Morphometric Analysis of Surface Utricles in Halimeda tuna (Bryopsidales, Ulvophyceae) Reveals Variation in Their Size and Symmetry within Individual Segments
Abstract
1. Introduction
2. Materials and Methods
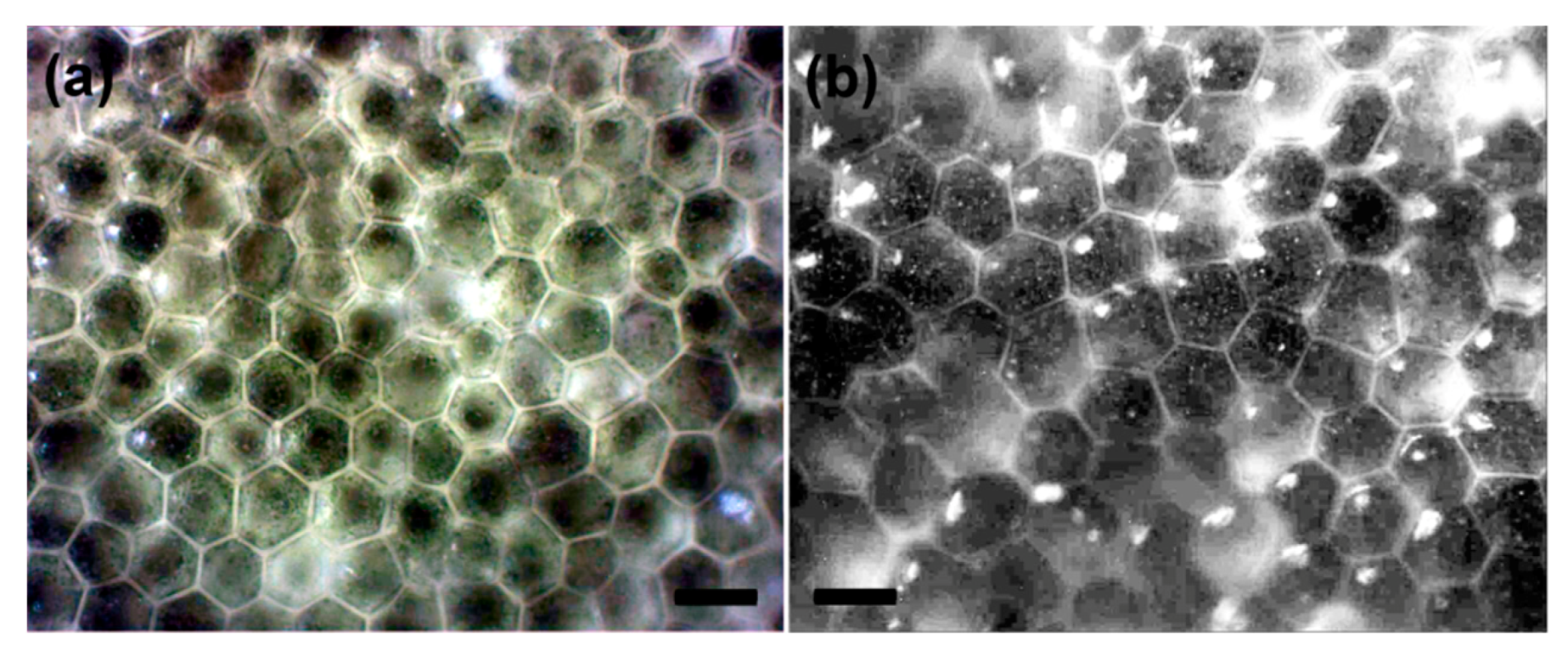
2.1. Sampling and Microscopy of Segments
2.2. Morphometric Analysis of Surface Utricles
2.3. Statistical Analysis of Utricle Parameters
2.4. Morphometric Analysis of Segments
3. Results
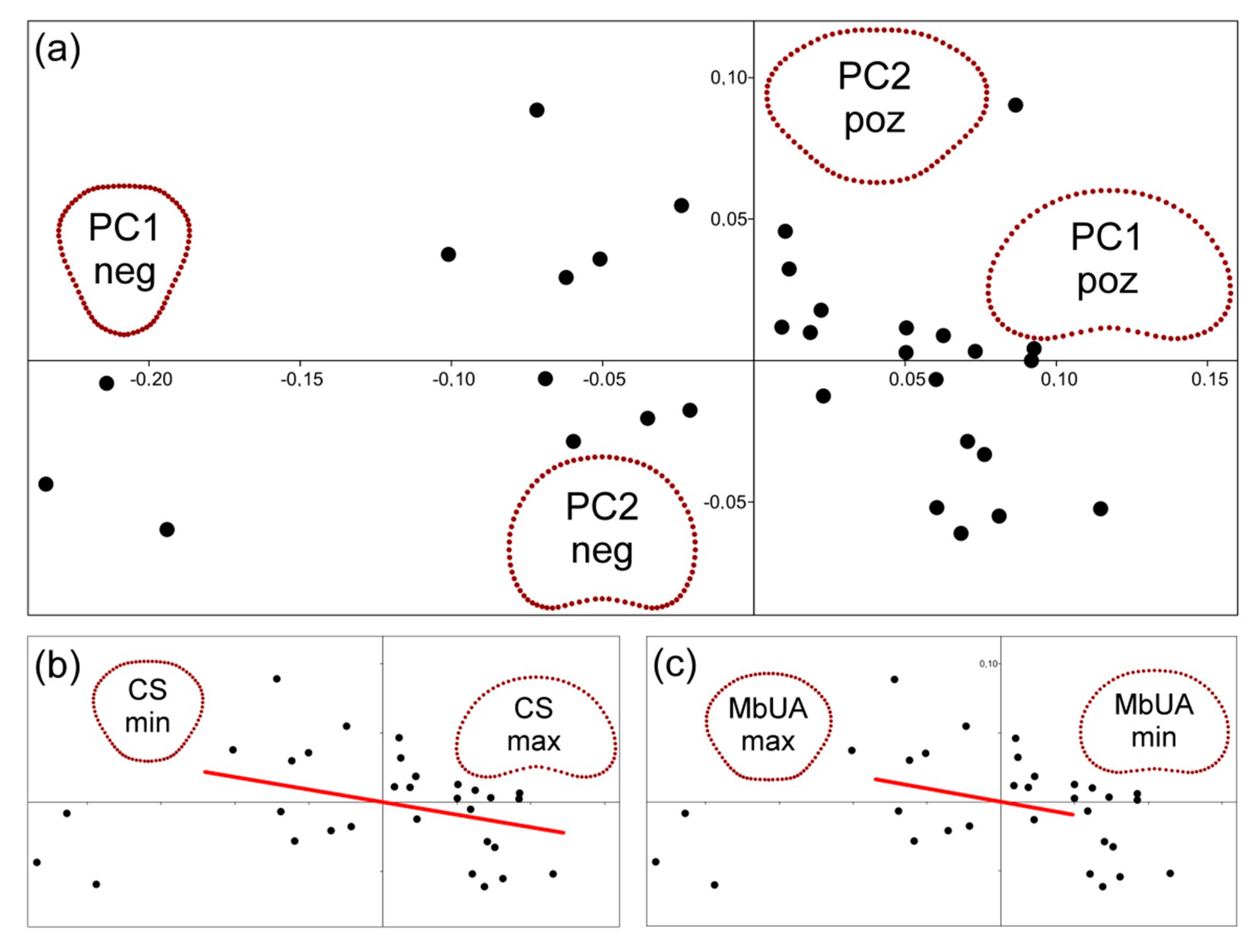
3.1. Correlation among Utricle Parameters
3.2. Utricle Size
3.3. Utricle Symmetry
3.4. Number of Polygon Vertices of Utricle Shapes
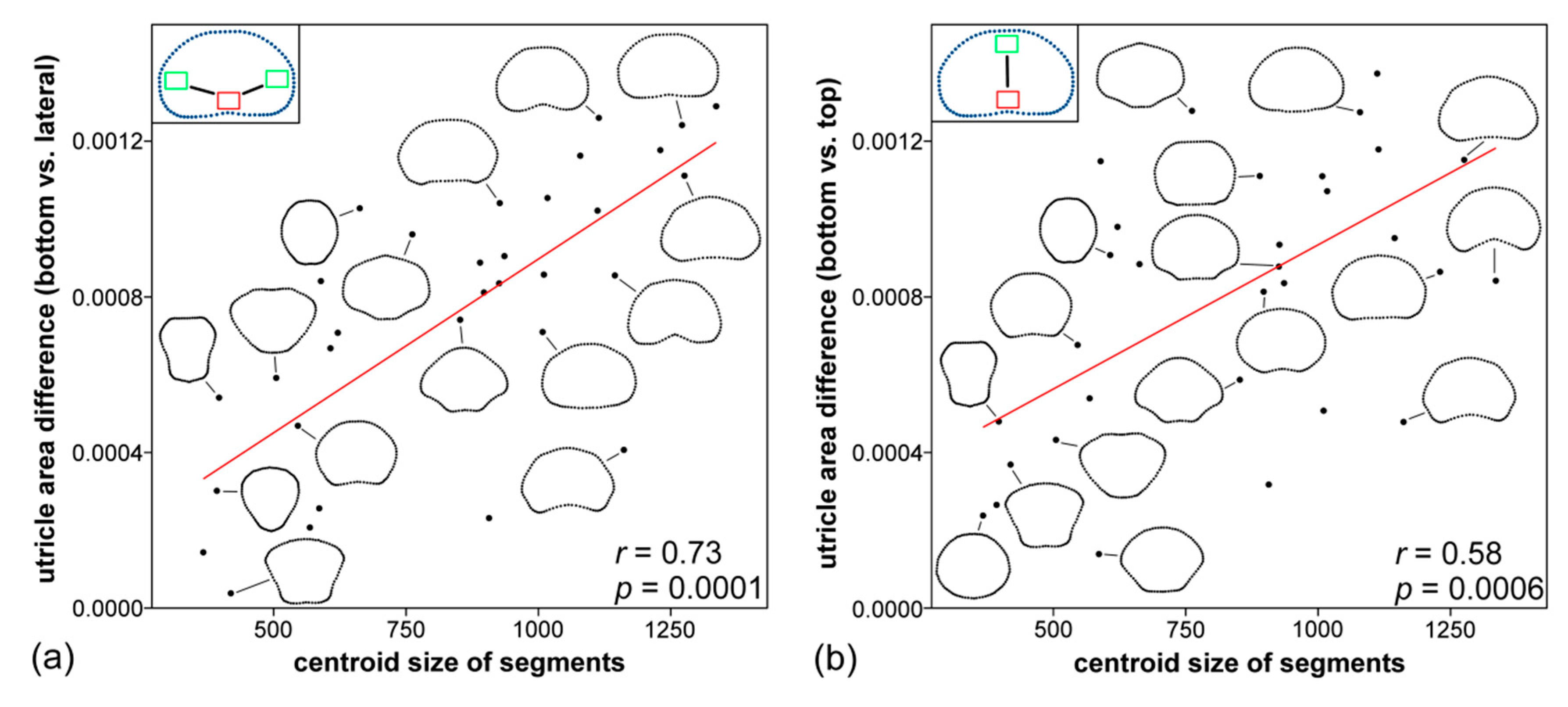
3.5. Morphology of Segments and Their Relation to Utricle Size and Symmetry
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hillis-Colinvaux, L. Ecology and Taxonomy of Halimeda: Primary Producer of Coral Reefs; Academic Press: London, UK, 1980; p. 327. [Google Scholar]
- Verbruggen, K.; Kooistra, W.H.C.F. Morphological characterization of lineages within the calcified tropical seaweed genus Halimeda (Bryopsidales, Chlorophyta). Eur. J. Phycol. 2004, 39, 213–228. [Google Scholar] [CrossRef]
- Borowitzka, M.A.; Larkum, A.W.D. Calcification in the green alga Halimeda. III. The sources of inorganic carbon for photosynthesis and calcification and a model of the mechanism of calcification. J. Exp. Bot. 1976, 27, 879–893. [Google Scholar] [CrossRef]
- Peach, K.E.; Koch, M.S.; Blackwelder, P.L.; Guerrero-Given, D.; Kamasawa, N. Primary utricle structure of six Halimeda species and potential relevance for ocean acidification tolerance. Bot. Mar. 2017, 60, 1–11. [Google Scholar] [CrossRef]
- Phillips, D. Tessellation. WIRES Comput. Stat. 2014, 6, 202–209. [Google Scholar] [CrossRef]
- Caër, G.L.; Ho, J.S. The Voronoi tessellation generated from eigenvalues of complex random matrices. J. Phys. A Math. Gen. 1990, 23, 3279–3295. [Google Scholar] [CrossRef]
- Verbruggen, H.; De Clerck, O.; Coppejans, E. Deviant segments hamper a morphometric approach towards Halimeda taxonomy. Crypt. Algol. 2005, 26, 259–274. [Google Scholar]
- Vroom, P.S.; Smith, C.M.; Coyer, J.A.; Walters, L.J.; Hunter, C.L.; Beach, K.S.; Smith, J.E. Field biology of Halimeda tuna (Bryopsidales, Chlorophyta) across a depth gradient: Comparative growth, survivorship, recruitment, and reproduction. Hydrobiologia 2003, 501, 149–166. [Google Scholar] [CrossRef]
- Pongparadon, S.; Zuccarello, G.C.; Prathep, A. High morpho-anatomical variability in Halimeda macroloba (Bryopsidales, Chlorophyta) in Thai waters. Phycol. Res. 2017, 65, 136–145. [Google Scholar] [CrossRef]
- Pongparadon, S.; Nooek, S.; Prathep, A. Phenotypic plasticity and morphological adaptation of Halimeda opuntia (Bryopsidales, Chlorophyta) to light intensity. Phycol. Res. 2020, 68, 115–125. [Google Scholar] [CrossRef]
- Neustupa, J.; Nemcova, Y. Morphological allometry constrains symmetric shape variation, but not asymmetry, of Halimeda tuna (Bryopsidales, Ulvophyceae) segments. PLoS ONE 2018, 13, e0206492. [Google Scholar] [CrossRef]
- Noble, J.M. Halimeda magnidisca (Caulerpales, Chlorophyta), a new species from the Great Barrier Reef, Australia. Phycologia 1986, 25, 331–339. [Google Scholar] [CrossRef]
- Rohlf, F.J. The tps series of software. Hystrix Ital. J. Mammal. 2015, 26, 9–12. [Google Scholar]
- Barber, C.B.; Dobkin, D.P.; Huhdanpaa, H.T. The Quickhull algorithm for convex hulls. ACM Trans. Math. Softw. 1996, 22, 469–483. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. 2018. Available online: https://www.r-project.org/ (accessed on 8 June 2020).
- Klingenberg, C.P. Size, shape, and form: Concepts of allometry in geometric morphometrics. Dev. Genes Evol. 2016, 226, 113–137. [Google Scholar] [CrossRef] [PubMed]
- Zabrodsky, H.; Peleg, S.; Avnir, D. Continuous symmetry measures. J. Am. Chem. Soc. 1992, 114, 7843–7851. [Google Scholar] [CrossRef]
- Zabrodsky, H.; Peleg, S.; Avnir, D. Symmetry as a continuous feature. IEEE Trans. Pattern Anal. Mach. Intell. 1995, 17, 1154–1166. [Google Scholar] [CrossRef]
- Zabrodsky, H.; Avnir, D. Continuous symmetry measures. 4. Chirality. J. Am. Chem. Soc. 1995, 117, 462–473. [Google Scholar] [CrossRef]
- Zahrt, A.F.; Denmark, S.E. Evaluating continuous chirality measure as a 3D descriptor in chemoinformatics applied to asymmetric catalysis. Tetrahedron 2019, 75, 1841–1851. [Google Scholar] [CrossRef]
- Graham, J.H.; Whitesell, M.J.; Fleming, M., II.; Hel-Or, H.; Nevo, E.; Raz, S. Fluctuating asymmetry of plant leaves: Batch processing with LAMINA and continuous symmetry measures. Symmetry 2015, 7, 255–268. [Google Scholar] [CrossRef]
- Iovita, R.; Tuvi-Arad, I.; Moncel, M.H.; Despriée, J.; Voinchet, P.; Bahain, J.J. High handaxe symmetry at the beginning of the European Acheulian: The data from la Noira (France) in context. PLoS ONE 2017, 12, e0177063. [Google Scholar] [CrossRef]
- Dryden, I.L. Shapes: Statistical Shape Analysis. R Package Version 1.2.5. 2019. Available online: https://CRAN.R-project.org/package=shapes (accessed on 8 June 2020).
- Klingenberg, C.P. Analyzing fluctuating asymmetry with geometric morphometrics: Concepts, methods, and applications. Symmetry 2015, 7, 843–934. [Google Scholar] [CrossRef]
- Li, K.; Gao, X.L.; Wang, J. Dynamic crushing behavior of honeycomb structures with irregular cell shapes and non-uniform cell wall thickness. Int. J. Sol. Struct. 2007, 44, 5003–5026. [Google Scholar] [CrossRef]
- Anderson, M.J.; Ter Braak, C.J.F. Permutation tests for multi-factorial analysis of variance. J. Stat. Comp. Sim. 2003, 73, 85–113. [Google Scholar] [CrossRef]
- Collyer, M.L.; Sekora, D.J.; Adams, D.C. A method for analysis of phenotypic change for phenotypes described by high-dimensional data. Heredity 2015, 115, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.C.; Otárola-Castillo, E. Geomorph: An R package for the collection and analysis of geometric morphometric shape data. Meth. Ecol. Evol. 2013, 4, 393–399. [Google Scholar] [CrossRef]
- Good, P.I. Permutation, Parametric, and Bootstrap Tests of Hypotheses, 3rd ed.; Springer: New York, NY, USA, 2005; p. 316. [Google Scholar]
- Bookstein, F.L. Landmark methods for forms without landmarks: Morphometrics of group differences in outline shape. Med. Image Anal. 1997, 1, 225–243. [Google Scholar] [CrossRef]
- Perez, S.I.; Bernal, V.; Gonzales, P.N. Differences between sliding semi-landmark methods in geometric morphometrics, with an application to human craniofacial and dental variation. J. Anat. 2006, 208, 769–784. [Google Scholar] [CrossRef]
- Klingenberg, C.P.; Barluenga, M.; Meyer, A. Shape analysis of symmetric structures: Quantifying variation among individuals and asymmetry. Evolution 2002, 56, 1909–1920. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological statistics software package for education and data analysis. Palaeont. Electron. 2001, 4, 1–9. [Google Scholar]
- Perrett, J.J.; Mundfrom, D.J. Bonferroni procedure. In Encyclopedia of Research Design; Salkind, N.J., Ed.; Sage Publ.: Thousand Oaks, CA, USA, 2010; Volume 1, pp. 98–101. [Google Scholar]
- Pongparadon, S.; Zuccarello, G.C.; Phang, S.M.; Kawai, H.; Hanyuda, T.; Prathep, A. Diversity of Halimeda (Chlorophyta) from the Thai–Malay Peninsula. Phycologia 2015, 54, 349–366. [Google Scholar] [CrossRef]
- Mariani Colombo, P.; Orsenigo, M. Sea depth effects on the algal photosynthetic apparatus II. An electron microscopic study of the photosynthetic apparatus of Halimeda tuna (Chlorophyta, Siphonales) at—0–5 m and—6–0 m sea depths. Phycologia 1977, 16, 9–17. [Google Scholar] [CrossRef]
- Blair, S.M.; Norris, J.N. The deep-water species of Halimeda Lamouroux (Halimedaceae, Chlorophyta) from San Salvador Island, Bahamas: Species composition, distribution and depth records. Coral Reefs 1988, 6, 227–236. [Google Scholar] [CrossRef]
- Verbruggen, H.; De Clerck, O.; Schils, T.; Kooistra, W.H.C.F.; Coppejans, E. Evolution and phylogeography of Halimeda section Halimeda (Bryopsidales, Chlorophyta). Mol. Phyl. Evol. 2005, 37, 789–803. [Google Scholar] [CrossRef] [PubMed]
- John, G.P.; Scoffoni, C.; Sack, L. Allometry of cells and tissues within leaves. Am. J. Bot. 2013, 100, 1936–1948. [Google Scholar] [CrossRef]
- Larkum, A.W.D.; Salih, A.; Kühl, M. Rapid mass movement of chloroplasts during segment formation of the calcifying siphonalean green alga, Halimeda macroloba. PLoS ONE 2011, 6, e20841. [Google Scholar] [CrossRef]
- Pérez-Pérez, J.M.; Rubio-Díaz, S.; Dhondt, S.; Hernández-Romero, D.; Sánchez-Soriano, J.; Beemster, G.T.S.; Rosa Ponce, M.; Micol, J.L. Whole organ, venation and epidermal cell morphological variations are correlated in the leaves of Arabidopsis mutants. Plant Cell Environ. 2011, 34, 2200–2211. [Google Scholar] [CrossRef]
- Kojima, R.; Hanyuda, T.; Kawai, H. Taxonomic re-examination of Japanese Halimeda species using genetic markers, and proposal of a new species Halimeda ryukyuensis (Bryopsidales, Chlorophyta). Phycol. Res. 2015, 63, 178–188. [Google Scholar] [CrossRef]
- Cremen, M.C.M.; Huisman, J.M.; Marcelino, V.R.; Verbruggen, H. Taxonomic revision of Halimeda (Bryopsidales, Chlorophyta) in south-western Australia. Aust. Syst. Bot. 2016, 29, 41–54. [Google Scholar] [CrossRef]
- Ximenes, C.F.; Cassano, V.; Oliveira-Carvalho, M.F.; Bandeira-Pedrosa, M.E.; Gurgel, C.F.D.; Verbruggen, H.; Barreto Pereira, S.M. Systematics of the genus Halimeda (Bryopsidales, Chlorophyta) in Brazil including the description of Halimeda jolyana sp. nov. Phycologia 2017, 56, 369–381. [Google Scholar] [CrossRef]
- Kooistra, W.H.C.F.; Coppejans, E.G.G.; Payri, C. Molecular systematics, historical ecology, and phylogeography of Halimeda (Bryopsidales). Mol. Phyl. Evol. 2002, 24, 121–138. [Google Scholar] [CrossRef]
- Lin, W.C.; Liu, Y. A lattice-based MRF model for dynamic near-regular texture tracking. IEEE Trans. Pattern Anal. Mach. Intell. 2007, 29, 777–792. [Google Scholar] [CrossRef] [PubMed]



| Area | CS | CSM | PDs | No. Vertices | |
|---|---|---|---|---|---|
| area | --- | 0.978 | 0.123 | 0.131 | 0.479 |
| CS | 0.957 | --- | 0.218 | 0.220 | 0.615 |
| CSM | 0.015 | 0.047 | --- | 0.971 | 0.132 |
| PDs | 0.017 | 0.049 | 0.944 | --- | 0.134 |
| no. vertices | 0.229 | 0.378 | 0.017 | 0.018 | --- |
| Utricle area [mm2] | Df | SS | MS | η2 | partial η2 | F | Z | p |
| Plant | 6 | 0.00018 | 3.057 × 10−5 | 0.091 | 0.148 | 4.415 | 1.769 | 0.021 |
| Position | 3 | 0.00044 | 1.452 × 10−4 | 0.217 | 0.297 | 652.88 | 6.892 | 0.001 |
| Segment (plant) | 25 | 0.00017 | 6.925 × 10−6 | 0.086 | 0.140 | 3.377 | 3.324 | 0.001 |
| Plant:position | 18 | 0.00003 | 1.402 × 10−6 | 0.013 | 0.028 | 6.306 | 5.337 | 0.001 |
| Position:segment (plant) | 75 | 0.00015 | 2.051 × 10−6 | 0.077 | 0.126 | 9.225 | 12.361 | 0.001 |
| Residuals | 4672 | 0.00104 | 2.220 × 10−7 | 0.517 | ||||
| Total | 4799 | 0.00201 | ||||||
| CSM | Df | SS | MS | η2 | partial η2 | F | Z | p |
| Plant | 6 | 0.02199 | 0.00367 | 0.025 | 0.028 | 2.987 | 1.749 | 0.029 |
| Position | 3 | 0.01034 | 0.00345 | 0.012 | 0.013 | 21.276 | 3.467 | 0.001 |
| Segment (plant) | 25 | 0.03068 | 0.00123 | 0.036 | 0.039 | 2.407 | 2.609 | 0.003 |
| Plant:position | 18 | 0.00451 | 0.00025 | 0.005 | 0.006 | 1.548 | 1.369 | 0.082 |
| Position:segment (plant) | 75 | 0.03824 | 0.00051 | 0.044 | 0.048 | 3.146 | 6.798 | 0.001 |
| Residuals | 4672 | 0.75722 | 0.00016 | 0.877 | ||||
| Total | 4799 | 0.86300 | ||||||
| No. polygon vertices | Df | SS | MS | η2 | partial η2 | F | Z | p |
| Plant | 6 | 2.09 | 0.34894 | 0.001 | 0.001 | 1.886 | 1.098 | 0.129 |
| Position | 3 | 2.55 | 0.85163 | 0.001 | 0.001 | 1.983 | 1.087 | 0.131 |
| Segment (plant) | 25 | 4.63 | 0.18505 | 0.002 | 0.002 | 0.956 | −0.080 | 0.550 |
| Plant:position | 18 | 3.07 | 0.17068 | 0.002 | 0.002 | 0.398 | −2.630 | 0.993 |
| Position:segment (plant) | 75 | 14.52 | 0.19364 | 0.007 | 0.007 | 0.451 | −4.496 | 1.000 |
| Residuals | 4672 | 2006.1 | 0.42939 | 0.987 | ||||
| Total | 4799 | 2032.99 |
| Utricle area [mm2] | mean value | bottom | centre | Top | lateral |
| bottom | 0.001543 | --- | 0.000458 | 0.000821 | 0.000736 |
| centre | 0.002001 | 0.001 | --- | 0.000363 | 0.000278 |
| top | 0.002364 | 0.001 | 0.001 | --- | 0.000085 |
| lateral | 0.002279 | 0.001 | 0.001 | 0.001 | --- |
| CSM | mean value | bottom | centre | top | lateral |
| bottom | 0.02028 | --- | 0.00402 | 0.00006 | 0.00098 |
| centre | 0.01626 | 0.001 | --- | 0.00395 | 0.00304 |
| top | 0.02022 | 0.905 | 0.001 | --- | 0.00091 |
| lateral | 0.01930 | 0.109 | 0.001 | 0.138 | --- |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Neustupa, J.; Nemcova, Y. Morphometric Analysis of Surface Utricles in Halimeda tuna (Bryopsidales, Ulvophyceae) Reveals Variation in Their Size and Symmetry within Individual Segments. Symmetry 2020, 12, 1271. https://doi.org/10.3390/sym12081271
Neustupa J, Nemcova Y. Morphometric Analysis of Surface Utricles in Halimeda tuna (Bryopsidales, Ulvophyceae) Reveals Variation in Their Size and Symmetry within Individual Segments. Symmetry. 2020; 12(8):1271. https://doi.org/10.3390/sym12081271
Chicago/Turabian StyleNeustupa, Jiri, and Yvonne Nemcova. 2020. "Morphometric Analysis of Surface Utricles in Halimeda tuna (Bryopsidales, Ulvophyceae) Reveals Variation in Their Size and Symmetry within Individual Segments" Symmetry 12, no. 8: 1271. https://doi.org/10.3390/sym12081271
APA StyleNeustupa, J., & Nemcova, Y. (2020). Morphometric Analysis of Surface Utricles in Halimeda tuna (Bryopsidales, Ulvophyceae) Reveals Variation in Their Size and Symmetry within Individual Segments. Symmetry, 12(8), 1271. https://doi.org/10.3390/sym12081271






