IMGT® and 30 Years of Immunoinformatics Insight in Antibody V and C Domain Structure and Function
Abstract
1. Introduction
2. IMGT Unique Numbering for V Domain
2.1. V Domain Strands and Loops
2.2. IMGT Gaps and Additional Positions
3. IMGT Unique Numbering for C Domain
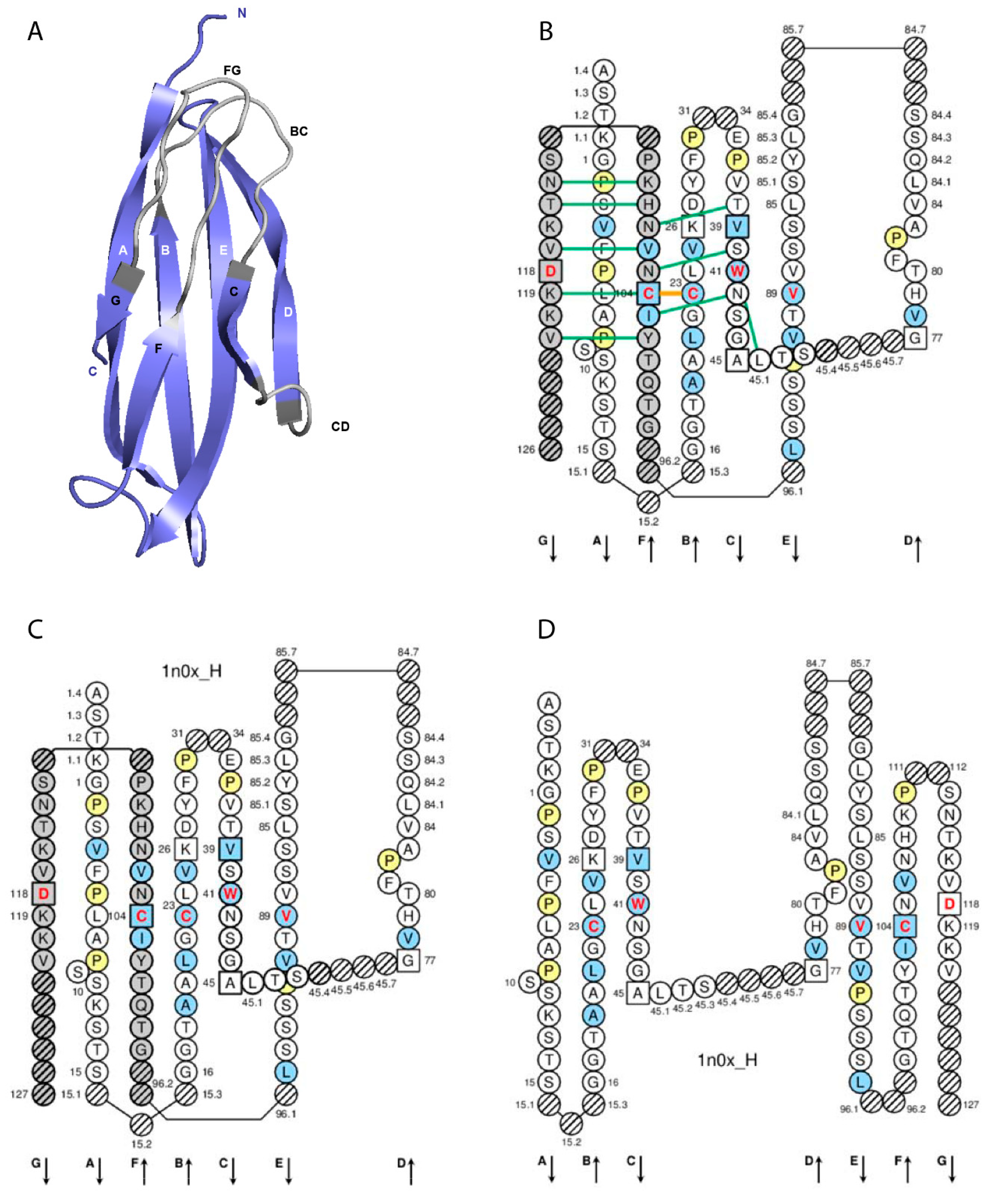
3.1. C Domain Strands, Loops, and Turns
3.2. C Domain and V Domain Comparison
3.3. IMGT Gaps and Additional Positions
4. IMGT® V and C Domain Insight for Antibody Humanization and Engineering
4.1. Antibody Humanization
4.1.1. CDR-IMGT Delimitation for Grafting
4.1.2. Amino Acid Interactions between FR-IMGT and CDR-IMGT
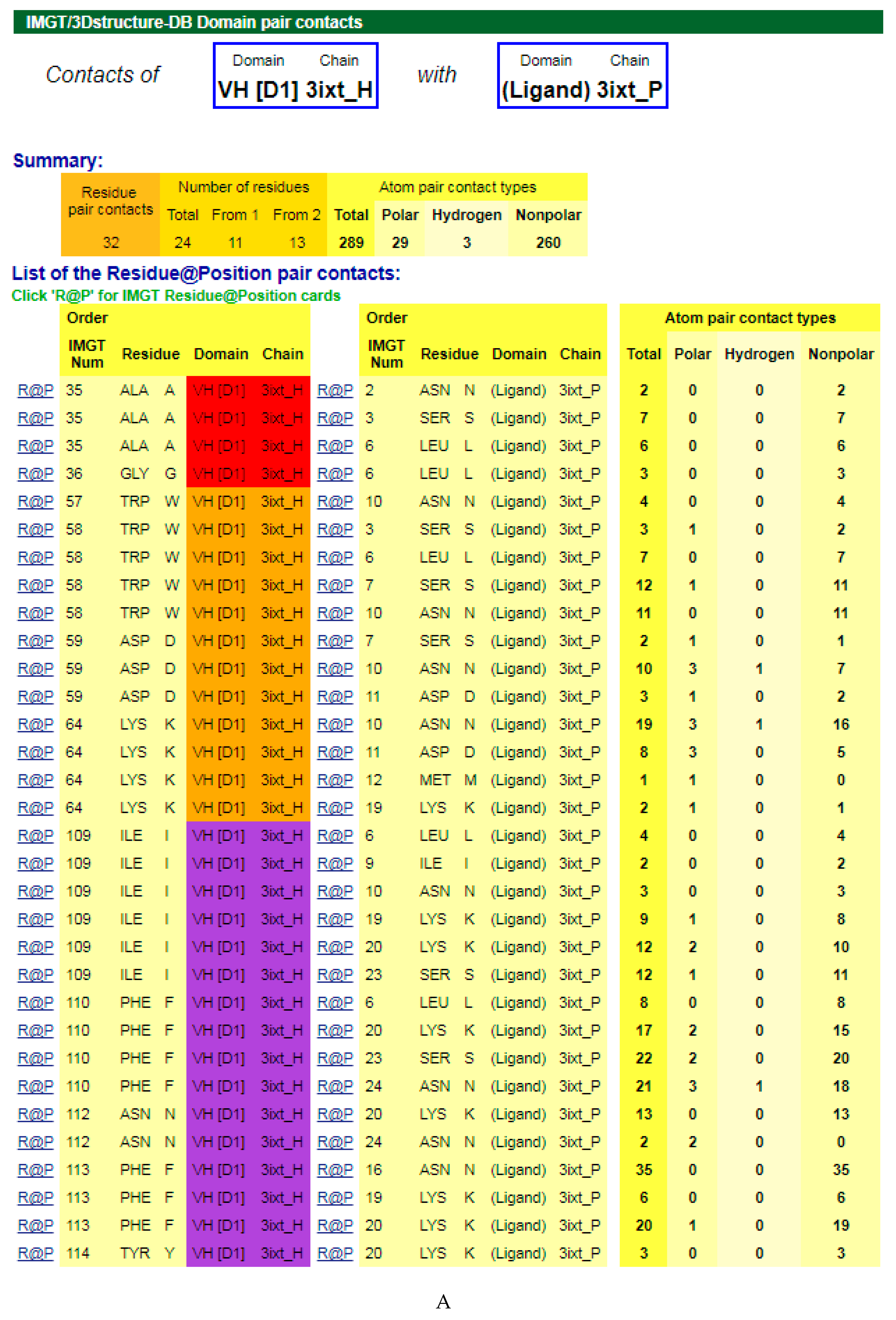
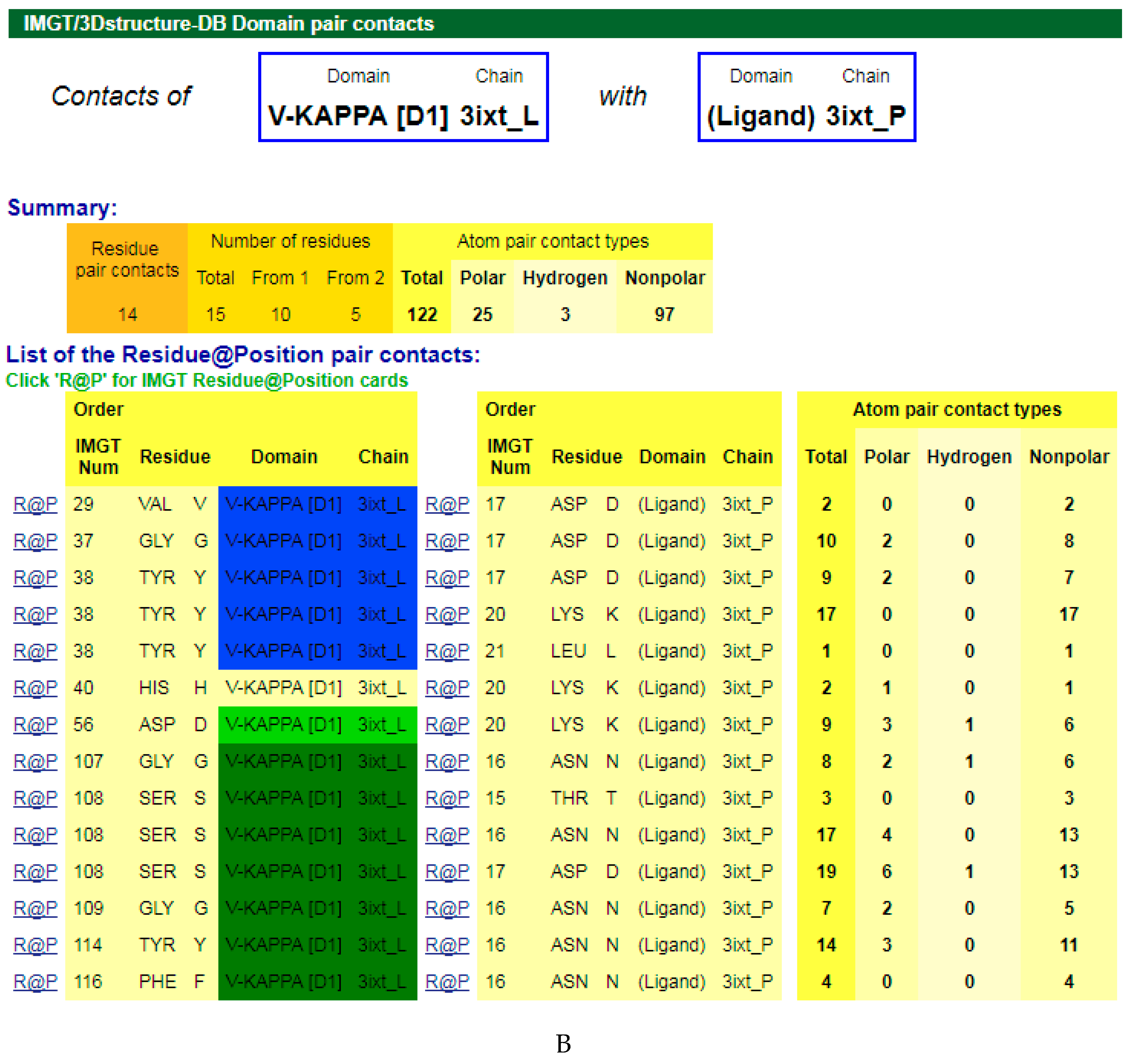
4.1.3. V-DOMAIN Contact Analysis and Paratope
4.1.4. Potential Immunogenicity and Physicochemical Properties
4.1.5. V-DOMAIN CDR-IMGT Lengths and Canonical Structures
4.2. IGHG1 Alleles and G1m Allotypes
4.3. Only-Heavy-Chain Antibodies
4.3.1. Dromedary IgG2 and IgG3 Only-Heavy-Chain Antibodies
4.3.2. Human Heavy Chain Diseases (HCD)
4.3.3. Nurse Shark IgN
5. IGHG CH Properties and Antibody Engineering
5.1. N-Linked Glycosylation Site CH2 N84.4
5.2. Interface Ball-and-Socket-Like Joints
5.3. Knobs-Into-Holes CH3 for the Obtaining of Bispecific Antibodies
5.4. IGHG Engineered Variants and Effector Properties
6. Conclusions
Availability and Citation
Funding
Acknowledgments
Conflicts of Interest
References
- Lefranc, M.-P.; Lefranc, G. The Immunoglobulin FactsBook; Academic Press: London, UK, 2001. [Google Scholar]
- Lefranc, M.-P.; Lefranc, G. The T Cell Receptor FactsBook; Academic Press: London, UK, 2001. [Google Scholar]
- Lefranc, M.-P. Immunoglobulin and T Cell Receptor Genes: IMGT® and the Birth and Rise of Immunoinformatics. Front. Immunol. 2014, 5, 22. [Google Scholar] [CrossRef] [PubMed]
- Lefranc, M.-P. Unique database numberings system for immunogenetic analysis. Immunol. Today 1997, 18, 509. [Google Scholar] [CrossRef]
- Lefranc, M.-P. The IMGT unique numbering for Immunoglobulins, T cell receptors and Ig-like domains. Immunologist 1999, 7, 132–136. [Google Scholar]
- Lefranc, M.-P.; Pommie, C.; Ruiz, M.; Giudicelli, V.; Foulquier, E.; Truong, L.; Thouvenin-Contet, V.; Lefranc, G. IMGT unique numbering for immunoglobulin and T cell receptor variable domains and Ig superfamily V-like domains. Dev. Comp. Immunol. 2003, 27, 55–77. [Google Scholar] [CrossRef]
- Lefranc, M.-P.; Pommie, C.; Kaas, Q.; Duprat, E.; Bosc, N.; Guiraudou, D.; Jean, C.; Ruiz, M.; Da Piedade, I.; Rouard, M.; et al. IMGT unique numbering for immunoglobulin and T cell receptor constant domains and Ig superfamily C-like domains. Dev. Comp. Immunol. 2005, 29, 185–203. [Google Scholar] [CrossRef]
- Lefranc, M.-P.; Duprat, E.; Kaas, Q.; Tranne, M.; Thiriot, A.; Lefranc, G. IMGT unique numbering for MHC groove G-DOMAIN and MHC superfamily (MhcSF) G-LIKE-DOMAIN. Dev. Comp. Immunol. 2005, 29, 917–938. [Google Scholar]
- Kaas, Q.; Ruiz, M.; Lefranc, M.-P. IMGT/3Dstructure-DB and IMGT/StructuralQuery, a database and a tool for immunoglobulin, T cell receptor and MHC structural data. Acids Res. 2004, 32, D208–D210. [Google Scholar] [CrossRef] [PubMed]
- Ehrenmann, F.; Kaas, Q.; Lefranc, M.-P. IMGT/3Dstructure-DB and IMGT/DomainGapAlign: A database and a tool for immunoglobulins or antibodies, T cell receptors, MHC, IgSF and MhcSF. Nucl. Acids Res. 2010, 38, D301–D307. [Google Scholar] [CrossRef] [PubMed]
- Ehrenmann, F.; Lefranc, M.-P. IMGT/3Dstructure-DB: Querying the IMGT Database for 3D Structures in Immunology and Immunoinformatics (IG or Antibodies, TR, MH, RPI, and FPIA). Cold Spring Harb. Protoc. 2011, 2011, 750–761. [Google Scholar] [CrossRef] [PubMed]
- Ehrenmann, F.; Lefranc, M.-P. IMGT/DomainGapAlign: IMGT Standardized Analysis of Amino Acid Sequences of Variable, Constant, and Groove Domains (IG, TR, MH, IgSF, MhSF). Cold Spring Harb. Protoc. 2011, 2011, 737–749. [Google Scholar] [CrossRef]
- Ehrenmann, F.; Lefranc, M.-P. IMGT/DomainGapAlign: The IMGT® tool for the analysis of IG, TR, MHC, IgSF and MhSF domain amino acid polymorphism. In Immunogenetics; Christiansen, F., Tait, B., Eds.; Humana Press: Totowa, NJ, USA, 2012; pp. 605–633. [Google Scholar]
- Lefranc, M.-P. IMGT unique numbering for the Variable (V), Constant (C), and Groove (G) domains of IG, TR, MH, IgSF, and MhSF. Cold Spring Harb. Protoc. 2011, 6, 633–642. [Google Scholar] [CrossRef]
- Pommie, C.; Levadoux, S.; Sabatier, R.; Lefranc, G.; Lefranc, M.-P. IMGT standardized criteria for statistical analysis of immunoglobulin V-REGION amino acid properties. J. Mol. Recognit. 2004, 17, 17–32. [Google Scholar] [CrossRef]
- Riechmann, L.; Clark, M.; Waldmann, H.; Winter, G. Reshaping human antibodies for therapy. Nature 1988, 332, 323–327. [Google Scholar] [CrossRef]
- Kabat, E.A.; Wu, T.T.; Perry, H.M.; Gottesman, K.S.; Foeller, C. Sequences of Proteins of Immunological Interest; U.S. Department of Health and Human Services (USDHHS), National Institute of Health NIH Publication: Washington, DC, USA, 1991.
- Lefranc, M.-P.; Lefranc, G. Human Gm, Km, and Am Allotypes and Their Molecular Characterization: A Remarkable Demonstration of Polymorphism. Methods Mol. Biol. 2012, 882, 635–680. [Google Scholar]
- Jefferis, R.; Lefranc, M.-P. Human immunoglobulin allotypes: Possible implications for immunogenicity. MAbs 2009, 1, 332–338. [Google Scholar] [CrossRef]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hammers, C.; Songa, E.B.; Bendahman, N.; Hammers, R. Naturally occurring antibodies devoid of light chains. Nature 1993, 363, 446–448. [Google Scholar] [CrossRef]
- Nguyen, V.K.; Hamers, R.; Wyns, L.; Muyldermans, S.; Niu, J.; Miao, R.; Huang, S.; Chang, J.; Davis-Dusenbery, B.N.; Kashima, R.; et al. Camel heavy-chain antibodies: Diverse germline VHH and specific mechanisms enlarge the antigen-binding repertoire. EMBO J. 2000, 19, 921–930. [Google Scholar] [CrossRef]
- Nguyen, V.K.; Hamers, R.; Wyns, L.; Muyldermans, S. Loss of splice consensus signal is responsible for the removal of the entire CH1 domain of the functional camel IgG2a heavy chain antibodies. Mol. Immunol. 1999, 36, 515–524. [Google Scholar] [CrossRef]
- Lefranc, M.-P.; Lefranc, G. The constant region genes of the immunoglobulin heavy chains. Mol. Genet. 1988, 7, 39–45. [Google Scholar]
- Greenberg, A.S.; Avila, D.; Hughes, M.; Hughes, A.; McKinney, E.C.; Flajnik, M.F. A new antigen receptor gene family that undergoes rearrangement and extensive somatic diversification in sharks. Nature 1995, 374, 168–173. [Google Scholar] [CrossRef]
- Lefranc, M.-P. How to Use IMGT® for Therapeutic Antibody Engineering. Handb. Ther. Antib. 2014, 1, 229–264. [Google Scholar]
- Lefranc, M.-P. IMGT® immunoglobulin repertoire analysis and antibody humanization. In Molecular Biology of B Cells, 2nd ed.; Alt, F.W., Honjo, T., Radbruch, A., Reth, M., Eds.; Academic Press: London, UK, 2014; Chapter 26; pp. 481–514. [Google Scholar]
- Shirai, H.; Prades, C.; Vita, R.; Marcatili, P.; Popovic, B.; Xu, J.; Overington, J.P.; Hirayama, K.; Soga, S.; Tsunoyama, K.; et al. Antibody informatics for drug discovery. Biochim. Biophys. BBA-Proteins Proteom. 2014, 1844, 2002–2015. [Google Scholar] [CrossRef]
- Teplyakov, A.; Zhao, Y.; Malia, T.J.; Obmolova, G.; Gilliland, G.L. IgG2 Fc structure and the dynamic features of the IgG CH2–CH3 interface. Mol. Immunol. 2013, 56, 131–139. [Google Scholar] [CrossRef]
- Lesk, A.M.; Chothia, C. Elbow motion in the immunoglobulins involves a molecular ball-and-socket joint. Nature 1988, 335, 188–190. [Google Scholar] [CrossRef]
- Ridgway, J.B.; Presta, L.G.; Carter, P. ‘Knobs-into-holes’ engineering of antibody CH3 domains for heavy chain heterodimerization. Eng. Des. Sel. 1996, 9, 617–621. [Google Scholar] [CrossRef]
- Lefranc, M.-P. IMGT, the international ImMunoGeneTics information system. In Immunoinformatics: Bioinformatic Strategies for Better Understanding of Immune Function; Bock, G., Goode, J., Eds.; Novartis Foundation Symposium; John Wiley and Sons: Chichester, UK, 2003; Volume 254, p. 126, discussion 136–142, 216–222, 250–252. [Google Scholar]
- Lefranc, M.-P.; Giudicelli, V.; Ginestoux, C.; Chaume, D. IMGT, the international ImMunoGeneTics information system: The reference in immunoinformatics. Stud. Health Technol. Inform. 2003, 95, 74–79. [Google Scholar]
- Lefranc, M.-P. IMGT databases, web resources and tools for immunoglobulin and T cell receptor sequence analysis. Leukemia 2003, 17, 260–266. [Google Scholar] [CrossRef]
- Lefranc, M.-P. IMGT, the international ImMunoGenetics information system®. In Antibody Engineering Methods and Protocols, 2nd ed.; Humana Press: Totowa, NJ, USA, 2004; pp. 27–49. [Google Scholar]
- Lefranc, M.-P. IMGT-ONTOLOGY and IMGT databases, tools and Web resources for immunogenetics and immunoinformatics. Mol. Immunol. 2004, 40, 647–659. [Google Scholar] [CrossRef]
- Lefranc, M.-P. IMGT, the international ImMunoGeneTics information system®: A standardized approach for immunogenetics and immunoinformatics. Immunome Res. 2005, 1, 3. [Google Scholar] [CrossRef][Green Version]
- Lefranc, M.-P. IMGT®, the International ImmunoGeneTics Information System® for Immunoinformatics. In Immunoinformatics: Predicting Immunogenicity In Silico; Flowers, D.R., Ed.; Humana Press: Totowa, NJ, USA, 2007; Volume 409, pp. 19–42. [Google Scholar]
- Lefranc, M.-P. IMGT-ONTOLOGY, IMGT® databases, tools and Web resources for Immunoinformatics. In Immunoinformatics; Schoenbach, C., Ranganathan, S., Brusic, V., Eds.; Immunomics Reviews, Series of Springer Science and Business Media LLC; Springer: New York, NY, USA, 2008; Volume 1, Chapter 1; pp. 1–18. [Google Scholar]
- Regnier, L.; Lefranc, M.-P.; Giudicelli, V.; Duroux, P. IMGT, a system and an ontology that bridge biological and computational spheres in bioinformatics. Brief. Bioinform. 2008, 9, 263–275. [Google Scholar]
- Lefranc, M.-P. IMGT®, the International ImMunoGeneTics Information System® for Immunoinformatics. Mol. Biotechnol. 2008, 40, 101–111. [Google Scholar] [CrossRef]
- Lefranc, M.-P. IMGT, the International ImMunoGeneTics Information System. Cold Spring Harb. Protoc. 2011, 2011, 595–603. [Google Scholar] [CrossRef]
- Lefranc, M.-P.; Giudicelli, V.; Duroux, P.; Jabado-Michaloud, J.; Folch, G.; Aouinti, S.; Carillon, E.; Duvergey, H.; Houles, A.; Paysan-Lafosse, T.; et al. IMGT®, the international ImMunoGeneTics information system® 25 years on. Nucleic Acids Res. 2015, 43, D413–D422. [Google Scholar] [CrossRef]
- Lefranc, M.-P. Antibody Informatics: IMGT, the International ImMunoGeneTics Information System. In Antibodies for Infectious Diseases; Crowe, J., Boraschi, D., Rappuoli, R., Eds.; ASM Press: Washington, DC, USA, 2015; pp. 363–379. [Google Scholar]
- Alamyar, E.; Giudicelli, V.; Shuo, L.; Duroux, P.; Lefranc, M.-P. IMGT/HighV-QUEST: The IMGT® web portal for immunoglobulin (IG) or antibody and T cell receptor (TR) analysis from NGS high throughput and deep sequencing. Immunome Res. 2012, 8, 26. [Google Scholar]
- Li, S.; Lefranc, M.-P.; Miles, J.J.; Alamyar, E.; Giudicelli, V.; Duroux, P.; Freeman, J.D.; Corbin, V.D.A.; Scheerlinck, J.-P.; Frohman, M.A.; et al. IMGT/HighV QUEST paradigm for T cell receptor IMGT clonotype diversity and next generation repertoire immunoprofiling. Nat. Commun. 2013, 4, 2333. [Google Scholar] [CrossRef]
- Giudicelli, V.; Duroux, P.; Lavoie, A.; Aouinti, S.; Lefranc, M.-P.; Kossida, S. From IMGT-ONTOLOGY to IMGT/HighV-QUEST for NGS immunoglobulin (IG) and T cell receptor (TR) repertoires in autoimmune and infectious diseases. Autoimmune Infec. Dis. 2015, 1, 1–15. [Google Scholar] [CrossRef]
- Aouinti, S.; Malouche, D.; Giudicelli, V.; Kossida, S.; Lefranc, M.-P. IMGT/HighV-QUEST statistical significance of IMGT clonotype (AA) diversity per gene for standardized comparisons of next generation sequencing immunoprofiles of immunoglobulins and T cell receptors. PLoS ONE 2015, 10, e0142353. [Google Scholar] [CrossRef]
- Aouinti, S.; Giudicelli, V.; Duroux, P.; Malouche, D.; Kossida, S.; Lefranc, M.-P. IMGT/StatClonotype for Pairwise Evaluation and Visualization of NGS IG and TR IMGT Clonotype (AA) Diversity or Expression from IMGT/HighV-QUEST. Front. Immunol. 2016, 7, 339. [Google Scholar] [CrossRef]
- Giudicelli, V.; Duroux, P.; Kossida, S.; Lefranc, M.-P. IG and TR single chain fragment variable (scFv) sequence analysis: A new advanced functionality of IMGT/V-QUEST and IMGT/HighV-QUEST. BMC Immunol. 2017, 18, 35. [Google Scholar] [CrossRef]
- Hemadou, A.; Giudicelli, V.; Smith, M.L.; Lefranc, M.-P.; Duroux, P.; Kossida, S.; Heiner, C.; Hepler, N.L.; Kuijpers, J.; Groppi, A.; et al. Pacific Biosciences Sequencing and IMGT/HighV-QUEST Analysis of Full-Length Single Chain Fragment Variable from an In Vivo Selected Phage-Display Combinatorial Library. Front. Immunol. 2017, 8, 1796. [Google Scholar] [CrossRef]
- Han, S.Y.; Antoine, A.; Howard, D.; Chang, B.; Chang, W.S.; Slein, M.; Deikus, G.; Kossida, S.; Duroux, P.; Lefranc, M.-P.; et al. Coupling of Single Molecule, Long Read Sequencing with IMGT/HighV-QUEST Analysis Expedites Identification of SIV gp140-Specific Antibodies from scFv Phage Display Libraries. Front. Immunol. 2018, 9, 329. [Google Scholar] [CrossRef]
- Lefranc, M.-P. Antibody Databases and Tools: The IMGT® Experience; Wiley: Hoboken, NJ, USA, 2009; Volume 4, pp. 91–114. [Google Scholar]
- Lefranc, M.-P. Antibody databases: IMGT®, a French platform of world-wide interest. [in French] Bases de données anticorps: IMGT®, une plate-forme française d’intérêt mondial. Médecine 2009, 25, 1020–1023. [Google Scholar] [CrossRef][Green Version]
- Ehrenmann, F.; Duroux, P.; Giudicelli, V.; Lefranc, M.-P. Standardized Sequence and Structure Analysis of Antibody Using IMGT®; Springer Nature: Basingstoke, UK, 2010; Volume 2, Chapter 2; pp. 11–31. [Google Scholar]
- Lefranc, M.-P.; Ehrenmann, F.; Ginestoux, C.; Giudicelli, V.; Duroux, P. Use of IMGT® Databases and Tools for Antibody Engineering and Humanization. In Antibody Engineering; Humana Press: Totowa, NJ, USA, 2012; pp. 3–37. [Google Scholar]
- Alamyar, E.; Giudicelli, V.; Duroux, P.; Lefranc, M.-P. Antibody V and C Domain Sequence, Structure, and Interaction Analysis with Special Reference to IMGT®. Methods Mol. Biol. 2014, 1131, 337–381. [Google Scholar]
- Lefranc, M.-P. Immunoinformatics of the V, C, and G Domains: IMGT® Definitive System for IG, TR and IgSF, MH, and MhSF. Methods Mol. Biol. 2014, 1184, 59–107. [Google Scholar]
- Marillet, S.; Lefranc, M.-P.; Boudinot, P.; Cazals, F. Novel Structural Parameters of Ig–Ag Complexes Yield a Quantitative Description of Interaction Specificity and Binding Affinity. Front. Immunol. 2017, 8, 34. [Google Scholar] [CrossRef]
- Lefranc, M.-P.; Ehrenmann, F.; Kossida, S.; Giudicelli, V.; Duroux, P. Use of IMGT® Databases and Tools for Antibody Engineering and Humanization. Microinjection 2018, 1827, 35–69. [Google Scholar] [CrossRef]
- Lefranc, M.-P. IMGT® Information System. In Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer Science+Business Media, LLC: New York, NY, USA, 2013; pp. 959–964. [Google Scholar] [CrossRef]
- Giudicelli, V.; Lefranc, M.-P. IMGT-ONTOLOGY. In Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer Science+Business Media, LLC: New York, NY, USA, 2013; pp. 964–972. [Google Scholar]
- Lefranc, M.-P. IMGT unique numbering. In Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer Science+Business Media, LLC: New York, NY, USA, 2013; pp. 952–959. [Google Scholar]
- Lefranc, M.-P. IMGT Collier de Perles. In Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer Science+Business Media, LLC: New York, NY, USA, 2013; pp. 944–952. [Google Scholar]
- Lefranc, M.-P. Antibody nomenclature: From IMGT-ONTOLOGY to INN definition. MAbs 2011, 3, 1–2. [Google Scholar] [CrossRef]
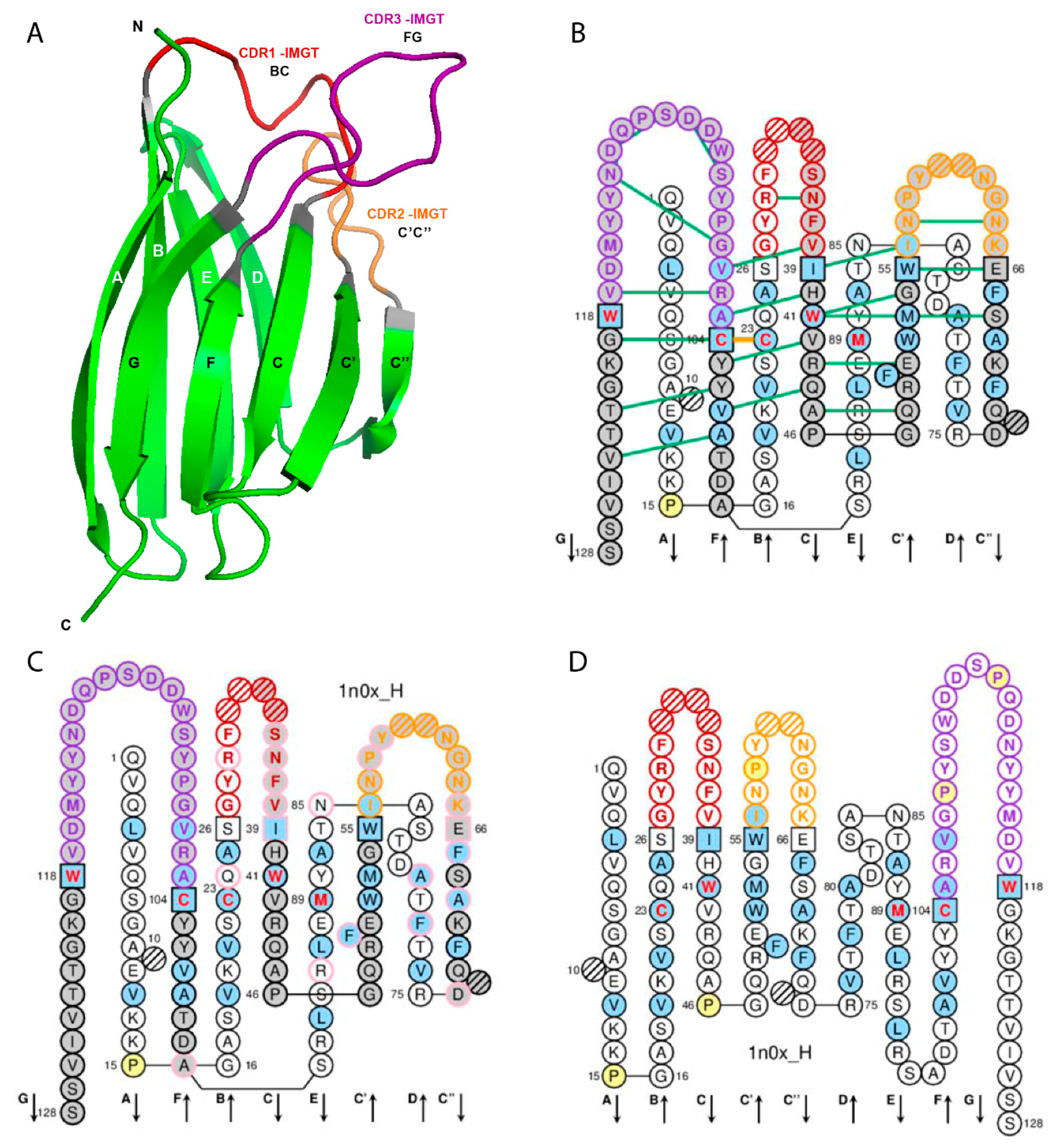
| V Domain Strands and Loops a | IMGT Positions | Lengths b | Characteristic Residue@Position c | V-DOMAIN FR-IMGT and CDR-IMGT |
|---|---|---|---|---|
| A-STRAND | 1–15 | 15 (14 if gap at 10) | FR1-IMGT | |
| B-STRAND | 16–26 | 11 | 1st-CYS 23 | |
| BC-LOOP | 27–38 | 12 (or less) | CDR1-IMGT | |
| C-STRAND | 39–46 | 8 | CONSERVED-TRP 41 | FR2-IMGT |
| C’-STRAND | 47–55 | 9 | ||
| C’C”-LOOP | 56–65 | 10 (or less) | CDR2-IMGT | |
| C”-STRAND | 66–74 | 9 (or 8 if gap at 73) | FR3-IMGT | |
| D-STRAND | 75–84 | 10 (or 8 if gaps at 81, 82) | ||
| E-STRAND | 85–96 | 12 | hydrophobic 89 | |
| F-STRAND | 97–104 | 8 | 2nd-CYS 104 | |
| FG-LOOP | 105–117 | 13 (or less, or more) | CDR3-IMGT | |
| G-STRAND | 118–128 | 11 (or 10) | V-DOMAIN J-PHE 118 or J-TRP 118 d | FR4-IMGT |
| C Domain Strands, Turns and Loops a | IMGT Positions | Lengths b | Characteristic Residue@Position c |
|---|---|---|---|
| A-STRAND | 1–15 | 15 (14 if gap at 10) | |
| AB-TURN | 15.1–15.3 | 0–3 | |
| B-STRAND | 16–26 | 11 | 1st-CYS 23 |
| BC-LOOP | 27–31 34–38 | 10 (or less) | |
| C-STRAND | 39–45 | 7 | CONSERVED-TRP 41 |
| CD-STRAND | 45.1–45.9 | 0–9 | |
| D-STRAND | 77–84 | 8 (or 7 if gap at 82) | |
| DE-TURN | 84.1–84.7 85.1–85.7 | 0–14 | |
| E-STRAND | 85–96 | 12 | hydrophobic 89 |
| EF-TURN | 96.1–96.2 | 0–2 | |
| F-STRAND | 97–104 | 8 | 2nd-CYS 104 |
| FG-LOOP | 105–117 | 13 (or less, or more) | |
| G-STRAND | 118–128 | 11 (or less) |
| IGHG1 Alleles | G1m Alleles a | IMGT Amino Acid Positions b | Populations [18] | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Allotypes | Isoallotypes c | CH1 | CH3 | ||||||||
| 103 | 120 | 12 | 14 | 101 | 110 | 115 | 116 | ||||
| G1m17/nG1m17 | G1m1/nG1m1 | /G1m27 | /G1m2 | /G1m28- | |||||||
| G1m3 d | |||||||||||
| IGHG1*01 e, IGHG1*02 e, IGHG1*05 e | G1m17,1 | I | K | D | L | V | A | H | Y | Caucasoid Negroid Mongoloid | |
| IGHG1*03 | G1m3 | nG1m1, nG1m17 | I | R | E | M | V | A | H | Y | Caucasoid |
| IGHG1*04 | G1m17,1,27 | I | K | D | L | I | A | H | Y | Negroid | |
| IGHG1*05p f | G1m17,1,28 | I | K | D | L | V | A | R | Y | Negroid | |
| IGHG1*06p f | G1m,17,1,27,28 | I | K | D | L | I | A | R | Y | Negroid | |
| IGHG1*07p f | G1m17,1,2 | I | K | D | L | V | G | H | Y | Caucasoid Mongoloid | |
| IGHG1*08p f | G1m3,1 | nG1m17 | I | R | D | L | V | A | H | Y | Mongoloid |
| IMGT Engineered Variant Nomenclature | IGHG Gene Variant Description | Property Modifications | |||||
|---|---|---|---|---|---|---|---|
| CH2 | AA and IMGT Position in CH2 of IGHG Gene Variant | CH3 | AA and IMGT Position in CH3 of IGHG Gene Variant | ADCC Enhancement or Reduction, ADCP Enhancement, B Cell Inhibition | CDC Enhancement or Reduction | Half-IG Exchange Reduction, Half-Life Increase, Knobs-Into-Holes | |
| Homsap IGHG1v1 | CH2 | P1.4 | ADCC reduction | ||||
| Homsap IGHG1v2 | CH2 | V1.3 | ADCC reduction | ||||
| Homsap IGHG1v3 | CH2 | A1.2 | ADCC reduction | ||||
| Homsap IGHG1v4 | CH2 | A114 | ADCC reduction | CDC reduction | |||
| Homsap IGHG1v5 | CH2 | W109 | ADCC reduction | CDC enhancement | |||
| Homsap IGHG1v6 | CH2 | A85.4, A118, A119 | ADCC enhancement | ||||
| Homsap IGHG1v7 | CH2 | D3, E117 | ADCC enhancement | ||||
| Homsap IGHG1v8 | CH2 | D3, L115, E117 | ADCC enhancement | CDC reduction | |||
| Homsap IGHG1v9 | CH2 | L7, P83, L85.2, I88 | CH3 | L83 | ADCC enhancement | ||
| Homsap IGHG1v10 | CH2 | Y1.3, Q1.2, W1.1, M3, D30, E34, A85.4 | ADCC enhancement | ||||
| Homsap IGHG1v11 | CH2 | E34, D109, M115, E119 | ADCC enhancement | ||||
| Homsap IGHG1v12 | CH2 | A1.1, D3, L115, E117 | ADCC enhancement | ||||
| Homsap IGHG1v13 | CH2 | A1.1, D3, E117 | ADCP enhancement | ||||
| Homsap IGHG1v14 | CH2 | A1.3, A1.2 | ADCC reduction | CDC reduction | |||
| Homsap IGHG1v15 | CH2 | S118 | CDC enhancement | ||||
| Homsap IGHG1v16 | CH2 | W109, S118 | CDC enhancement | ||||
| Homsap IGHG1v17 | CH2 | E29, F30, T107 | CDC enhancement | ||||
| Homsap IGHG1v18 | CH3 | R1, G109, Y120 | CDC enhancement | ||||
| Homsap IGHG1v19 | CH2 | A34 | CDC reduction | ||||
| Homsap IGHG1v20 | CH2 | A105 | CDC reduction | ||||
| Homsap IGHG1v21 | CH2 | Y15.1, T16, E18 | Half-life increase | ||||
| Homsap IGHG1v22 | CH2 | Y15.1, T16, E18 | CH3 | K113, F114, H116 | Half-life increase | ||
| Homsap IGHG1v23 | CH2 | E1.2 | ADCC reduction | CDC reduction | |||
| Homsap IGHG1v24 | CH3 | L107, S114 | Half-life increase | ||||
| Homsap IGHG1v25 | CH2 | E29, F113 | B cell inhibition | ||||
| Homsap IGHG1v26 | CH3 | Y22, T86 | Knobs-into-holes | ||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lefranc, M.-P.; Lefranc, G. IMGT® and 30 Years of Immunoinformatics Insight in Antibody V and C Domain Structure and Function. Antibodies 2019, 8, 29. https://doi.org/10.3390/antib8020029
Lefranc M-P, Lefranc G. IMGT® and 30 Years of Immunoinformatics Insight in Antibody V and C Domain Structure and Function. Antibodies. 2019; 8(2):29. https://doi.org/10.3390/antib8020029
Chicago/Turabian StyleLefranc, Marie-Paule, and Gérard Lefranc. 2019. "IMGT® and 30 Years of Immunoinformatics Insight in Antibody V and C Domain Structure and Function" Antibodies 8, no. 2: 29. https://doi.org/10.3390/antib8020029
APA StyleLefranc, M.-P., & Lefranc, G. (2019). IMGT® and 30 Years of Immunoinformatics Insight in Antibody V and C Domain Structure and Function. Antibodies, 8(2), 29. https://doi.org/10.3390/antib8020029





