The Effect of Chironomid Larvae on Nitrogen Cycling and Microbial Communities in Soft Sediments
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Setup
2.2. Benthic Flux Measurement
2.3. Oxygen Consumption by Individual Chironomid Larvae
2.4. Nucleic Acid Extraction
2.5. Synthesis of cDNA
2.6. Amplification and Sequencing of 16S rRNA Gene and Bioinformatics
2.7. Quantitative PCR Analyses
2.8. Statistical Analysis
3. Results
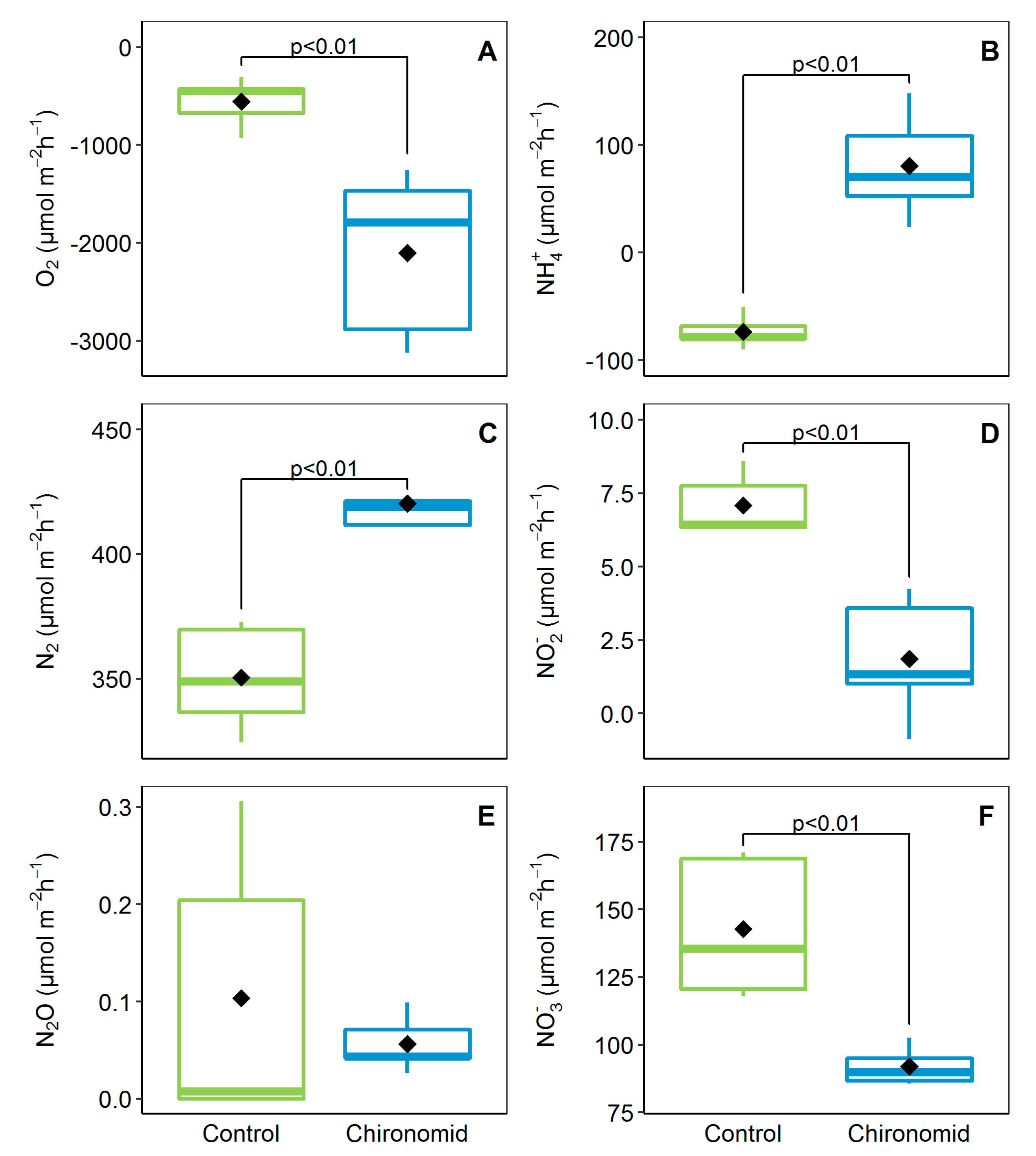
3.1. Benthic Fluxes at the Sediment–Water Interface and Animal O2 Consumption
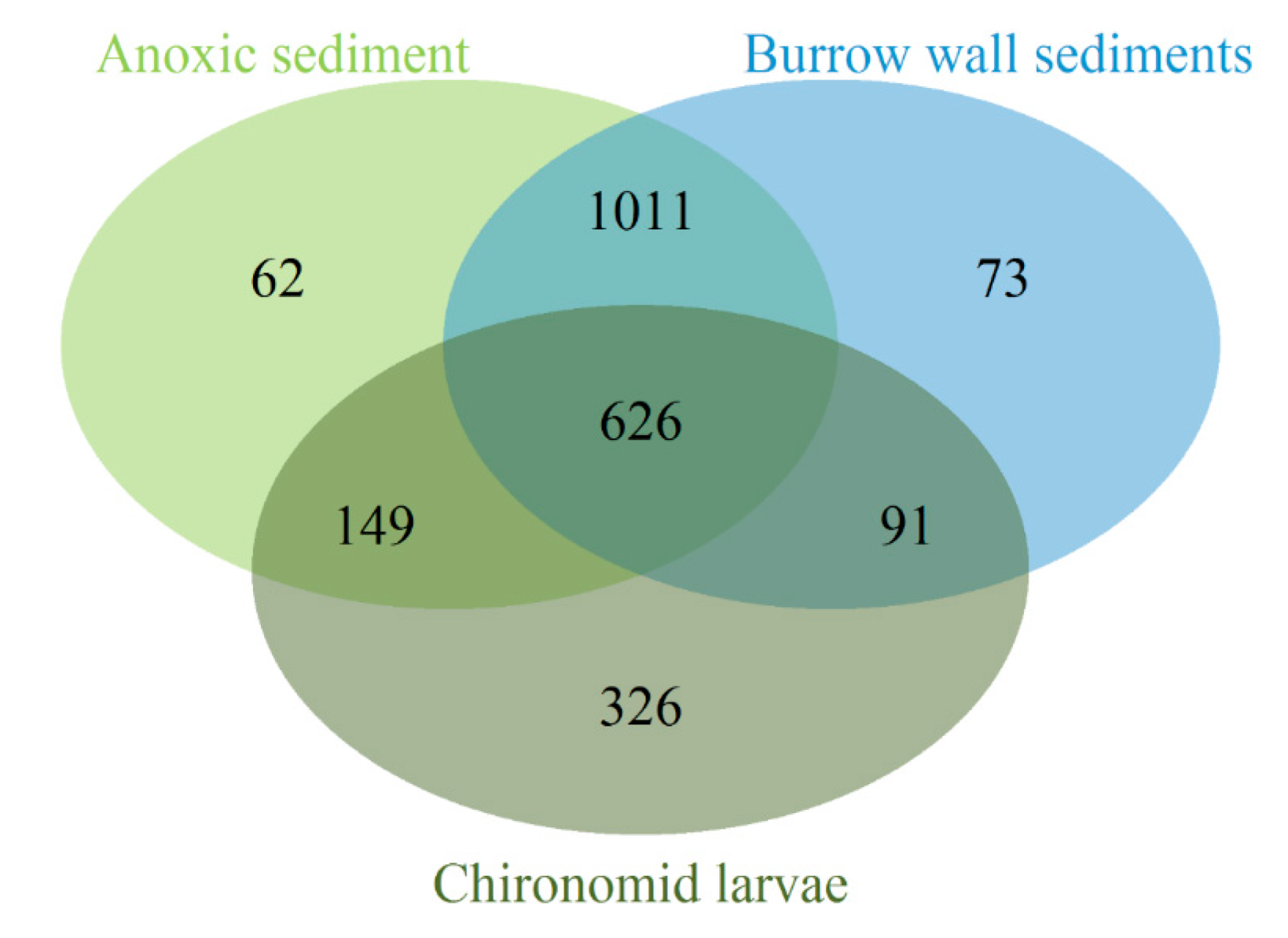
3.2. Bacterial Community Composition
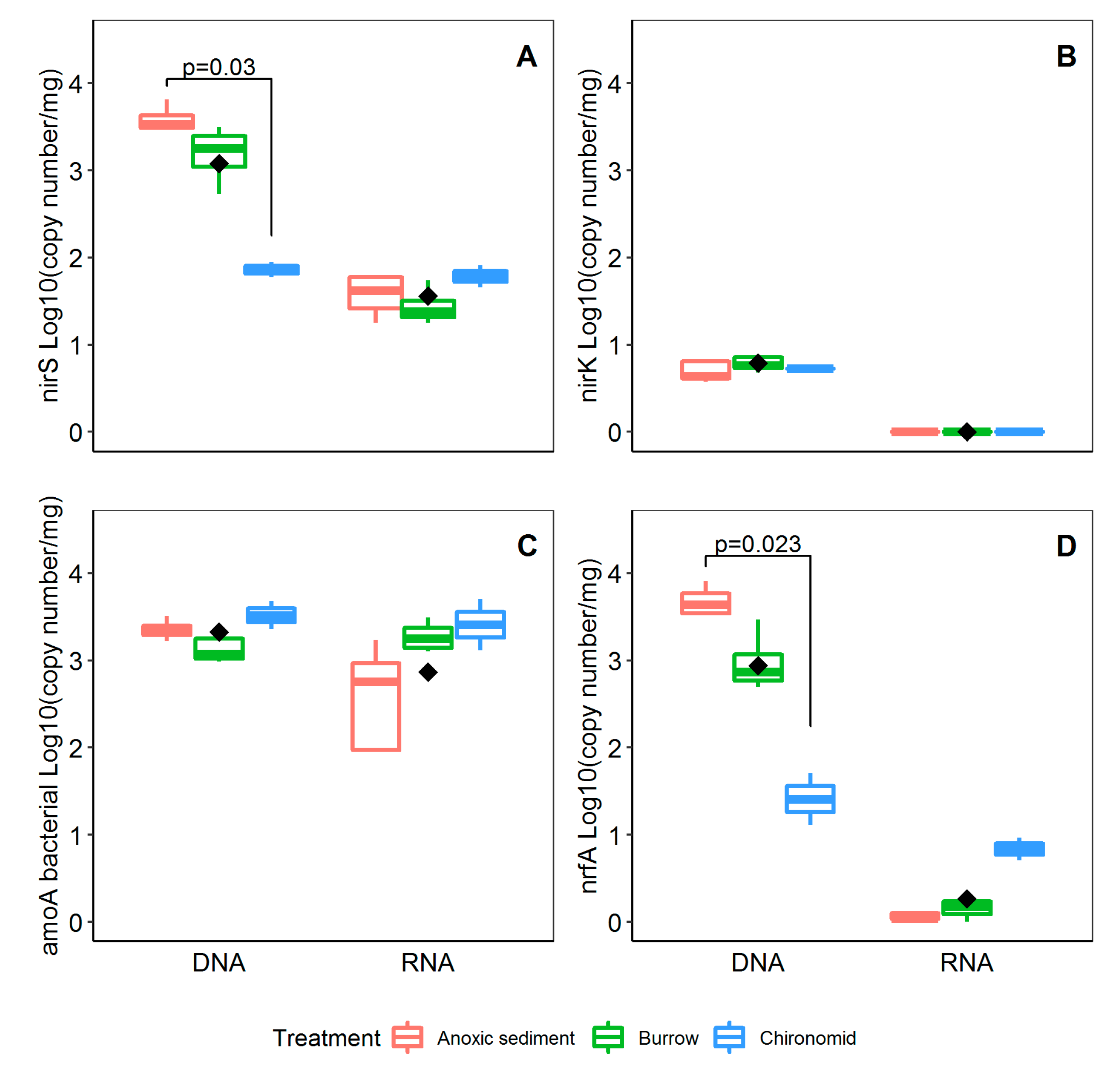
3.3. Abunadance and Activity of Nitrifying and Denitrifying Genes
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Asmala, E.; Carstensen, J.; Conley, D.J.; Slomp, C.P.; Stadmark, J.; Voss, M. Efficiency of the coastal filter: Nitrogen and phosphorus removal in the Baltic Sea. Limnol. Oceanogr. 2017, 62, S222–S238. [Google Scholar] [CrossRef]
- Zilius, M.; Vybernaite-Lubiene, I.; Vaiciute, D.; Petkuviene, J.; Zemlys, P.; Liskow, I.; Voss, M.; Bartoli, M.; Bukaveckas, P.A. The influence of cyanobacteria blooms on the attenuation of nitrogen throughputs in a Baltic coastal lagoon. Biogeochemistry 2018, 141, 143–165. [Google Scholar] [CrossRef]
- Bianchi, T.S. Biogeochemistry of Estuaries; Oxford University Press: New York, NY, USA, 2007. [Google Scholar]
- Brinson, M.M.; Christian, R.R.; Blum, L.K. Multiple states in the sea-level induced transition from terrestrial forest to estuary. Estuaries 1995, 18, 648–659. [Google Scholar] [CrossRef]
- Kristensen, E. Organic matter diagenesis at the oxic/anoxic interface in coastal marine sediments, with emphasis on the role of burrowing animals. Hydrobiologia 2000, 426, 1–24. [Google Scholar] [CrossRef]
- Mermillod-Blondin, F.; Rosenberg, R.; François-Carcaillet, F.; Norling, K.; Mauclaire, L. Influence of bioturbation by three benthic infaunal species on microbial communities and biogeochemical processes in marine sediment. Aquat. Microb. Ecol. 2004, 36, 271–284. [Google Scholar] [CrossRef]
- Magri, M.; Benelli, S.; Bondavalli, C.; Bartoli, M.; Christian, R.R.; Bodini, A. Benthic N pathways in illuminated and bioturbated sediments studied with network analysis. Limnol. Oceanogr. 2018, 63, S68–S84. [Google Scholar] [CrossRef]
- An, S.; Gardner, W.S. Dissimilatory nitrate reduction to ammonium (DNRA) as a nitrogen link, versus denitrification as a sink in a shallow estuary (Laguna Madre/Baffin Bay, Texas). Mar. Ecol. Prog. Ser. 2002, 237, 41–50. [Google Scholar] [CrossRef]
- Bonaglia, S.; Deutsch, B.; Bartoli, M.; Marchant, H.K.; Brüchert, V. Seasonal oxygen, nitrogen and phosphorus benthic cycling along an impacted Baltic Sea estuary: Regulation and spatial patterns. Biogeochemistry 2014, 119, 139–160. [Google Scholar] [CrossRef]
- Vybernaite-Lubiene, I.; Zilius, M.; Saltyte-Vaisiauske, L.; Bartoli, M. Recent trends (2012–2016) of N, Si, and P export from the Nemunas River Watershed: Loads, unbalanced stoichiometry, and threats for downstream aquatic ecosystems. Water 2018, 10, 1178. [Google Scholar] [CrossRef]
- Carstensen, J.; Conley, D.J.; Bonsdorff, E.; Gustafsson, B.G.; Hietanen, S.; Janas, U.; Jilbert, T.; Maximov, A.; Norkko, A.; Norkko, J.; et al. Hypoxia in the Baltic Sea: Biogeochemical cycles, benthic fauna, and management. Ambio 2014, 43, 26–36. [Google Scholar] [CrossRef]
- Benelli, S.; Bartoli, M.; Zilius, M.; Vybernaite–Lubiene, I.; Ruginis, T.; Petkuviene, J.; Fano, E.A. Microphytobenthos and chironomid larvae attenuate nutrient recycling in shallow-water sediments. Freshw. Biol. 2018, 63, 187–201. [Google Scholar] [CrossRef]
- Kristensen, E.; Penha-Lopes, G.; Delefosse, M.; Valdemarsen, T.; Quintana, C.O.; Banta, G.T. What is bioturbation? The need for a precise definition for fauna in aquatic sciences. Mar. Ecol. Prog. Ser. 2012, 446, 285–302. [Google Scholar] [CrossRef]
- Svensson, J.E.; Leonardson, L.G. Effects of bioturbation by tube-dwelling chironomid larvae on oxygen uptake and denitrification in eutrophic lake sediments. Freshw. Biol. 1996, 35, 289–300. [Google Scholar] [CrossRef]
- Stief, P.; Beer, D.D. Probing the microenvironment of freshwater sediment macrofauna: Implications of deposit-feeding and bioirrigation for nitrogen cycling. Limnol. Oceanogr. 2006, 51, 2538–2548. [Google Scholar] [CrossRef]
- Bonaglia, S.; Nascimento, F.A.; Bartoli, M.; Klawonn, I.; Brüchert, V. Meiofauna increases bacterial denitrification in marine sediments. Nat. Commun. 2014, 5, 5133. [Google Scholar] [CrossRef] [PubMed]
- Laverock, B.; Tait, K.; Gilbert, J.A.; Osborn, A.M.; Widdicombe, S. Impacts of bioturbation on temporal variation in bacterial and archaeal nitrogen–cycling gene abundance in coastal sediments. Environ. Microbiol. Rep. 2014, 6, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Foshtomi, M.Y.; Braeckman, U.; Derycke, S.; Sapp, M.; Van Gansbeke, D.; Sabbe, K.; Willems, A.; Vincx, M.; Vanaverbeke, J. The link between microbial diversity and nitrogen cycling in marine sediments is modulated by macrofaunal bioturbation. PLoS ONE 2015, 10, e0130116. [Google Scholar]
- Welsh, D.T.; Nizzoli, D.; Fano, E.A.; Viaroli, P. Direct contribution of clams (Ruditapes philippinarum) to benthic fluxes, nitrification, denitrification and nitrous oxide emission in a farmed sediment. Estuar. Coast. Shelf Sci. 2015, 154, 84–93. [Google Scholar] [CrossRef]
- Foshtomi, M.Y.; Leliaert, F.; Derycke, S.; Willems, A.; Vincx, M.; Vanaverbeke, J. The effect of bio-irrigation by the polychaete Lanice conchilega on active denitrifiers: Distribution, diversity and composition of nosZ gene. PLoS ONE 2018, 13, e0192391. [Google Scholar]
- Moraes, P.C.; Zilius, M.; Benelli, S.; Bartoli, M. Nitrification and denitrification in estuarine sediments with tube-dwelling benthic animals. Hydrobiologia 2018, 819, 217–230. [Google Scholar] [CrossRef]
- Murphy, A.E.; Kolkmeyer, R.; Song, B.; Anderson, I.C.; Bowen, J. Bioreactivity and Microbiome of Biodeposits from Filter-Feeding Bivalves. Microb. Ecol. 2019, 77, 343–357. [Google Scholar] [CrossRef]
- Stief, P. Stimulation of microbial nitrogen cycling in aquatic ecosystems by benthic macrofauna: Mechanisms and environmental implications. Biogeosciences 2013, 10, 7829–7846. [Google Scholar] [CrossRef]
- Laverock, B.; Gilbert, J.A.; Tait, K.; Osborn, A.M.; Widdicombe, S. Bioturbation: Impact on the marine nitrogen cycle. Biochem. Soc. Trans. 2011, 39, 315–320. [Google Scholar] [CrossRef]
- Gilbertson, W.W.; Solan, M.; Prosser, J.I. Differential effects of microorganism–invertebrate interactions on benthic nitrogen cycling. FEMS Microbiol. Ecol. 2012, 82, 11–22. [Google Scholar] [CrossRef]
- Bartoli, M.; Nizzoli, D.; Viaroli, P. Microphytobenthos activity and fluxes at the sediment-water interface: Interactions and spatial variability. Aquat. Ecol. 2003, 37, 341–349. [Google Scholar] [CrossRef]
- Moulton, O.M.; Altabet, M.A.; Beman, J.M.; Deegan, L.A.; Lloret, J.; Lyons, M.K.; Nelson, J.A.; Pfister, C.A. Microbial associations with macrobiota in coastal ecosystems: Patterns and implications for nitrogen cycling. Front. Ecol. Environ. 2016, 14, 200–208. [Google Scholar] [CrossRef]
- Stief, P.; Eller, G. The gut microenvironment of sediment-dwelling Chironomus plumosus larvae as characterised with O2, pH, and redox microsensors. J. Comp. Physiol. B 2006, 176, 673–683. [Google Scholar] [CrossRef]
- Poulsen, M.; Kofoed, M.V.; Larsen, L.H.; Schramm, A.; Stief, P. Chironomus plumosus larvae increase fluxes of denitrification products and diversity of nitrate-reducing bacteria in freshwater sediment. Syst. Appl. Microbiol. 2014, 37, 51–59. [Google Scholar] [CrossRef]
- Petersen, J.M.; Kemper, A.; Gruber-Vodicka, H.; Cardini, U.; van der Geest, M.; Kleiner, M.; Silvia Bulgheresi, S.; Mußmann, M.; Herbold, C.; Seah, B.K.B.; et al. Chemosynthetic symbionts of marine invertebrate animals are capable of nitrogen fixation. Nat. Microbiol. 2016, 2, 16196. [Google Scholar] [CrossRef]
- Hölker, F.; Vanni, M.J.; Kuiper, J.J.; Meile, C.; Grossart, H.P.; Stief, P.; Adrian, R.; Lorke, A.; Dellwig, O.; Brand, A.; et al. Tube-dwelling invertebrates: Tiny ecosystem engineers have large effects in lake ecosystems. Ecol. Monogr. 2015, 85, 333–351. [Google Scholar] [CrossRef]
- Stocum, E.T.; Plante, C.J. The eect of artificial defaunation on bacterial assemblages of intertidal sediments. J. Exp. Mar. Biol. Ecol. 2006, 337, 147–158. [Google Scholar] [CrossRef]
- Dalsgaard, T.; Nielsen, L.P.; Brotas, V.; Viaroli, P.; Underwood, G.; Nedwell, D.; Dong, L.D.; Sundbäck, K.; Rysgaard, S.; Miles, A.; et al. Protocol Handbook for NICE-Nitrogen Cycling in Estuaries: A Project under the EU Research Programme: Marine Science and Technology (MAST III); Ministry of Environment and Energy National Environmental Research Institute: Silkeborg, Denmark, 2000; pp. 1–62. [Google Scholar]
- Kana, T.M.; Darkangelo, C.; Hunt, M.D.; Oldham, J.B.; Bennett, G.E.; Cornwell, J.C. Membrane inlet mass spectrometer for rapid high-precision determination of N2, O2, and Ar in environmental water samples. Anal. Chem. 1994, 66, 4166–4170. [Google Scholar] [CrossRef]
- Grassshoff, K. Determination of nitrate. In Methods of Seawater Analysis; Grassoff, K., Ehrhardt, M., Kremling, K., Eds.; Verlag Chemie: Weinheimm, Germany, 1982; p. 143. [Google Scholar]
- Bonaglia, S.; Brüchert, V.; Callac, N.; Vicenzi, A.; Fru, E.C.; Nascimento, F.J.A. Methane fluxes from coastal sediments are enhanced by macrofauna. Sci. Rep. 2017, 7, 13145. [Google Scholar] [CrossRef]
- Milani, C.; Hevia, A.; Foroni, E.; Duranti, S.; Turroni, F.; Lugli, G.A.; Sanchez, B.; Martín, R.; Gueimonde, M.; van Sinderen, D.; et al. Assessing the fecal microbiota: An optimized ion torrent 16S rRNA gene-based analysis protocol. PLoS ONE 2013, 8, e68739. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef]
- Data-Driven Documents. Available online: https://d3js.org (accessed on 22 July 2019).
- Chen, H.; Boutros, P.C. VennDiagram: A package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinform. 2011, 12, 35. [Google Scholar] [CrossRef]
- Pohlert, T. The Pairwise Multiple Comparison of Mean Ranks Package (PMCMR); R Package, 2014; p. 27. [Google Scholar]
- R-Project. R: A Language and Environment for Statistical Computing [Online]. R Foundation for Statistical Computing: Vienna, Austria. 2014. Available online: http://www.R-project.org (accessed on 22 July 2019).
- Welsh, D.T. It’s a dirty job but someone has to do it: The role of marine benthic macrofauna in organic matter turnover and nutrient recycling to the water column. Chem. Ecol. 2003, 19, 321–342. [Google Scholar] [CrossRef]
- Stief, P.; Poulsen, M.; Nielsen, L.P.; Brix, H.; Schramm, A. Nitrous oxide emission by aquatic macrofauna. Proc. Natl. Acad. Sci. USA 2009, 106, 4296–4300. [Google Scholar] [CrossRef]
- Stief, P.; Schramm, A. Regulation of nitrous oxide emission associated with benthic invertebrates. Freshw. Biol. 2010, 55, 1647–1657. [Google Scholar] [CrossRef]
- Sun, X.; Hu, Z.; Jia, W.; Duan, C.; Yang, L. Decaying cyanobacteria decrease N2O emissions related to diversity of intestinal denitrifiers of Chironomus plumosus. J. Limnol. 2015, 74. [Google Scholar] [CrossRef][Green Version]
- Cabello, P.; Roldán, M.D.; Castillo, F.; Moreno-Vivián, C. Nitrogen cycle. In Encyclopaedia of Microbiology, 3rd ed.; Schaechter, M., Ed.; Academic Press: London, UK, 2009. [Google Scholar]
- Baggs, E.; Phillipot, L. Nitrous oxide production in the terrestrial environment. In Nitrogen Cycling in Bacteria: Molecular Analysis; Moir, J.W.B., Ed.; Caister Academic Press: Norfolk, UK, 2011. [Google Scholar]
- Rasigraf, O.; Schmitt, J.; Jetten, M.S.M.; Lüke, C. Metagenomic potential for and diversity of N-cycle driving microorganisms in the Bothnian Sea sediment. Microbiol. Open. 2017, 6, e475. [Google Scholar] [CrossRef]
- Svensson, J.M. Influence of Chironomus plumosus larvae on ammonium flux and denitrification (measured by the acetylene blockage- and the isotope pairing-technique) in eutrophic lake sediment. Hydrobiologia 1997, 346, 157–168. [Google Scholar] [CrossRef]
- Pelegrí, S.P.; Blackburn, T.H. Nitrogen cycling in lake sediments bioturbated by Chironomus plumosus larvae, under different degrees of oxygenation. Hydrobiologia 1996, 325, 231–238. [Google Scholar] [CrossRef]
- Xing, X.; Liu, L.; Yan, W.; Wu, T.; Zhao, L.; Wang, X. Bioturbation effects of Chironomid larvae on nitrogen release and ammonia-oxidizing bacteria abundance in sediments. Water 2018, 10, 512. [Google Scholar] [CrossRef]
- Nogaro, G.; Mermillod-Blondin, F.; Montuelle, B.; Boisson, J.C.; Gibert, J. Chironomid larvae stimulate biogeochemical and microbial processes in a riverbed covered with fine sediment. Aquat. Sci. 2008, 70, 156–168. [Google Scholar] [CrossRef]
- Song, B.; Lisa, J.A.; Tobias, C.R. Linking DNRA community structure and activity in a shallow lagoonal estuarine system. Front. Microbiol. 2014, 5, 460. [Google Scholar] [CrossRef]
- Herbert, R.A. Nitrogen cycling in coastal marine ecosystems. FEMS Microbiol. Rev. 1999, 23, 563–590. [Google Scholar] [CrossRef]
- Burgin, A.J.; Hamilton, S.K. Have we overemphasized the role of denitrification in aquatic ecosystems? A review of nitrate removal pathways. Front. Ecol. Environ. 2007, 5, 89–96. [Google Scholar] [CrossRef]



| Gene | Primer | Primer Sequence | Ann. Temp. | Reference Strain |
|---|---|---|---|---|
| nirS | F3nir | SCCGCACCCGGGBCGYGG | 60 °C | Pseudomonas stutzeri (DSM 4166) |
| R4bcd | CGTTGAAYTTRCCGGTSGG | |||
| nirK | F1aCu | ATCATGGTSCTGCCGCG | 60 °C | Achromobcter sp.
(DSM 30128) |
| R3Cu | GCCTCGATCAGRTTGTGGTT | |||
| AOA-amoA | AOA-amoA-f | CTGAYTGGGCYTGGACATC | 54–60 °C | Nitrosopumilus maritimus (NCIMB 15022) |
| AOA-amoA-r | TTCTTCTTTGTTGCCCAGTA | |||
| AOB-amoA | amoA-1F | GGGGHTTYTACTGGTGGT | 63 °C | Nitrosomonas europaea (DSM 28437) |
| amoA-2R | CCCCTCKGSAAAGCCTTCTTC | |||
| nrfA | nrfA-F2aw | CARTGYCAYGTBGARTA | 60 °C | Citrobacter freundii (DSM 30039) |
| nrfA-R1 | TWNGGCATRTGRCARTC |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samuiloviene, A.; Bartoli, M.; Bonaglia, S.; Cardini, U.; Vybernaite-Lubiene, I.; Marzocchi, U.; Petkuviene, J.; Politi, T.; Zaiko, A.; Zilius, M. The Effect of Chironomid Larvae on Nitrogen Cycling and Microbial Communities in Soft Sediments. Water 2019, 11, 1931. https://doi.org/10.3390/w11091931
Samuiloviene A, Bartoli M, Bonaglia S, Cardini U, Vybernaite-Lubiene I, Marzocchi U, Petkuviene J, Politi T, Zaiko A, Zilius M. The Effect of Chironomid Larvae on Nitrogen Cycling and Microbial Communities in Soft Sediments. Water. 2019; 11(9):1931. https://doi.org/10.3390/w11091931
Chicago/Turabian StyleSamuiloviene, Aurelija, Marco Bartoli, Stefano Bonaglia, Ulisse Cardini, Irma Vybernaite-Lubiene, Ugo Marzocchi, Jolita Petkuviene, Tobia Politi, Anastasija Zaiko, and Mindaugas Zilius. 2019. "The Effect of Chironomid Larvae on Nitrogen Cycling and Microbial Communities in Soft Sediments" Water 11, no. 9: 1931. https://doi.org/10.3390/w11091931
APA StyleSamuiloviene, A., Bartoli, M., Bonaglia, S., Cardini, U., Vybernaite-Lubiene, I., Marzocchi, U., Petkuviene, J., Politi, T., Zaiko, A., & Zilius, M. (2019). The Effect of Chironomid Larvae on Nitrogen Cycling and Microbial Communities in Soft Sediments. Water, 11(9), 1931. https://doi.org/10.3390/w11091931







