Abstract
Biosensors are indispensable tools to understand a plant’s immunity as its spatiotemporal dimension is key in withstanding complex plant immune signaling. The diversity of genetically encoded biosensors in plants is expanding, covering new analytes with ever higher sensitivity and robustness, but their assortment is limited in some respects, such as their use in following biotic stress response, employing more than one biosensor in the same chassis, and their implementation into crops. In this review, we focused on the available biosensors that encompass these aspects. We show that in vivo imaging of calcium and reactive oxygen species is satisfactorily covered with the available genetically encoded biosensors, while on the other hand they are still underrepresented when it comes to imaging of the main three hormonal players in the immune response: salicylic acid, ethylene and jasmonic acid. Following more than one analyte in the same chassis, upon one or more conditions, has so far been possible by using the most advanced genetically encoded biosensors in plants which allow the monitoring of calcium and the two main hormonal pathways involved in plant development, auxin and cytokinin. These kinds of biosensor are also the most evolved in crops. In the last section, we examine the challenges in the use of biosensors and demonstrate some strategies to overcome them.
1. Biosensors: Exploiting Molecular Hubs to Better Understand the Processes
A biosensor is a sensitive device that detects an analyte or event in a living organism and consequently produces a measurable output [1]. Thus, it consists of two functional parts: detector and reporter [2]. The detector domain must be specific and sensitive to enable detection of the biological concentration of analytes or events. The biosensor as a whole should enable detection with high signal to noise ratio (SNR), high spatiotemporal resolution on the organelle level, enable fast response, and must be functional in different cellular conditions (low pH, different redox states). Besides, it should not interfere with cellular processes and should not be toxic to the cells. Although each of available biosensors has its own limitations and cannot meet all of the abovementioned requirements, they are vital to obtain an insight into cellular events.
According to their mechanism of action, they can be separated into direct and indirect biosensors. Typical direct biosensors report protein activity upon ligand binding. Usually, they consist of two protein domains, one serving as a ligand receptor while the other can sense structural changes upon ligand binding and report it by measurable output. This is the mode of action of one of the first known biosensors named Cameleon, which is still widely used to measure Ca2+ concentration (Figure 1). It is translated as a single peptide chain, with two fluorescent proteins on each side, which are either blue and green or cyan and yellow, linked by calmodulin (CaM) and the M13 domain. When the concentration of Ca2+ in the cellular compartment increases, Ca2+ binds to CaM, which twists around the M13 domain and thereupon the two fluorescent proteins are close enough to enable Förster resonance energy transfer (FRET) and Ca2+ binding can be detected through the change of the fluorescence intensities [3]. Another kind of direct biosensors are degron-based biosensors, which undergo degradation as a result of analyte binding. They exploit the characteristics of the cellular signaling of some plant hormones. In the presence of the hormone in the cell, related transcriptional repressors are degraded and thus the transcription of the hormone-responsive genes is activated. This approach is used, for example, in a plant biosensor for auxin detection, known as DII-VENUS (Figure 1). This is a fusion protein of two domains, of which DII acts as a detector while fast-maturing fluorescent protein VENUS acts as a reporter. The DII domain is a part of the Aux/IAA repressor involved in the binding of auxin and subsequent degradation of the repressor via the ubiquitin/26S proteasome pathway. When auxin concentration in the cell increases, DII-VENUS fusion is degraded and the fluorescence intensity decreases [4]. Direct biosensors can be used for following changes in pH, redox state, ion and metabolite concentrations in the majority of plant cell compartments.
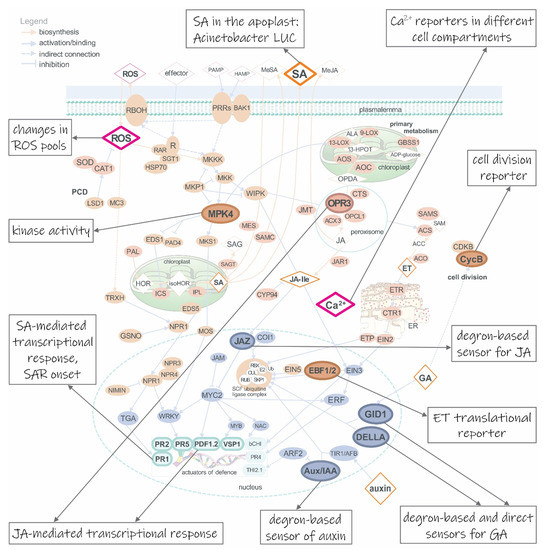
Figure 1.
Diversity of available biosensors for following immune response in plants. During the first stage of the immune response, Ca2+ influx in the cytosol can be monitored with diverse reporter proteins. They can be Förster resonance energy transfer (FRET) based, employing two FPs linked by calmodulin-sensing domain, e.g. Cameleons [3], or colourful GECOs (genetically encoded Ca2+ indicators for optical imaging) [28], employing one, circularly permutated fluorescent protein. Prominent sensors of reactive oxygen species (ROS) are redox-sensitive GFPs in fusion with oxidant receptor peroxidase-1 (roGFP2-Orp1) [29] and glutaredoxin 1 (Grx1-roGFP2) [30], H2O2 and glutathione sensors, respectively, that were used to distinguish the patterns of these two molecular species in the cytosol, chloroplast and mitochondria upon illumination [31]. The main three hormones of immune response, salicylic acid (SA), jasmonic acid (JA) and ethylene (ET), can be followed with a direct degron-based sensor Jas9-VENUS in case of JA or with indirect transcriptional reporters based on defense genes’ promoters, such as pathogen-responsive PR1, PR2 and PR5 for SA- and plant defensin 1.2 (PDF1.2) and vegetative storage protein (VSP) for JA-involving response. They can be additionally followed by transcriptional reporters that exploit promoters of genes involved in hormone biosynthesis, e.g. the promoter of 12-oxo-phytodienoic acid reductase 3 (OPR3) gene, a constituent of JA biosynthesis. However, these transcriptional reporters can only complement other biosensors as they are not sufficiently specific. SA in the apoplast can be followed through Acinetobacter sp. ADPWH_lux luciferase (LUC) activity [32]. ET presence can be followed with translational reporter, joining 3’-UTR mRNA of EBF2, repressor of ET-responsive genes, and coding sequence of FP [8,9]. Other hormones, such as auxin and gibberellins (GA), can also be followed with degron-based sensors, employing DII domain of Aux/IAA repressor (in case of auxin [4]) and DELLA repressor (in case of GA [33]). Auxin can also be monitored through more of its actions, not just directly with DII-FP, but also indirectly with transcriptional reporters of auxin-responsive genes. These can employ synthetic (DR5 [6]) promoter as a detector. On the other hand, GA can be followed with FRET-based direct biosensor exploiting interaction between GID1 and DELLA repressor [34]. Interplay of immune response and plant’s growth and development can be followed with biosensors of cell division, employing cyclins, e.g. CycB [35]. AtMPK4 and some other kinases’ activity can be detected with kinase localization reporters, employing a domain for kinase docking and a phosphorylation site. When phosphorylated, fluorescent reporter is exported from the nucleus [36]. Signaling proteins are presented as orange, enzymes as brown-red and transcription factors as grey ovals, respectively; metabolites are presented as rhombs and genes as squircles. Full and dotted arrows represent direct and indirect connection; orange and grey arrows represent biosynthetic pathways and binding; point and block arrows represent activation and inhibition, respectively. ARF2, auxin response factor 2; Aux/IAA, a member from the protein family of auxin-sensitive transcriptional repressors; CDPK, cyclin dependent phosphokinase; DELLA, a member of the protein family of GA-sensitive transcriptional repressors; GID1, gibberellin insensitive dwarf 1; JAZ, jasmonate ZIM-domain; MPK4, mitogen-activated protein kinase 4; PR1, 2 and 5, pathogenesis-related protein 1, 2 and 5, respectively. Figure was adapted on the basis of Figure 1c from Lukan et al. [37] with author’s permission.
Indirect biosensors are typically transcriptional reporters. Their detector domain is a promoter sequence that contains analyte-responding cis-elements and drives transcription of the reporter gene. The most commonly used reporters are beta-glucuronidase (GUS), fluorescent proteins (FPs) and luciferases. The signal produced by such biosensors is delayed, but amplified in comparison with direct biosensors. In order to gain higher SNR, specificity and sensitivity, native analyte-responsive promoter sequences are usually rebuilt by fusing cis-elements to the minimal Cauliflower mosaic virus (CaMV) 35S promoter sequence [5]. The most known example is the DR5 synthetic promoter, designed on the basis of GH3 gene promoter for auxin detection, which maintains its activity also in reverse orientation (DR5rev, Figure 1) [6]. The latter was improved to version DR5v2 with higher expression and sensitivity and therefore enables detection with better spatial resolution [7].
Recently, another type of indirect biosensor was developed to follow translational regulation of mRNA transcripts in the presence of ethylene. In this biosensor, the detector module is the ethylene-responsive 3’-UTR part of mRNA coding for EBF2, while the coding sequence is translated into reporter protein GFP (Figure 1). The mechanism of this biosensor is based on the action of C-terminal peptide of EIN2, which is cleaved when the concentration of ethylene increases. It can then bind to the 3’-UTR part from EBF2 mRNA fused with the GFP coding sequence, and thus represses its translation. In this way, GFP fluorescence decreases when the concentration of ET is increased [8,9,10].
During the last ten years, several informative reviews covering the topic of plant biosensors have been published, showing the advances in this technology and its importance for the plant research community. They revise the type of available biosensors and the recent results obtained [1], the use of biosensors to monitor plant hormones [11,12,13], the principles of most widely used biosensors in plants [14] and quantitative measurements with biosensors [15]. Some reviews are focused on biosensors of a single analyte such as abscisic acid (ABA) [16], auxin [17], Ca2+ [18], ethylene (ET) [10], gibberellins (GA) [19] and reactive oxygen species (ROS) [20]. The span of the developed biosensors goes hand in hand with advances of methodology, which exploits various principles and physical properties of the fluorescent proteins and other reporter proteins. The spectrum of advanced fluorescence imaging methods available nowadays for the use in plants is reviewed in Komis et al. [21]. The Fluorescent Biosensor Database [22] merges fluorescent genetically encoded biosensors regardless of the chassis and welcomes new updates by the community.
Variability of the biosensors available nowadays is broad. However, their use in plants is still limited to certain aspects. The majority of studies focus on the use of biosensors in roots to follow development in the model plant Arabidopsis thaliana. Consequently, the vast majority of biosensors were designed in this regard, providing even more than one sensor for a single analyte involved in a plant’s growth and development, each of them exploiting another cellular event. On the other hand, the span of available biosensors for immune response is narrow. While Ca2+ and ROS, involved in the first stages of signal transmission in general, are satisfactorily covered, the main hormones of immune response, salicylic acid (SA), jasmonic acid (JA) and ET, lack specific, sensitive and thus reliable biosensors to choose between. Leading research in plant development is also seen from the reports on following more than one analyte simultaneously and the transmission of the biosensors from model organisms to crops. Here, we report these findings. Finally, we also discuss difficulties associated with the application of the biosensors in plants with the aim of supporting the advancement of immune response biosensors and their application to crops.
2. Genetically Encoded Biosensors for Following Plant Immune Response
Plants respond to biotic stress by reprogramming a complex signaling network that results in gene activity and metabolic changes. Translation of pathogen recognition into effective defense response strongly depends on the action of several plant hormones and other signaling molecules [23,24,25]. Early signaling events include changes of intracellular Ca2+ levels and a rapid increase of reactive oxygen species. Among plant hormones, SA, JA and ET have been identified as main players [24,26]. In addition, recent evidence shows that the effects of these three hormonal signaling pathways are balanced by ABA, GA, auxins, cytokinins and brassinosteroids (reviewed by Verma et al. [27]), thus adding another layer of regulation. Therefore, these are the best candidate analytes to follow general immune response (Figure 1).
Ca2+ is the most versatile messenger regulating a wide range of responses, including biotic stress. After pathogen recognition, one of the earliest signaling events is the Ca2+ influx into the plant cell cytosol. Ca2+ signatures are also induced in the nucleus, mitochondria or chloroplasts and there is a complex interplay between Ca2+ and other messengers and their signaling pathways [38]. Moreover, it has been shown that Ca2+-mediated signaling is also involved in negative regulation of plant immunity [39]. Thus, there are still many questions that should be addressed. Ca2+ sensors are to date the most advanced among all biosensors. They have progressed from bioluminescent aequorin, which is natively anchoring calcium [40], to a wide range of designed fluorescent reporters employing calcium-binding domain CaM, e.g. FRET-based Cameleons (Figure 1) [3,41], single fluorophore GCaMPs (composed of circularly permutated enhanced GFP, CaM and M13 peptide) [42] and GECOs (genetically encoded Ca2+ indicators for optical imaging) [28] (Figure 1), and even GFP-aequorin, exploiting bioluminescence resonance energy transfer (BRET) [43]. Ca2+ response is fast and thus demands constitutively and strongly expressed direct biosensors, enabling subcellular [28], tissue and whole-plant imaging of calcium release reaching very high temporal resolution measured in seconds [44].
Despite the new insights that have been brought into the role of redox mechanisms in plant defence response, one of the major challenges still is to understand the spatial and temporal redox processes occurring during the defence response and to associate the transcriptional activity with the complex dynamics of these signaling molecules [45,46]. Several related biosensors have been successfully used in plants (reviewed in Choi et al. [47]) to help unravel the spatiotemporal redox signaling occurring during plant defense response. As an example, roGFPs are mutated GFP molecules sensitive to redox levels in the cell [48], which were later fused to signal sequences to allow targeting to different subcellular organelles, such as mitochondria [49] and chloroplasts [50]. Fusion partners known to be targets of redox transitions by glutathione (Figure 1) [30] or H2O2 (Figure 1) [29] were also designed and recently used for time-resolved measurements of both molecular species in the cytosol, chloroplasts and mitochondria [31]. This allowed for better understanding of the role of the mitochondria in sensing and signaling the cellular redox challenge in response to abiotic stress [51], deciphering the role of redox state in intercellular transport [50] or exploring the central role of glutathione in mediating redox signaling [52]. Another genetically encoded redox sensor used in plants is HyPer, based on circularly permutated YFP coupled with the H2O2-sensitive domain of OxyR, a transcription factor found in Escherichia coli [53]. A newly developed biosensor in plants, CROST (change in redox state of thioredoxin) employs a FRET-pair linked with redox-sensitive domain CP12 from A. thaliana, known to be reduced in vivo by thioredoxin [54]. Still, available H2O2 and redox sensors are frequently limited by extreme pH and redox conditions which must be considered when choosing the appropriate sensor for certain cellular compartments: apoplast, vacuole, endoplasmic reticulum, chloroplast, and mitochondria [20].
Recent advances in the development of plant biosensors (see review Novák et al. [11]) have helped to better understand dynamics of plant signaling, in particular in developmental processes [55]. Some of these can also be applied to monitor the immune response as some parts of the signaling network modules overlap. The most successful biosensors for hormones are transcriptional reporters, based on hormone-responsive promoter motifs fused to a reporter element. The first generation used the synthetic promoter fused to GUS, but, more recently, FPs have shown to be more versatile and have become the reporter of choice [56].
Transcriptional reporters have worked well not just in case of the abovementioned DR5 for auxin, but also in the case of the cytokinin Two Component System (TCS) [57] that was developed and used to uncover the roles of cytokinin signaling in A. thaliana root regeneration. It is named after the two-component phosphorelay cascade, the basis of the cytokinin signaling. The final target of the phosphorylation-triggered activation are transcription factors named B-type response regulators which activate the expression of cytokinin responsive genes. Their DNA binding sites are highly conserved and are thus exploited by TCS biosensors, joining six direct repeats to a minimal CaMV 35S promoter. The synthetic sensor TCS and its improved variants TCSnew (TCSn) [58] and TCS version 2 (TCSv2) [59] have been widely used in the model plant A. thaliana. Recently, a new synthetic transcriptional reporter for ABA was designed as six repeats of ABA-responsive elements (ABRE) from RD29A or ABI1 promoters fused to a minimal promoter, driving expression of either GUS or GFP targeted to endoplasmic reticulum. It was shown that both transcriptional reporters respond to osmotic stress [60].
Entanglement of the signaling pathways of the main three immune response hormones, SA, JA and ET, challenges the search for their specific transcriptional reporters. Promoters of pathogenesis-related proteins (PRs) are usually employed as transcriptional markers for SA signaling (Figure 1), while promoters of genes involved in JA biosynthesis (e.g. 12-oxo-phytodienoic acid reductase 3, OPR3), regulators of transcription (jasmonate-ZIM-domain 10, JAZ10) or target genes, e.g. plant defensin 1.2 (PDF1.2) and vegetative storage protein (VSP), are analyzed in connection with JA signaling (Figure 1). However, these promoters exhibit significant crosstalk between SA, JA and ET signaling pathways and are thus treated as defense-responsive genes. To our knowledge, specific genetically encoded promoter-based biosensors for SA and JA have not yet been developed in plants. ET transcriptional reporters are based on a synthetic promoter composed of five repeats of the Ethylene insensitive 3 (EIN3) binding site attached to the minimal CaMV 35S promoter [61]. So far, this has been used in combination with either luciferase or GUS due to its low strength [10]. Lately, some new promoters responding to ABA, auxin, cytokinin, SA and JA were identified in A. thaliana as promising candidates for transcriptional reporters [62].
Available direct biosensors that track plant hormone concentrations are based on either FRET or degrons. Recently, a major advance in high-resolution quantification of spatiotemporal GA distribution was achieved with the development of a sensor directly measuring GA. This exploits the interaction of GA receptor gibberellin insensitive dwarf 1 (GID1) with members of the DELLA family. Rizza et al. developed and implemented this FRET-based sensor (Gibberellin Perception Sensor 1, GPS1, Figure 1) in A. thaliana [34]. In fact, the first FRET-biosensors were developed for ABA measurement and were published in parallel by two different research groups: ABA concentration and uptake sensor (ABACUS) [63], and ABAleons [64].
Using a degron design, Larrieu et al. developed a biosensor for JA perception (Jas9-VENUS, Figure 1) and demonstrated its value for quantitative and dynamic analysis of JA response in A. thaliana roots [65]. The Jas9-VENUS biosensor uses the Jas motif of Jasmonate-ZIM-Domain (JAZ) proteins that are targeted to degradation via the ubiquitin/26S proteasome pathway in the presence of the bioactive form of JA. Another degron-based biosensor designed to monitor GA is the GFP-tagged DELLA protein repressor of GA1-3 (GFP-RGA, Figure 1) that was used in the model plant A. thaliana and revealed asymmetric distribution of GA and GA signaling during root gravitropic growth [33]. StrigoQuant is also a degron-based but luminescent reporter which includes two luciferases, firefly and Renilla luciferase (FLUC and RLUC, respectively) that are expressed under the same promoter. Co-translationally, fusion of FLUC and RLUC is cleaved by 2A self-cleaving peptide which links both reporters: one is degraded upon presence of strigolactons, the other is used for normalization of expression [66].
Both direct and indirect biosensors are now frequently designed along with their nonresponsive counterparts, which were constructed in the same way but are not responsive to the analyte. Thus, they show the background signal from the biosensor itself, and can be used directly for normalization when fused to another reporter protein. Some examples are nlsGPS1 and nlsGPS1-NR (gibberellin) [67], Jas9-VENUS and mJas9-VENUS [65], DII and mDII (e.g. R2D2: DII-3xVENUS and mDII-ntdTomato) [7], TCS and TCSm [57].
Other useful biosensors of immune response are those following the activity of mitogen-activated protein kinases (MAPKs) through the use of docking domain and phosphorylation site (FRET biosensors and KTRs, kinase translocation reporters, in which a reporter changes its intracellular localization when phosphorylated), namely A. thaliana’s MPK3, MPK4, MPK6, which take part in response to flg22, chitin and NaCl (Figure 1) [36,68]. The SnRK2 (sucrose nonfermenting-1-related kinase 2) activity sensor (SNACS) is another kinase activity FRET biosensor for ABA-responsive SnRK2 protein kinases involved in stomata closure, stably transformed in A. thaliana and its mutants [69].
To follow the outcome of defence and growth antagonism in the whole plant, it is possible to engage cell division reporters based on cyclins, for example, CYCB1;1, B-type cyclin that is present only in late G2 phase and early M phase of the cell cycle: AtpCYCB1;1::CYCB1;1-tYFPnls (Figure 1), also available in fusion with GUS [35], or CYCD6;1, driving S-phase of DNA replication: CYCD6;1::GFP [70]. Recently, a three-component biosensor was designed to follow the whole cell cycle in A. thaliana [71]. It contains a multigene cassette expressing fusions of CDT1a-eCFP, HTR13-mCherry and N-CYCB1;1-YFP, each under the control of the native promoter of the first fusion partner. CDT1a is involved in initiation of DNA replication and is expressed during the S and G2 phases, HTR13 is histone H3.1 protein and is expressed during the M and G1 phases, while N-CYCB1;1 is expressed during the late G2 and prophase and metaphase of mitosis [71]. Some possible solutions have not yet been applied to plants. Recently, another promising type of biosensor was established in the animalia kingdom, FlipGFP, for following protease activity. This is based on tripartite GFP, which fluoresces only when reconstituted with two missing beta-strands. These become available after the cleavage of their fusion by a specific protease [72].
3. Getting a Broader View and Deeper Understanding: More Than One Biosensor in the Same Chassis
In the field, plants are rarely exposed to a single stress. They often interact with different organisms, either simultaneously or sequentially, and can be affected by several abiotic stresses such as drought, heat or salinity. The effects of these adverse conditions in plant growth and yield can be devastating and are becoming more problematic with the advent of climate change. Understanding the response of plants to environmental conditions and how the different signaling pathways interact is crucial to guarantee efficient crop protection strategies. Several studies have reported that the involvement of different signaling pathways in response to multiple stresses and three-way interactions cannot be inferred from the response to a single stress [73,74]. Therefore, it is of high importance to gain better insights into the plant response to multiple stresses and the use of combined biosensors could be a promising tool to greatly advance this field.
Employing more than one biosensor simultaneously can provide information for more than one analyte or for the same analyte in more than one cell compartment. Although it demands additional efforts in cloning, transforming, imaging and analysis, examples have recently shown the value of this approach. Waadt et al. [75] investigated the interdependence of calcium and ABA signaling in A. thaliana roots. The authors performed multiparameter imaging of both analytes combining the red-emitting single-FP genetically encoded Ca2+ indicators for optical imaging (R-GECO1) [76] and the FRET-based ABA reporter ABAleon2.1 emitting in cyan/yellow [64]. Taking advantage of the high sensitivity of GECOs, a dual sensor for monitoring Ca2+ signal dynamics in the cytoplasm and the nuclear compartments was developed by assembling the nuclear-R-GECO1 (NR-GECO1) and the cytoplasmic green GECO1 (CG-GECO1) in a single construct [28]. The dual GECO sensor was shown to be a useful tool to monitor Ca2+ signal response to biotic and abiotic stress of Medicago truncatula and A. thaliana roots [28]. Another example of dual sensors is the use of transgenic A. thaliana plants simultaneously expressing DR5::3xVENUS-N7 [77] and TCS::GFP [57] reporters to study the spatial patterns of auxin and cytokinins, respectively [78].
The analysis of several analytes simultaneously requires the generation of transgenic plants that express several genetically encoded sensors by co-transformations with single transcriptional units or transformation with a multigene cassette following the assembly of single sensors. The generation of transgenic plants is time-consuming and the insertion of multiple transgenes into the A. thaliana genome could result in epigenetic silencing effects [79]. In contrast, it has been shown that the inducible co-expression of two interacting proteins, each tagged with FP, in a single multigene expression cassette reduces variability in expression of the proteins in a single cell, avoids mosaicism and can increase FRET [80]. By expressing two biosensors from one single mRNA Waadt et al. introduced the concept of dual-reporting transcriptionally linked genetically encoded fluorescent indicators (2-in-1-GEFIs). The two fluorescent proteins are separated by 2A self-cleaving peptide. This sensor was used for the multiparametric analysis of ABA, Ca2+, protons, chloride, the glutathione redox potential, and H2O2 in A. thaliana roots [81].
Many of these approaches are not specific to fluorescent proteins, but can be also applied to the constructs with luciferases. Recent research provided new luciferases that emit light of different wavelengths and can thus be used simultaneously (similar to FPs), or even in fusion with FPs in BRET experiments [82]. Apart from StrigoQuant [66], a pair of FLUC and RLUC was used as a sensor for miRNA silencing, where FLUC transcript was miRNA’s target while RLUC was used for normalization [83]. More recent luciferases that complement each other with respect to emission wavelength are the red firefly luciferase (redLUC) and Gaussia Dura luciferase (gLUC) that have already been used in a similar manner to StrigoQuant [84]. NanoLUC is smaller than other luciferases, does not use ATP and allows higher temporal resolution [85]. Light produced from another luciferase, GeNL, performs high transmittance through plant tissue when used with appropriate substrate [86].
In order to follow various molecules and more aspects of a response in a spatiotemporal manner, different approaches can be used simultaneously. We can exploit the options of non-genetically encoded biosensors such as nanosensors which are applied onto the surface, such as carbon nanotubes that change their fluorescence when exposed to higher H2O2 concentrations [87]. It has also been shown in A. thaliana, wheat and maize that plants can be fumigated with permeable fluorescent probes that irreversibly react with ROS [88]. Microbial biosensors, such as Acinetobacter sp. ADPWH_lux, can use SA as a sole carbon source and can therefore be exploited for SA detection (Figure 1) [32]. Such an approach is especially useful to gain appropriate time resolution when we want to avoid laborious and time-consuming stable transformation or when stably and constitutively expressed reporters cannot be obtained.
Single or multiple sensors can be used in parallel with reporter microorganisms that allow monitoring of the spatiotemporal response over the plant in relation to the signal from the genetically encoded sensor. Among these we can find GFP-coding viral genome or GFP-tagged infectious viral particles such as plum pox virus [89], potato virus X [90], cowpea mosaic virus [91] or potato virus Y [92]. Several FP-tagged parasitic and (endo)symbiotic bacteria have also been developed, while GUS reporter must be used cautiously because some microorganisms show strong GUS or GUS-like background activity [93,94]. However, the use of lacZ-labelled Rhizobium leguminosarum to study infection and nodule development in legumes [95] and the efficiency of luminescent Ralstonia solanacearum reporter [96] as a tool to assist potato breeding programs have been recently published. In the case of bacteria, an improved variant of self-assembling split super-folder green fluorescent protein system was optimized to investigate the spatiotemporal dynamics of effectors delivered by the bacterial type III secretion system into the plant cells [97]. Moreover, some FP-tagged fungi are also available, such as Magnaporthe oryzae [98,99], Fusarium graminearum [100] and Fusarium solani [101]. The availability of reporter-incorporated microorganisms is useful for studying biofilm formation, to follow plant-host interaction and, in case of fungi, it enables imaging of hyphae formation, from its passage through the apoplast to its entry into plant cells. Combined with multiple genetically encoded sensors, this approach would allow us to follow the plant response at the site of the interaction with spatiotemporal resolution.
4. Responses of the Cultivated: Biosensors in Crops
While A. thaliana serves as the playground where functionality of biosensors developed for animalia kingdom are usually first used, crops seem left behind, as only the most used examples reach them after a delay. The reason for this frequently lies in unsuccessful attempts to produce functionally stable transformants. Stable transformations of some plant species are demanding due to the larger genome and higher ploidy, higher content of repetitive regions and topologically associated domains (TADs), local intra-chromosomal contacts that are species-specific [102]. Added to the challenge of transforming crops is the fact that most can be unresponsive to tissue culture protocols and they have longer growing seasons. Some crops lack (fertile) seed production which also makes it harder to produce crossed lines that contain two or more stacked transgenes. Consequently, the analyses of responses of crops to various stimuli mostly depend on transcriptomics studies while the confirmation of results using biosensors is limited to the transient transformation of Nicotiana spp. or A. thaliana protoplasts, which can provide much faster but less accurate results. However, this restricts the span of the observable genes to those that have orthologues in further-related species while those with different functions often fail to be more closely observed in their native species.
Despite the limitations, there are some crops available with biosensory properties (Table 1). Ideally, the use of stable transformants is the most desirable approach but it is time consuming and frequently unsuccessful. In some species, mainly in the family of the legumes (but also many other species, e.g. barley [103]), Agrobacterium rhizogenes-mediated transformation of roots provides an alternative approach [104]. These can suffice for following various aspects of root development, interaction with microbiota and nodulation.

Table 1.
Examples of genetically encoded biosensors applied to crops. Comments are added to those biosensors that were used in multiparameter imaging.
Usually, the sensors that have been applied to crops are the widely used transcriptional reporters for cytokinin or auxin responses consisting of a synthetic promoter and a reporter. In contrast, from the wide range of remaining available sensors, to our knowledge, only the ones that enable monitoring Ca2+, ROS and cell division have been applied to a reduced number of plant species other than A. thaliana (Table 1).
Interestingly, multiparameter imaging has already been used in roots of legumes to follow relations between auxin and cytokinin signaling during root growth and nodulation. Fisher et al. assembled a multigene cassette carrying the GFP under the control of synthetic promoter DR5 and the tandem-dimer Tomato (tdTomato) under the control of the TCSn promoter, which enabled them to determine auxin and cytokinin response and their ratios in root and nodule tissues of soybean [125]. Similarly, Nadzieja et al. monitored auxin and cytokinin response of Lotus japonica roots, transformed with DR5::mCherry-NLS and TCSn::YFP-NLS sensors, expressed from the same multigene cassette [35]. To detect the spatial correlation between inoculation with symbiotic bacteria Mesorhizobium loti and cytokinin or auxin response, DsRed-marked bacteria were applied to roots of L. japonicus expressing either TCSn::YFP-NLS [131] or DR5::GUS [116] biosensor, respectively. DR5::tdTomato biosensor was co-expressed in soybean with GFP under constitutive promoter super ubiquitin (sUbi::GFP). Spatial overlap of the signals from FPs enabled discrimination between the red signal from the sensor and bright red autofluorescence [126] (Table 1).
5. Challenges of Plant Biosensors’ Development and Use
There are many successful reported uses of biosensors, though their development can be challenging and obtained results are not always straightforward to interpret. While the vast majority of biosensor detection has been performed in plant roots, protoplasts and transiently transformed tobacco pavement cells, imaging of photosynthetic tissue is far less commonly reported. It is often limited to whole plant imaging which does not allow high spatial resolution, while leaf tissue close-ups with subcellular resolution are rare. Besides some general instrumentation constraints, for instance focus drift and unstable laser power in the first couple of hours, uneven illumination and time-consuming high-quality image acquisition [134], imaging of FPs in plants is affected by fluorescent compounds found in cuticle, cell walls, plastids and vacuoles, such as lignin, chlorophyll and other pigments, etc. [135]. These components generate high background and cause low SNR in detection of fluorescent reporters. Apart from that, they can mislead our interpretation of subcellular organization and structures. For example, during their imaging of GFP-tagged plasmodesmata, Liu et al. reported strong reflection from the cell wall, which could be mistakenly interpreted as plasmodesmata structure in both of the neighboring cells [136]. They were able to avoid misinterpretation due to gene gun transformation with lower density of transformed cells compared to agrobacteria-mediated transformation. As microscopy is very time consuming it is in some cases not the method of choice, especially in the first phases of biosensor development, such as optimization of promoter sequences. Microplate reader fluorimetry/luminometry enables fast screening of many biological replicates simultaneously and detection of crucial time points for further more detailed observation [137,138], but lacks spatial resolution.
To follow a plant’s dynamic response through space and time with high resolution, the problem of SNR should be addressed. Partly, we can achieve optimal SNR with proper image acquisition and processing, and also region of interest (ROI) selection [134,139], but this is not always sufficient. There are different strategies to overcome the problem of low SNR at the level of biosensor construction. One option aims to overpower autofluorescence with high expression of fluorescent proteins. This is commonly achieved with the help of minimal CaMV 35S promoter. For transcriptional reporters the promoter can be coupled with CaMV 35S enhancer region and two or more repeats of inducible parts of a certain promoter, e.g. PR2 (from parsley) or AtCMPG1 [140,141]. Higher expression can also be achieved with the use of proper combinations of 5’-UTR and 3’-UTR enhancer regions [142], compatible with the plant of choice [143] as it is done for protein production in planta. When a transcriptional reporter is under the control of a weak promoter, an additional transcriptional regulator between the two can amplify its activity [144]. However, this effort is frequently opposed by silencing [137].
The proper choice of localization can also heighten SNR. The signal in plant cytoplasm is weak, variable among cells, hard to quantify and more prone to silencing. On the other hand, localization in other compartments can also have some constraints. Reporter protein localized in vacuole and apoplast should be pH stable, a condition refusing many widely used fluorescent proteins [145]. For this reason, red variants of redox sensors roGFP and HyPer were designed, namely Grx1-roCherry [146] and HyPerRed [147], respectively. Additionally, HyPer and HyPerRed have their redox-non-sensitive and pH-sensitive counterparts and can be used in parallel as a control of pH effect on measurements [147]. However, we have not found any report on their use in plants so far. Nucleus and mitochondrial localizations also have drawbacks. For example, small fluorescent proteins can exhibit leaking when tagged with either nuclear localization signal (NLS) or nuclear exportation signal (NES), while the mutants expressing fluorescent biosensors in the mitochondrial matrix were shown to grow more slowly than others [29].
Higher analyte specificity is also helpful when dealing with low SNR. This can be reached with a chimeric effector/detector module [148]. This often comes with lower constant of dissociation for the chosen analyte and can therefore interfere with the analyte’s availability for endogenous targets, which was the case in ABA sensors (reviewed in Isoda et al. [13]). Potential slow release of the analyte from the binding pocket of direct biosensors must also be considered to avoid misinterpretations of temporal dimension of the analyte availability in vivo [67].
Lower SNR in transformants after agrobacteria-mediated transient transformations can be caused by the expression of reporter proteins in agrobacteria, which can be overcome by the promoter exchange [149] or the insertion of an intron [150], the last resulting in higher expression in plant cells through intron-mediated enhancement [151].
Another common problem affecting the use of biosensors is their capacity for analyte quantification. Cell responses are not binary, but rather pattern- and concentration-dependent, so the need for biosensors enabling dynamic quantitative imaging arises, especially when high spatiotemporal resolution and quantitative data are needed for systems biology approaches [152]. Aequorin is a perfect example of absolute intensiometric biosensor for Ca2+ imaging as it enables measurement of absolute in vivo concentration [153]. Non-FRET ratiometric biosensors use a duet of reporter proteins, one as a reporter of analyte and another as a reporter of expression in a certain cell or tissue. The second can be expressed separately as a normalization transcriptional unit under constitutive promoter. Expression under strong viral constitutive promoter CaMV 35S can experience silencing or patterns in dividing cells [7] and is therefore often exchanged with plant constitutive promoters, such as rice actin and maize ubiquitin promoters. Still, variable expression in different species forces the exploration of novel options [154,155]. On the other hand, it is possible to opt for separation of two or more reporters at the protein stage. One possibility is co-translational separation by a self-cleaving peptide [84,156]. However, after the cleavage, the peptide is not excised but remains attached to the C-terminus of the upstream protein sequence which can affect its folding or function. This can be overcome by attachment of a peptide linker, which is a target of endogenous peptidases [157,158], or mini-intein with N-terminal autocleavage ability [159]. Synchronized expression of two proteins can also be obtained by using a polyprotein vector system that is based on a pair of self-excising mini-inteins, called dual-intein domain, which allow the release of both proteins (shown for tripartite sfGFP) [160]. In contrast, the use of internal ribosomal entry site (IRES) was not successful and is not recommended, as the level of IRES-driven translation can vary among cells [160].
To alleviate the problems associated with fluorescence imaging in plants, one can also use luciferases as the reporter domain in biosensor. One of the drawbacks of luciferases is the need for external application of the substrate which might not readily penetrate into plant tissue. To overcome this issue, autoluminescent N. benthamiana plants were engineered by the insertion of a fungal bioluminescence gene cluster (all with CaMV 35S promoter) [161]. Thus, the transgenic plants do not need any external substrate addition, as they produce fungal luciferin from caffeic acid. Treatment with methyl jasmonate, ethylene and wounding caused higher luminescence within seconds [161,162]. However, the use of the plant’s metabolite impacts the final luminescence produced according to availability. For example, older leaves showed lower luminescence [161,162]. A similar approach, avoiding exogenous substrate application, was explored in the reporter RUBY which produces red betalain pigment. The three enzymes that cooperate in its biosynthetic pathway from tyrosine were expressed under the control of various promoters in A. thaliana and were co-translationally separated due to the addition of 2A self-cleaving peptide [163].
6. Concluding Remarks
Biosensors have become indispensable tools to gain new insights in molecular biology with high spatiotemporal resolution. When being transferred to plants, especially crops, the community experiences challenges. However, overcoming these challenges is more and more supported by new achievements in synthetic biology, imaging and plant transformation fields, and can lead to new discoveries. Biosensors have so far been used individually, tracking only one analyte per experiment, with rare exceptions. We believe that the field of biosensors is now ready for multiparameter imaging. Therefore, this approach should now be used to obtain (quantitative) data with high spatiotemporal resolution that offer high-quality input for mathematical modeling of the dynamic network of plant responses to environment. Biosensors are promising tools to uncover the mysteries of a plant’s orchestrated signaling network that leads to discrimination between specific immune responses.
Author Contributions
All authors together conceptualized and wrote the manuscript. K.G. and A.C. acquired funding. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Slovenian Research Agency (research core funding No. P4-0165, projects J4-1777, J1-2467 and 1000-20-0105) and by the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 862858.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We wish to apologize to those colleagues whose work could not be cited due to space limitations.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Walia, A.; Waadt, R.; Jones, A.M. Genetically encoded biosensors in plants: Pathways to discovery. Annu. Rev. Plant Biol. 2018, 69, 497–524. [Google Scholar] [CrossRef]
- Sadanandom, A.; Napier, R.M. Biosensors in plants. Curr. Opin. Plant Biol. 2010, 13, 736–743. [Google Scholar] [CrossRef]
- Miyawaki, A.; Llopis, J.; Heim, R.; McCaffery, J.M.; Adams, J.A.; Ikura, M.; Tsien, R.Y.; Michael McCaffery, J.; Adams, J.A.; Ikura, M.; et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 1997, 388, 882–887. [Google Scholar] [CrossRef]
- Brunoud, G.; Wells, D.M.; Oliva, M.; Larrieu, A.; Mirabet, V.; Burrow, A.H.; Beeckman, T.; Kepinski, S.; Traas, J.; Bennett, M.J.; et al. A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 2012, 482, 103–106. [Google Scholar] [CrossRef]
- Ali, S.; Kim, W.C. A Fruitful Decade Using Synthetic Promoters in the Improvement of Transgenic Plants. Front. Plant Sci. 2019, 10, 1433. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Murfett, J.; Hagen, G.; Guilfoyle, T.J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 1997, 9, 1963–1971. [Google Scholar] [CrossRef]
- Liao, C.Y.; Smet, W.; Brunoud, G.; Yoshida, S.; Vernoux, T.; Weijers, D. Reporters for sensitive and quantitative measurement of auxin response. Nat. Methods 2015, 12, 207–210. [Google Scholar] [CrossRef]
- Li, W.; Ma, M.; Feng, Y.; Li, H.; Wang, Y.; Ma, Y.; Li, M.; An, F.; Guo, H. EIN2-directed translational regulation of ethylene signaling in Arabidopsis. Cell 2015, 163, 670–683. [Google Scholar] [CrossRef]
- Merchante, C.; Brumos, J.; Yun, J.; Hu, Q.; Spencer, K.R.; Enríquez, P.; Binder, B.M.; Heber, S.; Stepanova, A.N.; Alonso, J.M. Gene-specific translation regulation mediated by the hormone-signaling molecule EIN2. Cell 2015, 163, 684–697. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Moreno, J.P.; Stepanova, A.N. Monitoring ethylene in plants: Genetically encoded reporters and biosensors. Small Methods 2020, 4, 1900260. [Google Scholar] [CrossRef]
- Novák, O.; Napier, R.; Ljung, K. Zooming in on plant hormone analysis: Tissue- and cell-specific approaches. Annu. Rev. Plant Biol. 2017, 68, 323–348. [Google Scholar] [CrossRef] [PubMed]
- Waadt, R. Phytohormone signaling mechanisms and genetic methods for their modulation and detection. Curr. Opin. Plant Biol. 2020, 57, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Isoda, R.; Yoshinari, A.; Ishikawa, Y.; Sadoine, M.; Simon, R.; Frommer, W.B.; Nakamura, M. Sensors for the quantification, localization and analysis of the dynamics of plant hormones. Plant J. 2020. [Google Scholar] [CrossRef]
- Martin-Arevalillo, R.; Vernoux, T. Shining light on plant hormones with genetically encoded biosensors. Biol. Chem. 2019, 400, 477–486. [Google Scholar] [CrossRef]
- Okumoto, S.; Jones, A.; Frommer, W.B. Quantitative imaging with fluorescent biosensors. Annu. Rev. Plant Biol. 2012, 63, 663–706. [Google Scholar] [CrossRef]
- Jones, A.M. A new look at stress: Abscisic acid patterns and dynamics at high-resolution. New Phytol. 2016, 210, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Pařízková, B.; Pernisová, M.; Novák, O. What has been seen cannot be unseen—Detecting auxin in vivo. Int. J. Mol. Sci. 2017, 18, 2736. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; Navazio, L.; Szabo, I. The contribution of organelles to plant intracellular calcium signalling. J. Exp. Bot. 2018, 69, 4175–4193. [Google Scholar] [CrossRef] [PubMed]
- Rizza, A.; Jones, A.M. The makings of a gradient: Spatiotemporal distribution of gibberellins in plant development. Curr. Opin. Plant Biol. 2019, 47, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Villasante, C.; Burén, S.; Barón-Sola, Á.; Martínez, F.; Hernández, L.E. In vivo ROS and redox potential fluorescent detection in plants: Present approaches and future perspectives. Methods 2016, 109, 92–104. [Google Scholar] [CrossRef] [PubMed]
- Komis, G.; Novák, D.; Ovečka, M.; Šamajová, O.; Šamaj, J. Advances in imaging plant cell dynamics. Plant Physiol. 2018, 176, 80–93. [Google Scholar] [CrossRef]
- Greenwald, E.; Zhang, J. Fluorescent Biosensor Database. Available online: https://biosensordb.ucsd.edu/index.php (accessed on 20 November 2020).
- Pieterse, C.M.J.; Leon-Reyes, A.; Van Der Ent, S.; Van Wees, S.C.M. Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 2009, 5, 308–316. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Van der Does, D.; Zamioudis, C.; Leon-Reyes, A.; Van Wees, S.C.M. Hormonal modulation of plant immunity. Annu. Rev. Cell Dev. Biol. 2012, 28, 489–521. [Google Scholar] [CrossRef]
- Mandadi, K.K.; Scholthof, K.-B.G. Plant immune responses against viruses: How does a virus cause disease? Plant Cell 2013, 25, 1489–1505. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D.G. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef]
- Kelner, A.; Leitão, N.; Chabaud, M.; Charpentier, M.; de Carvalho-Niebel, F. Dual Color Sensors for Simultaneous Analysis of Calcium Signal Dynamics in the Nuclear and Cytoplasmic Compartments of Plant Cells. Front. Plant Sci. 2018, 9, 245. [Google Scholar] [CrossRef] [PubMed]
- Nietzel, T.; Elsässer, M.; Ruberti, C.; Steinbeck, J.; Ugalde, J.M.; Fuchs, P.; Wagner, S.; Ostermann, L.; Moseler, A.; Lemke, P.; et al. The fluorescent protein sensor roGFP2-Orp1 monitors in vivo H2O2 and thiol redox integration and elucidates intracellular H2O2 dynamics during elicitor-induced oxidative burst in Arabidopsis. New Phytol. 2019, 221, 1649–1664. [Google Scholar] [CrossRef] [PubMed]
- Aller, I.; Rouhier, N.; Meyer, A.J. Development of roGFP2-derived redox probes for measurement of the glutathione redox potential in the cytosol of severely glutathione-deficient rml1 seedlings. Front. Plant Sci. 2013, 4, 506. [Google Scholar] [CrossRef] [PubMed]
- Ugalde, J.M.; Fuchs, P.; Nietzel, T.; Cutolo, E.A.; Homagk, M.; Vothknecht, U.C.; Holuigue, L.; Schwarzländer, M.; Müller-Schüssele, S.J.; Meyer, A.J. Chloroplast-derived photo-oxidative stress causes changes in H2O2 and GSH in other subcellular compartments. Plant Physiol. 2021, kiaa095. [Google Scholar] [CrossRef]
- Huang, W.E.; Huang, L.; Preston, G.M.; Naylor, M.; Carr, J.P.; Li, Y.; Singer, A.C.; Whiteley, A.S.; Wang, H. Quantitative in situ assay of salicylic acid in tobacco leaves using a genetically modified biosensor strain of Acinetobacter sp. ADP1. Plant J. 2006, 46, 1073–1083. [Google Scholar] [CrossRef] [PubMed]
- Löfke, C.; Zwiewka, M.; Heilmann, I.; Van Montagu, M.C.E.; Teichmann, T.; Friml, J. Asymmetric gibberellin signaling regulates vacuolar trafficking of PIN auxin transporters during root gravitropism. Proc. Natl. Acad. Sci. USA 2013, 110, 3627–3632. [Google Scholar] [CrossRef] [PubMed]
- Rizza, A.; Walia, A.; Lanquar, V.; Frommer, W.B.; Jones, A.M. In vivo gibberellin gradients visualized in rapidly elongating tissues. Nat. Plants 2017, 3, 803–813. [Google Scholar] [CrossRef] [PubMed]
- Nadzieja, M.; Stougaard, J.; Reid, D. A toolkit for high resolution imaging of cell division and phytohormone signaling in legume roots and root nodules. Front. Plant Sci. 2019, 10, 1000. [Google Scholar] [CrossRef]
- Seitz, K.; Krysan, P.J. Expanding the toolkit of fluorescent biosensors for studying mitogen activated protein kinases in plants. Int. J. Mol. Sci. 2020, 21, 5350. [Google Scholar] [CrossRef]
- Lukan, T.; Pompe-Novak, M.; Baebler, Š.; Tušek-Žnidarič, M.; Kladnik, A.; Križnik, M.; Blejec, A.; Zagorščak, M.; Stare, K.; Dušak, B.; et al. Precision transcriptomics of viral foci reveals the spatial regulation of immune-signaling genes and identifies RBOHD as an important player in the incompatible interaction between potato virus Y and potato. Plant J. 2020, 104, 645–661. [Google Scholar] [CrossRef]
- Yuan, P.; Jauregui, E.; Du, L.; Tanaka, K.; Poovaiah, B.W. Calcium signatures and signaling events orchestrate plant–microbe interactions. Curr. Opin. Plant Biol. 2017, 38, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Seybold, H.; Trempel, F.; Ranf, S.; Scheel, D.; Romeis, T.; Lee, J. Ca2+ signalling in plant immune response: From pattern recognition receptors to Ca2+ decoding mechanisms. New Phytol. 2014, 204, 782–790. [Google Scholar] [CrossRef]
- Knight, M.R.; Campbell, A.K.; Smith, S.M.; Trewavas, A.J. Transgenic plant aequorin reports the effects of touch and cold-shock and elicitors on cytoplasmic calcium. Nature 1991, 352, 524–526. [Google Scholar] [CrossRef] [PubMed]
- Allen, G.J.; Kwak, J.M.; Chu, S.P.; Llopis, J.; Tsien, R.Y.; Harper, J.F.; Schroeder, J.I. Cameleon calcium indicator reports cytoplasmic calcium dynamics in Arabidopsis guard cells. Plant J. 1999, 19, 735–747. [Google Scholar] [CrossRef]
- Nakai, J.; Ohkura, M.; Imoto, K. A high signal-to-noise Ca2+ probe composed of a single green fluorescent protein. Nat. Biotechnol. 2001, 19, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Xiong, T.C.; Ronzier, E.; Sanchez, F.; Corratgé-Faillie, C.; Mazars, C.; Thibaud, J.B. Imaging long distance propagating calcium signals in intact plant leaves with the BRET-based GFP-aequorin reporter. Front. Plant Sci. 2014, 5, 43. [Google Scholar] [CrossRef]
- Toyota, M.; Spencer, D.; Sawai-Toyota, S.; Jiaqi, W.; Zhang, T.; Koo, A.J.; Howe, G.A.; Gilroy, S. Glutamate triggers long-distance, calcium-based plant defense signaling. Science 2018, 361, 1112–1115. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Vásquez, A.; Salinas, P.; Holuigue, L. Salicylic acid and reactive oxygen species interplay in the transcriptional control of defense genes expression. Front. Plant Sci. 2015, 6, 171. [Google Scholar] [CrossRef] [PubMed]
- Wrzaczek, M.; Brosché, M.; Kangasjärvi, J. ROS signaling loops—Production, perception, regulation. Curr. Opin. Plant Biol. 2013, 16, 575–582. [Google Scholar] [CrossRef]
- Choi, W.G.; Swanson, S.J.; Gilroy, S. High-resolution imaging of Ca2+, redox status, ROS and pH using GFP biosensors. Plant J. 2012, 70, 118–128. [Google Scholar] [CrossRef]
- Hanson, G.T.; Aggeler, R.; Oglesbee, D.; Cannon, M.; Capaldi, R.A.; Tsien, R.Y.; Remington, S.J. Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J. Biol. Chem. 2004, 279, 13044–13053. [Google Scholar] [CrossRef]
- Jiang, K.; Schwarzer, C.; Lally, E.; Zhang, S.; Ruzin, S.; Machen, T.; Remington, S.J.; Feldman, L. Expression and characterization of a redox-sensing green fluorescent protein (reduction-oxidation-sensitive green fluorescent protein) in Arabidopsis. Plant Physiol. 2006, 141, 397–403. [Google Scholar] [CrossRef]
- Stonebloom, S.; Brunkard, J.O.; Cheung, A.C.; Jiang, K.; Feldman, L.; Zambryski, P. Redox states of plastids and mitochondria differentially regulate intercellular transport via plasmodesmata. Plant Physiol. 2012, 158, 190–199. [Google Scholar] [CrossRef]
- Schwarzländer, M.; Fricker, M.D.; Sweetlove, L.J. Monitoring the in vivo redox state of plant mitochondria: Effect of respiratory inhibitors, abiotic stress and assessment of recovery from oxidative challenge. Biochim. Biophys. Acta Bioenerg. 2009, 1787, 468–475. [Google Scholar] [CrossRef] [PubMed]
- Meyer, A.J.; Brach, T.; Marty, L.; Kreye, S.; Rouhier, N.; Jacquot, J.P.; Hell, R. Redox-sensitive GFP in Arabidopsis thaliana is a quantitative biosensor for the redox potential of the cellular glutathione redox buffer. Plant J. 2007, 52, 973–986. [Google Scholar] [CrossRef]
- Hernández-Barrera, A.; Quinto, C.; Johnson, E.A.; Wu, H.M.; Cheung, A.Y.; Cárdenas, L. Using hyper as a molecular probe to visualize hydrogen peroxide in living plant cells: A method with virtually unlimited potential in plant biology. Methods Enzymol. 2013, 527, 275–290. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, K.; Yokochi, Y.; Fu, N.; Fukaya, Y.; Yoshida, K.; Mihara, S.; Hisabori, T. The thioredoxin (Trx) redox state sensor protein can visualize Trx activities in the light/dark response in chloroplasts. J. Biol. Chem. 2019, 294, 12091–12098. [Google Scholar] [CrossRef]
- Uslu, V.V.; Grossmann, G. The biosensor toolbox for plant developmental biology. Curr. Opin. Plant Biol. 2016, 29, 138–147. [Google Scholar] [CrossRef]
- Mylle, E.; Codreanu, M.C.; Boruc, J.; Russinova, E. Emission spectra profiling of fluorescent proteins in living plant cells. Plant Methods 2013, 9, 10. [Google Scholar] [CrossRef] [PubMed]
- Muller, B.; Sheen, J. Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 2008, 453, 1094–1097. [Google Scholar] [CrossRef]
- Zurcher, E.; Tavor-Deslex, D.; Lituiev, D.; Enkerli, K.; Tarr, P.T.; Muller, B. A robust and sensitive synthetic sensor to monitor the transcriptional output of the cytokinin signaling network in planta. Plant Physiol. 2013, 161, 1066–1075. [Google Scholar] [CrossRef] [PubMed]
- Steiner, E.; Livne, S.; Kobinson-Katz, T.; Tal, L.; Pri-tal, O.; Mosquna, A.; Tarkowská, D.; Muller, B.; Tarkowski, P.; Weiss, D. The putative O-linked N-acetylglucosamine transferase SPINDLY inhibits class I TCP proteolysis to promote sensitivity to cytokinin 1. Plant Physiol. 2016, 171, 1485–1494. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Duan, L.; Pruneda-Paz, J.L.; Oh, D.H.; Pound, M.; Kay, S.; Dinneny, J.R. The 6xABRE synthetic promoter enables the spatiotemporal analysis of ABA-mediated transcriptional regulation. Plant Physiol. 2018, 177, 1650–1665. [Google Scholar] [CrossRef]
- Stepanova, A.N.; Yun, J.; Likhacheva, A.V.; Alonso, J.M. Multilevel interactions between ethylene and auxin in Arabidopsis roots. Plant Cell 2007, 19, 2169–2185. [Google Scholar] [CrossRef]
- Lehmann, S.; Dominguez-Ferreras, A.; Huang, W.J.; Denby, K.; Ntoukakis, V.; Schäfer, P. Novel markers for high-throughput protoplast-based analyses of phytohormone signaling. PLoS ONE 2020, 15, e0234154. [Google Scholar] [CrossRef]
- Jones, A.M.; Danielson, J.Å.H.; Manojkumar, S.N.; Lanquar, V.; Grossmann, G.; Frommer, W.B. Abscisic acid dynamics in roots detected with genetically encoded FRET sensors. Elife 2014, e01741. [Google Scholar] [CrossRef]
- Waadt, R.; Hitomi, K.; Nishimura, N.; Hitomi, C.; Adams, S.R.; Getzoff, E.D.; Schroeder, J.I. FRET-based reporters for the direct visualization of abscisic acid concentration changes and distribution in Arabidopsis. Elife 2014, e01739. [Google Scholar] [CrossRef]
- Larrieu, A.; Champion, A.; Legrand, J.; Lavenus, J.; Mast, D.; Brunoud, G.; Oh, J.; Guyomarc’h, S.; Pizot, M.; Farmer, E.E.; et al. A fluorescent hormone biosensor reveals the dynamics of jasmonate signalling in plants. Nat. Commun. 2015, 6, 6043. [Google Scholar] [CrossRef] [PubMed]
- Samodelov, S.L.; Beyer, H.M.; Guo, X.; Augustin, M.; Jia, K.P.; Baz, L.; Ebenhöh, O.; Beyer, P.; Weber, W.; Al-Babili, S.; et al. StrigoQuant: A genetically encoded biosensor for quantifying strigolactone activity and specificity. Sci. Adv. 2016, 2, e1601266. [Google Scholar] [CrossRef] [PubMed]
- Rizza, A.; Walia, A.; Tang, B.; Jones, A.M. Visualizing cellular gibberellin levels using the nlsGPS1 Förster resonance energy transfer (FRET) biosensor. J. Vis. Exp. 2019, e58739. [Google Scholar] [CrossRef]
- Zaman, N.; Seitz, K.; Kabir, M.; George-Schreder, L.S.; Shepstone, I.; Liu, Y.; Zhang, S.; Krysan, P.J. A Förster resonance energy transfer sensor for live-cell imaging of mitogen-activated protein kinase activity in Arabidopsis. Plant J. 2019, 97, 970–983. [Google Scholar] [CrossRef]
- Zhang, L.; Takahashi, Y.; Hsu, P.K.; Kollist, H.; Merilo, E.; Krysan, P.J.; Schroeder, J.I. FRET kinase sensor development reveals SnRK2/OST1 activation by ABA but not by MeJA and high CO2 during stomatal closure. Elife 2020, 9, e56351. [Google Scholar] [CrossRef] [PubMed]
- Sozzani, R.; Cui, H.; Moreno-Risueno, M.A.; Busch, W.; Van Norman, J.M.; Vernoux, T.; Brady, S.M.; Dewitte, W.; Murray, J.A.H.; Benfey, P.N. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 2010, 466, 128–132. [Google Scholar] [CrossRef]
- Desvoyes, B.; Arana-Echarri, A.; Barea, M.D.; Gutierrez, C. A comprehensive fluorescent sensor for spatiotemporal cell cycle analysis in Arabidopsis. Nat. Plants 2020, 6, 1330–1334. [Google Scholar] [CrossRef]
- Zhang, Q.; Schepis, A.; Huang, H.; Yang, J.; Ma, W.; Torra, J.; Zhang, S.Q.; Yang, L.; Wu, H.; Nonell, S.; et al. Designing a green fluorogenic protease reporter by flipping a beta strand of GFP for imaging apoptosis in animals. J. Am. Chem. Soc. 2019, 141, 4526–4530. [Google Scholar] [CrossRef]
- Gruden, K.; Lidoy, J.; Petek, M.; Podpečan, V.; Flors, V.; Papadopoulou, K.K.; Pappas, M.L.; Martinez-Medina, A.; Bejarano, E.; Biere, A.; et al. Ménage à trois: Unraveling the mechanisms regulating plant–microbe–arthropod interactions. Trends Plant Sci. 2020, 25, 1215–1226. [Google Scholar] [CrossRef]
- Devireddy, A.R.; Zandalinas, S.I.; Fichman, Y.; Mittler, R. Integration of reactive oxygen species and hormone signaling during abiotic stress. Plant J. 2020. [Google Scholar] [CrossRef]
- Waadt, R.; Krebs, M.; Kudla, J.; Schumacher, K. Multiparameter imaging of calcium and abscisic acid and high-resolution quantitative calcium measurements using R-GECO1-mTurquoise in Arabidopsis. New Phytol. 2017, 216, 303–320. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Araki, S.; Wu, J.; Tramoto, T.; Chang, Y.-F.; Nakano, M.; Abderfattah, A.; Fujiwara, M.; Ishihara, T.; Nagai, T.; et al. An expanded palette of genetically encoded Ca2+ indicators. Science 2011, 557, 1888–1891. [Google Scholar] [CrossRef]
- Nakamura, A.; Higuchi, K.; Goda, H.; Fujiwara, M.T.; Sawa, S.; Koshiba, T.; Shimada, Y.; Yoshida, S. Brassinolide induces IAA5, IAA19, and DR5, a synthetic auxin response element in Arabidopsis, implying a cross talk point of brassinosteroid and auxin signaling. Plant Physiol. 2003, 133, 1843–1853. [Google Scholar] [CrossRef]
- Zhu, M.; Chen, W.; Mirabet, V.; Hong, L.; Bovio, S.; Strauss, S.; Schwarz, E.M.; Tsugawa, S.; Wang, Z.; Smith, R.S.; et al. Robust organ size requires robust timing of initiation orchestrated by focused auxin and cytokinin signalling. Nat. Plants 2020, 6, 686–698. [Google Scholar] [CrossRef] [PubMed]
- Behera, S.; Wang, N.; Zhang, C.; Schmitz-Thom, I.; Strohkamp, S.; Schültke, S.; Hashimoto, K.; Xiong, L.; Kudla, J. Analyses of Ca2+ dynamics using a ubiquitin-10 promoter-driven Yellow Cameleon 3.6 indicator reveal reliable transgene expression and differences in cytoplasmic Ca2+ responses in Arabidopsis and rice (Oryza sativa) roots. New Phytol. 2015, 206, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Denay, G.; Schultz, P.; Hänsch, S.; Weidtkamp-Peters, S.; Simon, R. Over the rainbow: A practical guide for fluorescent protein selection in plant FRET experiments. Plant Direct 2019, 3, e00189. [Google Scholar] [CrossRef]
- Waadt, R.; Köster, P.; Andrés, Z.; Waadt, C.; Bradamante, G.; Lampou, K.; Kudla, J.; Schumacher, K. Dual-reporting transcriptionally linked genetically encoded fluorescent indicators resolve the spatiotemporal coordination of cytosolic abscisic acid and second messenger dynamics in Arabidopsis. Plant Cell 2020, 32, 2582–2601. [Google Scholar] [CrossRef]
- Xu, X.; Soutto, M.; Xie, Q.; Servick, S.; Subramanian, C.; Von Arnim, A.G.; Johnson, C.H. Imaging protein interactions with bioluminescence resonance energy transfer (BRET) in plant and mammalian cells and tissues. Proc. Natl. Acad. Sci. USA 2007, 104, 10264–10269. [Google Scholar] [CrossRef]
- Moyle, R.L.; Carvalhais, L.C.; Pretorius, L.-S.; Nowak, E.; Subramaniam, G.; Dalton-Morgan, J.; Schenk, P.M. An optimized transient dual luciferase assay for quantifying microRNA directed repression of targeted sequences. Front. Plant Sci. 2017, 8, 1631. [Google Scholar] [CrossRef]
- Khosla, A.; Rodriguez-Furlan, C.; Kapoor, S.; Van Norman, J.M.; Nelson, D.C. A series of dual-reporter vectors for ratiometric analysis of protein abundance in plants. Plant Direct 2020, 4, e00231. [Google Scholar] [CrossRef]
- Urquiza-García, U.; Millar, A.J. Expanding the bioluminescent reporter toolkit for plant science with NanoLUC. Plant Methods 2019, 15, 68. [Google Scholar] [CrossRef]
- Furuhata, Y.; Sakai, A.; Murakami, T.; Nagasaki, A.; Kato, Y. Bioluminescent imaging of Arabidopsis thaliana using an enhanced Nano-lantern luminescence reporter system. PLoS ONE 2020, 15, e0227477. [Google Scholar] [CrossRef] [PubMed]
- Lew, T.T.S.; Koman, V.B.; Silmore, K.S.; Seo, J.S.; Gordiichuk, P.; Kwak, S.Y.; Park, M.; Ang, M.C.Y.; Khong, D.T.; Lee, M.A.; et al. Real-time detection of wound-induced H2O2 signalling waves in plants with optical nanosensors. Nat. Plants 2020, 6, 404–415. [Google Scholar] [CrossRef] [PubMed]
- Fichman, Y.; Miller, G.; Mittler, R. Whole-plant live imaging of reactive oxygen species. Mol. Plant 2019, 12, 1203–1210. [Google Scholar] [CrossRef]
- Pasin, F.; Kulasekaran, S.; Natale, P.; Simón-Mateo, C.; García, J.A. Rapid fluorescent reporter quantification by leaf disc analysis and its application in plant-virus studies. Plant Methods 2014, 10, 22. [Google Scholar] [CrossRef] [PubMed]
- Oparka, K.J.; Roberts, A.G.; Cruz, S.S.; Boevink, P.; Prior, D.A.M.; Smallcombe, A.; Oparka, K.J.; Roberts, A.G.; Cruz, S.S.; Boevink, P.; et al. Using GFP to study virus invasion and spread in plant tissues. Nature 1997, 388, 401–402. [Google Scholar] [CrossRef]
- Verver, J.; Wellink, J.; Van Lent, J.; Gopinath, K.; Van Kammen, A. Studies on the movement of cowpea mosaic virus using the jellyfish green fluorescent protein. Virology 1998, 242, 22–27. [Google Scholar] [CrossRef]
- Rupar, M.; Faurez, F.; Tribodet, M.; Gutiérrez-Aguirre, I.; Delaunay, A.; Glais, L.; Kriznik, M.; Dobnik, D.; Gruden, K.; Jacquot, E.; et al. Fluorescently tagged Potato Virus Y: A versatile tool for functional analysis of plant-virus interactions. Mol. Plant Microbe Interact. 2015, 28, 739–750. [Google Scholar] [CrossRef]
- Egener, T.; Hurek, T.; Reinhold-Hurek, B. Use of green fluorescent protein to detect expression of nif genes of Azoarcus sp. BH72, a grass-associated diazotroph, on rice roots. Mol. Plant Microbe Interact. 1998, 11, 71–75. [Google Scholar] [CrossRef][Green Version]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: Beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef] [PubMed]
- McAdam, E.L.; Reid, J.B.; Foo, E. Gibberellins promote nodule organogenesis but inhibit the infection stages of nodulation. J. Exp. Bot. 2018, 69, 2117–2130. [Google Scholar] [CrossRef]
- Cruz, A.P.Z.; Ferreira, V.; Pianzzola, M.J.; Siri, M.I.; Coll, N.S.; Valls, M. A novel, sensitive method to evaluate potato germplasm for bacterial wilt resistance using a luminescent Ralstonia solanacearum reporter strain. Mol. Plant Microbe Interact. 2014, 27, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Park, E.; Lee, H.Y.; Woo, J.; Choi, D.; Dinesh-Kumar, S.P. Spatiotemporal monitoring of Pseudomonas syringae effectors via type III secretion using split fluorescent protein fragments. Plant Cell 2017, 29, 1571–1584. [Google Scholar] [CrossRef]
- Campos-Soriano, L.; San Segundo, B. Assessment of blast disease resistance in transgenic PRms rice using a GFP-expressing Magnaporthe oryzae strain. Plant Pathol. 2009, 58, 677–689. [Google Scholar] [CrossRef]
- Jones, K.; Kim, D.W.; Park, J.S.; Khang, C.H. Live-cell fluorescence imaging to investigate the dynamics of plant cell death during infection by the rice blast fungus Magnaporthe oryzae. BMC Plant Biol. 2016, 16, 69. [Google Scholar] [CrossRef]
- Qiu, H.; Zhao, X.; Fang, W.; Wu, H.; Abubakar, Y.S.; Lu, G.; Wang, Z.; Zheng, W. Spatiotemporal nature of Fusarium graminearum-wheat coleoptile interactions. Phytopathol. Res. 2019, 1, 26. [Google Scholar] [CrossRef]
- Skiada, V.; Faccio, A.; Kavroulakis, N.; Genre, A.; Bonfante, P.; Papadopoulou, K.K. Colonization of legumes by an endophytic Fusarium solani strain FsK reveals common features to symbionts or pathogens. Fungal Genet. Biol. 2019, 127, 60–74. [Google Scholar] [CrossRef]
- Dong, P.; Tu, X.; Chu, P.Y.; Lü, P.; Zhu, N.; Grierson, D.; Du, B.; Li, P.; Zhong, S. 3D Chromatin Architecture of Large Plant Genomes Determined by Local A/B Compartments. Mol. Plant 2017, 10, 1497–1509. [Google Scholar] [CrossRef] [PubMed]
- Imani, J.; Li, L.; Schäfer, P.; Kogel, K.-H. STARTS—A stable root transformation system for rapid functional analyses of proteins of the monocot model plant barley. Plant J. 2011, 67, 726–735. [Google Scholar] [CrossRef]
- Chabaud, M.; Boisson-Dernier, A.; Zhang, J.; Taylor, C.G.; Yu, O.; Barker, D.G. Agrobacterium rhizogenes-mediated root transformation. In Medicago Truncatula Handbook; Mathesius, U., Journet, E.P., Sumner, L.W., Eds.; Noble Research Institute: Ardmore, OK, USA, 2006; ISBN 0-9754303-1-9. [Google Scholar]
- Krebs, M.; Held, K.; Binder, A.; Hashimoto, K.; Den Herder, G.; Parniske, M.; Kudla, J.; Schumacher, K. FRET-based genetically encoded sensors allow high-resolution live cell imaging of Ca2+ dynamics. Plant J. 2012, 69, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Sieberer, B.J.; Chabaud, M.; Timmers, A.C.; Monin, A.; Fournier, J.; Barker, D.G. A nuclear-targeted cameleon demonstrates intranuclear Ca2+ spiking in Medicago truncatula root hairs in response to rhizobial nodulation factors. Plant Physiol. 2009, 151, 1197–1206. [Google Scholar] [CrossRef]
- Moroz, N.; Tanaka, K. FlgII-28 is a major flagellin-derived defense elicitor in potato. Mol. Plant Microbe Interact. 2020, 33, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-R.; Xu, Y.-P.; Cai, X.-Z. SlCNGC1 and SlCNGC14 suppress Xanthomonas oryzae pv. oryzicola-induced hypersensitive response and non-host resistance in tomato. Front. Plant Sci. 2018, 9, 285. [Google Scholar] [CrossRef]
- Wang, R.; Li, R.; Xu, T.; Li, T. Optimization of the pollen-tube pathway method of plant transformation using the Yellow Cameleon 3.6 calcium sensor in Solanum lycopersicum. Biologia 2017, 72, 1147–1155. [Google Scholar] [CrossRef]
- Huang, W.J.; Liu, H.K.; McCormick, S.; Tang, W.H. Tomato pistil factor STIG1 promotes in vivo pollen tube growth by binding to phosphatidylinositol 3-phosphate and the extracellular domain of the pollen receptor kinase LePRK2. Plant Cell 2014, 26, 2505–2523. [Google Scholar] [CrossRef]
- Hipsch, M.; Lampl, N.; Zelinger, E.; Barda, O.; Rosenwasser, S. Sensing stress responses in potato with whole-plant redox imaging. bioRxiv 2020. [Google Scholar] [CrossRef]
- Andrio, E.; Marino, D.; Marmeys, A.; de Segonzac, M.D.; Damiani, I.; Genre, A.; Huguet, S.; Frendo, P.; Puppo, A.; Pauly, N. Hydrogen peroxide-regulated genes in the Medicago truncatula—Sinorhizobium meliloti symbiosis. New Phytol. 2013, 198, 179–189. [Google Scholar] [CrossRef]
- O’Connor, D.L.; Elton, S.; Ticchiarelli, F.; Hsia, M.M.; Vogel, J.P.; Leyser, T. Cross-species functional diversity within the PIN auxin efflux protein family. Elife 2017, 6, e31804. [Google Scholar] [CrossRef]
- Tucker, M.R.; Okada, T.; Johnson, S.D.; Takaiwa, F.; Koltunow, A.M.G. Sporophytic ovule tissues modulate the initiation and progression of apomixis in Hieracium. J. Exp. Bot. 2012, 63, 3229–3241. [Google Scholar] [CrossRef]
- Suzaki, T.; Yano, K.; Ito, M.; Umehara, Y.; Suganuma, N.; Kawaguchi, M. Positive and negative regulation of cortical cell division during root nodule development in Lotus japonicus is accompanied by auxin response. Development 2012, 139, 3997–4006. [Google Scholar] [CrossRef]
- Nadzieja, M.; Kelly, S.; Stougaard, J.; Reid, D. Epidermal auxin biosynthesis facilitates rhizobial infection in Lotus japonicus. Plant J. 2018, 95, 101–111. [Google Scholar] [CrossRef]
- Mir, R.; Aranda, L.Z.; Biaocchi, T.; Luo, A.; Sylvester, A.W.; Rasmussen, C.G. A DII domain-based auxin reporter uncovers low auxin signaling during telophase and early G1. Plant Physiol. 2017, 173, 863–871. [Google Scholar] [CrossRef] [PubMed]
- Gallavotti, A.; Yang, Y.; Schmidt, R.J.; Jackson, D. The relationship between auxin transport and maize branching. Plant Physiol. 2008, 147, 1913–1923. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Han, L.; Hou, C.; Metelli, A.; Qi, L.; Tadege, M.; Mysore, K.S.; Wang, Z.Y. Developmental analysis of a Medicago truncatula smooth leaf margin1 mutant reveals context-dependent effects on compound leaf development. Plant Cell 2011, 23, 2106–2124. [Google Scholar] [CrossRef] [PubMed]
- Herrbach, V.; Remblière, C.; Gough, C.; Bensmihen, S. Lateral root formation and patterning in Medicago truncatula. J. Plant Physiol. 2014, 171, 301–310. [Google Scholar] [CrossRef]
- Chen, Y.; Yordanov, Y.S.; Ma, C.; Strauss, S.; Busov, V.B. DR5 as a reporter system to study auxin response in Populus. Plant Cell Rep. 2013, 32, 453–463. [Google Scholar] [CrossRef] [PubMed]
- Joshi, M.; Fogelman, E.; Belausov, E.; Ginzberg, I. Potato root system development and factors that determine its architecture. J. Plant Physiol. 2016, 205, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yuan, Z.; Meng, Q.; Huang, G.; Périn, C.; Bureau, C.; Meunier, A.-C.; Ingouff, M.; Bennett, M.J.; Liang, W.; et al. Dynamic Regulation of Auxin Response during Rice Development Revealed by Newly Established Hormone Biosensor Markers. Front. Plant Sci. 2017, 8, 256. [Google Scholar] [CrossRef] [PubMed]
- Zoulias, N.; Duttke, S.H.C.; Garcês, H.; Spencer, V.; Kim, M. The role of auxin in the pattern formation of the Asteraceae flower head (Capitulum). Plant Physiol. 2019, 179, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Fisher, J.; Gaillard, P.; Fellbaum, C.R.; Subramanian, S.; Smith, S. Quantitative 3D imaging of cell level auxin and cytokinin response ratios in soybean roots and nodules. Plant Cell Environ. 2018, 41, 2080–2092. [Google Scholar] [CrossRef]
- Turner, M.; Nizampatnam, N.R.; Baron, M.; Coppin, S.; Damodaran, S.; Adhikari, S.; Arunachalam, S.P.; Yu, O.; Subramanian, S. Ectopic expression of miR160 results in auxin hypersensitivity, cytokinin hyposensitivity, and inhibition of symbiotic nodule development in soybean. Plant Physiol. 2013, 162, 2042–2055. [Google Scholar] [CrossRef] [PubMed]
- Dubrovsky, J.G.; Sauer, M.; Napsucialy-Mendivil, S.; Ivanchenko, M.G.; Friml, J.; Shishkova, S.; Celenza, J.; Benková, E. Auxin acts as a local morphogenetic trigger to specify lateral root founder cells. Proc. Natl. Acad. Sci. USA 2008, 105, 8790–8794. [Google Scholar] [CrossRef] [PubMed]
- Ben-Gera, H.; Shwartz, I.; Shao, M.-R.R.; Shani, E.; Estelle, M.; Ori, N. ENTIRE and GOBLET promote leaflet development in tomato by modulating auxin response. Plant J. 2012, 70, 903–915. [Google Scholar] [CrossRef] [PubMed]
- Pattison, R.J.; Catalá, C. Evaluating auxin distribution in tomato (Solanum lycopersicum) through an analysis of the PIN and AUX/LAX gene families. Plant J. 2012, 70, 585–598. [Google Scholar] [CrossRef]
- Kirschner, G.K.; Stahl, Y.; Imani, J.; von Korff, M.; Simon, R. Fluorescent reporter lines for auxin and cytokinin signalling in barley (Hordeum vulgare). PLoS ONE 2018, 13, e0196086. [Google Scholar] [CrossRef]
- Reid, D.; Nadzieja, M.; Novák, O.; Heckmann, A.B.; Sandal, N.; Stougaard, J. Cytokinin biosynthesis promotes cortical cell responses during nodule development. Plant Physiol. 2017, 175, 361–375. [Google Scholar] [CrossRef] [PubMed]
- Tao, J.; Sun, H.; Gu, P.; Liang, Z.; Chen, X.; Lou, J.; Xu, G.; Zhang, Y. A sensitive synthetic reporter for visualizing cytokinin signaling output in rice. Plant Methods 2017, 13, 89. [Google Scholar] [CrossRef] [PubMed]
- Farber, M.; Attia, Z.; Weiss, D. Cytokinin activity increases stomatal density and transpiration rate in tomato. J. Exp. Bot. 2016, 67, 6351–6362. [Google Scholar] [CrossRef] [PubMed]
- Jonkman, J.; Brown, C.M.; Wright, G.D.; Anderson, K.I.; North, A.J. Tutorial: Guidance for quantitative confocal microscopy. Nat. Protoc. 2020, 15, 1585–1611. [Google Scholar] [CrossRef]
- Donaldson, L. Autofluorescence in plants. Molecules 2020, 25, 2393. [Google Scholar] [CrossRef]
- Liu, D.Y.T.; Kuhlmey, B.T.; Smith, P.M.C.; Day, D.A.; Faulkner, C.R.; Overall, R.L. Reflection across plant cell boundaries in confocal laser scanning microscopy. J. Microsc. 2008, 231, 349–357. [Google Scholar] [CrossRef]
- De Col, V.; Fuchs, P.; Nietzel, T.; Elsässer, M.; Voon, C.P.; Candeo, A.; Seeliger, I.; Fricker, M.D.; Grefen, C.; Møller, I.M.; et al. ATP sensing in living plant cells reveals tissue gradients and stress dynamics of energy physiology. Elife 2017, 6, e26770. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Onai, K.; Abe, A.; Shimizu, M.; Takagi, H.; Tateda, C.; Utsushi, H.; Singkarabanit-Ogawa, S.; Kitakura, S.; Ono, E.; et al. Lumi-Map, a real-time luciferase bioluminescence screen of mutants combined with MutMap, reveals Arabidopsis genes involved in PAMP-triggered immunity. Mol. Plant Microbe Interact. 2020, 33, 1366–1380. [Google Scholar] [CrossRef]
- Vaz Martins, T.; Livina, V.N. What drives symbiotic calcium signalling in legumes? Insights and challenges of imaging. Int. J. Mol. Sci. 2019, 20, 2245. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Mazarei, M.; Rudis, M.R.; Fethe, M.H.; Stewart, C.N. Rapid in vivo analysis of synthetic promoters for plant pathogen phytosensing. BMC Biotechnol. 2011, 11, 108. [Google Scholar] [CrossRef]
- Shokouhifar, F.; Bahrabadi, M.; Bagheri, A.; Mamarabadi, M. Transient expression analysis of synthetic promoters containing F and D cis-acting elements in response to Ascochyta rabiei and two plant defense hormones. AMB Express 2019, 9, 195. [Google Scholar] [CrossRef] [PubMed]
- Peyret, H.; Brown, J.K.M.; Lomonossoff, G.P. Improving plant transient expression through the rational design of synthetic 5′ and 3′ untranslated regions. Plant Methods 2019, 15, 108. [Google Scholar] [CrossRef]
- Matsui, T.; Sawada, K.; Takita, E.; Kato, K. Compatibility of translational enhancers with various plant species. Plant Biotechnol. 2015, 32, 309–316. [Google Scholar] [CrossRef][Green Version]
- Persad, R.; Reuter, D.N.; Dice, L.T.; Nguyen, M.A.; Rigoulot, S.B.; Layton, J.S.; Schmid, M.J.; Poindexter, M.R.; Occhialini, A.; Stewart, C.N.; et al. The Q-System as a synthetic transcriptional regulator in plants. Front. Plant Sci. 2020, 11, 245. [Google Scholar] [CrossRef] [PubMed]
- Stoddard, A.; Rolland, V. I see the light! Fluorescent proteins suitable for cell wall/apoplast targeting in Nicotiana benthamiana leaves. Plant Direct 2019, 3, e00112. [Google Scholar] [CrossRef] [PubMed]
- Shokhina, A.G.; Kostyuk, A.I.; Ermakova, Y.G.; Panova, A.S.; Staroverov, D.B.; Egorov, E.S.; Baranov, M.S.; van Belle, G.J.; Katschinski, D.M.; Belousov, V.V.; et al. Red fluorescent redox-sensitive biosensor Grx1-roCherry. Redox Biol. 2019, 21, 101071. [Google Scholar] [CrossRef]
- Ermakova, Y.G.; Bilan, D.S.; Matlashov, M.E.; Mishina, N.M.; Markvicheva, K.N.; Subach, O.M.; Subach, F.V.; Bogeski, I.; Hoth, M.; Enikolopov, G.; et al. Red fluorescent genetically encoded indicator for intracellular hydrogen peroxide. Nat. Commun. 2014, 5, 5222. [Google Scholar] [CrossRef] [PubMed]
- De Paepe, B.; Maertens, J.; Vanholme, B.; De Mey, M. Chimeric LysR-type transcriptional biosensors for customizing ligand specificity profiles toward flavonoids. ACS Synth. Biol. 2019, 8, 318–331. [Google Scholar] [CrossRef]
- Raman, V.K.; Aggarwal, D.; Rao, R.S.; Sreevathsa, R.; Singh, A.; Abdin, M.; Mohapatra, T.; Pattanayak, D. Rapid and efficient Agrobacterium mediated transformation of early scutellum derived calli of indica rice. Indian J. Exp. Biol. 2018, 56, 20–28. [Google Scholar]
- Mankin, S.L.; Allen, G.C.; Thompson, W.F. Introduction of a plant intron into the luciferase gene of Photinus pyralis. Plant Mol. Biol. Rep. 1997, 15, 186–196. [Google Scholar] [CrossRef]
- Laxa, M. Intron-mediated enhancement: A tool for heterologous gene expression in plants? Front. Plant Sci. 2017, 7, 1977. [Google Scholar] [CrossRef] [PubMed]
- Machado, D.; Costa, R.S.; Rocha, M.; Ferreira, E.C.; Tidor, B.; Rocha, I. Modeling formalisms in systems biology. AMB Express 2011, 1, 1–14. [Google Scholar] [CrossRef]
- Mithöfer, A.; Mazars, C. Aequorin-based measurements of intracellular Ca2+-signatures in plant cells. Biol. Proc. Online 2002, 4, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Zhang, K.; Ding, Z.; He, Q.; Li, W.; Zhu, S.; Cheng, W.; Zhang, K.; Li, K. Characterization of a strong and constitutive promoter from the Arabidopsis serine carboxypeptidase-like gene AtSCPL30 as a potential tool for crop transgenic breeding. BMC Biotechnol. 2018, 18, 59. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.-M.; Kallam, K.; Tidd, H.; Gendarini, G.; Salzman, A.; Patron, N.J. Rational design of minimal synthetic promoters for plants. Nucleic Acids Res. 2020, 48, 11845–11856. [Google Scholar] [CrossRef]
- Liu, Z.; Chen, O.; Wall, J.B.J.; Zheng, M.; Zhou, Y.; Wang, L.; Ruth Vaseghi, H.; Qian, L.; Liu, J. Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci. Rep. 2017, 7, 2193. [Google Scholar] [CrossRef] [PubMed]
- François, I.E.J.A.; Van Hemelrijck, W.; Aerts, A.M.; Wouters, P.F.J.; Proost, P.; Broekaert, W.F.; Cammue, B.P.A. Processing in Arabidopsis thaliana of a heterologous polyprotein resulting in differential targeting of the individual plant defensins. Plant Sci. 2004, 166, 113–121. [Google Scholar] [CrossRef]
- Sun, H.; Zhou, N.; Wang, H.; Huang, D.; Lang, Z. Processing and targeting of proteins derived from polyprotein with 2A and LP4/2A as peptide linkers in a maize expression system. PLoS ONE 2017, 12, e0174804. [Google Scholar] [CrossRef]
- Zhang, B.; Rapolu, M.; Kumar, S.; Gupta, M.; Liang, Z.; Han, Z.; Williams, P.; Su, W.W. Coordinated protein co-expression in plants by harnessing the synergy between an intein and a viral 2A peptide. Plant Biotechnol. J. 2017, 15, 718–728. [Google Scholar] [CrossRef]
- Liu, T.-Y.; Chou, W.-C.; Chen, W.-Y.; Chu, C.-Y.; Dai, C.-Y.; Wu, P.-Y. Detection of membrane protein-protein interaction in planta based on dual-intein-coupled tripartite split-GFP association. Plant J. 2018, 94, 426–438. [Google Scholar] [CrossRef] [PubMed]
- Mitiouchkina, T.; Mishin, A.S.; Somermeyer, L.G.; Markina, N.M.; Chepurnyh, T.V.; Guglya, E.B.; Karataeva, T.A.; Palkina, K.A.; Shakhova, E.S.; Fakhranurova, L.I.; et al. Plants with genetically encoded autoluminescence. Nat. Biotechnol. 2020, 38, 944–946. [Google Scholar] [CrossRef]
- Khakhar, A.; Starker, C.G.; Chamness, J.C.; Lee, N.; Stokke, S.; Wang, C.; Swanson, R.; Rizvi, F.; Imaizumi, T.; Voytas, D.F. Building customizable auto-luminescent luciferase-based reporters in plants. Elife 2020, 9, e52786. [Google Scholar] [CrossRef]
- He, Y.; Zhang, T.; Sun, H.; Zhan, H.; Zhao, Y. A reporter for noninvasively monitoring gene expression and plant transformation. Hortic. Res. 2020, 7, 152. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).