Mathematical Modeling for Prediction Dynamics of the Coronavirus Disease 2019 (COVID-19) Pandemic, Quarantine Control Measures
Abstract
1. Introduction
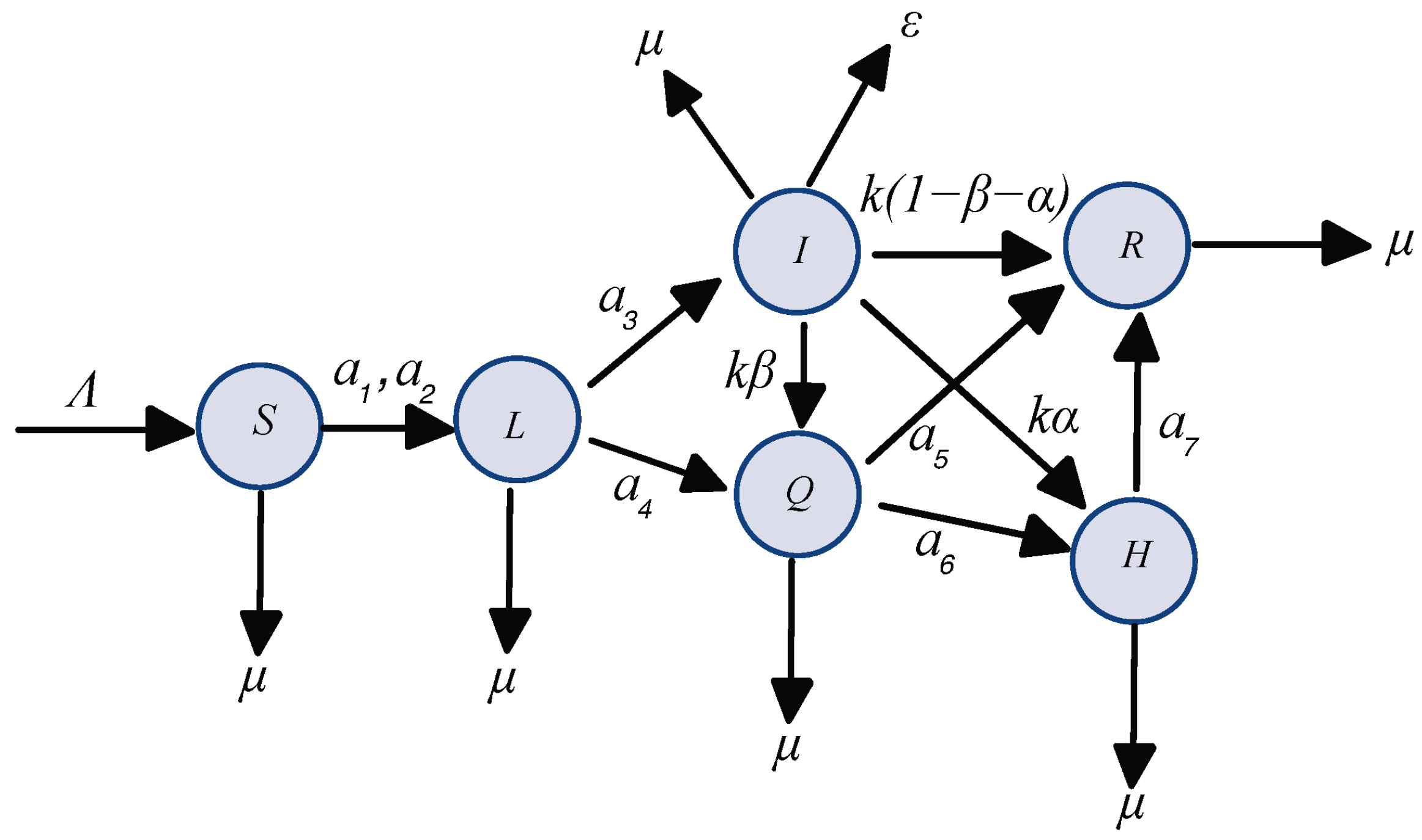
2. Mathematical Model
3. Analysis of the Model
3.1. Invariant Region
3.2. Existence of the Solution
3.3. Positivity of the Solution
3.4. Equilibria
- (i)
- The disease-free equilibrium (DFE)
- (ii)
- The endemic equilibriumwherewith and .
3.5. The Basic Reproduction Number ()
3.6. Stability of Disease-Free Equilibrium (DFE)
3.7. Stability of the Endemic Equilibrium
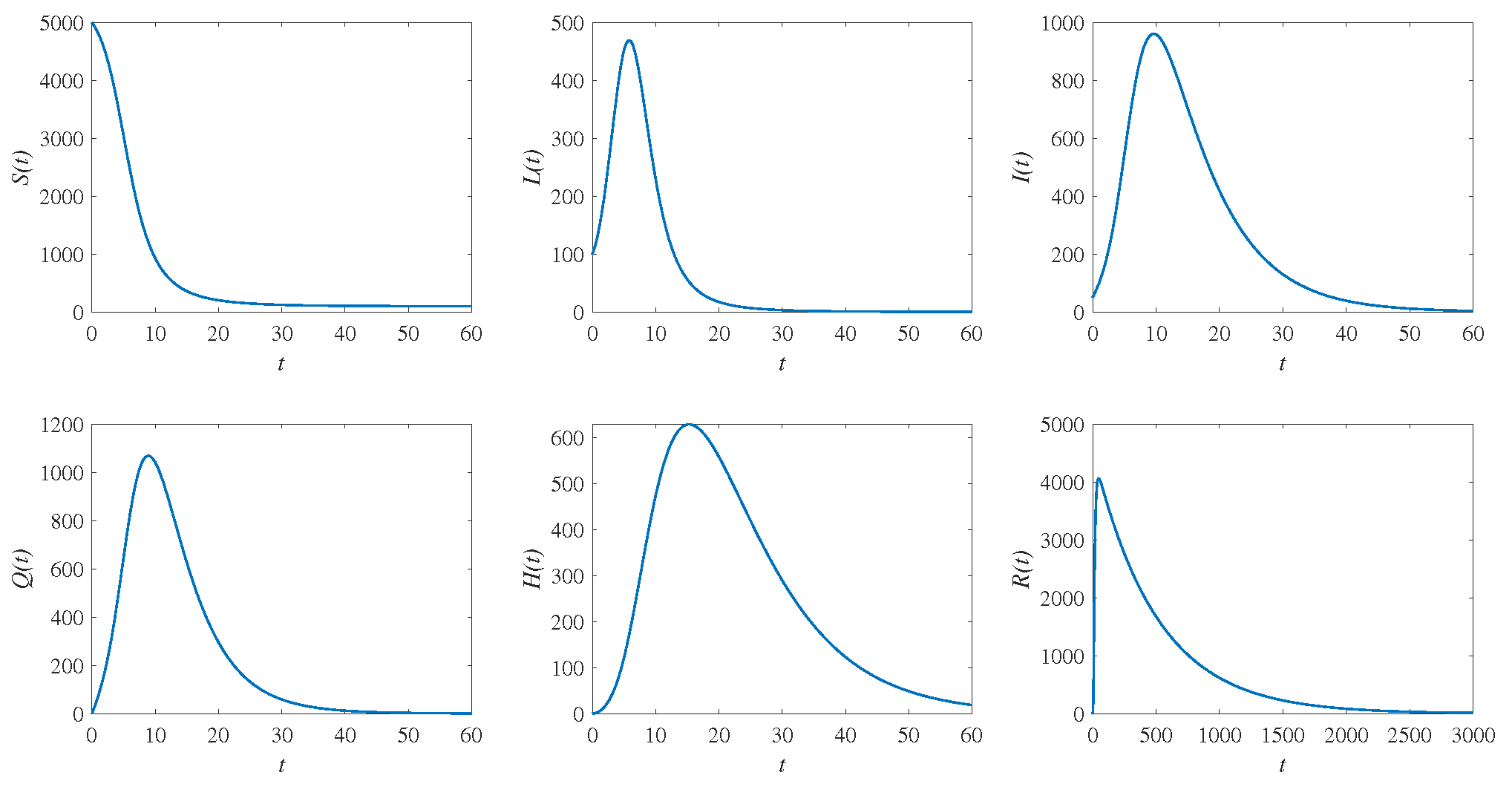
4. Numerical Simulations
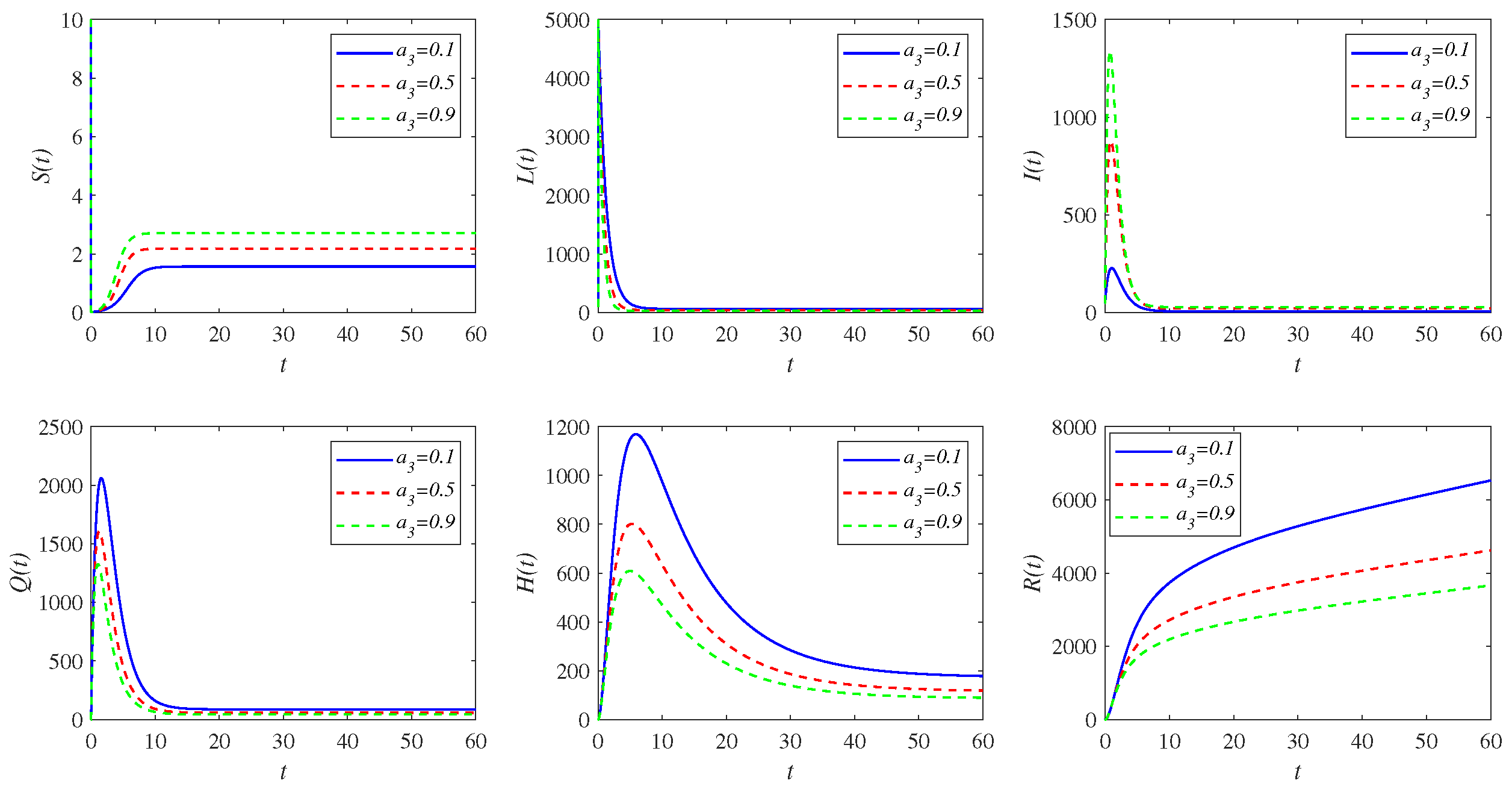
5. Sensitivity Analysis
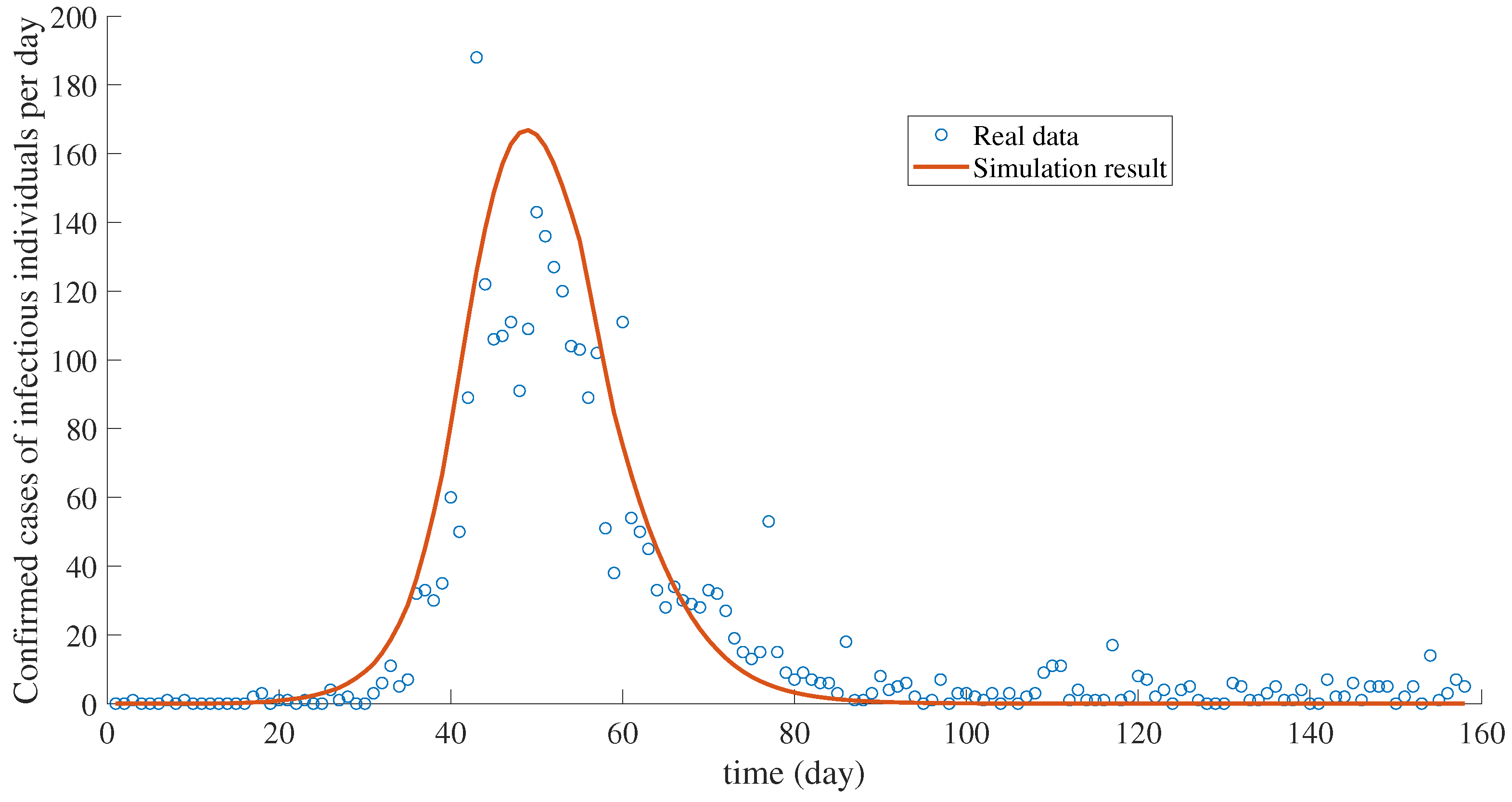
6. The Case Study of Thailand
7. Conclusions and Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Coronavirus Disease 2019 (Covid-19): Situation Report. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-remarks-at-the-media-briefing-on-2019-ncov-on-11-february-2020 (accessed on 1 July 2020).
- Fanelli, D.; Piazza, F. Analysis and forecast of covid-19 spreading in china, italy and france. Chaos Solitons Fractals 2020, 134, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Randhawa, G.; Soltysiak, M.; Roz, H.; de Souza, C.; Hill, K.; Kari, L. Machine learning using intrinsic genomic signatures for rapid classification of novel pathogens: Covid-19 case study. PLoS ONE 2020, 15, e0232391. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Chen, H.D. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Xiao, K.; Zhai, J.; Feng, Y.; Zhou, N.; Zhang, X.; Zou, J.J.; Zhang, Z. Isolation of sars-cov-2-related coronavirus from malayan pangolins. Nature 2020, 583, 286–289. [Google Scholar] [CrossRef]
- Zhang, T.; Wu, Q.; Zhang, Z. Probable pangolin origin of sars-cov-2 associated with the Covid-19 outbreak. Curr. Biol. 2020, 134, 1–5. [Google Scholar] [CrossRef]
- World Health Organization. Consensus Document on the Epidemiology of Severe Acute Respiratory Syndrome (Sars). Available online: https://www.who.int/csr/sars/WHOconsensus.pdf (accessed on 12 July 2020).
- World Health Organization. Middle East Respiratory Syndrome Coronavirus (Mers-Cov). Available online: https://www.who.int/news-room/fact-sheets/detail/middle-east-respiratory-syndrome-coronavirus-(mers-cov) (accessed on 12 July 2020).
- Atangana, A. Modelling the spread of Covid-19 with new fractal-fractional operators: Can the lockdown save mankind before vaccination? Chaos Solitons Fractals 2020, 136, 109860. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus Disease 2019 (Covid-19): Situation Report. 133. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200712-covid-19-sitrep-133.pdf (accessed on 12 July 2020).
- World Health Organization. Coronavirus Disease 2019 (Covid-19): Situation Report. 174. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200712-covid-19-sitrep-174.pdf (accessed on 12 July 2020).
- Suparit, P.; Wiratsudakul, A.; Modchang, C. A mathematical model for zika virus transmission dynamics with a time-dependent mosquito biting rate. Theor. Med. Model. 2018, 15, 1–11. [Google Scholar] [CrossRef]
- Bocharov, G.; Volpert, V.; Ludewig, B.; Meyerhans, A. Mathematical Immunology of Virus Infections; Springer International Publishing AG: Cham, Switzerland, 2018. [Google Scholar]
- Sardar, T.; Chattopadhyay, J. A mathematical model of dengue transmission with memory. Commun. Nonlinear Sci. Numer. Simul. 2015, 22, 511–525. [Google Scholar] [CrossRef]
- Wu, C.; Wong, P.Y. Dengue transmission: Mathematical model with discrete time delays and estimation of the reproduction number. J. Biol. Dyn. 2019, 13, 1–25. [Google Scholar] [CrossRef]
- Sazonov, I.; Grebennikov, D.; Kelbert, M.; Meyerhans, A.; Bocharov, G. Viral Infection Dynamics Model Based on a Markov Process with Time Delay between Cell Infection and Progeny Production. Mathematics 2020, 8, 1207. [Google Scholar] [CrossRef]
- Zheltkova, V.; Argilaguet, J.; Peligero, C.; Bocharov, G.; Meyerhans, A. Prediction of PD-L1 inhibition effects for HIV-infected individuals. PLoS Comput. Biol. 2019, 15, E1007401. [Google Scholar] [CrossRef]
- Shcherbatova, O.; Grebennikov, D.; Sazonov, I.; Meyerhans, A.; Bocharov, G. Modeling of the HIV-1 Life Cycle in Productively Infected Cells to Predict Novel Therapeutic Targets. Pathogens 2020, 9, 255. [Google Scholar] [CrossRef] [PubMed]
- Romanyukha, A.A.; Nosova, E.A. Modeling Spread of HIV as a Result of Social Maladjustment. Autom. Remote Control 2012, 73, 2071–2082. [Google Scholar] [CrossRef]
- Avilov, K.K.; Romanyukha, A.A.; Borisov, S.E.; Belilovsky, E.M.; Nechaeva, O.B.; Karkach, A.S. An approach to estimating tuberculosis incidence and case detection rate from routine notification data. Int. Tuberc. Lung Dis. 2015, 19, 288–294. [Google Scholar] [CrossRef]
- Jiang, S.; Wang, K.; Li, C.; Hong, G.; Zhang, X.; Shan, M.; Li, H.; Wang, J. Mathematical models for devising the optimal ebola virus disease eradication. J. Transl. Med. 2017, 1–10. [Google Scholar] [CrossRef]
- Rhoubari, Z.; Besbassi, H.; Hattaf, K.; Yousfi, N. Mathematical modeling of ebola virus disease in bat population. J. Transl. Med. 2018, 1–7. [Google Scholar] [CrossRef]
- Berge, T.; Lubuma, J.-S.; Moremedi, G.; Morris, N.; Kondera-Shava, R. A simple mathematical model for ebola in africa. J. Biol. Dyn. 2017, 11, 42–74. [Google Scholar] [CrossRef]
- Deepa, O.S.; Nallamalli, S.; SinghNaik, L.; Teja, G. Mathematical model for transmission of ebola. Procedia Comput. Sci. 2015, 48, 741–745. [Google Scholar] [CrossRef][Green Version]
- Carvalho, S.; da Silva, S.; da Cunha Charret, I. Mathematical modeling of dengue epidemic: Control methods and vaccination strategies. Theory Biosci. 2019, 138, 223–239. [Google Scholar] [CrossRef]
- Bonyah, E.; Okosun, K. Mathematical modeling of zika virus. Asian Pac. J. Trop. Dis. 2016, 6, 673–679. [Google Scholar] [CrossRef]
- Biswas, S.; Ghosh, U.; Sarkarb, S. Mathematical model of zika virus dynamics with vector control and sensitivity analysis. Infect. Dis. Model. 2020, 5, 23–41. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Ma, W.; Bai, P. A novel dynamic model describing the spread of the mers-cov and the expression of dipeptidyl peptidase 4. Comput. Math. Methods Med. 2017, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Liang, K. Mathematical model of infection kinetics and its analysis for Covid-19, sars and mers. Infect. Genet. Evol. 2020, 82, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Lamwong, J.; Tang, I.; Pongsumpun, P. Mers model of thai and south korean population. Curr. Appl. Sci. Technol. 2018, 18, 1–13. [Google Scholar]
- Herrera, G.; Fernandez-Merodo, J.; Mulas, J.; Pastor, M. A landslide forecasting model using ground based sar data: The portalet case study. Eng. Geol. 2013, 105, 220–230. [Google Scholar] [CrossRef]
- Goulard, M.; Laurent, T.; Thomas-Agnan, C. About predictions in spatial sar models: Optimal and almost optimal strategies. Spat. Econ. Anal. 2017, 12, 304–325. [Google Scholar] [CrossRef]
- Ndaïrou, F.; Torresa, D.M. Mathematical modeling of covid-19 transmission dynamics with a case study of wuhan. Chaos Solitons Fractals 2020, 135, 1–6. [Google Scholar] [CrossRef]
- Postnikov, E. Estimation of Covid-19 dynamics “on a back-of-envelope”: Does the simplest sir model provide quantitative parameters and predictions? Chaos Solitons Fractals 2020, 135, 109841. [Google Scholar] [CrossRef]
- Sarkar, K.; Khajanchi, S.; Nieto, J. Modeling and forecasting the covid-19 pandemic in india. Chaos Solitons Fractals 2020, 139, 110049. [Google Scholar] [CrossRef]
- Shereen, M.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. Covid-19 infection: Origin, transmission, and characteristics of human coronaviruses. J. Adv. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef]
- Arino, J.; Portet, S. A simple model for Covid-19. Infect. Dis. Model. 2020, 5, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Van Bavel, J.J.; Baicker, K.; Boggio, P.S.; Capraro, V.; Cichocka, A.; Cikara, M.; Drury, J. Using social and behavioural science to support Covid-19 pandemic response. Nat. Hum. Behav. 2020, 4, 460–471. [Google Scholar] [CrossRef] [PubMed]
- Derrick, N.; Grossman, S. Differential Equation with Application; Addision Wesley Publishing Company, Inc.: Reading, MA, USA, 1976. [Google Scholar]
- van den Driessche, P.; Watmough, J. Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math. Biosci. 2002, 180, 29–48. [Google Scholar] [CrossRef]
- Bhaskar, A.; Song, Y. Descartes’ rule of signs and the identifiability of population demographic models from genomic variation data. Ann. Stat. 2014, 42, 2469–2493. [Google Scholar] [CrossRef] [PubMed]
- Chitnisa, N.; Hymanb, J.; Cushing, J. Determining important parameters in the spread of malaria through the sensitivity analysis of a mathematical model. Bull. Math. Biol. 2008, 70, 1272–1296. [Google Scholar] [CrossRef]
- Ministry of Public Health, Department of Disease Control. COVID-19 Situation Reports. Available online: https://covid19.ddc.moph.go.th/en (accessed on 1 July 2020).



| Parameters Symbol | Sensitivity Indices |
|---|---|
| 1 | |
| 0.8236331570 | |
| 0.1763668430 | |
| −0.07342499706 | |
| −0.7493755204 | |
| k | −0.06294319877 |
| −1.000958526 | |
| −0.1798043729 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prathumwan, D.; Trachoo, K.; Chaiya, I. Mathematical Modeling for Prediction Dynamics of the Coronavirus Disease 2019 (COVID-19) Pandemic, Quarantine Control Measures. Symmetry 2020, 12, 1404. https://doi.org/10.3390/sym12091404
Prathumwan D, Trachoo K, Chaiya I. Mathematical Modeling for Prediction Dynamics of the Coronavirus Disease 2019 (COVID-19) Pandemic, Quarantine Control Measures. Symmetry. 2020; 12(9):1404. https://doi.org/10.3390/sym12091404
Chicago/Turabian StylePrathumwan, Din, Kamonchat Trachoo, and Inthira Chaiya. 2020. "Mathematical Modeling for Prediction Dynamics of the Coronavirus Disease 2019 (COVID-19) Pandemic, Quarantine Control Measures" Symmetry 12, no. 9: 1404. https://doi.org/10.3390/sym12091404
APA StylePrathumwan, D., Trachoo, K., & Chaiya, I. (2020). Mathematical Modeling for Prediction Dynamics of the Coronavirus Disease 2019 (COVID-19) Pandemic, Quarantine Control Measures. Symmetry, 12(9), 1404. https://doi.org/10.3390/sym12091404





