRef]
- Neunhoeffer, M.; Sternberg, S. How Cross-Validation Can Go Wrong and What to Do About It. Political Anal. 2019, 27, 101–106. [Google Scholar] [CrossRef] [Green Version]
- Cokelaer, T.; Hasch, J. ‘Spectrum‘: Spectral Analysis in Python. J. Open Source Softw. 2017, 2, 348. [Google Scholar] [CrossRef]
- Song, C.; Liu, K.; Zhang, X.; Chen, L.; Xian, X. An Obstructive Sleep Apnea Detection Approach Using a Discriminative Hidden Markov Model From ECG Signals. IEEE Trans. Biomed. Eng. 2016, 63, 1532–1542. [Google Scholar] [CrossRef]
- Zhang, Y.; Sun, Y.; Phillips, P.; Liu, G.; Zhou, X.; Wang, S. A Multilayer Perceptron Based Smart Pathological Brain Detection System by Fractional Fourier Entropy. J. Med. Syst. 2016, 40, 1–11. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016. [Google Scholar]
- Lin, T.Y.; Goyal, P.; Girshick, R.; He, K.; Dollár, P. Focal loss for dense object detection. In Proceedings of the 2017 IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2999–3007. [Google Scholar]
- Martín-González, S.; Navarro-Mesa, J.L.; Juliá-Serdá, G.; Kraemer, J.F.; Wessel, N.; Ravelo-García, A.G. Heart rate variability feature selection in the presence of sleep apnea: An expert system for the characterization and detection of the disorder. Comput. Biol. Med. 2017, 91, 47–58. [Google Scholar] [CrossRef]
- Surrel, G.; Aminifar, A.; Rincon, F.; Murali, S.; Atienza, D. Online Obstructive Sleep Apnea Detection on Medical Wearable Sensors. IEEE Trans. Biomed. Circuits Syst. 2018, 12, 762–773. [Google Scholar] [CrossRef]
- Chang, H.-Y.; Yeh, C.-Y.; Lee, C.-T.; Lin, C.-C. A Sleep Apnea Detection System Based on a One-Dimensional Deep Convolution Neural Network Model Using Single-Lead Electrocardiogram. Sensors 2020, 20, 4157. [Google Scholar] [CrossRef]
- Sinam, A.; Majumder, S. A novel approach osa detection using single-lead ecg scalogram based on deep neural network. J. Mech. Med. Biol. 2019, 19, 1950026. [Google Scholar]

| Layer | Output Size | Network Architecture |
|---|---|---|
| conv1 | 100 × 1 | Convolutional layer: 7 × 1, 64, Stride: 3 |
| conv2_ms | 50 × 1 | Pooling layer: 3 × 1, Stride: 2 |
| conv3_ms | 25 × 1 | |
| conv4_ms | 13 × 1 | |
| conv5_ms | 7 × 1 | |
| 1 × 1 | Dropout: 0.5, | |
| Computing power | 0.144 × 109 | |
| Forecast Result | Accuracy/% | Sensitivity/% | Specificity/% | ||||
|---|---|---|---|---|---|---|---|
| N | AH | Total | |||||
| Realitylabel | N | 9158 | 1353 | 10,511 | 86.0 | 84.1 | 87.1 |
| AH | 1036 | 5462 | 6498 | ||||
| Total | 10,194 | 6815 | 17,009 | ||||
| Method | Accuracy/% | Sensitivity/% | Specificity/% | AUC% | F1-Score/% |
|---|---|---|---|---|---|
| ResNet | 84.6 | 82.2 | 86.1 | 0.918 | 80.3 |
| ResNet + Multiscale | 86.0 | 84.1 | 87.1 | 0.931 | 82.1 |
| Method | Accuracy/% | Sensitivity/% | Specificity/% | AUC | Corr/% |
|---|---|---|---|---|---|
| ResNet | 91.2 | 100 | 75 | 0.985 | 0.945 |
| ResNet + Multiscale | 97.1 | 100 | 91.7 | 1 | 0.956 |
| Method | Accuracy/% | Sensitivity/% | Specificity/% |
|---|---|---|---|
| ResNet | 67.1 | 35.5 | 72.2 |
| ResNet + Multiscale | 72.4 | 36.5 | 83.6 |
| Work | Method | Accuracy/% | Sensitivity/% | Specificity/% |
|---|---|---|---|---|
| Sharma and Sharma | LS-SVM | 83.4 | 79.5 | 88.4 |
| Pinho et al. | ANN/SVM | 82.1 | 88.4 | 72.3 |
| Viswabhargav et al. | SVM | 78.1 | 78.0 | 78.1 |
| Surrel et al. | LS-SVM | 82.2 | 73.3 | 87.6 |
| Li et al. | DNN + HMM | 84.7 | 88.9 | 82.1 |
| Feng et al. | TDCS | 85.1 | 86.2 | 84.4 |
| Martin-Gonzalez et al. | LDA + QDA + LR | 84.8 | 81.5 | 86.8 |
| Chang et al. | 1D CNN | 87.9 | 81.1 | 92.0 |
| Singh et al. | CNN + Decision Fusion | 86.2 | 90.0 | 83.8 |
| Our method | ResNet + Multiscale | 86.0 | 84.1 | 87.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fang, H.; Lu, C.; Hong, F.; Jiang, W.; Wang, T. Sleep Apnea Detection Based on Multi-Scale Residual Network. Life 2022, 12, 119. https://doi.org/10.3390/life12010119
Fang H, Lu C, Hong F, Jiang W, Wang T. Sleep Apnea Detection Based on Multi-Scale Residual Network. Life. 2022; 12(1):119. https://doi.org/10.3390/life12010119
Chicago/Turabian StyleFang, Hengyang, Changhua Lu, Feng Hong, Weiwei Jiang, and Tao Wang. 2022. "Sleep Apnea Detection Based on Multi-Scale Residual Network" Life 12, no. 1: 119. https://doi.org/10.3390/life12010119
APA StyleFang, H., Lu, C., Hong, F., Jiang, W., & Wang, T. (2022). Sleep Apnea Detection Based on Multi-Scale Residual Network. Life, 12(1), 119. https://doi.org/10.3390/life12010119







