MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation
Abstract
1. Introduction
2. Background
3. Materials and Methods
3.1. Cell Lines
3.2. Cell Colony Feature Extraction
- is the normalized gray level histogram counts for the current sliding window (obtained from the native RGB color space using the rgb2gray MatLab built-in function);
- and are the dimensions of the pixel matrix (, considering that the well, automatically detected, is circular);
- is the number of gray levels considered after the quantization (in our case, );
- is the local mean considering a sliding window of size .
3.3. Circle Hough Transform
3.4. Spatial FCM Clustering
- is the fuzzification constant (weighting exponent: ) that determines the fuzziness degree of the classification process. If , the FCM algorithm approximates hard K-means. In general, the higher the value, the greater the fuzziness degree will be (the most common choice is );
- is the fuzzy -partition of the data matrix ;
- is the squared distance between and , computed through an induced norm on (usually the Euclidean norm).
3.5. Surviving Fraction Quantification
- The Plating Efficiency (η) represents the fraction of colonies formed from cells that were not exposed to the treatment, considering the number of grown colonies and the number of plated cells, and must be determined for each experiment;
- The Survival Fraction (SF) represents the surviving fraction of cells after any treatment (i.e., irradiation or cytotoxic agent), calculated taking into account the PE of untreated cells, the number of colonies grown after treatment and the number of plated cells.
3.6. MF2C3 Parameter Settings
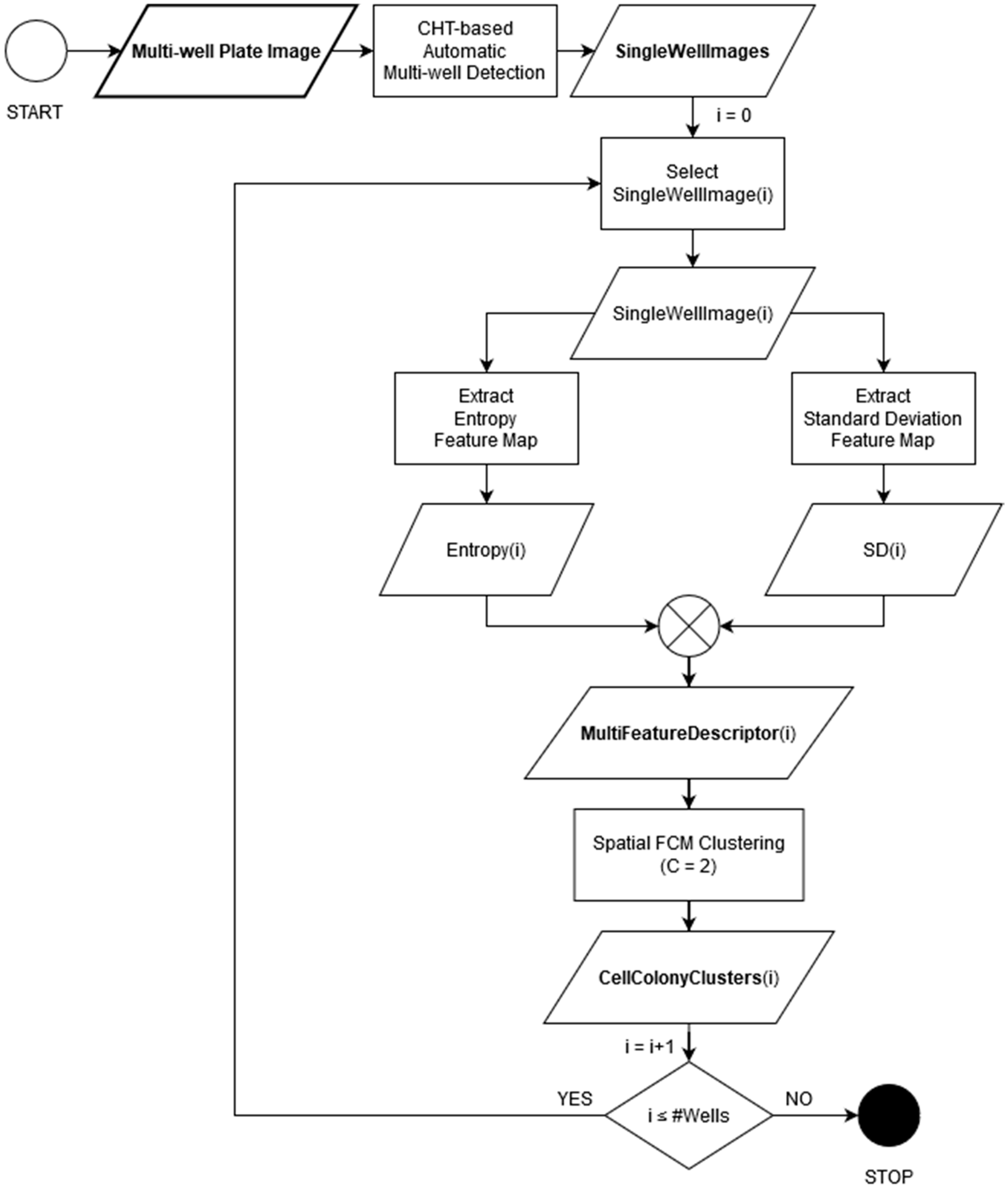
4. The Proposed Multi-Feature Automatic Clustering Approach
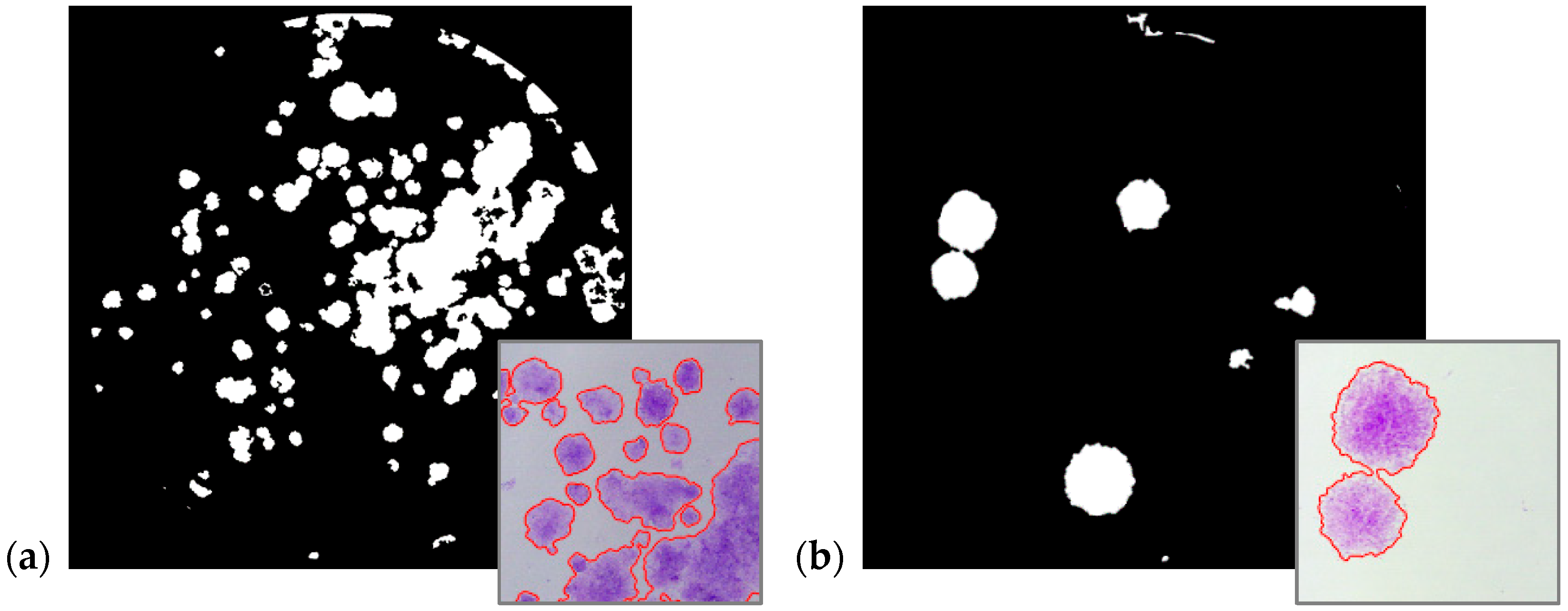
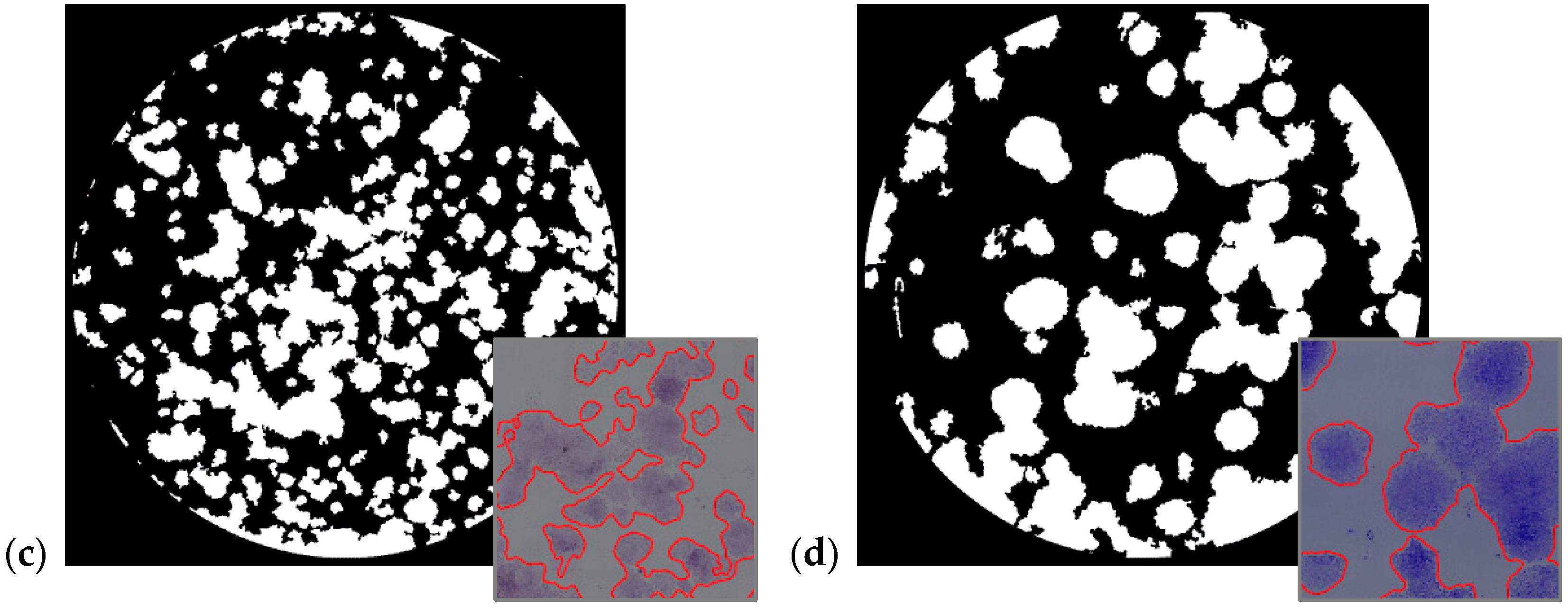
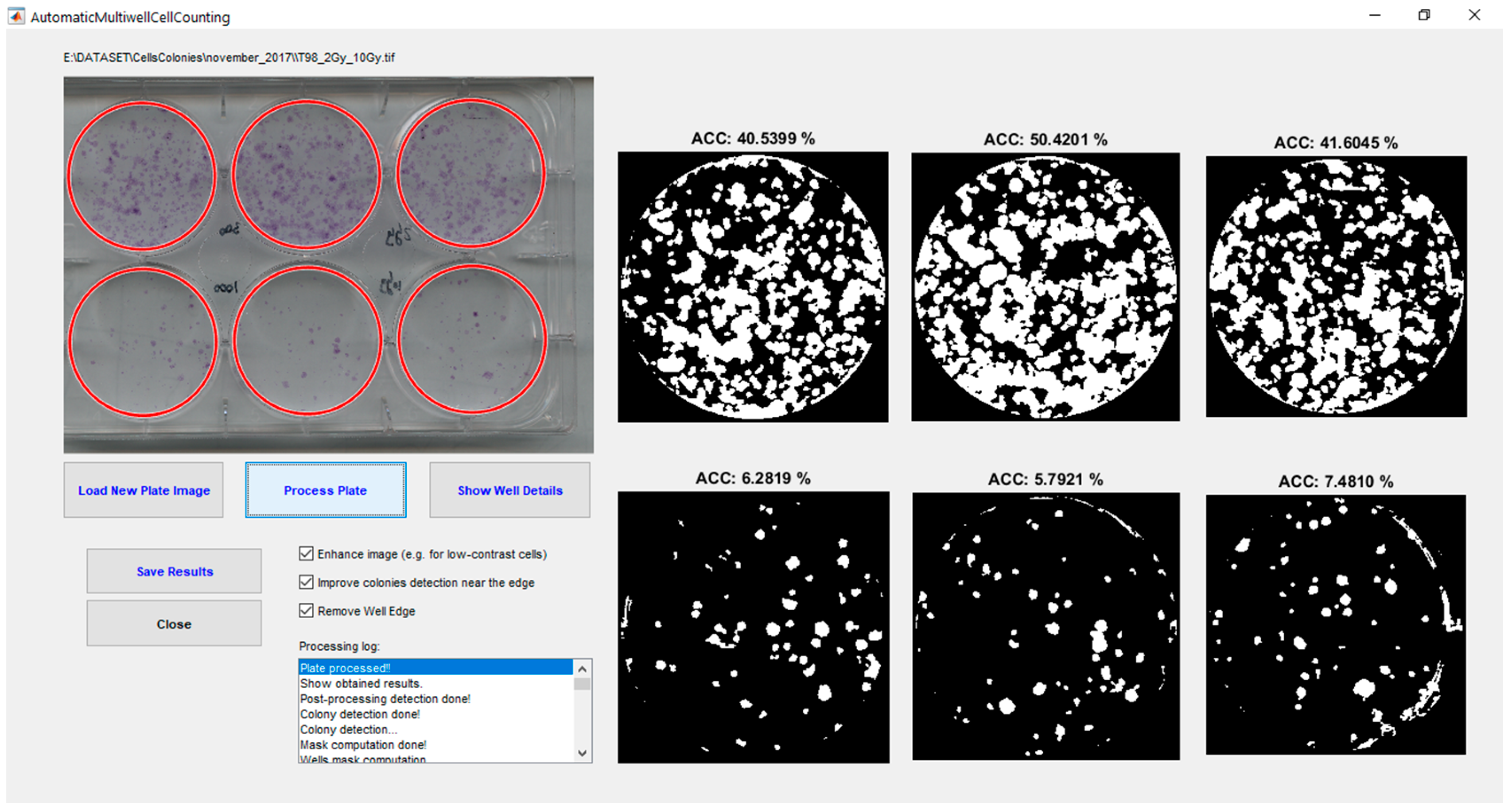
5. Experimental Results
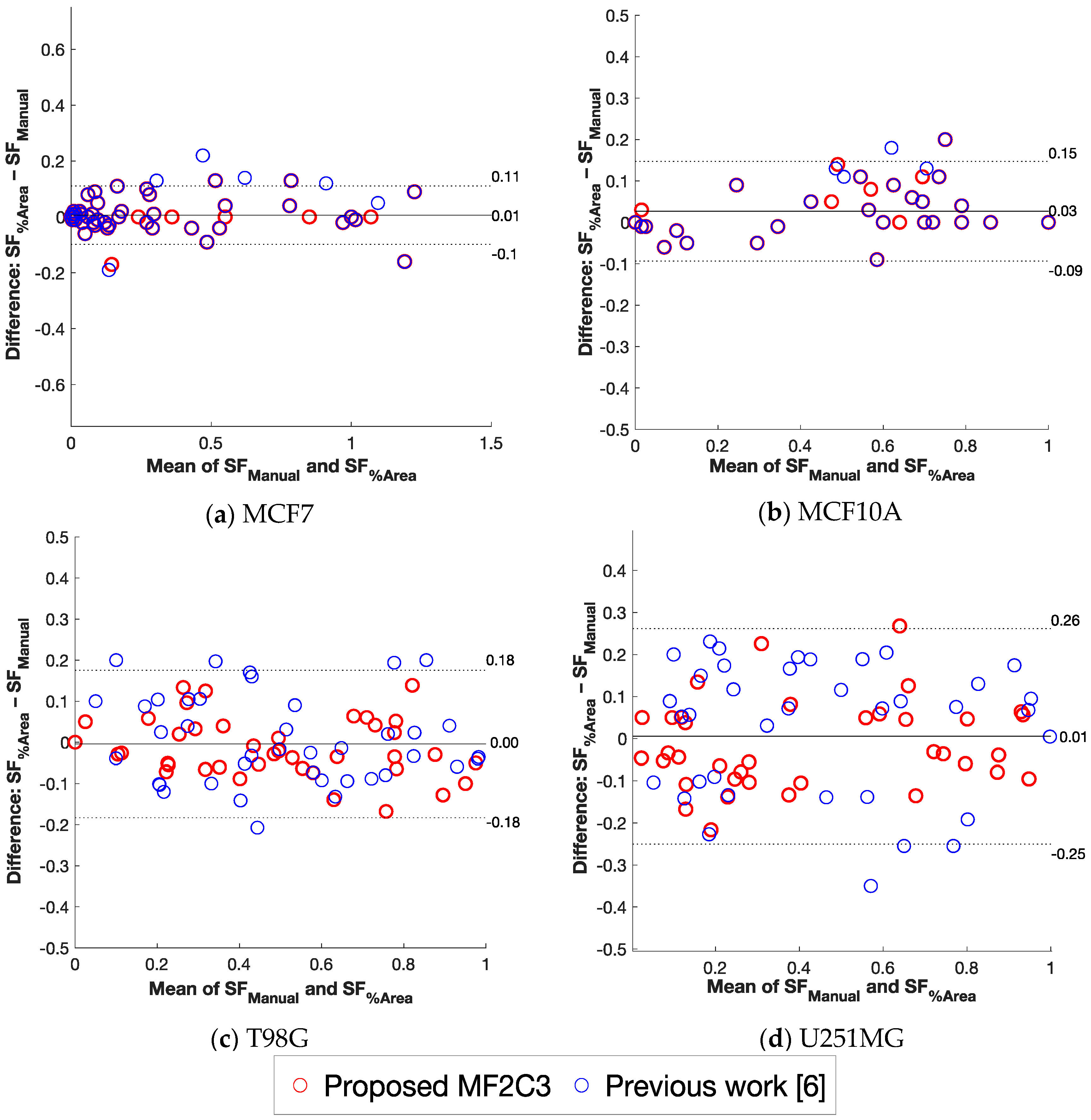
6. Discussion and Comparisons
- Fully automatic approach that does not require any human intervention, allowing for eliminating the subjectivity of the quantification;
- No need for specific hardware during the images acquisition phase. Since just a general-purpose flat-bed scanner is required, every laboratory can use our approach, without having to acquire any additional hardware;
- Using a multi-feature local descriptor as input of the sFCM clustering algorithm for cell colony detection. This novel approach, exploiting entropy and SD, was tested on four human cell lines and obtained excellent results.
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Puck, T.T.; Marcus, P.I. Action of x-rays on mammalian cells. J. Exp. Med. 1956, 103, 653–666. [Google Scholar] [CrossRef] [PubMed]
- Chapman, J.D. Can the two mechanisms of tumor cell killing by radiation be exploited for therapeutic gain? J. Radiat. Res. 2014, 55, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Franken, N.A.; Rodermond, H.M.; Stap, J.; Haveman, J.; van Bree, C. Clonogenic assay of cells in vitro. Nat. Protoc. 2006, 1, 2315–2319. [Google Scholar] [CrossRef] [PubMed]
- Lumley, M.A.; Burgess, R.; Billingham, L.J.; McDonald, D.F.; Milligan, D.W. Colony counting is a major source of variation in CFU-GM results between centres. Br. J. Haematol. 1997, 97, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Matsui, T.; Nuryadi, E.; Komatsu, S.; Hirota, Y.; Shibata, A.; Oike, T.; Nakano, T. Robustness of clonogenic assays as a biomarker for cancer cell radiosensitivity. Int. J. Mol. Sci. 2019, 20, 4148. [Google Scholar] [CrossRef]
- Militello, C.; Rundo, L.; Conti, V.; Minafra, L.; Cammarata, F.P.; Mauri, G.; Gilardi, M.C.; Porcino, N. Area-based cell colony surviving fraction evaluation: A novel fully automatic approach using general-purpose acquisition hardware. Comput. Biol. Med. 2017, 89, 454–465. [Google Scholar] [CrossRef]
- Chuang, K.S.; Tzeng, H.L.; Chen, S.; Wu, J.; Chen, T.J. Fuzzy c-means clustering with spatial information for image segmentation. Comput. Med. Imaging Graph. 2006, 30, 9–15. [Google Scholar] [CrossRef]
- Li, Y.l.; Shen, Y. An automatic fuzzy c-means algorithm for image segmentation. Soft Comput. 2010, 14, 123–128. [Google Scholar] [CrossRef]
- Li, B.N.; Chui, C.K.; Chang, S.; Ong, S.H. Integrating spatial fuzzy clustering with level set methods for automated medical image segmentation. Comput. Biol. Med. 2011, 41, 1–10. [Google Scholar] [CrossRef]
- Hough, P.V.C. Method and Means for Recognizing Complex Pattern. U.S. Patent 3,069,654, 18 December 1962. [Google Scholar]
- Duda, R.O.; Hart, P.E. Use of the Hough transformation to detect lines and curves in pictures. Commun. ACM 1972, 15, 11–15. [Google Scholar] [CrossRef]
- Thielmann, H.W.; Hagedorn, R.; Freber, W. Evaluation of colony-forming ability experiments using normal and DNA repair-deficient human fibroblast strains and an automatic colony counter. Cytometry 1985, 6, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Parry, R.L.; Chin, T.W.; Donahoe, P.K. Computer-aided cell colony counting. Biotechniques 1991, 10, 772–774. [Google Scholar] [PubMed]
- Wilson, I.G. Use of the IUL Countermat automatic colony counter for spiral plated total viable counts. Appl. Environ. Microbiol. 1995, 61, 3158–3160. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, D.P.; Pal, A.; Sarma, S.E.; Majumder, D.D. Bacterial colony counting using distance transform. Int. J. Biomed. Comput. 1995, 38, 131–140. [Google Scholar] [CrossRef]
- Hoekstra, S.J.; Tarka, D.K.; Kringle, R.O.; Hincks, J.R. Development of an automated bone marrow colony counting system. In Vitro Toxicol. 1998, 11, 207–213. [Google Scholar]
- Dobson, K.; Reading, L.; Scutt, A. A cost-effective method for the automatic quantitative analysis of fibroblastic colony-forming units. Calcif. Tissue Int. 1999, 65, 166–172. [Google Scholar] [CrossRef]
- Barber, P.R.; Vojnovic, B.; Kelly, J.; Mayes, C.R.; Boulton, P.; Woodcock, M.; Joiner, M.C. Automated counting of mammalian cell colonies. Phys. Med. Biol. 2001, 46, 63–76. [Google Scholar] [CrossRef]
- Bernard, R.; Kanduser, M.; Pernus, F. Model-based automated detection of mammalian cell colonies. Phys. Med. Biol. 2001, 46, 3061–3072. [Google Scholar] [CrossRef]
- Dahle, J.; Kakar, M.; Steen, H.B.; Kaalhus, O. Automated counting of mammalian cell colonies by means of a flat bed scanner and image processing. Cytom. Part A 2004, 60, 182–188. [Google Scholar] [CrossRef]
- Siragusa, M.; Dall’Olio, S.; Fredericia, P.M.; Jensen, M.; Groesser, T. Cell colony counter called CoCoNut. PLoS ONE 2018, 13, e0205823. [Google Scholar] [CrossRef]
- Clarke, M.L.; Burton, R.L.; Hill, A.N.; Litorja, M.; Nahm, M.H.; Hwang, J. Low-cost, high-throughput, automated counting of bacterial colonies. Cytom. Part A 2010, 77, 790–797. [Google Scholar] [CrossRef] [PubMed]
- Geissmann, Q. OpenCFU, a new free and open-source software to count cell colonies and other circular objects. PLoS ONE 2013, 8, e54072. [Google Scholar] [CrossRef] [PubMed]
- Chiang, P.J.; Tseng, M.J.; He, Z.S.; Li, C.H. Automated counting of bacterial colonies by image analysis. J. Microbiol. Methods 2015, 108, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.U.M.; Torelli, A.; Wolf, I.; Gretz, N. AutoCellSeg: Robust automatic colony forming unit (CFU)/cell analysis using adaptive image segmentation and easy-to-use post-editing techniques. Sci. Rep. 2018, 8, 7302. [Google Scholar] [CrossRef] [PubMed]
- Boukouvalas, D.T.; Prates, R.A.; Leal, C.R.L.; de Araújo, S.A. Automatic segmentation method for CFU counting in single plate-serial dilution. Chemometr. Intell. Lab. Syst. 2019, 195, 103889. [Google Scholar] [CrossRef]
- Najman, L.; Couprie, M.; Bertrand, G. Watersheds, mosaics, and the emergence paradigm. Discret. Appl. Math. 2005, 147, 301–324. [Google Scholar] [CrossRef]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Boukouvalas, D.T.; Belan, P.; Leal, C.R.L.; Prates, R.A.; de Araújo, S.A. Automated colony counter for single plate serial dilution spotting. In Proceedings of the Iberoamerican Congress on Pattern Recognition (CIARP), Madrid, Spain, 19–22 November 2018; Springer International Publishing: Cham, Switzerland, 2018; Volume 11401, pp. 410–418. [Google Scholar]
- Bewes, J.M.; Suchowerska, N.; McKenzie, D.R. Automated cell colony counting and analysis using the circular Hough image transform algorithm (CHiTA). Phys. Med. Biol. 2008, 53, 5991–6008. [Google Scholar] [CrossRef]
- Marotz, J.; Lubbert, C.; Eisenbeiss, W. Effective object recognition for automated counting of colonies in Petri dishes (automated colony counting). Comput. Methods Programs Biomed. 2001, 66, 183–198. [Google Scholar] [CrossRef]
- Niyazi, M.; Niyazi, I.; Belka, C. Counting colonies of clonogenic assays by using densitometric software. Radiat. Oncol. 2007, 2, 4. [Google Scholar] [CrossRef]
- Wouters, A.; Pauwels, B.; Lambrechts, H.A.J.; Pattyn, G.G.O.; Ides, J.; Baay, M.; Meijnders, P.; Lardon, F.; Vermorken, J.B. Counting clonogenic assays from normoxic and anoxic irradiation experiments manually or by using densitometric software. Phys. Med. Biol. 2010, 55, N167–N178. [Google Scholar] [CrossRef] [PubMed]
- Choudhry, P. High-throughput method for automated colony and cell counting by digital image analysis based on edge detection. PLoS ONE 2016, 11, e0148469. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Chattopadhyay, N.; Liu, W.J.; Chan, C.; Pignol, J.P.; Reilly, R.M. Optimized digital counting colonies of clonogenic assays using ImageJ software and customized macros: Comparison with manual counting. Int. J. Radiat. Biol. 2011, 87, 1135–1146. [Google Scholar] [CrossRef] [PubMed]
- Guzman, C.; Bagga, M.; Kaur, A.; Westermarck, J.; Abankwa, D. ColonyArea: An ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS ONE 2014, 9, e92444. [Google Scholar] [CrossRef]
- Szénási, S.; Vámossy, Z.; Kozlovszky, M. Evaluation and comparison of cell nuclei detection algorithms. In Proceedings of the 16th IEEE International Conference on Intelligent Engineering Systems (INES), Lisbon, Portugal, 13–15 June 2012; pp. 469–475. [Google Scholar] [CrossRef]
- Neve, R.M.; Chin, F.; Fridlyand, J.; Yeh, J.; Baehner, F.L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.P.; Tong, F.; et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef]
- Bravatà, V.; Minafra, L.; Russo, G.; Forte, G.I.; Cammarata, F.P.; Ripamonti, M.; Casarino, C.; Augello, G.; Costantini, F.; Barbieri, G.; et al. High dose ionizing radiation regulates gene expression changes in MCF7 breast cancer cell line. Anticancer Res. 2015, 35, 2577–2591. [Google Scholar]
- Minafra, L.; Bravatà, V.; Russo, G.; Forte, G.I.; Cammarata, F.P.; Ripamonti, M.; Candiano, G.; Cervello, M.; Giallongo, A.; Perconti, G.; et al. Gene expression profiling of MCF10A breast epithelial cells exposed to IOERT. Anticancer Res. 2015, 35, 3223–3234. [Google Scholar]
- Rennerfeldt, D.A.; Van Vliet, K.J. Concise review: When colonies are not clones: Evidence and implications of intracolony heterogeneity in mesenchymal stem cells. Stem Cells 2016, 34, 1135–1141. [Google Scholar] [CrossRef]
- Amsterdam, A.; Shezen, E.; Raanan, C.; Schreiber, L.; Slilat, Y.; Fabrikant, Y.; Melzer, E. Differential staining of γ synuclein in poorly differentiated compared to highly differentiated colon cancer cells. Oncol. Rep. 2012, 27, 1451–1454. [Google Scholar] [CrossRef][Green Version]
- Rundo, L.; Tangherloni, A.; Galimberti, S.; Cazzaniga, P.; Woitek, R.; Sala, E.; Nobile, M.S.; Mauri, G. HaraliCU: GPU-powered Haralick feature extraction on medical images exploiting the full dynamics of gray-scale levels. In Proceedings of the 14th International Conference on Parallel Computing Technologies (PaCT), Almaty, Kazakhstan, 19–23 August 2019; Springer International Publishing: Cham, Switzerland, 2019; Volume 11657, pp. 304–318. [Google Scholar] [CrossRef]
- Illingworth, J.; Kittler, J. The adaptive Hough transform. IEEE Trans. Pattern Anal. Mach. Intell. 1987, PAMI-9, 690–698. [Google Scholar] [CrossRef]
- Atherton, T.J.; Kerbyson, D.J. Size invariant circle detection. Image Vis. Comput. 1999, 17, 795–803. [Google Scholar] [CrossRef]
- Bezdek, J.C. Objective function clustering. In Pattern Recognition with Fuzzy Objective Function Algorithms, 1st ed.; Springer-Verlag New York, Inc.: Secaucus, NJ, USA, 1981; pp. 43–93. ISBN 978-1-4757-0452-5. [Google Scholar] [CrossRef]
- Rundo, L.; Militello, C.; Russo, G.; Garufi, A.; Vitabile, S.; Gilardi, M.C.; Mauri, G. Automated prostate gland segmentation based on an unsupervised fuzzy C-means clustering technique using multispectral T1w and T2w MR imaging. Information 2017, 8, 49. [Google Scholar] [CrossRef]
- Bezdek, J.C.; Ehrlich, R.; Full, W. FCM: The fuzzy c-means clustering algorithm. Comput. Geosci. 1984, 10, 191–203. [Google Scholar] [CrossRef]
- Lloyd, S. Least squares quantization in PCM. IEEE Trans. Inform. Theory 1982, 28, 129–137. [Google Scholar] [CrossRef]
- Udupa, J.K.; Samarasekera, S. Fuzzy connectedness and object definition: Theory, algorithms, and applications in image segmentation. Graph. Models Image Process. 1996, 58, 246–261. [Google Scholar] [CrossRef]
- Zhang, D.Q.; Chen, S.C. A novel kernelized fuzzy c-means algorithm with application in medical image segmentation. Artif. Intell. Med. 2004, 32, 37–50. [Google Scholar] [CrossRef]
- Caponetti, L.; Castellano, G.; Corsini, V. MR brain image segmentation: A framework to compare different clustering techniques. Information 2017, 8, 138. [Google Scholar] [CrossRef]
- Pham, D.L. Spatial models for fuzzy clustering. Comput. Vis. Image Underst. 2001, 84, 285–297. [Google Scholar] [CrossRef]
- Gong, M.; Liang, Y.; Shi, J.; Ma, W.; Ma, J. Fuzzy c-means clustering with local information and kernel metric for image segmentation. IEEE Trans. Image Process. 2012, 22, 573–584. [Google Scholar] [CrossRef]
- Rundo, L.; Militello, C.; Russo, G.; Vitabile, S.; Gilardi, M.C.; Mauri, G. GTVcut for neuro-radiosurgery treatment planning: An MRI brain cancer seeded image segmentation method based on a cellular automata model. Nat. Comput. 2017, 17, 521–536. [Google Scholar] [CrossRef]
- Bouteldja, S.; Kourgli, A. An efficient CBIR system for high resolution remote sensing images. In Proceedings of the Iberoamerican Congress on Pattern Recognition (CIARP), Madrid, Spain, 19–22 November 2018; Springer International Publishing: Cham, Switzerland, 2018; Volume 11401, pp. 392–400. [Google Scholar] [CrossRef]
- Rundo, L.; Tangherloni, A.; Nobile, M.S.; Militello, C.; Besozzi, D.; Mauri, G.; Cazzaniga, P. MedGA: A novel evolutionary method for image enhancement in medical imaging systems. Expert Syst. Appl. 2019, 119, 387–399. [Google Scholar] [CrossRef]
- Nanni, L.; Brahnam, S.; Salvatore, C.; Castiglioni, I. Texture descriptors and voxels for the early diagnosis of Alzheimer’s disease. Artif. Intell. Med. 2019, 97, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.L.; Rosenthal, R.; Rubin, D.B. Comparing correlated correlation coefficients. Psychol. Bull. 1992, 111, 172. [Google Scholar] [CrossRef]
- Bland, J.M.; Altman, D.G. Measuring agreement in method comparison studies. Stat. Methods Med. Res. 1999, 8, 135–160. [Google Scholar] [CrossRef]
- Minafra, L.; Porcino, N.; Bravatà, V.; Gaglio, D.; Bonanomi, M.; Amore, E.; Cammarata, F.P.; Russo, G.; Militello, C.; Savoca, G.; et al. Radiosensitizing effect of curcumin-loaded lipid nanoparticles in breast cancer cells. Sci. Rep. 2019, 9, 11134. [Google Scholar] [CrossRef]
- Gamarra, M.; Zurek, E.; Escalante, H.J.; Hurtado, L.; San-Juan-Vergara, H. Split and merge watershed: A two-step method for cell segmentation in fluorescence microscopy images. Biomed. Signal Process. Control 2019, 53, 101575. [Google Scholar] [CrossRef]
- Harris, L.A.; Frick, P.L.; Garbett, S.P.; Hardeman, K.N.; Paudel, B.B.; Lopez, C.F.; Quaranta, V.; Tyson, D.R. An unbiased metric of antiproliferative drug effect in vitro. Nat. Methods 2016, 13, 497–500. [Google Scholar] [CrossRef]
- Ortega, O.O.; Lopez, C.F. Interactive multiresolution visualization of cellular network processes. iScience 2020, 23, 100748. [Google Scholar] [CrossRef]


| Cell Line | Description | Number of Analyzed Wells |
|---|---|---|
| MCF7 1 | human breast epithelial carcinoma estrogen receptor-positive | 79 |
| MCF10A 1 | human non-tumorigenic breast epithelial | 40 |
| T98G | human glioblastoma | 40 |
| U251MG | human glioblastoma-astrocytoma | 40 |
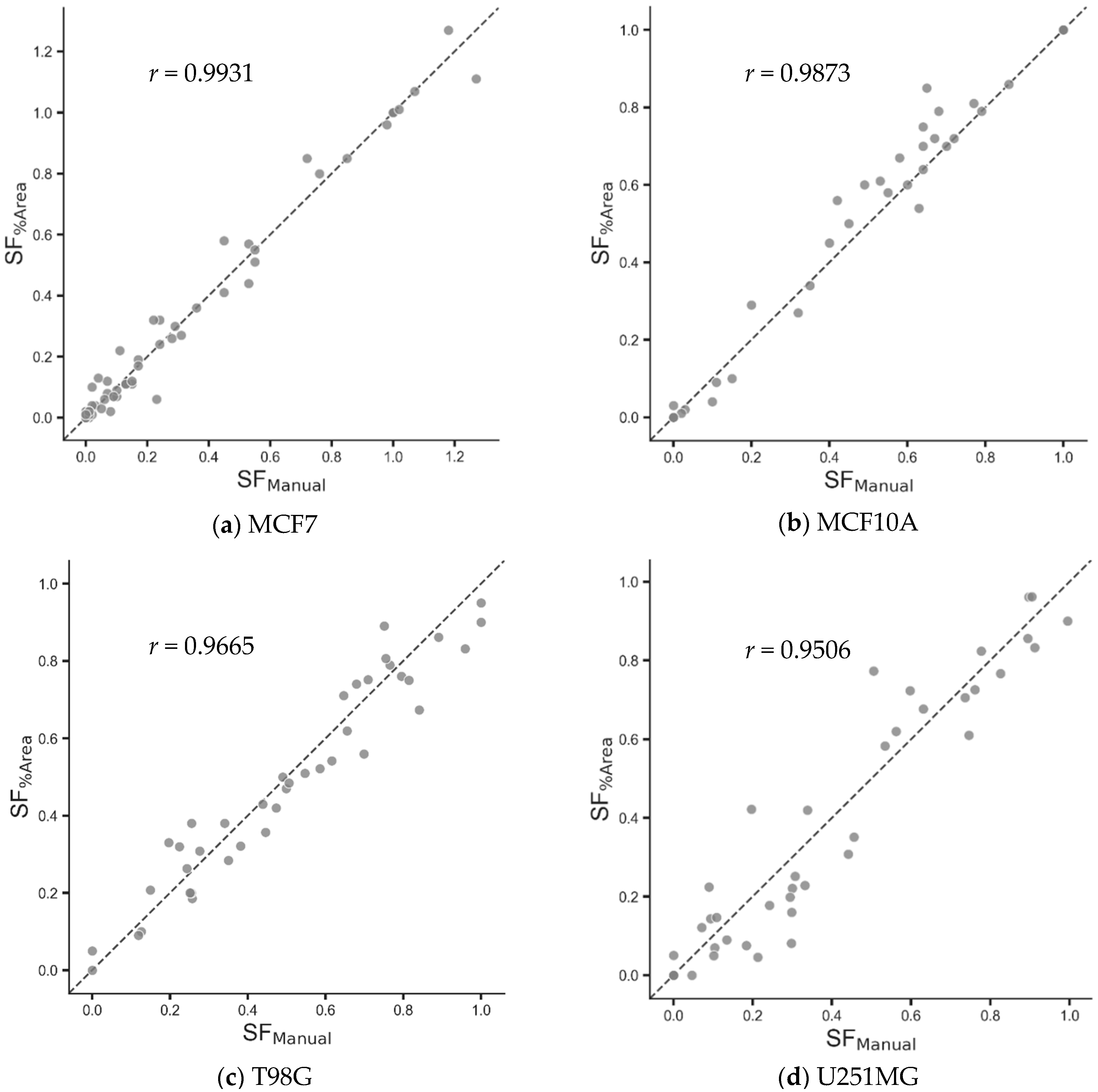
| Cell Line | Number of Samples | Pearson’s Correlation Coefficient (p Value) [6] | Pearson’s Correlation Coefficient (p Value) [MF2C3] | Fisher’s z-Transformation (p Value) |
|---|---|---|---|---|
| MCF7 | 79 | 0.9791 (<0.0001) | 0.9931(<0.0001) | 5.87 × 10−4 |
| MCF10A | 40 | 0.9682 (<0.0001) | 0.9873 (<0.0001) | 4.61 × 10−3 |
| T98G | 40 | 0.9237 (<0.0001) | 0.9665 (<0.0001) | 6.94 × 10−3 |
| U251MG | 40 | 0.8777 (<0.0001) | 0.9506 (<0.0001) | 4.23 × 10−3 |
| Method | Approach Type | Human Intervention | Specific Hardware Needed | Plate Position Detection | Cells Type |
|---|---|---|---|---|---|
| Barber et al., 2001 [18] | Semi-automatic | processing method selection | Yes | fixed plate position | HT29, A172, U118, IN1265 |
| Bernard et al., 2001 [19] | Supervised model | parameters tuning | Yes | fixed positioning mask | DC3F |
| Dahle et al., 2004 [20] | Semi-automatic | thresholding interaction | No | fixed positioning mask | V79, HaCaT |
| Siragusa et al., 2018 [21] | Semi-automatic | one parameter setting | Yes | fixed positioning mask | V79, HeLa |
| Clarke et al., 2010 [22] | Semi-automatic | parameters selection | No | fixed position | Escherichia coli, Streptococcus pneumoniae |
| Geissmann, 2013 [23] | Semi-automatic | interactive parameters initialization | No | automatic Petri dish detection | Staphylococcus aureus |
| Chiang et al., 2015 [24] | Automatic | N.A. | Yes | fixed position | Escherichia coli K12 |
| Khan et al., 2018 [25] | Automatic | N.A. | No | fixed positioning mask | Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, Staphylococcus aureus |
| Boukouvalas et al., 2019 [26] | Automatic | N.A. | No | automatic ROIs detection | Streptococcus mutans, Candida albicans, Staphylococcus aureus |
| Bewes et al., 2008 [30] | Automatic | N.A. | No | fixed flask position | MM576, NCI-H460, IMR-32 |
| Guzmán et al., 2014 [36] | Semi-automatic | alignment and cropping of the image | No | fixed positioning mask | T98G |
| Militello et al., 2017 [6] | Automatic | N.A. | No | automatic multi-well detection | MFC7, MCF10A |
| MF2C3 | Automatic | N.A. | No | automatic multi-well detection | MCF7, MCF10A, T98G, U251MG |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Militello, C.; Rundo, L.; Minafra, L.; Cammarata, F.P.; Calvaruso, M.; Conti, V.; Russo, G. MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation. Symmetry 2020, 12, 773. https://doi.org/10.3390/sym12050773
Militello C, Rundo L, Minafra L, Cammarata FP, Calvaruso M, Conti V, Russo G. MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation. Symmetry. 2020; 12(5):773. https://doi.org/10.3390/sym12050773
Chicago/Turabian StyleMilitello, Carmelo, Leonardo Rundo, Luigi Minafra, Francesco Paolo Cammarata, Marco Calvaruso, Vincenzo Conti, and Giorgio Russo. 2020. "MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation" Symmetry 12, no. 5: 773. https://doi.org/10.3390/sym12050773
APA StyleMilitello, C., Rundo, L., Minafra, L., Cammarata, F. P., Calvaruso, M., Conti, V., & Russo, G. (2020). MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation. Symmetry, 12(5), 773. https://doi.org/10.3390/sym12050773











