Aptamers, Riboswitches, and Ribozymes in S. cerevisiae Synthetic Biology
Abstract
1. Introduction
2. Ligand-Responsive Aptamers in S. cerevisiae
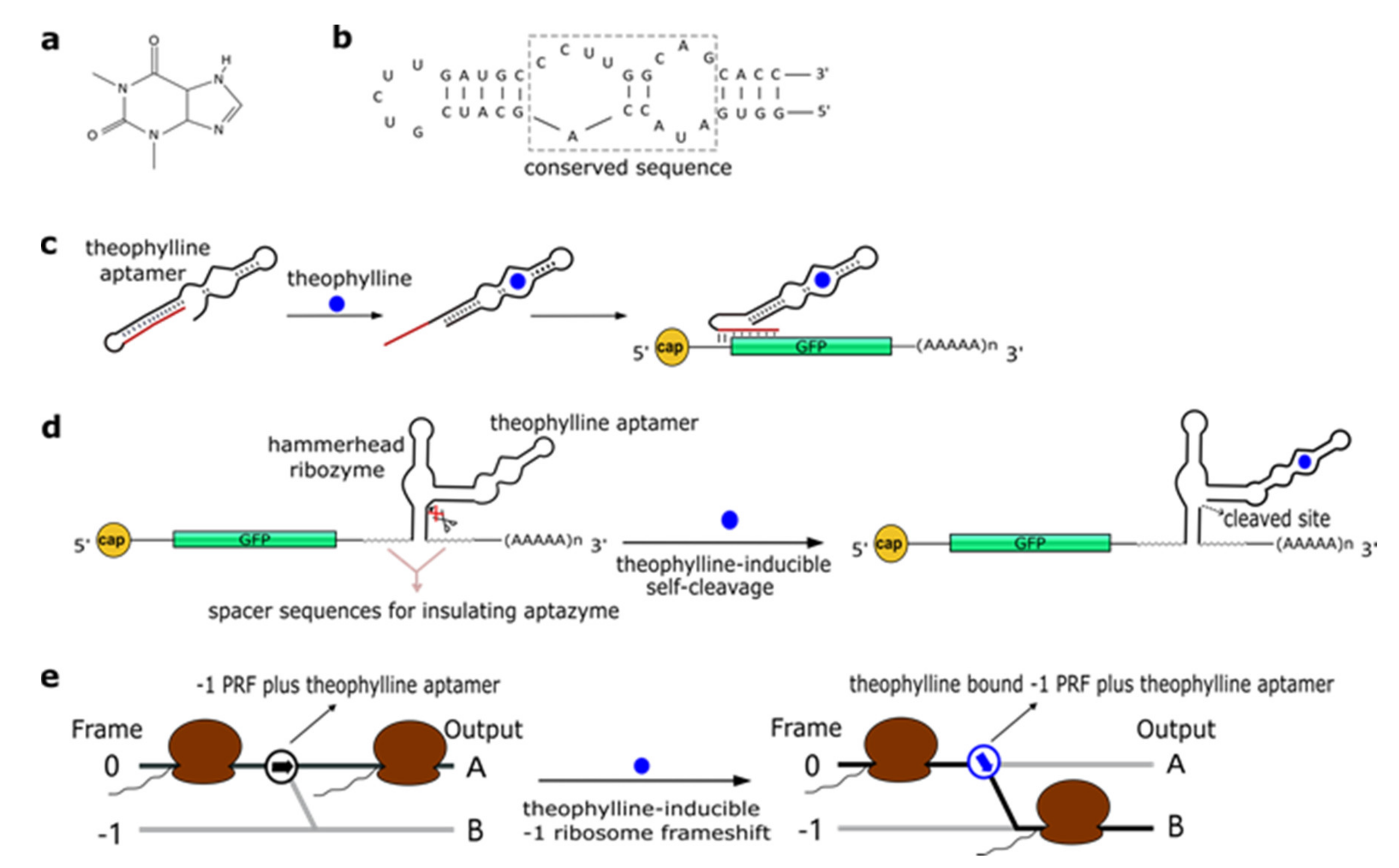
2.1. Theophylline Aptamers
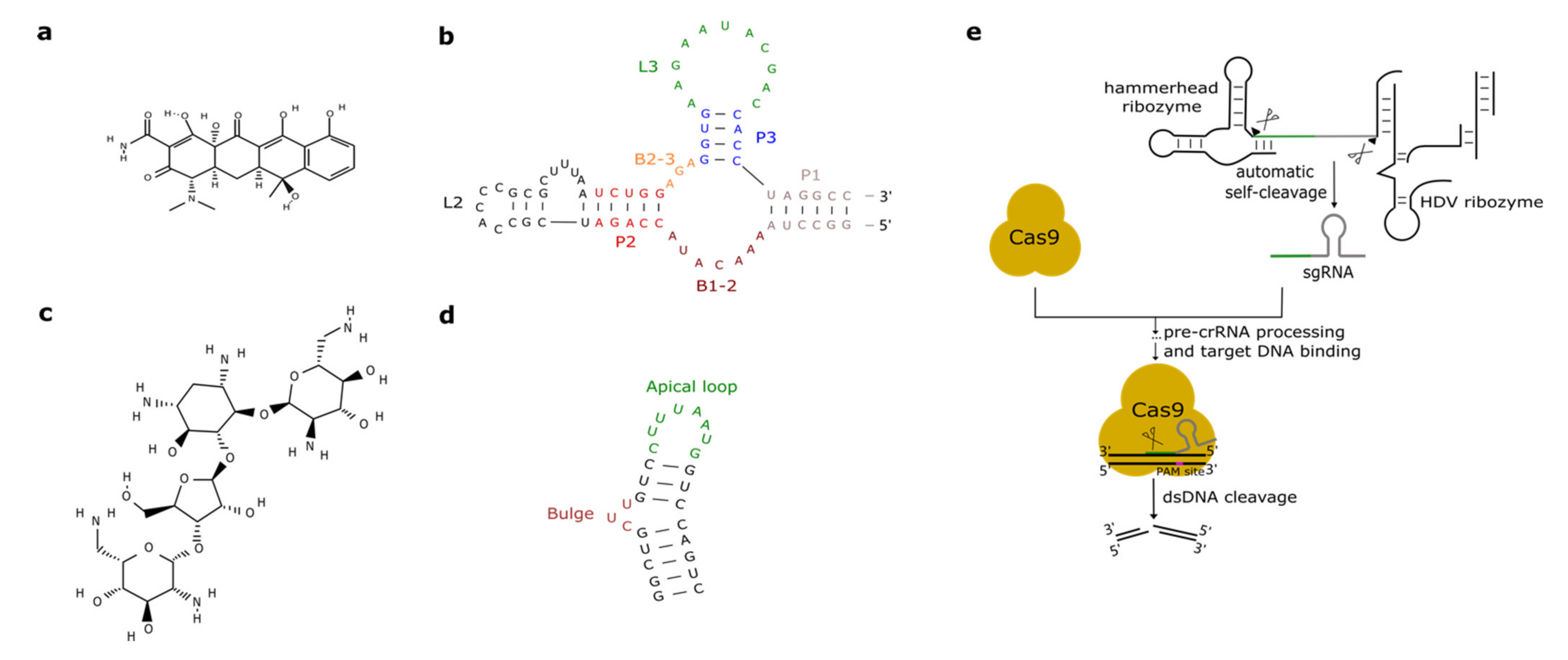
2.2. Tetracycline Aptamers
2.3. Aminoglycosides-Responsive Aptamers
2.4. Other Aptamers Used in S. cerevisiae
3. CRISPR-Cas, Ribozymes, and Hairpins
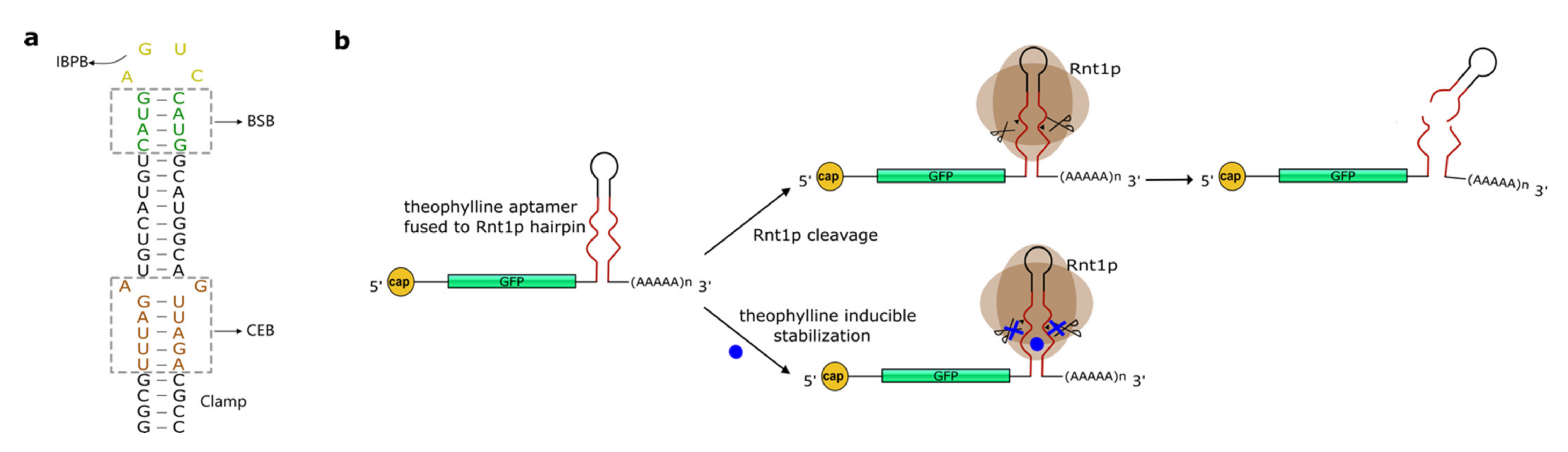
4. Hairpins Cleaved by RNase III
5. Others RNA Structures
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Benenson, Y. Synthetic biology with RNA: Progress report. Curr. Opin. Chem. Biol. 2012, 16, 278–284. [Google Scholar] [CrossRef]
- Marchisio, M.A.; Stelling, J. Automatic design of digital synthetic gene circuits. PLoS Comput. Biol. 2011, 7, e1001083. [Google Scholar] [CrossRef] [PubMed]
- Serganov, A.; Nudler, E. A decade of riboswitches. Cell 2013, 152, 17–24. [Google Scholar] [CrossRef]
- Mironov, A.S.; Gusarov, I.; Rafikov, R.; Lopez, L.E.; Shatalin, K.; Kreneva, R.A.; Perumov, D.A.; Nudler, E. Sensing small molecules by nascent RNA: A mechanism to control transcription in bacteria. Cell 2002, 111, 747–756. [Google Scholar] [CrossRef]
- Nahvi, A.; Sudarsan, N.; Ebert, M.S.; Zou, X.; Brown, K.L.; Breaker, R.R. Genetic control by a metabolite binding mRNA. Chem. Biol. 2002, 9, 1043–1049. [Google Scholar] [CrossRef]
- Winkler, W.; Nahvi, A.; Breaker, R.R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 2002, 419, 952–956. [Google Scholar] [CrossRef]
- Schroeder, K.T.; Daldrop, P.; Lilley, D.M.J. RNA Tertiary Interactions in a Riboswitch Stabilize the Structure of a Kink Turn. Structure 2011, 19, 1233–1240. [Google Scholar] [CrossRef] [PubMed]
- Hollands, K.; Proshkin, S.; Sklyarova, S.; Epshtein, V.; Mironov, A.; Nudler, E.; Groisman, E.A. Riboswitch control of Rho-dependent transcription termination. Proc. Natl. Acad. Sci. USA 2012, 109, 5376–5381. [Google Scholar] [CrossRef]
- Breaker, R.R. Riboswitches and Translation Control. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.S.; Gusti, V.; Pillai, S.G.; Gaur, R.K. An artificial riboswitch for controlling pre-mRNA splicing. RNA 2005, 11, 1667–1677. [Google Scholar] [CrossRef]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Hermann, T.; Patel, D.J. Biochemistry—Adaptive recognition by nucleic acid aptamers. Science 2000, 287, 820–825. [Google Scholar] [CrossRef]
- Kruger, K.; Grabowski, P.J.; Zaug, A.J.; Sands, J.; Gottschling, D.E.; Cech, T.R. Self-splicing RNA: Autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 1982, 31, 147–157. [Google Scholar] [CrossRef]
- Baskerville, S.; Bartel, D.P. A ribozyme that ligates RNA to protein. Proc. Natl. Acad. Sci. USA 2002, 99, 9154–9159. [Google Scholar] [CrossRef] [PubMed]
- Doudna, J.A.; Cech, T.R. The chemical repertoire of natural ribozymes. Nature 2002, 418, 222–228. [Google Scholar] [CrossRef] [PubMed]
- Serganov, A.; Keiper, S.; Malinina, L.; Tereshko, V.; Skripkin, E.; Hobartner, C.; Polonskaia, A.; Phan, A.T.; Wombacher, R.; Micura, R.; et al. Structural basis for Diels-Alder ribozyme-catalyzed carbon-carbon bond formation. Nat. Struct. Mol. Biol. 2005, 12, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Park, S.V.; Yang, J.S.; Jo, H.; Kang, B.; Oh, S.S.; Jung, G.Y. Catalytic RNA, ribozyme, and its applications in synthetic biology. Biotechnol. Adv. 2019, 37, 107452. [Google Scholar] [CrossRef]
- O’Rourke, S.M.; Scott, W.G. Structural Simplicity and Mechanistic Complexity in the Hammerhead Ribozyme. Prog. Mol. Biol. Transl. Sci. 2018, 159, 177–202. [Google Scholar] [CrossRef]
- Ren, A.; Micura, R.; Patel, D.J. Structure-based mechanistic insights into catalysis by small self-cleaving ribozymes. Curr. Opin. Chem. Biol. 2017, 41, 71–83. [Google Scholar] [CrossRef]
- Lacroix-Labonte, J.; Girard, N.; Dagenais, P.; Legault, P. Rational engineering of the Neurospora VS ribozyme to allow substrate recognition via different kissing-loop interactions. Nucleic Acids Res. 2016, 44, 6924–6934. [Google Scholar] [CrossRef]
- Been, M.D.; Wickham, G.S. Self-cleaving ribozymes of hepatitis delta virus RNA. Eur. J. Biochem. 1997, 247, 741–753. [Google Scholar] [CrossRef]
- Savinov, A.; Block, S.M. Self-cleavage of the glmS ribozyme core is controlled by a fragile folding element. Proc. Natl. Acad. Sci. USA 2018, 115, 11976–11981. [Google Scholar] [CrossRef]
- Roth, A.; Weinberg, Z.; Chen, A.G.; Kim, P.B.; Ames, T.D.; Breaker, R.R. A widespread self-cleaving ribozyme class is revealed by bioinformatics. Nat. Chem. Biol. 2014, 10, 56–60. [Google Scholar] [CrossRef]
- Tang, J.; Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 1997, 4, 453–459. [Google Scholar] [CrossRef]
- Soukup, G.A.; Breaker, R.R. Engineering precision RNA molecular switches. Proc. Natl. Acad. Sci. USA 1999, 96, 3584–3589. [Google Scholar] [CrossRef] [PubMed]
- Zhong, G.; Wang, H.; Bailey, C.C.; Gao, G.; Farzan, M. Rational design of aptazyme riboswitches for efficient control of gene expression in mammalian cells. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Jenison, R.D.; Gill, S.C.; Pardi, A.; Polisky, B. High-resolution molecular discrimination by RNA. Science 1994, 263, 1425–1429. [Google Scholar] [CrossRef]
- Wrist, A.; Sun, W.; Summers, R.M. The Theophylline Aptamer: 25 Years as an Important Tool in Cellular Engineering Research. ACS Synth. Biol. 2020, 9, 682–697. [Google Scholar] [CrossRef]
- Lee, S.W.; Zhao, L.; Pardi, A.; Xia, T. Ultrafast Dynamics Show That the Theophylline and 3-Methylxanthine Aptamers Employ a Conformational Capture Mechanism for Binding Their Ligands. Biochemistry 2010, 49, 2943–2951. [Google Scholar] [CrossRef]
- Zimmermann, G.R.; Jenison, R.D.; Wick, C.L.; Simorre, J.P.; Pardi, A. Interlocking structural motifs mediate molecular discrimination by a theophylline-binding RNA. Nat. Struct. Biol. 1997, 4, 644–649. [Google Scholar] [CrossRef] [PubMed]
- Bayer, T.S.; Smolke, C.D. Programmable ligand-controlled riboregulators of eukaryotic gene expression. Nat. Biotechnol. 2005, 23, 337–343. [Google Scholar] [CrossRef]
- Win, M.N.; Smolke, C.D. A modular and extensible RNA-based gene-regulatory platform for engineering cellular function. Proc. Natl. Acad. Sci. USA 2007, 104, 14283–14288. [Google Scholar] [CrossRef] [PubMed]
- Win, M.N.; Smolke, C.D. Higher-order cellular information processing with synthetic RNA devices. Science 2008, 322, 456–460. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.C.; Chang, A.L.; Kennedy, A.B.; Smolke, C.D. A high-throughput, quantitative cell-based screen for efficient tailoring of RNA device activity. Nucleic Acids Res. 2012, 40, e154. [Google Scholar] [CrossRef] [PubMed]
- Michener, J.K.; Smolke, C.D. High-throughput enzyme evolution in Saccharomyces cerevisiae using a synthetic RNA switch. Metab. Eng. 2012, 14, 306–316. [Google Scholar] [CrossRef]
- Anzalone, A.V.; Lin, A.J.; Zairis, S.; Rabadan, R.; Cornish, V.W. Reprogramming eukaryotic translation with ligand-responsive synthetic RNA switches. Nat. Methods 2016, 13, 453–458. [Google Scholar] [CrossRef]
- Brierley, I. Ribosomal frameshifting viral RNAs. J. Gen. Virol. 1995, 76 Pt 8, 1885–1892. [Google Scholar] [CrossRef]
- Zilio, N.; Wehrkamp-Richter, S.; Boddy, M.N. A new versatile system for rapid control of gene expression in the fission yeast Schizosaccharomyces pombe. Yeast 2012, 29, 425–434. [Google Scholar] [CrossRef]
- Belli, G.; Gari, E.; Piedrafita, L.; Aldea, N.; Herrero, E. An activator/repressor dual system allows tight tetracycline-regulated gene expression in budding yeast. Nucleic Acids Res. 1998, 26, 942–947. [Google Scholar] [CrossRef]
- Wishart, J.A.; Hayes, A.; Wardleworth, L.; Zhang, N.S.; Oliver, S.G. Doxycycline, the drug used to control the tet-regulatable promoter system, has no effect on global gene expression in Saccharomyces cerevisiae. Yeast 2005, 22, 565–569. [Google Scholar] [CrossRef]
- Groher, A.C.; Jager, S.; Schneider, C.; Groher, F.; Hamacher, K.; Suess, B. Tuning the Performance of Synthetic Riboswitches using Machine Learning. ACS Synth. Biol. 2019, 8, 34–44. [Google Scholar] [CrossRef]
- Foerster, U.; Weigand, J.E.; Trojanowski, P.; Suess, B.; Wachtveitl, J. Conformational dynamics of the tetracycline-binding aptamer. Nucleic Acids Res. 2012, 40, 1807–1817. [Google Scholar] [CrossRef] [PubMed]
- Suess, B.; Hanson, S.; Berens, C.; Fink, B.; Schroeder, R.; Hillen, W. Conditional gene expression by controlling translation with tetracycline-binding aptamers. Nucleic Acids Res. 2003, 31, 1853–1858. [Google Scholar] [CrossRef] [PubMed]
- Hanson, S.; Berthelot, K.; Fink, B.; McCarthy, J.E.; Suess, B. Tetracycline-aptamer-mediated translational regulation in yeast. Mol. Microbiol. 2003, 49, 1627–1637. [Google Scholar] [CrossRef] [PubMed]
- Kotter, P.; Weigand, J.E.; Meyer, B.; Entian, K.D.; Suess, B. A fast and efficient translational control system for conditional expression of yeast genes. Nucleic Acids Res. 2009, 37, e120. [Google Scholar] [CrossRef]
- Weigand, J.E.; Suess, B. Tetracycline aptamer-controlled regulation of pre-mRNA splicing in yeast. Nucleic Acids Res. 2007, 35, 4179–4185. [Google Scholar] [CrossRef]
- Wittmann, A.; Suess, B. Selection of tetracycline inducible self-cleaving ribozymes as synthetic devices for gene regulation in yeast. Mol. Biosyst. 2011, 7, 2419–2427. [Google Scholar] [CrossRef]
- Walter, F.; Vicens, Q.; Westhof, E. Aminoglycoside—RNA interactions. Curr. Opin. Chem. Biol. 1999, 3, 694–704. [Google Scholar] [CrossRef]
- Schroeder, R.; Waldsich, C.; Wank, H. Modulation of RNA function by aminoglycoside antibiotics. EMBO J. 2000, 19, 1–9. [Google Scholar] [CrossRef]
- Zhao, F.; Zhao, Q.; Blount, K.F.; Han, Q.; Tor, Y.; Hermann, T. Molecular recognition of RNA by neomycin and a restricted neomycin derivative. Angew. Chem. Int. Ed. 2005, 44, 5329–5334. [Google Scholar] [CrossRef] [PubMed]
- Stage, T.K.; Hertel, K.J.; Uhlenbeck, O.C. Inhibition of the hammerhead ribozyme by neomycin. RNA 1995, 1, 95–101. [Google Scholar] [PubMed]
- Von Ahsen, U.; Davies, J.; Schroeder, R. Antibiotic inhibition of group I ribozyme function. Nature 1991, 353, 368–370. [Google Scholar] [CrossRef]
- Weigand, J.E.; Sanchez, M.; Gunnesch, E.B.; Zeiher, S.; Schroeder, R.; Suess, B. Screening for engineered neomycin riboswitches that control translation initiation. RNA 2008, 14, 89–97. [Google Scholar] [CrossRef]
- Weigand, J.E.; Schmidtke, S.R.; Will, T.J.; Duchardt-Ferner, E.; Hammann, C.; Woehnert, J.; Suess, B. Mechanistic insights into an engineered riboswitch: A switching element which confers riboswitch activity. Nucleic Acids Res. 2011, 39, 3363–3372. [Google Scholar] [CrossRef]
- Duchardt-Ferner, E.; Weigand, J.E.; Ohlenschlaeger, O.; Schtnidtke, S.R.; Suess, B.; Woehnert, J. Highly Modular Structure and Ligand Binding by Conformational Capture in a Minimalistic Riboswitch. Angew. Chem. Int. Ed. 2010, 49, 6216–6219. [Google Scholar] [CrossRef] [PubMed]
- Schmidtke, S.R.; Duchardt-Ferner, E.; Weigand, J.E.; Suess, B.; Woehnert, J. NMR resonance assignments of an engineered neomycin-sensing riboswitch RNA bound to ribostamycin and tobramycin. Biomol. Nmr Assign. 2010, 4, 115–118. [Google Scholar] [CrossRef] [PubMed]
- Krstic, I.; Frolow, O.; Sezer, D.; Endeward, B.; Weigand, J.E.; Suess, B.; Engels, J.W.; Prisner, T.F. PELDOR spectroscopy reveals preorganization of the neomycin-responsive riboswitch tertiary structure. J. Am. Chem. Soc. 2010, 132, 1454–1455. [Google Scholar] [CrossRef]
- Kulik, M.; Mori, T.; Sugita, Y.; Trylska, J. Molecular mechanisms for dynamic regulation of N1 riboswitch by aminoglycosides. Nucleic Acids Res. 2018, 46, 9960–9970. [Google Scholar] [CrossRef]
- Klauser, B.; Atanasov, J.; Siewert, L.K.; Hartig, J.S. Ribozyme-based aminoglycoside switches of gene expression engineered by genetic selection in S. cerevisiae. ACS Synth. Biol. 2015, 4, 516–525. [Google Scholar] [CrossRef]
- Klauser, B.; Rehm, C.; Summerer, D.; Hartig, J.S. Engineering of Ribozyme-Based Aminoglycoside Switches of Gene Expression by In Vivo Genetic Selection in Saccharomyces cerevisiae. Methods Enzymol. 2015, 550, 301–320. [Google Scholar] [PubMed]
- Felletti, M.; Stifel, J.; Wurmthaler, L.A.; Geiger, S.; Hartig, J.S. Twister ribozymes as highly versatile expression platforms for artificial riboswitches. Nat. Commun. 2016, 7, 12834. [Google Scholar] [CrossRef]
- Sack, M.; Stifel, J.; Kreft, S.G.; Deuerling, E.; Hartig, J.S. Neomycin-dependent hammerhead ribozymes for the direct control of gene expression in Saccharomyces cerevisiae. Methods 2019, 161, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Wieland, M.; Gfell, M.; Hartig, J.S. Expanded hammerhead ribozymes containing addressable three-way junctions. RNA 2009, 15, 968–976. [Google Scholar] [CrossRef] [PubMed]
- Murphy, G.J.; Mostoslavsky, G.; Kotton, D.N.; Mulligan, R.C. Exogenous control of mammalian gene expression via modulation of translational termination. Nat. Med. 2006, 12, 1093–1099. [Google Scholar] [CrossRef]
- Boussebayle, A.; Torka, D.; Ollivaud, S.; Braun, J.; Bofill-Bosch, C.; Dombrowski, M.; Groher, F.; Hamacher, K.; Suess, B. Next-level riboswitch development-implementation of Capture-SELEX facilitates identification of a new synthetic riboswitch. Nucleic Acids Res. 2019, 47, 4883–4895. [Google Scholar] [CrossRef]
- Stoltenburg, R.; Nikolaus, N.; Strehlitz, B. Capture-SELEX: Selection of DNA Aptamers for Aminoglycoside Antibiotics. J. Anal. Methods Chem. 2012, 2012, 415697. [Google Scholar] [CrossRef]
- Groher, F.; Bofill-Bosch, C.; Schneider, C.; Braun, J.; Jager, S.; Geissler, K.; Hamacher, K.; Suess, B. Riboswitching with ciprofloxacin-development and characterization of a novel RNA regulator. Nucleic Acids Res. 2018, 46, 2121–2132. [Google Scholar] [CrossRef]
- Grate, D.; Wilson, C. Inducible regulation of the S-cerevisiae cell cycle mediated by an RNA aptamer-ligand complex. Bioorg. Med. Chem. 2001, 9, 2565–2570. [Google Scholar] [CrossRef]
- Lotz, T.S.; Halbritter, T.; Kaiser, C.; Rudolph, M.M.; Kraus, L.; Groher, F.; Steinwand, S.; Wachtveitl, J.; Heckel, A.; Suess, B. A light-responsive RNA aptamer for an azobenzene derivative. Nucleic Acids Res. 2019, 47, 2029–2040. [Google Scholar] [CrossRef]
- Wachter, A.; Tunc-Ozdemir, M.; Grove, B.C.; Green, P.J.; Shintani, D.K.; Breaker, R.R. Riboswitch control of gene expression in plants by splicing and alternative 3′ end processing of mRNAs. Plant. Cell 2007, 19, 3437–3450. [Google Scholar] [CrossRef]
- Cheah, M.T.; Wachter, A.; Sudarsan, N.; Breaker, R.R. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 2007, 447, 497–500. [Google Scholar] [CrossRef]
- McRose, D.; Guo, J.; Monier, A.; Sudek, S.; Wilken, S.; Yan, S.; Mock, T.; Archibald, J.M.; Begley, T.P.; Reyes-Prieto, A.; et al. Alternatives to vitamin B-1 uptake revealed with discovery of riboswitches in multiple marine eukaryotic lineages. ISME J. 2014, 8, 2517–2529. [Google Scholar] [CrossRef]
- Donovan, P.D.; Holland, L.M.; Lombardi, L.; Coughlan, A.Y.; Higgins, D.G.; Wolfe, K.H.; Butler, G. TPP riboswitch-dependent regulation of an ancient thiamin transporter in Candida. PLoS Genet. 2018, 14, e1007429. [Google Scholar] [CrossRef]
- Marchisio, M.A.; Huang, Z. CRISPR-Cas type II-based Synthetic Biology applications in eukaryotic cells. RNA Biol. 2017, 14, 1286–1293. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- DiCarlo, J.E.; Norville, J.E.; Mali, P.; Rios, X.; Aach, J.; Church, G.M. Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Res. 2013, 41, 4336–4343. [Google Scholar] [CrossRef]
- Gao, Y.; Zhao, Y. Self- processing of ribozyme-flanked RNAs into guide RNAs in vitro and in vivo for CRISPR-mediated genome editing. J. Integr. Plant Biol. 2014, 56, 343–349. [Google Scholar] [CrossRef]
- Li, J.; Xu, Z.; Chupalov, A.; Marchisio, M.A. Anti-CRISPR-based biosensors in the yeast S. cerevisiae. J. Biol. Eng. 2018, 12, 11. [Google Scholar] [CrossRef] [PubMed]
- Pley, H.W.; Flaherty, K.M.; McKay, D.B. Three-dimensional structure of a hammerhead ribozyme. Nature 1994, 372, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Ferre-D’Amare, A.R.; Zhou, K.H.; Doudna, J.A. Crystal structure of a hepatitis delta virus ribozyme. Nature 1998, 395, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Gander, M.W.; Vrana, J.D.; Voje, W.E.; Carothers, J.M.; Klavins, E. Digital logic circuits in yeast with CRISPR-dCas9 NOR gates. Nat. Commun. 2017, 8, 15459. [Google Scholar] [CrossRef]
- Kundert, K.; Lucas, J.E.; Watters, K.E.; Fellmann, C.; Ng, A.H.; Heineike, B.M.; Fitzsimmons, C.M.; Oakes, B.L.; Qu, J.; Prasad, N.; et al. Controlling CRISPR-Cas9 with ligand-activated and ligand-deactivated sgRNAs. Nat. Commun. 2019, 10, 2127. [Google Scholar] [CrossRef]
- Qi, L.; Haurwitz, R.E.; Shao, W.; Doudna, J.A.; Arkin, A.P. RNA processing enables predictable programming of gene expression. Nat. Biotechnol. 2012, 30, 1002–1006. [Google Scholar] [CrossRef] [PubMed]
- Haurwitz, R.E.; Jinek, M.; Wiedenheft, B.; Zhou, K.; Doudna, J.A. Sequence- and Structure-Specific RNA Processing by a CRISPR Endonuclease. Science 2010, 329, 1355–1358. [Google Scholar] [CrossRef] [PubMed]
- Cady, K.C.; O’Toole, G.A. Non-Identity-Mediated CRISPR-Bacteriophage Interaction Mediated via the Csy and Cas3 Proteins. J. Bacteriol. 2011, 193, 3433–3445. [Google Scholar] [CrossRef]
- Wu, H.H.; Yang, P.K.; Butcher, S.E.; Kang, S.; Chanfreau, G.; Feigon, J. A novel family of RNA tetraloop structure forms the recognition site for Saccharomyces cerevisiae RNase III. EMBO J. 2001, 20, 7240–7249. [Google Scholar] [CrossRef]
- Abou Elela, S.; Ji, X. Structure and function of Rnt1p: An alternative to RNAi for targeted RNA degradation. Wiley Interdiscip. Rev. RNA 2019, 10. [Google Scholar] [CrossRef]
- Lamontagne, B.; Ghazal, G.; Lebars, I.; Yoshizawa, S.; Fourmy, D.; Abou Elela, S. Sequence Dependence of Substrate Recognition and Cleavage by Yeast RNase III. J. Mol. Biol. 2003, 327, 985–1000. [Google Scholar] [CrossRef]
- Wu, H.H.; Henras, A.; Chanfreau, G.; Feigon, J. Structural basis for recognition of the AGNN tetraloop RNA fold by the double-stranded RNA-binding domain of Rnt1p RNase III. Proc. Natl. Acad. Sci. USA 2004, 101, 8307–8312. [Google Scholar] [CrossRef]
- Lamontagne, B.; Abou Elela, S. Short RNA guides cleavage by eukaryotic RNase III. PLoS ONE 2007, 2, e472. [Google Scholar] [CrossRef]
- Babiskin, A.H.; Smolke, C.D. A synthetic library of RNA control modules for predictable tuning of gene expression in yeast. Mol. Syst Biol. 2011, 7, 471. [Google Scholar] [CrossRef] [PubMed]
- Babiskin, A.H.; Smolke, C.D. Synthetic RNA modules for fine-tuning gene expression levels in yeast by modulating RNase III activity. Nucleic Acids Res. 2011, 39, 8651–8664. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, J.; Sonenberg, N. Insertion mutagenesis to increase secondary structure within the 5’ noncoding region of a eukaryotic mRNA reduces translational efficiency. Cell 1985, 40, 515–526. [Google Scholar] [CrossRef]
- Babiskin, A.H.; Smolke, C.D. Engineering ligand-responsive RNA controllers in yeast through the assembly of RNase III tuning modules. Nucleic Acids Res. 2011, 39, 5299–5311. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Samarsky, D.A.; Ferbeyre, G.; Bertrand, E.; Singer, R.H.; Cedergren, R.; Fournier, M.J. A small nucleolar RNA:ribozyme hybrid cleaves a nucleolar RNA target in vivo with near-perfect efficiency. Proc. Natl. Acad. Sci. USA 1999, 96, 6609–6614. [Google Scholar] [CrossRef] [PubMed]
- Hirao, I.; Spingola, M.; Peabody, D.; Ellington, A.D. The limits of specificity: An experimental analysis with RNA aptamers to MS2 coat protein variants. Mol. Divers. 1998, 4, 75–89. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, A.B.; Vowles, J.V.; d’Espaux, L.; Smolke, C.D. Protein-responsive ribozyme switches in eukaryotic cells. Nucleic Acids Res. 2014, 42, 12306–12321. [Google Scholar] [CrossRef]
- Weenink, T.; van der Hilst, J.; McKiernan, R.M.; Ellis, T. Design of RNA hairpin modules that predictably tune translation in yeast. Synth. Biol. 2018, 3, ysy019. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Siederdissen, C.H.Z.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Biol. 2011, 6. [Google Scholar] [CrossRef]
- Grilly, C.; Stricker, J.; Pang, W.L.; Bennett, M.R.; Hasty, J. A synthetic gene network for tuning protein degradation in Saccharomyces cerevisiae. Mol. Syst. Biol. 2007, 3, 127. [Google Scholar] [CrossRef] [PubMed]
- Reichel, C.; Johnsson, N. The split-ubiquitin sensor: Measuring interactions and conformational alterations of proteins in vivo. Methods Enzymol. 2005, 399, 757–776. [Google Scholar] [CrossRef] [PubMed]
- Kiga, D.; Futamura, Y.; Sakamoto, K.; Yokoyama, S. An RNA aptamer to the xanthine/guanine base with a distinctive mode of purine recognition. Nucleic Acids Res. 1998, 26, 1755–1760. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.S.; Kaplan, M.; Dykstra, P.; Hinks, M.; McKeague, M.; Smolke, C.D. Massively parallel RNA device engineering in mammalian cells with RNA-Seq. Nat. Commun. 2019, 10, 4327. [Google Scholar] [CrossRef] [PubMed]
- Yokobayashi, Y. Aptamer-based and aptazyme-based riboswitches in mammalian cells. Curr. Opin. Chem. Biol. 2019, 52, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Wurmthaler, L.A.; Klauser, B.; Hartig, J.S. Highly motif- and organism-dependent effects of naturally occurring hammerhead ribozyme sequences on gene expression. RNA Biol. 2018, 15, 231–241. [Google Scholar] [CrossRef] [PubMed]
| RNA Structure | Kd | Additional RNA | Action Triggered by the Chemicals | Performance | Reference(s) |
|---|---|---|---|---|---|
| Theophylline aptamer mTCT8-4 | 0.1 μM | Short antisense RNA | Increase/decrease in gene expression upon binding near the START codon | 90% reduction (OFF switch) or increase (ON switch) in fluorescence expression (1 to 10 mM of theophylline) | [32] |
| HHR | Induction/inhibition of ribozyme self-cleavage | 14-fold increase in fluorescence expression from an AND gate (5–10 mM theophylline and 0.25–0.5 mM tetracycline) | [33,34] | ||
| - | Translational frame shift (-1 PRF) | 7.0-fold increase or 5.9-fold decrease in the translation frame shift (40 mM theophylline) | [38] | ||
| Tetracycline aptamer | 0.8 nM | - | Translation inhibition upon placement on the 5’ UTR of gfp | Single aptamer: 6-to 9-fold fluorescence repression; 2 and 3 aptamers: 21-fold and 37-fold fluorescence repression, respectively. In every case, 250 μM tetracycline were used | [44,45] |
| Intron | Pre-mRNA splicing | Unquantified fluorescence reduction | [47] | ||
| HHR | Ribozyme self-cleavage | Complete self-cleaved (1 μM tetracycline) | [48] | ||
| Neomycin aptamer | - | - | Translational frame shift (-1 PRF) | 5.0-fold enhancement or 4.2-fold reduction in the translation frame shift (550 µM neomycin) | [37] |
| Neomycin aptamer N1 Neomycin aptamer N1-based riboswitch | - - | S. mansoni HHR | Translation inhibition upon insertion on the 3’UTR of gal4 | Around 25-fold lacZ expression downregulation (100 μg/mL) | [60,61] |
| Inactive env-9 twister ribozyme | Translation inhibition upon insertion on the 3’UTR of gal4 | About 10-fold decrease in lacZ expression | [62] | ||
| Neomycin aptamer M4 or M7 | - | S. mansoni HRR | Translation inhibition upon insertion on the 3’UTR of gfp | Around 2-fold fluorescence upregulation (ON switch) and 3-fold fluorescence downregulation (OFF switch) (100 μg/mL neomycin) | [64] |
| Neomycin aptamer M4 | - | - | Translation inhibition | 7.5-fold fluorescence repression (100 µM neomycin) | [54] |
| PARO riboswitch (paromomycin) | 20 nM | - | Translation inhibition | 8.5-fold decrease in gene expression (250 μM paromomycin) | [66,67] |
| 13.8-fold decrease in gene expression from a NOR gate (250 μM of both paromomycin and neomycin) | |||||
| fluoroquinolone ciprofloxacin riboswitch | 64.2 nM | - | Translation inhibition | 7.5-fold fluorescence downregulation | [68] |
| Tetra- methylrosamine aptamer | - | - | Translation inhibition upon placement on the 5’ UTR of CLB2 | Reduction in cell growth | [69] |
| azoCm aptamer | - | - | Configurational change | Unquantified control of gene expression | [70] |
| TPP (thiamine pyrophosphate) riboswitch | - | Intron | Splicing inhibition | pre-mRNA is not spliced in the presence of 10 µM thiamine | [74] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ge, H.; Marchisio, M.A. Aptamers, Riboswitches, and Ribozymes in S. cerevisiae Synthetic Biology. Life 2021, 11, 248. https://doi.org/10.3390/life11030248
Ge H, Marchisio MA. Aptamers, Riboswitches, and Ribozymes in S. cerevisiae Synthetic Biology. Life. 2021; 11(3):248. https://doi.org/10.3390/life11030248
Chicago/Turabian StyleGe, Huanhuan, and Mario Andrea Marchisio. 2021. "Aptamers, Riboswitches, and Ribozymes in S. cerevisiae Synthetic Biology" Life 11, no. 3: 248. https://doi.org/10.3390/life11030248
APA StyleGe, H., & Marchisio, M. A. (2021). Aptamers, Riboswitches, and Ribozymes in S. cerevisiae Synthetic Biology. Life, 11(3), 248. https://doi.org/10.3390/life11030248







