Evolution of Marine Organisms under Climate Change at Different Levels of Biological Organisation
Abstract
:1. Introduction
2. Role of Molecular and Cellular Processes in Evolutionary Responses
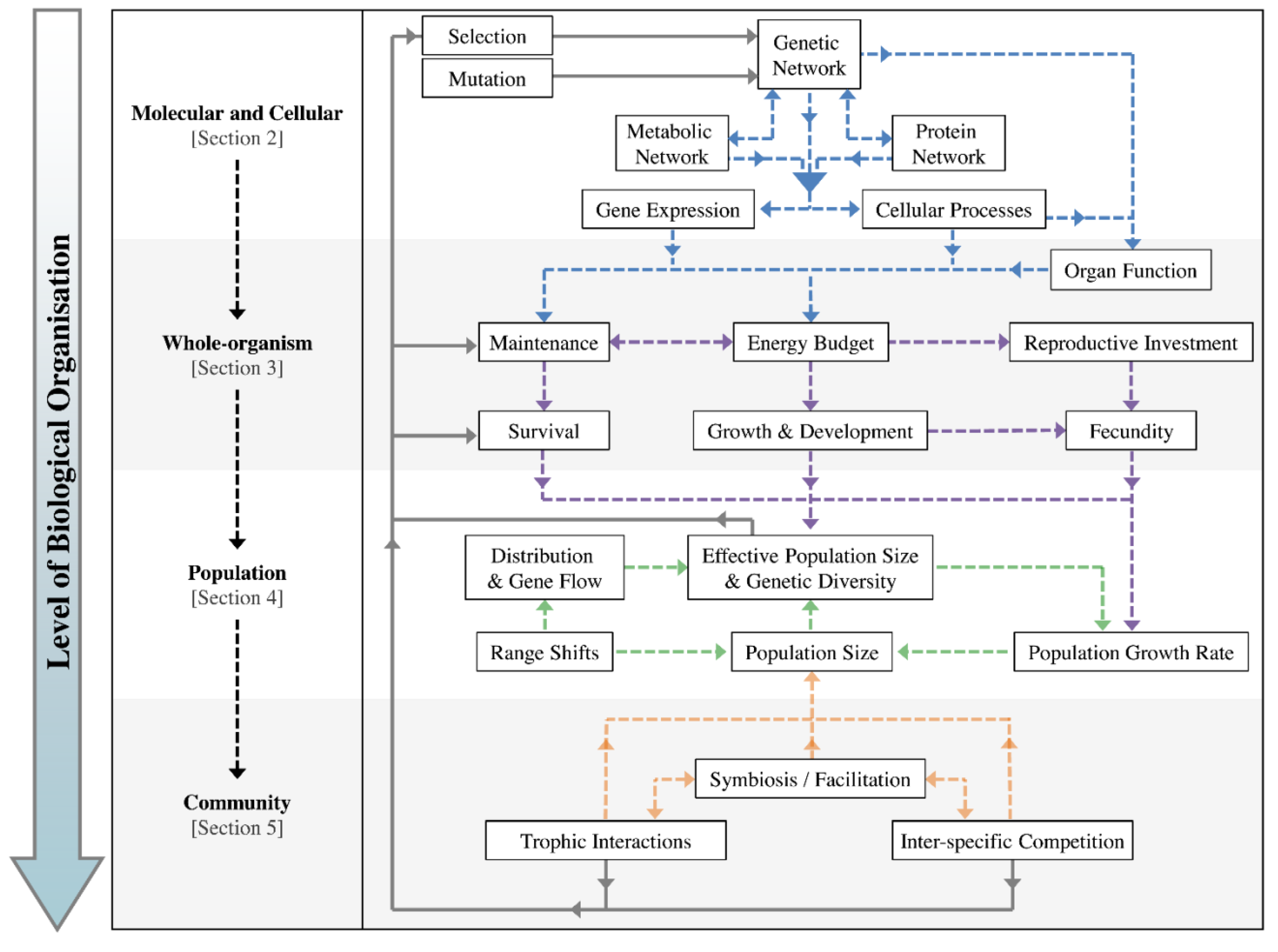
2.1. Biochemical Reactions and Gene Expression
2.2. Cellular Processes and Organ Function
2.3. Epigenetics and Trans-Generational Plasticity
3. Role of Whole-Organism Physiological and Behavioural Responses
3.1. Maintenance and Energetic Trade-Offs
3.2. Life-History Stages
3.3. Behavioural Responses
4. Role of Population-Level Responses
4.1. Demographic Processes
| Taxonomic Affiliation | Response Variable(s) | Driver | Method(s) | Evolutionary Response | Ref. |
|---|---|---|---|---|---|
| Spermatophyta: Zostera marina | Growth rate Survival | T | F | Genotypic complementarity | [107] |
| Coccolithophyceae: Emiliania huxleyi | Growth rate Production rate: (PIC) | OA | LS | Selection of genotypes Direct positive adaptation | [108] |
| Gephyrocapsa oceanica | Growth rate Carbon fixation | OA | LS | Selection of genotypes (Adaptation) | [109] |
| Diatomophyceae: Thalassiosira pseudonana | Phytosynthetic efficiency | OA | LS | No adaptation | [110] |
| Anthozoa: Acropora millepora | Thermal and physiological tolerance | T | F | Natural selection | [111] |
| Pocillopora damicornis | Coral bleaching (thermal tolerance) | T ES | CG | Local adaptation or acclimation | [112] |
| Bivalvia: Mytilus trossulus | Growth rate Survival | T | TE | Possible thermal adaptation | [113] |
| Gastropoda: Haliotis rufescens | Genetic polymorphism | T | SNP | Local adaptation Genetic differentiation | [114] |
| Polychaeta: Platynereis dumerilii | Body size | OA | TE | Genetic adaptation | [26] |
| Amphiglena mediterranea | Body size | OA | TE | Physiological plasticity | [26] |
| Amphipoda: Orchestia gammarellus | Growth Thermal tolerance | T | LS | Selection | [115] |
| Cirripedia: Semibalanus balanoides | Genetic polymorphism | T D | TE | Balancing selection Local adaptation | [116] |
| Copepoda: Tigriopus californicus | Survival (LT50) Thermal plasticity | T | LS | Low adaptation potential | [117] |
| Decapoda: Uca pugnax | Developmental rate | T | CG | Selection on variation Local adaptation | [118] |
| Echinoidea: Heliocidaris erythrogramma armigera | Hatching success | T | QG | Genotype-by- environment interaction | [119] |
| Strongylocentrotus purpuratus | Gene expression: thermal resistance | T | CG | Selection of thermally sensitive genes | [120] |
| Strongylocentrotus purpuratus | Larval body size | OA | CG | Heritability correlates with high-pCO2 | [121] |
| Centrostephanus rodgersii | Cleavage and gastrulation stage | T OA | QG | Heritable genetic variation for sires | [75] |
| Centrostephanus rodgersii | Embryonic development | T OA | CG | Varying expansion of population | [122] |
| Teleostei: Gadus morhua | Body shape | T | CG | Counter-gradient variation | [123] |
| Fundulus heteroclitus | Thermal tolerance | T | LS | Selection Regulation of heat shock proteins | [124] |
4.2. Environmental Variability
4.3. Modes of Population-Level Response
5. Community Composition and Interactions
5.1. Changes to Community Dynamics
5.2. Habitat Fragmentation and Biological Invasions
6. Future Directions
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Schmitz, O.J. Global climate change and the evolutionary ecology of ecosystem functioning. Annu. N. Y. Acad. Sci. 2013, 1297, 61–72. [Google Scholar]
- Bell, G.; Gonzalez, A. Evolutionary rescue can prevent extinction following environmental change. Ecol. Lett. 2009, 12, 942–948. [Google Scholar] [CrossRef] [PubMed]
- Burrows, M.T.; Schoeman, D.S.; Buckley, L.B.; Moore, P.; Poloczanska, E.S.; Brander, K.M.; Brown, C.; Bruno, J.F.; Duarte, C.M.; Halpern, B.S.; et al. The pace of shifting climate in marine and terrestrial ecosystems. Science 2011, 334, 652–655. [Google Scholar] [CrossRef] [PubMed]
- Doney, S.C.; Schimel, D.S. Carbon and climate system coupling on timescales from the precambrian to the anthropocene. Annu. Rev. Environ. Resour. 2007, 32, 31–66. [Google Scholar] [CrossRef]
- Halpern, B.S.; Walbridge, S.; Selkoe, K.A.; Kappel, C.V.; Micheli, F.; D’Agrosa, C.; Bruno, J.F.; Casey, K.S.; Ebert, C.; Fox, H.E.; et al. A global map of human impact on marine ecosystems. Science 2008, 319, 948–952. [Google Scholar] [CrossRef] [PubMed]
- Dupont, S.; Dorey, N.; Thorndyke, M. What meta-analysis can tell us about vulnerability of marine biodiversity to ocean acidification? Estuar. Coast. Shelf Sci. 2010, 89, 182–185. [Google Scholar] [CrossRef]
- Kroeker, K.; Kordas, R.; Crim, R.; Singh, G. Meta-analysis reveals negative yet variable effects of ocean acidification on marine organisms. Ecol. Lett. 2010, 13, 1419–1434. [Google Scholar] [CrossRef] [PubMed]
- Harvey, B.P.; Gwynn-Jones, D.; Moore, P.J. Meta-analysis reveals complex marine biological responses to the interactive effects of ocean acidification and warming. Ecol. Evol. 2013, 3, 1016–1030. [Google Scholar] [CrossRef] [PubMed]
- Kroeker, K.J.; Kordas, R.L.; Crim, R.; Hendriks, I.E.; Ramajo, L.; Singh, G.S.; Duarte, C.M.; Gattuso, J.-P. Impacts of ocean acidification on marine organisms: Quantifying sensitivities and interaction with warming. Glob. Change Biol. 2013, 19, 1884–1896. [Google Scholar] [CrossRef]
- Pörtner, H.-O. Physiological basis of temperature-dependent biogeography: Trade-offs in muscle design and performance in polar ectotherms. J. Exp. Biol. 2002, 205, 2217–2230. [Google Scholar] [PubMed]
- Somero, G.N. Linking biogeography to physiology: Evolutionary and acclimatory adjustments of thermal limits. Front. Zool. 2005, 2, 1. [Google Scholar] [CrossRef] [PubMed]
- Widdicombe, S.; Spicer, J. Predicting the impact of ocean acidification on benthic biodiversity: What can animal physiology tell us? J. Exp. Mar. Biol. Ecol. 2008, 366, 187–197. [Google Scholar] [CrossRef]
- Somero, G.N. The physiology of climate change: How potentials for acclimatization and genetic adaptation will determine ‘winners’ and ‘losers’. J. Exp. Biol. 2010, 213, 912–920. [Google Scholar] [CrossRef] [PubMed]
- Van der Putten, W.H.; Macel, M.; Visser, M.E. Predicting species distribution and abundance responses to climate change: Why it is essential to include biotic interactions across trophic levels. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2010, 365, 2025–2034. [Google Scholar] [CrossRef] [PubMed]
- Stockwell, C.A.; Hendry, A.P.; Kinnison, M.T. Contemporary evolution meets conservation biology. Trends Ecol. Evol. 2003, 18, 94–101. [Google Scholar] [CrossRef]
- Munday, P.L.; Warner, R.R.; Monro, K.; Pandolfi, J.M.; Marshall, D.J. Predicting evolutionary responses to climate change in the sea. Ecol. Lett. 2013, 16, 1488–1500. [Google Scholar] [CrossRef] [PubMed]
- Sunday, J.M.; Calosi, P.; Dupont, S.; Munday, P.L.; Stillman, J.H.; Reusch, T.B.H. Evolution in an acidifying ocean. Trends Ecol. Evol. 2014, 29, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Dupont, S.; Portner, H. Marine science: Get ready for ocean acidification. Nature 2013, 498, 429. [Google Scholar] [CrossRef] [PubMed]
- Gienapp, P.; Teplitsky, C.; Alho, J.S.; Mills, J.A.; Merilä, J. Climate change and evolution: Disentangling environmental and genetic responses. Mol. Ecol. 2008, 17, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Merilä, J.; Hendry, A.P. Climate change, adaptation, and phenotypic plasticity: The problem and the evidence. Evol. Appl. 2014, 7, 1–14. [Google Scholar] [CrossRef]
- Chevin, L.-M.; Lande, R.; Mace, G.M. Adaptation, plasticity, and extinction in a changing environment: Towards a predictive theory. PLoS Biol. 2010, 8. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.A.; Sgrò, C.M. Climate change and evolutionary adaptation. Nature 2011, 470, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Reusch, T.B.H. Climate change in the oceans: Evolutionary versus phenotypically plastic responses of marine animals and plants. Evol. Appl. 2013, 7, 104–122. [Google Scholar] [CrossRef] [PubMed]
- Blondel, J. From biogeography to life history theory: A multithematic approach illustrated by the biogeography of vertebrates. J. Biogeogr. 1987, 14, 405–422. [Google Scholar] [CrossRef]
- Palumbi, S.R.; Barshis, D.J.; Traylor-Knowles, N.; Bay, R.A. Mechanisms of reef coral resistance to future climate change. Science 2014, 344, 895–898. [Google Scholar] [CrossRef] [PubMed]
- Calosi, P.; Rastrick, S.P.S.; Lombardi, C.; de Guzman, H.J.; Davidson, L.; Jahnke, M.; Giangrande, A.; Hardege, J.D.; Schulze, A.; Spicer, J.I.; et al. Adaptation and acclimatization to ocean acidification in marine ectotherms: An in situ transplant experiment with polychaetes at a shallow CO2 vent system. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368. [Google Scholar] [CrossRef] [PubMed]
- Queirós, A.M.; Fernandes, J.A.; Faulwetter, S.; Nunes, J.; Rastrick, S.P.S.; Mieszkowska, N.; Artioli, Y.; Yool, A.; Calosi, P.; Arvanitidis, C.; et al. Scaling up experimental ocean acidification and warming research: From individuals to the ecosystem. Glob. Change Biol. 2014. [Google Scholar] [CrossRef]
- Prosser, C.L. Physiological variation in animals. Biol. Rev. 1955, 30, 229–261. [Google Scholar] [CrossRef]
- Somero, G.N. The physiology of global change: Linking patterns to mechanisms. Annu. Rev. Mar. Sci. 2012, 4, 39–61. [Google Scholar] [CrossRef]
- Feder, M.E.; Walser, J.C. The biological limitations of transcriptomics in elucidating stress and stress responses. J. Evol. Biol. 2005, 18, 901–910. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Biochemistry; WH Freeman: New York, NY, USA, 2002; p. 1026. [Google Scholar]
- Griffiths, A.J.F.; Miller, J.H.; Suzuki, D.T.; Lewontin, R.C.; Gelbart, W.M. Gene-protein relations. In An Introduction to Genetic Analysis, 7th ed.; Griffiths, A.J.F., Ed.; WH Freeman: New York, NY, USA, 2000. [Google Scholar]
- Kassahn, K.S.; Crozier, R.H.; Pörtner, H.O.; Caley, M.J. Animal performance and stress: Responses and tolerance limits at different levels of biological organisation. Biol. Rev. Camb. Philos. Soc. 2009, 84, 277–292. [Google Scholar] [CrossRef] [PubMed]
- Fields, P.A.; Rudomin, E.L.; Somero, G.N. Temperature sensitivities of cytosolic malate dehydrogenases from native and invasive species of marine mussels (genus Mytilus): Sequence-function linkages and correlations with biogeographic distribution. J. Exp. Biol. 2006, 209, 656–667. [Google Scholar] [CrossRef] [PubMed]
- Powers, D.A.; Schulte, P.M. Evolutionary adaptations of gene structure and expression in natural populations in relation to a changing environment: A multidisciplinary approach to address the million-year saga of a small fish. J. Exp. Zool. 1998, 282, 71–94. [Google Scholar] [CrossRef] [PubMed]
- Norry, F.M.; Larsen, P.F.; Liu, Y.; Loeschcke, V. Combined expression patterns of QTL-linked candidate genes best predict thermotolerance in Drosophila melanogaster. J. Insect Physiol. 2009, 55, 1050–1057. [Google Scholar] [CrossRef] [PubMed]
- Franks, S.J.; Hoffmann, A.A. Genetics of climate change adaptation. Annu. Rev. Genet. 2012, 46, 185–208. [Google Scholar] [CrossRef] [PubMed]
- Davidson, E.H. The Regulatory Genome: Gene Regulatory Networks in Development and Evolution; Academic Press: San Diego, CA, USA, 2006. [Google Scholar]
- Davidson, E.; Levin, M. Gene regulatory networks. Proc. Natl. Acad. Sci. USA 2005, 102, 4935. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.E.; Shoo, L.P.; Isaac, J.L.; Hoffmann, A.A.; Langham, G. Towards an integrated framework for assessing the vulnerability of species to climate change. PLoS Biol. 2008, 6, e325. [Google Scholar] [CrossRef]
- Bradshaw, W.E.; Holzapfel, C.M. Light, time, and the physiology of biotic response to rapid climate change in animals. Annu. Rev. Physiol. 2010, 72, 147–166. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, N.J.; Urwin, P.E. The interaction of plant biotic and abiotic stresses: From genes to the field. J. Exp. Bot. 2012, 63, 3523–3543. [Google Scholar] [CrossRef] [PubMed]
- Gardner, J.L.; Peters, A.; Kearney, M.R.; Joseph, L.; Heinsohn, R. Declining body size: A third universal response to warming? Trends Ecol. Evol. 2011, 26, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Kovach, R.P.; Gharrett, A.J.; Tallmon, D.A. Genetic change for earlier migration timing in a pink salmon population. Proc. R. Soc. Biol. Sci. Ser. B 2012, 279, 3870–3878. [Google Scholar] [CrossRef]
- Karell, P.; Ahola, K.; Karstinen, T.; Valkama, J.; Brommer, J.E. Climate change drives microevolution in a wild bird. Nat. Commun. 2011, 2, 208. [Google Scholar] [CrossRef] [PubMed]
- Runcie, D.E.; Garfield, D.A.; Babbitt, C.C.; Wygoda, J.A.; Mukherjee, S.; Wray, G.A. Genetics of gene expression responses to temperature stress in a sea urchin gene network. Mol. Ecol. 2012, 21, 4547–4562. [Google Scholar] [CrossRef] [PubMed]
- Keller, B.; Feuillet, C. Colinearity and gene density in grass genomes. Trends Plant Sci. 2000, 5, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Barshis, D.J.; Ladner, J.T.; Oliver, T.A.; Seneca, F.O.; Traylor-Knowles, N.; Palumbi, S.R. Genomic basis for coral resilience to climate change. Proc. Natl. Acad. Sci. USA 2013, 110, 1387–1392. [Google Scholar] [CrossRef]
- Todgham, A.E.; Hofmann, G.E. Transcriptomic response of sea urchin larvae Strongylocentrotus purpuratus to CO2-driven seawater acidification. J. Exp. Biol. 2009, 212, 2579–2594. [Google Scholar] [CrossRef] [PubMed]
- Wray, G.A. The evolutionary significance of cis-regulatory mutations. Nat. Rev. Genet. 2007, 8, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Barrett, R.D.; Hoekstra, H.E. Molecular spandrels: Tests of adaptation at the genetic level. Nat. Rev. Genet. 2011, 12, 767–780. [Google Scholar] [CrossRef] [PubMed]
- Davidson, W.S. Adaptation genomics: next generation sequencing reveals a shared haplotype for rapid early development in geographically and genetically distant populations of rainbow trout. Mol. Ecol. 2012, 21, 219–222. [Google Scholar] [CrossRef] [PubMed]
- Edwards, S.V. Next-generation QTL mapping: Crowdsourcing SNPs, without pedigrees. Mol. Ecol. 2013, 22, 3885–3887. [Google Scholar] [CrossRef] [PubMed]
- Kaniewska, P.; Campbell, P.R.; Kline, D.I.; Rodriguez-Lanetty, M.; Miller, D.J.; Dove, S.; Hoegh-Guldberg, O. Major cellular and physiological impacts of ocean acidification on a reef building coral. PLoS ONE 2012, 7, e34659. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, I.M.; Frederich, M.; Bagwe, R.; Lannig, G.; Sukhotin, A.A. Energy homeostasis as an integrative tool for assessing limits of environmental stress tolerance in aquatic invertebrates. Mar. Environ. Res. 2012, 79, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Seebacher, F.; Franklin, C.E. Determining environmental causes of biological effects: The need for a mechanistic physiological dimension in conservation biology. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 1607–1614. [Google Scholar] [CrossRef] [PubMed]
- Buckley, B.A.; Gracey, A.Y.; Somero, G.N. The cellular response to heat stress in the goby Gillichthys mirabilis: A cDNA microarray and protein-level analysis. J. Exp. Biol. 2006, 209, 2660–2677. [Google Scholar] [CrossRef] [PubMed]
- Wapinski, I.; Pfeffer, A.; Friedman, N.; Regev, A. Natural history and evolutionary principles of gene duplication in fungi. Nature 2007, 449, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Raffaele, S.; Farrer, R.A.; Cano, L.M.; Studholme, D.J.; MacLean, D.; Thines, M.; Jiang, R.H.Y.; Zody, M.C.; Kunjeti, S.G.; Donofrio, N.M.; et al. Genome evolution following host jumps in the Irish potato famine pathogen lineage. Science 2010, 330, 1540–1543. [Google Scholar] [CrossRef] [PubMed]
- Hartl, D.L. Essential Genetics: A Genomics Perspective; Jones and Bartlett Publishers: London, UK, 2014. [Google Scholar]
- Pires-daSilva, A.; Sommer, R.J. The evolution of signalling pathways in animal development. Nat. Rev. Genet. 2003, 4, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Moreno, B. Adaptations of proteins to cellular and subcellular pH. J. Biol. 2009, 8, 98. [Google Scholar] [CrossRef] [PubMed]
- Salinas, S.; Munch, S.B. Thermal legacies: transgenerational effects of temperature on growth in a vertebrate. Ecol. Lett. 2012, 15, 159–163. [Google Scholar] [CrossRef] [PubMed]
- Jaenisch, R.; Bird, A. Epigenetic regulation of gene expression: How the genome integrates intrinsic and environmental signals. Nat. Genet. 2003, 33, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Turner, B.M. Epigenetic responses to environmental change and their evolutionary implications. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 3403–3418. [Google Scholar] [CrossRef]
- Parker, L.M.; Ross, P.M.; O’Connor, W.A.; Borysko, L.; Raftos, D.A.; Pörtner, H.-O. Adult exposure influences offspring response to ocean acidification in oysters. Glob. Change Biol. 2012, 18, 82–92. [Google Scholar] [CrossRef]
- Garfield, D.A.; Runcie, D.E.; Babbitt, C.C.; Haygood, R.; Nielsen, W.J.; Wray, G.A. The Impact of gene expression variation on the robustness and evolvability of a developmental Gene Regulatory Network. PLoS Biol. 2013, 11. [Google Scholar] [CrossRef] [PubMed]
- Suchanek, T.H. The role of disturbance in the evolution of life history strategies in the intertidal mussels Mytilus edulis and Mytilus californianus. Oecologia 1981, 50, 143–152. [Google Scholar] [CrossRef]
- Dawson, M.N. Phylogeography in coastal marine animals: A solution from California? J. Biogeogr. 2001, 28, 723–736. [Google Scholar] [CrossRef]
- Sherman, C.D.H.; Hunt, A.; Ayre, D.J. Is life history a barrier to dispersal? Contrasting patterns of genetic differentiation along an oceanographically complex coast. Biol. J. Linn. Soc. 2008, 95, 106–116. [Google Scholar] [CrossRef]
- Sunday, J.M.; Crim, R.N.; Harley, C.D.G.; Hart, M.W. Quantifying rates of evolutionary adaptation in response to ocean acidification. PLoS ONE 2011, 6, e22881. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.H.; Gillooly, J.F.; Allen, A.P.; Savage, V.M.; West, G.B. Toward a metabolic theory of ecology. Ecology 2004, 85, 1771–1789. [Google Scholar] [CrossRef]
- Kooijman, S.A.L.M. Dynamic Energy Budget Theory for Metabolic Organisation; Cambridge University Press: Cambridge, UK, 2010; p. 514. [Google Scholar]
- Holcomb, M.; McCorkle, D.C.; Cohen, A.L. Long-term effects of nutrient and CO2 enrichment on the temperate coral astrangia poculata (Ellis and Solander, 1786). J. Exp. Mar. Biol. Ecol. 2010, 386, 27–33. [Google Scholar]
- Foo, S.A.; Dworjanyn, S.A.; Poore, A.G.B.; Byrne, M. Adaptive capacity of the habitat modifying sea urchin Centrostephanus rodgersii to ocean warming and ocean acidification: Performance of early embryos. PLoS ONE 2012, 7, e42497. [Google Scholar] [CrossRef] [PubMed]
- Langenbuch, M.; Pörtner, H.O. High sensitivity to chronically elevated CO2 levels in a eurybathic marine sipunculid. Aquat. Toxicol. 2004, 70, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, H. Effects of CO2-driven ocean acidification on the early developmental stages of invertebrates. Mar. Ecol. Prog. Ser. 2008, 373, 275–284. [Google Scholar] [CrossRef]
- Pörtner, H.-O. Ecosystem effects of ocean acidification in times of ocean warming: A physiologist’s view. Mar. Ecol. Prog. Ser. 2008, 373, 203–217. [Google Scholar] [CrossRef]
- O'Connor, M.I.; Bruno, J.F.; Gaines, S.D.; Halpern, B.S.; Lester, S.E.; Kinlan, B.P.; Weiss, J.M. Temperature control of larval dispersal and the implications for marine ecology, evolution, and conservation. Proc. Natl. Acad. Sci. USA 2007, 104, 1266–1271. [Google Scholar] [CrossRef] [PubMed]
- Arnold, K.; Findlay, H.; Spicer, J.; Daniels, C.; Boothroyd, D. Effect of CO2-related acidification on aspects of the larval development of the European lobster, Homarus gammarus (L.). Biogeosciences 2009, 6, 1747–1754. [Google Scholar] [CrossRef]
- McDonald, M.R.; McClintock, J.B.; Amsler, C.D.; Rittschof, D.; Angus, R.A.; Orihuela, B.; Lutostanski, K. Effects of ocean acidification over the life history of the barnacle Amphibalanus amphitrite. Mar. Ecol. Prog. Ser. 2009, 385, 179–187. [Google Scholar] [CrossRef]
- Byrne, M. Impact of ocean warming and ocean acidification on marine invertebrate life history stages: Vulnerabilities and potential for persistence in a changing ocean. Oceanogr. Mar. Biol. 2011, 49, 1–42. [Google Scholar]
- Miller, N.A.; Paganini, A.W.; Stillman, J.H. Differential thermal tolerance and energetic trajectories during ontogeny in porcelain crabs, genus Petrolisthes. J. Therm. Biol. 2013, 38, 79–85. [Google Scholar] [CrossRef]
- Emlet, R.B.; Sadro, S.S. Linking stages of life history: How larval quality translates into juvenile performance for an intertidal barnacle (Balanus glandula). Integr. Comp. Biol. 2006, 46, 334–346. [Google Scholar] [CrossRef] [PubMed]
- Werner, E.E.; Gilliam, J.F. The ontogenetic niche and species interactions in size-structured populations. Annu. Rev. Ecol. Syst. 1984, 15, 393–425. [Google Scholar] [CrossRef]
- Dupont, S.; Dorey, N.; Stumpp, M.; Melzner, F.; Thorndyke, M.C. Long-term and trans-life-cycle effects of exposure to ocean acidification in the green sea urchin Strongylocentrotus droebachiensis. Mar. Biol. 2012, 160, 1835–1843. [Google Scholar] [CrossRef]
- Ferrari, M.C.O.; Manassa, R.P.; Dixson, D.L.; Munday, P.L.; McCormick, M.I.; Meekan, M.G.; Sih, A.; Chivers, D.P. Effects of ocean acidification on learning in coral reef fishes. PLoS ONE 2012, 7, e31478. [Google Scholar] [CrossRef] [PubMed]
- Candolin, U.; Wong, B.B.M. Behavioural Responses to a Changing World: Mechanisms and Consequences; Oxford University Press: Oxford, UK, 2012; pp. 208–210. [Google Scholar]
- Dissanayake, A.; Ishimatsu, A. Synergistic effects of elevated CO2 and temperature on the metabolic scope and activity in a shallow-water coastal decapod (Metapenaeus joyneri; Crustacea: Penaeidae). ICES J. Mar. Sci. 2011, 68, 1147–1154. [Google Scholar] [CrossRef]
- Biro, P.A.; Beckmann, C.; Stamps, J.A. Small within-day increases in temperature affects boldness and alters personality in coral reef fish. Proc. R. Soc. Biol. Sci. B 2010, 277, 71–77. [Google Scholar] [CrossRef]
- Watson, S.-A.; Lefevre, S.; McCormick, M.I.; Domenici, P.; Nilsson, G.E.; Munday, P.L. Marine mollusc predator-escape behaviour altered by near-future carbon dioxide levels. Proc. R. Soc. Biol. Sci. B 2014, 281. [Google Scholar] [CrossRef]
- Nilsson, G.E.; Dixson, D.L.; Domenici, P.; McCormick, M.I.; Sørensen, C.; Watson, S.-A.; Munday, P.L. Near-future carbon dioxide levels alter fish behaviour by interfering with neurotransmitter function. Nat. Clim. Change 2012, 2, 201–204. [Google Scholar] [CrossRef]
- Chivers, D.P.; McCormick, M.I.; Nilsson, G.E.; Munday, P.L.; Watson, S.-A.; Meekan, M.G.; Mitchell, M.D.; Corkill, K.C.; Ferrari, M.C.O. Impaired learning of predators and lower prey survival under elevated CO2: A consequence of neurotransmitter interference. Glob. Change Biol. 2014, 20, 515–522. [Google Scholar] [CrossRef]
- Munday, P.L.; Dixson, D.L.; Donelson, J.M.; Jones, G.P.; Pratchett, M.S.; Devitsina, G.V.; Døving, K.B. Ocean acidification impairs olfactory discrimination and homing ability of a marine fish. Proc. Natl. Acad. Sci. USA 2009, 106, 1848. [Google Scholar] [CrossRef] [PubMed]
- Goh, C.S.; Bogan, A.A.; Joachimiak, M.; Walther, D.; Cohen, F.E. Co-evolution of proteins with their interaction partners. J. Mol. Biol. 2000, 299, 283–293. [Google Scholar] [CrossRef]
- Greenwood, A.K.; Wark, A.R.; Yoshida, K.; Peichel, C.L. Genetic and neural modularity underlie the evolution of schooling behavior in threespine sticklebacks. Curr. Biol. 2013, 23, 1884–1888. [Google Scholar] [CrossRef] [PubMed]
- Blomberg, S.P.; Garland, T.; Ives, A.R. Testing for phylogenetic signal in comparative data: Behavioral traits are more labile. Evolution 2003, 57, 717–745. [Google Scholar] [CrossRef] [PubMed]
- Bradshaw, A.D.; McNeilly, T. Evolutionary response to global climatic change. Ann. Bot. Lond. 1991, 67, 5–14. [Google Scholar]
- Bohonak, A.J. Dispersal, gene flow, and population structure. Q. Rev. Biol. 1999, 74, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Hellberg, M.E.; Burton, R.S.; Neigel, J.E.; Palumbi, S.R. Genetic assessment of connectivity among marine populations. Bull. Mar. Sci. 2002, 70, 273–290. [Google Scholar]
- Sanford, E.; Kelly, M.W. Local adaptation in marine invertebrates. Ann. Rev. Mar. Sci. 2011, 3, 509–535. [Google Scholar] [CrossRef] [PubMed]
- Merilä, J. Evolution in response to climate change: In pursuit of the missing evidence. BioEssays 2012, 34, 811–818. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Ronce, O.; Ferriere, R.; Hochberg, M.E. Evolutionary rescue: An emerging focus at the intersection between ecology and evolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368. [Google Scholar] [CrossRef] [PubMed]
- Johannesson, K.; Smolarz, K.; Grahn, M.; André, C. The future of Baltic Sea populations: Local extinction or evolutionary rescue? Ambio 2011, 40, 179–190. [Google Scholar] [CrossRef] [PubMed]
- Bell, G. Evolutionary rescue and the limits of adaptation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368. [Google Scholar] [CrossRef] [PubMed]
- Cuveliers, E.L.; Volckaert, F.A.M.; Rijnsdorp, A.D.; Larmuseau, M.H.D.; Maes, G.E. Temporal genetic stability and high effective population size despite fisheries-induced life-history trait evolution in the North Sea sole. Mol. Ecol. 2011, 20, 3555–3568. [Google Scholar] [PubMed]
- Reusch, T.B.H.; Ehlers, A.; Hämmerli, A.; Worm, B. Ecosystem recovery after climatic extremes enhanced by genotypic diversity. Proc. Natl. Acad. Sci. USA 2005, 102, 2826–2831. [Google Scholar] [CrossRef] [PubMed]
- Lohbeck, K.T.; Riebesell, U.; Reusch, T.B.H. Adaptive evolution of a key phytoplankton species to ocean acidification. Nat. Geosci. 2012, 5, 346–351. [Google Scholar] [CrossRef]
- Jin, P.; Gao, K.; Beardall, J. Evolutionary responses of a coccolithophorid Gephyrocapsa oceanica to ocean acidification. Evolution 2013, 67, 1869–1878. [Google Scholar] [CrossRef]
- Crawfurd, K.J.; Raven, J.A.; Wheeler, G.L.; Baxter, E.J.; Joint, I. The response of Thalassiosira pseudonana to long-term exposure to increased CO2 and decreased pH. PLoS ONE 2011, 6, e26695. [Google Scholar] [CrossRef] [PubMed]
- Smith-Keune, C.; Oppen, M. Genetic structure of a reef-building coral from thermally distinct environments on the Great Barrier Reef. Coral Reefs 2006, 25, 493–502. [Google Scholar] [CrossRef]
- D’Croz, L.; Maté, J.L. Experimental responses to elevated water temperature in genotypes of the reef coral Pocillopora damicornis from upwelling and non-upwelling environments in Panama. Coral Reefs 2004, 23, 473–483. [Google Scholar] [CrossRef]
- Yanick, J.F.; Heath, J.W.; Heath, D.D. Survival and growth of local and transplanted blue mussels (Mytilus trossulus, Lamark). Aquac. Res. 2003, 34, 869–875. [Google Scholar]
- De Wit, P.; Palumbi, S.R. Transcriptome-wide polymorphisms of red abalone (Haliotis rufescens) reveal patterns of gene flow and local adaptation. Mol. Ecol. 2013, 22, 2884–2897. [Google Scholar] [CrossRef] [PubMed]
- Gaston, K.J.; Spicer, J.I. Do upper thermal tolerances differ in geographically separated populations of the beachflea Orchestia gammarellus (Crustacea: Amphipoda)? J. Exp. Mar. Biol. Ecol. 1998, 229, 265–276. [Google Scholar] [CrossRef]
- Schmidt, C.; Heinz, P.; Kucera, M.; Uthicke, S. Temperature-induced stress leads to bleaching in larger benthic foraminifera hosting endosymbiotic diatoms. Limnol. Oceanogr. 2011, 56, 1587–1602. [Google Scholar] [CrossRef]
- Kelly, M.W.; Sanford, E.; Grosberg, R.K. Limited potential for adaptation to climate change in a broadly distributed marine crustacean. Proc. R. Soc. Biol. Sci. B 2012, 279, 349–356. [Google Scholar] [CrossRef]
- Sanford, E.; Holzman, S.B.; Haney, R.A.; Rand, D.M.; Bertness, M.D. Larval tolerance, gene flow, and the northern geographic range limit of fiddler crabs. Ecology 2006, 87, 2882–2894. [Google Scholar] [CrossRef] [PubMed]
- Lymbery, R.A.; Evans, J.P. Genetic variation underlies temperature tolerance of embryos in the sea urchin Heliocidaris erythrogramma armigera. J. Evol. Biol. 2013, 26, 2271–2282. [Google Scholar] [CrossRef] [PubMed]
- Osovitz, C.J.; Hofmann, G.E. Thermal history-dependent expression of the hsp70 gene in purple sea urchins: Biogeographic patterns and the effect of temperature acclimation. J. Exp. Mar. Biol. Ecol. 2005, 327, 134–143. [Google Scholar] [CrossRef]
- Kelly, M.W.; Padilla-Gamiño, J.L.; Hofmann, G.E. Natural variation and the capacity to adapt to ocean acidification in the keystone sea urchin Strongylocentrotus purpuratus. Glob. Change Biol. 2013, 19, 2536–2546. [Google Scholar] [CrossRef]
- Pecorino, D.; Barker, M.F.; Dworjanyn, S.A.; Byrne, M.; Lamare, M.D. Impacts of near future sea surface pH and temperature conditions on fertilisation and embryonic development in Centrostephanus rodgersii from northern New Zealand and northern New South Wales, Australia. Mar. Biol. 2014, 161, 101–110. [Google Scholar] [CrossRef]
- Marcil, J.; Swain, D.P.; Hutchings, J.A. Countergradient variation in body shape between two populations of Atlantic cod (Gadus morhua). Proc. R. Soc. Biol. Sci. B 2006, 273, 217–223. [Google Scholar] [CrossRef]
- Fangue, N.A.; Hofmeister, M.; Schulte, P.M. Intraspecific variation in thermal tolerance and heat shock protein gene expression in common killifish, Fundulus heteroclitus. J. Exp. Biol. 2006, 209, 2859–2872. [Google Scholar] [CrossRef] [PubMed]
- Schaum, E.; Rost, B.; Millar, A.J.; Collins, S. Variation in plastic responses of a globally distributed picoplankton species to ocean acidification. Nature Clim. Change 2012, 3, 298–302. [Google Scholar] [CrossRef]
- Pistevos, J.C.A.; Calosi, P.; Widdicombe, S.; Bishop, J.D.D. Will variation among genetic individuals influence species responses to global climate change? Oikos 2011, 120, 675–689. [Google Scholar] [CrossRef]
- Willi, Y.; Van Buskirk, J.; Hoffmann, A.A. Limits to the adaptive potential of small populations. Annu. Rev. Ecol. Evol. Syst. 2006, 37, 433–458. [Google Scholar] [CrossRef]
- Lande, R. Adaptation to an extraordinary environment by evolution of phenotypic plasticity and genetic assimilation. J. Evol. Biol. 2009, 22, 1435–1446. [Google Scholar] [CrossRef] [PubMed]
- Feely, R.A.; Sabine, C.L.; Hernandez-Ayon, J.M.; Ianson, D.; Hales, B. Evidence for upwelling of corrosive “acidified” water onto the continental shelf. Science 2008, 320, 1490–1492. [Google Scholar] [CrossRef] [PubMed]
- Evans, T.G.; Chan, F.; Menge, B.A.; Hofmann, G.E. Transcriptomic responses to ocean acidification in larval sea urchins from a naturally variable pH environment. Mol. Ecol. 2013, 22, 1609–1625. [Google Scholar] [CrossRef] [PubMed]
- Saderne, V.; Wahl, M. Differential responses of calcifying and non-calcifying epibionts of a brown macroalga to present-day and future upwelling pCO2. PLoS ONE 2013, 8, e70455. [Google Scholar] [PubMed]
- Leal, S.M. Genetics and Analysis of Quantitative Traits. Am. J. Hum. Genet. 2001, 68, 548–549. [Google Scholar] [CrossRef]
- Hughes, A.R.; Inouye, B.D.; Johnson, M.T.J.; Underwood, N.; Vellend, M. Ecological consequences of genetic diversity. Ecol. Lett. 2008, 11, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Pespeni, M.H.; Sanford, E.; Gaylord, B.; Hill, T.M.; Hosfelt, J.D.; Jaris, H.K.; LaVigne, M.; Lenz, E.A.; Russell, A.D.; Young, M.K.; et al. Evolutionary change during experimental ocean acidification. Proc. Natl. Acad. Sci. USA 2013, 110, 6937–6942. [Google Scholar] [CrossRef] [PubMed]
- Hedrick, P.W. Genetic Polymorphism in Heterogeneous Environments: The Age of Genomics. Annu. Rev. Ecol., Evol., Syst. 2006, 37, 67–93. [Google Scholar] [CrossRef]
- Intergovernmental Panel on Climate Change. Climate Change 2013—The Physical Science Basis; Cambridge University Press: Cambridge, UK, 2014. [Google Scholar]
- Thomsen, J.; Casties, I.; Pansch, C.; Körtzinger, A.; Melzner, F. Food availability outweighs ocean acidification effects in juvenile Mytilus edulis: Laboratory and field experiments. Glob. Change Biol. 2012, 19, 1017–1027. [Google Scholar] [CrossRef]
- Thomsen, J.; Gutowska, M.A.; Saphörster, J.; Heinemann, A.; Trübenbach, K.; Fietzke, J.; Hiebenthal, C.; Eisenhauer, A.; Körtzinger, A.; Wahl, M.; et al. Calcifying invertebrates succeed in a naturally CO2-rich coastal habitat but are threatened by high levels of future acidification. Biogeosciences 2010, 7, 3879–3891. [Google Scholar]
- Dam, H.G. Evolutionary adaptation of marine zooplankton to global change. Annu. Rev. Mar. Sci. 2013, 5, 349–370. [Google Scholar] [CrossRef]
- Conover, D.O.; Present, T.M.C. Countergradient variation in growth rate: Compensation for length of the growing season among Atlantic silversides from different latitudes. Oecologia 1990, 83, 316–324. [Google Scholar]
- Hairston, N.G.; Ellner, S.P.; Geber, M.A.; Yoshida, T.; Fox, J.A. Rapid evolution and the convergence of ecological and evolutionary time. Ecol. Lett. 2005, 8, 1114–1127. [Google Scholar] [CrossRef]
- Olsen, E.M.; Heino, M.; Lilly, G.R.; Morgan, M.J.; Brattey, J.; Ernande, B.; Dieckmann, U. Maturation trends indicative of rapid evolution preceded the collapse of northern cod. Nature 2004, 428, 932–935. [Google Scholar] [CrossRef] [PubMed]
- Schoener, T.W. The newest synthesis: understanding the interplay of evolutionary and ecological dynamics. Science 2011, 331, 426–429. [Google Scholar] [CrossRef] [PubMed]
- Palkovacs, E.P.; Marshall, M.C.; Lamphere, B.A.; Lynch, B.R.; Weese, D.J.; Fraser, D.F.; Reznick, D.N.; Pringle, C.M.; Kinnison, M.T. Experimental evaluation of evolution and coevolution as agents of ecosystem change in Trinidadian streams. Philos. Trans. R. Soc. B 2009, 364, 1617–1628. [Google Scholar] [CrossRef]
- Janzen, D.H. When is it coevolution. Evolution 1980, 34, 611–612. [Google Scholar] [CrossRef]
- Turcotte, M.M.; Corrin, M.S.; Johnson, M.T. Adaptive evolution in ecological communities. PLoS Biol. 2012, 10. [Google Scholar] [CrossRef] [PubMed]
- Strauss, S.Y.; Sahli, H.; Conner, J.K. Toward a more trait-centered approach to diffuse (co)evolution. New Phytol. 2005, 165, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Lai, S.-M.; Liu, W.-C.; Jordán, F. On the centrality and uniqueness of species from the network perspective. Biol. Lett. 2012, 8, 570–573. [Google Scholar] [CrossRef] [PubMed]
- Hale, R.; Calosi, P.; McNeill, L.; Mieszkowska, N.; Widdicombe, S. Predicted levels of future ocean acidification and temperature rise could alter community structure and biodiversity in marine benthic communities. Oikos 2011, 120, 661–674. [Google Scholar] [CrossRef]
- Diaz-Pulido, G.; Gouezo, M.; Tilbrook, B.; Dove, S.; Anthony, K. High CO2 enhances the competitive strength of seaweeds over corals. Ecol. Lett. 2011, 14, 156–162. [Google Scholar] [CrossRef]
- Ferrari, M.C.O.; McCormick, M.I.; Munday, P.L.; Meekan, M.G.; Dixson, D.L.; Lonnstedt, Ö.; Chivers, D.P. Putting prey and predator into the CO2 equation—Qualitative and quantitative effects of ocean acidification on predator–prey interactions. Ecol. Lett. 2011, 14, 1143–1148. [Google Scholar] [CrossRef] [PubMed]
- Sanford, E.; Gaylord, B.; Hettinger, A.; Lenz, E.A.; Meyer, K.; Hill, T.M. Ocean acidification increases the vulnerability of native oysters to predation by invasive snails. Proc. R. Soc. Biol. Sci. B 2014, 281. [Google Scholar] [CrossRef]
- O’Connor, M.I. Warming strengthens an herbivore-plant interaction. Ecology 2009, 90, 388–398. [Google Scholar] [CrossRef] [PubMed]
- Poore, A.G.B.; Graba-Landry, A.; Favret, M.; Sheppard Brennand, H.; Byrne, M.; Dworjanyn, S.A. Direct and indirect effects of ocean acidification and warming on a marine plant–herbivore interaction. Oecologia 2013, 173, 1113–1124. [Google Scholar] [CrossRef] [PubMed]
- Collins, S.; Rost, B.; Rynearson, T.A. Evolutionary potential of marine phytoplankton under ocean acidification. Evol. Appl. 2014, 7, 140–155. [Google Scholar] [CrossRef] [PubMed]
- Falkowski, P.G.; Fenchel, T.; Delong, E.F. The microbial engines that drive Earth’s biogeochemical cycles. Science 2008, 320, 1034–1039. [Google Scholar] [CrossRef] [PubMed]
- Iglesias-Rodriguez, M.; Halloran, P.; Rickaby, R.; Hall, I.; Colmenero-Hidalgo, E.; Gittins, J.; Green, D.; Tyrrell, T.; Gibbs, S.; von Dassow, P.; et al. Phytoplankton calcification in a high-CO2 world. Science 2008, 320, 336–340. [Google Scholar] [CrossRef] [PubMed]
- Harley, C.D.G.; Anderson, K.M.; Demes, K.W.; Jorve, J.P.; Kordas, R.L.; Coyle, T.A.; Graham, M.H. Effects of climate change on global seaweed communities. J. Phycol. 2012, 48, 1064–1078. [Google Scholar] [CrossRef]
- Hurd, C.L.; Hepburn, C.D.; Currie, K.I.; Raven, J.A.; Hunter, K.A. Testing the effects of ocean acidification on algal metabolism: Considerations for experimental designs. J. Phycol. 2009, 45, 1236–1251. [Google Scholar] [CrossRef]
- Tatters, A.O.; Schnetzer, A.; Fu, F.; Lie, A.Y.A.; Caron, D.A.; Hutchins, D.A. Short- versus long-term responses to changing CO2 in a coastal dinoflagellate bloom: Implications for interspecific competitive interactions and community structure. Evolution 2013, 67, 1879–1891. [Google Scholar] [CrossRef] [PubMed]
- Kremp, A.; Godhe, A.; Egardt, J.; Dupont, S.; Suikkanen, S.; Casabianca, S.; Penna, A. Intraspecific variability in the response of bloom-forming marine microalgae to changed climate conditions. Ecol. Evol. 2012, 2, 1195–1207. [Google Scholar] [CrossRef] [PubMed]
- Boyd, P.W.; Rynearson, T.A.; Armstrong, E.A.; Fu, F.; Hayashi, K.; Hu, Z.; Hutchins, D.A.; Kudela, R.M.; Litchman, E.; Mulholland, M.R.; et al. Marine phytoplankton temperature versus growth responses from polar to tropical waters—Outcome of a scientific community-wide study. PLoS ONE 2013, 8, e63091. [Google Scholar] [CrossRef]
- Alsterberg, C.; Eklöf, J.S.; Gamfeldt, L.; Havenhand, J.N.; Sundbäck, K. Consumers mediate the effects of experimental ocean acidification and warming on primary producers. Proc. Natl. Acad. Sci. USA 2013, 110, 8603–8608. [Google Scholar] [CrossRef] [PubMed]
- Witt, V.; Wild, C.; Anthony, K.R.N.; Diaz-Pulido, G.; Uthicke, S. Effects of ocean acidification on microbial community composition of, and oxygen fluxes through, biofilms from the Great Barrier Reef. Environ. Microbiol. 2011, 13, 2976–2989. [Google Scholar] [CrossRef] [PubMed]
- Beaugrand, G.; Brander, K.M.; Alistair Lindley, J.; Souissi, S.; Reid, P.C. Plankton effect on cod recruitment in the North Sea. Nature 2003, 426, 661–664. [Google Scholar] [CrossRef] [PubMed]
- Melzner, F.; Stange, P.; Trübenbach, K.; Thomsen, J.; Casties, I.; Panknin, U.; Gorb, S.N.; Gutowska, M.A. Food supply and seawater pCO2 impact calcification and internal shell dissolution in the blue mussel Mytilus edulis. PLoS ONE 2011, 6, e24223. [Google Scholar] [CrossRef]
- Calosi, P.; Rastrick, S.P.S.; Graziano, M.; Thomas, S.C.; Baggini, C.; Carter, H.A.; Hall-Spencer, J.M.; Milazzo, M.; Spicer, J.I. Distribution of sea urchins living near shallow water CO2 vents is dependent upon species acid-base and ion-regulatory abilities. Mar. Pollut. Bull. 2013, 73, 470–484. [Google Scholar] [CrossRef] [PubMed]
- Connell, S.D.; Kroeker, K.J.; Fabricius, K.E.; Kline, D.I.; Russell, B.D. The other ocean acidification problem: CO2 as a resource among competitors for ecosystem dominance. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368. [Google Scholar] [CrossRef] [PubMed]
- Kroeker, K.J.; Micheli, F.; Gambi, M.C. Ocean acidification causes ecosystem shifts via altered competitive interactions. Nat. Clim. Change 2012, 3, 156–159. [Google Scholar] [CrossRef]
- Connell, S.; Russell, B. The direct effects of increasing CO2 and temperature on non-calcifying organisms: increasing the potential for phase shifts in kelp forests. Proc. R. Soc. Biol. Sci. B 2010, 277, 1409–1415. [Google Scholar] [CrossRef]
- de Mazancourt, C.; Johnson, E.; Barraclough, T.G. Biodiversity inhibits species’ evolutionary responses to changing environments. Ecol. Lett. 2008, 11, 380–388. [Google Scholar] [CrossRef]
- Johansson, J. Evolutionary responses to climate changes: how does competition affect adaptation? Evolution 2008, 62, 421–435. [Google Scholar] [CrossRef] [PubMed]
- Fabricius, K.E.; Langdon, C.; Uthicke, S.; Humphrey, C.; Noonan, S.; De’ath, G.; Okazaki, R.; Muehllehner, N.; Glas, M.S.; Lough, J.M. Losers and winners in coral reefs acclimatized to elevated carbon dioxide concentrations. Nat. Clim. Change 2011, 1, 165–169. [Google Scholar] [CrossRef]
- Dunn, R.R.; Harris, N.C.; Colwell, R.K.; Koh, L.P.; Sodhi, N.S. The sixth mass coextinction: Are most endangered species parasites and mutualists? Proc. R. Soc. Biol. Sci. B 2009, 276, 3037–3045. [Google Scholar] [CrossRef]
- Stenseth, N.C.; Smith, J.M. Coevolution in ecosystems: Red queen evolution or stasis? Evolution 1984, 38, 870–880. [Google Scholar] [CrossRef]
- Lawrence, D.; Fiegna, F.; Behrends, V.; Bundy, J.G.; Phillimore, A.B.; Bell, T.; Barraclough, T.G. Species interactions alter evolutionary responses to a novel environment. PLoS Biol. 2012, 10. [Google Scholar] [CrossRef]
- Chesson, P. A need for niches? Trends Ecol. Evol. 1991, 6, 26–28. [Google Scholar] [CrossRef] [PubMed]
- Byrnes, J.E.; Gamfeldt, L.; Isbell, F.; Lefcheck, J.S.; Griffin, J.N.; Hector, A.; Cardinale, B.J.; Hooper, D.U.; Dee, L.E.; Emmett Duffy, J. Investigating the relationship between biodiversity and ecosystem multifunctionality: Challenges and solutions. Meth. Ecol. Evol. 2014, 5, 111–124. [Google Scholar] [CrossRef]
- Pereira, H.M.; Leadley, P.W.; Proença, V.; Alkemade, R.; Scharlemann, J.P.W.; Fernandez-Manjarrés, J.F.; Araújo, M.B.; Balvanera, P.; Biggs, R.; Cheung, W.W.L.; et al. Scenarios for global biodiversity in the 21st century. Science 2010, 330, 1496–1501. [Google Scholar] [CrossRef] [PubMed]
- Burrows, M.T.; Schoeman, D.S.; Richardson, A.J.; Molinos, J.G.; Hoffmann, A.; Buckley, L.B.; Moore, P.J.; Brown, C.J.; Bruno, J.F.; Duarte, C.M.; et al. Geographical limits to species-range shifts are suggested by climate velocity. Nature 2014, 507, 492–495. [Google Scholar] [CrossRef] [PubMed]
- Pespeni, M.H.; Chan, F.; Menge, B.A.; Palumbi, S.R. Signs of adaptation to local pH conditions across an environmental mosaic in the California current ecosystem. Integr. Comp. Biol. 2013, 53, 857–870. [Google Scholar] [CrossRef]
- Ricevuto, E.; Kroeker, K.J.; Ferrigno, F.; Micheli, F.; Gambi, M.C. Spatio-temporal variability of polychaete colonization at CO2 vents indicates high tolerance to ocean acidification. Mar. Biol. 2014, 161, 2909–2919. [Google Scholar] [CrossRef]
- Hughes, T.P.; Baird, A.H.; Bellwood, D.R.; Card, M.; Connolly, S.R.; Folke, C.; Grosberg, R.; Hoegh-Guldberg, O.; Jackson, J.B.C.; Kleypas, J.; et al. Climate change, human impacts, and the resilience of coral reefs. Science 2003, 301, 929–933. [Google Scholar] [CrossRef] [PubMed]
- Bellantuono, A.J.; Hoegh-Guldberg, O.; Rodriguez-Lanetty, M. Resistance to thermal stress in corals without changes in symbiont composition. Proc. R. Soc. Biol. Sci. B 2012, 279, 1100–1107. [Google Scholar] [CrossRef]
- Barshis, D.; Ladner, J.; Oliver, T.; Palumbi, S.R. Lineage specific transcriptional profiles of Symbiodinium spp. unaltered by heat stress in a coral host. Mol. Biol. Evol. 2014, 31, 1343–1352. [Google Scholar] [CrossRef] [PubMed]
- Benton, M.J. The Red Queen and the Court Jester: Species diversity and the role of biotic and abiotic factors through time. Science 2009, 323, 728–732. [Google Scholar] [CrossRef] [PubMed]
- Drake, J.M.; Lodge, D.M. Hull fouling is a risk factor for intercontinental species exchange in aquatic ecosystems. Aq. Inv. 2007, 2, 121–131. [Google Scholar] [CrossRef]
- Walther, G.-R.; Roques, A.; Hulme, P.E.; Sykes, M.T.; Pysek, P.; Kühn, I.; Zobel, M.; Bacher, S.; Botta-Dukát, Z.; Bugmann, H.; et al. Alien species in a warmer world: Risks and opportunities. Trends Ecol. Evol. 2009, 24, 686–693. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, C.N. Biodiversity issues for the forthcoming tropical Mediterranean Sea. Hydrobiologia 2007, 580, 7–21. [Google Scholar] [CrossRef]
- Irwin, A.J.; Finkel, Z.V.; Schofield, O.M.E.; Falkowski, P.G. Scaling-up from nutrient physiology to the size-structure of phytoplankton communities. J. Plankton Res. 2006, 28, 459–471. [Google Scholar] [CrossRef]
- Levine, J.M.; Vilà, M.; Antonio, C.M.D.; Dukes, J.S.; Grigulis, K.; Lavorel, S. Mechanisms underlying the impacts of exotic plant invasions. Proc. R. Soc. Biol. Sci. B 2003, 270, 775–781. [Google Scholar] [CrossRef]
- Schluter, D. Experimental evidence that competition promotes divergence in adaptive radiation. Science 1994, 266, 798–801. [Google Scholar] [CrossRef] [PubMed]
- Halpern, B.S.; Selkoe, K.A.; Micheli, F.; Kappel, C.V. Evaluating and ranking the vulnerability of global marine ecosystems to anthropogenic threats. Conserv. Biol. 2007, 21, 1301–1315. [Google Scholar] [CrossRef] [PubMed]
- Barton, A.D.; Pershing, A.J.; Litchman, E.; Record, N.R.; Edwards, K.F.; Finkel, Z.V.; Kiørboe, T.; Ward, B.A. The biogeography of marine plankton traits. Ecol. Lett. 2013, 16, 522–534. [Google Scholar] [CrossRef] [PubMed]
- Reusch, T.B.H.; Boyd, P.W. Experimental evolution meets marine phytoplankton. Evolution 2013, 67, 1849–1859. [Google Scholar] [CrossRef]
- Russell, B.D.; Wernberg, T.; Mieszkowska, N.; Widdicombe, S.; Hall-Spencer, J.; Connell, S. Predicting ecosystem shifts requires new approaches that integrate the effects of climate change across entire systems. Biol. Lett. 2012, 8, 164–166. [Google Scholar] [CrossRef] [PubMed]
- Hendriks, I.E.; Duarte, C.M. Ocean acidification: Separating evidence from judgment—A reply to Dupont et al. Estuar. Coast. Shelf Sci. 2010, 89, 186–190. [Google Scholar] [CrossRef]
- Gaylord, B.; Hill, T.M.; Sanford, E.; Lenz, E.A.; Jacobs, L.A.; Sato, K.N.; Russell, A.D.; Hettinger, A. Functional impacts of ocean acidification in an ecologically critical foundation species. J. Exp. Biol. 2011, 214, 2586–2594. [Google Scholar] [CrossRef] [PubMed]
- Vihtakari, M.; Hendriks, I.; Holding, J.; Renaud, P.; Duarte, C.; Havenhand, J. Effects of ocean acidification and warming on sperm activity and early life stages of the Mediterranean mussel (Mytilus galloprovincialis). Water 2013, 5, 1890–1915. [Google Scholar] [CrossRef]
- Van Valen, L. A new evolutionary law. Evol. Theor. 1973, 1, 1–30. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harvey, B.P.; Al-Janabi, B.; Broszeit, S.; Cioffi, R.; Kumar, A.; Aranguren-Gassis, M.; Bailey, A.; Green, L.; Gsottbauer, C.M.; Hall, E.F.; et al. Evolution of Marine Organisms under Climate Change at Different Levels of Biological Organisation. Water 2014, 6, 3545-3574. https://doi.org/10.3390/w6113545
Harvey BP, Al-Janabi B, Broszeit S, Cioffi R, Kumar A, Aranguren-Gassis M, Bailey A, Green L, Gsottbauer CM, Hall EF, et al. Evolution of Marine Organisms under Climate Change at Different Levels of Biological Organisation. Water. 2014; 6(11):3545-3574. https://doi.org/10.3390/w6113545
Chicago/Turabian StyleHarvey, Ben P., Balsam Al-Janabi, Stefanie Broszeit, Rebekah Cioffi, Amit Kumar, Maria Aranguren-Gassis, Allison Bailey, Leon Green, Carina M. Gsottbauer, Emilie F. Hall, and et al. 2014. "Evolution of Marine Organisms under Climate Change at Different Levels of Biological Organisation" Water 6, no. 11: 3545-3574. https://doi.org/10.3390/w6113545
APA StyleHarvey, B. P., Al-Janabi, B., Broszeit, S., Cioffi, R., Kumar, A., Aranguren-Gassis, M., Bailey, A., Green, L., Gsottbauer, C. M., Hall, E. F., Lechler, M., Mancuso, F. P., Pereira, C. O., Ricevuto, E., Schram, J. B., Stapp, L. S., Stenberg, S., & Rosa, L. T. S. (2014). Evolution of Marine Organisms under Climate Change at Different Levels of Biological Organisation. Water, 6(11), 3545-3574. https://doi.org/10.3390/w6113545





