A Genetic XK-Means Algorithm with Empty Cluster Reassignment
Abstract
1. Introduction
2. Algorithms
2.1. Notations
2.2. Evaluation Strategies
2.3. XK-Means
2.4. EXK-Means
- Let be the number of empty clusters, .
- Find the most marginal point of each non-empty cluster: , where is the set of genes in the k-th cluster.
- Sort in descending order according to their distances to the corresponding centroids to get .
- Take the first genes from to form new centers .
- Adjust the partition of genes according to original centers and the new centers.
2.5. Genetic Operations
2.5.1. Label Vectors
2.5.2. Initialization
2.5.3. Selection
2.5.4. Mutation
2.5.5. Three Steps EXK-Means
2.6. Genetic XK-Means ( GXK-Means )
- Initialization: Set the population size N, the maximum number of iterations T, the mutation probability , the number of clusters K and the error tolerance . Let , and choose the initial population according to Section 2.5.2. In addition, choose the best individual from and denote it as super individual .
- Selection: Select a new population from according to Section 2.5.3, and denote it by .
- Mutation: Mutate each individual in according to Section 2.5.4, and get a new population denoted by .
- XK-Means: Perform XK-Means on three times to get the next generation population denoted by .
- Update the super individual: choose the best individual from and compare it with to get .
- Stop if either or (see ), otherwise go to 2 with .
2.7. GEXK-Means (Genetic EXK-Means)
- Initialization: Set the population size N, the maximum number of iterations T, the mutation probability , the number of clusters K and the error tolerance . Let , and choose the initial population according to Section 2.5.2. In addition, choose the best individual from and denote it as super individual .
- Selection: Select a new population from according to Section 2.5.3, and denote it by .
- Mutation: Mutate each individual in according to Section 2.5.4, and get a new population denoted by .
- EXK-Means: Perform the three steps EXK-Means on according to Section 2.5.5 to get the next generation population denoted by .
- Update the super individual: choose the best individual from and compare it with to get .
- Stop if either or (see ), otherwise go to 2 with .
3. Experimental Evaluation and Results
3.1. Data Sets and Parameters
| Population size | (cf. Section 2.6 and Section 2.7) |
| Mutation probability | (cf. Section 2.6 and Section 2.7) |
| Error tolerance | (cf. Section 2.2, Section 2.6 and Section 2.7) |
| (cf. , ) | |
| (cf. , ) | |
| (cf. Section 2.6 and Section 2.7) |
3.2. Experimental Results and Discussion
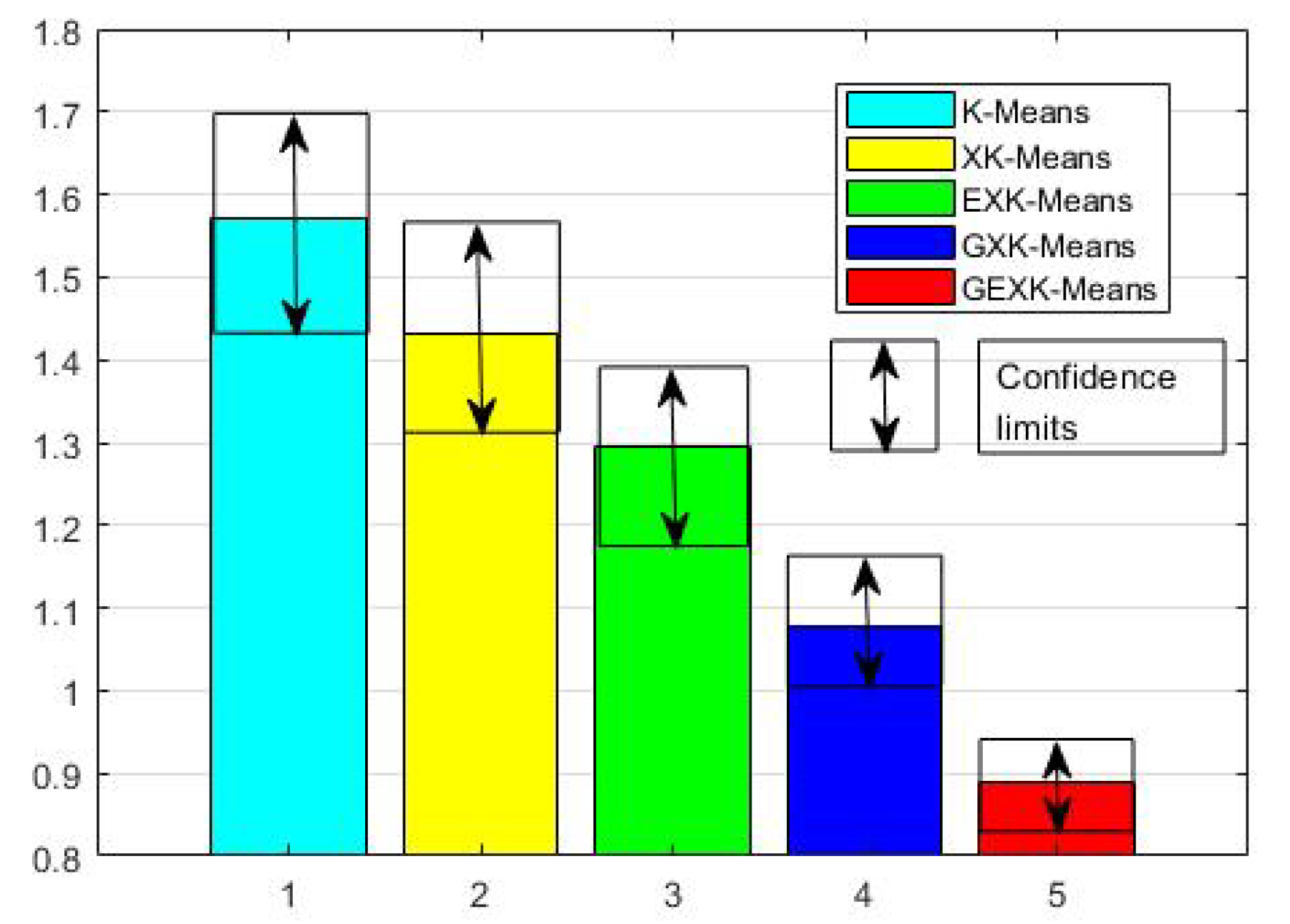
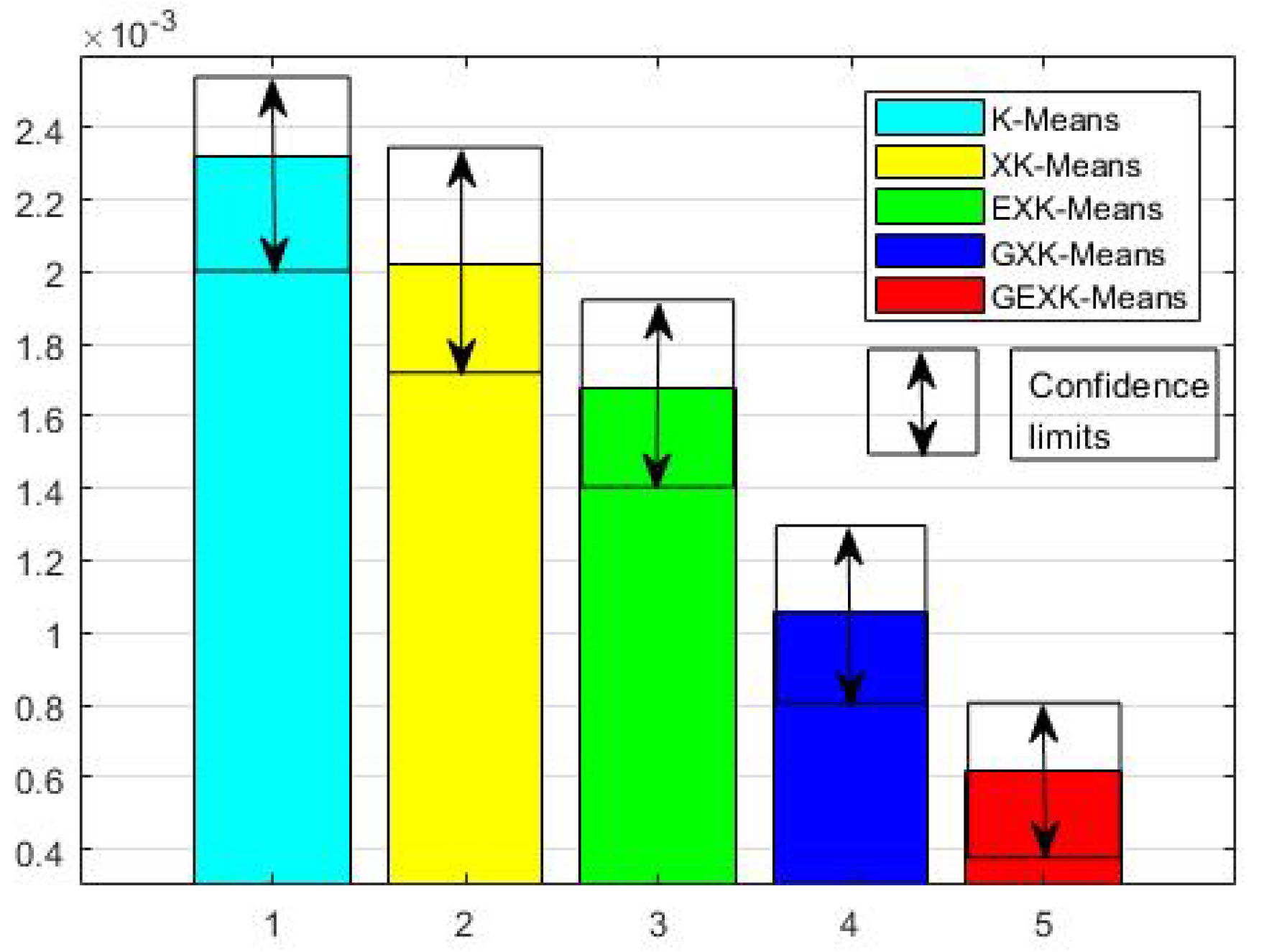
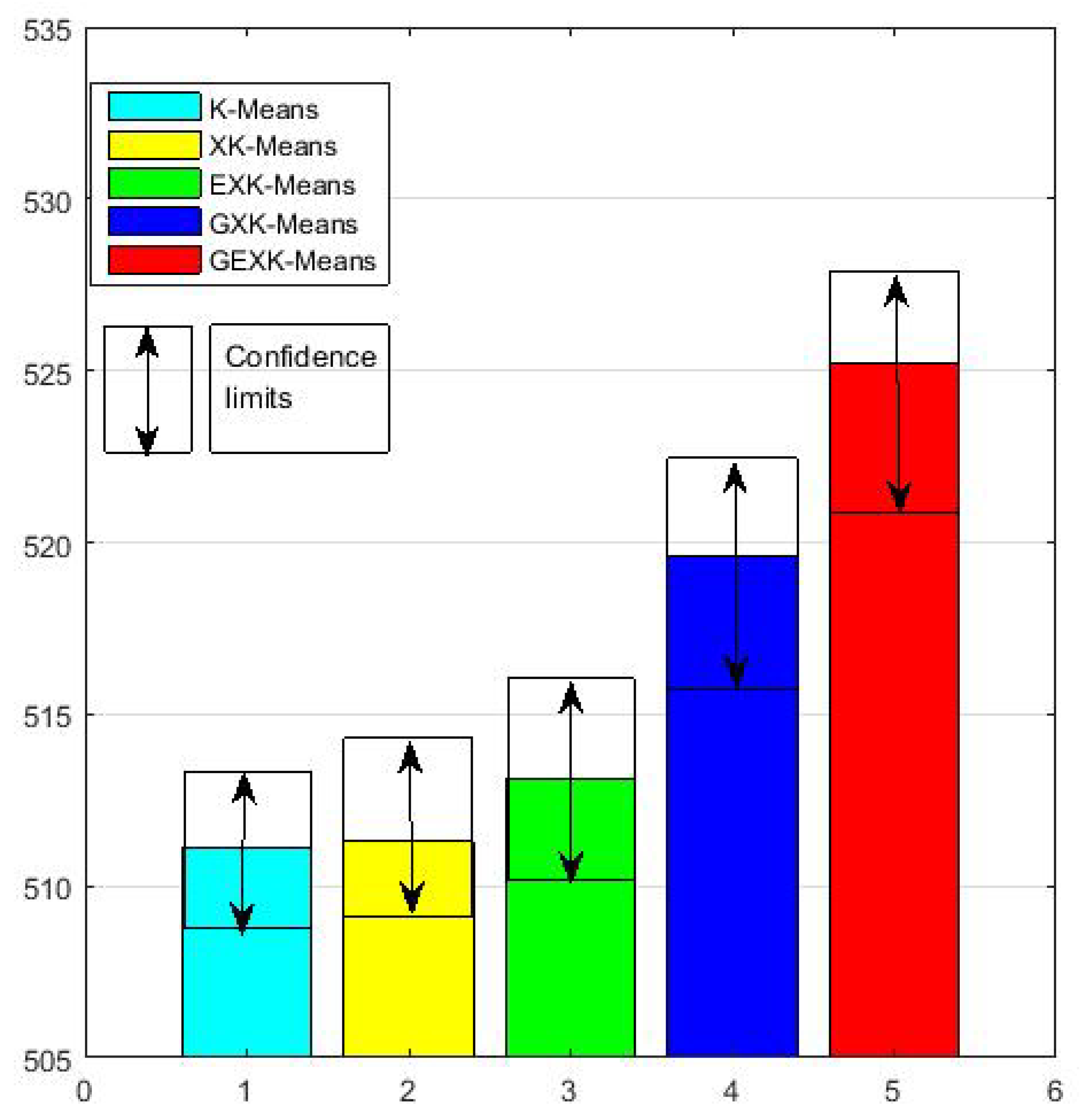
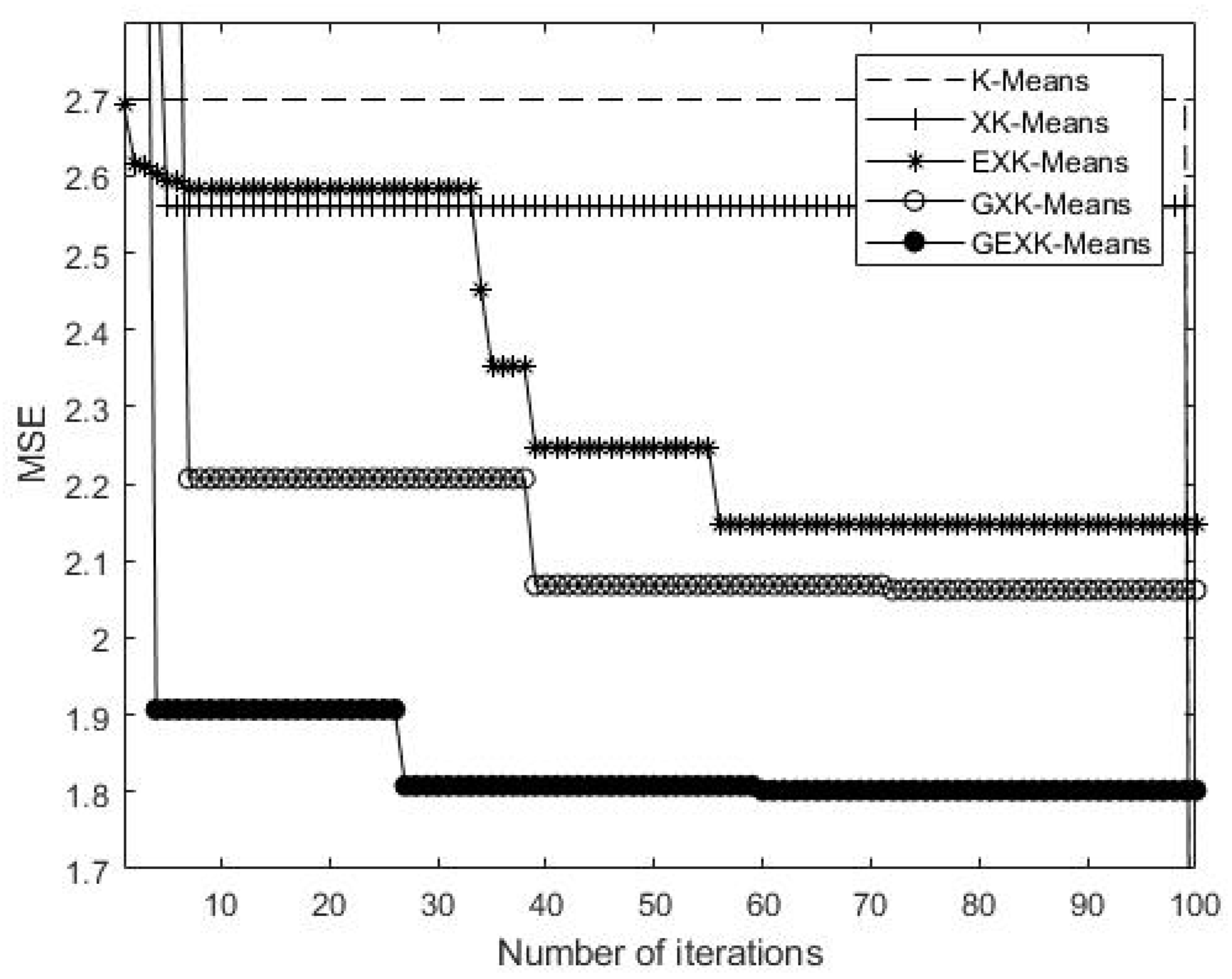
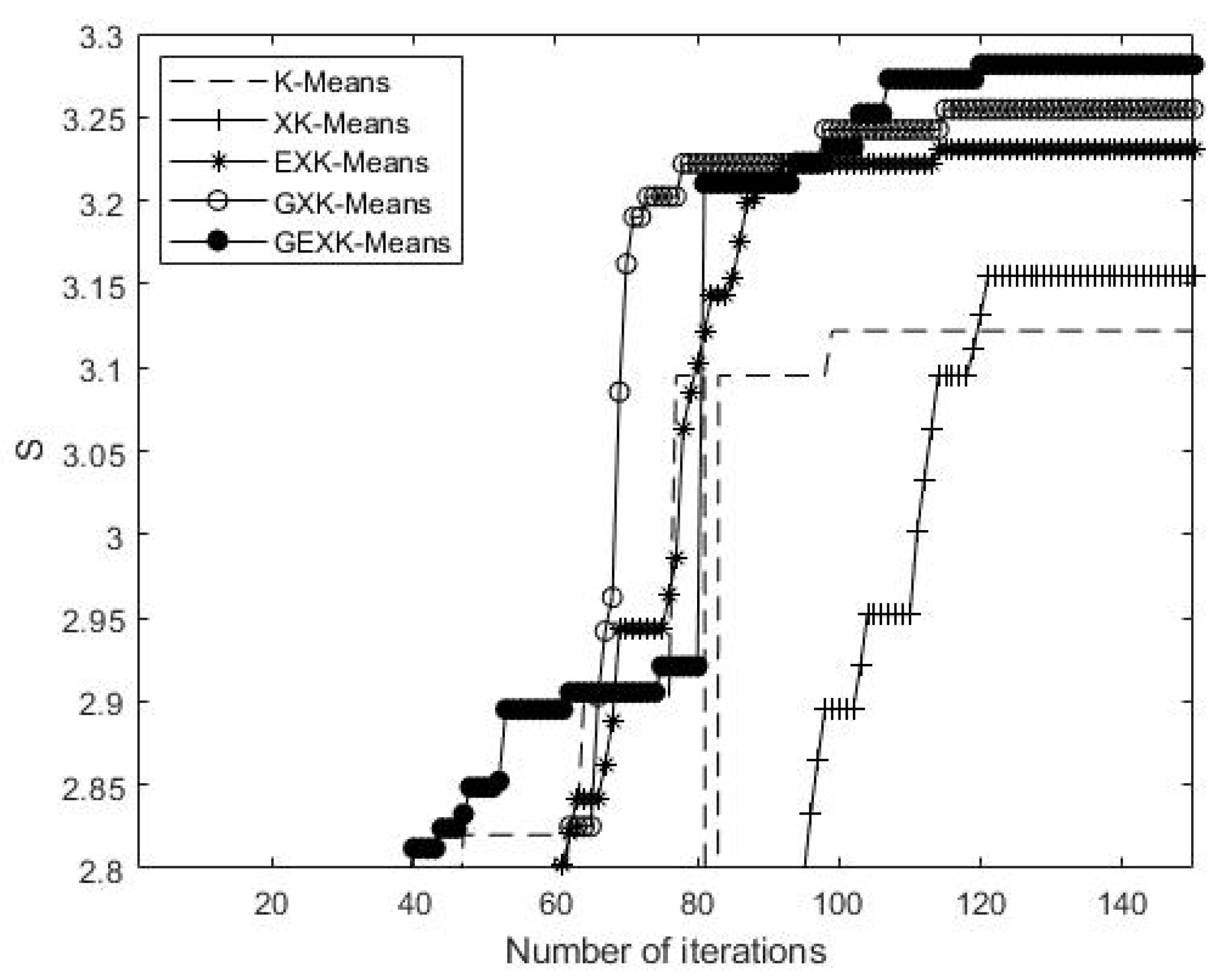
3.2.1. MSE, S, DB and XB Performances
3.2.2. Nemenyi Test
3.2.3. Computational Time
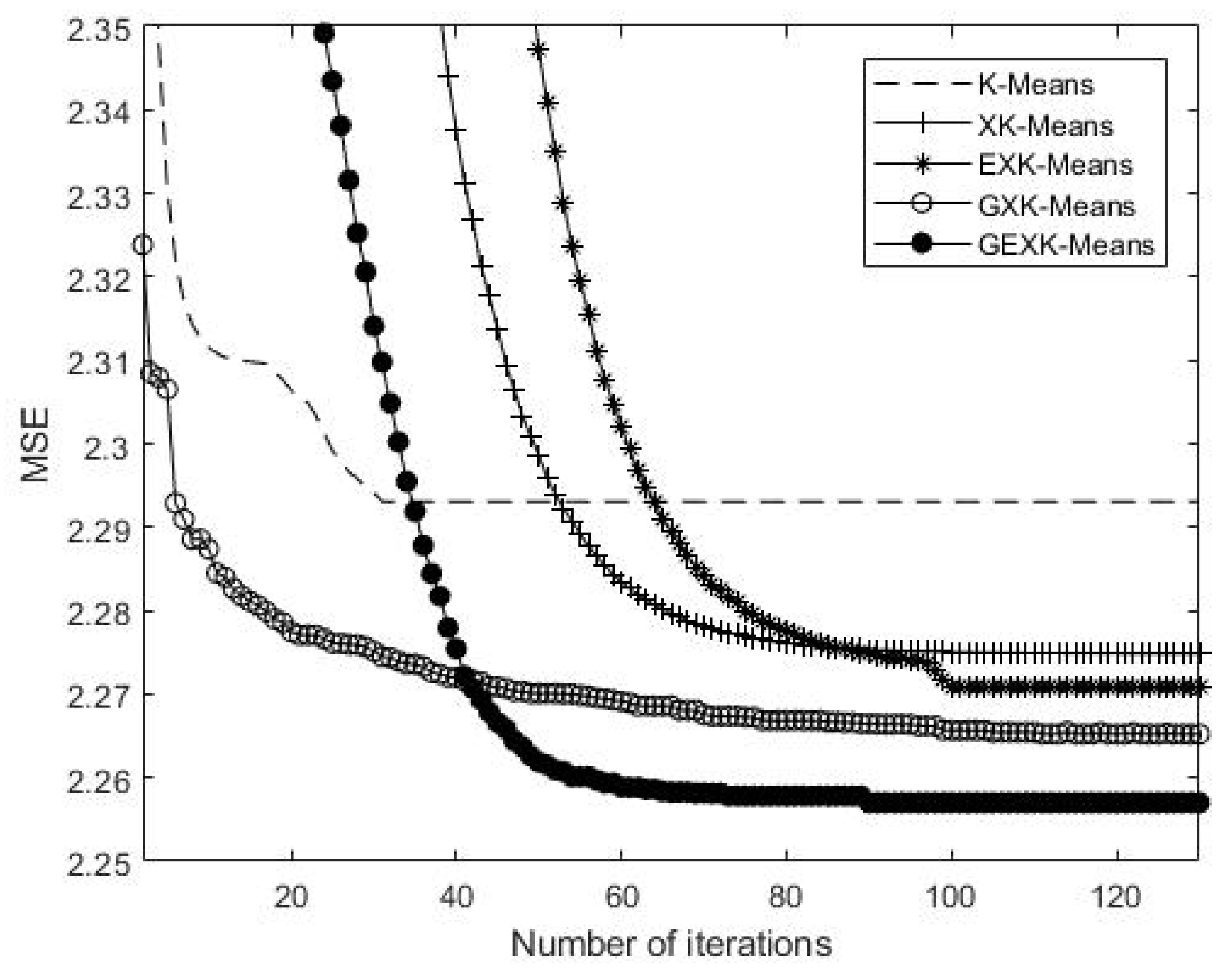
4. Convergence
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Steinhaus, H. Sur la division des corp materiels en parties. Bull. Acad. Polon. Sci. 1956, 3, 801–804. [Google Scholar]
- Macqueen, J. Some Methods for Classification and Analysis of MultiVariate Observations. In Proceedings of the 5th Berkeley Symposium on Mathematical Statistics and Probability, Los Angeles, CA, USA, 21 June–18 July 1967. [Google Scholar]
- Wlodarczyk-Sielicka, M. Importance of Neighborhood Parameters During Clustering of Bathymetric Data Using Neural Network. In Proceedings of the 22nd International Conference, Duruskininkai, Lithuania, 13–15 October 2016. [Google Scholar]
- Du, Z.; Wang, Y. PK-Means: A New Algorithm for Gene Clustering. Comput. Biol. Chem. 2008, 32, 243–247. [Google Scholar] [CrossRef] [PubMed]
- Lin, F.; Du, Z. A Novel Parallelization Approach for Hierarchical Clustering. Parallel Comput. 2005, 31, 523–527. [Google Scholar]
- Santhanam, T.; Padmavathi, M.S. Application of K-Means and Genetic Algorithms for Dimension Reduction by Integrating SVM for Diabetes Diagnosis. Procedia Comput. Sci. 2015, 47, 76–83. [Google Scholar] [CrossRef]
- Deep, K.; Thakur, M. A New Mutation Operator for Real Coded Genetic Algorithms. Appl. Math. Comput. 2007, 193, 211–230. [Google Scholar] [CrossRef]
- Ming, L.; Wang, Y. On Convergence Rate of a Class of Genetic Algorithms. In Proceedings of the World Automation Congress, Budapest, Hungary, 24–26 July 2006. [Google Scholar]
- Maulik, U. Genetic Algorithm Based Clustering Technique. Pattern Recognit. 2000, 33, 1455–1465. [Google Scholar] [CrossRef]
- Jones, D.R.; Beltramo, M.A. Solving Partitioning Problems with Genetic Algorithms. In Proceedings of the 4th International Conference on Genetic Algorithms, San Diego, CA, USA, 13–16 July 1991. [Google Scholar]
- Zheng, Y.; Jia, L.; Cao, H. Multi-Objective Gene Expression Programming for Clustering. Inf. Technol. Control 2012, 41, 283–294. [Google Scholar] [CrossRef][Green Version]
- Xie, X.L.; Beni, G. A Validity Measure for Fuzzy Clustering. IEEE Trans. Pattern Anal. Mach. Intell. 1991, 13, 841–847. [Google Scholar] [CrossRef]
- Liu, Y.G. Automatic Clustering Using Genetic Algorithms. Appl. Math. Comput. 2011, 218, 1267–1279. [Google Scholar] [CrossRef]
- Krishna, K.; Murty, M.N. Genetic K-Means Algorithm. IEEE Trans. Syst. Man Cybern. 1999, 29, 433–439. [Google Scholar] [CrossRef]
- Bouhmala, N.; Viken, A. Enhanced Genetic Algorithm with K-Means for the Clustering Problem. Int. J. Model. Optim. 2015, 5, 150–154. [Google Scholar] [CrossRef][Green Version]
- Sheng, W.G.; Tucker, A. Clustering with Niching Genetic K-means algorithm. In Proceedings of the 6th Annual Genetic and Evolutionary Computation Conference (GECCO 2004), Seattle, WA, USA, 26–30 June 2004. [Google Scholar]
- Zhou, X.B.; Gu, J.G. An Automatic K-Means Clustering Algorithm of GPS Data Combining a Novel Niche Genetic Algorithm with Noise and Density. ISPRS Int. J. Geo-Inf. 2017, 6, 392. [Google Scholar] [CrossRef]
- Islam, M.Z.; Estivill-Castro, V.; Rahman, M.A.; Bossomaier, T. Combining K-Means and a Genetic Algorithm through a Novel Arrangement of Genetic Operators for High Quality Clustering. Expert Syst. Appl. 2018, 91, 402–417. [Google Scholar] [CrossRef]
- Michael, L.; Sumitra, M. A Genetic Algorithm that Exchanges Neighboring Centers for k-means clustering. Pattern Recognit. Lett. 2007, 28, 2359–2366. [Google Scholar]
- Ishibuchi, H.; Yamamoto, T. Fuzzy Rule Selection by Multi-objective Genetic Local Search Algorithms and Rule Evaluation Measures in Data Mining. Fuzzy Sets Syst. 2004, 141, 59–88. [Google Scholar] [CrossRef]
- Zubova, J.; Kurasova, O. Dimensionality Reduction Methods: The Comparison of Speed and Accuracy. Inf. Technol. Control 2018, 47, 151–160. [Google Scholar] [CrossRef]
- Wozniak, M.; Polap, D. Object Detection and Recognition via Clustered Features. Neurocomputing 2018, 320, 76–84. [Google Scholar] [CrossRef]
- Anusha, M.; Sathiaseelan, G.R. Feature Selection Using K-Means Genetic Algorithm for Multi-objective Optimization. Procedia Comput. Sci. 2015, 57, 1074–1080. [Google Scholar] [CrossRef]
- Bezdek, J.C.; Ehrlich, R. FCM: The Fuzzy C-Means Clustering Algorithm. Comput. Geosci. 1984, 10, 191–203. [Google Scholar] [CrossRef]
- Indrajit, S.; Ujjwal, M. A New Multi-objective Technique for Differential Fuzzy Clustering. Appl. Soft Comput. 2011, 11, 2765–2776. [Google Scholar]
- Lam, Y.K.; Tsang, P.W.M. eXplotatory K-Means: A New Simple and Efficient Algorithm for Gene Clustering. Appl. Soft. Comput. 2012, 12, 1149–1157. [Google Scholar] [CrossRef]
- Davies, D.L.; Bouldin, D.W. A Cluster Separation Measure. IEEE Trans. Pattern Anal. Mach. Intell. 1979, PAMI-1, 224–227. [Google Scholar] [CrossRef]
- Liu, Y.W.; Chen, W.H. A SAS Macro for Testing Differences among Three or More Independent Groups Using Kruskal-Wallis and Nemenyi Tests. J. Huazhong Univ. Sci. Tech.-Med. 2012, 32, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Nemenyi, P. Distribution-Free Multiple Comparisons. Ph.D. Thesis, Princeton University, Princeton, NJ, USA, 1963. [Google Scholar]
- Fan, Y.; Hao, Z.O. Applied Statistics Analysis Using SPSS, 1st ed.; China Water Conservancy and Hydroelectricity Publishing House: Beijing, China, 2003; pp. 138–152. [Google Scholar]
- Chu, S.; DeRisi, J. The Transcriptional Program of Sporulation in Budding Yeast. Science 1998, 282, 699–705. [Google Scholar] [CrossRef] [PubMed]
- Spellman, P.T. Comprehensive Identification of Cell Cycle-regulated Genes of the Yeast Saccharomyces Cerevisiae by Microarray Hybridization. Mol. Biol. 1998, 9, 3273–3297. [Google Scholar] [CrossRef]
- Alizadeh, A.A.; Eisen, M.B. Distinct Types of Diffuse Large B-cell Lymphoma Identified by Gene Expression Profiling. Nature 2000, 403, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Yoon, D.; Lee, E.K. Robust Imputation Method for Missing Values in Microarray Data. BMC Bioinform. 2007, 8, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Troyanskaya, O.; Cantor, M. Missing Value Estimation Methods for DNA Microarrays. Bioinformatics 2001, 17, 520–525. [Google Scholar] [CrossRef]
- Corso, D.E.; Cerquitelli, T. METATECH: Meteorological Data Analysis for Thermal Energy CHaracterization by Means of Self-Learning Transparent Models. Energies 2018, 11, 1336. [Google Scholar] [CrossRef]
- Liu, G.G.; Zhuang, Z.; Guo, W.Z. A novel particle swarm optimizer with multi-stage transformation and genetic operation for VLSI routing. Energies 2018, 11, 1336. [Google Scholar]
- Rudolph, G. Convergence Analysis of Canonical Genetic Algorithms. IEEE Trans. Neural Netw. 1994, 5, 96–101. [Google Scholar] [CrossRef] [PubMed]


| Data Sets | No. of Vectors n | No. of Vectors with Missing Components <20% | No. of Vectors with Missing Component ≥20% | No. of Attributes D | No. of Classes K |
|---|---|---|---|---|---|
| Sporulation | 6023 | 413 | 198 | 7 | 16 |
| Yeast Cell Cycle | 6078 | 5498 | 680 | 77 | 256 |
| Lymphoma | 4022 | 3166 | 3 | 96 | 150 |
| Yeast | 1484 | 0 | 0 | 8 | 10 |
| Ecoli | 336 | 0 | 0 | 8 | 7 |
| Dermatology | 366 | 8 | 0 | 34 | 6 |
| Glass Identification | 214 | 0 | 0 | 10 | 7 |
| Image Segmentation | 2310 | 0 | 0 | 20 | 7 |
| Wine Quality | 4898 | 0 | 0 | 12 | 7 |
| Wireless Indoor Localization | 2000 | 0 | 0 | 7 | 4 |
| Statlog Vehicle | 946 | 0 | 0 | 18 | 4 |
| Page Blocks Classification | 5473 | 0 | 0 | 10 | 6 |
| Wine | 178 | 0 | 0 | 13 | 3 |
| Data Sets | Algorithm | MSE | S | DB | XB |
|---|---|---|---|---|---|
| (Lower the Better) | (Higher the Better) | (Lower the Better) | (Lower the Better) | ||
| Yeast Cell Cycle | K-Means | 2.3092 | 3.1637 | 2.6990 | |
| XK-Means | 2.2753 | 3.1773 | 2.5595 | ||
| EXK-Means | 2.2728 | 3.2170 | 2.0686 | ||
| GXK-Means | 2.2623 | 3.2560 | 2.4612 | ||
| GEXK-Means | 2.2572 | 3.2625 | 1.8070 | ||
| Sporulation | K-Means | 0.8959 | 2.7612 | 1.5781 | |
| XK-Means | 0.8968 | 2.7556 | 1.5261 | ||
| EXK-Means | 0.8987 | 2.7315 | 1.6226 | ||
| GXK-Means | 0.8961 | 2.7510 | 1.5207 | ||
| GEXK-Means | 0.8951 | 2.7674 | 1.3285 | ||
| Lymphoma | K-Means | 4.8762 | 7.1725 | 2.8532 | |
| XK-Means | 4.7683 | 7.2998 | 2.7294 | ||
| EXK-Means | 4.7764 | 7.2890 | 2.5627 | ||
| GXK-Means | 4.7520 | 7.3296 | 2.3285 | ||
| GEXK-Means | 4.7244 | 7.3787 | 2.1489 | ||
| Yeast | K-Means | 0.1613 | 0.3106 | 1.6854 | |
| XK-Means | 0.1586 | 0.3192 | 1.3592 | ||
| EXK-Means | 0.1607 | 0.3234 | 1.3701 | ||
| GXK-Means | 0.1581 | 0.3194 | 1.2621 | ||
| GEXK-Means | 0.1560 | 0.3296 | 0.9154 | ||
| Ecoli | K-Means | 0.2914 | 3.3151 | 1.4452 | |
| XK-Means | 0.2880 | 3.2775 | 1.0973 | ||
| EXK-Means | 0.2382 | 3.7191 | 0.7127 | ||
| GXK-Means | 0.2321 | 3.4022 | 0.3364 | ||
| GEXK-Means | 0.2268 | 3.7791 | 0.3021 |
| Data Sets | Algorithm | MSE | S | DB | XB |
|---|---|---|---|---|---|
| (Lower the Better) | (Higher the Better) | (Lower the Better) | (Lower the Better) | ||
| Glass Identification | K-Means | 1.2886 | 3.8306 | 2.0521 | |
| XK-Means | 1.1556 | 4.1060 | 1.3489 | ||
| EXK-Means | 1.2113 | 4.1161 | 1.4594 | ||
| GXK-Means | 1.1268 | 4.1218 | 0.9689 | ||
| GEXK-Means | 1.0250 | 4.1293 | 0.7230 | ||
| Image Segmentation | K-Means | 63.8058 | 168.3603 | 1.4437 | |
| XK-Means | 66.8793 | 169.0878 | 1.3625 | ||
| EXK-Means | 59.8254 | 169.7355 | 1.2243 | ||
| GXK-Means | 59.7865 | 186.6527 | 1.0697 | ||
| GEXK-Means | 59.5037 | 187.7843 | 1.0269 | ||
| Page Blocks Classification | K-Means | 645.1506 | 1.3881 | ||
| XK-Means | 643.3151 | 1.6223 | |||
| EXK-Means | 640.8521 | 1.1316 | |||
| GXK-Means | 605.8574 | 0.8693 | |||
| GEXK-Means | 601.7767 | 0.7865 | |||
| Wireless Indoor Localization | K-Mean | 10.2066 | 28.9495 | 1.6324 | |
| XK-Means | 10.1989 | 28.9495 | 1.7512 | ||
| EXK-Means | 10.1962 | 28.9490 | 1.14556 | ||
| GXK-Means | 10.1849 | 28.9210 | 0.9275 | ||
| GEXK-Means | 10.0854 | 28.9840 | 0.8816 | ||
| Dermatology | K-Mean | 5.7441 | 20.8749 | 1.4462 | |
| XK-Means | 5.8551 | 20.6550 | 1.2770 | ||
| EXK-Means | 5.7397 | 20.7767 | 1.2425 | ||
| GXK-Means | 5.7420 | 20.7639 | 1.2201 | ||
| GEXK-Means | 5.7252 | 20.9325 | 0.8816 | ||
| Statlog (Vehicle Silhouettes) | K-Mean | 53.8433 | 271.5335 | 0.8871 | |
| XK-Means | 53.8433 | 271.5335 | 1.2066 | ||
| EXK-Means | 53.6535 | 269.1440 | 1.0323 | ||
| GXK-Means | 53.5880 | 270.6111 | 0.6289 | ||
| GEXK-Means | 53.4423 | 301.5472 | 0.4893 | ||
| Wine Quality | K-Mean | 14.2767 | 58.1880 | 0.9625 | |
| XK-Means | 14.2021 | 58.2228 | 0.9658 | ||
| EXK-Means | 14.2090 | 58.5258 | 0.9420 | ||
| GXK-Means | 14.1382 | 58.5540 | 0.8499 | ||
| GEXK-Means | 14.1032 | 58.6269 | 0.6973 | ||
| Wine | K-Means | 93.0094 | 470.2573 | 0.8236 | |
| XK-Means | 93.2120 | 470.2573 | 0.5436 | ||
| EXK-Means | 93.0092 | 469.4700 | 0.6360 | ||
| GXK-Means | 92.9745 | 470.3015 | 0.5275 | ||
| GEXK-Means | 92.8682 | 504.1908 | 0.3211 |
| Groups for Comparison | Evaluation Technique | Pr |
|---|---|---|
| XK-Means vs. EXK-Means | MSE | 0.8994 |
| S | 0.6691 | |
| DB | 0.0421 | |
| XB | 0.4539 | |
| GXK-Means vs. GEXK-Means | MSE | 0.0492 |
| S | 0.0412 | |
| DB | 0.0409 | |
| XB | 0.0403 |
| Data Sets | Evaluation Technique | K -Means | XK -Means | EXK -Means | GXK -Means | GEXK -Means | Machine |
|---|---|---|---|---|---|---|---|
| Sporulation | MSE | 11.368 | 11.689 | 12.56 | 289.656 | 688.552 | M2 |
| S | 11.464 | 11.792 | 12.61 | 256.782 | 689.457 | M2 | |
| DB | 10.424 | 10.569 | 10.480 | 239.529 | 662.183 | M2 | |
| XB | 9.632 | 9.881 | 9.878 | 269.364 | 396.33 | M2 | |
| Yeast Cell Cycle | MSE | 58.65 | 60.213 | 64.52 | 4867.560 | 5432.23 | M2 |
| S | 58.87 | 60.351 | 65.11 | 4789.108 | 5433.69 | M2 | |
| DB | 69.425 | 72.37 | 73.06 | 4965.682 | 5654.75 | M2 | |
| XB | 52.584 | 56.321 | 59.66 | 4754.538 | 5014.56 | M2 | |
| Lymphoma | MSE | 69.425 | 70.43 | 74.332 | 4962.224 | 5321.6 | M2 |
| S | 69.661 | 71.34 | 74.994 | 4987.300 | 5324.1 | M2 | |
| DB | 72.135 | 77.483 | 74.286 | 4753.626 | 5332.42 | M2 | |
| XB | 62.526 | 66.36 | 68.665 | 4665.186 | 5026.559 | M2 | |
| Glass Identification | MSE | 0.3675 | 0.368 | 0.2975 | 19.566 | 20.824 | M1 |
| S | 0.3678 | 0.371 | 0.2980 | 19.960 | 20.885 | M1 | |
| DB | 0.505 | 0.535 | 0.536 | 19.325 | 27.687 | M1 | |
| XB | 0.267 | 0.303 | 0.304 | 16.263 | 19.806 | M1 | |
| Image Segmentation | MSE | 4.552 | 4.230 | 4.336 | 298.128 | 361.755 | M1 |
| S | 4.578 | 4.257 | 4.402 | 296.867 | 363.455 | M1 | |
| DB | 4.564 | 4.663 | 4.718 | 286.692 | 382.924 | M1 | |
| XB | 4.565 | 4.131 | 4.183 | 263.960 | 332.092 | M1 | |
| Page Blocks Classification | MSE | 4.039 | 4.078 | 4.122 | 382.663 | 395.273 | M1 |
| S | 4.165 | 4.216 | 4.269 | 378.632 | 397.421 | M1 | |
| DB | 7.859 | 7.649 | 7.947 | 376.657 | 520.118 | M1 | |
| XB | 4.596 | 4.622 | 4.264 | 296.186 | 390.57 | M1 | |
| Yeast | MSE | 1.810 | 1.964 | 1.870 | 159.200 | 165.08 | M1 |
| S | 1.914 | 1.998 | 1.978 | 161.230 | 167.1 | M1 | |
| DB | 3.630 | 3.784 | 3.800 | 163.620 | 241.346 | M1 | |
| XB | 1.718 | 1.817 | 1.847 | 148.960 | 161.346 | M1 | |
| Wireless Indoor Localization | MSE | 1.428 | 1.430 | 1.433 | 119.630 | 122.278 | M1 |
| S | 1.473 | 1.459 | 1.62 | 120.775 | 124.512 | M1 | |
| DB | 2.843 | 2.760 | 2.840 | 124.360 | 176.689 | M1 | |
| XB | 1.436 | 1.445 | 1.418 | 98.641 | 120.628 | M1 | |
| Ecoli | MSE | 0.408 | 0.437 | 0.437 | 29.611 | 30.494 | M1 |
| S | 0.409 | 0.441 | 0.439 | 29.845 | 30.569 | M1 | |
| DB | 0.736 | 0.858 | 0.882 | 28.010 | 43.321 | M1 | |
| XB | 0.399 | 0.447 | 0.490 | 29.380 | 31.22 | M1 | |
| Dermatology | MSE | 0.634 | 0.635 | 0.710 | 23.450 | 24.829 | M1 |
| S | 0.639 | 0.651 | 0.786 | 24.687 | 25.854 | M1 | |
| DB | 0.78 | 0.832 | 0.857 | 24.312 | 26.758 | M1 | |
| XB | 0.406 | 0.542 | 0.516 | 23.720 | 26.811 | M1 | |
| Statlog (Vehicle Silhouettes) | MSE | 0.705 | 0.726 | 0.768 | 53.856 | 57.061 | M1 |
| S | 0.712 | 0.732 | 0.798 | 55.289 | 58.067 | M1 | |
| DB | 1.298 | 1.303 | 1.351 | 58.450 | 77.966 | M1 | |
| XB | 0.590 | 0.634 | 0.642 | 43.892 | 51.137 | M1 | |
| Wine Quality | MSE | 4.538 | 4.658 | 4.758 | 386.680 | 444.963 | M1 |
| S | 4.563 | 4.713 | 4.799 | 379.668 | 446.921 | M1 | |
| DB | 8.435 | 8.512 | 8.571 | 412.654 | 565.498 | M1 | |
| XB | 4.463 | 4.499 | 4.635 | 356.215 | 421.432 | M1 | |
| Wine | MSE | 0.159 | 0.182 | 0.179 | 10.226 | 11.133 | M1 |
| S | 0.161 | 0.188 | 0.182 | 10.129 | 11.186 | M1 | |
| DB | 0.267 | 0.304 | 0.309 | 11.636 | 13.852 | M1 | |
| XB | 0.152 | 0.172 | 0.174 | 8.385 | 10.130 | M1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hua, C.; Li, F.; Zhang, C.; Yang, J.; Wu, W. A Genetic XK-Means Algorithm with Empty Cluster Reassignment. Symmetry 2019, 11, 744. https://doi.org/10.3390/sym11060744
Hua C, Li F, Zhang C, Yang J, Wu W. A Genetic XK-Means Algorithm with Empty Cluster Reassignment. Symmetry. 2019; 11(6):744. https://doi.org/10.3390/sym11060744
Chicago/Turabian StyleHua, Chun, Feng Li, Chao Zhang, Jie Yang, and Wei Wu. 2019. "A Genetic XK-Means Algorithm with Empty Cluster Reassignment" Symmetry 11, no. 6: 744. https://doi.org/10.3390/sym11060744
APA StyleHua, C., Li, F., Zhang, C., Yang, J., & Wu, W. (2019). A Genetic XK-Means Algorithm with Empty Cluster Reassignment. Symmetry, 11(6), 744. https://doi.org/10.3390/sym11060744




