Comparison of the ColiPlate™ Kit with Two Common E. coli Enumeration Methods for Water
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Analysis Preparation
2.2. ColiPlate Analysis
2.3. Colilert Analysis
2.4. E. coli Enumeration by Membrane Filtration with Modified mTEC Agar
2.5. Statistical Analysis
3. Results
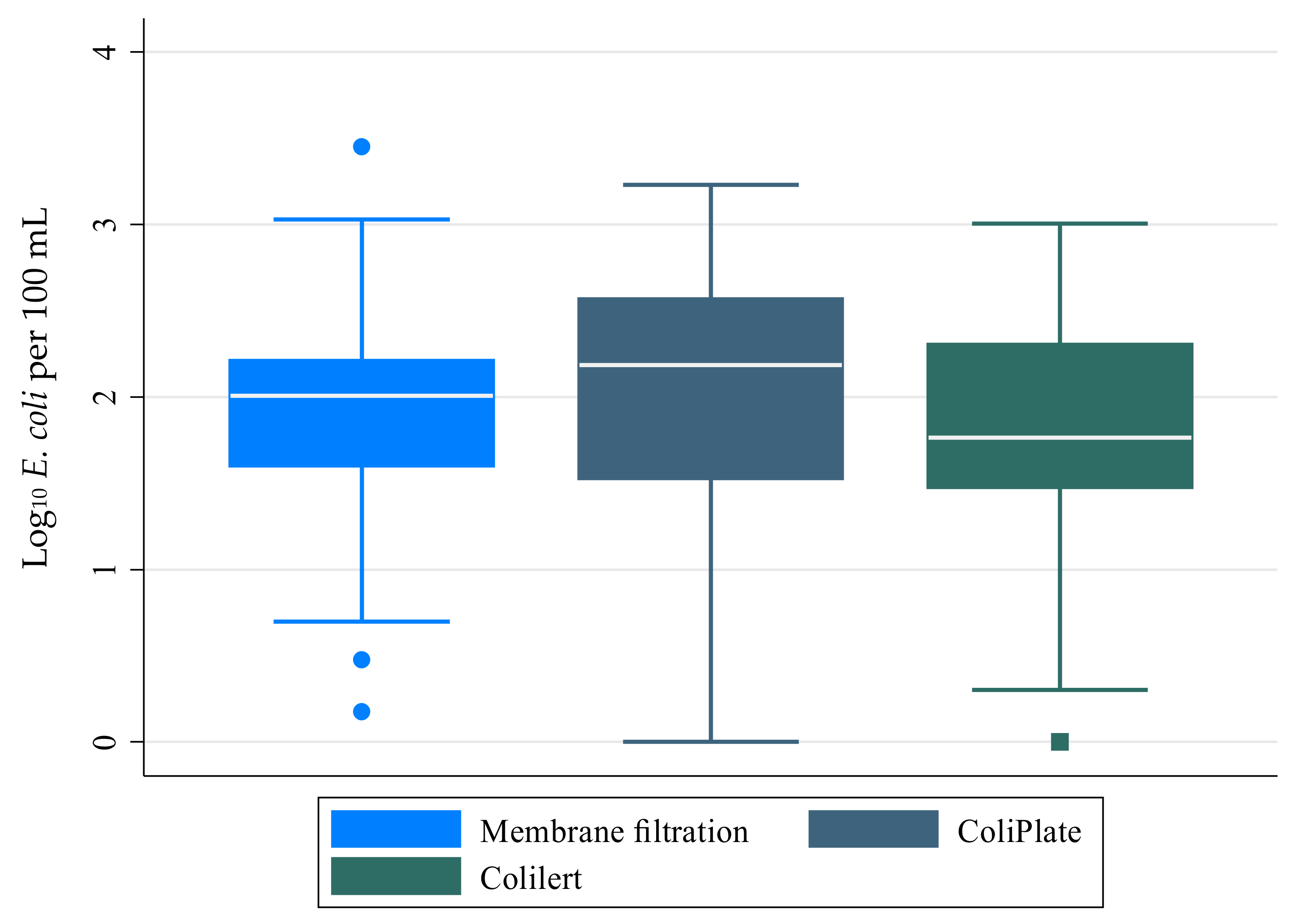
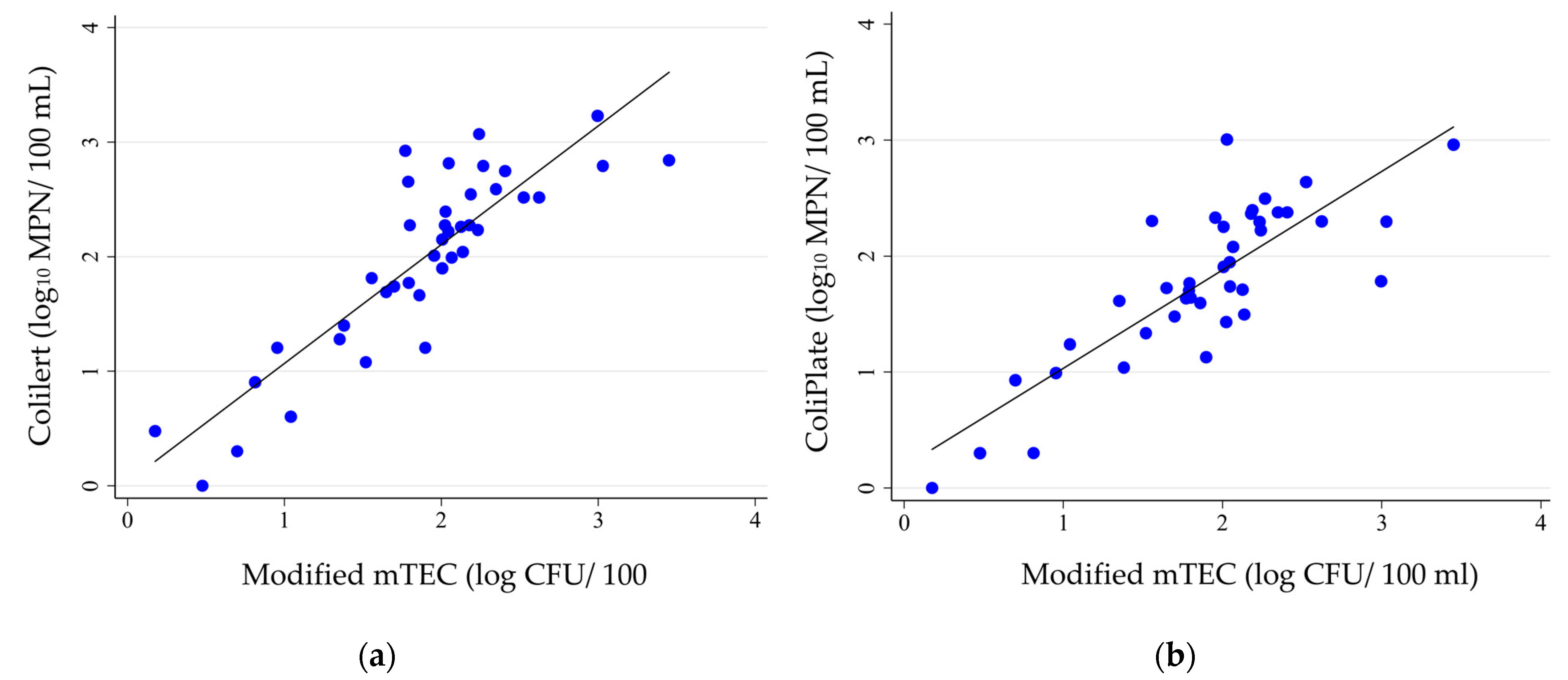
3.1. Comparison of E. coli Density Results by Enumeration Method
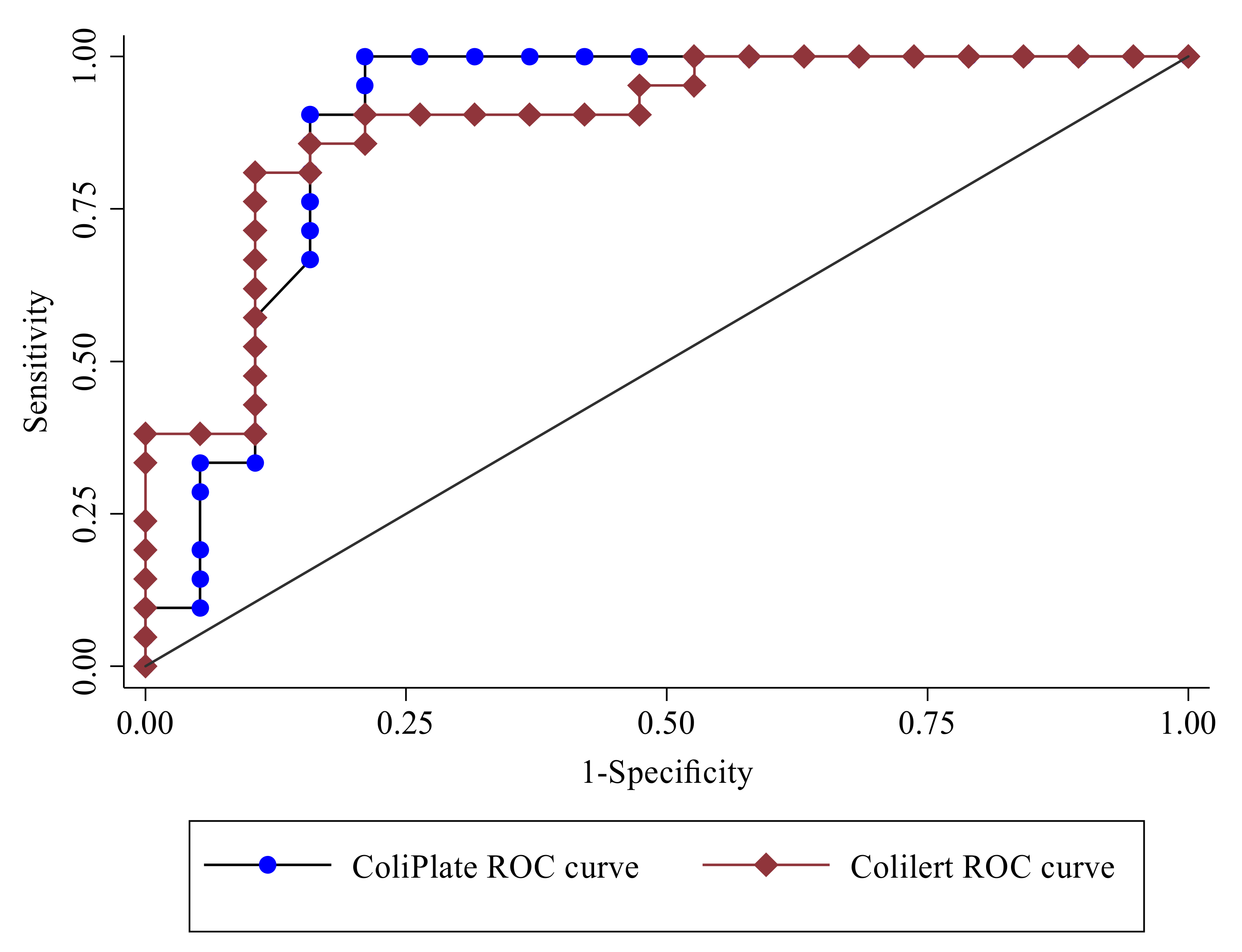
3.2. ColiPlate and Colilert for the Detection of Very High-Risk E. coli Density
4. Discussion
4.1. Comparison of the Observed Correlations with Related Studies
4.2. Application of Results to Water Quality Analyses in Low-Resource Environments
4.3. ColiPlate E. coli Densities Greater Than Densities from Membrane Filtration and Colilert
4.4. Study Limitations and Challenges Presented
5. Conclusions
- Strong correlations were observed between the E. coli densities obtained by ColiPlate and Colilert with the MF method using modified mTEC agar.
- The ColiPlate and Colilert methods provided excellent discrimination in properly classifying the WHO high-risk conditions for drinking water-related illnesses.
- ColiPlate had higher sensitivity but lower specificity than Colilert in this study for determining the WHO high-risk conditions for drinking water-related illnesses.
- The data from these Kentucky (USA) water samples suggest ColiPlate continues to effectively assess fecal indicator bacteria in natural water relative to Colilert, and the evidence suggests that ColiPlate could potentially be an effective tool for citizen scientists evaluating untreated drinking waters and natural waters.
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kay, D.; Crowther, J.; Davies, C.; Edwards, A.; Fewtrell, L.; Francis, C.; Kay, C.; McDonald, A.; Stapleton, C.; Watkins, J.; et al. Faecal indicators and pathogens: Expanding opportunities for the microbiology community. In The Significance of Faecal Indicators in Water; Kay, D., Fricker, C., Eds.; Royal Society of Chemistry: Cambridge, UK, 2012; pp. 1–17. [Google Scholar] [CrossRef]
- WHO & UNICEF. Progress on Drinking Water, Sanitation and Hygiene: 2017 Update and SDG Baselines; WHO & UNICEF: Geneva, Switzerland, 2017; p. 26. [Google Scholar]
- An, Y.J.; Breindenbach, G.P. Monitoring E. coli and total coliforms in natural spring water as related to recreational mountain areas. Environ. Monit. Assess. 2005, 102, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Giampaoli, S.; Spica, V.R. Health and safety in recreational waters. Bull. World Health Organ. 2014, 92, 79. [Google Scholar] [CrossRef] [PubMed]
- Edberg, S.C.L.; Rice, E.W.; Karlin, R.J.; Allen, M.J. Escherichia coli: The best biological drinking water indicator for public health protection. J. Appl. Microbiol. 2000, 88, 106–116. [Google Scholar] [CrossRef] [PubMed]
- McGuinness, M. Faecal indicators in drinking water—is it time to move on? In The Significance of Faecal Indicators in Water; Kay, D., Fricker, C., Eds.; Royal Society of Chemistry: Cambridge, UK, 2012; pp. 18–26. [Google Scholar] [CrossRef]
- Prüss, A. Review of epidemiological studies on health effects from exposure to recreational water. Int. J. Epidemiol. 1998, 27, 1–9. [Google Scholar] [CrossRef]
- Wade, T.J.; Pai, N.; Eisenberg, J.N.; Colford, J.M., Jr. Do U.S. Environmental Protection Agency water quality guidelines for recreational waters prevent gastrointestinal illness? A systematic review and meta-analysis. Environ. Health Perspect. 2003, 111, 1102–1109. [Google Scholar] [CrossRef]
- Wiedenmann, A.; Krüger, P.; Dietz, K.; López-Pila, J.M.; Szewzyk, R.; Botzenhart, K. A randomized controlled trial assessing infectious disease risks from bathing in fresh recreational waters in relation to the concentration of Escherichia coli, intestinal enterococci, Clostridium perfringens, and somatic coliphages. Environ. Health Perspect. 2006, 114, 228–236. [Google Scholar] [CrossRef]
- Marion, J.W.; Lee, J.; Lemeshow, S.; Buckley, T.J. Association of gastrointestinal illness and recreational water exposure at an inland US beach. Water Res. 2010, 44, 4796–4804. [Google Scholar] [CrossRef] [PubMed]
- European Commission. Directive 2006/7/EC of the European Parliament, concerning the management of bathing water quality and repealing Directive 76/160/EEC. Off. J. Eur. Union 2006, 64, 37–51. [Google Scholar]
- U.S. EPA. Recreational Water Quality Criteria; Office of Water 820-F-12-058; United States Environmental Protection Agency: Washington, DC, USA, 2012.
- Levy, K. Does Poor Water Quality Cause Diarrheal Disease? Am. J. Trop. Med. Hyg. 2015, 93, 899–900. [Google Scholar] [CrossRef][Green Version]
- Gundry, S.; Wright, J.; Conroy, R. A systematic review of the health outcomes related to household water quality in developing countries. J. Water Health 2004, 2, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Gruber, J.S.; Ercumen, A.; Colford, J.M., Jr. Coliform bacteria as indicators of diarrheal risk in household drinking water: Systematic review and meta-analysis. PLoS ONE 2014, 9, e107429. [Google Scholar] [CrossRef]
- Profitós, J.M.H.; Mouhaman, A.; Lee, S.; Garabed, R.; Moritz, M.; Piperata, B.; Tien, J.; Bisesi, M.; Lee, J. Muddying the waters: A new area of concern for drinking water contamination in Cameroon. Int. J. Environ. Res. Public Health 2014, 11, 12454–12472. [Google Scholar] [CrossRef]
- Luby, S.P.; Halder, A.K.; Huda, T.M.; Unicomb, L.; Islam, M.S.; Arnold, B.F.; Johnston, R.B. Microbiological contamination of drinking water associated with subsequent child diarrhea. Am. J. Trop. Med. Hyg. 2015, 93, 904–911. [Google Scholar] [CrossRef] [PubMed]
- Levy, K.; Nelson, K.L.; Hubbard, A.; Eisenberg, J.N. Rethinking indicators of microbial drinking water quality for health studies in tropical developing countries: Case study in northern coastal Ecuador. Am. J. Trop. Med. Hyg. 2012, 86, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Weppelmann, T.A.; Alam, M.T.; Widmer, J.; Morrissey, D.; Rashid, M.H.; De Rochars, V.M.B.; Morris, J.G.; Ali, A.; Johnson, J.A. Feasibility of the hydrogen sulfide test for the assessment of drinking water quality in post-earthquake Haiti. Environ. Monit. Assess. 2014, 186, 8509–8516. [Google Scholar] [CrossRef][Green Version]
- Sobsey, M.D.; Pfaender, F.K. Evaluation of the H2S Method for Detection of Fecal Contamination of Drinking Water; World Health Organization Report WHO/SDE/WSH/02.08. World Health Organization: Geneva, Switzerland, 2002; p. 44. Available online: http://www.who.int/water_sanitation_health/dwq/WSH02.08.pdf (accessed on 20 June 2021).
- Gupta, S.K.; Sheikh, M.A.; Islam, M.S.; Rahman, K.S.; Jahan, N.; Rahman, M.M.; Hoekstra, R.M.; Johnston, R.; Ram, P.K.; Luby, S. Usefulness of the hydrogen sulfide test for assessment of water quality in Bangladesh. J. Appl. Microbiol. 2008, 104, 388–395. [Google Scholar] [CrossRef]
- Bernardes, C.; Bernardes, R.; Zimmer, C.; Dorea, C.C. A simple off-grid incubator for microbiological water quality analysis. Water 2020, 12, 240. [Google Scholar] [CrossRef]
- IDEXX. Colilert*. Available online: https://www.idexx.com/en/water/water-products-services/colilert/ (accessed on 20 June 2021).
- Environmental Bio-Detection Products Inc. ColiPlate™ and WaterCheck™ Handout. Available online: http://www.biotoxicity.org/images/Toxicity%20PDF/ColiPlate%20and%20WaterCheck%20Handout.pdf (accessed on 20 June 2021).
- Bishop, I.J.; Warner, S.; van Noordwijk, T.C.G.E.; Nyoni, F.C.; Loiselle, S. Citizen science monitoring for sustainable development goal indicator 6.3.2 in England and Zambia. Sustainability 2020, 12, 10271. [Google Scholar] [CrossRef]
- Lifshitz, R.; Joshi, R. Comparison of a novel ColiPlate™ kit and the standard membrane filter technique for enumerating total coliforms and Escherichia coli bacteria in Water. Environ. Toxicol. Water Qual. 1998, 13, 157–164. [Google Scholar] [CrossRef]
- Bluewater Biosciences. Available online: http://bluewaterbiosciences.com (accessed on 14 May 2021).
- Reed, B.C.; Rasnake, M.S. An assessment of coliform bacteria in water sources near Appalachian Trail shelters within the Great Smoky Mountains National Park. Wilderness Environ. Med. 2016, 27, 107–110. [Google Scholar] [CrossRef]
- Milne, J.; Gilpin, S.; Fortuna, J. A review of Hamilton Harbour beaches: Towards delisting 2020, successes and challenges. Aquat. Ecosyst. Health Manag. 2017, 20, 278–284. [Google Scholar] [CrossRef]
- Bain, R.; Bartram, J.; Elliott, M.; Matthews, R.; McMahan, L.; Tung, R.; Chuang, P.; Gundry, S. A summary catalogue of microbial drinking water tests for low and medium resource settings. Inter. J. Environ. Res. Public Health 2012, 9, 1609–1625. [Google Scholar] [CrossRef] [PubMed]
- Bluewater Biosciences. MPN Table. Available online: https://bluewaterbiosciences.com/index.php?main_page=page&id=13 (accessed on 20 June 2021).
- IDEXX. IDEXX Quanti-Tray®/2000 MPN Table (per 100 mL). Available online: https://www.idexx.com/files/qt97mpntable.pdf (accessed on 20 June 2021).
- U.S. EPA. Method 1603: Escherichia coli (E. coli) in Water by Membrane Filtration Using Modified Membrane-Thermotolerant Escherichia Coli agar (Modified mTEC); Office of Water EPA-821-R-14-010; United States Environmental Protection Agency: Washington, DC, USA, 2014.
- WHO. Guidelines for Drinking-Water Quality: Fourth Edition Incorporating the First Addendum; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- U.S. EPA. Evaluation of Multiple Indicator Combinations to Develop Relationships; Office of Water EPA-822-R-10-004; United States Environmental Protection Agency: Washington, DC, USA, 2010. Available online: https://www.epa.gov/sites/production/files/2015-11/documents/evaluation-multiple-indicator-relationships.pdf (accessed on 20 June 2021).
- APHA/AWWA/WEF. Standard Methods for the Examination of Water and Wastewater, 22nd ed.; American Public Health Association/American Water Works Association, Water Environment Federation: Washington, DC, USA, 2012. [Google Scholar]
- Hosmer, D.W., Jr.; Lemeshow, S.; Sturdivant, R.X. Applied Logistic Regression, 3rd ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2013. [Google Scholar]
- Kinzelman, J.L.; Singh, A.; Ng, C.; Pond, K.R.; Bagley, R.C.; Gradus, S. Use of IDEXX Colilert-18® and Quanti-Tray/2000 as a rapid and simple enumeration method for the implementation of recreational water monitoring and notification programs. Lake Reserv. Manag. 2005, 21, 73–77. [Google Scholar] [CrossRef]
- Buckalew, D.W.; Hartman, L.J.; Grimsley, G.A.; Martin, A.E.; Register, K.M. A long-term study comparing membrane filtration with Colilert® defined substrates in detecting fecal coliforms and Escherichia coli in natural waters. J. Environ. Manag. 2006, 80, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, W.P.; Kim, M.; Thackston, E.L. Comparison of commercially available Escherichia coli enumeration tests: Implications for attaining water quality standards. Water Res. 2005, 39, 4869–4878. [Google Scholar] [CrossRef] [PubMed]
- Bain, R.E.; Woodall, C.; Elliott, J.; Arnold, B.F.; Tung, R.; Morley, R.; du Preez, M.; Bartram, J.K.; Davis, A.P.; Gundry, S.W.; et al. Evaluation of an inexpensive growth medium for direct detection of Escherichia coli in temperate and sub-tropical waters. PLoS ONE 2015, 10, e0140997. [Google Scholar] [CrossRef]
- Genter, F.; Marks, S.J.; Clair-Caliot, G.; Mugume, D.S.; Johnston, R.B.; Bain, R.E.; Julian, T.R. Evaluation of the novel substrate RUG™ for the detection of Escherichia coli in water from temperate (Zurich, Switzerland) and tropical (Bushenyi, Uganda) field sites. Environ. Sci. Water Res. Technol. 2019, 5, 1082–1091. [Google Scholar] [CrossRef]
- Ramoutar, S. The use of Colilert-18, Colilert and Enterolert for the detection of faecal coliform, Escherichia coli and Enterococci in tropical marine waters, Trinidad and Tobago. Region. Stud. Mar. Sci. 2020, 40, 101490. [Google Scholar] [CrossRef]
- Pisciotta, J.M.; Rath, D.F.; Stanek, P.A.; Flanery, D.M.; Harwood, V.J. Marine bacteria cause false-positive results in the Colilert-18 rapid identification test for Escherichia coli in Florida waters. Appl. Environ. Microbiol. 2002, 68, 539–544. [Google Scholar] [CrossRef]
- Chao, W.L. Evaluation of Colilert-18 for the detection of coliforms and Escherichia coli in tropical fresh water. Lett. Appl. Microbiol. 2006, 42, 115–120. [Google Scholar] [CrossRef]
- Shadix, L.C.; Rice, E.W. Evaluation of β-glucuronidase assay for the detection of Escherichia coli from environmental waters. Can. J. Microbiol. 1991, 37, 908–911. [Google Scholar] [CrossRef]
- Bissonnette, G.K.; Jezeski, J.J.; McFeters, G.A.; Stuart, D. Influence of environmental stress on enumeration of indicator bacteria from natural waters. Appl. Microbiol. 1975, 29, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, N.J.; Zeigler, W.L.; Reed, F.C.; Stukel, T.A.; Rice, E.W. Comparison of membrane filter, multiple-fermentation-tube, and presence-absence techniques for detecting total coliforms in small community water systems. Appl. Environ. Microbiol. 1986, 51, 1007–1012. [Google Scholar] [CrossRef] [PubMed]
- Davies, C.M.; Apte, S.C.; Peterson, S.M.; Stauber, J.L. Plant and algal interference in bacterial beta-D-galactosidase and beta-D-glucuronidase assays. Appl. Environ. Microbiol. 1994, 60, 3959–3964. [Google Scholar] [CrossRef] [PubMed]
- Chao, K.K.; Chao, C.C.; Chao, W.L. Evaluation of Colilert-18 for detection of coliforms and Eschericha coli in subtropical freshwater. Appl. Environ. Microbiol. 2004, 70, 1242–1244. [Google Scholar] [CrossRef]
| Method | Units | n | Mean | SE | Median | Range |
|---|---|---|---|---|---|---|
| ColiPlate | log10 MPN | 40 | 1.98 | 0.13 | 2.18 | 0.0–3.23 |
| Colilert | log10 MPN | 40 | 1.78 | 0.11 | 1.75 | 0.0–3.01 |
| MF + mTEC 1 | log10 CFU | 40 | 1.88 | 0.11 | 2.01 | 0.2–3.45 |
| Method/covariate | Units | β 1 | SE β 2 | OR 3 | OR, 95% CI 4 | p |
|---|---|---|---|---|---|---|
| ColiPlate | log10 MPN | 3.54 | 1.14 | 34.51 | 3.72–320.49 | 0.002 |
| Constant | −7.18 | |||||
| Colilert | log10 MPN | 3.79 | 1.20 | 44.47 | 4.24–465.50 | 0.002 |
| Constant | −6.75 |
| Method | Probability Cutoff | Sensitivity (%) | Specificity (%) | Positive Predictive Value (%) | Negative Predictive Value (%) | Correctly Classified (%) |
|---|---|---|---|---|---|---|
| ColiPlate | 0.40 | 95.00 | 78.95 | 83.33 | 93.75 | 87.50 |
| Colilert | 0.40 | 90.48 | 73.68 | 79.17 | 87.50 | 82.50 |
| ColiPlate | 0.45 | 95.24 | 78.95 | 83.33 | 93.75 | 87.50 |
| Colilert | 0.45 | 85.71 | 84.21 | 85.71 | 84.21 | 85.00 |
| ColiPlate | 0.50 | 90.48 | 84.21 | 86.36 | 88.89 | 87.50 |
| Colilert | 0.50 | 80.95 | 89.47 | 89.47 | 80.95 | 85.00 |
| ColiPlate | 0.55 | 85.71 | 84.21 | 85.71 | 84.21 | 85.00 |
| Colilert | 0.55 | 76.19 | 89.47 | 88.89 | 77.27 | 82.50 |
| ColiPlate | 0.60 | 85.71 | 84.21 | 85.71 | 84.21 | 85.00 |
| Colilert | 0.60 | 76.19 | 89.47 | 88.89 | 77.27 | 82.80 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gibson, C.J.; Maritim, A.K.; Marion, J.W. Comparison of the ColiPlate™ Kit with Two Common E. coli Enumeration Methods for Water. Water 2021, 13, 1804. https://doi.org/10.3390/w13131804
Gibson CJ, Maritim AK, Marion JW. Comparison of the ColiPlate™ Kit with Two Common E. coli Enumeration Methods for Water. Water. 2021; 13(13):1804. https://doi.org/10.3390/w13131804
Chicago/Turabian StyleGibson, Cassi J., Abraham K. Maritim, and Jason W. Marion. 2021. "Comparison of the ColiPlate™ Kit with Two Common E. coli Enumeration Methods for Water" Water 13, no. 13: 1804. https://doi.org/10.3390/w13131804
APA StyleGibson, C. J., Maritim, A. K., & Marion, J. W. (2021). Comparison of the ColiPlate™ Kit with Two Common E. coli Enumeration Methods for Water. Water, 13(13), 1804. https://doi.org/10.3390/w13131804






