Effects of Selected Root Exudate Components on Nitrogen Removal and Development of Denitrifying Bacteria in Constructed Wetlands
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Design
2.2. Sampling and Water Quality Analysis
2.3. DNA Extraction and qPCR Analysis
2.4. Illumina High-Throughput Sequencing
2.5. Statistical Analysis
3. Results
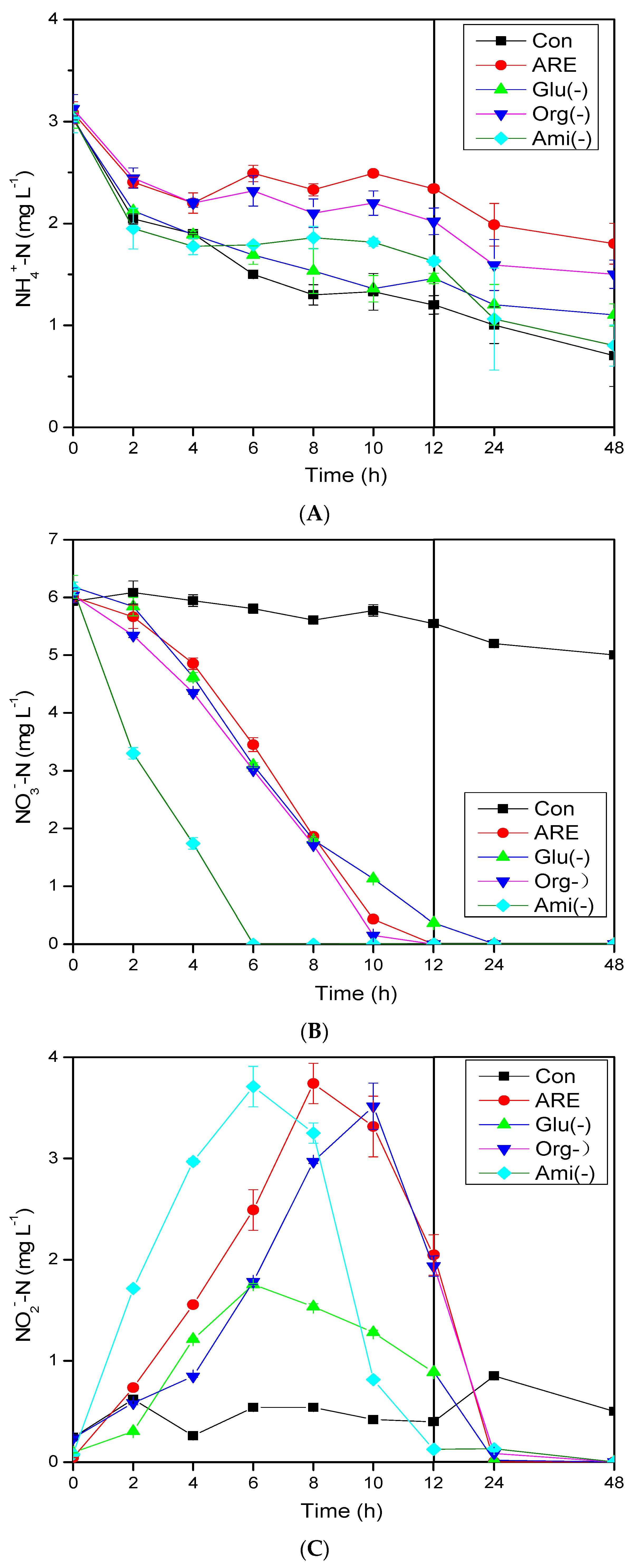
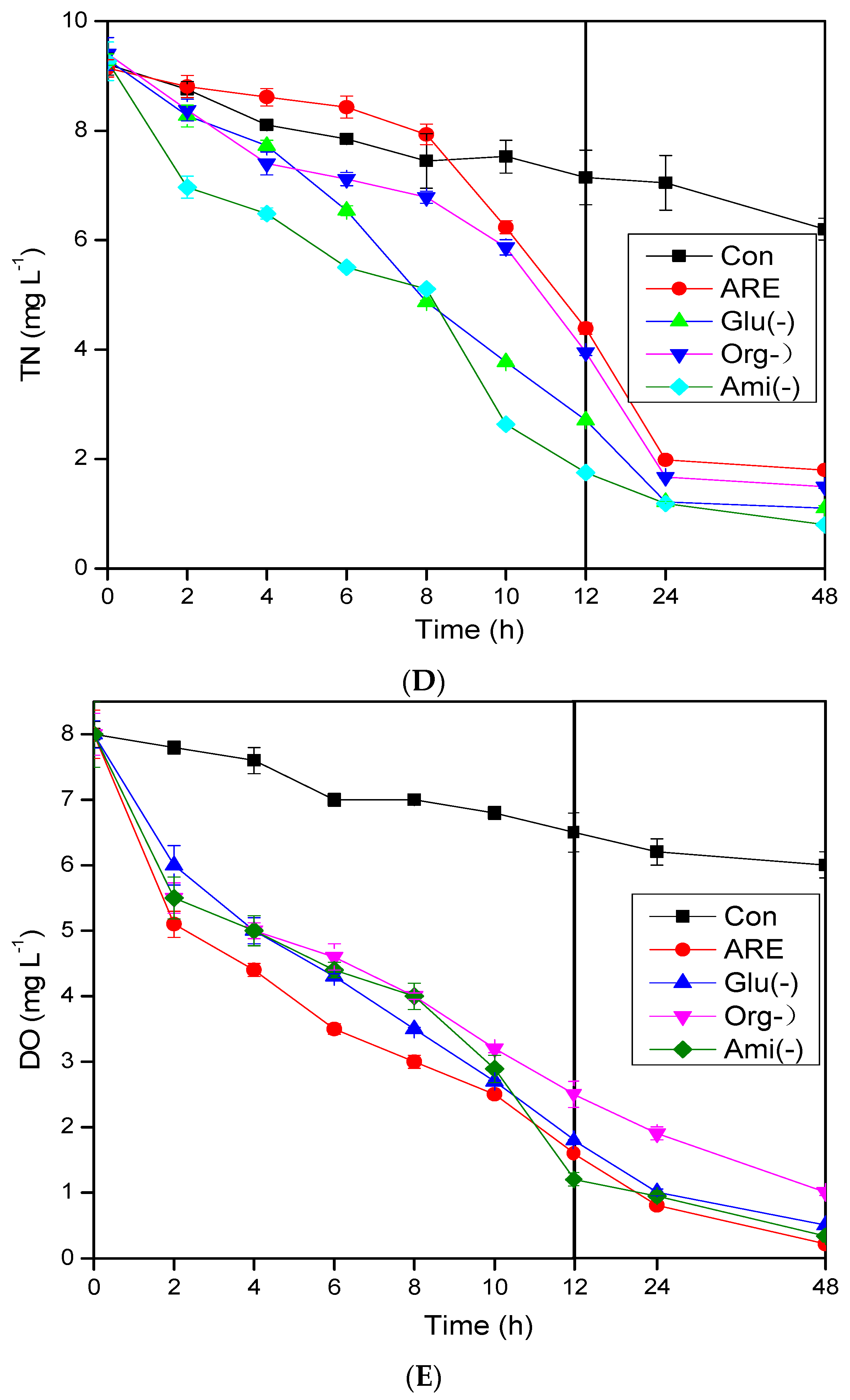
3.1. Pollutant Removal Performance
3.2. Quantification of Denitrifying Bacteria
3.3. Community Composition of Denitrifying Bacteria
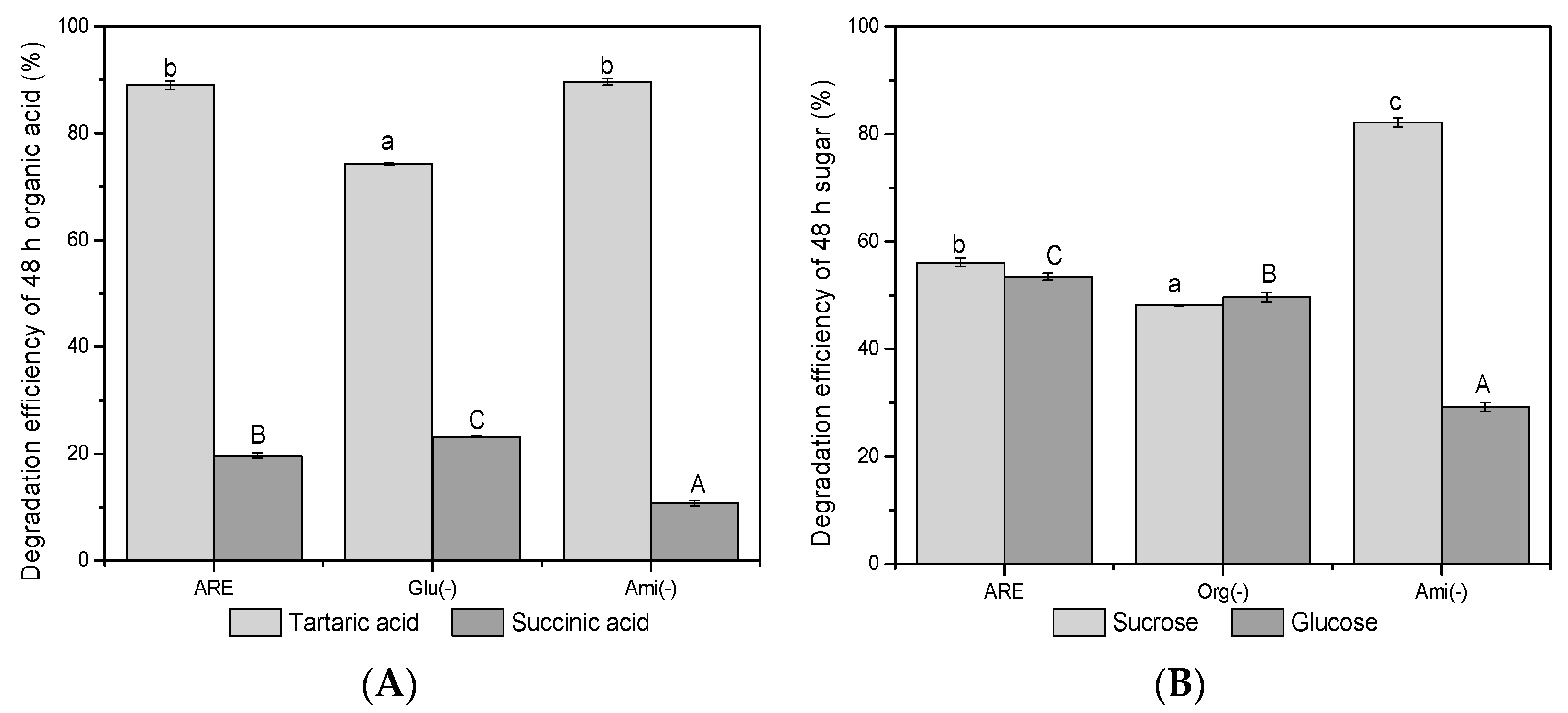
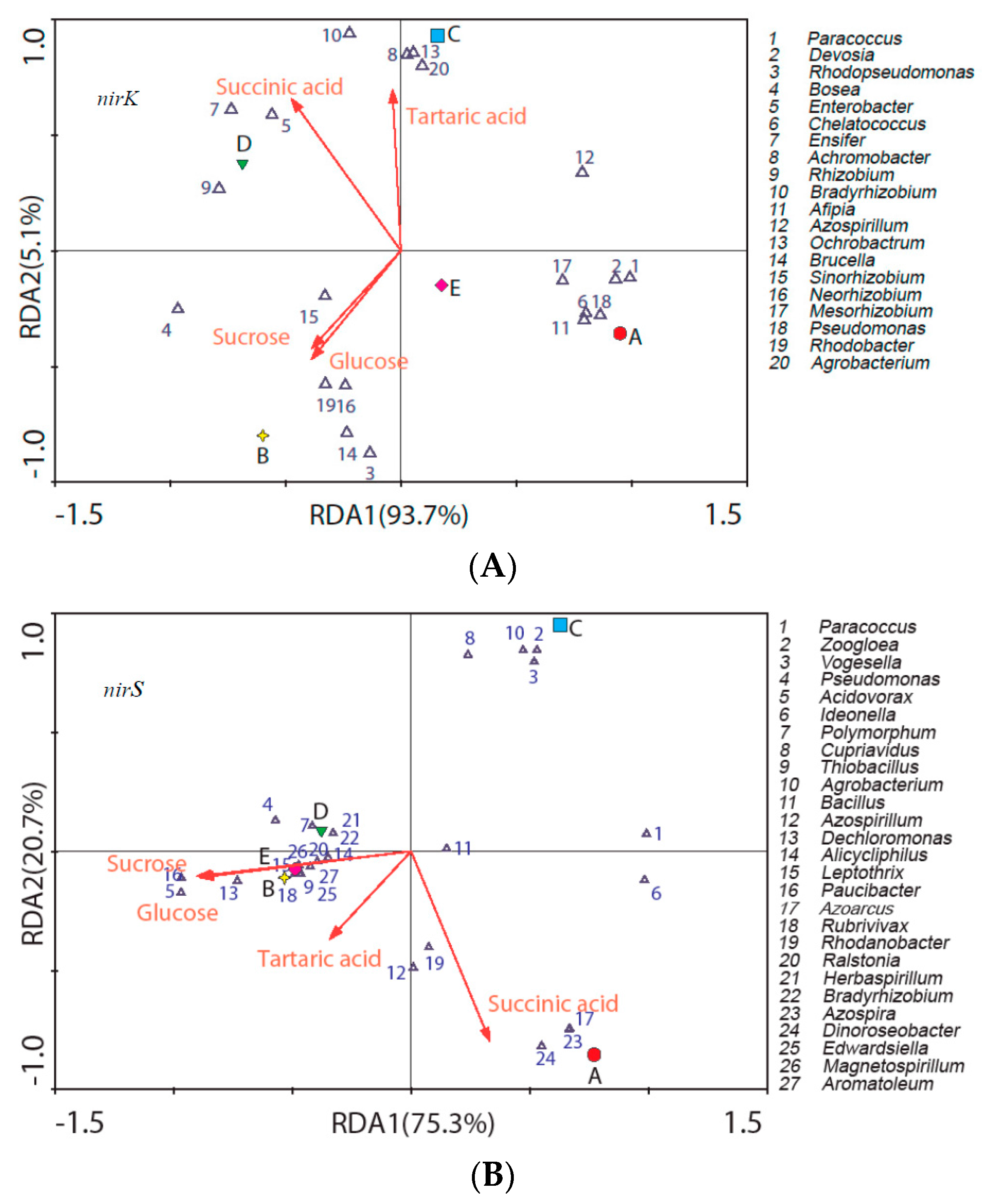
3.4. LMWC Substrates Degradation and Their Relevance on Denitrifying Microbes
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Anderson, D.M.; Glibert, P.M.; Burkholder, J.M. Harmful algal blooms and eutrophication: Nutrient sources, composition, and consequences. Estuaries 2002, 25, 704–726. [Google Scholar] [CrossRef]
- Liu, D.; Ge, Y.; Chang, J.; Peng, C.; Gu, B.; Chan, G.Y.S.; Wu, X. Constructed wetlands in China: Recent developments and future challenges. Front. Ecol. Environ. 2009, 7, 261–268. [Google Scholar] [CrossRef]
- Vymazal, J. Removal of nutrients in various types of constructed wetlands. Sci. Total Environ. 2007, 380, 48–65. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Song, X.; Wang, Y.; Yan, D. Effects of dissolved oxygen and influent COD/N ratios on nitrogen removal in horizontal subsurface flow constructed wetland. Ecol. Eng. 2012, 46, 107–111. [Google Scholar] [CrossRef]
- Fan, J.; Wang, W.; Zhang, B.; Guo, Y.; Ngo, H.H.; Guo, W.; Zhang, J.; Wu, H. Nitrogen removal in intermittently aerated vertical flow constructed wetlands: Impact of influent COD/N ratios. Bioresour. Technol. 2013, 143, 461–466. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Liu, J.; Ma, B.; Liu, Y.; Wang, B.; Peng, Y. Improving municipal wastewater nitrogen and phosphorous removal by feeding sludge fermentation products to sequencing batch reactor (SBR). Bioresour. Technol. 2016, 222, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Torresi, E.; Casas, M.E.; Polesel, F.; Plosz, B.G.; Christensson, M.; Bester, K. Impact of external carbon dose on the removal of micropollutants using methanol and ethanol in post-denitrifying Moving Bed Biofilm Reactors. Water Res. 2017, 108, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Zhai, X.; Piwpuan, N.; Arias, C.A.; Headley, T.; Brix, H. Can root exudates from emergent wetland plants fuel denitrification in subsurface flow constructed wetland systems? Ecol. Eng. 2013, 61, 555–563. [Google Scholar] [CrossRef]
- Zhao, C.; Xie, H.; Xu, J.; Xu, X.; Zhang, J.; Hu, Z.; Liu, C.; Liang, S.; Wang, Q.; Wang, J. Bacterial community variation and microbial mechanism of triclosan (TCS) removal by constructed wetlands with different types of plants. Sci. Total Environ. 2015, 505, 633–639. [Google Scholar] [CrossRef] [PubMed]
- Meng, P.; Hu, W.; Pei, H.; Hou, Q.; Ji, Y. Effect of different plant species on nutrient removal and rhizospheric microorganisms distribution in horizontal-flow constructed wetlands. Environ. Technol. 2014, 35, 808–816. [Google Scholar] [CrossRef] [PubMed]
- Salvato, M.; Borin, M.; Doni, S.; Macci, C.; Ceccanti, B.; Marinari, S.; Masciandaro, G. Wetland plants, micro-organisms and enzymatic activities interrelations in treating N polluted water. Ecol. Eng. 2012, 47, 36–43. [Google Scholar] [CrossRef]
- Huang, X.F.; Chaparro, J.M.; Reardon, K.F.; Zhang, R.; Shen, Q.; Vivanco, J.M. Rhizosphere interactions: Root exudates, microbes, and microbial communities. Botany-Botanique 2014, 92, 267–275. [Google Scholar] [CrossRef]
- Osaka, T.; Shirotani, K.; Yoshie, S.; Tsuneda, S. Effects of carbon source on denitrification efficiency and microbial community structure in a saline wastewater treatment process. Water Res. 2008, 42, 3709–3718. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.M.; Liao, M.; Yang, J.; Chai, J.J.; Fang, S.; Wang, R.H. Influence of root-exudates concentration on pyrene degradation and soil microbial characteristics in pyrene contaminated soil. Chemosphere 2012, 88, 1190–1195. [Google Scholar] [CrossRef] [PubMed]
- Bais, H.P.; Weir, T.L.; Perry, L.G.; Gilroy, S.; Vivanco, J.M. The role of root exudates in rhizosphere interations with plants and other organisms. Annu. Rev. Plant Biol. 2006, 57, 233–266. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.L.; Wang, X.Z.; He, X.J.; Zhang, S.B.; Liang, R.B.; Shen, J. Effects of root exudates on denitrifier gene abundance, community structure and activity in a micro-polluted constructed wetland. Sci. Total Environ. 2017, 598, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Xu, K.Q.; He, X.J.; Wang, X.Z. Removal of Nitrogen by Three Plant Species in Hydroponic Culture: Plant Uptake and Microbial Degradation. Water Air Soil Pollut. 2016, 227, 324. [Google Scholar] [CrossRef]
- Di, H.J.; Cameron, K.C.; Shen, J.P.; Winefield, C.S.; O’Callaghan, M.; Bowatte, S.; He, J.Z. Ammonia-oxidizing bacteria and archaea grow under contrasting soil nitrogen conditions. FEMS Microbiol. Ecol. 2010, 72, 386–394. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Mothur Analysis. Available online: https://www.mothur.org/ (accessed on 12 June 2017).
- Valentin-Vargas, A.; Root, R.A.; Neilson, J.W.; Chorover, J.; Maier, R.M. Environmental factors influencing the structural dynamics of soil microbial communities during assisted phytostabilization of acid-generating mine tailings: A mesocosm experiment. Sci. Total Environ. 2014, 500, 314–324. [Google Scholar] [CrossRef] [PubMed]
- Baudoin, E.; Benizri, E.; Guckert, A. Impact of artificial root exudates on the bacterial community structure in bulk soil and maize rhizosphere. Soil Biol. Biochem. 2003, 35, 1183–1192. [Google Scholar] [CrossRef]
- Jones, C.M.; Spor, A.; Brennan, F.P.; Breuil, M.C.; Bru, D.; Lemanceau, P.; Griffiths, B.; Hallin, S.; Philippot, L. Recently identified microbial guild mediates soil N2O sink capacity. Nat. Clim. Chang. 2014, 4, 801–805. [Google Scholar] [CrossRef]
- Hallin, S.; Hellman, M.; Choudhury, M.I.; Ecke, F. Relative importance of plant uptake and plant associated denitrification for removal of nitrogen from mine drainage in sub-arctic wetlands. Water Res. 2015, 85, 377–383. [Google Scholar] [CrossRef] [PubMed]
- Shrewsbury, L.H.; Smith, J.L.; Huggins, D.R.; Carpenter-Boggs, L.; Reardon, C.L. Denitrifier abundance has a greater influence on denitrification rates at larger landscape scales but is a lesser driver than environmental variables. Soil Biol. Biochem. 2016, 103, 221–231. [Google Scholar] [CrossRef]
- Broeckling, C.D.; Broz, A.K.; Bergelson, J.; Manter, D.K.; Vivanco, J.M. Root exudates regulate soil fungal community composition and diversty. Appl. Environ. Microbiol. 2008, 74, 738–744. [Google Scholar] [CrossRef] [PubMed]
- Houlden, A.; Timms-Wilson, T.M.; Day, M.J.; Bailey, M.J. Influence of plant developmental stage on microbial community structure and activity in the rhizosphere of three field crops. FEMS Microbiol. Ecol. 2008, 65, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Rudrappa, T.; Czymmek, K.J.; Pare, P.W.; Bais, H.P. Root-Secreted Malic Acid Recruits Beneficial Soil Bacteria. Plant Physiol. 2008, 148, 1547–1556. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Wang, S.; Lv, J.; Xu, X.; Zhang, S. Influences of artificial root exudate components on the behaviors of BDE-28 and BDE-47 in soils: Desorption, availability, and biodegradation. Environ. Sci. Pollut. Res. 2016, 23, 7702–7711. [Google Scholar] [CrossRef] [PubMed]
- Haichar, F.Z.; Santaella, C.; Heulin, T.; Achouak, W. Root exudates mediated interactions belowground. Soil Biol. Biochem. 2014, 77, 69–80. [Google Scholar] [CrossRef]
- Michalet, S.; Rohr, J.; Warshan, D.; Bardon, C.; Rogy, J.C.; Domenach, A.M.; Czarnes, S.; Pommier, T.; Combourieu, B.; Guillaumaud, N.; et al. Phytochemical analysis of mature tree root exudates in situ and their role in shaping soil microbial communities in relation to tree N-acquisition strategy. Plant Physiol. Biochem. 2013, 72, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Richardson, A.E.; O’Callaghan, M.; DeAngelis, K.M.; Jones, E.E.; Stewart, A.; Firestone, M.K.; Condron, L.M. Effects of selected root exudate components on soil bacterial communities. FEMS Microbiol. Ecol. 2011, 77, 600–610. [Google Scholar] [CrossRef] [PubMed]
- Eilers, K.G.; Lauber, C.L.; Knight, R.; Fierer, N. Shifts in bacterial community structure associated with inputs of low molecular weight carbon compounds to soil. Soil Biol. Biochem. 2010, 42, 896–903. [Google Scholar] [CrossRef]
- Torrijos, V.; Gonzalo, O.G.; Trueba-Santiso, A.; Ruiz, I.; Soto, M. Effect of by-pass and effluent recirculation on nitrogen removal in hybrid constructed wetlands for domestic and industrial wastewater treatment. Water Res. 2016, 103, 92–100. [Google Scholar] [CrossRef] [PubMed]

| Treatment | Sugars (g L−1) | Organic Acids (g L−1) | Amino Acid (mg L−1) | C input (g L−1) | N Input (mg L−1) |
|---|---|---|---|---|---|
| Con | 0 | 0 | 0 | 0 | 0 |
| ARE | Glucose-0.637 Sucrose-0.624 | Tartaric acid-0.704 Succinic acid-0.633 | Serine-0.5 glutamate-0.5 alanine-0.5 | 1 | 0.2 |
| Glu(-) | 0 | Tartaric acid-1.513 Succinic acid-1.269 | Serine-0.5 glutamate-0.5 alanine-0.5 | 1 | 0.2 |
| Org(-) | Glucose-1.241 Sucrose-1.197 | 0 | Serine-0.5 glutamate-0.5 alanine-0.5 | 1 | 0.2 |
| Ami(-) | Glucose-0.637 Sucrose-0.624 | Tartaric acid-0.704 Succinic acid-0.633 | 0 | 1 | 0 |
| Gene | Con | ARE | Glu(-) | Org(-) | Ami(-) |
|---|---|---|---|---|---|
| nirK | 8.68 × 102 ± 96 a | 1.78 × 105 ± 8.51 × 103 c | 1.46 × 105 ± 6.97 × 104 b | 8.29 × 103 ± 6.66 × 102 a | 1.79 × 103 ± 1.27 × 102 a |
| nirS | 2.43 × 103 ± 2.15 × 102 a | 1.75 × 105 ± 3.23 × 104 c | 6.40 × 104 ± 9.21 × 103 b | 6.24 × 103 ± 5.69 × 102 a | 1.59 × 103 ± 2.54 × 102 a |
| Gene | Sample | Reads | OTU | Ace | Chao | Shannon | Simpson |
|---|---|---|---|---|---|---|---|
| nirK | Con | 14,720 | 72 | 76 | 77 | 1.96 | 0.2579 |
| ARE | 31,869 | 86 | 89 | 89 | 2.67 | 0.1064 | |
| Glu(-) | 16,534 | 63 | 69 | 69 | 2.55 | 0.1216 | |
| Org(-) | 13,833 | 68 | 80 | 78 | 2.64 | 0.1211 | |
| Ami(-) | 31,068 | 89 | 91 | 90 | 2.34 | 0.2151 | |
| nirS | Con | 22,229 | 69 | 73 | 73 | 2.49 | 0.1254 |
| ARE | 22,114 | 72 | 73 | 73 | 2.56 | 0.1452 | |
| Glu(-) | 16,167 | 55 | 57 | 57 | 2.3 | 0.165 | |
| Org(-) | 14,427 | 72 | 78 | 79 | 3.03 | 0.0934 | |
| Ami(-) | 21,320 | 76 | 77 | 77 | 2.96 | 0.0889 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, H.; Wang, X.; He, X. Effects of Selected Root Exudate Components on Nitrogen Removal and Development of Denitrifying Bacteria in Constructed Wetlands. Water 2017, 9, 430. https://doi.org/10.3390/w9060430
Wu H, Wang X, He X. Effects of Selected Root Exudate Components on Nitrogen Removal and Development of Denitrifying Bacteria in Constructed Wetlands. Water. 2017; 9(6):430. https://doi.org/10.3390/w9060430
Chicago/Turabian StyleWu, Hailu, Xinze Wang, and Xiaojuan He. 2017. "Effects of Selected Root Exudate Components on Nitrogen Removal and Development of Denitrifying Bacteria in Constructed Wetlands" Water 9, no. 6: 430. https://doi.org/10.3390/w9060430
APA StyleWu, H., Wang, X., & He, X. (2017). Effects of Selected Root Exudate Components on Nitrogen Removal and Development of Denitrifying Bacteria in Constructed Wetlands. Water, 9(6), 430. https://doi.org/10.3390/w9060430