Succession of Microbial Community in a Small Water Body within the Alluvial Aquifer of a Large River
Abstract
1. Introduction
2. Materials and Methods
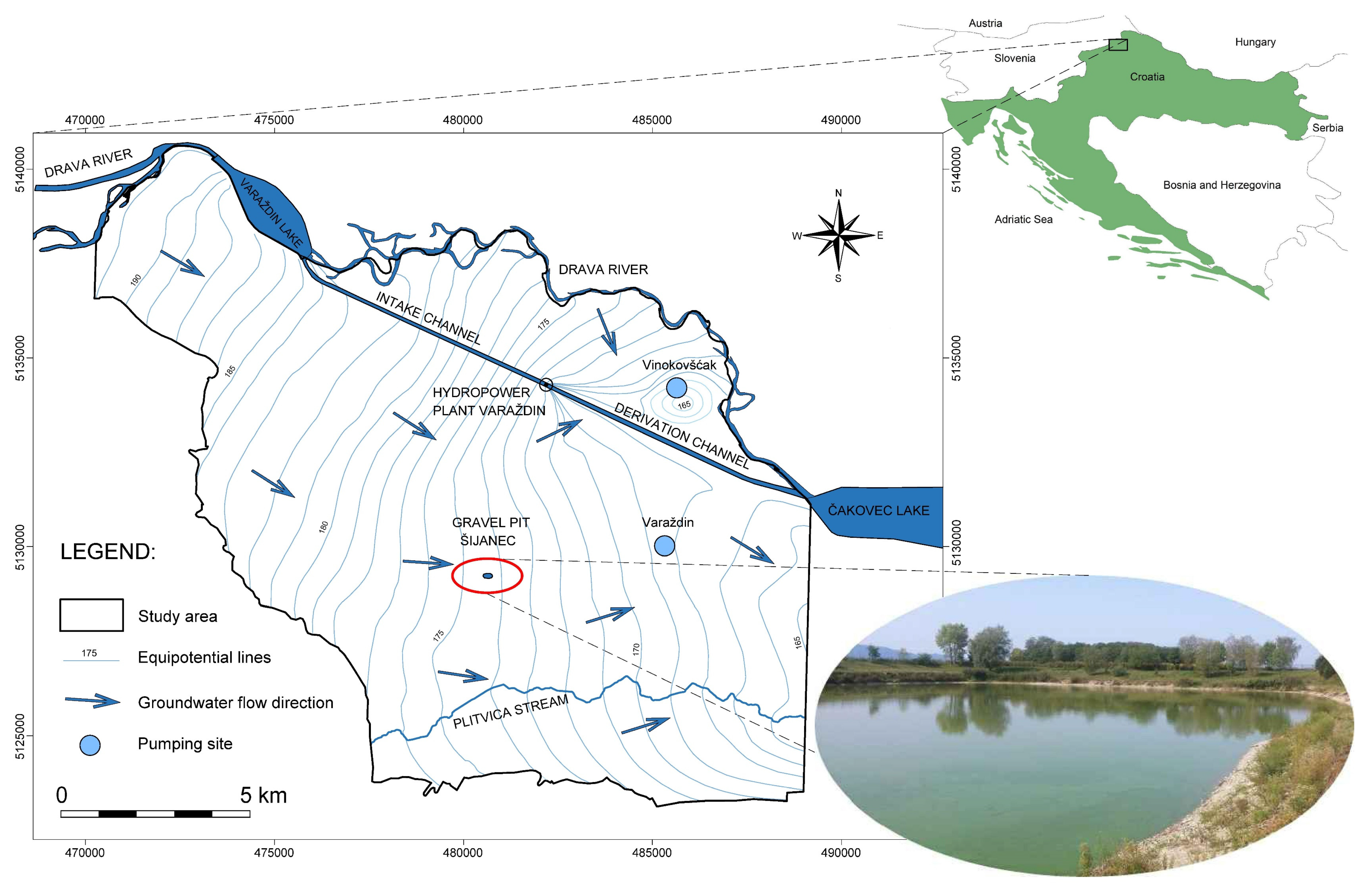
2.1. Study Area
2.2. Sampling and Laboratory Analysis
2.3. Microbial Community Analysis
2.4. PCR of the Phytoplankton (Eukaryotic) Community
2.5. PCR for the Bacterial Community
2.6. Bioinformatic Analysis of the Phytoplankton (Eukaryotic) Community
2.7. Bioinformatic Analysis of the Bacterial Community
2.8. Statistical Analysis
3. Results
3.1. Analysis of Environmental Parameters
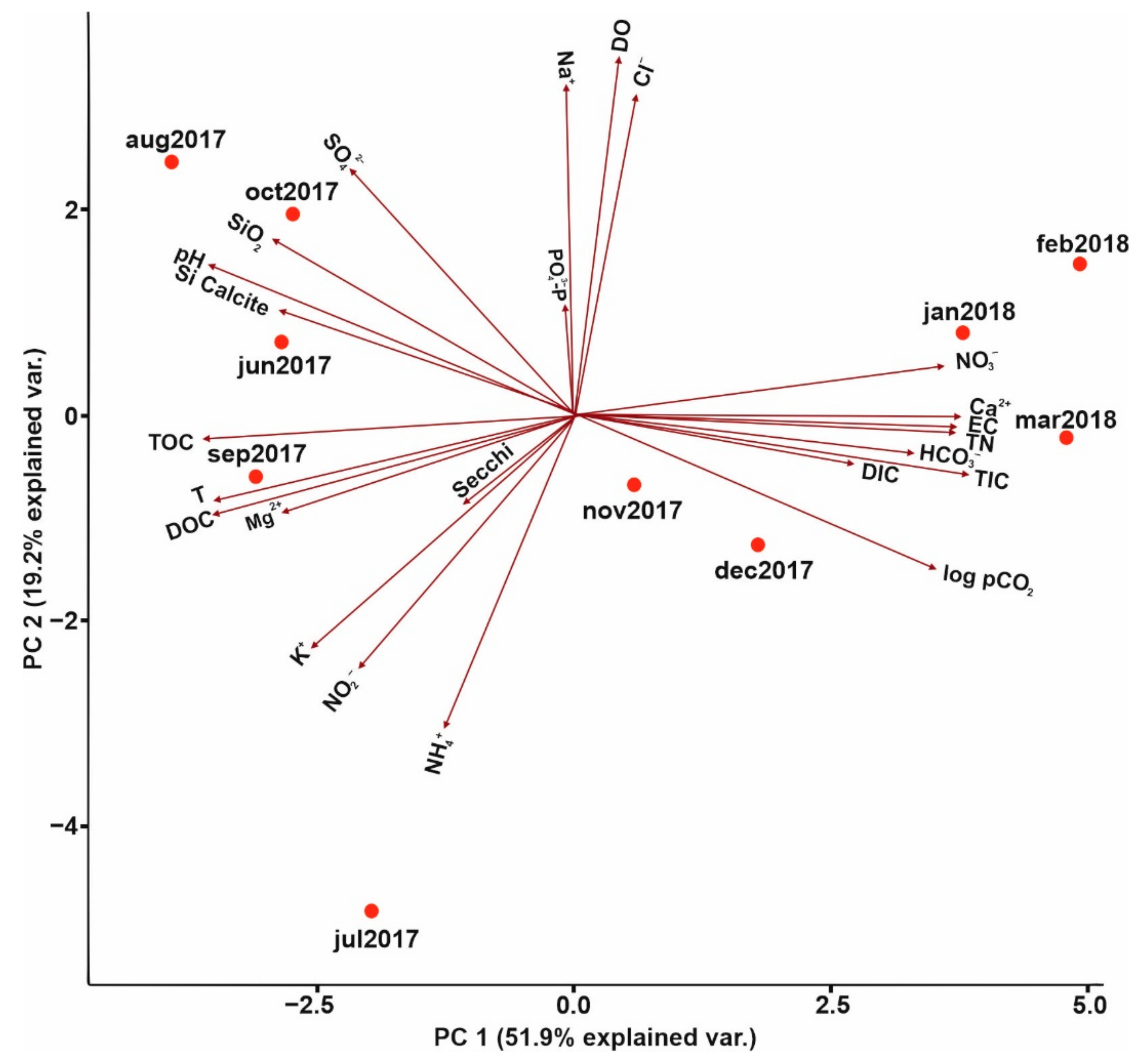
PCA Analysis
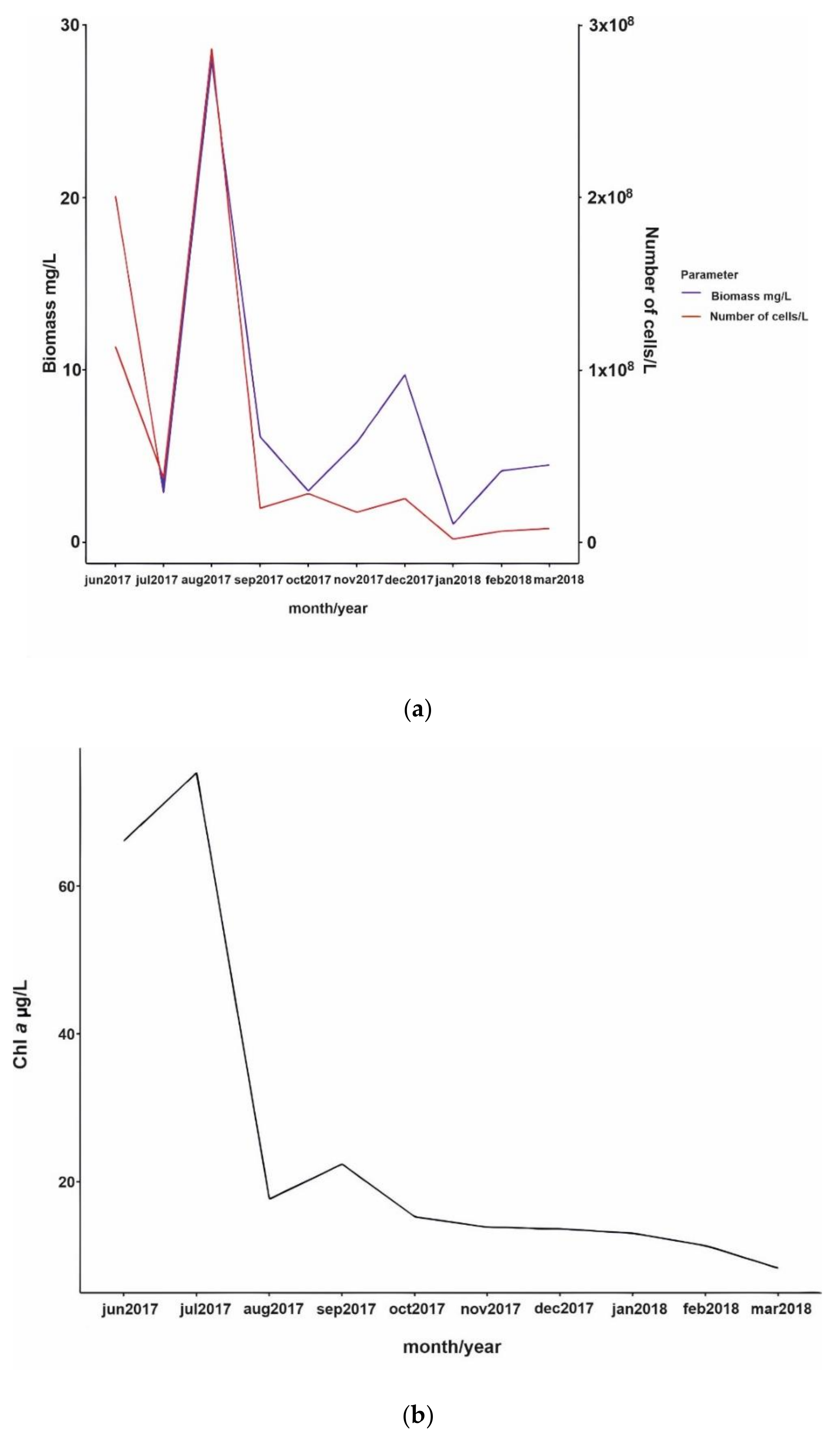
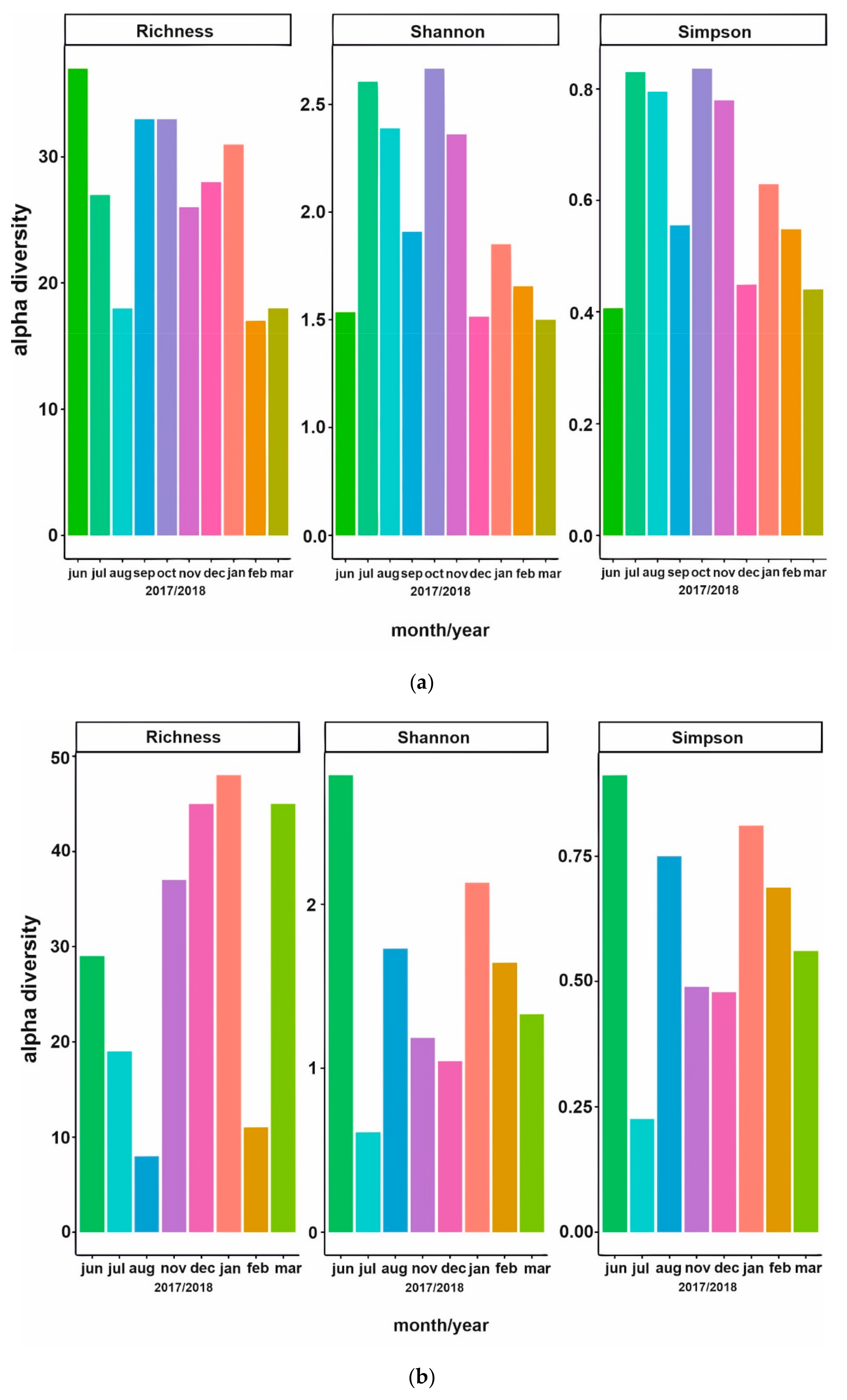
3.2. Phytoplankton Succession
3.3. Bacterial Community Composition
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Thingstad, T.; Sakshaug, E. Control of phytoplankton growth in nutrient recycling ecosystems. Theory and terminology. Mar. Ecol. Prog. Ser. 1990, 63, 261–272. [Google Scholar] [CrossRef]
- Chan, J.L.; Mantzoros, C.S. Role of leptin in energy-deprivation states: Normal human physiology and clinical implications for hypothalamic amenorrhoea and anorexia nervosa. Lancet 2005, 366, 74–85. [Google Scholar] [CrossRef]
- Green, P.; Vörösmarty, C.J.; Meybeck, M.; Galloway, J.N.; Peterson, B.J.; Boyer, E.W. Pre-industrial and contemporary fluxes of nitrogen through rivers: A global assessment based on typology. Biogeochemistry 2004, 68, 71–105. [Google Scholar] [CrossRef]
- Li, Z.G.; Lin, B.-L.; Sagisaka, M.; Yang, P.; Wu, W. Global-scale modelling of potential changes in terrestrial nitrogen cycle from a growing nitrogen deposition. Proc. Environ. Sci. 2012, 13, 1057–1068. [Google Scholar] [CrossRef]
- Galloway, J.N.; Cowling, E.B.; Seitzinger, S.; Socolow, R.H. Reactive nitrogen: Too much of a good thing? Ambio 2002, 31, 60–63. [Google Scholar] [CrossRef]
- Paerl, H.W.; Fulton, R.S.; Moisander, P.H.; Dyble, J. Harmful freshwater algal blooms, with an emphasis on Cyanobacteria. Sci. World J. 2001, 1, 76–113. [Google Scholar] [CrossRef]
- Erisman, J.W.; Galloway, J.; Seitzinger, S.; Bleeker, A.; Butterbach-Bahl, K. Reactive nitrogen in the environment and its effect on climate change. Curr. Opin. Environ. Sustain. 2011, 3, 281–290. [Google Scholar] [CrossRef]
- Dupas, R.; Delmas, M.; Dorioz, J.-M.; Garnier, J.; Moatar, F.; Gascuel-Odoux, C. Assessing the impact of agricultural pressures on N and P loads and eutrophication risk. Ecol. Indic. 2015, 48, 396–407. [Google Scholar] [CrossRef]
- Erisman, J.W.; Galloway, J.N.; Seitzinger, S.; Bleeker, A.; Dise, N.B.; Petrescu, A.M.R.; Leach, A.M.; de Vries, W. Consequences of human modification of the global nitrogen cycle. Philos. Trans. R. Soc. B Biol. Sci. 2013, 368, 20130116. [Google Scholar] [CrossRef]
- Mekonnen, M.M.; Hoekstra, A.Y. Global gray water footprint and water pollution levels related to anthropogenic nitrogen loads to fresh water. Environ. Sci. Technol. 2015, 49, 12860–12868. [Google Scholar] [CrossRef]
- le Doux, E.; Gomez, E.; Monget, J.; Viavattene, C.; Viennot, P.; Ducharne, A.; Benoit, M.; Mignolet, C.; Schott, C.; Mary, B. Agriculture and groundwater nitrate contamination in the Seine basin. The STICS–MODCOU modelling chain. Sci. Total Environ. 2007, 375, 33–47. [Google Scholar] [CrossRef] [PubMed]
- Mengel, K.; Hütsch, B.; Kane, Y. Nitrogen fertilizer application rates on cereal crops according to available mineral and organic soil nitrogen. Eur. J. Agron. 2006, 24, 343–348. [Google Scholar] [CrossRef]
- Sete, P.B.; Comin, J.J.; Ciotta, M.N.; Salume, J.A.; Thewes, F.; Brackmann, A.; Toselli, M.; Nava, G.; Rozane, D.-E.; Loss, A.; et al. Nitrogen fertilization affects yield and fruit quality in pear. Sci. Hortic. 2019, 258, 108782. [Google Scholar] [CrossRef]
- Jégo, G.; Sánchez-Pérez, J.M.; Justes, E. Predicting soil water and mineral nitrogen contents with the STICS model for estimating nitrate leaching under agricultural fields. Agric. Water Manag. 2012, 107, 54–65. [Google Scholar] [CrossRef]
- Bernard-Jannin, L.; Sun, X.; Teissier, S.; Sauvage, S.; Sánchez-Pérez, J.M. Spatio-temporal analysis of factors controlling nitrate dynamics and potential denitrification hot spots and hot moments in groundwater of an alluvial floodplain. Ecol. Eng. 2017, 103, 372–384. [Google Scholar] [CrossRef]
- Singer, G.; Besemer, K.; Schmitt-Kopplin, P.; Hödl, I.; Battin, T.J. Physical heterogeneity increases biofilm resource use and its molecular diversity in stream mesocosms. PLoS ONE 2010, 5, e9988. [Google Scholar] [CrossRef]
- Cardinale, B.J.; Duffy, J.E.; Gonzalez, A.; Hooper, D.U.; Perrings, C.; Venail, P.; Narwani, A.; Mace, G.M.; Tilman, D.; Wardle, D.A.; et al. Biodiversity loss and its impact on humanity. Nat. Cell Biol. 2012, 486, 59–67. [Google Scholar] [CrossRef]
- Besemer, K.; Singer, G.; Quince, C.; Bertuzzo, E.; Sloan, W.; Battin, T.J. Headewaters are critical reservoirs of microbial diversity for fluvial networks. Proc. R. Soc. B 2013, 280, 20131760. [Google Scholar] [CrossRef]
- Widder, S.; Besemer, K.; Singer, G.A.; Ceola, S.; Bertuzzo, E.; Quince, C.; Sloan, W.T.; Rinaldo, A.; Battin, T.J. Fluvial network organization imprints on microbial co-occurrence networks. Proc. Natl. Acad. Sci. USA 2014, 111, 12799–12804. [Google Scholar] [CrossRef]
- Peterson, B.J.; Wollheim, W.M.; Mulholland, P.J.; Webster, J.R.; Meyer, J.L.; Tank, J.L.; Martí, E.; Bowden, W.B.; Valett, H.M.; Hershey, A.E.; et al. Control of nitrogen export from watersheds by headwater streams. Science 2001, 292, 86–90. [Google Scholar] [CrossRef]
- Paerl, H.W. Mitigating harmful cyanobacterial blooms in a human- and climatically impacted world. Life 2014, 4, 988–1012. [Google Scholar] [CrossRef]
- Pal, R.; Choudhury, A.K. An Introduction to Phytoplanktons: Diversity and Ecology; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2014; pp. 25–52. [Google Scholar]
- Herrero, A.; Muro-Pastor, A.M.; Flores, E. Nitrogen control in Cyanobacteria. J. Bacteriol. 2001, 183, 411–425. [Google Scholar] [CrossRef] [PubMed]
- Stal, L. Cyanobacteria: Diversity and versatility. In Algae and Cyanobacteria in Extreme Environments: Cellular Origin, Life in Extreme Habitats and Astrobiology; Sedbach, J., Ed.; Springer: Dordrecht, The Netherlands, 2007; Volume 11, pp. 659–680. [Google Scholar]
- Sellner, K.G. Physiology, ecology, and toxic properties of marine cyanobacteria blooms. Limnol. Oceanogr. 1997, 42, 1089–1104. [Google Scholar] [CrossRef]
- Hemida, M.; Ohyam, T. Nitrogen fixing Cyanobacteria: Future prospect. Nitrogen Fixat. 2014, 23–48. [Google Scholar] [CrossRef]
- Bakker, E.S.; Hilt, S. Impact of water-level fluctuations on cyanobacterial blooms: Options for management. Aquat. Ecol. 2016, 50, 485–498. [Google Scholar] [CrossRef]
- Hamilton, L.C. Public awareness of the scientific consensus on climate. SAGE Open 2016, 6. [Google Scholar] [CrossRef]
- van Gremberghe, I.; Leliaert, F.; Mergeay, J.; Vanormelingen, P.; van der Gucht, K.; Debeer, A.-E.; Lacerot, G.; de Meester, L.; Vyverman, W. Lack of phylogeographic structure in the freshwater Cyanobacterium Microcystis aeruginosa suggests global dispersal. PLoS ONE 2011, 6, e19561. [Google Scholar] [CrossRef]
- Gijzen, H.J.; Mulder, A. The nitrogen cycle out of balance. Water 2001, 21, 38–40. [Google Scholar]
- Moss, B.; Barker, T.; Stephen, D.; Williams, A.E.; Balayla, D.J.; Beklioglu, M.; de Carvalho, L.P.S. Consequences of reduced nutrient loading on a lake system in a lowland catchment: Deviations from the norm? Freshw. Biol. 2005, 50, 1687–1705. [Google Scholar] [CrossRef]
- Rosset, V.; Angélibert, S.; Arthaud, F.; Bornette, G.; Robin, J.; Wezel, A.; Vallod, D.; Oertli, B. Is eutrophication really a major impairment for small waterbody biodiversity? J. Appl. Ecol. 2014, 51, 415–425. [Google Scholar] [CrossRef]
- Menetrey, N.; Sager, L.; Oertli, B.; Lachavanne, J.-B. Looking for metrics to assess the trophic state of ponds. Macroinvertebrates and amphibians. Aquat. Conserv. Mar. Freshw. Ecosyst. 2005, 15, 653–664. [Google Scholar] [CrossRef]
- Søndergaard, M.; Jeppesen, E.; Jensen, J.P.; Amsinck, S.L. Water Framework Directive: Ecological classification of Danish lakes. J. Appl. Ecol. 2005, 42, 616–629. [Google Scholar] [CrossRef]
- Williams, P.; Whitfield, M.; Biggs, J.; Bray, S.; Fox, G.; Nicolet, P.; Sear, D. Comparative biodiversity of rivers, streams, ditches and ponds in an agricultural landscape in Southern England. Biol. Conserv. 2004, 115, 329–341. [Google Scholar] [CrossRef]
- Gvozdić, V.; Brana, J.; Puntarić, D.; Vidosavljević, D.; Roland, D. Changes in the Lower Drava River water quality parameters over 24 years. Arch. Ind. Hyg. Toxicol. 2011, 62, 325–333. [Google Scholar] [CrossRef]
- Villanueva, J.; Rosell, A.; Grimalt, J.O.; Navarro, A.; Rosell-Melé, A. Chemical characterization of polycyclic aromatic hydrocarbon mixtures in uncontrolled hazardous waste dumps. Chemosphere 1991, 22, 317–326. [Google Scholar] [CrossRef]
- Navarro, A.; Carbonell, M. Assessment of groundwater contamination caused by uncontrolled dumping in old gravel quarries in the Besòs aquifers (Barcelona, Spain). Environ. Geochem. Health 2007, 30, 273–289. [Google Scholar] [CrossRef]
- Preziosi, E.; Frollini, E.; Zoppini, A.; Ghergo, S.; Melita, M.; Parrone, D.; Rossi, D.; Amalfitano, S. Disentangling natural and anthropogenic impacts on groundwater by hydrogeochemical, isotopic and microbiological data: Hints from a municipal solid waste landfill. Waste Manag. 2019, 84, 245–255. [Google Scholar] [CrossRef]
- Ondrasek, G.; Begić, H.B.; Zovko, M.; Filipović, L.; Meriño-Gergichevich, C.; Savić, R.; Rengel, Z. Biogeochemistry of soil organic matter in agroecosystems & environmental implications. Sci. Total Environ. 2019, 658, 1559–1573. [Google Scholar] [CrossRef]
- Marković, T.; Karlović, I.; Tadić, M.P.; Larva, O. Application of stable water isotopes to improve conceptual model of alluvial aquifer in the Varaždin area. Water 2020, 12, 379. [Google Scholar] [CrossRef]
- Mandel, S.; Shiftan, Z.L. Groundwater Resources: Investigation and Development, 1st ed.; Academic Press: New York, NY, USA, 1980; pp. 180–200. [Google Scholar]
- Domenico, P.A.; Schwartz, F.W. Physical and Chemical Hydrogeology, 2nd ed.; John Willey and Sons: New York, NY, USA, 1990. [Google Scholar]
- Parkhurst, D.L.; Appelo, C. User’s guide to PHREEQC (Version 2): A computer program for speciation, batch-reaction, one-dimensional transport, and inverse geochemical calculations. Water Res. Investig. Rep. 1999, 99, 4259. [Google Scholar]
- ISO/TC 147 Water Quality. Available online: https://www.iso.org/committee/52834/x/catalogue/ (accessed on 16 April 2018).
- Utermöhl, H. Zur vervollkomnung der quantitativen phytoplankton-methodik. Mitteilungen Internationale Vereiningung für Theoretische und Angewandte. Limnologie 1958, 9, 1–38. [Google Scholar]
- Stock, A.; Jürgens, K.; Bunge, J.; Stoeck, T. Protistan diversity in suboxic and anoxic waters of the Gotland Deep (Baltic Sea) as revealed by 18S rRNA clone libraries. Aquat. Microb. Ecol. 2009, 55, 267–284. [Google Scholar] [CrossRef]
- Stoeck, T.; Bass, D.; Nebel, M.; Christen, R.; Jones, M.D.M.; Breiner, H.-W.; Richards, T.A. Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol. Ecol. 2010, 19, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Amaral-Zettler, L.; McCliment, E.A.; Ducklow, H.W.; Huse, S.M. A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS ONE 2009, 4, e6372. [Google Scholar] [CrossRef]
- Herlemann, D.P.R.; Labrenz, M.; Jürgens, K.; Bertilsson, S.; Waniek, J.J.; Andersson, A.F. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 2011, 5, 1571–1579. [Google Scholar] [CrossRef]
- de Barba, M.; Miquel, C.; Boyer, F.; Mercier, C.; Rioux, D.; Coissac, E.; Taberlet, P. DNA metabarcoding multiplexing and validation of data accuracy for diet assessment: Application to omnivorous diet. Mol. Ecol. Resour. 2013, 14, 306–323. [Google Scholar] [CrossRef] [PubMed]
- Bellemain, E.; Carlsen, T.; Brochmann, C.; Coissac, E.; Taberlet, P.; Kauserud, H. ITS as an environmental DNA barcode for fungi: An in silico approach reveals potential PCR biases. BMC Microbiol. 2010, 10, 189. [Google Scholar] [CrossRef]
- Ficetola, G.F.; Coissac, E.; Zundel, S.; Riaz, T.; Shehzad, W.; Bessière, J.; Taberlet, P.; Pompanon, F. An in-silico approach for the evaluation of DNA barcodes. BMC Genom. 2010, 11, 434. [Google Scholar] [CrossRef]
- BBMap. Available online: https://sourceforge.net/projects/bbmap/ (accessed on 19 February 2019).
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Mahé, F.; Rognes, T.; Quince, C.; de Vargas, C.; Dunthorn, M. Swarm v2: Highly scalable and high-resolution amplicon clustering. PeerJ 2015, 3, e1420. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Vienna, Austria, 2020. [Google Scholar]
- Tapolczai, K.; Vasselon, V.; Lefrançois, E.; Stenger-Kovács, C.; Padisák, J.; Rimet, F. The impact of OTU sequence similarity threshold on diatom-based bioassessment: A case study of the rivers of Mayotte (France, Indian Ocean). Ecol. Evol. 2019, 9, 166–179. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.J. Distance-based tests for homogeneity of multivariate dispersions. Biometrics 2005, 62, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Packag. Vers. 22-1. 2015, 2, 1–2. [Google Scholar]
- Bolpagni, R.; Poikane, S.; Laini, A.; Bagella, S.; Bartoli, M.; Cantonati, M. Ecological and conservation value of small standing-water ecosystems: A systematic review of current knowledge and future challenges. Water 2019, 11, 402. [Google Scholar] [CrossRef]
- Céréghino, R.; Boix, D.; Cauchie, H.-M.; Martens, K.; Oertli, B. The ecological role of ponds in a changing world. Hydrobiologia 2014, 723, 1–6. [Google Scholar] [CrossRef]
- Brönmark, C.; Hansson, L.-A. Environmental issues in lakes and ponds: Current state and perspectives. Environ. Conserv. 2002, 29, 290–307. [Google Scholar] [CrossRef]
- Hering, D.; Haidekker, A.; Schmidt-Kloiber, A.; Barker, T.; Buisson, L.; Graf, W.; Grenouillet, G.; Lorenz, A.; Sandin, L.; Stendera, S. Monitoring the responses of freshwater ecosystems to climate change. Clim. Chang. Impacts Freshw. Ecosyst. 2010, 84–118. [Google Scholar] [CrossRef]
- Boix, D.; Biggs, J.; Céréghino, R.; Hull, A.P.; Kalettka, T.; Oertli, B. Pond research and management in Europe: “Small is Beautiful”. Hydrobiologia 2012, 689, 1–9. [Google Scholar] [CrossRef]
- Bagella, S.; Gascón, S.; Filigheddu, R.; Cogoni, A.; Boix, D. Mediterranean temporary ponds: New challenges from a neglected habitat. Hydrobiologia 2016, 782, 1–10. [Google Scholar] [CrossRef]
- Stanković, I.; Vlahović, T.; Udovič, M.G.; Várbíró, G.; Borics, G. Phytoplankton functional and morpho-functional approach in large floodplain rivers. Hydrobiologia 2012, 698, 217–231. [Google Scholar] [CrossRef]
- Kovács, A.D.; Tóth, G.; Lóczy, D. Water Quality of the Lower Drava River; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2018; pp. 231–245. [Google Scholar]
- Gao, K.; Helbling, E.; Häder, D.; Hutchins, D. Responses of marine primary producers to interactions between ocean acidification, solar radiation, and warming. Mar. Ecol. Prog. Ser. 2012, 470, 167–189. [Google Scholar] [CrossRef]
- van Verseveld, W.J.; McDonnell, J.J.; Lajtha, K. The role of hillslope hydrology in controlling nutrient loss. J. Hydrol. 2009, 367, 177–187. [Google Scholar] [CrossRef]
- Jiang, R.; Woli, K.P.; Kuramochi, K.; Hayakawa, A.; Shimizu, M.; Hatano, R. Hydrological process controls on nitrogen export during storm events in an agricultural watershed. Soil Sci. Plant Nutr. 2010, 56, 72–85. [Google Scholar] [CrossRef]
- Serio, F.; Miglietta, P.P.; Lamastra, L.; Ficocelli, S.; Intini, F.; de Leo, F.; de Donno, A. Groundwater nitrate contamination and agricultural land use: A grey water footprint perspective in Southern Apulia Region (Italy). Sci. Total Environ. 2018, 645, 1425–1431. [Google Scholar] [CrossRef] [PubMed]
- Filipovic, V.; Kodešová, R.; Petosic, D. Experimental and mathematical modeling of water regime and nitrate dynamics on zero tension plate lysimeters in soil influenced by high groundwater table. Nutr. Cycl. Agroecosyst. 2012, 95, 23–42. [Google Scholar] [CrossRef]
- Filipović, V.; Romić, D.; Romić, M.; Borošić, J.; Filipović, L.; Mallmann, F.J.K.; Robinson, D.A. Plastic mulch and nitrogen fertigation in growing vegetables modify soil temperature, water and nitrate dynamics: Experimental results and a modeling study. Agric. Water Manag. 2016, 176, 100–110. [Google Scholar] [CrossRef]
- Hardie, M.; Ridges, J.; Swarts, N.; Close, D.C. Drip irrigation wetting patterns and nitrate distribution: Comparison between electrical resistivity (ERI), dye tracer, and 2D soil–water modelling approaches. Irrig. Sci. 2018, 36, 97–110. [Google Scholar] [CrossRef]
- Paredes, I.; Otero, N.; Soler, A.; Green, A.J.; Soto, D.X. Agricultural and urban delivered nitrate pollution input to Mediterranean temporary freshwaters. Agric. Ecosyst. Environ. 2020, 294, 106859. [Google Scholar] [CrossRef]
- Wetzel, R.G. Limnology: Lake and River Ecosystems, 3rd ed.; Academic Press: San Diego, CA, USA, 2001; pp. 239–288. [Google Scholar]
- Kanda, F.; Yagi, E.; Fukuda, M.; Nakajima, K.; Ohta, T.; Nakata, O. Elucidation of chemical compounds responsible for foot malodour. Br. J. Dermatol. 1990, 122, 771–776. [Google Scholar] [CrossRef]
- Strom, S. Novel interactions between phytoplankton and microzooplankton: Their influence on the coupling between growth and grazing rates in the sea. Hydrobiologia 2002, 480, 41–54. [Google Scholar] [CrossRef]
- Budzyńska, A.; Gołdyn, R.; Zagajewski, P.; Dondajewska, R.; Kowalczewska-Madura, K. The dynamics of a Planktothrix agardhii population in a shallow dimictic lake. Oceanol. Hydrobiol. Stud. 2009, 38, 1–6. [Google Scholar]
- Meffert, M.F. Planktic unsheathed filaments (Cyanophyceae) with polar and central gas-vacuoles. II. Biology population dynamics and biotopes of Limnothrix redekei (Van Goor) Meffert. Arch. Hydrobiol. 1989, 116, 257–282. [Google Scholar]
- Trifonova, I.S. Ecology and Succession of Lake Phytoplankton; Nauka Press: Leningrad, Russia, 1990. [Google Scholar]
- Wiedner, C.; Nixdorf, B. Success of chrysophytes, cryptophytes and dinoflagellates over blue green (cyanobacteria) during an extreme winter (1995/96) in eutrophic shallow lakes. Hydrobiologia 1998, 369, 229–235. [Google Scholar] [CrossRef]
- Romanov, R.E. Limnothrix redeckei (Van Goor) Meffert (Cyanoprocaryota) in the potamoplankton. Int. J. Algae 2007, 9, 105–116. [Google Scholar] [CrossRef]
- Licursi, M.; Sierra, M.V.; Gómez, N. Diatom assemblages from a turbid coastal plain estuary: Río de la Plata (South America). J. Mar. Syst. 2006, 62, 35–45. [Google Scholar] [CrossRef]
- Meister, F. Die Kieselalgen der Schweiz. In Beiträge zur Kryptogamenflora der Schweiz; Wyss, K.J., Ed.; K.J. Wyss: Bern, Switzerland, 1912; Volume 4, p. 254. [Google Scholar]
- Bottinelli, M. Fioriture di Cianobatteri della specie Microcystis wesenbergii nel Lago di Muzzano. Ph.D. Thesis, Tesi Sperimentale di Laurea in Scienze Naturali, Universita degli Studi di Pavia, Pavia, Italy, 1999; p. 172. [Google Scholar]
- Bottinelli, M.; Tonolla, M.; Forlani, G.; Crivelli, C.; Sanangelantoni, A.M.; e Peduzzi, R. Fioriture di Cianobatteri della specie Microcystis wesenbergii nel Lago di Muzzano (Svizzera). Boll. Della Soc. Tic. Di Sci. Nat. 2000, 88, 53–61. [Google Scholar]
- Isenburg, C.; Loizeau, J.L.; Tonolla, M.; Peduzzi, R. Aspetti limnologici e microbiologici del Laghetto di Muzzano (TI). Boll. Soc. Tic. Sci. Nat. 2000, 88, 41–51. [Google Scholar]
- Pedrotta, T. Analisi qualitativa degli immissari e dell’emissario del Laghetto di Muzzano. Tesi di Laurea in AGRN. Front. Ecol. Evolut. 2009, 3, 70. [Google Scholar]
- Reynolds, C. Ecology of Phytoplankton; Cambridge University Press: Cambridge, UK, 2006. [Google Scholar]
- Glibert, P.M.; Wilkerson, F.P.; Dugdale, R.C.; Raven, J.A.; Dupont, C.L.; Leavitt, P.R.; Parker, A.E.; Burkholder, J.M.; Kana, T.M. Pluses and minuses of ammonium and nitrate uptake and assimilation by phytoplankton and implications for productivity and community composition, with emphasis on nitrogen-enriched conditions. Limnol. Oceanogr. 2016, 61, 165–197. [Google Scholar] [CrossRef]
- Eppley, R.W.; Coatsworth, J.L.; Solórzano, L. Studies of nitrate reductase in marine phytoplankton. Limnol. Oceanogr. 1969, 14, 194–205. [Google Scholar] [CrossRef]
- Sommer, U. Disturbance-diversity relationships in two lakes of similar nutrient chemistry but contrasting disturbance regimes. Hydrobiologia 1993, 249, 59–65. [Google Scholar] [CrossRef]
- Miyazaki, T.; Honjo, Y.; Ichimura, S. Applicability of the stable isotope method using 13C and 15N simultaneously to the estimation of carbon and nitrogen assimilation in a eutrophic, freshwater lake, Lake Nakanuma, Japan. Arch. Hydrobiol. 1985, 102, 355–365. [Google Scholar]
- Karya, N.; van der Steen, N.; Lens, P. Photo-oxygenation to support nitrification in an algal–bacterial consortium treating artificial wastewater. Bioresour. Technol. 2013, 134, 244–250. [Google Scholar] [CrossRef]
- Hou, X.L.; Ma, D.L.; Yang, K.F.; Wang, R.; Rong, D.L.; Yin, X.T. The physiological responses of one algae species Scenedesmus quadricauda to ammonium and alanine. IOP Conf. Ser. Earth Environ. Sci. 2019, 344, 012167. [Google Scholar] [CrossRef]
- Watanabe, T.; Miyazaki, T. Maximum ammonium uptake rates of Scenedesmus quadricauda (Chlorophyta) and Microcystis novacekii (Cyanobacteria) grown under nitrogen limitation and implications for competition. J. Phycol. 1996, 32, 243–249. [Google Scholar] [CrossRef]
- Trommer, G.; Poxleitner, M.; Stibor, H. Responses of lake phytoplankton communities to changing inorganic nitrogen supply forms. Aquat. Sci. 2020, 82, 22. [Google Scholar] [CrossRef]
- Barone, R.; Naselli-Flores, L. Distribution and seasonal dynamics of cryptomonads in Sicilian water bodies. Hydrobiologia 2003, 502, 325–329. [Google Scholar] [CrossRef]
- Paerl, H.W. Mitigating toxic planktonic cyanobacterial blooms in aquatic ecosystems facing increasing anthropogenic and climatic pressures. Toxins 2018, 10, 76. [Google Scholar] [CrossRef]
- Hutchinson, G.E. The paradox of the plankton. Am. Nat. 2002, 95, 137–145. [Google Scholar] [CrossRef]
- Wilhelm, S.W.; Bullerjahn, G.S.; McKay, R.M.L. The complicated and confusing ecology of Microcystis blooms. mBio 2020, 11, 00529-20. [Google Scholar] [CrossRef] [PubMed]
- Belykh, O.I.; Dmitrieva, O.A.; Gladkikh, A.S.; Sorokovikova, E.G. Identification of toxigenic Cyanobacteria of the genus Microcystis in the Curonian Lagoon (Baltic Sea). Oceanology 2013, 53, 71–79. [Google Scholar] [CrossRef]
- Fuerst, J.A. Planctomycetes—new models for microbial cells and activities. Microb. Res. 2017, 1–27. [Google Scholar] [CrossRef]
- Strous, M.; Kuenen, J.G.; Jetten, M.S.M. Key physiology of anaerobic ammonium oxidation. Appl. Environ. Microbiol. 1999, 65, 3248–3250. [Google Scholar] [CrossRef] [PubMed]
- Jetten, M.S.; Wagner, M.; Fuerst, J.; van Loosdrecht, M.; Kuenen, G.; Strous, M. Microbiology and application of the anaerobic ammonium oxidation (‘anammox’) process. Curr. Opin. Biotechnol. 2001, 12, 283–288. [Google Scholar] [CrossRef]
- Jetten, M.S.; Sliekers, O.; Kuypers, M.M.M.; Dalsgaard, T.K.; van Niftrik, L.; Cirpus, I.; van de Pas-Schoonen, K.; Lavik, G.; Thamdrup, B.; le Paslier, D.; et al. Anaerobic ammonium oxidation by marine and freshwater planctomycete-like bacteria. Appl. Microbiol. Biotechnol. 2003, 63, 107–114. [Google Scholar] [CrossRef]
- Schmid, M.; Schmitz-Esser, S.; Jetten, M.; Wagner, M. 16S-23S rDNA intergenic spacer and 23S rDNA of anaerobic ammonium-oxidizing bacteria: Implications for phylogeny and in situ detection . Environ. Microbiol. 2001, 3, 450–459. [Google Scholar] [CrossRef]
- Kuypers, M.M.M.; Sliekers, A.O.; Lavik, G.; Schmid, M.; Jørgensen, B.B.; Kuenen, J.G.; Damsté, J.S.S.; Strous, M.; Jetten, M.S.M. Anaerobic ammonium oxidation by anammox bacteria in the Black Sea. Nat. Cell Biol. 2003, 422, 608–611. [Google Scholar] [CrossRef]
- van de Graaf, A.A.; de Bruijn, P.; Robertson, L.A.; Jetten, M.S.M.; Kuenen, J.G. Metabolic pathway of anaerobic ammonium oxidation on the basis of 15N studies in a fluidized bed reactor. Microbiology 1997, 143, 2415–2421. [Google Scholar] [CrossRef]
- Newton, P.N.; Amin, A.A.; E Bird, C.; Passmore, P.; Dukes, G.; Tomson, G.; Simons, B.; Bate, R.; Guerin, P.J.; White, N.J. The primacy of public health considerations in defining poor quality medicines. PLoS Med. 2011, 8, e1001139. [Google Scholar] [CrossRef] [PubMed]
- Russo, D.A.; Couto, N.; Beckerman, A.P.; Pandhal, J. A metaproteomic analysis of the response of a freshwater microbial community under nutrient enrichment. Front. Microbiol. 2016, 7, 1172. [Google Scholar] [CrossRef] [PubMed]
- Martin-Jezequel, V.; Hildebrand, M.; Brzezinski, M.A. Silicon metabolism in diatoms: Implications for growth. J. Phycol. 2000, 36, 821–840. [Google Scholar] [CrossRef]
- Lundholm, N.; Moestrup, Ø.; Kotaki, Y.; Hoef-Emden, K.; Scholin, C.; Miller, P. Inter- and intraspecific variation of the Pseudo-nitzschia delicatissima complex (Bacillariophyceae) illustrated by rRNA probes, morphological data and phylogenetic analyses. J. Phycol. 2006, 42, 464–481. [Google Scholar] [CrossRef]
- Nelson, D.M.; Dortch, Q. Silicic acid depletion in the plume of the Mississippi River and limitation of Si availability to diatoms in the northern Gulf of Mexico: Evidence from kinetic studies in spring and summer. Mar. Ecol. Prog. Ser. 1996, 136, 163–178. [Google Scholar] [CrossRef]
- Gibson, C.E.; Wang, G.; Foy, R.H. Silica and diatom growth in Lough Neagh: The importance of internal recycling. Freshw. Biol. 2000, 45, 285–293. [Google Scholar] [CrossRef]
- Glibert, P.M.; Wilkerson, F.P.; Dugdale, R.C.; Parker, A.E.; Alexander, J.A.; Blaser, S.; Murasko, S. Phytoplankton communities from San Francisco Bay Delta respond differently to oxidized and reduced nitrogen substrates-even under conditions that would otherwise suggest nitrogen sufficiency. Front. Mar. Sci. 2014, 1, 17. [Google Scholar] [CrossRef]
- Dortch, Q. The interaction between ammonium and nitrate uptake in phytoplankton. Mar. Ecol. Prog. Ser. 1990, 61, 183–201. [Google Scholar] [CrossRef]
- Kamjunke, N.; Henrichs, T.; Gaedke, U. Phosphorus gain by bacterivory promotes the mixotrophic flagellate Dinobryon spp. during re-oligotrophication. J. Plankton Res. 2006, 29, 39–46. [Google Scholar] [CrossRef]
- Žutinić, P.; Gligora Udovič, M.; Kralj Borojević, K.; Plenković-Moraj, A.; Padisák, J. Morpho-functional classifications of phytoplankton assemblages of two deep karstic lakes. Hydrobiologia 2014, 740, 147–166. [Google Scholar] [CrossRef]
- Eiler, A.; Bertilsson, S. Flavobacteria blooms in four eutrophic lakes: Linking population dynamics of freshwater bacterioplankton to resource availability. Appl. Environ. Microbiol. 2007, 73, 3511–3518. [Google Scholar] [CrossRef] [PubMed]
- Ghai, R.; Mizuno, C.M.; Picazo, A.; Camacho, A.; Rodriguez-Valera, F. Key roles for freshwater Actinobacteria revealed by deep metagenomic sequencing. Mol. Ecol. 2014, 23, 6073–6090. [Google Scholar] [CrossRef] [PubMed]
- Lauro, F.M.; McDougald, D.; Thomas, T.; Williams, T.J.; Egan, S.; Rice, S.; DeMaere, M.Z.; Ting, L.; Ertan, H.; Johnson, J.; et al. The genomic basis of trophic strategy in marine bacteria. Proc. Natl. Acad. Sci. USA 2009, 106, 15527–15533. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.K.; Sommerfeld, K.; Bullerjahn, G.S.; Matteson, A.R.; Wilhelm, S.W.; Jezbera, J.; Brandt, U.; Doolittle, W.F.; Hahn, M.W. Actinorhodopsin genes discovered in diverse freshwater habitats and among cultivated freshwater Actinobacteria. ISME J. 2009, 3, 726–737. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Moestrup, Ø.; Jordan, R.W.; Iwataki, M. Two new freshwater Woloszynskioids Asulcocephalium miricentonis gen. et sp. nov. and Leiocephalium pseudosanguineum gen. et sp. nov. (Suessiaceae, Dinophyceae) lacking an apical furrow apparatus. Protist 2015, 166, 638–658. [Google Scholar] [CrossRef] [PubMed]
- Reveal, J.L. Wislouchiella planctonica Skvortz. (Chlorophyta, Volvocales), a new algal record for Nevada. Great Basin Nat. 1969, 29, 3–4. [Google Scholar]
- Broady, P.A.; Flint, E.A.; Nelson, W.A.; Cassie Cooper, V.; de Winton, M.D.; Novis, P.M. Phylum Chlorophyta and Charophyta: Green algae. In New Zealand Inventory of Biodiversity, Kingdoms Bacteria, Protozoa, Chromista, Plantae, Fungi; Gordon, D.P., Ed.; Canterbury University Press: Christchurch, New Zealand, 2012; Volume 3, pp. 347–381. [Google Scholar]
- Felip, M.; Sattler, B.; Psenner, R.; Catalan, J. Highly active microbial communities in the ice and snow cover of high mountain lakes. Appl. Environ. Microbiol. 1995, 61, 2394. [Google Scholar] [CrossRef]
- Callieri, C.; Stockner, J.G. Freshwater autotrophic picoplankton: A review. J. Limnol. 2002, 61, 1–14. [Google Scholar] [CrossRef]
- Fawley, M.W.; Fawley, K.P.; Buchheim, M.A. Molecular diversity among communities of freshwater microchlorophytes. Microb. Ecol. 2004, 48, 489–499. [Google Scholar] [CrossRef]
- Muramoto, K.; Nakada, T.; Shitara, T.; Hara, Y.; Nozaki, H. Re-examination of the snow algal species Chloromonas miwae (Fukushima) Muramoto et al., comb. nov. (Volvocales, Chlorophyceae) from Japan, based on molecular phylogeny and cultured material. Eur. J. Phycol. 2010, 45, 27–37. [Google Scholar] [CrossRef]
- Mikhailyuk, T.; Glaser, K.; Tsarenko, P.; Demchenko, E.; Karsten, U. Composition of biological soil crusts from sand dunes of the Baltic Sea coast in the context of an integrative approach to the taxonomy of microalgae and cyanobacteria. Eur. J. Phycol. 2019, 54, 263–290. [Google Scholar] [CrossRef]
- Medinger, R.; Nolte, V.; Pandey, R.V.; Jost, S.; Ottenwälder, B.; Schlötterer, C.; Boenigk, J. Diversity in a hidden world: Potential and limitation of next-generation sequencing for surveys of molecular diversity of eukaryotic microorganisms. Mol. Ecol. 2010, 19, 32–40. [Google Scholar] [CrossRef]
- Mackintosh, C.; Beattie, K.A.; Klumpp, S.; Cohen, P.; Codd, G.A. Cyanobacterial microcystin-LR is a potent and specific inhibitor of protein phosphatases 1 and 2A from both mammals and higher plants. FEBS Lett. 1990, 264, 187–192. [Google Scholar] [CrossRef]
- Eland, L.E.; Davenport, R.J.; Mota, C. Evaluation of DNA extraction methods for freshwater eukaryotic microalgae. Water Res. 2012, 46, 5355–5364. [Google Scholar] [CrossRef] [PubMed]
- Rashidan, K.K.; Bird, D.F. Role of predatory bacteria in the termination of a cyanobacterial bloom. Microb. Ecol. 2001, 41, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Love, N.G. Application of rbcL based molecular diversity analysis to algae in wastewater treatment plants. Bioresour. Technol. 2011, 102, 3619–3622. [Google Scholar] [CrossRef]
- van Elsas, J.D.; Boersma, F.G.H. A review of molecular methods to study the microbiota of soil and the mycosphere. Eur. J. Soil Biol. 2011, 47, 77–87. [Google Scholar] [CrossRef]
| Variable | Min | Max | Mean | Med | SD |
|---|---|---|---|---|---|
| T (°C) | 1.2 | 24.2 | 13.0 | 11.5 | 9.1 |
| EC (μS cm−1) | 252 | 497 | 347 | 316 | 99 |
| pH | 7.83 | 9.48 | 8.49 | 8.23 | 0.60 |
| DO (mg L−1) | 7.1 | 17.1 | 12.5 | 12.7 | 2.9 |
| logpCO2 | −4.69 | −2.54 | −3.44 | −3.15 | 0.74 |
| HCO3− (mg L−1) | 107 | 249 | 159 | 138 | 54 |
| PO43−-P (mg L−1) | 0.01 | 0.32 | 0.10 | 0.05 | 0.11 |
| TN (mg L−1) | 0.28 | 10.15 | 4.57 | 4.45 | 3.54 |
| NH4+ (mg L−1) | 0.01 | 2.75 | 0.42 | 0.09 | 0.84 |
| NO2− (mg L−1) | 0.05 | 0.17 | 0.08 | 0.07 | 0.04 |
| NO3− (mg L−1) | 0.6 | 38.4 | 15.3 | 11.1 | 14.0 |
| TIC (mg L−1) | 18.44 | 29.36 | 23.76 | 23.06 | 4.62 |
| DIC (mg L−1) | 14.50 | 27.68 | 21.61 | 20.19 | 4.33 |
| TOC (mg L−1) | 7.20 | 24.66 | 16.61 | 16.25 | 6.68 |
| DOC (mg L−1) | 6.13 | 20.22 | 14.08 | 13.82 | 5.57 |
| Ca2+ (mg L−1) | 20.0 | 66.1 | 38.2 | 31.7 | 18.3 |
| Mg2+ (mg L−1) | 15.2 | 20.0 | 17.4 | 16.7 | 1.7 |
| Na+ (mg L−1) | 6.9 | 17.4 | 12.2 | 13.3 | 3.7 |
| K+ (mg L−1) | 0.9 | 1.6 | 1.1 | 1.1 | 0.2 |
| Cl− (mg L−1) | 11.8 | 24.7 | 17.5 | 17.3 | 4.2 |
| SO42− (mg L−1) | 16.0 | 33.1 | 24.3 | 24.8 | 5.6 |
| SiO2 (mg L−1) | 7.2 | 26.8 | 15.6 | 13.0 | 6.2 |
| ZSD (m) | 0.125 | 0.5 | 0.28 | 0.25 | 0.111 |
| SICalcite | −0.2 | 1.1 | 0.5 | 0.5 | 0.4 |
| PCA Axis | PC1 | PC2 |
|---|---|---|
| Standard deviation | 3.531 | 2.145 |
| Proportion of variance (%) | 51.9 | 19.2 |
| Cumulative proportion (%) | 51.9 | 71.1 |
| Eigenvalues | ||
| T | −0.246 | −0.094 |
| EC | 0.268 | −0.012 |
| pH | −0.254 | 0.173 |
| DO | 0.030 | 0.407 |
| logpCO2 | 0.256 | −0.184 |
| HCO3− | 0.237 | −0.045 |
| PO43−-P | −0.008 | −0.126 |
| TN | 0.265 | −0.021 |
| NH4+ | −0.089 | −0.355 |
| NO2− | −0.154 | −0.294 |
| NO3− | 0.266 | 0.059 |
| TIC | 0.271 | −0.067 |
| DIC | 0.193 | −0.054 |
| TOC | −0.260 | −0.028 |
| DOC | −0.249 | −0.106 |
| Ca2+ | 0.269 | −0.0003 |
| Mg2+ | −0.205 | −0.103 |
| Na+ | −0.006 | 0.377 |
| K+ | −0.182 | −0.263 |
| Cl− | 0.041 | 0.368 |
| SO42− | −0.156 | 0.285 |
| SiO2 | −0.208 | 0.200 |
| ZSD (Secchi) | −0.079 | −0.103 |
| SICalcite | −0.205 | 0.127 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kulaš, A.; Marković, T.; Žutinić, P.; Kajan, K.; Karlović, I.; Orlić, S.; Keskin, E.; Filipović, V.; Gligora Udovič, M. Succession of Microbial Community in a Small Water Body within the Alluvial Aquifer of a Large River. Water 2021, 13, 115. https://doi.org/10.3390/w13020115
Kulaš A, Marković T, Žutinić P, Kajan K, Karlović I, Orlić S, Keskin E, Filipović V, Gligora Udovič M. Succession of Microbial Community in a Small Water Body within the Alluvial Aquifer of a Large River. Water. 2021; 13(2):115. https://doi.org/10.3390/w13020115
Chicago/Turabian StyleKulaš, Antonija, Tamara Marković, Petar Žutinić, Katarina Kajan, Igor Karlović, Sandi Orlić, Emre Keskin, Vilim Filipović, and Marija Gligora Udovič. 2021. "Succession of Microbial Community in a Small Water Body within the Alluvial Aquifer of a Large River" Water 13, no. 2: 115. https://doi.org/10.3390/w13020115
APA StyleKulaš, A., Marković, T., Žutinić, P., Kajan, K., Karlović, I., Orlić, S., Keskin, E., Filipović, V., & Gligora Udovič, M. (2021). Succession of Microbial Community in a Small Water Body within the Alluvial Aquifer of a Large River. Water, 13(2), 115. https://doi.org/10.3390/w13020115








