Structure-Based Long-Term Biodegradation of the Azo Dye: Insights from the Bacterial Community Succession and Efficiency Comparison
Abstract
:1. Introduction
2. Materials and Methods
2.1. Activated Sludge
2.2. Dyes and Reagents
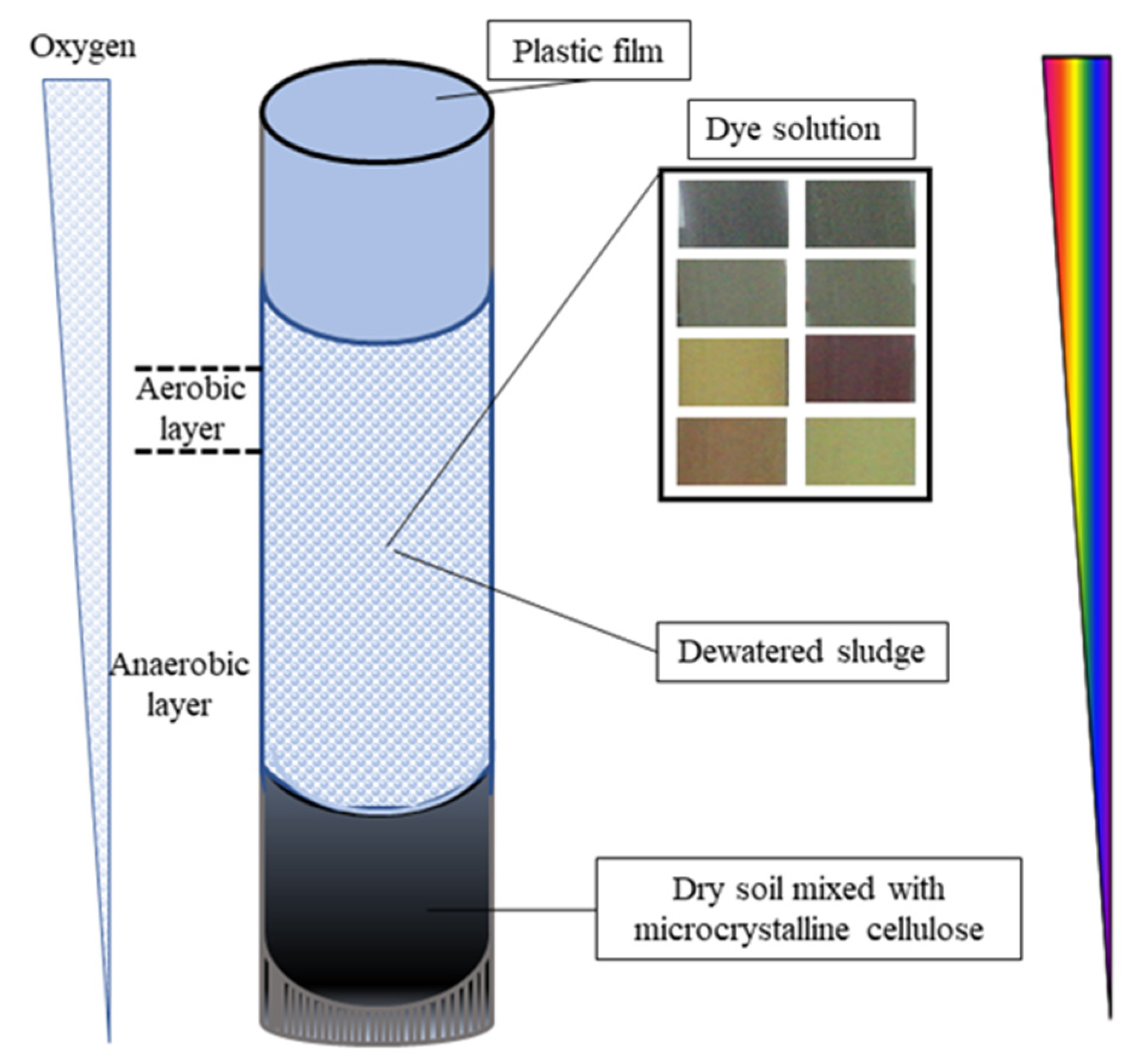
2.3. Sludge Acclimatization
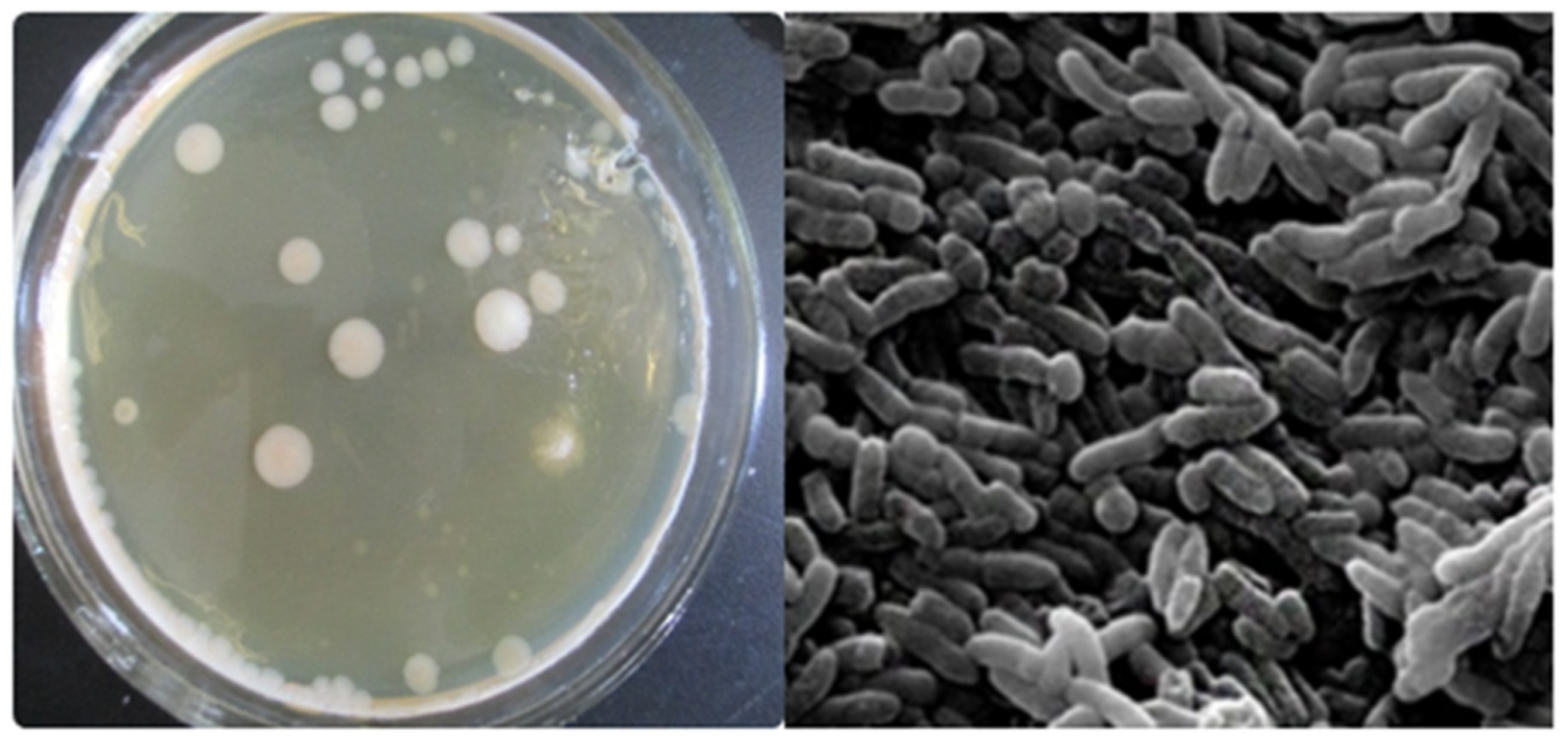
2.4. Strain Isolation
2.5. Strain Identification
2.6. Decolorization Test of Isolated Strains
2.7. T-RFLP Analysis of Acclimated Sludge
2.8. CCA Analysis
3. Results and Discussion
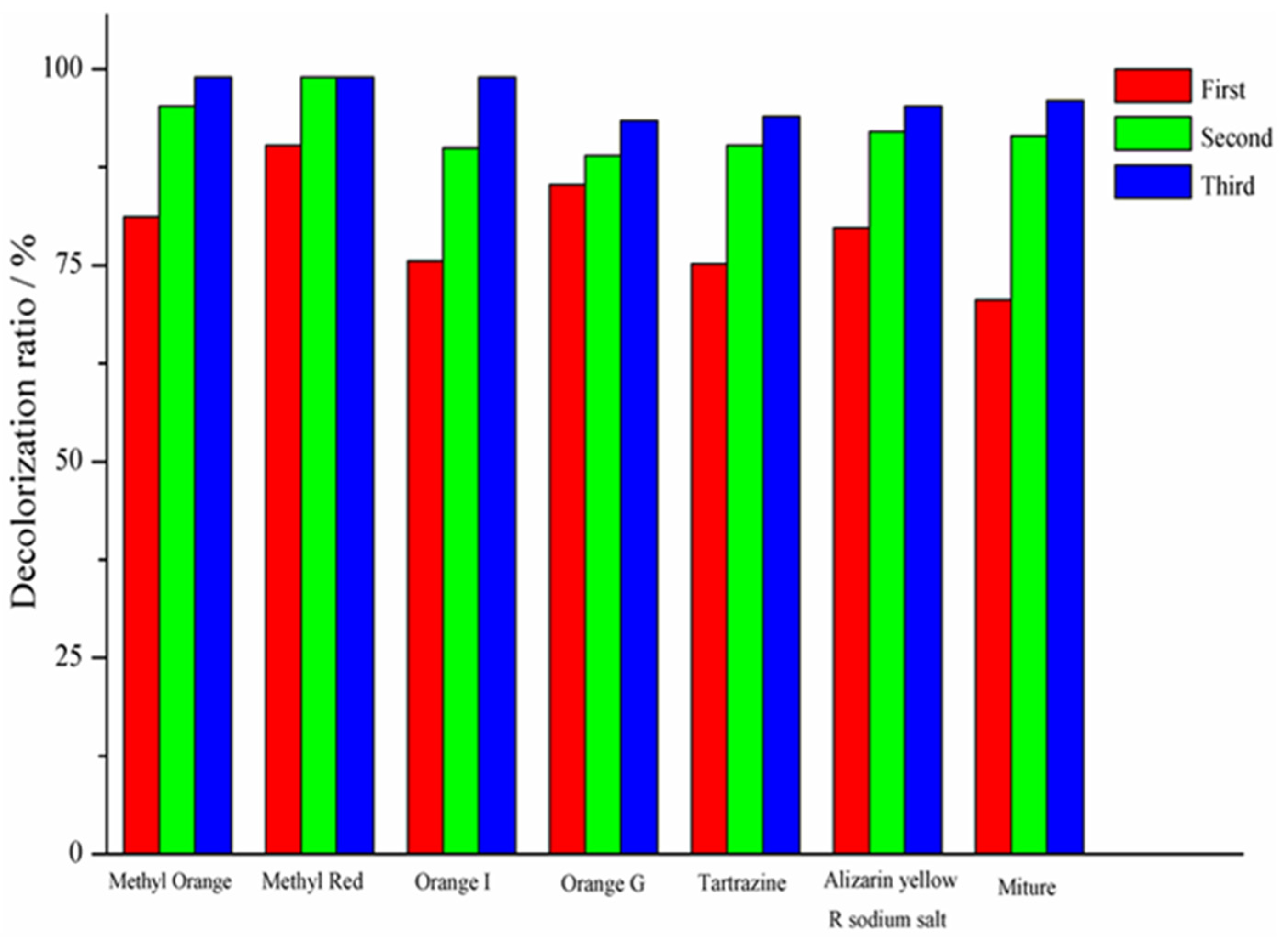
3.1. Decolorization of Azo Dyes
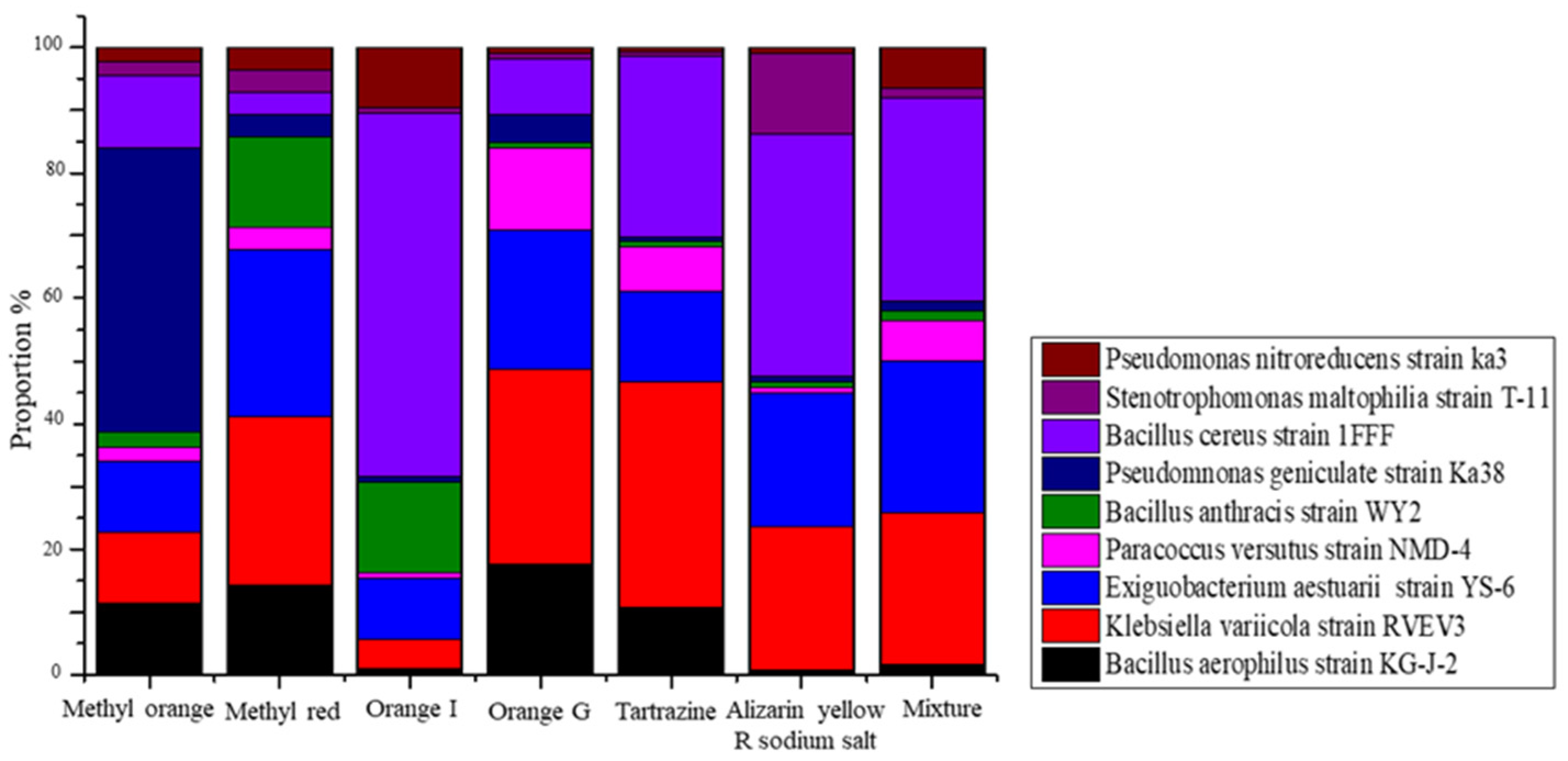
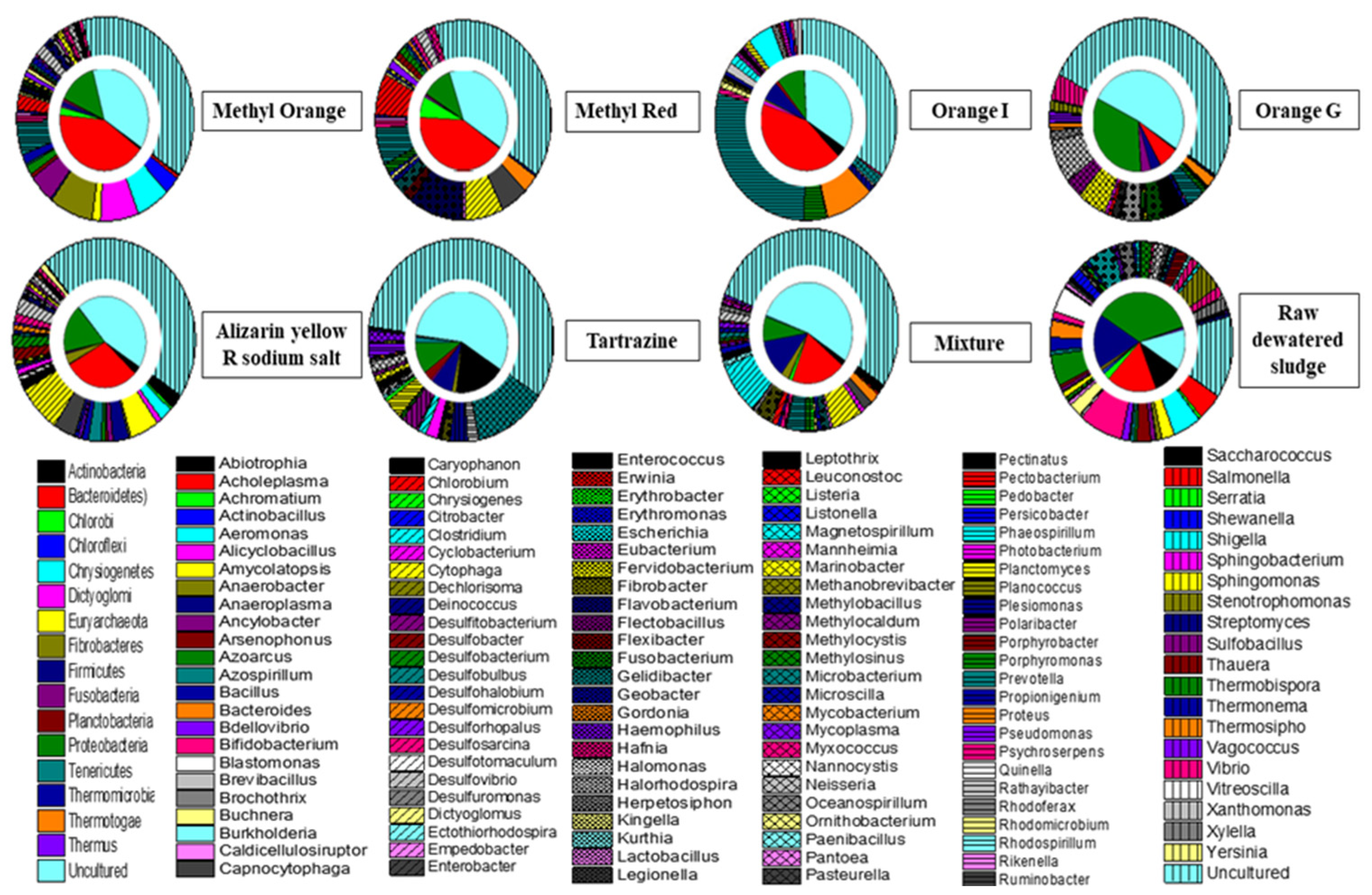
3.2. Cultured Bacterial Composition of Azo Dye in Acclimatized Sludge
3.3. Taxonomical and Functional Evaluation of the Acclimatized Sludge Based on T-RFLP Analysis
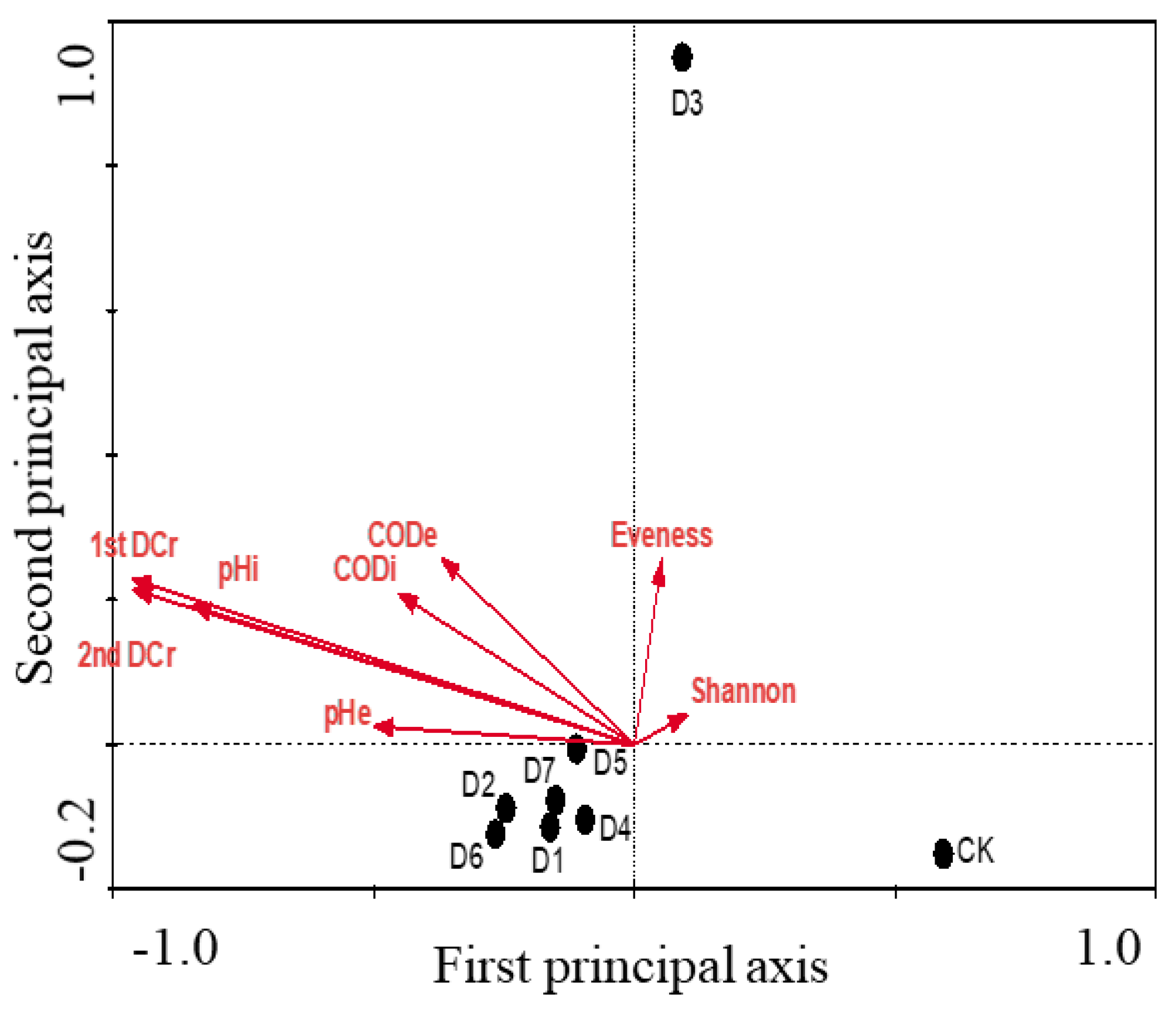
3.4. CCA Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Saratale, R.G.; Saratale, G.D.; Kalyani, D.C.; Chang, J.S.; Govindwar, S.P. Enhanced decolorization and biodegradation of textile azo dye Scarlet R by using developed microbial consortium-GR. Bioresour. Technol. 2009, 100, 2493–2500. [Google Scholar] [CrossRef] [PubMed]
- Rawat, D.; Sharma, R.S.; Karmakar, S.; Arora, L.S.; Mishra, V. Ecotoxic potential of a presumably non-toxic azo dye. Ecotoxicol Environ. Saf. 2018, 148, 528–537. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Wang, H.; Zhang, S.; Nishimura, O.; Li, X. Azo dye degradation pathway and bacterial community structure in biofilm electrode reactors. Chemosphere 2018, 208, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Koyani, R.D.; Sanghvi, G.V.; Sharma, R.K.; Rajput, K.S. Contribution of lignin degrading enzymes in decolourisation and degradation of reactive textile dyes. Int. Biodeterior. Biodegrad. 2013, 77, 1–9. [Google Scholar] [CrossRef]
- Waghmode, T.R.; Kurade, M.B.; Sapkal, R.T.; Bhosale, C.H.; Jeon, B.H.; Govindwar, S.P. Sequential photocatalysis and biological treatment for the enhanced degradation of the persistent azo dye methyl red. J. Hazard Mater. 2019, 371, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Liu, G.; Zhou, J.; Shiang Fu, Q.; Wang, G. Azo dye decolorization by Shewanella aquimarina under saline conditions. Bioresour. Technol. 2012, 114, 95–101. [Google Scholar] [CrossRef]
- Cui, D.; Cui, M.H.; Liang, B.; Liu, W.Z.; Tang, Z.E.; Wang, A.J. Mutual effect between electrochemically active bacteria (EAB) and azo dye in bio-electrochemical system (BES). Chemosphere 2020, 239, 124787. [Google Scholar] [CrossRef] [PubMed]
- Mezohegyi, G.; Bengoa, C.; Stuber, F.; Font, J.; Fabregat, A.; Fortuny, A. Novel bioreactor design for decolorization of azo dye effluents. Chem. Eng. J. 2008, 143, 293–298. [Google Scholar] [CrossRef]
- Balapure, K.H.; Jain, K.; Chattaraj, S.; Bhatt, N.S.; Madamwar, D. Co-metabolic degradation of diazo dye—Reactive blue 160 by enriched mixed cultures BDN. J. Hazard. Mater. 2014, 279, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Zhang, X.Y.; Wang, S.; Tang, Q.W.; Xie, T.; Lei, N.Y.; Lam, M.H.W. Microbial community structure associated with treatment of azo dye in a start-up anaerobic sequenced batch reactor. J. Taiwan Inst. Chem. Eng. 2015, 54, 118–124. [Google Scholar] [CrossRef]
- Ye, D.D.; Liang, H.B.; Zhou, W.; Yan, J.W.; Zhou, S.Q.; Luo, L.X. Total and and active microbial communities in a full-scale system treating wastewater from soy sauce production. Int. Biodeterior. Biodegrad. 2017, 123, 206–215. [Google Scholar] [CrossRef]
- Xie, X.H.; Liu, N.; Yang, B.; Yu, C.Z.; Zhang, Q.Y.; Zheng, X.L.; Xu, L.Y.; Li, R.; Liu, J.S. Comparison of microbial community in hydrolysis acidification reactor depending on different structure dyes by Illumina MiSeq sequencing. Int. Biodeterior. Biodegrad. 2016, 111, 14–21. [Google Scholar] [CrossRef]
- Köchling, T.; Ferraz, A.D.N., Jr.; Florencio, L.; Kato, M.T.; Gavazza, S. 454-Pyrosequencing analysis of highly adapted azo dye-degrading microbial communities in a two-stage anaerobic–aerobic bioreactor treating textile effluent. Environ. Technol. 2017, 38, 687–693. [Google Scholar] [CrossRef]
- Hsueh, C.C.; Chen, B.Y.; Yen, C.Y. Understanding effects of chemical structure on azo dye decolorization characteristics by Aeromonas hydrophila. J. Hazard Mater. 2009, 167, 995–1001. [Google Scholar] [CrossRef] [PubMed]
- Ayed, L.; Mandhi, A.; Cheref, A.; Bakhrouf, A. Decolorization and degradation of azo dye Methyl Red by an isolated Sphingomonas paucimobilis: Biotoxicity and metabolites characterization. Desalination 2011, 274, 272–277. [Google Scholar] [CrossRef]
- Jadhav, J.P.; Kalyani, D.C.; Telke, A.A.; Phugare, S.S.; Govindwar, S.P. Evaluation of the efficacy of a bacterial consortium for the removal of color, reduction of heavy metals, and toxicity from textile dye effluent. Bioresour. Technol. 2010, 101, 165–173. [Google Scholar] [CrossRef]
- Solis, M.; Solis, A.; Perez, H.I.; Manjarrez, N.; Flores, M. Microbial decolouration of azo dyes: A review. Process Biochem. 2012, 47, 1723–1748. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, H.; Liu, P.; Hu, H.; Yao, H. Pyrolysis characteristics and kinetic analysis of different dewatered sludge. Bioresour. Technol. 2014, 170, 325–330. [Google Scholar] [CrossRef]
- Zhu, C.; Wang, H.; Mahmood, Z.; Wang, Q.; Ma, H. Biocompatibility and biodegradability of polyacrylate/ZnO nanocomposite during the activated sludge treatment process. PLoS ONE 2018, 13, e0205990. [Google Scholar] [CrossRef]
- Holt, J.; Krieg, N.; Sneath, P.; Staley, J.; Williams, S. Family Enterobacteriaceae. Bergey’s Manual of Determinative Bacteriology, 9th ed.; Hensyl, W.R., Ed.; Lippincott Williams & Wilkins: Baltimore, MD, USA, 1994; pp. 175–194. [Google Scholar]
- Barinova, S.; Tavassi, M. Study of seasonal influences on algal biodiversity in the River Yarqon (central Israel) by bio-indication and canonical correspondence analysis (CCA). Turk. J. Bot. 2009, 33, 353–372. [Google Scholar] [CrossRef]
- Tan, L.; Qu, Y.; Zhou, J.; Li, A.; Zhang, H.; Su, Z. Microbial community dynamics in SBR for azo dyes treatment by PCR-DGGE technique. In Proceedings of the 13th International Biotechnology Symposium 2008, Dalian, China, 12–17 October 2008; p. S609. [Google Scholar]
- Georgiou, D.; Metallinou, C.; Aivasidis, A.; Voudrias, E.; Gimouhopoulos, K. Decolorization of azo-reactive dyes and cotton-textile wastewater using anaerobic digestion and acetate-consuming bacteria. Biochem. Eng. J. 2004, 19, 75–79. [Google Scholar] [CrossRef]
- Benkhaya, S.; M’Rabet, S.; El Harfi, A. Classifications, properties, recent synthesis and applications of azo dyes. Heliyon 2020, 6, e03271. [Google Scholar] [CrossRef] [Green Version]
- Chenini, H.; Djebbar, K.; Zendaoui, S. Removal of an Azo Dye (Orange G) by Various Methods in Homogenious Phase: Comparative Study. Jordan J. Chem. 2011, 146, 1–13. [Google Scholar]
- Hamous, H.; Khenifi, A.; Bouberka, Z.; Derriche, Z. Electrochemical degradation of Orange G in K2SO4 and KCl medium. Environ. Eng. Res. 2020, 25, 571–578. [Google Scholar] [CrossRef] [Green Version]
- Kai-Hui, Z.T.-Q.L.; ZHANG, J. Microbial Ecology Principle of Activated Sludge Acclimation. Microbiology 2008, 6. [Google Scholar]
- Lade, H.; Kadam, A.; Paul, D.; Govindwar, S. Biodegradation and detoxification of textile azo dyes by bacterial consortium under sequential microaerophilic/aerobic processes. EXCLI J. 2015, 14, 158–174. [Google Scholar] [CrossRef] [PubMed]
- Yoshie, S.; Makino, H.; Hirosawa, H.; Shirotani, K.; Tsuneda, S.; Hirata, A. Molecular analysis of halophilic bacterial community for high-rate denitrification of saline industrial wastewater. Appl. Microbiol. Biotechnol. 2006, 72, 182–189. [Google Scholar] [CrossRef]
- Kadam, S.K.; Chandanshive, V.V.; Rane, N.R.; Patil, S.M.; Gholave, A.R.; Khandare, R.V.; Bhosale, A.R.; Jeon, B.H.; Govindwar, S.P. Phytobeds with Fimbristylis dichotoma and Ammannia baccifera for treatment of real textile effluent: An in situ treatment, anatomical studies and toxicity evaluation. Environ. Res. 2018, 160, 1–11. [Google Scholar] [CrossRef]
- Fernando, E.; Keshavarz, T.; Kyazze, G. External resistance as a potential tool for influencing azo dye reductive decolourisation kinetics in microbial fuel cells. Int. Biodeterior. Biodegrad. 2014, 89, 7–14. [Google Scholar] [CrossRef]
- Su, Y.; Zhang, Y.; Wang, J.; Zhou, J.; Lu, X.; Lu, H. Enhanced bio-decolorization of azo dyes by co-immobilized quinone-reducing consortium and anthraquinone. Bioresour. Technol. 2009, 100, 2982–2987. [Google Scholar] [CrossRef]
- Sarkar, S.; Banerjee, A.; Halder, U.; Biswas, R.; Bandopadhyay, R. Degradation of synthetic azo dyes of textile industry: A sustainable approach using microbial enzymes. Water Conserv. Sci. Eng. 2017, 2, 121–131. [Google Scholar] [CrossRef] [Green Version]
- Rathod, J.; Archana, G. Molecular fingerprinting of bacterial communities in enriched azo dye (Reactive Violet 5R) decolorising native acclimatised bacterial consortia. Bioresour. Technol. 2013, 142, 436–444. [Google Scholar] [CrossRef]
- Cui, D.; Zhang, H.; He, R.; Zhao, M. The Comparative Study on the Rapid Decolorization of Azo, Anthraquinone and Triphenylmethane Dyes by Anaerobic Sludge. Int. J. Environ. Res. Public Health 2016, 13, 1053. [Google Scholar] [CrossRef] [PubMed]
- Shabbir, S.; Faheem, M.; Ali, N.; Kerr, P.G.; Wu, Y. Evaluating role of immobilized periphyton in bioremediation of azo dye amaranth. Bioresour. Technol. 2017, 225, 395–401. [Google Scholar] [CrossRef]
- Brander, S.; Jørn, D.M.; Kasper, P.K. Characterization of an alkali-and halide-resistant laccase expressed in E. coli: CotA from Bacillus clausii. PLoS ONE 2014, 9, e99402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuberan, T.; Anburaj, J.; Sundaravadivelan, C.; Kumar, P. Biodegradation of azo dye by Listeria sp. Int. J. Environ. Sci. 2011, 1, 1760–1769. [Google Scholar]
- Onat, T.A.; Gumusdere, H.T.; Guvenc, A.; Donmez, G.; Mehmetoglu, U. Decolorization of textile azo dyes by ultrasonication and microbial removal. Desalination 2010, 255, 154–158. [Google Scholar] [CrossRef]
- Hong, Y.G.; Gu, J.D. Physiology and biochemistry of reduction of azo compounds by Shewanella strains relevant to electron transport chain. Appl. Microbiol. Biotechnol. 2010, 88, 637–643. [Google Scholar] [CrossRef] [Green Version]
- Saratale, G.D.; Chien, L.-J.; Chang, J.S. Enzymatic treatment of lignocellulosic wastes for anaerobic digestion and bioenergy production. In Environmental Anaerobic Technology: Applications and New Developments; World Scientific: Singapore, 2010; pp. 279–308. [Google Scholar]
- Bodaghabadi, M.B.; Salehi, M.H.; Martinez-Casasnovas, J.A.; Mohammadi, J.; Toomanian, N.; Borujeni, I.E. Using Canonical Correspondence Analysis (CCA) to identify the most important DEM attributes for digital soil mapping applications. Catena 2011, 86, 66–74. [Google Scholar] [CrossRef]
- Ali, L.; Algaithi, R.; Habib, H.M.; Souka, U.; Rauf, M.A.; Ashraf, S.S. Soybean peroxidase-mediated degradation of an azo dye–a detailed mechanistic study. BMC Biochem. 2013, 14, 35. [Google Scholar] [CrossRef] [Green Version]
- Miao, Y.; Wang, Z.; Liao, R.H.; Shi, P.; Li, A.M. Assessment of phenol effect on microbial community structure and function in an anaerobic denitrifying process treating high concentration nitrate wastewater. Chem. Eng. J. 2017, 330, 757–763. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, C.; Mahmood, Z.; Siddique, M.S.; Wang, H.; Anqi, H.; Sillanpää, M. Structure-Based Long-Term Biodegradation of the Azo Dye: Insights from the Bacterial Community Succession and Efficiency Comparison. Water 2021, 13, 3017. https://doi.org/10.3390/w13213017
Zhu C, Mahmood Z, Siddique MS, Wang H, Anqi H, Sillanpää M. Structure-Based Long-Term Biodegradation of the Azo Dye: Insights from the Bacterial Community Succession and Efficiency Comparison. Water. 2021; 13(21):3017. https://doi.org/10.3390/w13213017
Chicago/Turabian StyleZhu, Chao, Zarak Mahmood, Muhammad Saboor Siddique, Heyou Wang, He Anqi, and Mika Sillanpää. 2021. "Structure-Based Long-Term Biodegradation of the Azo Dye: Insights from the Bacterial Community Succession and Efficiency Comparison" Water 13, no. 21: 3017. https://doi.org/10.3390/w13213017






