Multi-Scale and Multi-Branch Convolutional Neural Network for Retinal Image Segmentation
Abstract
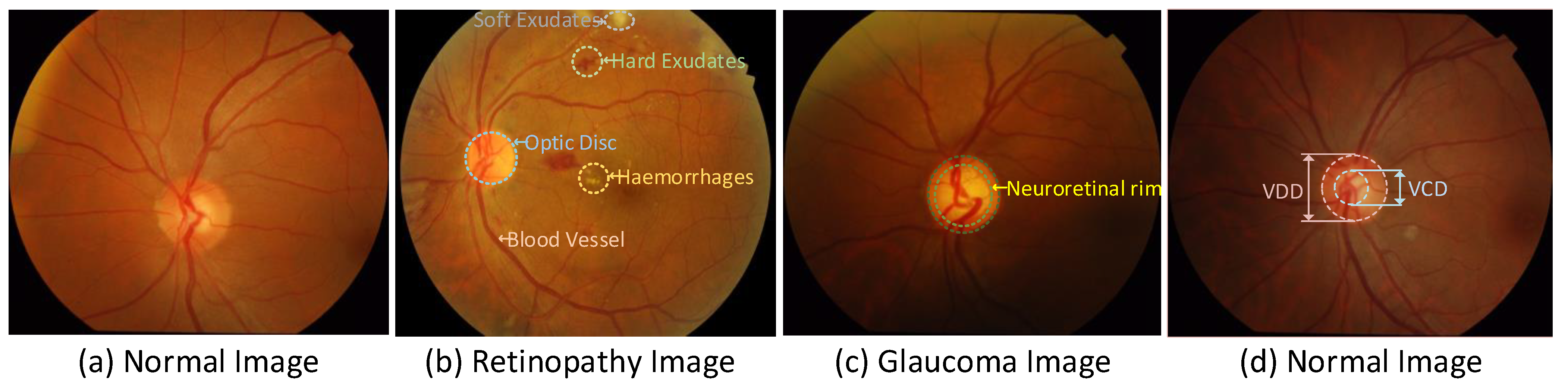
1. Introduction
- We propose an effective multi-scale and multi-branch network (MSMB-Net) model for the automatic segmentation of retinal vessels. The proposed network model is similarly used for the accurate joint segmentation of optic disc and optic cup;
- MSMB-Net has the following advantages: (a) The multi-scale context information fusion module uses skip connections and different expansion ratios of atrous convolution to improve the model’s full understanding of local context information. It improves the feature extraction ability of the network structure and maintains the correlation of features in the receptive field; (b) The multi-branch convolution module combines convolutions of different receptive field sizes to improve the sensitivity to global context information; (c) Side-out rebuilding layer aggregates the effective features of different stages to improve the network learning ability without adding additional parameters and calculations;
- The network model proposed in this paper is tested on the DRIVE, STARE, CHASE_DB1 and Drishti-GS1 datasets. The proposed MSMB-Net can obtain the most advanced results, which proves the robustness and effectiveness of the method.
2. Method
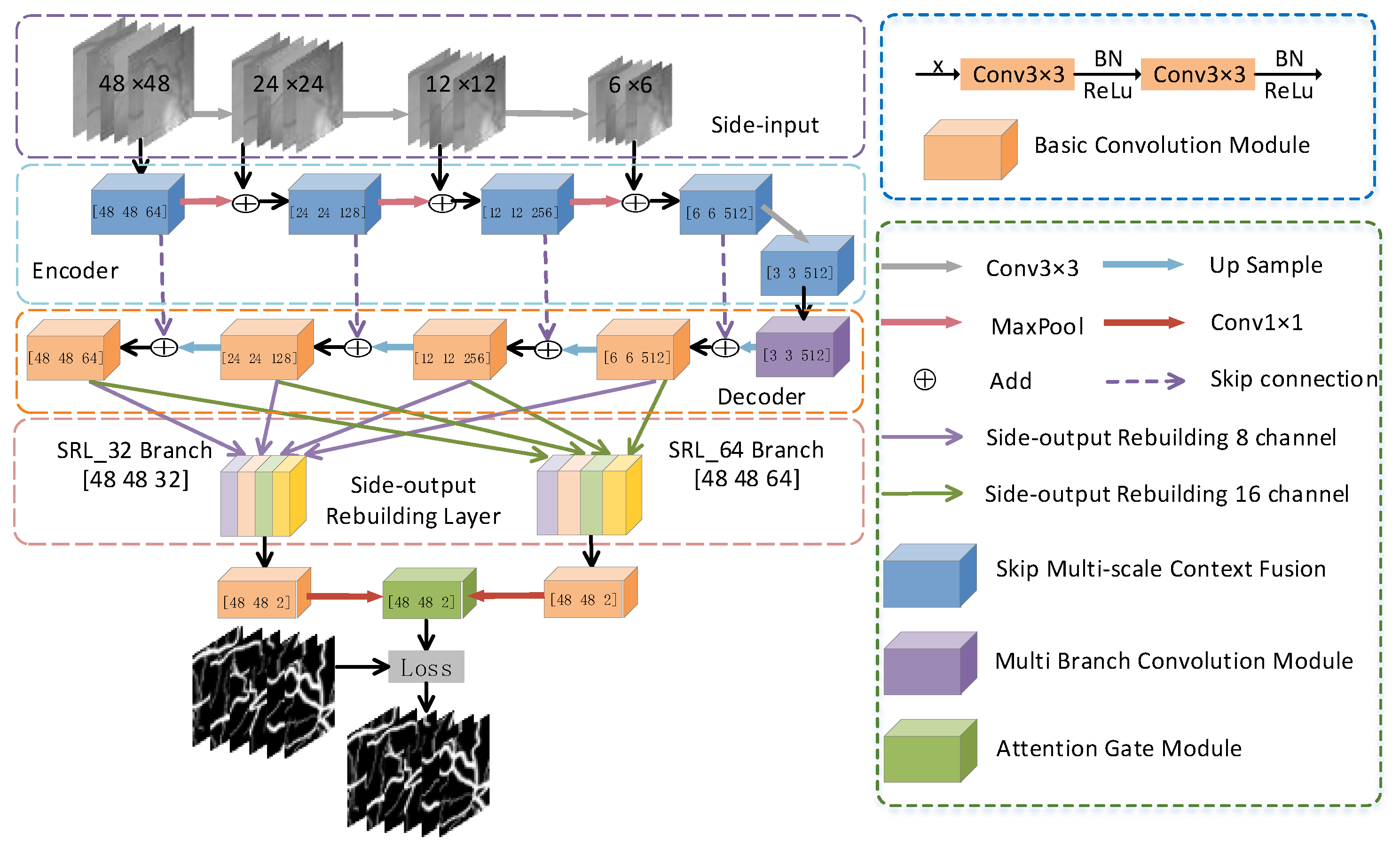
2.1. Network Structure
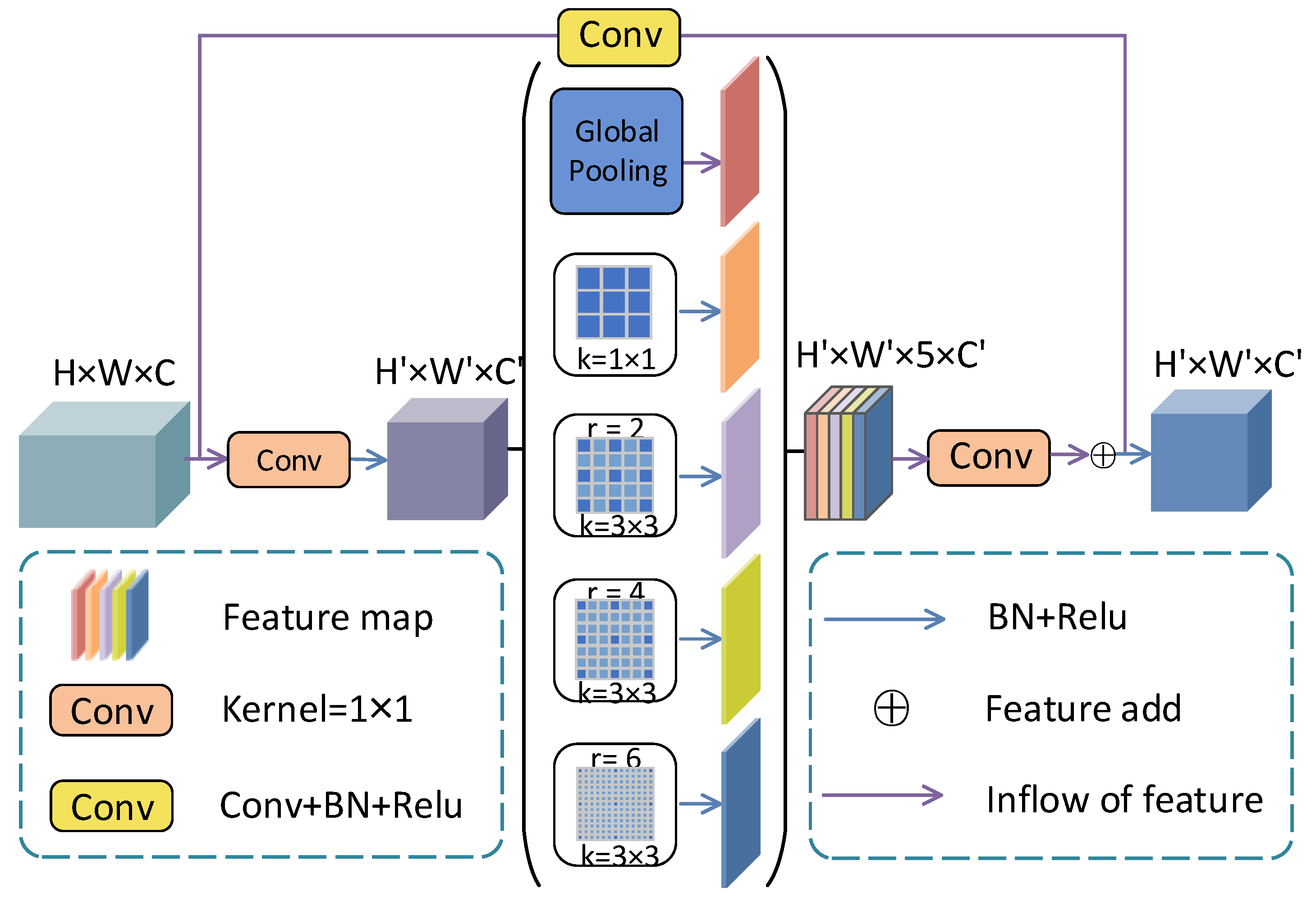
2.2. Multi-Scale Context Fusion Module
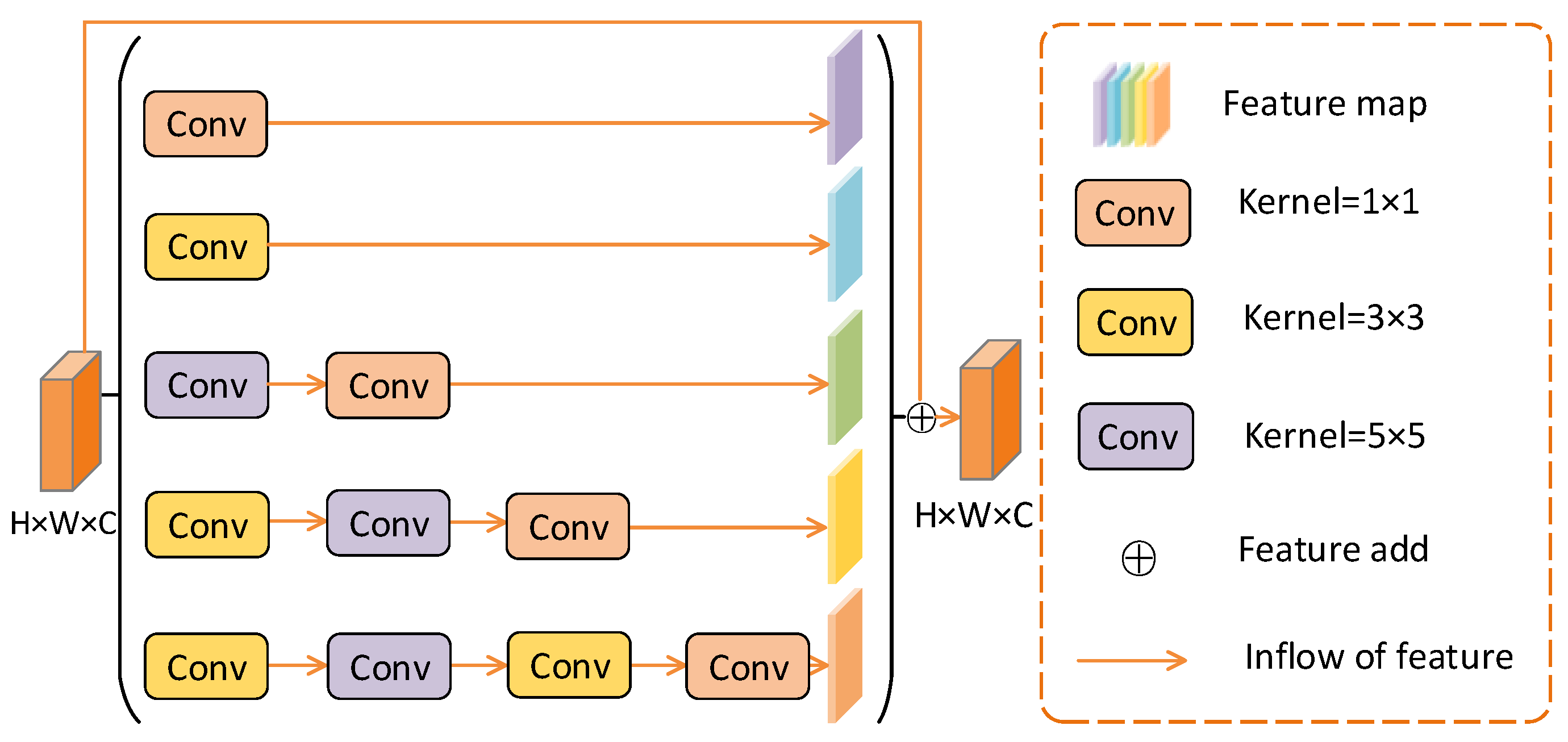
2.3. Multi-Branch Convolution Module
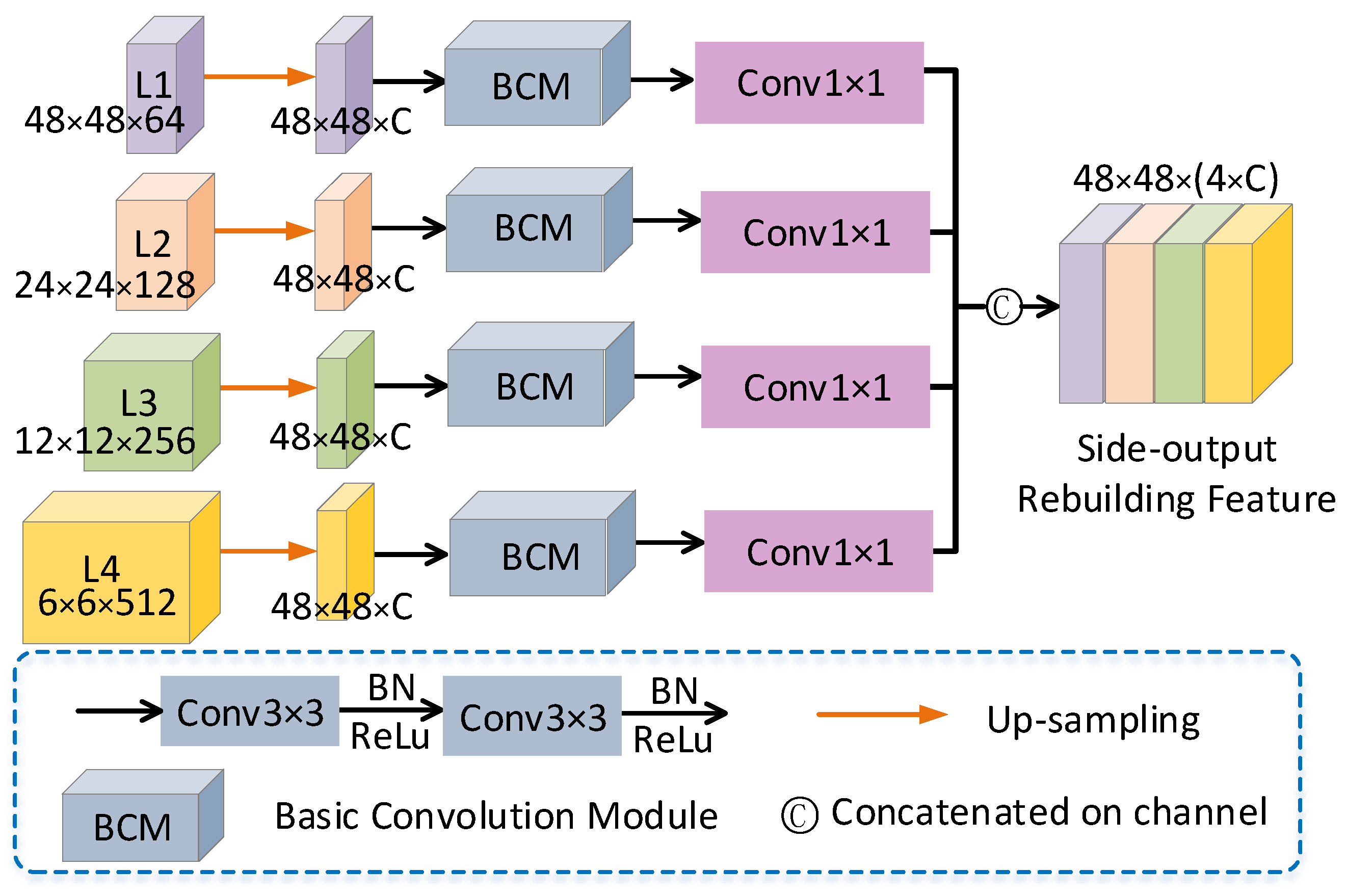
2.4. Side-Output Rebuilding Layer
| Algorithm 1: Side-output Rebuilding Layer |
Input: Feature map:. Batch of feature maps: N. Height of the feature map:h. Width of the feature map:w. Channel c of the feature map:c. Downsampling factor:d. Output: Feature map after scale rebuilding:
|
2.5. Attention Module
3. Dataset and Evaluation
3.1. Dataset
3.2. Implementation Details
3.3. Performance Evaluation
4. Experimental Results and Discussion
4.1. Compare the Results of the Improved Model
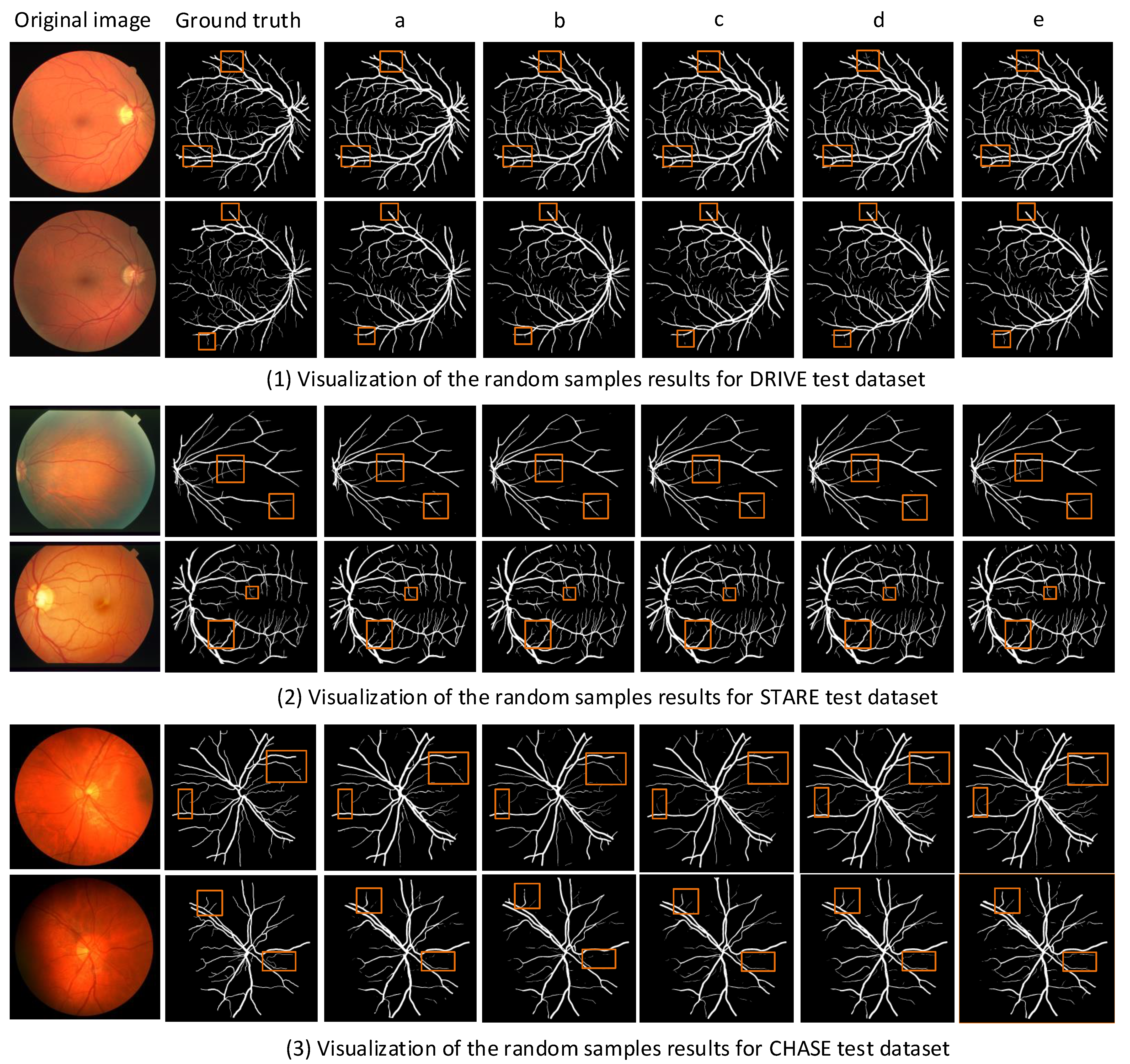
4.2. Retinal Vessel Segmentation
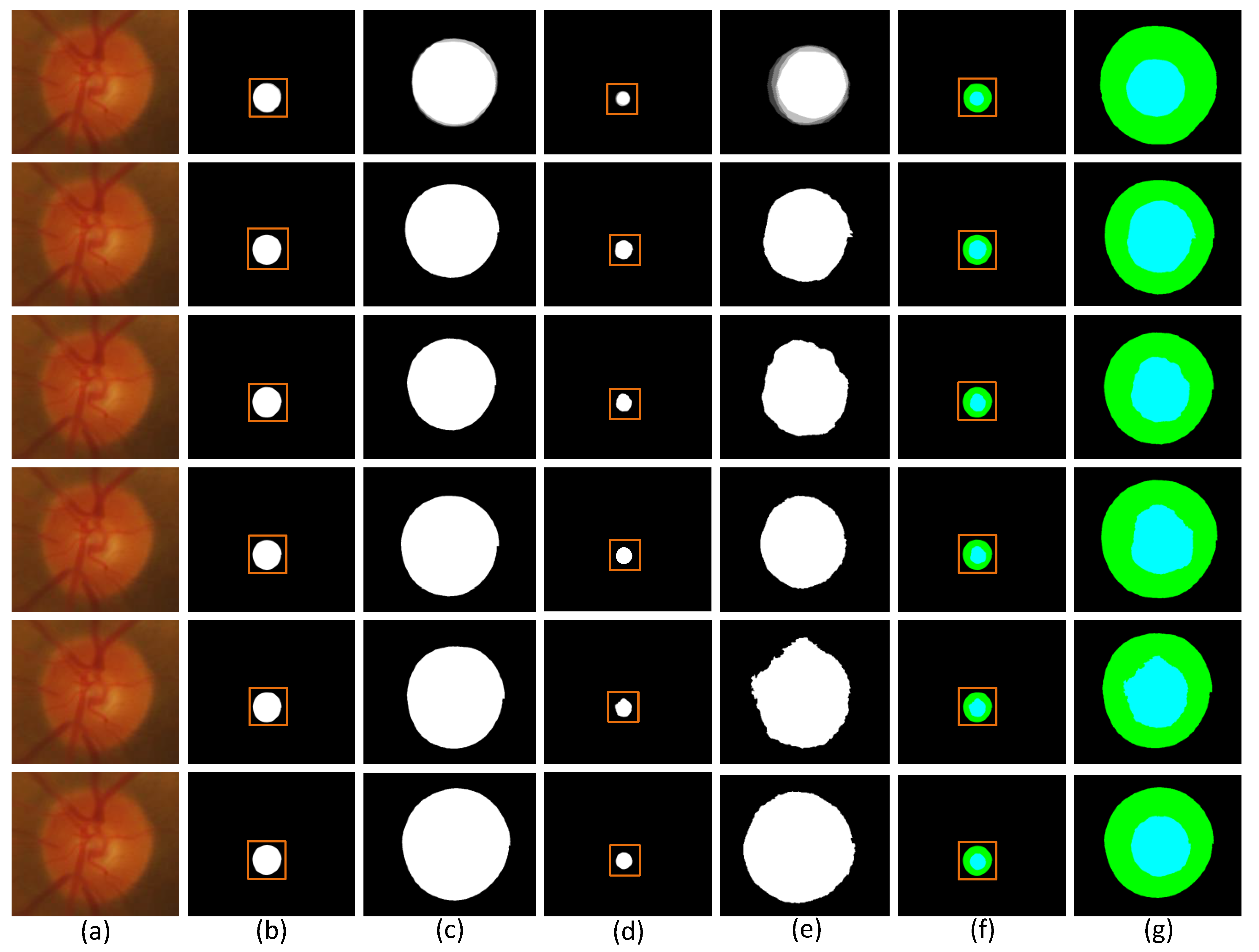
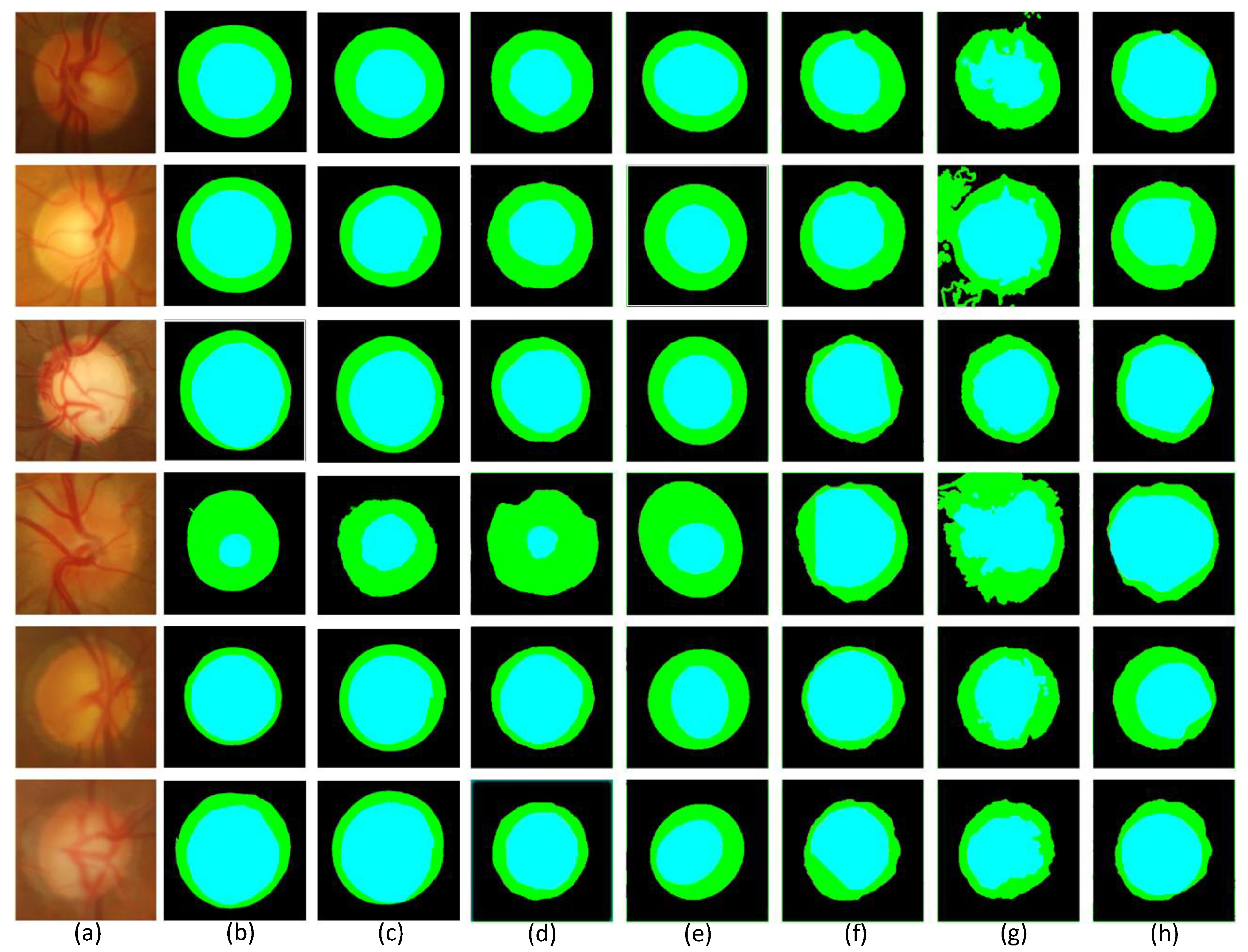
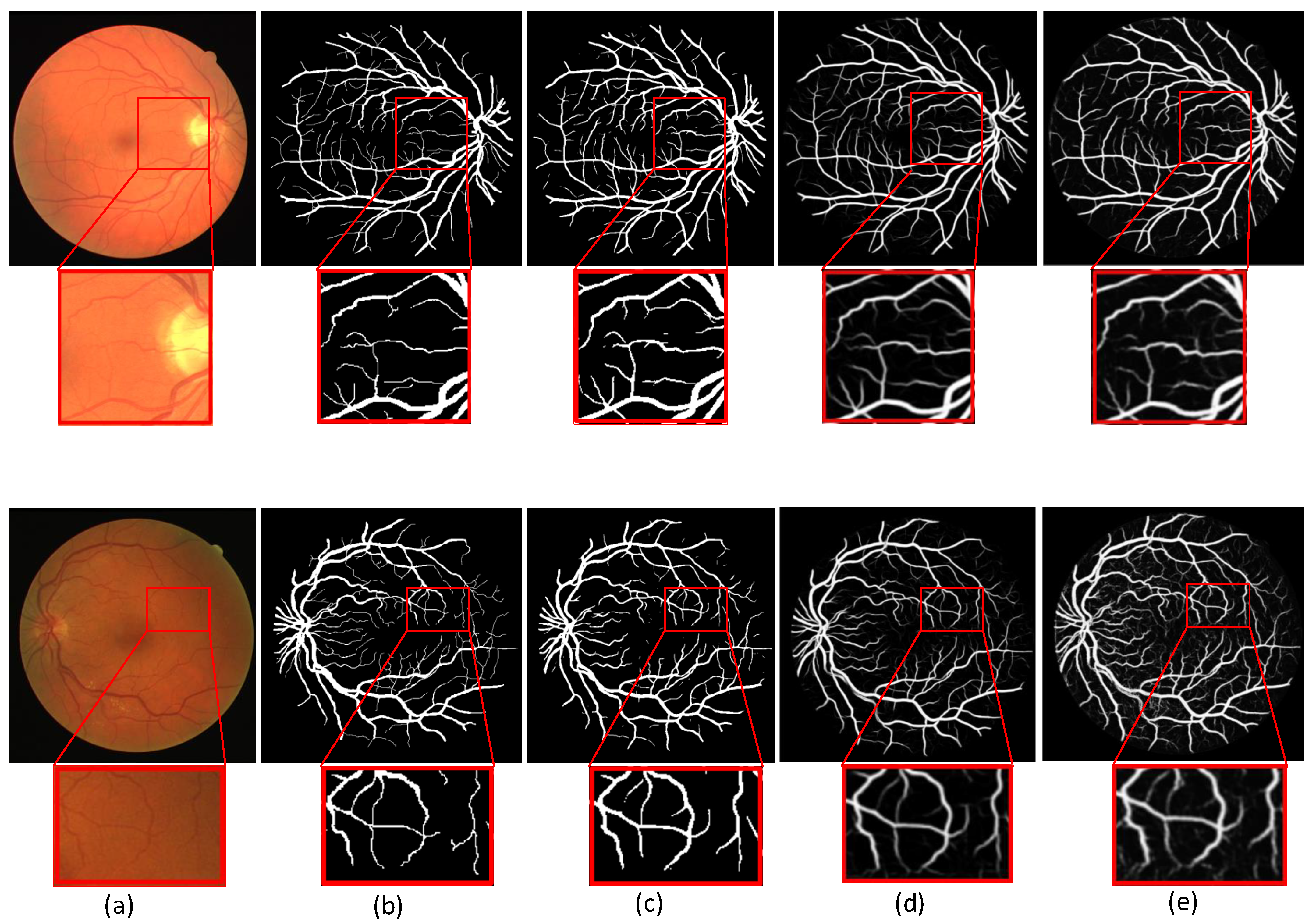
4.3. Optic Disc and Optic Cup Comparison of Different Methods
4.4. Different Segmentation Quantitative Analysis of the Results
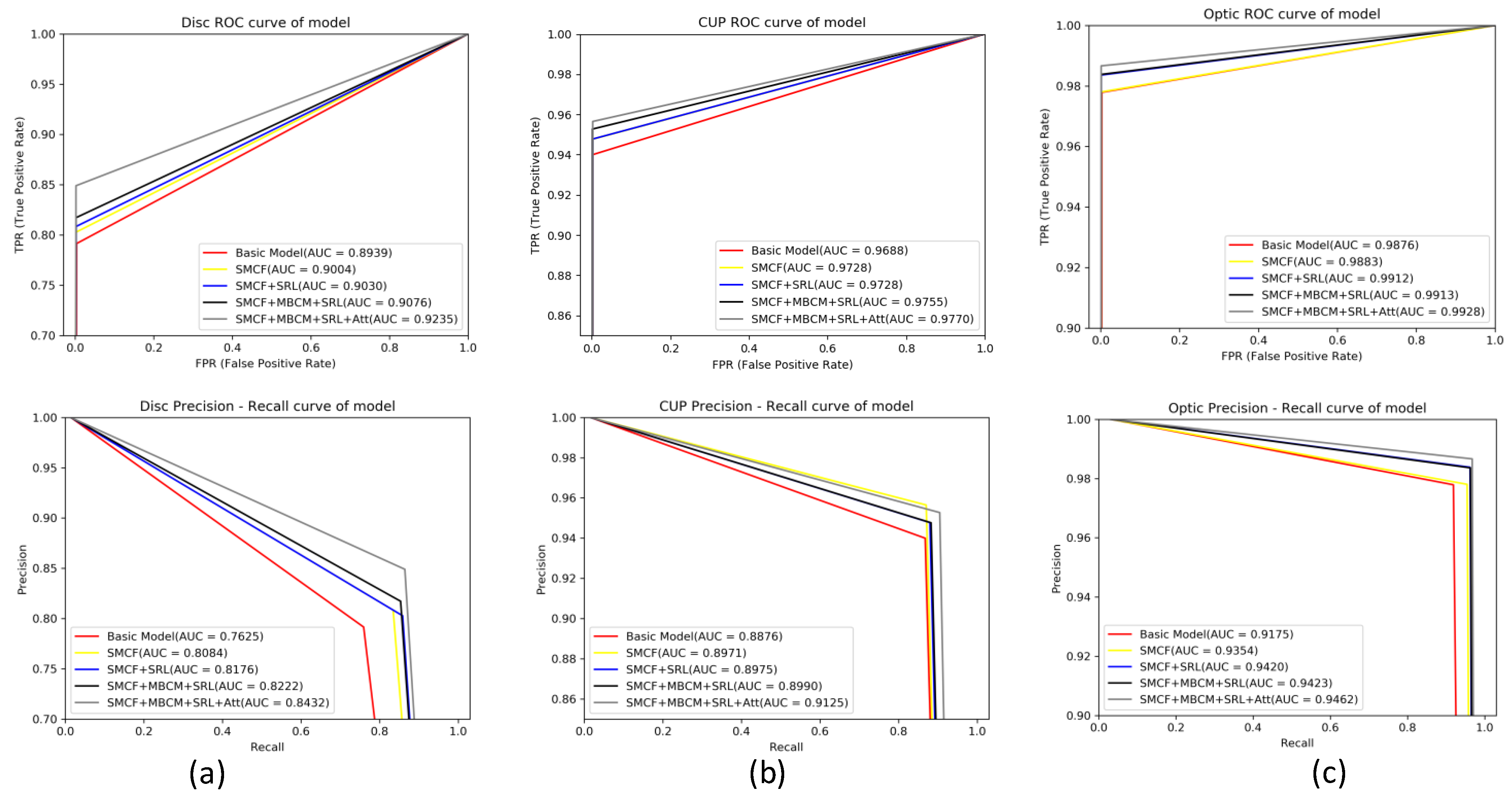
4.5. Evaluation of ROC Curve and PR Curve
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Khan, K.B.; Khaliq, A.A.; Jalil, A.; Iftikhar, M.A.; Ullah, N.; Aziz, M.W.; Ullah, K.; Shahid, M. A review of retinal blood vessels extraction techniques: challenges, taxonomy, and future trends. Pattern Anal. Appl. 2019, 22, 767–802. [Google Scholar] [CrossRef]
- Franklin, S.W.; Rajan, S.E. Computerized screening of diabetic retinopathy employing blood vessel segmentation in retinal images. Biocybern. Biomed. Eng. 2014, 34, 117–124. [Google Scholar] [CrossRef]
- Jonas, J.B.; Bergua, A.; Schmitz-Valckenberg, P.; Papastathopoulos, K.I.; Budde, W.M. Ranking of optic disc variables for detection of glaucomatous optic nerve damage. Investig. Ophthalmol. Vis. Sci. 2000, 41, 1764. [Google Scholar]
- Miao, Y.; Cheng, Y. Automatic extraction of retinal blood vessel based on matched filtering and local entropy thresholding. In Proceedings of the 8th International Conference on Biomedical Engineering and Informatics (BMEI), Shenyang, China, 14–16 October 2015; pp. 62–67. [Google Scholar]
- Kundu, A.; Chatterjee, R.K. Retinal vessel segmentation using Morphological Angular Scale-Space. In Proceedings of the 2012 Third International Conference on Emerging Applications of Information Technology, Kolkata, India, 30 November–1 December 2012; pp. 316–319. [Google Scholar] [CrossRef]
- Palomera-Perez, M.A.; Martinez-Perez, M.E.; Benitez-Perez, H. Parallel Multiscale Feature Extraction and Region Growing: Application in Retinal Blood Vessel Detection. IEEE Trans. Inf. Technol. Biomed. 2010, 14, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Azzopardi, G.; Strisciuglio, N.; Vento, M.; Petkov, N. Trainable COSFIRE filters for vessel delineation with application to retinal images. Med. Image Anal. 2015, 19, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Kumar, K.; Samal, D. Automated retinal vessel segmentation based on morphological preprocessing and 2D-Gabor wavelets. In Advanced Computing and Intelligent Engineering; Springer: Singapore, 2020; pp. 411–423. [Google Scholar]
- Tian, C.; Fang, T.; Fan, Y.; Wu, W. Multi-path convolutional neural network in fundus segmentation of blood vessels. Biocybern. Biomed. Eng. 2020, 40, 583–595. [Google Scholar] [CrossRef]
- Jainish, G.R.; Jiji, G.W.; Infant, P.A. A novel automatic retinal vessel extraction using maximum entropy based EM algorithm. Multimed. Tools Appl. 2020, 79, 22337–22353. [Google Scholar] [CrossRef]
- Marín, D.; Aquino, A.; Gegúndez-Arias, M.E.; Bravo, J.M. A new supervised method for blood vessel segmentation in retinal images by using gray-level and moment invariants-based features. IEEE Trans. Med. Imaging 2010, 30, 146–158. [Google Scholar] [CrossRef] [PubMed]
- Aslani, S.; Sarnel, H. A new supervised retinal vessel segmentation method based on robust hybrid features. Biomed. Signal Process. Control 2016, 30, 1–12. [Google Scholar] [CrossRef]
- Feng, Z.; Yang, J.; Yao, L. Patch-based fully convolutional neural network with skip connections for retinal blood vessel segmentation. In Proceedings of the IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 1742–1746. [Google Scholar]
- Mo, J.; Zhang, L. Multi-level deep supervised networks for retinal vessel segmentation. Int. J. Comput. Assist. Radiol. Surg. 2017, 12, 2181–2193. [Google Scholar] [CrossRef]
- Hu, K.; Zhang, Z.; Niu, X.; Zhang, Y.; Cao, C.; Xiao, F.; Gao, X. Retinal vessel segmentation of color fundus images using multiscale convolutional neural network with an improved cross-entropy loss function. Neurocomputing 2018, 39, 179–191. [Google Scholar] [CrossRef]
- Yan, Z.; Yang, X.; Cheng, K.T. A three-stage deep learning model for accurate retinal vessel segmentation. IEEE J. Biomed. Health Inform. 2018, 23, 1427–1436. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2015; pp. 234–241. [Google Scholar]
- Alom, M.Z.; Hasan, M.; Yakopcic, C.; Taha, T.M.; Asari, V.K. Recurrent residual convolutional neural network based on u-net (r2u-net) for medical image segmentation. arXiv 2018, arXiv:1802.06955. [Google Scholar]
- Li, L.; Verma, M.; Nakashima, Y.; Nagahara, H.; Kawasaki, R. Iternet: Retinal image segmentation utilizing structural redundancy in vessel networks. In Proceedings of the IEEE Winter Conference on Applications of Computer Vision (WACV), Snowmass, CO, USA, 1–5 March 2020; pp. 3656–3665. [Google Scholar]
- Jin, Q.; Meng, Z.; Pham, T.D.; Chen, Q.; Wei, L.; Su, R. DUNet: A deformable network for retinal vessel segmentation. Knowl.-Based Syst. 2019, 178, 149–162. [Google Scholar] [CrossRef]
- Atli, İ.; Gedik, O.S. Sine-Net: A fully convolutional deep learning architecture for retinal blood vessel segmentation. Eng. Sci. Technol. Int. J. 2020, in press. [Google Scholar]
- Wang, D.; Haytham, A.; Pottenburgh, J.; Saeedi, O.; Tao, Y. Hard Attention Net for Automatic Retinal Vessel Segmentation. IEEE J. Biomed. Health Inform. 2020, 24, 3384–3396. [Google Scholar] [CrossRef]
- Zilly, J.G.; Buhmann, J.M.; Mahapatra, D. Boosting convolutional filters with entropy sampling for optic cup and disc image segmentation from fundus images. In International Workshop on Machine Learning in Medical Imaging; Springer: Cham, Switzerland, 2015; pp. 136–143. [Google Scholar]
- Sevastopolsky, A.; Drapak, S.; Kiselev, K.; Snyder, B.M.; Keenan, J.D.; Georgievskaya, A. Stack-u-net: Refinement network for image segmentation on the example of optic disc and cup. arXiv 2018, arXiv:1804.11294. [Google Scholar]
- Chakravarty, A.; Sivaswamy, J. RACE-net: A recurrent neural network for biomedical image segmentation. IEEE J. Biomed. Health Inform. 2018, 23, 1151–1162. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.; Kasukurthi, N.; Pande, H. Dynamic Region Proposal Networks For Semantic Segmentation In Automated Glaucoma Screening. In Proceedings of the IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Venice, Italy, 8–11 April 2019; pp. 578–582. [Google Scholar]
- Yu, S.; Xiao, D.; Frost, S.; Kanagasingam, Y. Robust optic disc and cup segmentation with deep learning for glaucoma detection. Comput. Med. Imaging Graph. 2019, 74, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Ding, F.; Yang, G.; Liu, J.; Wu, J.; Ding, D.; Xv, J.; Cheng, G.; Li, X. Hierarchical Attention Networks for Medical Image Segmentation. arXiv 2019, arXiv:1911.08777. [Google Scholar]
- Gu, Z.; Cheng, J.; Fu, H.; Zhou, K.; Hao, H.; Zhao, Y.; Zhang, T.; Gao, S.; Liu, J. Ce-net: Context encoder network for 2d medical image segmentation. IEEE Trans. Med. Imaging 2019, 38, 2281–2292. [Google Scholar] [CrossRef] [PubMed]
- Kadambi, S.; Wang, Z.; Xing, E. WGAN domain adaptation for the joint optic disc-and-cup segmentation in fundus images. Int. J. Comput. Assist. Radiol. Surg. 2020. [Google Scholar] [CrossRef]
- Tabassum, M.; Khan, T.M.; Arslan, M.; Naqvi, S.S. CDED-Net: Joint Segmentation of Optic Disc and Optic Cup for Glaucoma Screening. IEEE Access 2020. [Google Scholar] [CrossRef]
- Chen, L.C.; Papandreou, G.; Schroff, F.; Adam, H. Rethinking atrous convolution for semantic image segmentation. arXiv 2017, arXiv:1706.05587. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016. [Google Scholar]
- Chen, L.-C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 40, 834–848. [Google Scholar] [CrossRef]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2881–2890. [Google Scholar]
- Woo, S.; Park, J.; Lee, J.-Y.; Kweon, I.S. Cbam: Convolutional block attention module. In Proceedings of the European Conference on Computer Vision (ECCV); Springer: Cham, Switzerland, 2018. [Google Scholar] [CrossRef]
- Staal, J.; Abràmoff, M.D.; Niemeijer, M.; Viergever, M.A.; van Ginneken, B. Ridge-based vessel segmentation in color images of the retina. IEEE Trans. Med. Imaging 2004, 23, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Hoover, A.D.; Kouznetsova, V.; Goldbaum, M. Locating blood vessels in retinal images by piecewise threshold probing of a matched filter response. IEEE Trans. Med Imaging 2000, 19, 203–210. [Google Scholar] [CrossRef]
- Owen, C.G.; Rudnicka, A.R.; Mullen, R.; Barman, S.A.; Monekosso, D.; Whincup, P.H.; Ng, J.; Paterson, C. Measuring retinal vessel tortuosity in 10-year-old children: Validation of the computer-assisted image analysis of the retina (CAIAR) program. Investig. Ophthalmol. Vis. Sci. 2009, 50, 2004–2010. [Google Scholar] [CrossRef]
- Chakravarty, A.; Sivaswamy, J. Glaucoma classification with a fusion of segmentation and image-based features. In Proceedings of the IEEE 13th International Symposium on Biomedical Imaging (ISBI), Prague, Czech Republic, 13–16 April 2016; pp. 689–692. [Google Scholar]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; De Vito, Z.; Lin, Z.; Desmaison, A.; Antiga, L.; Lerer, A.; et al. Automatic Differentiation in Pytorch. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Loshchilov, I.; Hutter, F. Decoupled weight decay regularization. arXiv 2017, arXiv:1711.05101. [Google Scholar]
- Zhuang, J. Laddernet: Multi-path networks based on u-net for medical image segmentation. arXiv 2018, arXiv:1810.07810. [Google Scholar]
- Soares, J.V.B.; Leandro, J.J.G.; Cesar, R.M.; Jelinek, H.F.; Cree, M.J. Retinal vessel segmentation using the 2-D Gabor wavelet and supervised classification. IEEE Trans. Med. Imaging 2006, 25, 1214–1222. [Google Scholar] [CrossRef]
- Jiang, Y.; Zhang, H.; Tan, N.; Chen, L. Automatic retinal blood vessel segmentation based on fully convolutional neural networks. Symmetry 2019, 11, 1112. [Google Scholar] [CrossRef]
- Dharmawan, D.A.; Li, D.; Ng, B.P.; Rahardja, S. A new hybrid algorithm for retinal vessels segmentation on fundus images. IEEE Access 2019, 7, 41885–41896. [Google Scholar] [CrossRef]
- Fu, H.; Cheng, J.; Xu, Y.; Wong, D.W.K.; Liu, J.; Cao, X. Joint optic disc and cup segmentation based on multi-label deep network and polar transformation. IEEE Trans. Med. Imaging 2018, 37, 1597–1605. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yang, S.; Yang, S.; Zhao, C.; Tian, G.; Gao, Y.; Chen, Y.; Lu, Y. Automatic thyroid nodule recognition and diagnosis in ultrasound imaging with the YOLOv2 neural network. World J. Surg. Oncol. 2019, 17, 12. [Google Scholar] [CrossRef] [PubMed]
- Chakravarty, A.; Sivaswamy, J. Joint optic disc and cup boundary extraction from monocular fundus images. Comput. Methods Programs Biomed. 2017, 147, 51–61. [Google Scholar] [CrossRef]
- Joshi, G.D.; Sivaswamy, J.; Krishnadas, S.R. Optic disk and cup segmentation from monocular color retinal images for glaucoma assessment. IEEE Trans. Med. Imaging 2011, 30, 1192–1205. [Google Scholar] [CrossRef]
- Joshi, G.D.; Sivaswamy, J.; Krishnadas, S.R. Depth discontinuity-based cup segmentation from multiview color retinal images. IEEE Trans. Biomed. Eng. 2012, 59, 1523–1531. [Google Scholar] [CrossRef]
- Cheng, J.; Liu, J.; Xu, Y.; Yin, F.; Wong, D.W.K.; Tan, N.-M.; Tao, D.; Cheng, C.-Y.; Aung, T.; Wong, T.Y. Superpixel classification based optic disc and optic cup segmentation for glaucoma screening. IEEE Trans. Med. Imaging 2013, 32, 1019–1032. [Google Scholar] [CrossRef]
- Zheng, Y.; Stambolian, D.; O’Brien, J.; Gee, C.J. Optic disc and cup segmentation from color fundus photograph using graph cut with priors. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Berlin/Heidelberg, Germany, 2013; pp. 75–82. [Google Scholar]





| Methods | DRIVE | STARE | CHASE | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | Acc | Se | Sp | F1 | Acc | Se | Sp | F1 | Acc | Se | Sp | |
| Basic | 0.8263/0.0189 | 0.9707/0.0039 | 0.7957/0.0519 | 0.9864/0.0032 | 0.8307/0.0211 | 0.9742/0.0044 | 0.8470/0.0527 | 0.9848/0.0045 | 0.8081/ 0.0181 | 0.9754/0.0038 | 0.8220/0.0331 | 0.9857/0.0025 |
| SMCF | 0.8280/0.0182 | 0.9705/ 0.0032 | 0.8093/0.0518 | 0.9860/0.0034 | 0.8317/0.0179 | 0.9743/0.0040 | 0.8476/0.0452 | 0.9848/0.0034 | 0.8140/ 0.0190 | 0.9762/0.0036 | 0.8256/0.0391 | 0.9863/0.0022 |
| SMCF+SRL | 0.8292/0.0156 | 0.9702/0.0026 | 0.8240/0.0477 | 0.9843/0.0039 | 0.8336/0.0135 | 0.9747/0.0035 | 0.8512/0.0311 | 0.9849/0.0029 | 0.8150/0.0136 | 0.9763/ 0.0031 | 0.8273/0.0344 | 0.9863/0.0023 |
| SMCF+MBCM+SRL | 0.8301/0.0143 | 0.9704/0.0028 | 0.8246/0.0487 | 0.9844/0.0038 | 0.8341/0.0119 | 0.9747/0.0032 | 0.8534/0.0299 | 0.9847/0.0032 | 0.8161/0.0168 | 0.9757/0.0045 | 0.9844/0.0020 | |
| MSMB-Net (ours) | 0.8371/0.0280 | |||||||||||
| Methods | Disc | Cup | Optic | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | Acc | Se | BLE | F1 | Acc | Se | BLE | F1 | Acc | Se | Sp | |
| Basic | 0.9604/0.101 | 0.9935/0.003 | 0.8056/0.125 | 7.327/6.191 | 0.8834/0.112 | 0.9935/0.001 | 0.9417/0.084 | 17.528/11.964 | 0.9596/0.031 | 0.9968/0.002 | 0.9766/0.031 | 0.9974/0.002 |
| SMCF | 0.8280/0.072 | 0.9949/0.002 | 0.8180/0.110 | 6.196/5.388 | 0.8959/0.096 | 0.9970/0.001 | 0.9498/0.063 | 16.289/10.575 | 0.9687/0.017 | 0.9979/0.001 | 0.9775/0.027 | 0.9985/0.001 |
| SMCF+SRL | 0.9741/0.069 | 0.9952/0.002 | 0.8114/0.110 | 5.410/4.871 | 0.8999/0.109 | 0.9968/0.001 | 15.086/11.286 | 0.9736/0.015 | 0.9983/0.0009 | 0.9831/0.021 | 0.9987/0.001 | |
| SMCF+MBCM+SRL | 0.9750/0.055 | 0.9953/0.002 | 0.8355/0.081 | 4.459/2.203 | 0.8995/0.106 | 0.9969/0.001 | 0.9558/0.063 | 13.354/10.111 | 0.9735/0.011 | 0.9983/0.0007 | 0.9833/0.016 | 0.9988/0.0009 |
| MSMB-Net (ours) | 0.9560/0.050 | |||||||||||
| Type | Methods | Year | Se | Sp | Acc | F1 |
|---|---|---|---|---|---|---|
| Unsupervised methods | 2nd human expert | 0.7743 | 0.9819 | 0.9637 | 0.7889 | |
| Miao et al. [4] | 2015 | 0.7481 | 0.9748 | 0.9597 | - | |
| Kumar et al. [8] | 2019 | 0.7503 | 0.9717 | 0.9432 | - | |
| Tian et al. [9] | 2019 | 0.8639 | 0.9690 | 0.9580 | - | |
| Jainish et al. [10] | 2020 | - | - | 0.9657 | - | |
| Supervised methods | Marín et al. [11] | 2010 | 0.7607 | 0.9801 | 0.9452 | - |
| Aslani et al. [12] | 2016 | 0.7545 | 0.9801 | 0.9513 | - | |
| Feng et al. [13] | 2017 | 0.7811 | 0.9839 | 0.9560 | - | |
| U-Net [17] | 2018 | 0.7537 | 0.9820 | 0.9531 | 0.8142 | |
| R2U-Net [18] | 2018 | 0.7792 | 0.9813 | 0.9556 | 0.8171 | |
| IterNet [19] | 2019 | 0.7735 | 0.9838 | 0.9573 | 0.8205 | |
| Ce-net [29] | 2019 | 0.8309 | - | 0.9545 | - | |
| Sine-Net [21] | 2020 | 0.8260 | 0.9824 | 0.9685 | - | |
| HAnet [22] | 2020 | 0.7991 | 0.9813 | 0.9581 | 0.8293 | |
| MSMB-Net (ours) | 2020 | 0.8283 | 0.9864 | 0.9708 | 0.8315 |
| Type | Methods | Year | Se | Sp | Acc | F1 |
|---|---|---|---|---|---|---|
| Unsupervised methods | 2nd human expert | 0.9017 | 0.9564 | 0.9522 | 0.7417 | |
| Miao et al. [4] | 2015 | 0.7298 | 0.9831 | 0.9532 | - | |
| Azzopardi et al. [7] | 2015 | 0.7716 | 0.9701 | 0.9497 | - | |
| Jainish et al. [10] | 2020 | - | - | 0.9703 | - | |
| Supervised methods | Marín et al. [11] | 2010 | 0.6944 | 0.9819 | 0.9526 | - |
| Aslani et al. [12] | 2016 | 0.7556 | 0.9837 | 0.9605 | - | |
| Mo et al. [14] | 2017 | 0.8147 | 0.9844 | 0.9674 | - | |
| Hu et al. [15] | 2018 | 0.7543 | 0.9814 | 0.9632 | - | |
| U-Net [17] | 2018 | 0.8270 | 0.9842 | 0.9690 | 0.8373 | |
| IterNet [19] | 2019 | 0.7715 | 0.9886 | 0.9701 | 0.8146 | |
| DUNet [20] | 2019 | 0.8369 | 0.9888 | 0.9773 | 0.8485 | |
| Sine-Net [21] | 2020 | 0.6776 | 0.9946 | 0.9711 | - | |
| HAnet [22] | 2020 | 0.8186 | 0.9844 | 0.9673 | 0.8379 | |
| MSMB-Net (ours) | 2020 | 0.8760 | 0.9899 | 0.9753 | 0.8469 |
| Type | Methods | Year | Se | Sp | Acc | F1 |
|---|---|---|---|---|---|---|
| Unsupervised methods | 2nd human expert | 0.6776 | 0.9946 | 0.9711 | - | |
| Azzopardi et al. [7] | 2015 | 0.7585 | 0.9587 | 0.9387 | - | |
| Tian et al. [9] | 2019 | 0.8778 | 0.9680 | 0.9601 | - | |
| Supervised methods | Mo et al. [14] | 2017 | 0.7661 | 0.9816 | 0.9599 | - |
| Yan et al. [16] | 2018 | 0.7641 | 0.9806 | 0.9607 | - | |
| U-Net [17] | 2018 | 0.8288 | 0.9701 | 0.9578 | 0.7783 | |
| R2U-Net [18] | 2018 | 0.7756 | 0.9820 | 0.9634 | 0.7928 | |
| IterNet [19] | 2019 | 0.7970 | 0.9823 | 0.9655 | 0.8073 | |
| DUNet [20] | 2019 | 0.8155 | 0.9752 | 0.9610 | 0.7883 | |
| Sine-Net [21] | 2020 | 0.7856 | 0.9845 | 0.9676 | - | |
| MSMB-Net (ours) | 2020 | 0.8331 | 0.9864 | 0.9767 | 0.8190 |
| Methods | Year | Se | Sp | Acc | Dice | BLE |
|---|---|---|---|---|---|---|
| Vessel Bend [51] | 2011 | - | - | - | 0.9600/0.02 | 8.93/2.96 |
| Multiview [52] | 2012 | - | - | - | 0.9600/0.02 | 8.93/2.96 |
| Superpixel [53] | 2013 | - | - | - | 0.9500/0.02 | 9.38/5.75 |
| Graph Cut [54] | 2013 | - | - | - | 0.9400/0.06 | 14.74/15.66 |
| U-Net [17] | 2015 | 0.9600 | 0.9800 | 0.9700 | 0.9500 | - |
| Zilly et al. [23] | 2015 | - | - | - | 0.9470 | - |
| BCRF [50] | 2017 | - | - | - | 0.9700/0.02 | 6.61/3.55 |
| Stack-u-net [24] | 2018 | - | - | - | 0.9700/0.02 | 6.47/3.51 |
| RACE-net [25] | 2018 | - | - | - | 0.9700/0.02 | 6.06/3.84 |
| Shah et al. [26] | 2019 | - | - | - | 0.9600 | - |
| Yu et al. [27] | 2019 | - | - | - | 0.9738 | - |
| Ding et al. [28] | 2019 | - | - | - | 0.9721 | - |
| Ce-net [29] | 2019 | 0.9759 | 0.9990 | - | 0.9642 | - |
| WGAN [30] | 2020 | - | - | - | 0.9540 | - |
| CDED-Net [31] | 2020 | 0.9754 | 0.9973 | - | 0.9597 | - |
| MSMB-Net (ours) | 2020 | 0.9610 | 0.9984 | 0.9959 | 0.9782 | 3.98/1.82 |
| Methods | Year | Se | Sp | Acc | Dice | BLE |
|---|---|---|---|---|---|---|
| Vessel Bend [51] | 2011 | - | - | - | 0.7700/0.20 | 30.51/24.80 |
| Multiview [52] | 2012 | - | - | - | 0.7900/0.18 | 25.28/18.00 |
| Superpixel [53] | 2013 | - | - | - | 0.8000/0.14 | 22.04/12.57 |
| Graph Cut [54] | 2013 | - | - | - | 0.7700/0.16 | 26.70/16.67 |
| U-Net [17] | 2015 | 0.9600 | 0.9800 | 0.9700 | 0.8500/0.10 | 19.53/13.98 |
| Zilly et al. [23] | 2015 | - | - | - | 0.8300 | - |
| BCRF [50] | 2017 | - | - | - | 0.8300/0.15 | 18.61/13.02 |
| Stack-u-net [24] | 2018 | - | - | - | 0.8900/0.09 | 14.39/7.18 |
| RACE-net [25] | 2018 | - | - | - | 0.8700/0.09 | 16.13/7.63 |
| Shah et al. [26] | 2019 | - | - | - | 0.8900 | - |
| Yu et al. [27] | 2019 | - | - | - | 0.8877 | - |
| Ding et al. [28] | 2019 | - | - | - | 0.8513 | - |
| Ce-net [29] | 2019 | 0.8819 | 0.9909 | - | 0.8818 | - |
| WGAN [30] | 2020 | - | - | - | 0.8400 | - |
| CDED-Net [31] | 2020 | 0.9567 | 0.9981 | - | 0.9240 | - |
| MSMB-Net (ours) | 2020 | 0.9560 | 0.9983 | 0.9975 | 0.9184 | 13.01/9.24 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, Y.; Liu, W.; Wu, C.; Yao, H. Multi-Scale and Multi-Branch Convolutional Neural Network for Retinal Image Segmentation. Symmetry 2021, 13, 365. https://doi.org/10.3390/sym13030365
Jiang Y, Liu W, Wu C, Yao H. Multi-Scale and Multi-Branch Convolutional Neural Network for Retinal Image Segmentation. Symmetry. 2021; 13(3):365. https://doi.org/10.3390/sym13030365
Chicago/Turabian StyleJiang, Yun, Wenhuan Liu, Chao Wu, and Huixiao Yao. 2021. "Multi-Scale and Multi-Branch Convolutional Neural Network for Retinal Image Segmentation" Symmetry 13, no. 3: 365. https://doi.org/10.3390/sym13030365
APA StyleJiang, Y., Liu, W., Wu, C., & Yao, H. (2021). Multi-Scale and Multi-Branch Convolutional Neural Network for Retinal Image Segmentation. Symmetry, 13(3), 365. https://doi.org/10.3390/sym13030365








