1. Introduction
Precision agriculture, which focuses on optimizing production by accounting for variabilities and dealing with uncertainties in agricultural systems, has been under active research in recent years [
1]. Feature monitoring and plant phenotyping are essential parts of precision agriculture. They can help in modeling the growth process of plants and guide farmers to obtain higher yields with appropriate fertilizer, irrigation, and disease control [
2,
3].
Traditional plant phenotyping, involves a large number of manual measurements, and this has been identified as the current bottleneck in modern plant breeding and research programs [
4]. The number of leaves of a plant is considered one of the critical phenotypic metrics related to its development and growth stages [
5,
6], flowering time [
7], and water condition. The traditional manual measurement is slow, tedious, and expensive. Therefore, several image-based and machine learning technologies have been introduced for leaf counting. However, counting leaves automatically is challenging [
8], due to a plant’s rapid growth and leaf occlusion and illumination problems. Moreover, most study on leaf counting are based on rosette plants, and the relevant algorithms are not suitable for maize plants. Considering this, we designed a model suitable for counting maize leaves.
In this study, we estimate the number of leaves on a maize plant at different growth stages. The problem is posed as a nonlinear regression problem, which does not require segmenting individual leaf instances. First, features are extracted from each sample image. Then these feature vectors are used to regress the number of leaves. For this model, the input is a maize image, and the output is the number of leaves.
Effective feature extraction is an important step for leaf counting regression and plant phenotyping research. Over the past years, substantial efforts have been dedicated to developing robust feature representation methods in different domains. The histogram of oriented gradients (HOG) has been used to detect the tasseling stage of maize [
9]. Then, mid-level feature methods, such as wavelet transform and the Fisher vector, are used as feature descriptors since they attract much attention. In [
10], the authors used wavelet transform to extract energy features and detect maize water stress. FV coding is combined with scale-invariant feature transforms (SIFT) for object detection [
11].
Recently, deep learning, particularly the use of deep convolutional neural networks (CNN), has become the new state-of-the-art solution for object detection, recognition, and regression. Compared with traditional feature descriptors, the convolutional layer of CNN can extract low-level to high-level features. As the number of convolutional layers increases, more abstract features are extracted. Furthermore, more convolutional layers mean more parameters to be trained, but when the training samples are far fewer than the parameters, the risk of model over-fitting will increase. Therefore, some strategies of reducing parameters have been proposed such as Google Inception Net V3 [
12] and residual networks [
13]. For instance, in [
14] Google Inception Net was used to identify leaf species. In addition, the traditional method is used to optimize network parameters. Such as adding constraint to optimize the parameters in the CNN output to improve the accuracy of low-accuracy classes [
15].
Few recent works have demonstrated that the middle layer of CNN contains a large amount of useful information, which can improve the discrimination of feature representation. One example is that it can improve the discrimination of feature representation. In [
16], the authors converted the input image into the multi-scale image and fixed-size image. Then the multi-scale image and the fixed-size image was fed into CNN with the same structure separately. Subsequently, the features of each convolutional layer were extracted from the multi-scale image and the features of the full connection layer are extracted from the fixed-size image. After Fisher vector coding and principal component analysis dimensionality reduction, these features were fed into the support vector machine.
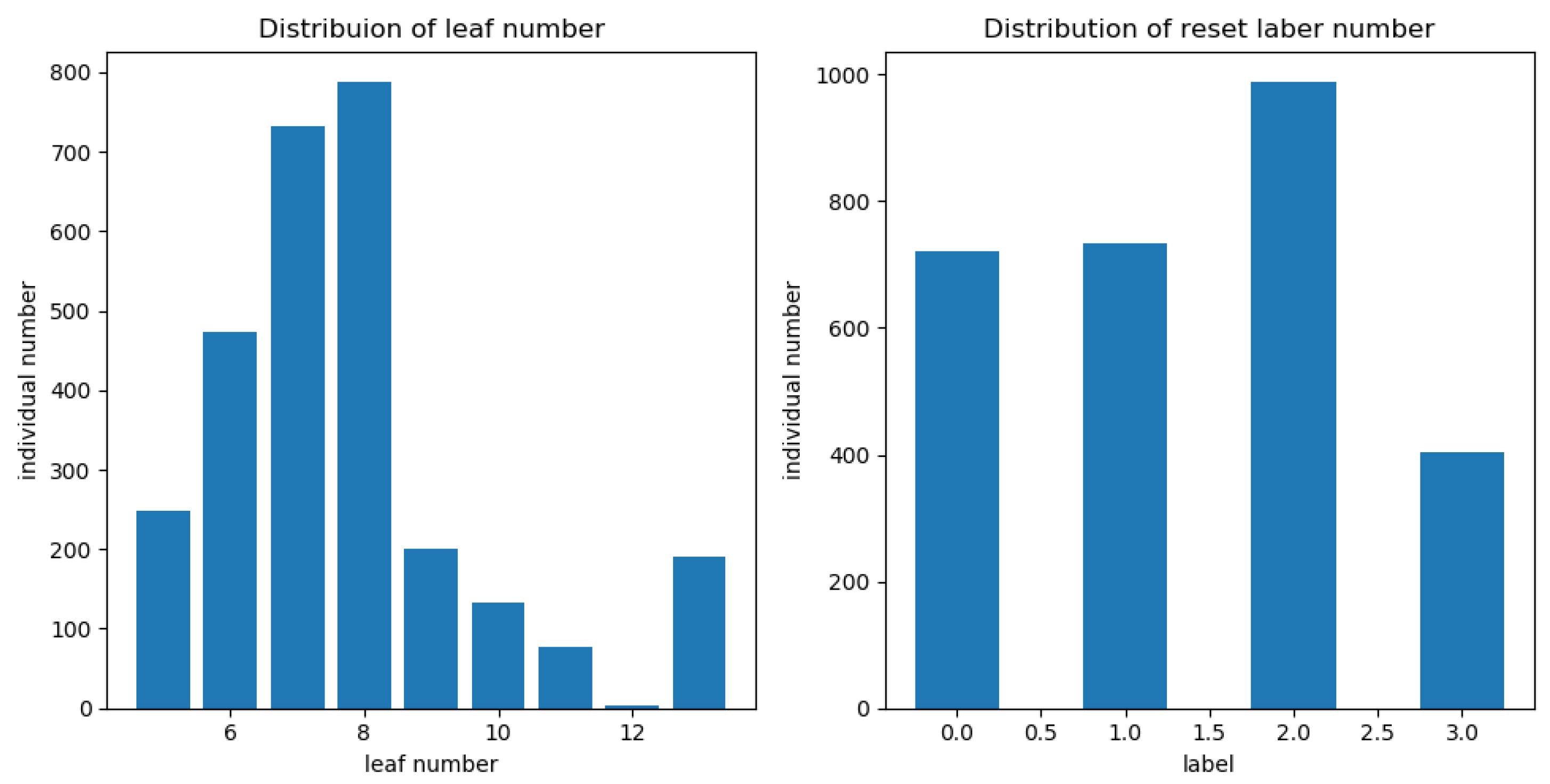
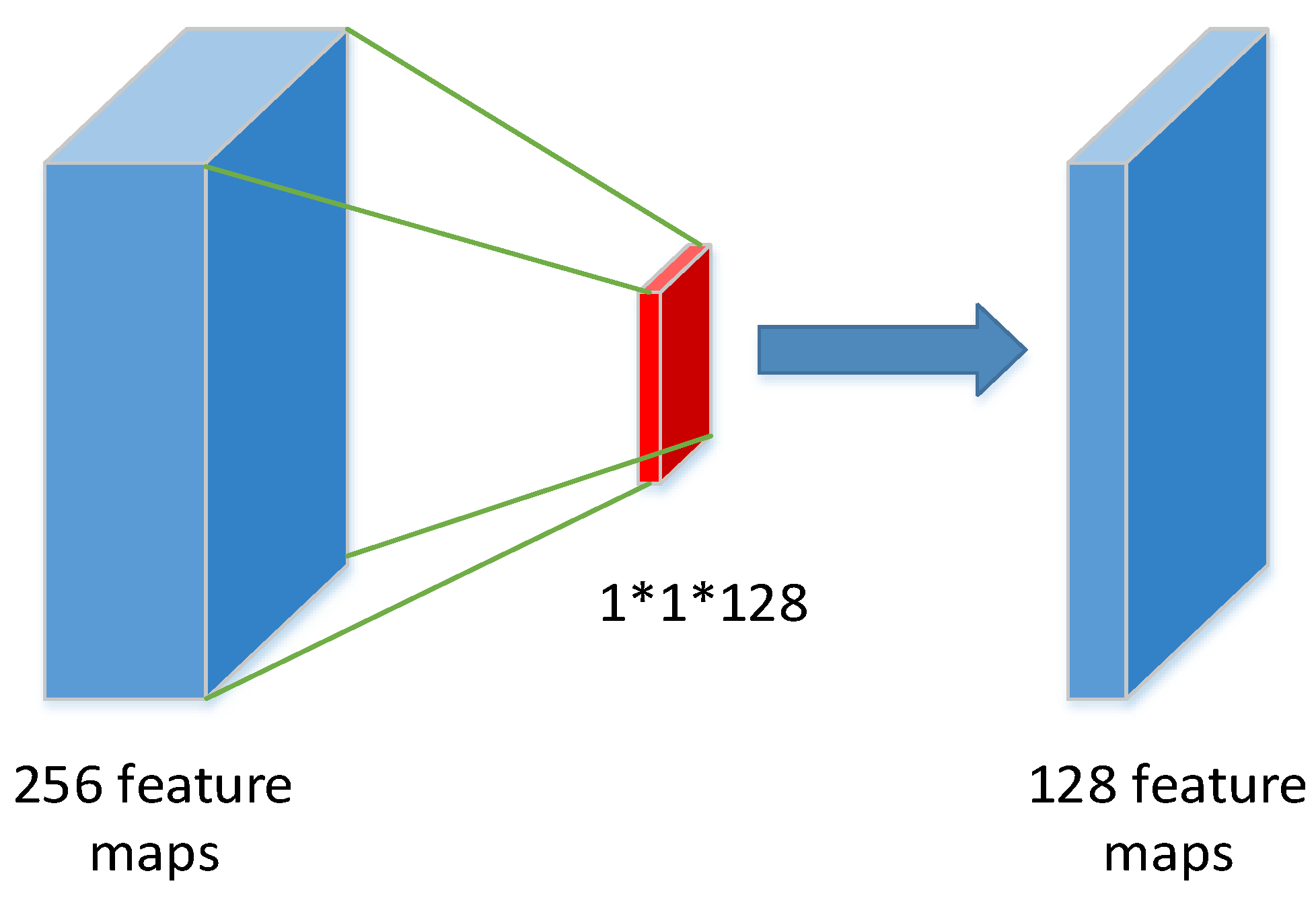
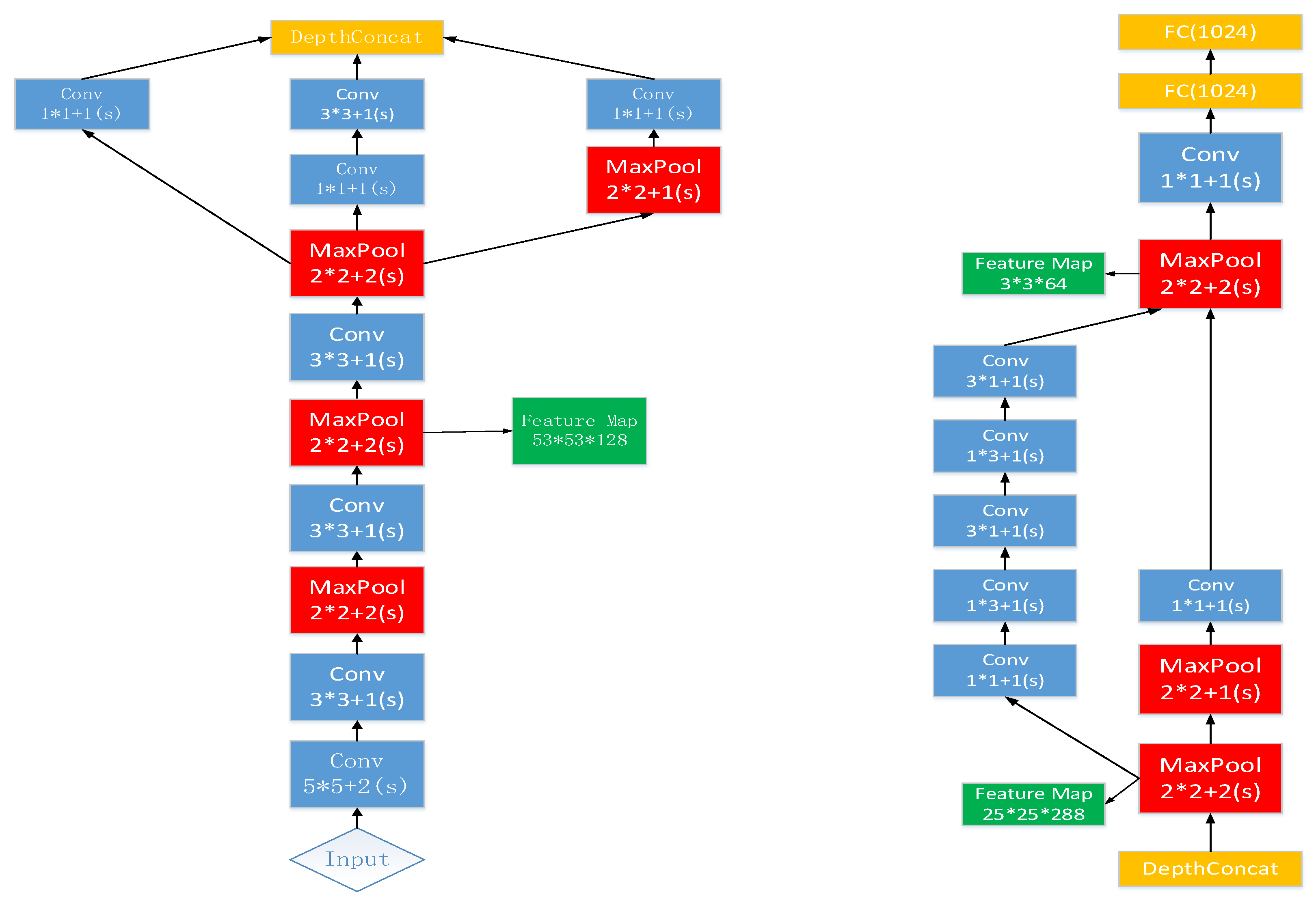
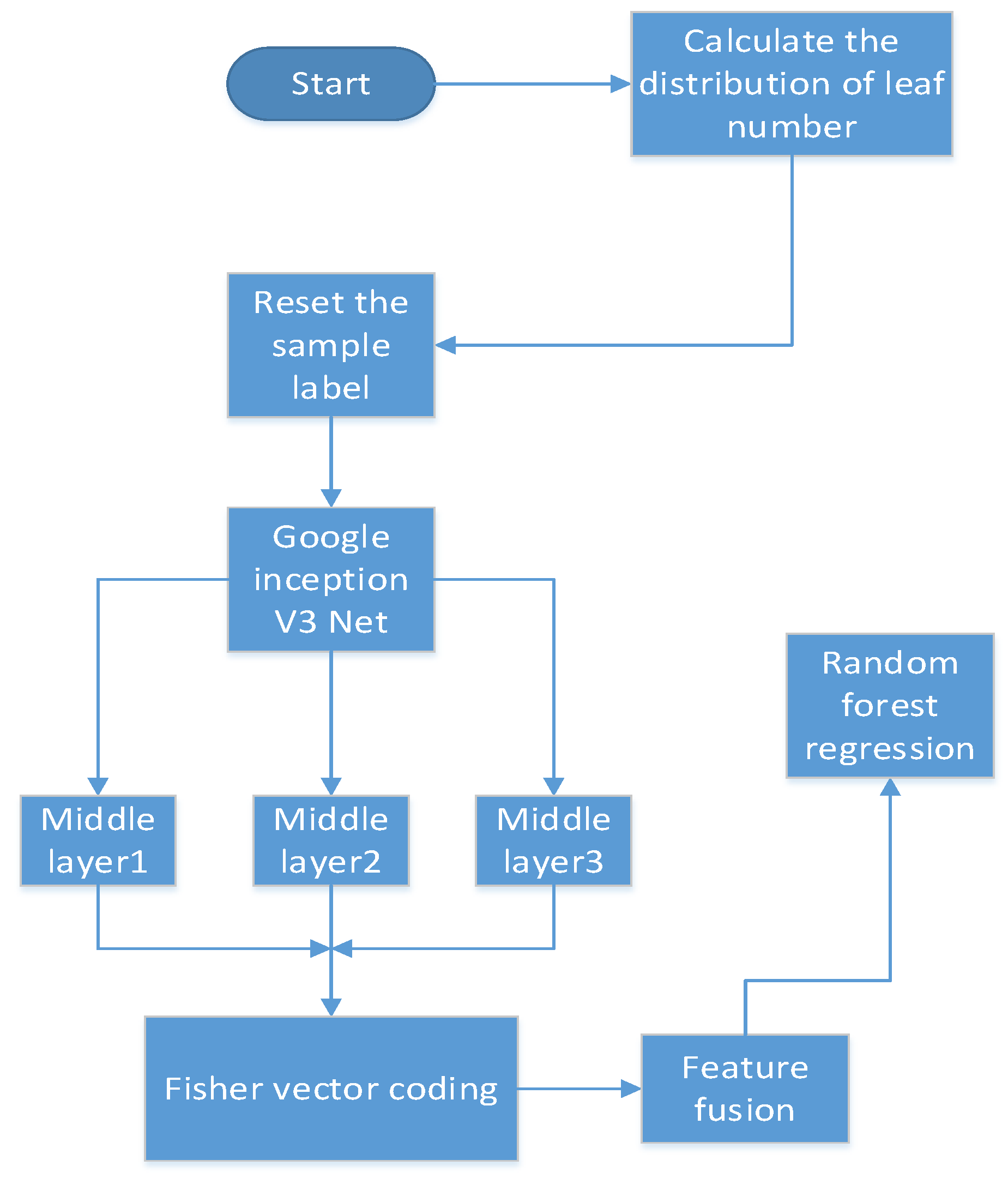
Compared to the existing work, we used the inception structure from GoogLeNet. The multi-scale convolution kernel was used in one layer instead of inputting multi-scale images, and information loss may be caused by compressing the original image when generating multi-scale images. Before feeding the features into CNN, we divided the number of leaves into different ranges and reset the label of each image sample. In fact, during training, the CNN regresses the range of leaf numbers. Extracting feature maps from each layer is computationally intensive, therefore, we obtain feature maps from three layers. In the feature extraction layer, the number of convolution kernels is reduced, which plays a role in compressing the feature map. These feature maps of the three layers are encoded by FV as fixed length feature vectors, which deduces dimensionality. Moreover, the FV can count the frequency of visual words in the feature maps and count the difference between visual dictionaries with local features.
2. Related Work
It has been proved that second-order statistics significantly improves classification performance [
17]. Some methods of CNN architectures that combine second-order statistics or coding method have been proposed. Symmetric positive definite matrix network (SPDNet) was proposed in [
18]. Referenced by the structure of the CNN, it is designed with bilinear mapping layers and eigenvalue layers, instead of convolution layers and rectified linear units. In [
19], the authors proposed a hybrid deep-learning architecture which allows to encode CNN features with log-Euclidean Fisher Vector (LE FV).
The leaf counting methods used in recent studies are mainly of two types: counting via object segmentation and direct counting via nonlinear regression model. Counting via object segmentation. This method involves segmenting the foreground and background points of the image and filtering the background before counting. Especially, the end-to-end instance segmentation method [
20] combined with long short-term memory [
21] segments one leaf at one time. In [
22], the authors used a segmented image mask to generate a plant skeleton and then extracted some skeleton features such as skeleton length, convex hull circularity and the number of skeleton branch points. These features were used to regress the leaf count. A 3D color histogram has always been used to segment plants, and therefore, threshold was selected from the color histogram to segment the plant [
23]. Then the segmented plant generates a distance map with highlighted peaks, which serve as leaf center points. The number of center points is the number of leaves. Influenced by illumination and the image shooting angle, some background points may remain in the segmented images. These noise points have a high probability to affect the leaf counting result.
Direct count. Here segmentation is not need, and the original image is directly used to count. In previous research, the architecture of the Resnet50 model has been modified. In [
24], the modified network took as input an RGB(red, green, blue three channel) image of a rosette plant and outputted a leaf count prediction. Aich and Stavness [
25] used VGGNet architecture to regress the number of leaves. The input data had four channels (segmentation+RGB; leaf counting competition offered segmented samples). There are many other similar approaches, which have different structures of the selected network. The advantage of these methods is that the image segmentation step is omitted and the original image information can be used directly to regress the leaf numbers. No additional noise will be introduced.
3. Materials
This experiment has been done in a laboratory environment. The samples selected in this study were Zhengdan No.958 grown in a pot under an in-house condition. Zhengdan which is the most popular cultivated in China and growing in Henan, Hebei and Shandong province. The average plant height is 240 cm, and the average panicle height is 100 cm. The pot’s height we selected in this study is 0.5 m and the upper diameter is 0.4 m, the bottom diameter is 0.3 m. The soil type is medium loam.
In this research, different water content degree has been set. In a field environment, maize is likely to be drought because of the water shortage. The purpose of this research is to detect the number of leaves in different environments. Therefore, setting experimental samples of different water content degree can better simulate the natural environment and show the effect of different water content degree on the number of leaves.
Table 1 shows the moisture control.
The growth stage of the maize selected in this paper was V8 (eight visible leaves) ~ VT (the last spike was visible). According to some research, the water supply of maize in two weeks before and after the pollination period will determine the final yield [
26]. Therefore, it is more meaningful to study and detect the phenotype of maize in this period.
Table 2 shows the growth stage and description.
Figure 1 shows some example images of maize at different growing stage. The selected image collection equipment was Canon Eos 700D. This camera has 18 million effective pixels. The actual collection of the maize image samples had a resolution of 5184 × 3456. However, the original picture resolution was compressed to 441 × 441 in the calculation to improve the computational efficiency. The camera angle and focal length were adjusted with the growth of the maize. A picture was taken every 5 minutes at 5:30 am to 18:30 pm.
5. Results and Discussion
In this section, we evaluate the proposed leaf counting approach on an image data set of maize plant. First, we discuss the experimental settings and parameters. Next, we present the details of the training and testing samples. Then we show a comparison between the experimental results and existing methods to prove the effectiveness of our algorithm. In this paper, we assume that samples with similar numbers of leaves have similar leaf features. There was a high similarity of features learned by CNN for samples with the same label. When only the feature map of the last layer in the network was used, the feature discrimination of samples in the same label was small, and this is not conducive to the subsequent leaf number fitting (when setting labels for samples, samples with the same label may not have the same leaf number). Therefore, we extracted the feature maps from the middle layer in the same way, and the difference of these feature maps was greater than that of the last layer. Meanwhile, these features are a good complement to detailed features, which also explains why we used multi-scale features.
5.1. Implentation Details
(1). The framework of Python+Tensorflow have been used to build the network. Some training parameters are shown in
Table 4.
(2). Our CNN-net trained and tested under Windows 10 64-operation system on Intel Core i7-8700 at 3.2GHZ with 32-GB RAM. The GPU is GTX 1080Ti.
(3). Finally, we use random forests to classify the number of maize leaves, the number of trees in the forest was set 70 and the max features was set 44(sqrt(features)).
5.2. Image Data
In 2.4, we present the method to get image data samples. The samples of the four water levels were 701,644,851 and 649 in number (because of the problem of shooting angle and illumination, we removed some poor-quality samples). The total number of samples was 2845, of which 80% were training samples and 20% are testing samples. The numbers of final training samples and testing samples were 2276 and 569.
5.3. Experimental Results and Comparison with Other Methods
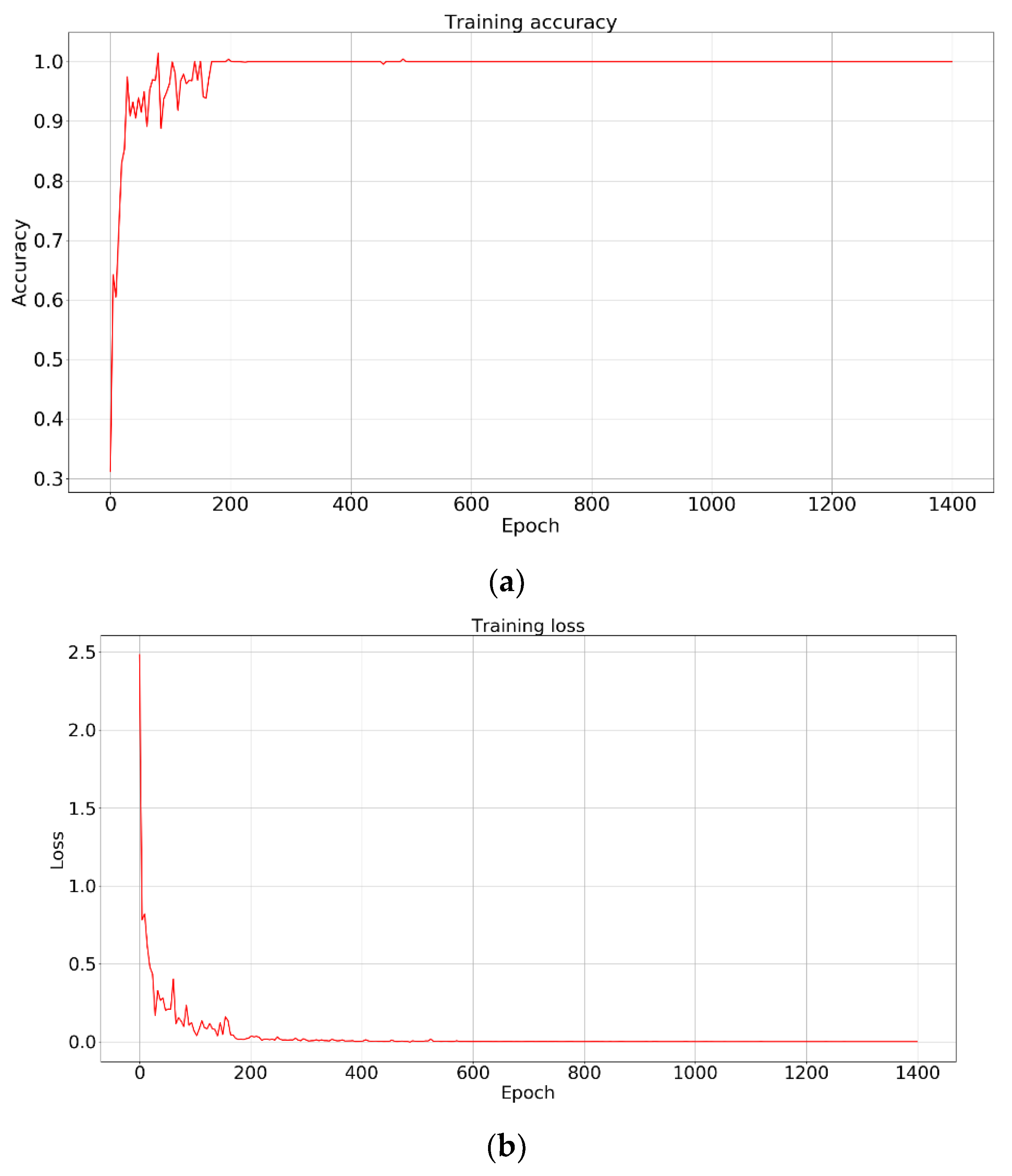
The training accuracy and training loss are shown in
Figure 7. With an increase in training epochs, the model gradually converged. As we can see from
Figure 7, with an increase in training epochs, the training loss converged quickly and training accuracy was close to 100% in 200 epochs. This indicates that the features learned by CNN are suitable for classification. It is reasonable to assign the same label for maize samples with similar leaf numbers. By training the classification of maize samples, the model can roughly determine the range of leaf numbers of a single maize sample. The label assigned to the sample represents the range of the leaf numbers. Therefore, the CNN model extracts feature by learning to predict the range of plant leaf numbers. As we know, a high accuracy rate is very helpful for further encoding feature maps. The correct rate directly reflects the feature extraction ability of our network. Test samples were not reserved because our aim was not to classify them. Finally, to avoid over-fitting, the weight model obtained by the 200th iteration was saved to extract the feature from the middle layer.
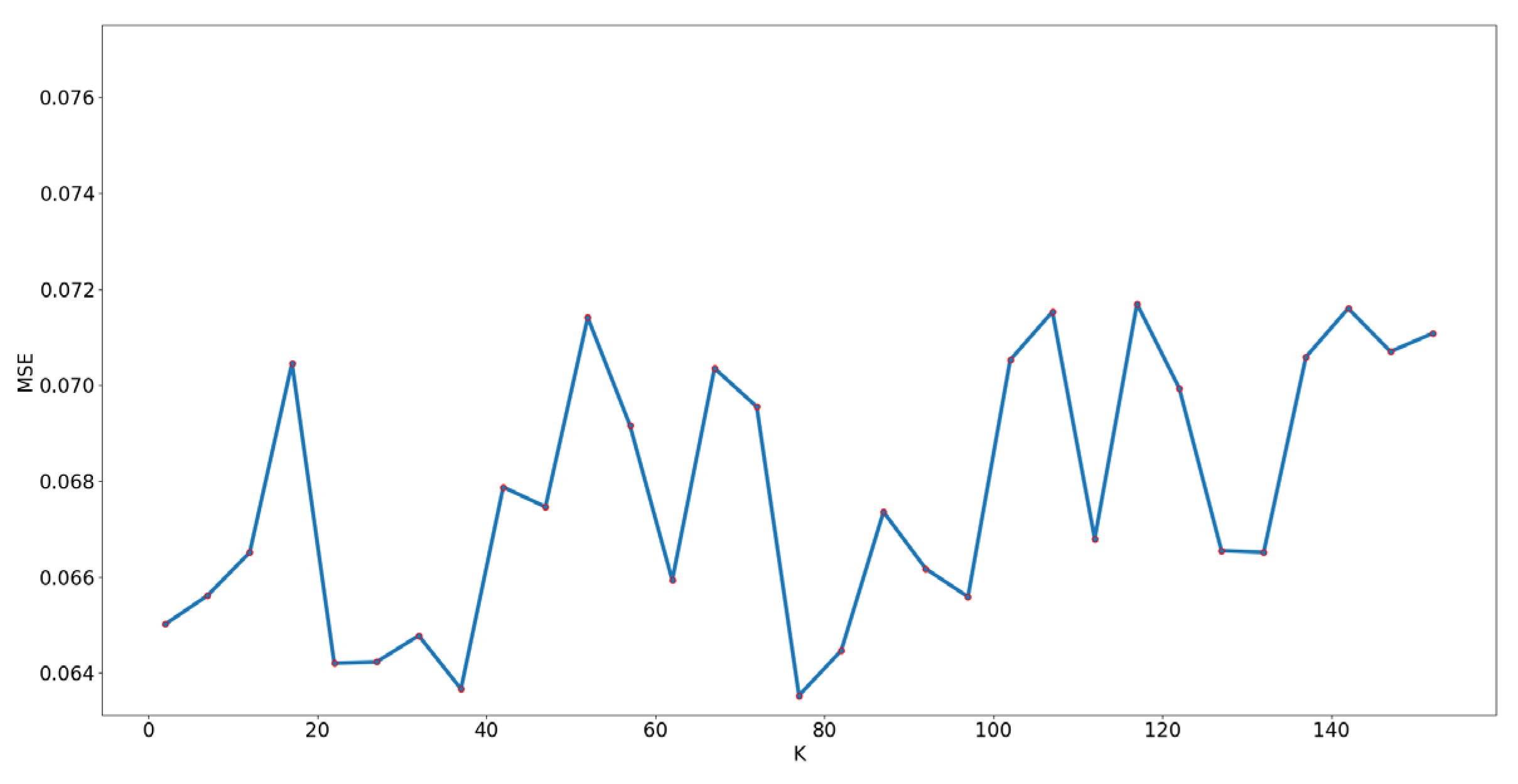
In FV coding, there exists an important parameter, parameter k (the number of Gaussian distribution), which should be assigned during the whole process. To select a reasonable parameter k, four water level samples from different growth stage were selected. The number of this part is 650. The result can be seen in
Figure 8. We can see k = 77 has the best performance.
The results of the comparison are shown in
Table 5, “CountDiff” refers to the mean and standard deviation of the difference in count averaged over images. “AbsCountDiff” is the absolute of “CountDiff.” “MSE” is the abbreviation for mean-square error [
25].
Table 5 compares the results of our algorithm and the method of directly fitting with the deep neural network. Method (1) [
32] and method (2) [
33] shows that the method of directly fitting the leaf number with depth network has a high mean-square error. In the experiment, we found that the result of these method is close to the mean of the training samples, especially for samples with a large number of leaves. It can be seen from (3) and (4) that, the deep neural network is more powerful for sample feature extraction than that traditional local feature extraction algorithms, such as SIFT. In method (4), there is a large gap between the training result and test result, the over-fitting is serious. These imply that extracting multi-scale features from CNN combined with the traditional machine learning is more advantageous than the single CNN method for estimating the number of maize leaves.
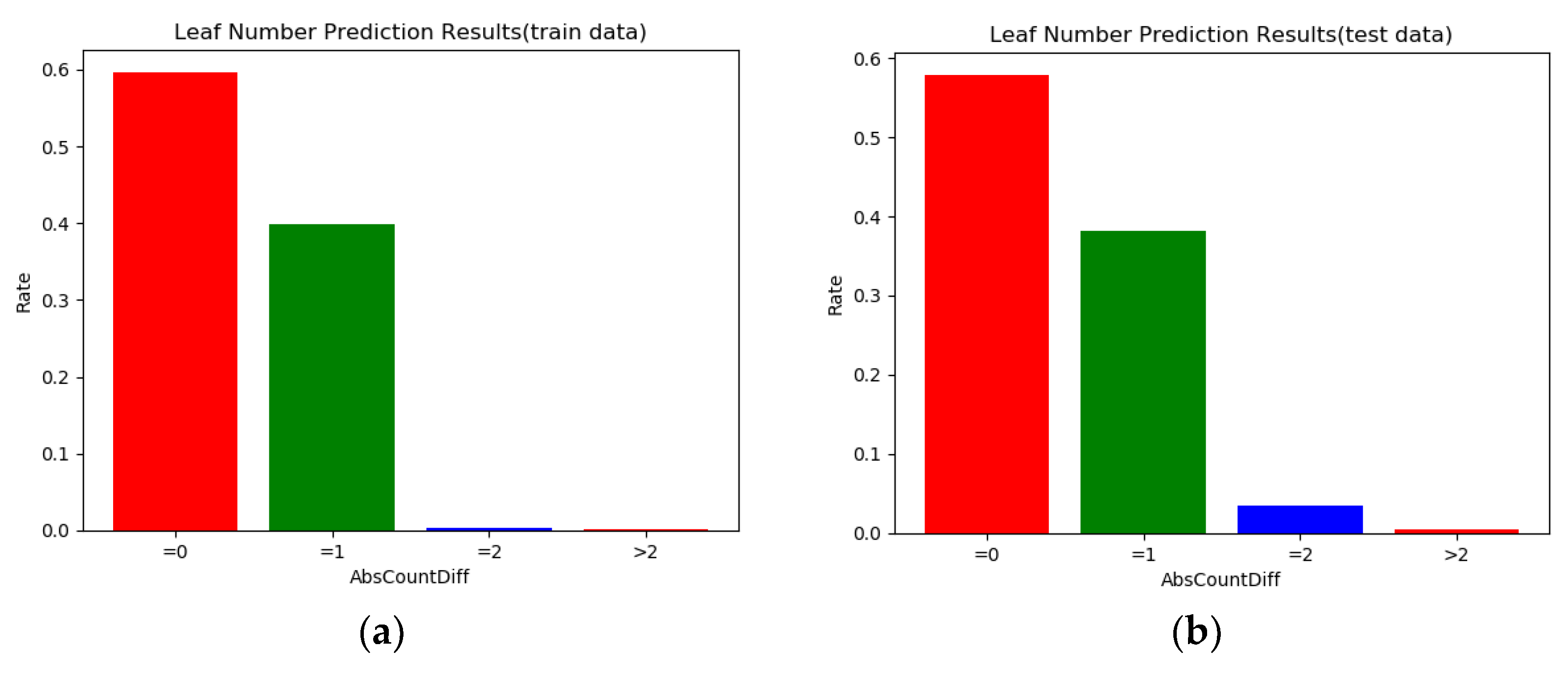
From
Figure 9, we can see that most of the prediction errors are within one leaf. Comparing the (a) and (b), the range of error distribution of the training set and test set was consistent, and there was no large fluctuation in the distribution, which proves that our model is stable and has practical application value.
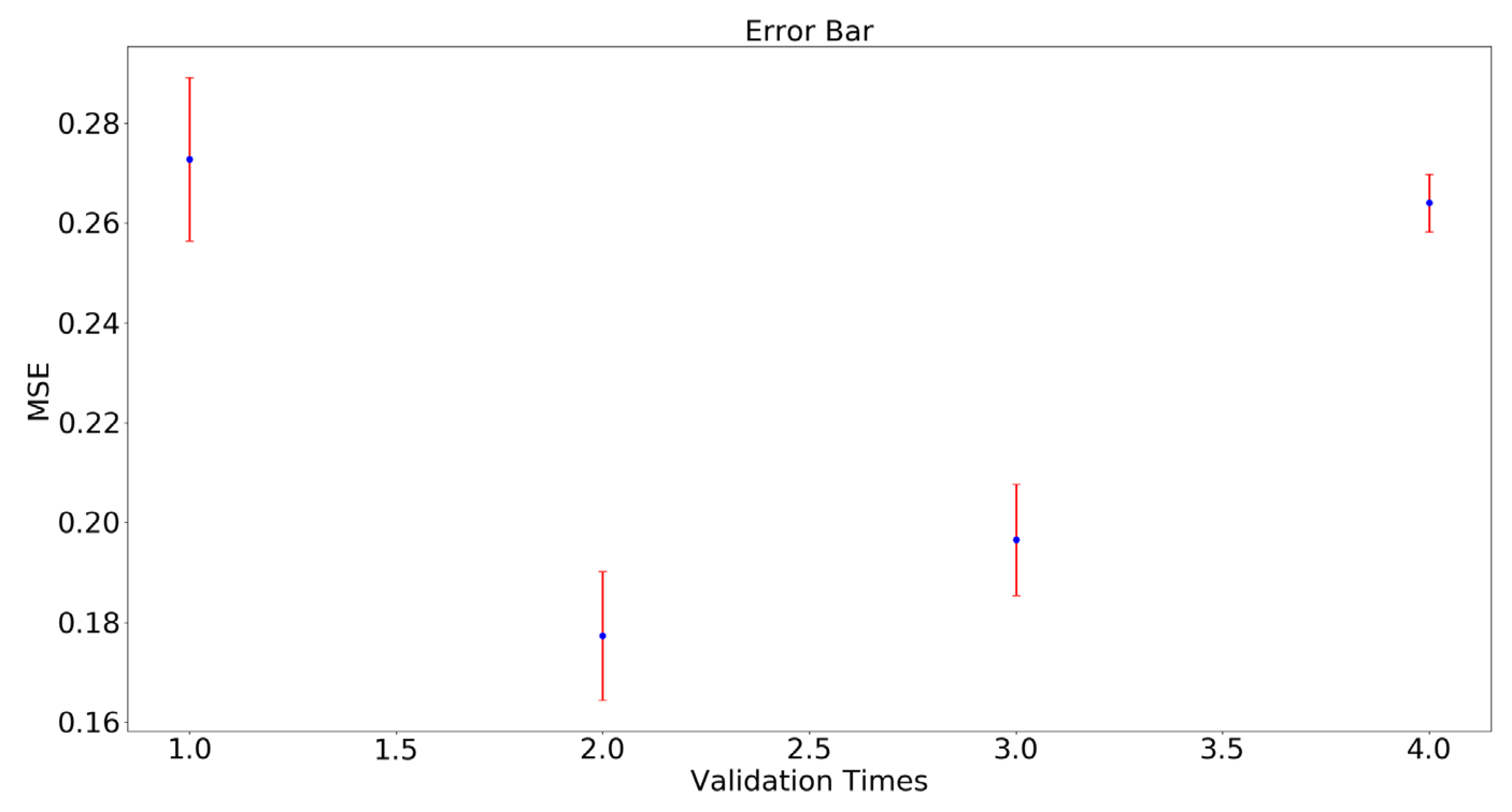
To verify the robustness of the proposed approach, a cross-validation experiment was designed. One type sample was reserved as the validation set and the other three samples were the training set. Therefore, each of the four water level samples was treated as a validation set. Then each group of experiments was repeated five times. The error bar of MSE was shown in
Figure 10.
As can be seen from
Figure 10, our model performed well for different water level samples, and performance was worst in the first validation compared to other times. In the first validation, samples 1, 2 and 3 were the training set and sample 4 was the validation set. Sample 4 was the most severely affected by drought stress and its leaves were fewer than those of other samples in the same period. This indicates that the feature vectors of sample 4 were quite different from those of the other three type samples.
5.4. Misclassified Image Analysis
In this study, some incorrect regress samples have been shown in
Figure 11a–f are the samples of incorrect leaf count at different maize growth stages. The counting error of (a)–(d) are within 2 leaves. As can be seen from the samples, influenced by illumination there are some leaves in (a)–(d) are hard to distinguish with white pots. These leaves are located in the lower part of the maize. To (c) and (d), some leaves are withered because of the water shortage. These factors increase the difficulty of leaf counting and lead to counting errors. The counting error of (e) and (f) are more than 2 leaves. These samples have more leaves than (a)–(d) and the leaves are shaded from each other, therefore the counting error increases dramatically.
5.5. The Relationship Between Maize Leaf Number and Water Content
The number of leaves can reflect the water content of a maize plant.
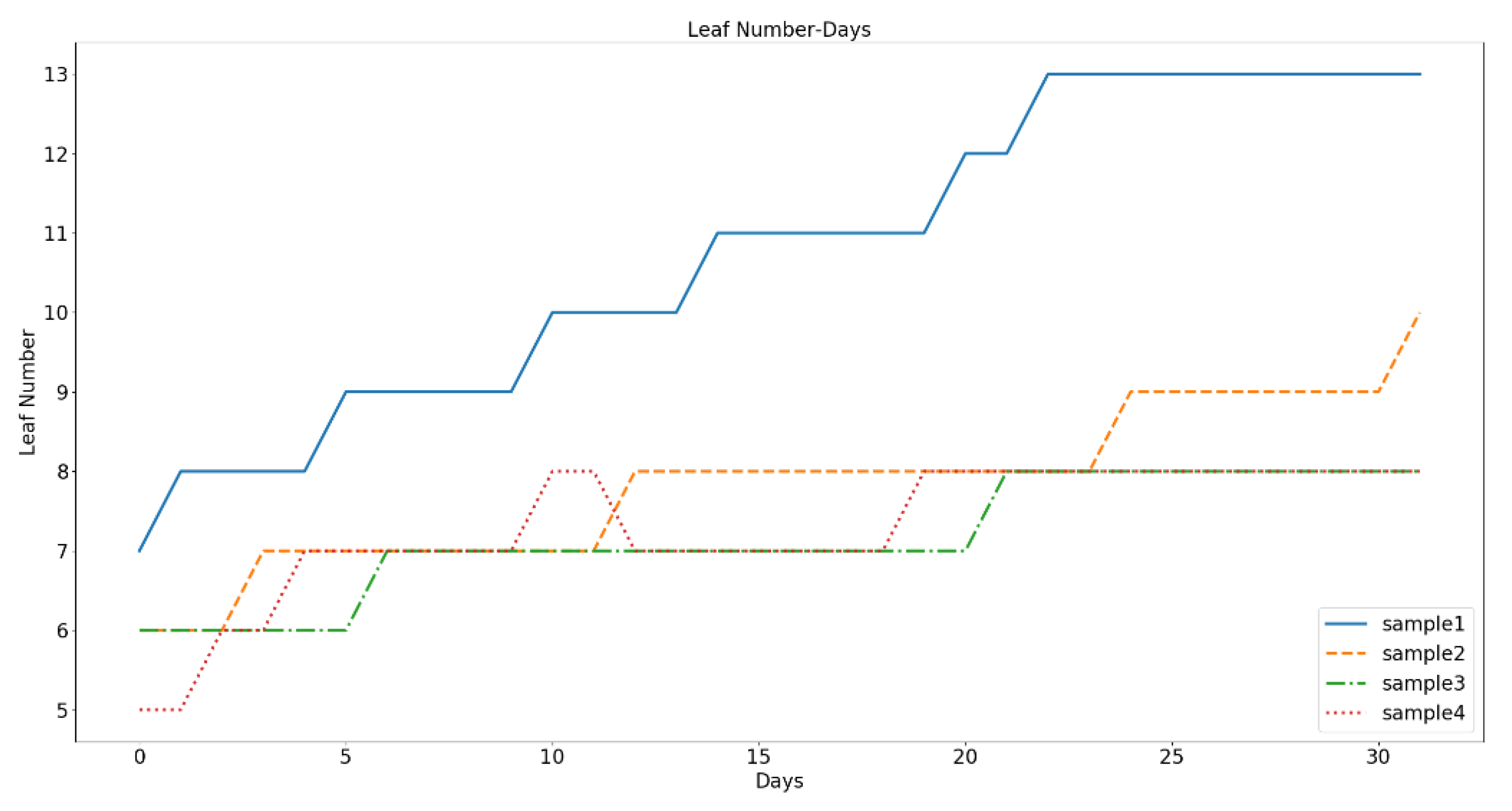
Figure 12 shows a line chart of the changes in the number of leaves over time for the four samples. The observation period was 32 days. The blue line represents the sample with suitable moisture. We can see that the number of leaves increase stepwise with time, and there is a clear demarcation line between the other three water-deficient samples. The green line represents moderate drought. Although it overlaps with the other two water-deficient samples, the total leaf number also rises stepwise; the rising rate is much lower than that of the suitable moisture sample and higher than that of the other two water-deficient samples. The discrimination between samples 3 and 4 was small, the number of leaves in sample 4 first increased and then decreased with time. By observing the actual image of the sample, it was found that the leaves in Sample 4 dropped seriously due to dry up. Sample 3 also exhibited the same condition. Our algorithm does not detect the drooping leaves, because their color is close to that of the soil. Therefore, the distributions of samples 3 and 4 are similar in
Figure 12.
6. Conclusions
In this paper, a deep learning approach combined with the traditional machine learning is been proposed. The CNN is responsible for extracting multi-scale features from different layers. Multi-scale features extraction can compensate for the loss of features caused by pooling layers. A FV maps the multi-scale features to a higher dimensional space. This can enhance the expression ability of the CNN and make the model perform well. Our method does not require segmentation, and new noise regions, which are associated with the error segmentation algorithm, are not introduced. The experimental results demonstrate that this method effectively counts maize leaves. However, for the samples with abnormal illumination and leaf occlusion, there are still large errors in counting. This indicates that some work still needs to be done.
In future extensions of this paper, we plan to enrich our data set by collecting more images of new maize species. Moreover, the current related studies are mainly conducted in a laboratory environment; future works should focus on field environment. In the preprocessing, the sample labels for different leaf numbers need to be manually marked to ensure the partitioned samples have similar morphological features and a uniform distribution. However, manual operation is inconvenient, and the automatic partitioning method needs be developed. Furthermore, to avoid redundancy of information, only three-layer feature maps were extracted according to the CNN structure. We cannot guarantee that these three-level feature maps are the best combination. In subsequent works, we will continue to study how to select the optimal combination.