Origin of Species before Origin of Life: The Role of Speciation in Chemical Evolution
Abstract
1. Introduction
2. Species Concepts and Definitions
3. Speciation in Biology
4. Prebiotic Compartments
5. Prebiotic Compartment Speciation
6. Droplet-Based Microfluidics: Speciation in Action
7. Summary and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Mora, C.; Tittensor, D.P.; Adl, S.; Simpson, A.G.B.; Worm, B. How Many Species are There on Earth and in the Ocean? PLoS Biol. 2011, 9, e1001127. [Google Scholar] [CrossRef]
- Tilman, D.; Reich, P.B.; Knops, J.; Wedin, D.; Mielke, T.; Lehman, C. Diversity and Productivity in a Long-Term Grassland Experiment. Science 2001, 294, 843–845. [Google Scholar] [CrossRef]
- Cardinale, B.J.; Srivastava, D.S.; Duffy, J.E.; Wright, J.P.; Downing, A.L.; Sankaran, M.; Jouseau, C. Effects of biodiversity on the functioning of trophic groups and ecosystems. Nature 2006, 443, 989–992. [Google Scholar] [CrossRef]
- Balvanera, P.; Pfisterer, A.B.; Buchmann, N.; He, J.-S.; Nakashizuka, T.; Raffaelli, D.; Schmid, B. Quantifying the evidence for biodiversity effects on ecosystem functioning and services. Ecol. Lett. 2006, 9, 1146–1156. [Google Scholar] [CrossRef]
- Zavaleta, E.S.; Pasari, J.R.; Hulvey, K.B.; Tilman, G.D. Sustaining multiple ecosystem functions in grassland communities requires higher biodiversity. Proc. Natl. Acad. Sci. USA 2010, 107, 1443–1446. [Google Scholar] [CrossRef]
- Darwin, C. On the Origin of Species, 5th ed.; D. Appleton and Company: New York, NY, USA, 1871. [Google Scholar]
- Tilman, D.; Lehman, C.L.; Thomson, K.T. Plant diversity and ecosystem productivity: Theoretical considerations. Proc. Natl. Acad. Sci. USA 1997, 94, 1857–1861. [Google Scholar] [CrossRef]
- Forrester, D.I.; Bauhus, J. A Review of Processes Behind Diversity—Productivity Relationships in Forests. Curr. For. Rep. 2016, 2, 45–61. [Google Scholar] [CrossRef]
- Ives, A.R.; Carpenter, S.R. Stability and Diversity of Ecosystems. Science 2007, 317, 58–62. [Google Scholar] [CrossRef]
- Yachi, S.; Loreau, M. Biodiversity and ecosystem productivity in a fluctuating environment: The insurance hypothesis. Proc. Natl. Acad. Sci. USA 1999, 96, 1463–1468. [Google Scholar] [CrossRef]
- Elton, C.C. The Ecology of Invasions by Animals and Plants; Springer: Dordrecht, The Netherlands, 1977; ISBN 9780412214608. [Google Scholar]
- Tilman, D. Niche tradeoffs, neutrality, and community structure: A stochastic theory of resource competition, invasion, and community assembly. Proc. Natl. Acad. Sci. USA 2004, 101, 10854–10861. [Google Scholar] [CrossRef]
- Bascompte, J. Disentangling the Web of Life. Science 2009, 325, 416–419. [Google Scholar] [CrossRef]
- Wächtershäuser, G. Before enzymes and templates: Theory of surface metabolism. Microbiol. Rev. 1988, 52, 452–484. [Google Scholar] [CrossRef]
- Russell, M.J.; Hall, A.J. The emergence of life from iron monosulphide bubbles at a submarine hydrothermal redox and pH front. J. Geol. Soc. Lond. 1997, 154, 377–402. [Google Scholar] [CrossRef]
- Fox, S.W.; Nakashima, T. The assembly and properties of protobiological structures: The beginnings of cellular peptide synthesis. Biosystems 1980, 12, 155–166. [Google Scholar] [CrossRef]
- Wächtershäuser, G. From volcanic origins of chemoautotrophic life to Bacteria, Archaea and Eukarya. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 1787–1806. [Google Scholar] [CrossRef]
- Oparin, A.I. The Origin of Life; Dover Publications: New York, NY, USA, 1965; ISBN 9780486602134. [Google Scholar]
- Haldane, J.B.S. Origin of Life. Ration. Annu. 1929, 148, 3–10. [Google Scholar]
- Oparin, A.I.; Braunshtein, A.E.; Pasynskii, A.G.; Pavlovskaya, T.E. The Origin of Life on the Earth; Elsevier: Amsterdam, The Netherlands, 1959; ISBN 9781483197371. [Google Scholar]
- Guttenberg, N.; Virgo, N.; Chandru, K.; Scharf, C.; Mamajanov, I. Bulk measurements of messy chemistries are needed for a theory of the origins of life. Philos. Trans. A Math. Phys. Eng. Sci. 2017, 375, 20160347. [Google Scholar] [CrossRef]
- Muchowska, K.B.; Varma, S.J.; Moran, J. Nonenzymatic Metabolic Reactions and Life’s Origins. Chem. Rev. 2020, 120, 7708–7744. [Google Scholar] [CrossRef] [PubMed]
- Morowitz, H.J. Energy Flow in Biology; Biological Organization as Problem in Thermal Physics; Academic Press: New York, NY, USA, 1968; ISBN 9780918024138. [Google Scholar]
- Kauffman, S.A. Autocatalytic sets of proteins. J. Theor. Biol. 1986, 119, 1–24. [Google Scholar] [CrossRef]
- Shapiro, R. Small molecule interactions were central to the origin of life. Q. Rev. Biol. 2006, 81, 105–125. [Google Scholar] [CrossRef] [PubMed]
- Dadon, Z.; Wagner, N.; Ashkenasy, G. The road to non-enzymatic molecular networks. Angew. Chem. Int. Ed. Engl. 2008, 47, 6128–6136. [Google Scholar] [CrossRef] [PubMed]
- Joyce, G.F.; Szostak, J.W. Protocells and RNA Self-Replication. Cold Spring Harb. Perspect. Biol. 2018, 10, a034801. [Google Scholar] [CrossRef]
- Koonin, E.V. Comparative genomics, minimal gene-sets and the last universal common ancestor. Nat. Rev. Microbiol. 2003, 1, 127–136. [Google Scholar] [CrossRef]
- Delaye, L.; Becerra, A.; Lazcano, A. The Last Common Ancestor: What’s in a name? Orig. Life Evol. Biosph. 2005, 35, 537–554. [Google Scholar] [CrossRef]
- Cornish-Bowden, A.; Luz Cárdenas, M. Life before LUCA. J. Theor. Biol. 2017, 434, 68–74. [Google Scholar] [CrossRef]
- Raup, D.M.; Valentine, J.W. Multiple origins of life. Proc. Natl. Acad. Sci. USA 1983, 80, 2981–2984. [Google Scholar] [CrossRef]
- Mallet, J. Alternative views of biological species: Reproductively isolated units or genotypic clusters? Natl. Sci. Rev. 2020, 7, 1401–1407. [Google Scholar] [CrossRef]
- Mallet, J. Species, Encyclopedia of Biodiversity, 2nd ed.; Levin, S.A., Ed.; Academic Press: Waltham, MA, USA, 2013; Volume 6, pp. 679–691. [Google Scholar]
- De Queiroz, K. Species Concepts and Species Delimitation. Syst. Biol. 2007, 56, 879–886. [Google Scholar] [CrossRef]
- De Queiroz, K. Ernst Mayr and the modern concept of species. Proc. Natl. Acad. Sci. USA 2005, 102, 6600–6607. [Google Scholar] [CrossRef] [PubMed]
- Zirkle, C. Species before Darwin. Proc. Am. Philos. Soc. 1959, 103, 636–644. [Google Scholar]
- Ray, J.; Linnaeus, C. Synopsis Methodica Stirpium Britannicarum: Editio Tertia, 1724 [and] Flora Anglica, 1754 & 1759, 3rd ed.; Ray Society: London, UK, 1973; ISBN 9780903874007. [Google Scholar]
- Mayr, E. The Growth of Biological Thought: Diversity, Evolution, and Inheritance; Harvard University Press: Cambridge, MA, USA, 1982; ISBN 9780674364462. [Google Scholar]
- Linnaeus, C. Species Plantarum; Ray Society: London, UK, 2014; ISBN 9780903874472. [Google Scholar]
- Larson, J.L. The Species Concept of Linnaeus. Isis 1968, 59, 291–299. [Google Scholar] [CrossRef]
- Mayr, E. Systematics and the Origin of Species, from the Viewpoint of a Zoologist; Harvard University Press: Cambridge, MA, USA, 1999; ISBN 9780674862500. [Google Scholar]
- Dobzhansky, T. Genetics and the Origin of Species; Columbia University Press: New York, NY, USA, 1982; ISBN 9780231054751. [Google Scholar]
- Thorp, J.H.; Rogers, D.C. Thorp and Covich’s Freshwater Invertebrates: Ecology and General Biology; Elsevier: Amsterdam, The Netherlands, 2014; ISBN 9780123850270. [Google Scholar]
- Häuser, C.L. The debate about the biological species concept—A review. J. Zool. Syst. Evol. Res. 2009, 25, 241–257. [Google Scholar] [CrossRef]
- Sokal, R.R.; Crovello, T.J. The Biological Species Concept: A Critical Evaluation. Am. Nat. 1970, 104, 127–153. [Google Scholar] [CrossRef]
- Velasco, J.D. Species concepts should not conflict with evolutionary history, but often do. Stud. Hist. Philos. Biol. Biomed. Sci. 2008, 39, 407–414. [Google Scholar] [CrossRef]
- Wilkins, J.S. Philosophically speaking, how many species concepts are there? Zootaxa 2011, 2765, 58–60. [Google Scholar] [CrossRef]
- Woese, C.R.; Kandler, O.; Wheelis, M.L. Towards a natural system of organisms: Proposal for the domains Archaea, Bacteria, and Eucarya. Proc. Natl. Acad. Sci. USA 1990, 87, 4576–4579. [Google Scholar] [CrossRef] [PubMed]
- DeSalle, R.; Goldstein, P. Review and Interpretation of Trends in DNA Barcoding. Front. Ecol. Evol. 2019, 7, 302. [Google Scholar] [CrossRef]
- Van Valen, L. Ecological Species, Multispecies, and Oaks. Taxon 1976, 25, 233–239. [Google Scholar] [CrossRef]
- Clark, M.R.; Rowden, A.A.; Schlacher, T.A.; Guinotte, J.; Dunstan, P.K.; Williams, A.; O’Hara, T.D.; Watling, L.; Niklitschek, E.; Tsuchida, S. Identifying Ecologically or Biologically Significant Areas (EBSA): A systematic method and its application to seamounts in the South Pacific Ocean. Ocean. Coast. Manag. 2014, 91, 65–79. [Google Scholar] [CrossRef]
- Balakrishnan, R. Species Concepts, Species Boundaries and Species Identification: A View from the Tropics. Syst. Biol. 2005, 54, 689–693. [Google Scholar] [CrossRef]
- Mayr, E. What Is a Species, and What Is Not? Philos. Sci. 1996, 63, 262–277. [Google Scholar] [CrossRef]
- Coyne, J.A.; Orr, H.A. Speciation; Sinauer: Sunderland, MA, USA, 2004; ISBN 9780878930890. [Google Scholar]
- Blanckaert, A.; Bank, C.; Hermisson, J. The limits to parapatric speciation 3: Evolution of strong reproductive isolation in presence of gene flow despite limited ecological differentiation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2020, 375, 20190532. [Google Scholar] [CrossRef] [PubMed]
- Lamichhaney, S.; Berglund, J.; Almén, M.S.; Maqbool, K.; Grabherr, M.; Martinez-Barrio, A.; Promerová, M.; Rubin, C.-J.; Wang, C.; Zamani, N.; et al. Evolution of Darwin’s finches and their beaks revealed by genome sequencing. Nature 2015, 518, 371–375. [Google Scholar] [CrossRef]
- Sulloway, F.J. Tantalizing Tortoises and the Darwin-Galápagos Legend. J. Hist. Biol. 2009, 42, 3–31. [Google Scholar] [CrossRef]
- Lawson, L.P.; Bates, J.M.; Menegon, M.; Loader, S.P. Divergence at the edges: Peripatric isolation in the montane spiny throated reed frog complex. BMC Evol. Biol. 2015, 15, 128. [Google Scholar] [CrossRef]
- Feder, J.L.; Roethele, J.B.; Filchak, K.; Niedbalski, J.; Romero-Severson, J. Evidence for inversion polymorphism related to sympatric host race formation in the apple maggot fly, Rhagoletis pomonella. Genetics 2003, 163, 939–953. [Google Scholar]
- Jiggins, C.D.; Bridle, J.R. Speciation in the apple maggot fly: A blend of vintages? Trends Ecol. Evol. 2004, 19, 111–114. [Google Scholar] [CrossRef]
- Smith, J.M. Sympatric Speciation. Am. Nat. 1966, 100, 637–650. [Google Scholar] [CrossRef]
- Dieckmann, U.; Doebeli, M. On the origin of species by sympatric speciation. Nature 1999, 400, 354–357. [Google Scholar] [CrossRef]
- William Birky, C.; Barraclough, T.G. Asexual Speciation. In Lost Sex; Springer: Dordrecht, The Netherlands, 2009; pp. 201–216. ISBN 9789048127702. [Google Scholar]
- Soucy, S.M.; Huang, J.; Gogarten, J.P. Horizontal gene transfer: Building the web of life. Nat. Rev. Genet. 2015, 16, 472–482. [Google Scholar] [CrossRef]
- Shapiro, B.J.; Leducq, J.-B.; Mallet, J. What Is Speciation? PLoS Genet. 2016, 12, e1005860. [Google Scholar] [CrossRef]
- Hanage, W.P.; Fraser, C.; Spratt, B.G. Fuzzy species among recombinogenic bacteria. BMC Biol. 2005, 3, 1–7. [Google Scholar] [CrossRef]
- Shapiro, B.J.; Polz, M.F. Microbial Speciation. Cold Spring Harb. Perspect. Biol. 2015, 7, a018143. [Google Scholar] [CrossRef]
- Cleaves, H.J. Prebiotic Chemistry: Geochemical Context and Reaction Screening. Life 2013, 3, 331. [Google Scholar] [CrossRef]
- Patel, B.H.; Percivalle, C.; Ritson, D.J.; Duffy, C.D.; Sutherland, J.D. Common origins of RNA, protein and lipid precursors in a cyanosulfidic protometabolism. Nat. Chem. 2015, 7, 301–307. [Google Scholar] [CrossRef]
- Adam, Z.R.; Fahrenbach, A.C.; Kacar, B.; Aono, M. Prebiotic Geochemical Automata at the Intersection of Radiolytic Chemistry, Physical Complexity, and Systems Biology. Complexity 2018, 2018. [Google Scholar] [CrossRef]
- Vincent, L.; Berg, M.; Krismer, M.; Saghafi, S.T.; Cosby, J.; Sankari, T.; Vetsigian, K.; Cleaves, H.J.; Baum, D.A. Chemical Ecosystem Selection on Mineral Surfaces Reveals Long-Term Dynamics Consistent with the Spontaneous Emergence of Mutual Catalysis. Life 2019, 9, 80. [Google Scholar] [CrossRef]
- Segré, D.; Shenhav, B.; Kafri, R.; Lancet, D. The molecular roots of compositional inheritance. J. Theor. Biol. 2001, 213, 481–491. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gutierrez, J.M.P.; Hinkley, T.; Taylor, J.W.; Yanev, K.; Cronin, L. Evolution of oil droplets in a chemorobotic platform. Nat. Commun. 2014, 5, 380. [Google Scholar] [CrossRef]
- Adamala, K.; Szostak, J.W. Competition between model protocells driven by an encapsulated catalyst. Nat. Chem. 2013, 5, 495–501. [Google Scholar] [CrossRef] [PubMed]
- Shirt-Ediss, B.J. Modelling Early Transitions toward Autonomous Protocells. arXiv 2016, arXiv:1606.03620. [Google Scholar]
- Spoelstra, W.K.; Deshpande, S.; Dekker, C. Tailoring the appearance: What will synthetic cells look like? Curr. Opin. Biotechnol. 2018, 51, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Adam, Z.R.; Hongo, Y.; Cleaves, H.J., 2nd; Yi, R.; Fahrenbach, A.C.; Yoda, I.; Aono, M. Estimating the capacity for production of formamide by radioactive minerals on the prebiotic Earth. Sci. Rep. 2018, 8, 265. [Google Scholar] [CrossRef]
- Jia, T.Z.; Hentrich, C.; Szostak, J.W. Rapid RNA exchange in aqueous two-phase system and coacervate droplets. Orig. Life Evol. Biosph. 2014, 44, 1–12. [Google Scholar] [CrossRef]
- Poudyal, R.R.; Guth-Metzler, R.M.; Veenis, A.J.; Frankel, E.A.; Keating, C.D.; Bevilacqua, P.C. Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef]
- Tagami, S.; Attwater, J.; Holliger, P. Simple peptides derived from the ribosomal core potentiate RNA polymerase ribozyme function. Nat. Chem. 2017, 9, 325–332. [Google Scholar] [CrossRef]
- Szostak, J.W. The eightfold path to non-enzymatic RNA replication. J. Syst. Chem. 2012, 3, 2. [Google Scholar] [CrossRef]
- Szostak, J.W.; Bartel, D.P.; Luisi, P.L. Synthesizing life. Nature 2001, 409, 387–390. [Google Scholar] [CrossRef] [PubMed]
- Schrum, J.P.; Zhu, T.F.; Szostak, J.W. The origins of cellular life. Cold Spring Harb. Perspect. Biol. 2010, 2, a002212. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Szostak, J.W. Lipid constituents of model protocell membranes. Emerg. Top. Life Sci. 2019, 3, 537–542. [Google Scholar]
- Mansy, S. Model Protocells from Single-Chain Lipids. Int. J. Mol. Sci. 2009, 10, 835–843. [Google Scholar] [CrossRef]
- Deamer, D.W.; Pashley, R.M. Amphiphilic components of the Murchison carbonaceous chondrite: Surface properties and membrane formation. Orig. Life Evol. Biosph. 1989, 19, 21–38. [Google Scholar] [CrossRef]
- Fiore, M.; Strazewski, P. Prebiotic Lipidic Amphiphiles and Condensing Agents on the Early Earth. Life 2016, 6, 17. [Google Scholar] [CrossRef] [PubMed]
- Sahai, N.; Kaddour, H.; Dalai, P.; Wang, Z.; Bass, G.; Gao, M. Mineral Surface Chemistry and Nanoparticle-aggregation Control Membrane Self-Assembly. Sci. Rep. 2017, 7, 435. [Google Scholar] [CrossRef]
- Milshteyn, D.; Damer, B.; Havig, J.; Deamer, D. Amphiphilic Compounds Assemble into Membranous Vesicles in Hydrothermal Hot Spring Water but Not in Seawater. Life 2018, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Damer, B.; Deamer, D. The Hot Spring Hypothesis for an Origin of Life. Astrobiology 2020, 20, 429–452. [Google Scholar] [CrossRef]
- Forsythe, J.G.; Yu, S.-S.; Mamajanov, I.; Grover, M.A.; Krishnamurthy, R.; Fernández, F.M.; Hud, N.V. Ester-Mediated Amide Bond Formation Driven by Wet-Dry Cycles: A Possible Path to Polypeptides on the Prebiotic Earth. Angew. Chem. Int. Ed. Engl. 2015, 127, 10009–10013. [Google Scholar] [CrossRef]
- Campbell, T.D.; Febrian, R.; McCarthy, J.T.; Kleinschmidt, H.E.; Forsythe, J.G.; Bracher, P.J. Prebiotic condensation through wet-dry cycling regulated by deliquescence. Nat. Commun. 2019, 10, 4508. [Google Scholar] [CrossRef]
- Mamajanov, I. Wet-Dry Cycling Delays the Gelation of Hyperbranched Polyesters: Implications to the Origin of Life. Life 2019, 9, 56. [Google Scholar] [CrossRef] [PubMed]
- Chandru, K.; Guttenberg, N.; Giri, C.; Hongo, Y.; Butch, C.; Mamajanov, I.; Cleaves, H.J. Simple prebiotic synthesis of high diversity dynamic combinatorial polyester libraries. Commun. Chem. 2018, 1, 30. [Google Scholar] [CrossRef]
- Mamajanov, I.; MacDonald, P.J.; Ying, J.; Duncanson, D.M.; Dowdy, G.R.; Walker, C.A.; Engelhart, A.E.; Fernández, F.M.; Grover, M.A.; Hud, N.V.; et al. Ester Formation and Hydrolysis during Wet–Dry Cycles: Generation of Far-from-Equilibrium Polymers in a Model Prebiotic Reaction. Macromolecules 2014, 47, 1334–1343. [Google Scholar] [CrossRef]
- Higgs, P.G. The Effect of Limited Diffusion and Wet-Dry Cycling on Reversible Polymerization Reactions: Implications for Prebiotic Synthesis of Nucleic Acids. Life 2016, 6, 24. [Google Scholar] [CrossRef]
- Chen, I.A.; Walde, P. From self-assembled vesicles to protocells. Cold Spring Harb. Perspect. Biol. 2010, 2, a002170. [Google Scholar] [CrossRef]
- Mansy, S.S.; Szostak, J.W. Thermostability of model protocell membranes. Proc. Natl. Acad. Sci. USA 2008, 105, 13351–13355. [Google Scholar] [CrossRef]
- Apel, C.L.; Deamer, D.W.; Mautner, M.N. Self-assembled vesicles of monocarboxylic acids and alcohols: Conditions for stability and for the encapsulation of biopolymers. Biochim. Biophys. Acta Biomembr. 2002, 1559, 1–9. [Google Scholar] [CrossRef]
- Adamala, K.P.; Engelhart, A.E.; Szostak, J.W. Collaboration between primitive cell membranes and soluble catalysts. Nat. Commun. 2016, 7, 11041. [Google Scholar] [CrossRef]
- Jin, L.; Engelhart, A.E.; Adamala, K.P.; Szostak, J.W. Preparation, Purification, and Use of Fatty Acid-containing Liposomes. J. Vis. Exp. 2018, 132, e57324. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S.; Dagar, S.; Verma, A.; Rajamani, S. Compositional heterogeneity confers selective advantage to model protocellular membranes during the origins of cellular life. Sci. Rep. 2020, 10, 4483. [Google Scholar] [CrossRef]
- Budin, I.; Szostak, J.W. Physical effects underlying the transition from primitive to modern cell membranes. Proc. Natl. Acad. Sci. USA 2011, 108, 5249–5254. [Google Scholar] [CrossRef]
- Yoshizawa, T.; Nozawa, R.-S.; Jia, T.Z.; Saio, T.; Mori, E. Biological phase separation: Cell biology meets biophysics. Biophys. Rev. 2020, 12, 519–539. [Google Scholar] [CrossRef] [PubMed]
- Koga, S.; Williams, D.S.; Perriman, A.W.; Mann, S. Peptide-nucleotide microdroplets as a step towards a membrane-free protocell model. Nat. Chem. 2011, 3, 720–724. [Google Scholar] [CrossRef]
- Aumiller, W.M., Jr.; Keating, C.D. Phosphorylation-mediated RNA/peptide complex coacervation as a model for intracellular liquid organelles. Nat. Chem. 2016, 8, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Poudyal, R.R.; Pir Cakmak, F.; Keating, C.D.; Bevilacqua, P.C. Physical Principles and Extant Biology Reveal Roles for RNA-Containing Membraneless Compartments in Origins of Life Chemistry. Biochemistry 2018, 57, 2509–2519. [Google Scholar] [CrossRef]
- Pir Cakmak, F.; Keating, C.D. Combining Catalytic Microparticles with Droplets Formed by Phase Coexistence: Adsorption and Activity of Natural Clays at the Aqueous/Aqueous Interface. Sci. Rep. 2017, 7, 3215. [Google Scholar] [CrossRef]
- Crowe, C.D.; Keating, C.D. Liquid-liquid phase separation in artificial cells. Interface Focus 2018, 8, 20180032. [Google Scholar] [CrossRef]
- Ianeselli, A.; Mast, C.B.; Braun, D. Periodic Melting of Oligonucleotides by Oscillating Salt Concentrations Triggered by Microscale Water Cycles Inside Heated Rock Pores. Angew. Chem. Int. Ed. Engl. 2019, 58, 13155–13160. [Google Scholar] [CrossRef]
- Kreysing, M.; Keil, L.; Lanzmich, S.; Braun, D. Heat flux across an open pore enables the continuous replication and selection of oligonucleotides towards increasing length. Nat. Chem. 2015, 7, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Jia, T.Z.; Chandru, K.; Hongo, Y.; Afrin, R.; Usui, T.; Myojo, K.; Cleaves, H.J. Membraneless polyester microdroplets as primordial compartments at the origins of life. Proc. Natl. Acad. Sci. USA 2019, 116, 15830–15835. [Google Scholar] [CrossRef]
- Chandru, K.; Mamajanov, I.; Cleaves, H.J.; Jia, T.Z. Polyesters as a Model System for Building Primitive Biologies from Non-Biological Prebiotic Chemistry. Life 2020, 10, 6. [Google Scholar] [CrossRef]
- Mayer, C.; Schreiber, U.; Dávila, M.J. Selection of Prebiotic Molecules in Amphiphilic Environments. Life 2017, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, U.; Locker-Grütjen, O.; Mayer, C. Hypothesis: Origin of life in deep-reaching tectonic faults. Orig. Life Evol. Biosph. 2012, 42, 47–54. [Google Scholar] [CrossRef]
- Tang, T.-Y.D.; Rohaida Che Hak, C.; Thompson, A.J.; Kuimova, M.K.; Williams, D.S.; Perriman, A.W.; Mann, S. Fatty acid membrane assembly on coacervate microdroplets as a step towards a hybrid protocell model. Nat. Chem. 2014, 6, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Pir Cakmak, F.; Grigas, A.T.; Keating, C.D. Lipid Vesicle-Coated Complex Coacervates. Langmuir 2019, 35, 7830–7840. [Google Scholar] [CrossRef]
- Shakya, A.; King, J.T. DNA Local-Flexibility-Dependent Assembly of Phase-Separated Liquid Droplets. Biophys. J. 2018, 115, 1840–1847. [Google Scholar] [CrossRef]
- Mountain, G.A.; Keating, C.D. Formation of Multiphase Complex Coacervates and Partitioning of Biomolecules within them. Biomacromolecules 2020, 21, 630–640. [Google Scholar] [CrossRef]
- Fraccia, T.P.; Jia, T.Z. Liquid Crystal Coacervates Composed of Short Double-Stranded DNA and Cationic Peptides. ACS Nano 2020, 14, 15071–15082. [Google Scholar] [CrossRef]
- Robertson, M.P.; Joyce, G.F. The origins of the RNA world. Cold Spring Harb. Perspect. Biol. 2012, 4, a003608. [Google Scholar] [CrossRef]
- Le Vay, K.; Mutschler, H. The difficult case of an RNA-only origin of life. Emerg. Top. Life Sci. 2019, 3, 469–475. [Google Scholar] [PubMed]
- Hud, N.V. Searching for lost nucleotides of the pre-RNA World with a self-refining model of early Earth. Nat. Commun. 2018, 9, 5171. [Google Scholar] [CrossRef]
- Hud, N.V.; Cafferty, B.J.; Krishnamurthy, R.; Williams, L.D. The Origin of RNA and “My Grandfather’s Axe.”. Chem. Biol. 2013, 20, 466–474. [Google Scholar] [CrossRef]
- Segre, D.; Ben-Eli, D.; Lancet, D. Compositional genomes: Prebiotic information transfer in mutually catalytic noncovalent assemblies. Proc. Natl. Acad. Sci. USA 2000, 97, 4112–4117. [Google Scholar] [CrossRef] [PubMed]
- Strom, K.R.; Szostak, J.W.; Prywes, N. Transfer of Sequence Information and Replication of Diimine Duplexes. J. Org. Chem. 2019, 84, 3754–3761. [Google Scholar] [CrossRef]
- Turk, R.M.; Chumachenko, N.V.; Yarus, M. Multiple translational products from a five-nucleotide ribozyme. Proc. Natl. Acad. Sci. USA 2010, 107, 4585–4589. [Google Scholar] [CrossRef]
- Rufo, C.M.; Moroz, Y.S.; Moroz, O.V.; Stöhr, J.; Smith, T.A.; Hu, X.; DeGrado, W.F.; Korendovych, I.V. Short peptides self-assemble to produce catalytic amyloids. Nat. Chem. 2014, 6, 303–309. [Google Scholar] [CrossRef]
- Kroiss, D.; Aramini, J.M.; McPhee, S.A.; Tuttle, T.; Ulijn, R.V. Unbiased Discovery of Dynamic Peptide-ATP Complexes. ChemSystemsChem 2019, 1, 7–11. [Google Scholar] [CrossRef]
- Bonfio, C.; Valer, L.; Scintilla, S.; Shah, S.; Evans, D.J.; Jin, L.; Szostak, J.W.; Sasselov, D.D.; Sutherland, J.D.; Mansy, S.S. UV-light-driven prebiotic synthesis of iron-sulfur clusters. Nat. Chem. 2017, 9, 1229–1234. [Google Scholar] [CrossRef]
- Bonfio, C.; Godino, E.; Corsini, M.; Fabrizi de Biani, F.; Guella, G.; Mansy, S.S. Prebiotic iron–sulfur peptide catalysts generate a pH gradient across model membranes of late protocells. Nat. Catal. 2018, 1, 616–623. [Google Scholar] [CrossRef]
- Cleaves, H.J.; Scott, A.M.; Hill, F.C.; Leszczynski, J.; Sahai, N.; Hazen, R. Mineral-organic interfacial processes: Potential roles in the origins of life. Chem. Soc. Rev. 2012, 41, 5502–5525. [Google Scholar] [CrossRef]
- Gillams, R.J.; Jia, T.Z. Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research. Life 2018, 8, 10. [Google Scholar] [CrossRef]
- Lopez, A.; Fiore, M. Investigating Prebiotic Protocells for A Comprehensive Understanding of the Origins of Life: A Prebiotic Systems Chemistry Perspective. Life 2019, 9, 49. [Google Scholar] [CrossRef]
- Chen, I.A. The emergence of cells during the origin of life. Science 2006, 314, 1558–1559. [Google Scholar] [CrossRef]
- Mann, S. Systems of creation: The emergence of life from nonliving matter. Acc. Chem. Res. 2012, 45, 2131–2141. [Google Scholar] [CrossRef]
- Keating, C.D. Aqueous phase separation as a possible route to compartmentalization of biological molecules. Acc. Chem. Res. 2012, 45, 2114–2124. [Google Scholar] [CrossRef]
- Toparlak, O.D.; Mansy, S.S. Progress in synthesizing protocells. Exp. Biol. Med. 2019, 244, 304–313. [Google Scholar] [CrossRef]
- Silva, E.; Lazcano, A. Membranes and prebiotic evolution: Compartments, spatial isolation and the origin of life. In Molecules in Time and Space; Vicente, M., Tamames, J., Valencia, A., Mingorance, J., Eds.; Kluwer Academic Publishers: Boston, MA, USA, 2004; Volume 26, pp. 13–25. ISBN 9780306485787. [Google Scholar]
- Monnard, P.-A.; Walde, P. Current Ideas about Prebiological Compartmentalization. Life 2015, 5, 1239–1263. [Google Scholar] [CrossRef] [PubMed]
- Dzieciol, A.J.; Mann, S. Designs for life: Protocell models in the laboratory. Chem. Soc. Rev. 2012, 41, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Song, S.; van Hest, J.; Abdelmohsen, L.K.E.A.; Huang, X.; Sánchez, S. Biomimicry of Cellular Motility and Communication Based on Synthetic Soft-Architectures. Small 2020, 16, 1907680. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, S.; Das, S.; Dagar, S.; Joshi, M.P.; Mungi, C.V.; Sawant, A.A.; Patki, G.M.; Rajamani, S. Prebiological Membranes and Their Role in the Emergence of Early Cellular Life. J. Membr. Biol. 2020, 253, 589–608. [Google Scholar] [CrossRef]
- Okihana, H.; Ponnamperuma, C. Functions of the coacervate droplets. Orig. Life Evol. Biosph. 1982, 12, 347–353. [Google Scholar] [CrossRef]
- Monnard, P.-A.; Deamer, D.W. Membrane self-assembly processes: Steps toward the first cellular life. Anat. Rec. 2002, 268, 196–207. [Google Scholar] [CrossRef]
- Hall, D.; Minton, A.P. Macromolecular crowding: Qualitative and semiquantitative successes, quantitative challenges. Biochim. Biophys. Acta Proteins Proteom. 2003, 1649, 127–139. [Google Scholar] [CrossRef]
- Paudel, B.P.; Rueda, D. Molecular crowding accelerates ribozyme docking and catalysis. J. Am. Chem. Soc. 2014, 136, 16700–16703. [Google Scholar] [CrossRef]
- Saha, R.; Pohorille, A.; Chen, I.A. Molecular Crowding and Early Evolution. Orig. Life Evol. Biosph. 2014, 44, 319–324. [Google Scholar] [CrossRef]
- Ameta, S.; Arsène, S.; Foulon, S.; Saudemont, B.; Clifton, B.E.; Griffiths, A.D.; Nghe, P. Darwinian properties and their trade-offs in autocatalytic RNA reaction networks. Nat. Commun. 2021, 12, 842. [Google Scholar] [CrossRef]
- Hordijk, W.; Steel, M. Autocatalytic Networks at the Basis of Life’s Origin and Organization. Life 2018, 8, 62. [Google Scholar] [CrossRef]
- Chen, I.A.; Roberts, R.W.; Szostak, J.W. The emergence of competition between model protocells. Science 2004, 305, 1474–1476. [Google Scholar] [CrossRef]
- Scott, S.K. Chemical Chaos; Clarendon Press: Oxford, UK, 1993; ISBN 9780198556589. [Google Scholar]
- Field, R.J.; Györgyi, L. Chaos in Chemistry and Biochemistry; World Scientific: Singapore, 1993; ISBN 9789810210243. [Google Scholar]
- Matsumura, S.; Kun, Á.; Ryckelynck, M.; Coldren, F.; Szilágyi, A.; Jossinet, F.; Rick, C.; Nghe, P.; Szathmáry, E.; Griffiths, A.D. Transient compartmentalization of RNA replicators prevents extinction due to parasites. Science 2016, 354, 1293–1296. [Google Scholar] [CrossRef]
- Bansho, Y.; Furubayashi, T.; Ichihashi, N.; Yomo, T. Host-parasite oscillation dynamics and evolution in a compartmentalized RNA replication system. Proc. Natl. Acad. Sci. USA 2016, 113, 4045–4050. [Google Scholar] [CrossRef]
- Hogeweg, P.; Takeuchi, N. Multilevel Selection in Models of Prebiotic Evolution: Compartments and Spatial Self-organization. Orig. Life Evol. Biosph. 2003, 33, 375–403. [Google Scholar] [CrossRef]
- Takeuchi, N.; Hogeweg, P. Multilevel selection in models of prebiotic evolution II: A direct comparison of compartmentalization and spatial self-organization. PLoS Comput. Biol. 2009, 5, e1000542. [Google Scholar] [CrossRef]
- Kurihara, K.; Tamura, M.; Shohda, K.-I.; Toyota, T.; Suzuki, K.; Sugawara, T. Self-reproduction of supramolecular giant vesicles combined with the amplification of encapsulated DNA. Nat. Chem. 2011, 3, 775–781. [Google Scholar] [CrossRef]
- Yeh Martín, N.; Valer, L.; Mansy, S.S. Toward long-lasting artificial cells that better mimic natural living cells. Emerg. Top. Life Sci. 2019, 3, 597–607. [Google Scholar] [PubMed]
- Tawfik, D.S.; Griffiths, A.D. Man-made cell-like compartments for molecular evolution. Nat. Biotech. 1998, 16, 652–656. [Google Scholar] [CrossRef]
- Doran, D.; Abul-Haija, Y.M.; Cronin, L. Emergence of Function and Selection from Recursively Programmed Polymerisation Reactions in Mineral Environments. Angew. Chem. Int. Ed. Engl. 2019, 58, 11253–11256. [Google Scholar] [CrossRef]
- Koonin, E.V.; Martin, W. On the origin of genomes and cells within inorganic compartments. Trends Genet. 2005, 21, 647–654. [Google Scholar] [CrossRef]
- Martin, N. Dynamic Synthetic Cells Based on Liquid-Liquid Phase Separation. ChemBioChem 2019, 20, 2553–2568. [Google Scholar] [CrossRef]
- Sato, Y.; Sakamoto, T.; Takinoue, M. Sequence-based engineering of dynamic functions of micrometer-sized DNA droplets. Sci. Adv. 2020, 6, eaba3471. [Google Scholar] [CrossRef]
- Kapoor, S.; Berghaus, M.; Suladze, S.; Prumbaum, D.; Grobelny, S.; Degen, P.; Raunser, S.; Winter, R. Prebiotic Cell Membranes that Survive Extreme Environmental Pressure Conditions. Angew. Chem. Int. Ed. Engl. 2014, 126, 8537–8541. [Google Scholar] [CrossRef]
- Weiss, M.C.; Preiner, M.; Xavier, J.C.; Zimorski, V.; Martin, W.F. The last universal common ancestor between ancient Earth chemistry and the onset of genetics. PLoS Genet. 2018, 14, e1007518. [Google Scholar] [CrossRef]
- Ruiz-Mirazo, K.; Shirt-Ediss, B.; Escribano-Cabeza, M.; Moreno, A. The Construction of Biological “Inter-Identity” as the Outcome of a Complex Process of Protocell Development in Prebiotic Evolution. Front. Physiol. 2020, 11, 530. [Google Scholar] [CrossRef] [PubMed]
- Woese, C. The universal ancestor. Proc. Natl. Acad. Sci. USA 1998, 95, 6854–6859. [Google Scholar] [CrossRef]
- Weiss, M.C.; Sousa, F.L.; Mrnjavac, N.; Neukirchen, S.; Roettger, M.; Nelson-Sathi, S.; Martin, W.F. The physiology and habitat of the last universal common ancestor. Nat. Microbiol. 2016, 1, 16116. [Google Scholar] [CrossRef]
- Sohrabi, S.; Kassir, N.; Keshavarz Moraveji, M. Droplet microfluidics: Fundamentals and its advanced applications. RSC Adv. 2020, 10, 27560–27574. [Google Scholar] [CrossRef]
- Castro-Hernández, E.; Gundabala, V.; Fernández-Nieves, A.; Gordillo, J.M. Scaling the drop size in coflow experiments. New J. Phys. 2009, 11, 075021. [Google Scholar] [CrossRef]
- Teh, S.-Y.; Lin, R.; Hung, L.-H.; Lee, A.P. Droplet microfluidics. Lab Chip 2008, 8, 198–220. [Google Scholar] [CrossRef]
- Chong, Z.Z.; Tan, S.H.; Gañán-Calvo, A.M.; Tor, S.B.; Loh, N.H.; Nguyen, N.-T. Active droplet generation in microfluidics. Lab Chip 2016, 16, 35–58. [Google Scholar] [CrossRef]
- Dertinger, S.K.W.; Chiu, D.T.; Jeon, N.L.; Whitesides, G.M. Generation of Gradients Having Complex Shapes Using Microfluidic Networks. Anal. Chem. 2001, 73, 1240–1246. [Google Scholar] [CrossRef]
- Um, E.; Rogers, M.E.; Stone, H.A. Combinatorial generation of droplets by controlled assembly and coalescence. Lab Chip 2013, 13, 4674–4680. [Google Scholar] [CrossRef]
- Lucchetta, E.M.; Munson, M.S.; Ismagilov, R.F. Characterization of the local temperature in space and time around a developing Drosophila embryo in a microfluidic device. Lab Chip 2006, 6, 185–190. [Google Scholar] [CrossRef]
- Song, H.; Tice, J.D.; Ismagilov, R.F. A microfluidic system for controlling reaction networks in time. Angew. Chem. Int. Ed. Engl. 2003, 42, 768–772. [Google Scholar] [CrossRef]
- Gu, H.; Duits, M.H.G.; Mugele, F. Droplets formation and merging in two-phase flow microfluidics. Int. J. Mol. Sci. 2011, 12, 2572–2597. [Google Scholar] [CrossRef] [PubMed]
- Abate, A.R.; Hung, T.; Mary, P.; Agresti, J.J.; Weitz, D.A. High-throughput injection with microfluidics using picoinjectors. Proc. Natl. Acad. Sci. USA 2010, 107, 19163–19166. [Google Scholar] [CrossRef] [PubMed]
- O’Donovan, B.; Eastburn, D.J.; Abate, A.R. Electrode-free picoinjection of microfluidic drops. Lab Chip 2012, 12, 4029–4032. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.-C.; Ho, Y.L.; Lee, A.P. Microfluidic sorting of droplets by size. Microfluid. Nanofluid. 2008, 4, 343–348. [Google Scholar] [CrossRef]
- Yoon, D.H.; Tanaka, D.; Sekiguchi, T.; Shoji, S. Size-Dependent and Property-Independent Passive Microdroplet Sorting by Droplet Transfer on Dot Rails. Micromachines 2018, 9, 513. [Google Scholar] [CrossRef]
- Huh, D.; Bahng, J.H.; Ling, Y.; Wei, H.-H.; Kripfgans, O.D.; Fowlkes, J.B.; Grotberg, J.B.; Takayama, S. Gravity-Driven Microfluidic Particle Sorting Device with Hydrodynamic Separation Amplification. Anal. Chem. 2007, 79, 1369–1376. [Google Scholar] [CrossRef]
- Wang, L.; Flanagan, L.A.; Monuki, E.; Jeon, N.L.; Lee, A.P. Dielectrophoresis switching with vertical sidewall electrodes for microfluidic flow cytometry. Lab Chip 2007, 7, 1114–1120. [Google Scholar] [CrossRef]
- Turk-MacLeod, R.; Henson, A.; Rodriguez-Garcia, M.; Gibson, G.M.; Aragon Camarasa, G.; Caramelli, D.; Padgett, M.J.; Cronin, L. Approach to classify, separate, and enrich objects in groups using ensemble sorting. Proc. Natl. Acad. Sci. USA 2018, 115, 5681–5685. [Google Scholar] [CrossRef]
- Theberge, A.B.; Mayot, E.; El Harrak, A.; Kleinschmidt, F.; Huck, W.T.S.; Griffiths, A.D. Microfluidic platform for combinatorial synthesis in picolitre droplets. Lab Chip 2012, 12, 1320–1326. [Google Scholar] [CrossRef]
- Lucchetta, E.M.; Lee, J.H.; Fu, L.A.; Patel, N.H.; Ismagilov, R.F. Dynamics of Drosophila embryonic patterning network perturbed in space and time using microfluidics. Nature 2005, 434, 1134–1138. [Google Scholar] [CrossRef]
- Joensson, H.N.; Andersson Svahn, H. Droplet microfluidics—A tool for single-cell analysis. Angew. Chem. Int. Ed. Engl. 2012, 51, 12176–12192. [Google Scholar] [CrossRef]
- Zheng, B.; Roach, L.S.; Ismagilov, R.F. Screening of protein crystallization conditions on a microfluidic chip using nanoliter-size droplets. J. Am. Chem. Soc. 2003, 125, 11170–11171. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhu, L.-N.; Guo, R.; Cui, H.-J.; Ye, S.; Fang, Q. Nanoliter-scale protein crystallization and screening with a microfluidic droplet robot. Sci. Rep. 2014, 4, 5046. [Google Scholar] [CrossRef]
- Ahrberg, C.D.; Manz, A.; Chung, B.G. Polymerase chain reaction in microfluidic devices. Lab Chip 2016, 16, 3866–3884. [Google Scholar] [CrossRef]
- Chiu, F.W.Y.; Stavrakis, S. High-throughput droplet-based microfluidics for directed evolution of enzymes. Electrophoresis 2019, 40, 2860–2872. [Google Scholar] [CrossRef]
- Agresti, J.J.; Antipov, E.; Abate, A.R.; Ahn, K.; Rowat, A.C.; Baret, J.-C.; Marquez, M.; Klibanov, A.M.; Griffiths, A.D.; Weitz, D.A. Ultrahigh-throughput screening in drop-based microfluidics for directed evolution. Proc. Natl. Acad. Sci. USA 2010, 107, 4004–4009. [Google Scholar] [CrossRef]
- Ma, F.; Chung, M.T.; Yao, Y.; Nidetz, R.; Lee, L.M.; Liu, A.P.; Feng, Y.; Kurabayashi, K.; Yang, G.-Y. Efficient molecular evolution to generate enantioselective enzymes using a dual-channel microfluidic droplet screening platform. Nat. Commun. 2018, 9, 1030. [Google Scholar] [CrossRef]
- Hanczyc, M.M. Metabolism and motility in prebiotic structures. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2011, 366, 2885–2893. [Google Scholar] [CrossRef] [PubMed]
- Do Nascimento Vieira, A.; Kleinermanns, K.; Martin, W.F.; Preiner, M. The ambivalent role of water at the origins of life. FEBS Lett. 2020, 594, 2717–2733. [Google Scholar] [CrossRef]
- Allmon, W.D. Species, speciation and palaeontology up to the Modern Synthesis: Persistent themes and unanswered questions. Palaeontology 2013, 56, 1199–1223. [Google Scholar] [CrossRef]
- Ringo, J.; Wood, D.; Rockwell, R.; Dowse, H. An Experiment Testing Two Hypotheses of Speciation. Am. Nat. 1985, 126, 642–661. [Google Scholar] [CrossRef]
- Rice, W.R.; Hostert, E.E. Laboratory Experiments on Speciation: What Have We Learned in 40 Years? Evolution 1993, 47, 1637–1653. [Google Scholar] [CrossRef]

| Term | Definition |
|---|---|
| Allele | A version of a gene which may contain variations different from other versions of the same gene. In some biological systems, more than one gene variant exists (for example, two of each chromosome exists in biploid organisms like humans, while multiple forms of each chromosome exist in polyploid organisms like wheat). An allele is simply one of these gene variants. |
| Allopatry or Allopatric Speciation | Speciation resulting from geographical isolation of two populations of the same original species. |
| Autotrophic | A process by which a system obtains energy that drives its own function (such as replication, growth, evolution, etc.) directly from an inorganic source (such as minerals, geological processes, sun, etc.). |
| Autotrophic Origin of Life | A theory assuming that life originated from geochemical processes patterned after non-enzymatic metabolic reactions, most commonly, the reverse Krebs cycle. |
| Biological Concept of Species (BCS) | Groups of actually or potentially interbreeding natural populations, which are reproductively isolated from other such groups. |
| Biological Speciation | The process by which biological lineages diverge from their ancestors to become new biological species as a result of differential selective pressures, geographical isolation, or other processes. |
| Chemical Speciation | The process by which delimited chemical systems of similar makeup and properties are subjected to differential environments or chemical/physical conditions, resulting in emergence of more than one distinct delimited chemical system with different makeup and properties from each other. |
| Coacervates | Macromolecule-rich dispersed droplets in liquid–liquid phase separated systems. According to Oparin and Haldane, coacervates constitute protocells. |
| Compartmentalization | The physical process by which an analyte stably resides within a compartment for long periods of time and does not rapidly depart from that compartment on short timescales. |
| Complex Coacervates | A coacervate, but specifically one resulting from phase separation caused by electrostatic binding of oppositely charged polyions. |
| Composome | The compositional information of a collection of molecules before the emergence of life, often referred to as part of the graded autocatalysis replication domain (GARD) model, which assumes that primitive functions such as replication and catalysis could have existed before the emergence of replicating polymers. |
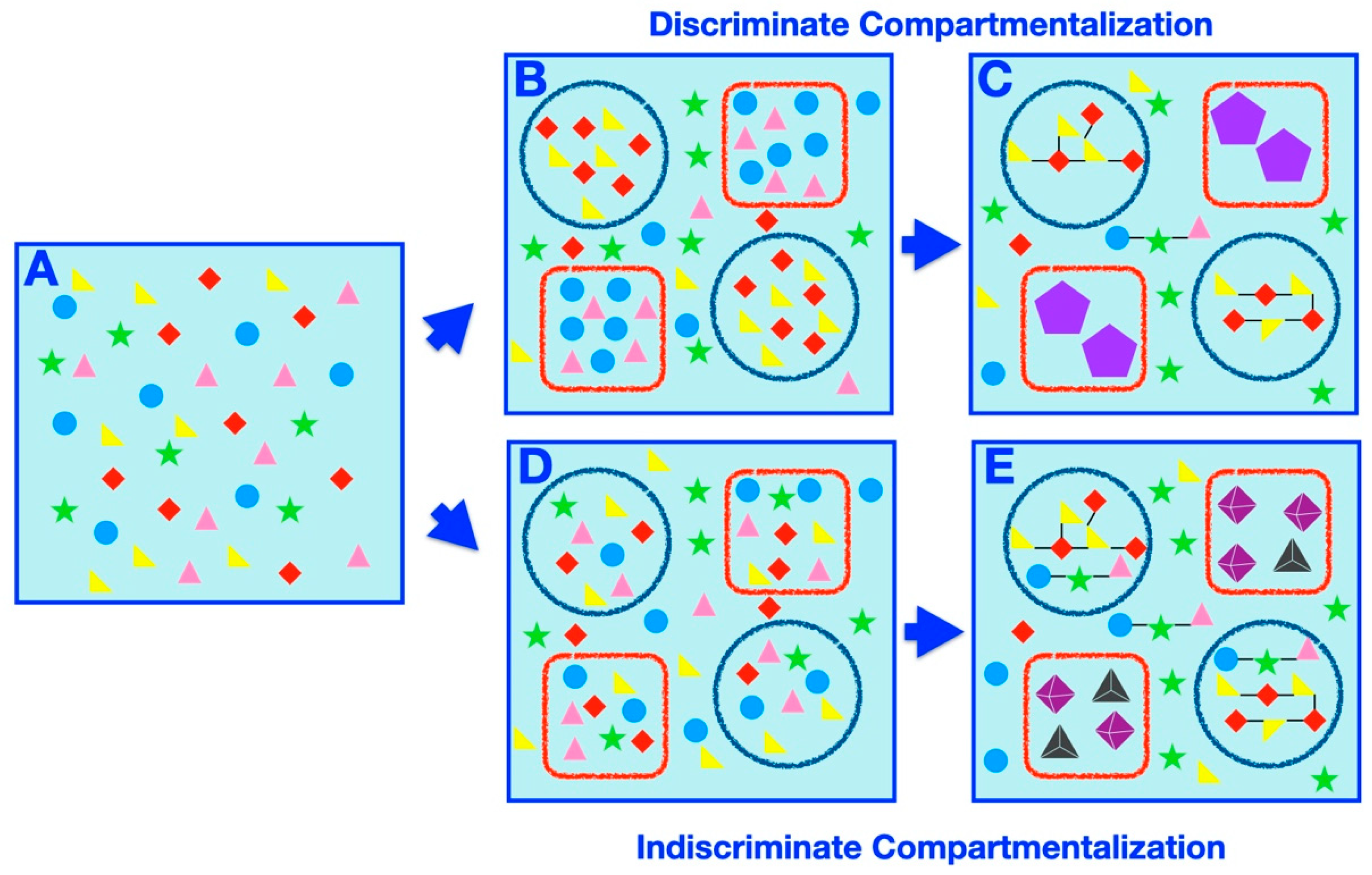
| Discriminate Compartmentalization | The process by which specific subsets of molecules are placed within specific compartments due to systematic factors (such as physicochemical properties of the molecules and/or the compartments). |
| Ecological Niches | The placement of an organism within a specific ecological community, and the various interactions this organism has with the environment and other members of the community. |
| GARD (Graded Autocatalysis Replication Domain) | A general kinetic model in evolutionary biology for homeostatic growth and fission of compositional assemblies with specific application toward lipids. |
| Heterotrophic | A process by which a system obtains energy that drives its own function (such as replication, growth, evolution, etc.) from other non-inorganic sources (such as other chemicals, polymers, organisms, etc.). |
| Heterotrophic Origin of Life | A theory assuming that early organisms depended on abiotically synthesized organic molecules for their structural components and as an energy source. |
| Homologous Recombination | Genetic information exchange occurring between two identical (or nearly identical) gene-carrying forms (alleles). For example, this would occur between two different alleles of homologous (identical) genes between similar organisms, akin to sexual reproduction. |
| Horizontal Gene Transfer | The transfer of genetic information not acquired directly from a parent but rather from other members of the same (or similar) community of organisms that may come from the same lineage. |
| Indiscriminate Compartmentalization | The process by which random samplings of molecules are partitioned within compartments arbitrarily, independent of the physicochemical properties of the molecules or the compartments. |
| Liquid–liquid Phase Separation (LLPS) | The process by which a mixture two liquids separate from each other (due to thermodynamic reasons such as immiscibility, differences in chemical affinities, entropic effects, etc.), resulting in two immiscible liquids co-existing. |
| LUCA (Last Common Universal Ancestor) | The earliest single-celled form of life on Earth which eventually evolved into all known forms of modern life. |
| Membrane Liposome/Vesicle | A specific compartment type that has been proposed as a protocell and primitive compartment model. |
| Messy Chemistry | A field of prebiotic chemistry which assumes that the prebiotic milieu is sufficiently complex (i.e., “messy”), and whose experiments focus on bulk properties of complex prebiotic chemical mixtures (such as emergent functions or reactions) as opposed to characterizing exact mechanisms of primitive processes (it is impossible to study such processes on the actual prebiotic Earth). |
| Nonhomologous Recombination | Genetic information exchange which occurs as a result of ligation of two (or more) essentially unrelated genes or nucleic acids. |
| Parapatry or Parapatric Speciation | Speciation resulting from partial geographical separation afforded to diverging populations. |
| Peripatry or Peripatric Speciation | A form of allopatric speciation in which a small population is isolated on the “periphery” of a large population, and the reduced population eventually can no longer breed with the main population. |
| Phenotypic | An observable or expressed trait in any organism, species, system, etc., that may be governed by its genetic, chemical, structural, etc. composition. |
| Primitive Compartment | A compartment that can be assembled using prebiotically available building blocks on early Earth; this compartment need not have directly evolved into modern cells and may have provided other functions to primitive chemical systems. |
| Progenote | A proposed form that early life on Earth could have taken on. Rather than a single-celled LUCA, a progenote was a collection of cells which communicated and evolved through horizontal transfer of encapsulated components (i.e., genetic/catalytic biopolymers). |
| Protocell | A primitive compartment that necessarily evolved directly into modern cells. A protocell is a specific type of primitive compartment, but a primitive compartment is not necessarily a protocell. |
| Protolife Systems | These are systems which emerged before the origin of life and necessarily evolved into modern life. They may have been composed of protocells, or of a collection of protocells, or simply of a collection of chemicals. |
| Random Genetic Drift | Random fluctuations in the frequency of expression of genes within a population (such as variation of alleles) resulting from random statistical occurrences (such as sudden reduction of population size resulting in more variation of gene expression). |
| Sympatry or Sympatric Speciation | Speciation resulting from an absence of geographical separation of any form. |
| Taxonomy | The study of classifying organisms based on (generally) patterns of their genetic inheritance. Taxonomy can also be applied to classifying chemical systems based on chemical properties or structures. |
| Vertical Transmission | The transfer of genetic information directly from a parent organism to an offspring organism. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, T.Z.; Caudan, M.; Mamajanov, I. Origin of Species before Origin of Life: The Role of Speciation in Chemical Evolution. Life 2021, 11, 154. https://doi.org/10.3390/life11020154
Jia TZ, Caudan M, Mamajanov I. Origin of Species before Origin of Life: The Role of Speciation in Chemical Evolution. Life. 2021; 11(2):154. https://doi.org/10.3390/life11020154
Chicago/Turabian StyleJia, Tony Z., Melina Caudan, and Irena Mamajanov. 2021. "Origin of Species before Origin of Life: The Role of Speciation in Chemical Evolution" Life 11, no. 2: 154. https://doi.org/10.3390/life11020154
APA StyleJia, T. Z., Caudan, M., & Mamajanov, I. (2021). Origin of Species before Origin of Life: The Role of Speciation in Chemical Evolution. Life, 11(2), 154. https://doi.org/10.3390/life11020154